Retinal Image Analysis for Diabetes-Based Eye Disease Detection Using Deep Learning
Abstract
1. Introduction
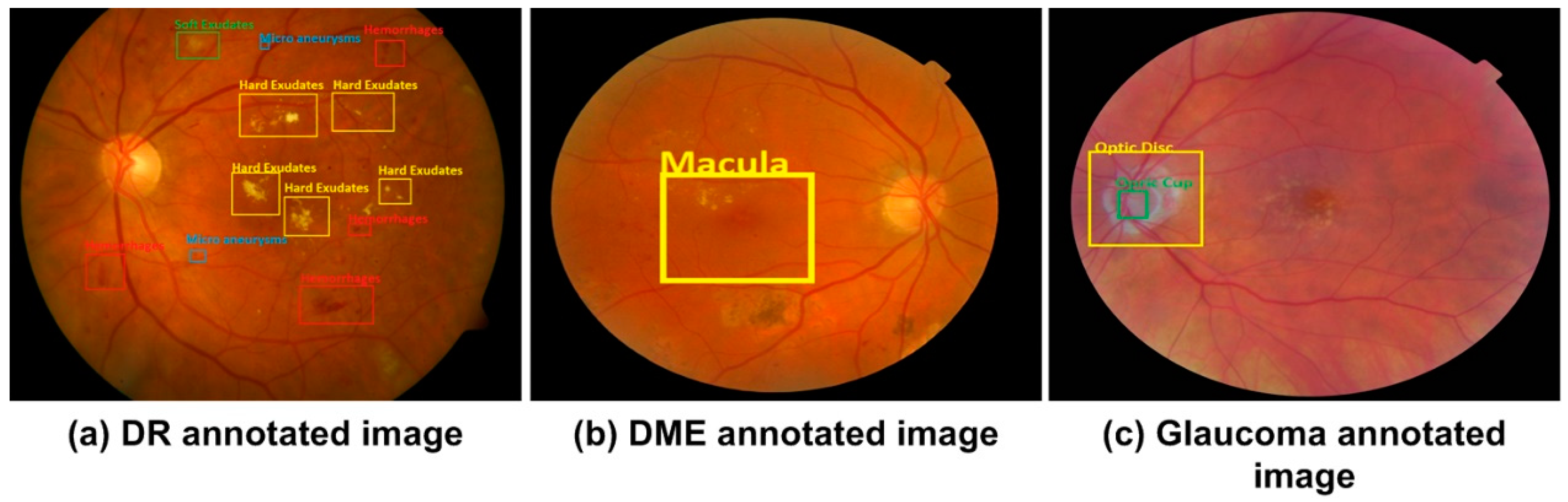
- Hard exudates are bright yellow-colored spots with a waxy appearance on the retina, which are formed because of the leakage of blood from vessels.
- Soft exudates are white lesions on the retina that occur due to occlusion of the arteriole.
- Hemorrhages develop due to blood leakage from damaged vessels and appear as dark red spots.
- Microaneurysms developed due to distortions in the boundary of blood vessels and appear as small red dots on the retina.
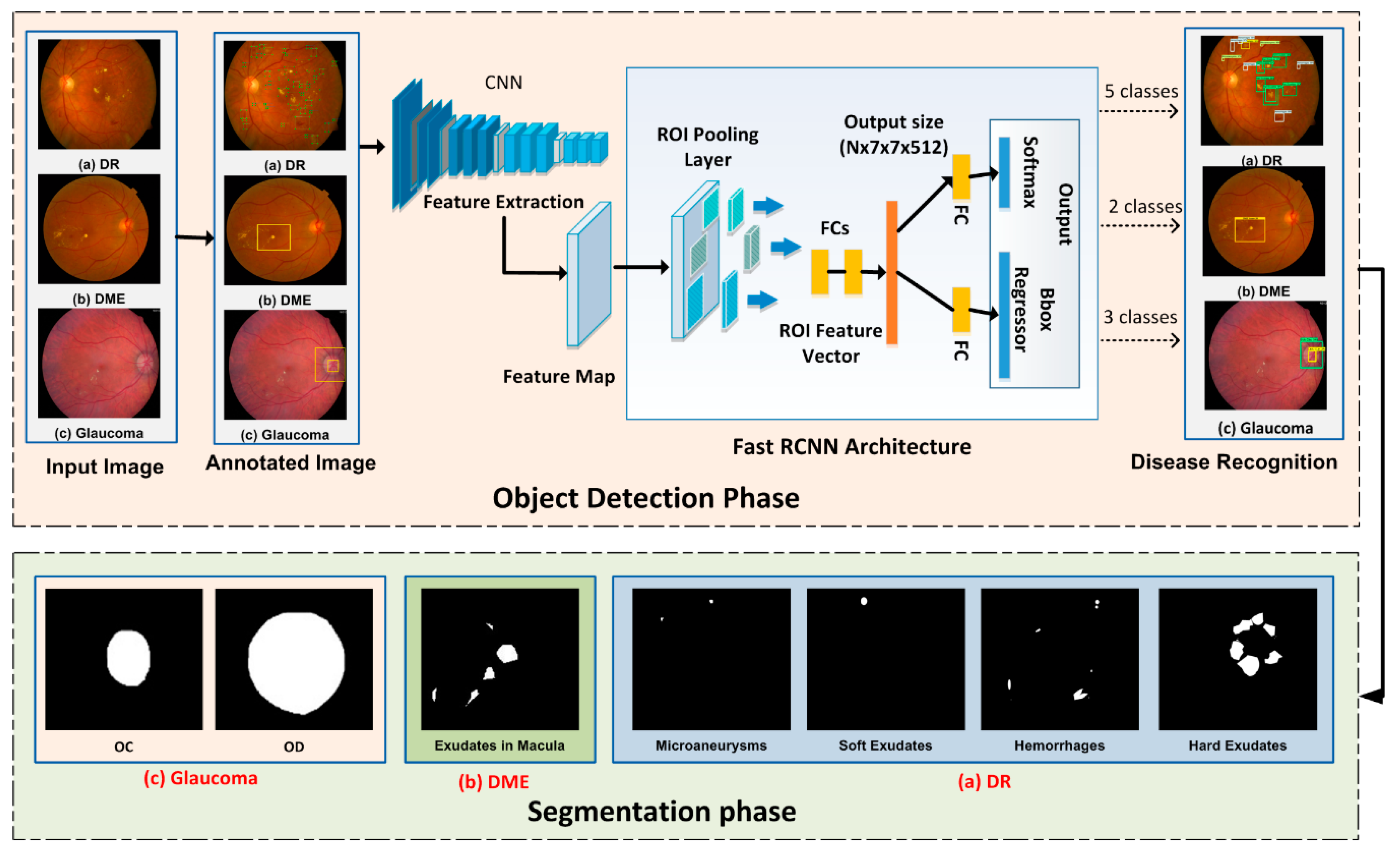
- Early and automated detection of diabetes-based eye diseases regions using machine learning-based segmentation is a complex task. In the presented methodology, we used the FRCNN-based method for localization of disease regions. Our findings conclude that the combination of FRCNN with FKM clustering results in accurate localization of the affected areas, which ensure the precise recognition of the disease in an automated manner.
- To accomplish the human-level performance over the challenging dataset i.e., ORIGA and MESSIDOR, the retinal images are represented by the FRCNN deep features, that are then segmented through the FKM clustering.
- The proposed method can detect the signs of disease including early signs simultaneously and has no issue in learning to detect an image of a healthy eye.
