Molecular and Antigenic Characterization of GI-13 and GI-16 Avian Infectious Bronchitis Virus Isolated in Chile from 2009 to 2017 Regarding 4/91 Vaccine Introduction
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Embryonated Chicken Eggs and Chicks
2.2. Viruses
2.3. Molecular Characterization
2.4. Virus Neutralization Test
2.5. Protection Test
2.6. Design of RT-qPCR for IBV
2.7. Statistical Analysis
2.8. Accession Number of Sequence Data
3. Results
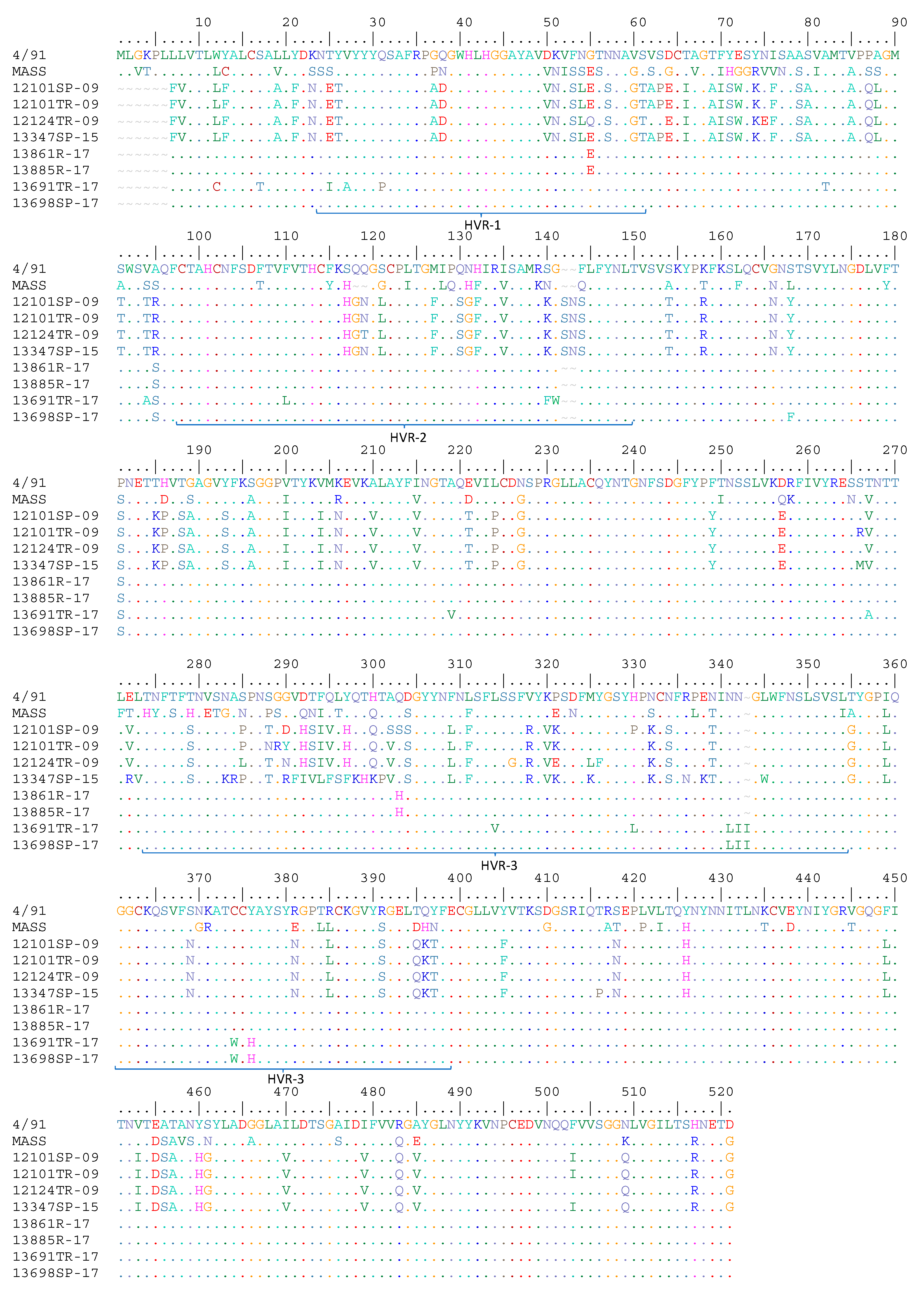
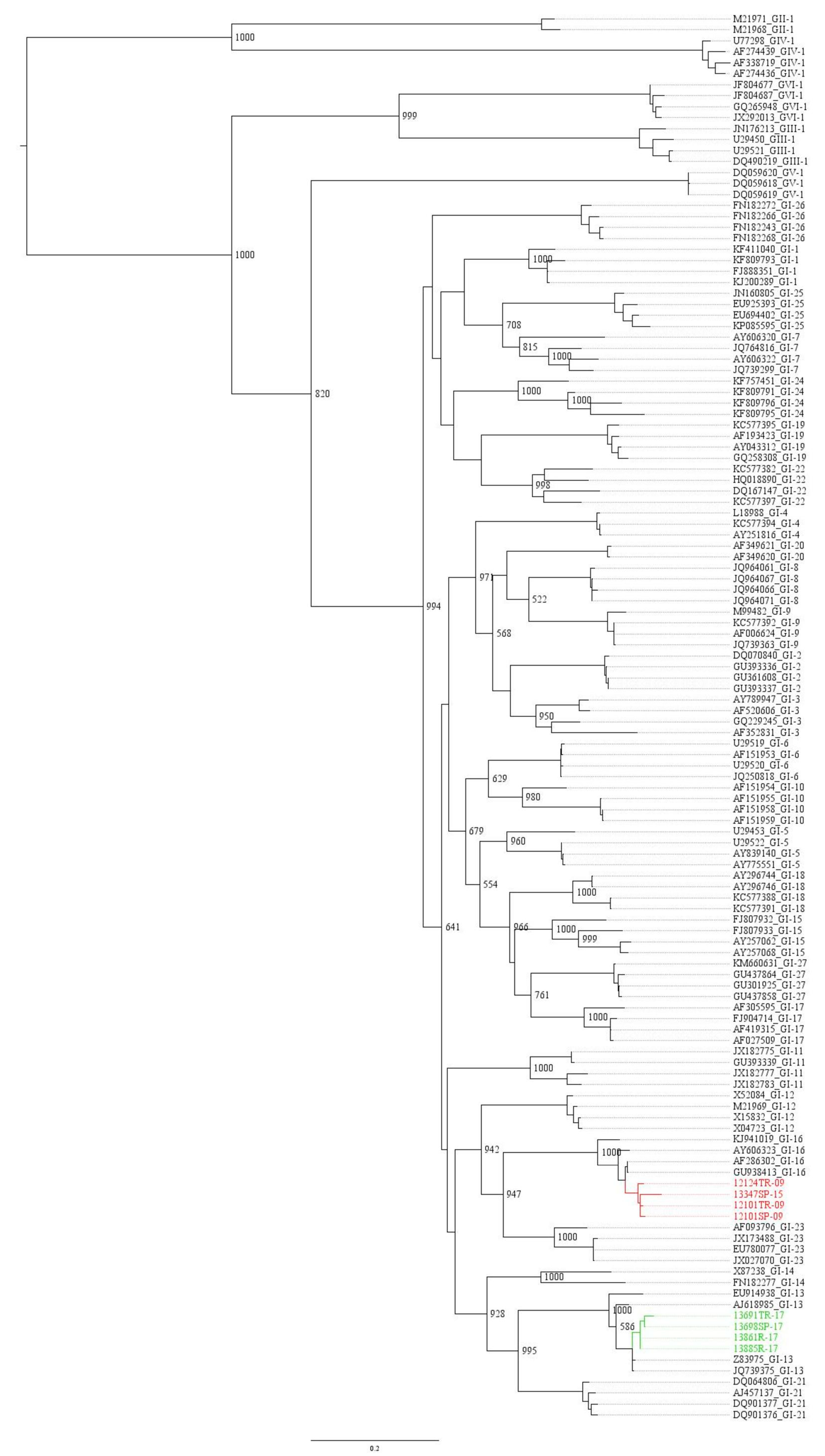
3.1. Molecular Characterization
3.2. Virus Neutralization Test
3.3. Protection Test
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Colvero, L.P.; Villarreal, L.Y