- The available datasets do not have bbox ground truths, so first, we developed the bbox annotations from given ground truths of the dataset which are necessary for the training of FRCNN.
2. Proposed Methodology
2.1. Ground Truth Generation
2.2. Localization Phase
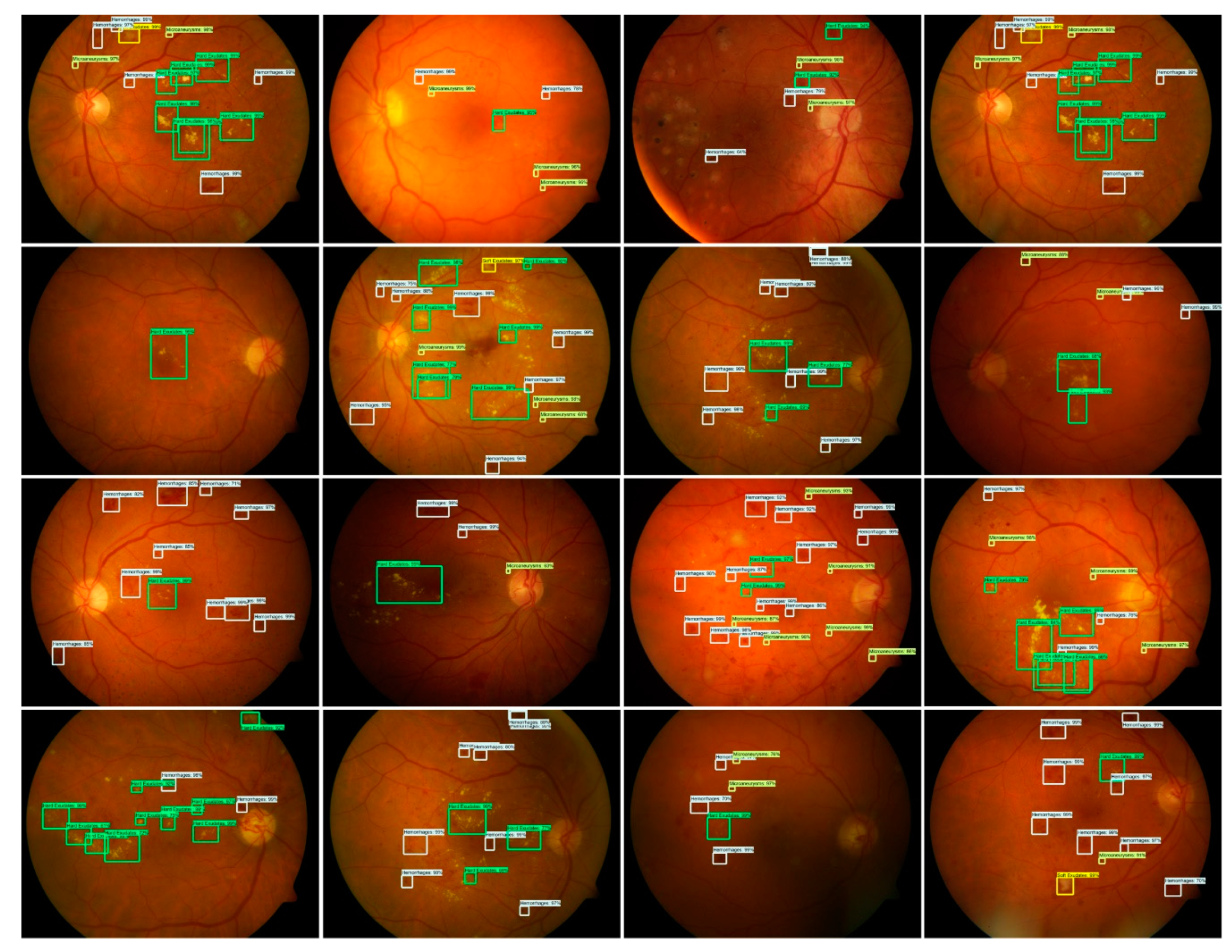
2.2.1. Localization of DR Regions
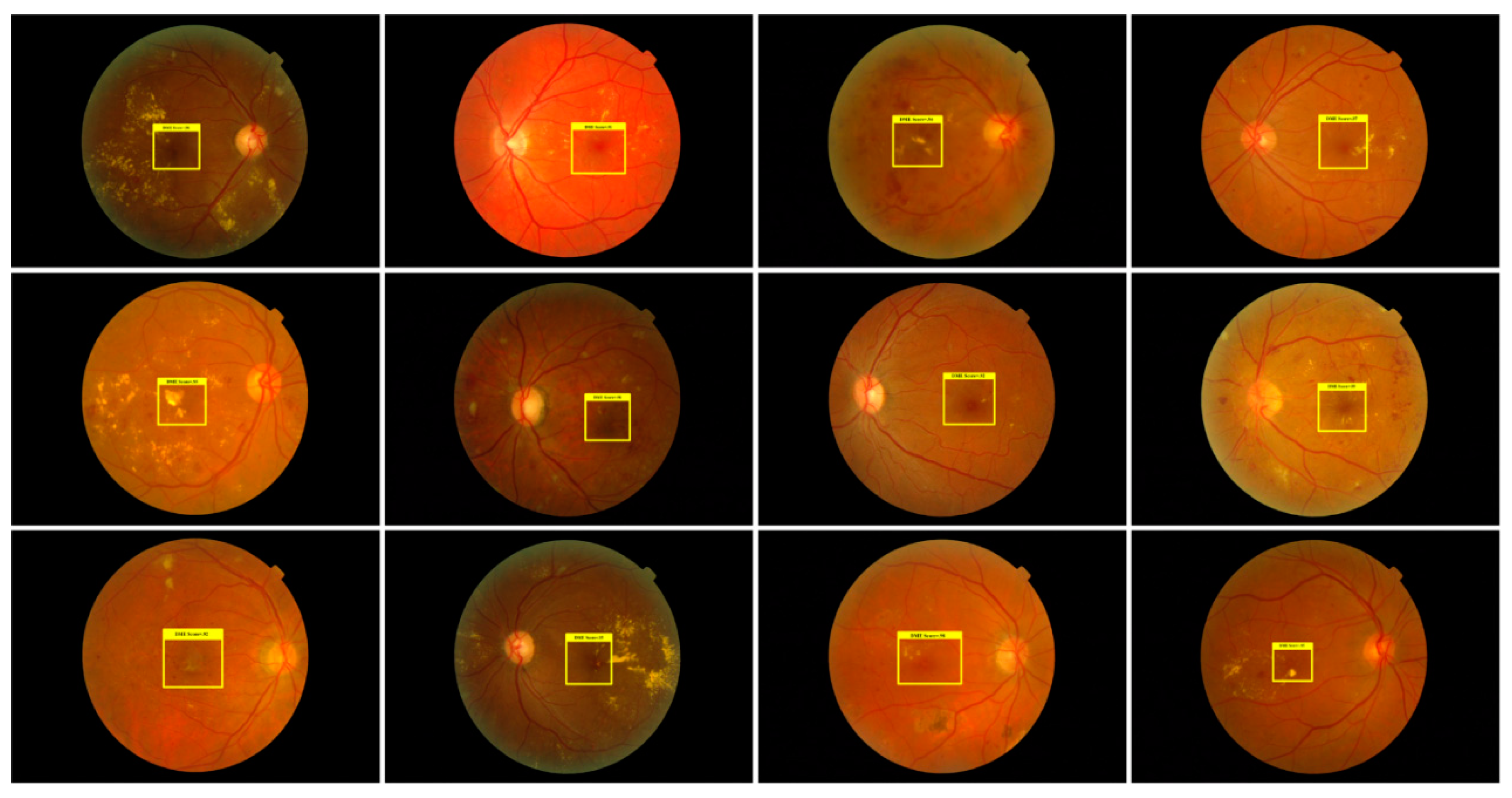
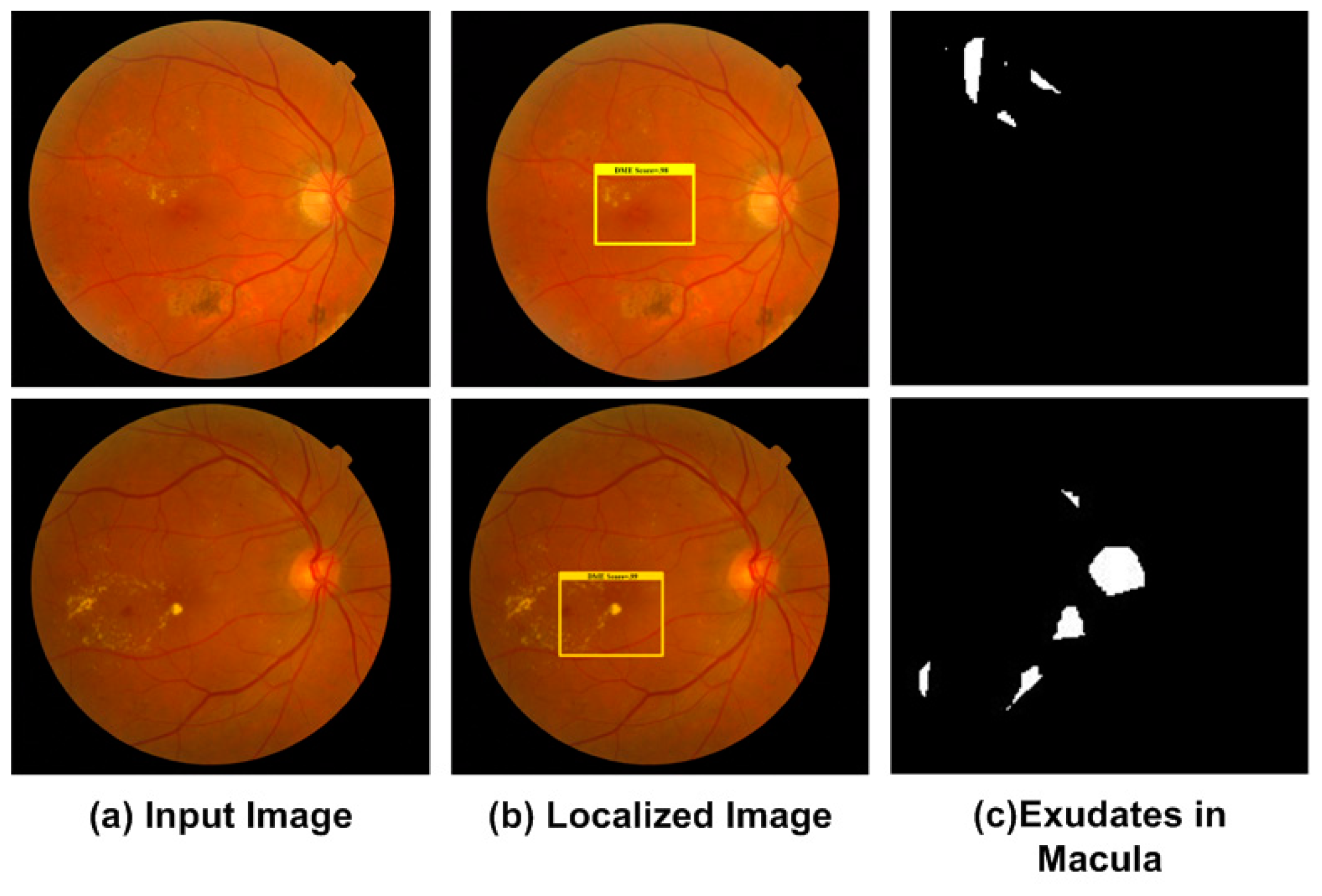
2.2.2. Localization of DME Region
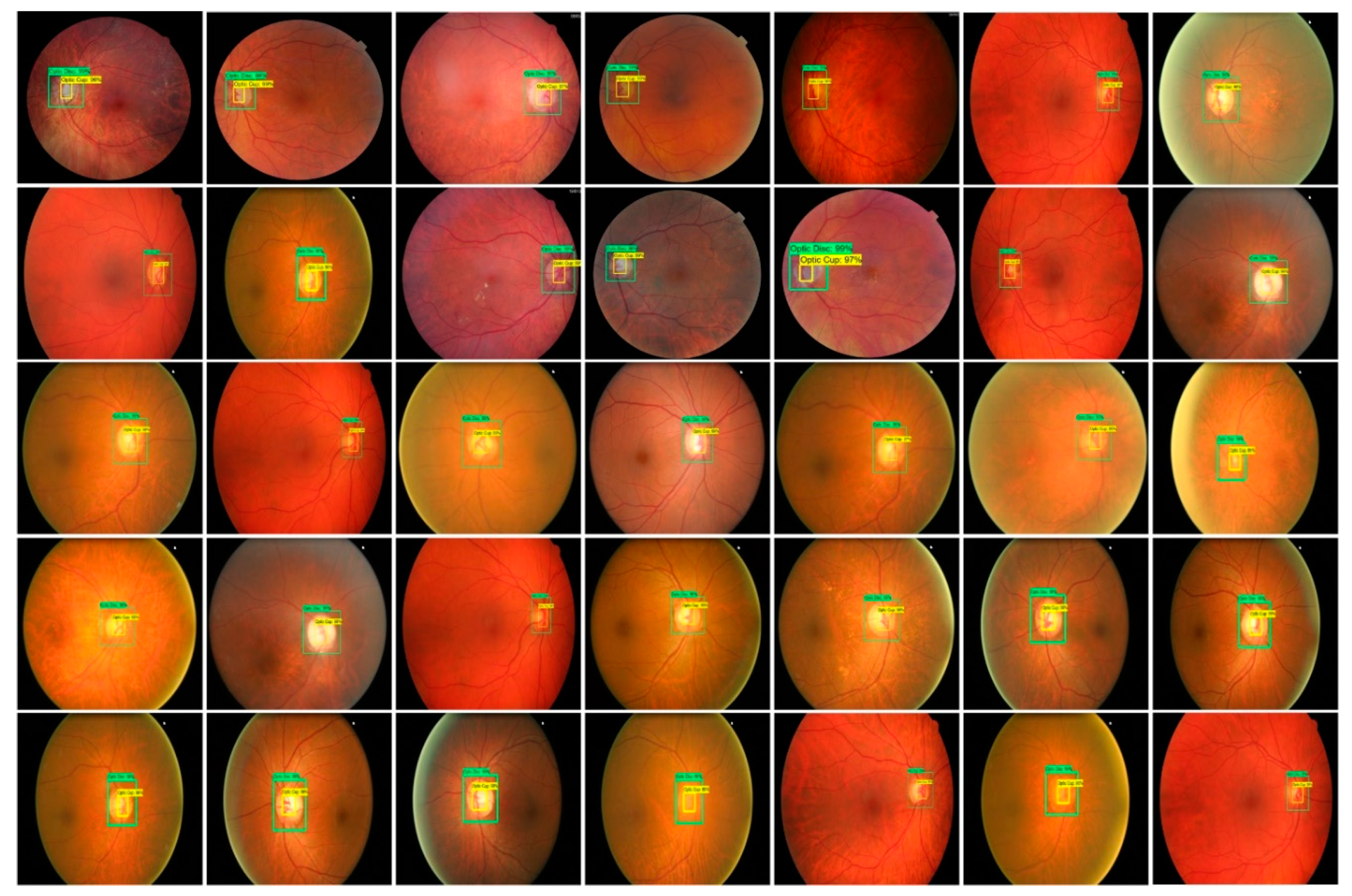
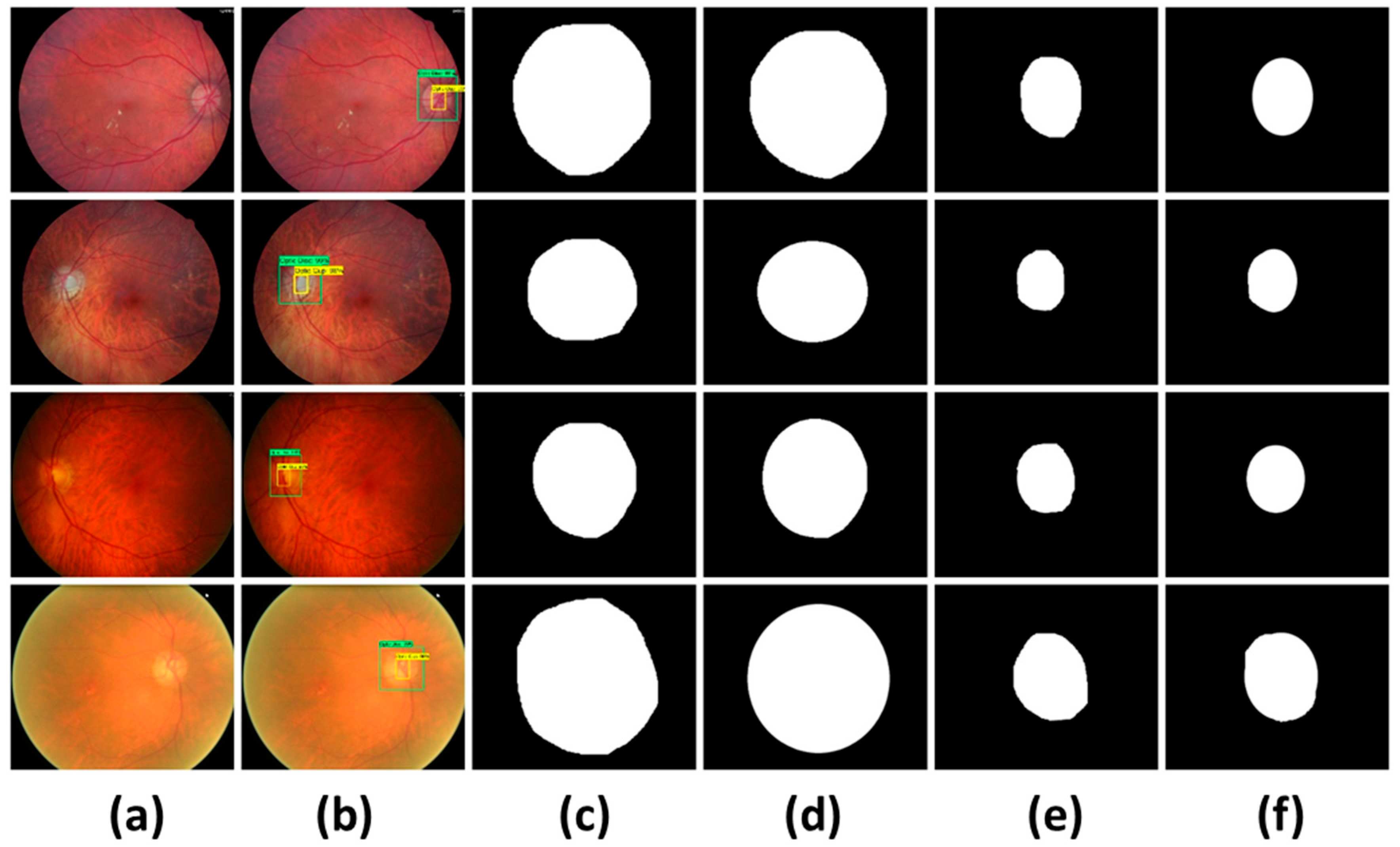
2.2.3. Localization of Glaucoma Regions
Feature Extraction
Multi-Task Loss
SGD Hypermeters
Testing through FRCNN
2.3. Segmentation of Regions through Fuzzy K-Means Clustering
- (1)
- Specify the number of clusters
- (2)
- Set Cj(0) to the initial clusters.
- (3)
- Compute the membership of all datapoints for each using following equation:where m is the fuzzification coefficient.
- (4)
- Update cluster centers using following equation:
- (5)
- Repeat from step 3, till the FKM is converged (the centroids updated between two passes is not greater than ε, the defined sensitivity threshold).
3. Experimental Results
3.1. Datasets
3.2. Evaluation Metrics
3.3. Results
3.3.1. Evaluation of FRCNN
3.3.2. Localization of DR Regions
3.3.3. Localization of DME Regions
3.3.4. Localization of Glaucoma Regions
3.3.5. Segmentation Results
3.4. Comparative Studies
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Mateen, M.; Wen, J.; Hassan, M.; Nasrullah, N.; Sun, S.; Hayat, S. Automatic Detection of Diabetic Retinopathy: A Review on Datasets, Methods and Evaluation Metrics. IEEE Access 2020, 8, 48784–48811. [Google Scholar] [CrossRef]
- Rekhi, R.S.; Issac, A.; Dutta, M.K. Automated detection and grading of diabetic macular edema from digital colour fundus images. In Proceedings of the 2017 4th IEEE Uttar Pradesh Section International Conference on Electrical, Computer and Electronics (UPCON), Mathura, India, 26–28 October 2017; pp. 482–486. [Google Scholar]
- Al-Bander, B.; Al-Nuaimy, W.; Al-Taee, M.A.; Zheng, Y. Automated glaucoma diagnosis using deep learning approach. In Proceedings of the 2017 14th International Multi-Conference on Systems, Signals & Devices (SSD), Marrakech, Morocco, 28–31 March 2017; pp. 207–210. [Google Scholar]
- Zago, G.; Andreão, R.V.; Dorizzi, B.; Salles, E.O.T. Diabetic retinopathy detection using red lesion localization and convolutional neural networks. Comput. Biol. Med. 2020, 116, 103537. [Google Scholar] [CrossRef] [PubMed]
- Kunwar, A.; Magotra, S.; Sarathi, M.P. Detection of high-risk macular edema using texture features and classification using SVM classifier. In Proceedings of the 2015 International Conference on Advances in Computing, Communications and Informatics (ICACCI), Kochi, India, 10–13 August 2015; pp. 2285–2289. [Google Scholar]
- Quigley, H.; Broman, A.T. The number of people with glaucoma worldwide in 2010 and 2020. Br. J. Ophthalmol. 2006, 90, 262–267. [Google Scholar] [CrossRef]
- Bourne, R.R.; Flaxman, S.R.; Braithwaite, T.; Cicinelli, M.V.; Das, A.; Jonas, J.B.; Keeffe, J.; Kempen, J.H.; Leasher, J.; Limburg, H.; et al. Magnitude, temporal trends, and projections of the global prevalence of blindness and distance and near vision impairment: A systematic review and meta-analysis. Lancet Glob. Health 2017, 5, e888–e897. [Google Scholar] [CrossRef]
- Al-Bander, B.; Williams, B.M.; Al-Nuaimy, W.; Al-Taee, M.A.; Pratt, H.; Zheng, Y. Dense Fully Convolutional Segmentation of the Optic Disc and Cup in Colour Fundus for Glaucoma Diagnosis. Symmetry 2018, 10, 87. [Google Scholar] [CrossRef]
- Chen, X.; Xu, Y.; Wong, D.W.K.; Wong, T.Y.; Liu, J. Glaucoma detection based on deep convolutional neural network. In Proceedings of the 2015 37th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Milan, Italy, 25–29 August 2015; Volume 2015, pp. 715–718. [Google Scholar]
- Mary, M.C.V.S.; Rajsingh, E.B.; Naik, G.R. Retinal Fundus Image Analysis for Diagnosis of Glaucoma: A Comprehensive Survey. IEEE Access 2016, 4, 4327–4354. [Google Scholar] [CrossRef]
- Sinthanayothin, C. Image Analysis for Automatic Diagnosis of Diabetic Retinopathy; University of London: London, UK, 1999. [Google Scholar]
- Krishna, B.S.; Gnanasekaran, T. Unsupervised Automated Retinal Vessel Extraction Framework Using Enhanced Filtering and Hessian Based Method with Hysteresis Thresholding. J. Med. Imaging Health Inf. 2019, 9, 1000–1010. [Google Scholar] [CrossRef]
- Khansari, M.M.; O’Neill, W.D.; Penn, R.D.; Blair, N.P.; Shahidi, M. Detection of Subclinical Diabetic Retinopathy by Fine Structure Analysis of Retinal Images. J. Ophthalmol. 2019, 2019, 5171965. [Google Scholar] [CrossRef]
- Qummar, S.; Khan, F.G.; Shah, S.; Khan, A.; Shamshirband, S.; Rehman, Z.U.; Khan, I.A.; Jadoon, W. A Deep Learning Ensemble Approach for Diabetic Retinopathy Detection. IEEE Access 2019, 7, 150530–150539. [Google Scholar] [CrossRef]
- Pan, X.; Jin, K.; Cao, J.; Liu, Z.; Wu, J.; You, K.; Lu, Y.; Xu, Y.; Su, Z.; Jiang, J.; et al. Multi-label classification of retinal lesions in diabetic retinopathy for automatic analysis of fundus fluorescein angiography based on deep learning. Graefe’s Arch. Clin. Exp. Ophthalmol. 2020, 258, 779–785. [Google Scholar] [CrossRef]
- Shankar, K.; Sait, A.R.W.; Gupta, D.; Lakshmanaprabu, S.; Khanna, A.; Pandey, H.M.; Kathiresan, S. Automated detection and classification of fundus diabetic retinopathy images using synergic deep learning model. Pattern Recognit. Lett. 2020, 133, 210–216. [Google Scholar] [CrossRef]
- Riaz, H.; Park, J.; Choi, H.; Kim, H.; Kim, J. Deep and Densely Connected Networks for Classification of Diabetic Retinopathy. Diagnostics 2020, 10, 24. [Google Scholar] [CrossRef] [PubMed]
- Agurto, C.; Murray, V.; Yu, H.; Wigdahl, J.; Pattichis, C.S.; Nemeth, S.; Barriga, E.S.; Soliz, P. A Multiscale Optimization Approach to Detect Exudates in the Macula. IEEE J. Biomed. Health Inf. 2014, 18, 1328–1336. [Google Scholar] [CrossRef] [PubMed]
- Rekhi, R.S.; Issac, A.; Dutta, M.K.; Travieso, C.M. Automated classification of exudates from digital fundus images. In Proceedings of the 2017 International Conference and Workshop on Bioinspired Intelligence (IWOBI), Funchal, Portugal, 10–12 July 2017; pp. 1–6. [Google Scholar]
- Dey, A.; Dey, K.N. Automated Glaucoma Detection from Fundus Images of Eye Using Statistical Feature Extraction Methods and Support Vector Machine Classification. In Industry Interactive Innovations in Science, Engineering and Technology; Lecture Notes in Networks and Systems; Springer: Singapore, 2016; Volume 11, pp. 511–521. [Google Scholar]
- Krishnan, M.M.R.; Faust, O. Automated glaucoma detection using hybrid feature extraction in retinal fundus images. J. Mech. Med. Biol. 2013, 13, 1350011. [Google Scholar] [CrossRef]
- Khan, M.W.; Sharif, M.; Yasmin, M.; Fernandes, S.L. A new approach of cup to disk ratio based glaucoma detection using fundus images. J. Integr. Des. Process. Sci. 2016, 20, 77–94. [Google Scholar] [CrossRef]
- Nayak, J.; Acharya, R.; Bhat, P.S.; Shetty, N.; Lim, T.C. Automated diagnosis of glaucoma using digital fundus images. J. Med. Syst. 2009, 33, 337–346. [Google Scholar] [CrossRef]
- Salam, A.A.; Akram, M.U.; Wazir, K.; Anwar, S.M.; Majid, M. Autonomous Glaucoma detection from fundus image using cup to disc ratio and hybrid features. In Proceedings of the 2015 IEEE International Symposium on Signal Processing and Information Technology (ISSPIT), Abu Dhabi, UAE, 7–10 December 2015; pp. 370–374. [Google Scholar] [CrossRef]
- Agrawal, D.K.; Kirar, B.S.; Pachori, R.B. Automated glaucoma detection using quasi-bivariate variational mode decomposition from fundus images. IET Image Process. 2019, 13, 2401–2408. [Google Scholar] [CrossRef]
- Dromain, C.; Boyer, B.; Ferre, R.; Canale, S.; Delaloge, S.; Balleyguier, C. Computed-aided diagnosis (CAD) in the detection of breast cancer. Eur. J. Radiol. 2013, 82, 417–423. [Google Scholar] [CrossRef]
- Abràmoff, M.D.; Lou, Y.; Erginay, A.; Clarida, W.; Amelon, R.; Folk, J.C.; Niemeijer, M. Improved Automated Detection of Diabetic Retinopathy on a Publicly Available Dataset Through Integration of Deep Learning. Investig. Opthalmol. Vis. Sci. 2016, 57, 5200. [Google Scholar] [CrossRef]
- Zeng, X.; Chen, H.; Luo, Y.; Bin Ye, W. Automated Diabetic Retinopathy Detection Based on Binocular Siamese-Like Convolutional Neural Network. IEEE Access 2019, 7, 30744–30753. [Google Scholar] [CrossRef]
- Ren, F.; Cao, P.; Zhao, D.; Wan, C. Diabetic macular edema grading in retinal images using vector quantization and semi-supervised learning. Technol. Health Care 2018, 26, 389–397. [Google Scholar] [CrossRef] [PubMed]
- Marín, D.; Gegundez-Arias, M.E.; Ponte, B.; Alvarez, F.; Garrido, J.; Ortega, C.; Vasallo, M.J.; Bravo, J.M. An exudate detection method for diagnosis risk of diabetic macular edema in retinal images using feature-based and supervised classification. Med. Biol. Eng. 2018, 56, 1379–1390. [Google Scholar] [CrossRef] [PubMed]
- Perdomo, O.; Otalora, S.; Rodríguez, F.; Arevalo, J.; González, F.A.; Chen, X.; Garvin, M.K.; Liu, J.; Trucco, E.; Xu, Y. A Novel Machine Learning Model Based on Exudate Localization to Detect Diabetic Macular Edema. In Proceedings of the Ophthalmic Medical Image Analysis Third International Workshop, Athens, Greece, 21 October 2016; pp. 137–144. [Google Scholar] [CrossRef]
- Jiang, Y.; Duan, L.; Cheng, J.; Gu, Z.; Xia, H.; Fu, H.; Li, C.; Liu, J. JointRCNN: A Region-Based Convolutional Neural Network for Optic Disc and Cup Segmentation. IEEE Trans. Biomed. Eng. 2020, 67, 335–343. [Google Scholar] [CrossRef]
- Shankaranarayana, S.M.; Ram, K.; Mitra, K.; Sivaprakasam, M. Joint optic disc and cup segmentation using fully convolutional and adversarial networks. In Fetal, Infant and Ophthalmic Medical Image Analysis; Springer: Berlin/Heidelberg, Germany, 2017; pp. 168–176. [Google Scholar]
- Raghavendra, U.; Fujita, H.; Bhandary, S.V.; Gudigar, A.; Tan, J.H.; Acharya, U.R. Deep convolution neural network for accurate diagnosis of glaucoma using digital fundus images. Inf. Sci. 2018, 441, 41–49. [Google Scholar] [CrossRef]
- Sevastopolsky, A. Optic disc and cup segmentation methods for glaucoma detection with modification of U-Net convolutional neural network. Pattern Recognit. Image Anal. 2017, 27, 618–624. [Google Scholar] [CrossRef]
- Abbas, Q. Glaucoma-Deep: Detection of Glaucoma Eye Disease on Retinal Fundus Images using Deep Learning. Int. J. Adv. Comput. Sci. Appl. 2017, 8, 41–45. [Google Scholar] [CrossRef]
- Bajwa, M.N.; Malik, M.I.; Siddiqui, S.A.; Dengel, A.; Shafait, F.; Neumeier, W.; Sheraz Ahmed, S. Two-stage framework for optic disc localization and glaucoma classification in retinal fundus images using deep learning. BMC Med. Inf. Decis. Mak. 2019, 19, 136. [Google Scholar]
- Lu, Z.; Chen, D. Weakly Supervised and Semi-Supervised Semantic Segmentation for Optic Disc of Fundus Image. Symmetry 2020, 12, 145. [Google Scholar] [CrossRef]
- Ramani, R.G.; Shanthamalar, J.J. Improved image processing techniques for optic disc segmentation in retinal fundus images. Biomed. Signal Process. Control. 2020, 58, 101832. [Google Scholar] [CrossRef]
- Zhang, W.; Zhong, J.; Yang, S.; Gao, Z.; Hu, J.; Chen, Y.; Yi, Z. Automated identification and grading system of diabetic retinopathy using deep neural networks. Know. Based Syst. 2019, 175, 12–25. [Google Scholar] [CrossRef]
- De La Torre, J.; Valls-Mateu, A.; Puig, D. A deep learning interpretable classifier for diabetic retinopathy disease grading. Neurocomputing 2020, 396, 465–476. [Google Scholar] [CrossRef]
- Tzutalin. LabelImg. Available online: https://github.com/tzutalin/labelImg (accessed on 11 January 2020).
- Wang, X.; Shrivastava, A.; Gupta, A. A-Fast-RCNN: Hard Positive Generation via Adversary for Object Detection. In Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017; pp. 3039–3048. [Google Scholar]
- He, G.K.; Zhang, X.; Ren, S.; Sun, J. Spatial pyramid pooling in deep convolutional networks for visual recognition. IEEE Trans. Pattern Anal. Mach. Intell. 2015, 37, 1904–1916. [Google Scholar] [CrossRef] [PubMed]
- Girshick, R.; Donahue, J.; Darrell, T.; Malik, J.; Malik, J. Rich Feature Hierarchies for Accurate Object Detection and Semantic Segmentation. In Proceedings of the 2014 IEEE Conference on Computer Vision and Pattern Recognition, Columbus, OH, USA, 24–27 June 2014; pp. 580–587. [Google Scholar]
- Denton, E.L.; Zaremba, W.; Bruna, J.; LeCun, Y.; Fergus, R. Exploiting linear structure within convolutional networks for efficient evaluation. In Advances in Neural Information Processing Systems; MIT Press: Cambridge, MA, USA, 2014; pp. 1269–1277. [Google Scholar]
- Girshick, R. Fast R-CNN. In Proceedings of the 2015 IEEE International Conference on Computer Vision (ICCV), Las Condes, Chile, 11–18 December 2015; pp. 1440–1448. [Google Scholar]
- Qin, Z.; Kim, D. Rethinking Softmax with Cross-Entropy: Neural Network Classifier as Mutual Information Estimator. 2019. Available online: https://arxiv.org/abs/1911.10688 (accessed on 25 January 2020).
- Heil, J.; Häring, V.; Marschner, B.; Stumpe, B. Advantages of fuzzy k-means over k-means clustering in the classification of diffuse reflectance soil spectra: A case study with West African soils. Geoderma 2019, 337, 11–21. [Google Scholar] [CrossRef]
- Rashid, J.; Shah, S.M.A.; Irtaza, A.; Mahmood, T.; Nisar, M.W.; Shafiq, M.; Gardezi, A. Topic Modeling Technique for Text Mining Over Biomedical Text Corpora through Hybrid Inverse Documents Frequency and Fuzzy K-Means Clustering. IEEE Access 2019, 7, 146070–146080. [Google Scholar] [CrossRef]
- Kauppi, T.; Kalesnykiene, V.; Kamarainen, J.-K.; Lensu, L.; Sorri, I.; Raninen, A.; Voutilainen, R.; Uusitalo, H.; Kalviainen, H.; Pietilä, J. The DIARETDB1 diabetic retinopathy database and evaluation protocol. BMVC 2007, 1, 1–10. [Google Scholar]
- Odstrcilik, J.; Kolar, R.; Kubena, T.; Cernosek, P.; Budai, A.; Hornegger, J.; Gazarek, J.; Svoboda, O.; Jan, J.; Angelopoulou, E. Retinal vessel segmentation by improved matched filtering: Evaluation on a new high-resolution fundus image database. IET Image Process. 2013, 7, 373–383. [Google Scholar] [CrossRef]
- Holm, S.; Russell, G.; Nourrit, V.; McLoughlin, N. DR HAGIS—A fundus image database for the automatic extraction of retinal surface vessels from diabetic patients. J. Med. Imaging 2017, 4, 14503. [Google Scholar] [CrossRef]
- Zhang, Z.Z.; Yin, F.S.; Liu, J.J.; Wong, W.K.; Tan, N.M.; Lee, B.H.; Cheng, J.; Wong, T.Y. ORIGA-light: An online retinal fundus image database for glaucoma analysis and research. In Proceedings of the 2010 Annual International Conference of the IEEE Engineering in Medicine and Biology, Buenos Aires, Argentina, 31 August–4 September 2010; Volume 2010, pp. 3065–3068. [Google Scholar]
- Arafat, Y.; Iqbal, M.J. Urdu-Text Detection and Recognition in Natural Scene Images Using Deep Learning. IEEE Access 2020, 8, 96787–96803. [Google Scholar] [CrossRef]
- Girshick, R.; Donahue, J.; Darrell, T.; Malik, J. Region-Based Convolutional Networks for Accurate Object Detection and Segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2015, 38, 142–158. [Google Scholar] [CrossRef]
- Gulshan, V.; Peng, L.; Coram, M.; Stumpe, M.C.; Wu, D.; Narayanaswamy, A.; Venugopalan, S.; Widner, K.; Madams, T.; Cuadros, J.; et al. Development and Validation of a Deep Learning Algorithm for Detection of Diabetic Retinopathy in Retinal Fundus Photographs. JAMA 2016, 316, 2402–2410. [Google Scholar] [CrossRef]
- Zhou, L.; Zhao, Y.; Yang, J.; Yu, Q.; Xu, X. Deep multiple instance learning for automatic detection of diabetic retinopathy in retinal images. IET Image Process. 2018, 12, 563–571. [Google Scholar] [CrossRef]
- Kaur, J.; Mittal, D. A generalized method for the segmentation of exudates from pathological retinal fundus images. Biocybern. Biomed. Eng. 2018, 38, 27–53. [Google Scholar] [CrossRef]
- Abbas, Q.; Fondon, I.; Sarmiento, A.; Jiménez, S.; Alemany, P. Automatic recognition of severity level for diagnosis of diabetic retinopathy using deep visual features. Med. Biol. Eng. 2017, 55, 1959–1974. [Google Scholar] [CrossRef] [PubMed]
- Granero, A.C.; Igual, J.; Naranjo, V. Detection of Early Signs of Diabetic Retinopathy Based on Textural and Morphological Information in Fundus Images. Sensors 2020, 20, 1005. [Google Scholar] [CrossRef]
- Li, X.; Hu, X.; Yu, L.; Zhu, L.; Fu, C.-W.; Heng, P. CANet: Cross-Disease Attention Network for Joint Diabetic Retinopathy and Diabetic Macular Edema Grading. IEEE Trans. Med. Imaging 2020, 39, 1483–1493. [Google Scholar] [CrossRef]
- Deepak, K.S.; Medathati, N.K.; Sivaswamy, J. Detection and discrimination of disease-related abnormalities based on learning normal cases. Pattern Recognit. 2012, 45, 3707–3716. [Google Scholar] [CrossRef]
- Medhi, J.P.; Dandapat, S. An effective fovea detection and automatic assessment of diabetic maculopathy in color fundus images. Comput. Biol. Med. 2016, 74, 30–44. [Google Scholar] [CrossRef]
- Lim, S.; Zaki, W.M.D.W.; Hussain, A.; Kusalavan, S.; Lim, S.L. Automatic classification of diabetic macular edema in digital fundus images. In Proceedings of the 2011 IEEE Colloquium on Humanities, Science and Engineering, Penang, Malaysia, 5–6 December 2011; pp. 265–269. [Google Scholar]
- Rahim, S.; Palade, V.; Jayne, C.; Holzinger, A.; Shuttleworth, J. Detection of diabetic retinopathy and maculopathy in eye fundus images using fuzzy image processing. In International Conference on Brain Informatics and Health; Springer: Cham, Switzerland, 2015; pp. 379–388. [Google Scholar]
- Syed, A.M.; Akram, M.; Akram, T.; Muzammal, M.; Khalid, S.; Khan, M.A. Fundus Images-Based Detection and Grading of Macular Edema Using Robust Macula Localization. IEEE Access 2018, 6, 58784–58793. [Google Scholar] [CrossRef]
- Varadarajan, A.V.; Bavishi, P.; Ruamviboonsuk, P.; Chotcomwongse, P.; Venugopalan, S.; Narayanaswamy, A.; Cuadros, J.; Kanai, K.; Bresnick, G.; Tadarati, M.; et al. Predicting optical coherence tomography-derived diabetic macular edema grades from fundus photographs using deep learning. Nat. Commun. 2020, 11, 130–138. [Google Scholar] [CrossRef]
- He, X.; Zhou, Y.; Wang, B.; Cui, S.; Shao, L. DME-Net: Diabetic Macular Edema Grading by Auxiliary Task Learning. In Medical Image Computing and Computer Assisted Intervention—MICCAI 2019. MICCAI 2019; Lecture Notes in Computer Science; Springer: Cham, Switzerland, 2019; Volume 11764, pp. 788–796. [Google Scholar]
- Liao, W.; Zou, B.; Zhao, R.; Chen, Y.; He, Z.; Zhou, M. Clinical Interpretable Deep Learning Model for Glaucoma Diagnosis. IEEE J. Biomed. Health Informatics 2019, 24, 1405–1412. [Google Scholar] [CrossRef]
- Chen, X.; Xu, Y.; Yan, S.; Wong, D.W.K.; Wong, T.Y.; Liu, J. Automatic Feature Learning for Glaucoma Detection Based on Deep Learning. In Proceedings of the Bioinformatics Research and Applications; Springer Science and Business Media LLC: Berlin/Heidelberg, Germany, 2015; pp. 669–677. [Google Scholar]
- Xu, Y.; Lin, S.; Wong, D.W.K.; Liu, J.; Xu, D. Efficient Reconstruction-Based Optic Cup Localization for Glaucoma Screening. In Proceedings of the Computer Vision; Springer Science and Business Media LLC: Berlin/Heidelberg, Germany, 2013; pp. 445–452. [Google Scholar]
- Li, A.; Cheng, J.; Wong, D.W.K.; Liu, J. Integrating holistic and local deep features for glaucoma classification. In Proceedings of the 2016 38th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Orlando, FL, USA, 16–20 August 2016; pp. 1328–1331. [Google Scholar]
- Prakash, N.B.; Selvathi, D. An Efficient Detection System for Screening Glaucoma InRetinal Images. Biomed. Pharmacol. J. 2017, 10, 459–465. [Google Scholar] [CrossRef]


| Reference | Method | Findings | Shortcomings |
|---|---|---|---|
| Pan et al. [15] | The model is based on DenseNet, ResNet50, and VGG16 models | Detection of DR lesions and the method is computationally robust. | The method may not classify microaneurysms efficiently because these are easily misclassified under the pervading presence of fluorescein |
| Zeng et al. [28] | A Siamese-like CNN framework by employing the concept of weigh-sharing layers based on Inception V3. | The work exhibits promising results for DR prediction with kappa score of 0.829. | It may not perform well for those sample databases where paired fundus images are not available. |
| Qummar et al. [14] | Five different frameworks of CNN named Dense169, Inception V3, Resnet50, Dense121, and Xception were employed. | To locate and classify the DR lesions into different classes according to the severity level of moles. | The method suffers from high computational cost. |
| Zhang et al. [40] | A DL framework named DeepDR was presented for DR detection. In addition, a new database for DR labelled DR images was also introduced. | The proposed network has attained the sensitivity value of 97.5%, along with the specificity value of 97.7%. | The introduced model needs to be evaluated on more complex and larger dataset. |
| Torre et al. [41] | A DL based method was used to predict the expected DR class and assign scores to individual pixels to exhibit their relevance in each input sample. The assigned score was employed to take final classification decision. | The introduced DL framework acquired more than 90% of sensitivity and specificity values. | The evaluation performance of the presented algorithm can be improved through appropriate measures. |
| Rekhi et al. [2] | The method was based on geometrical, morphological, and orientation features. The classification was performed through SVM. | Grading and classification of DME from fundus images with an accuracy of 92.11%. | The detection accuracy needs further improvement. |
| Kunwar et al. [5] | The method was based on texture features and the SVM classifier. | high-risk DME detection with accuracy of 86%. | Experiments were performed on small dataset. |
| Marin et al. [30] | The method was based on thresholding and regularized regression techniques. | DME risk detection with 0.90 sensitivity. | The detection performance requires improvement. |
| Perdomo et al. [31] | The presented method was composed of two-stage CNNs | The method detects regions of interest in the retinal image and then predicts its class of DME | The technique is computationally complex. |
| Jiang et al. [32] | The end-to-end Region- based Convolutional Neural Network was used for OD and OC segmentation. | OD and OC segmentation with AUC of 0.901. The method is robust to glaucoma detection. | The method is computationally complex because it employs two separate RCNNs to compute the bboxes of the OC and OD, respectively. |
| Bajwa et al. [37] | The localization was achieved through RCNN, while the other stage used deep CNN to classify the computed disc into glaucomatous or healthy. | Localization and classification of glaucoma with AUC of 0.874. | The method is computationally complex as it takes two-stage framework to localize and classify the glaucoma. The performance is affected by increasing the network hierarchy as it results in losing the discriminative set of features. |
| Zheng Lu et al. [38] | The Modified U-Net model was improved by minimizing the original U-shape structure through adding 2-dimensional convolutional layer. | Before OD segmentation, the ground-truths were generated through the GrabCut method. | The presented technique requires less training, however, shows lower segmentation accuracy as compare latest approaches because of missing ground truths. |
| Ramani et al. [39] | The region-based pixel density calculation method based on Circular Hough Transform with Hough Peak Value Selection and Red Channel Super-pixel method. | The technique is robust and efficient to optic disc segmentation. | The detection accuracy is affected over the images having pathological distractions. |
| DR | DME | Glaucoma |
|---|---|---|
| 0. Background 1. Microaneurysms 2. Soft Exudates 3. Hard Exudates 4. Hemorrhages | 0. Background 1. Macula Region | 0. Background 1. OD 2. OC |
| Technique | mAP |
|---|---|
| SPPnet [44] | 0.85 |
| RCNN [56] | 0.89 |
| FRCNN (Proposed) | 0.94 |
| Dataset | Img1 | Img2 | Img3 | Img4 | Img5 | Img6 | Img7 | Img8 | mAP |
|---|---|---|---|---|---|---|---|---|---|
| HRF | 0.949 | 0.942 | 0.963 | 0.935 | 0.99 | 0.891 | 0.953 | 0.941 | 0.946 |
| DR HAGIS | 0.937 | 0.94 | 0.975 | 0.891 | 0.912 | 0.939 | 0.99 | 0.933 | 0.940 |
| ORIGA | 0.941 | 0.935 | 0.899 | 0.876 | 0.97 | 0.99 | 0.94 | 0.95 | 0.938 |
| DR Signs | Acc | SP | SE |
|---|---|---|---|
| Hard Exudates | 0.958 | 0.941 | 0.957 |
| Soft Exudates | 0.943 | 0.961 | 0.955 |
| Micro aneurysms | 0.957 | 0.954 | 0.943 |
| Hemorrhages | 0.952 | 0.96 | 0.951 |
| Database | Acc | Dc | SP | SE |
|---|---|---|---|---|
| HRF | 0.958 | 0.952 | 0.97 | 0.957 |
| DR HAGIS | 0.943 | 0.89 | 0.961 | 0.955 |
| ORIGA | 0.957 | 0.943 | 0.954 | 0.943 |
| Average | 0.9526 | 0.9283 | 0.961 | 0.951 |
| Technique | SP | SE | Acc | AUC |
|---|---|---|---|---|
| Zeng et al. [28] | 0.635 | 0.77 | - | 0.94 |
| Gulshan et al. [57] | 0.91 | - | 0.913 | 0.96 |
| Zhou et al. [58] | 0.863 | 0.995 | - | - |
| Kaur et al. [59] | 0.96 | 0.88 | 0.93 | - |
| Colomer et al. [61] | 0.818 | 0.81 | 0.93 | - |
| Abbas et al. [60] | 0.94 | 0.92 | - | 0.92 |
| Proposed | 0.965 | 0.961 | 0.95 | 0.967 |
| Method | SE | SP | Acc |
|---|---|---|---|
| Li et al. [62] | 0.70 | 0.76 | 0.912 |
| Deepak et al. [63] | 0.95 | 0.90 | - |
| Medhi et al. [64] | 0.95 | 0.95 | - |
| Rekhi et al. [2] | - | - | 0.921 |
| Lim et al. [65] | 0.80 | 0.90 | - |
| Rahim et al. [66] | 0.85 | 0.55 | - |
| Syed et al. [67] | 0.96 | 0.95 | 0.935 |
| Varadarajan et al. [68] | 0.85 | 0.80 | - |
| Xiaodong et al. [69] | 0.959 | 0.97 | - |
| Proposed | 0.96 | 0.958 | 0.958 |
| Method | Year | Dataset | SE | SP | AUC | Dc | Time |
|---|---|---|---|---|---|---|---|
| Liao et al. [70] | 2019 | ORIGA | - | - | 0.88 | 0.9 | - |
| Chen et al. [71] | 2015 | ORIGA | - | - | 0.838 | - | - |
| Xu et al. [72] | 2013 | ORIGA | 0.58 | - | 0.823 | - | - |
| Li et al. [73] | 2016 | ORIGA | - | - | 0.8384 | - | - |
| Bajwa et al. [37] | 2019 | ORIGA | 0.71 | - | 0.868 | - | - |
| Ramani et al. [39] | 2020 | HRF | 0.849 | 0.999 | - | - | 1.49 s |
| N.B Parakash et al. [74] | 2017 | HRF | 0.7025 | 0.997 | - | - | - |
| Krishna et al. [12] | 2019 | DR HAGIS | 0.94 | - | - | - | - |
| Proposed | ORIGA | 0.941 | 0.945 | 0.94 | 0.943 | 0.9 s | |
| HRF | 0.95 | 0.96 | 0.963 | 0.952 | 0.9 s | ||
| DR HAGIS | 0.945 | 0.941 | 0.94 | 0.89 | 0.9 s |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nazir, T.; Irtaza, A.; Javed, A.; Malik, H.; Hussain, D.; Naqvi, R.A. Retinal Image Analysis for Diabetes-Based Eye Disease Detection Using Deep Learning. Appl. Sci. 2020, 10, 6185. https://doi.org/10.3390/app10186185
Nazir T, Irtaza A, Javed A, Malik H, Hussain D, Naqvi RA. Retinal Image Analysis for Diabetes-Based Eye Disease Detection Using Deep Learning. Applied Sciences. 2020; 10(18):6185. https://doi.org/10.3390/app10186185
Chicago/Turabian StyleNazir, Tahira, Aun Irtaza, Ali Javed, Hafiz Malik, Dildar Hussain, and Rizwan Ali Naqvi. 2020. "Retinal Image Analysis for Diabetes-Based Eye Disease Detection Using Deep Learning" Applied Sciences 10, no. 18: 6185. https://doi.org/10.3390/app10186185
APA StyleNazir, T., Irtaza, A., Javed, A., Malik, H., Hussain, D., & Naqvi, R. A. (2020). Retinal Image Analysis for Diabetes-Based Eye Disease Detection Using Deep Learning. Applied Sciences, 10(18), 6185. https://doi.org/10.3390/app10186185









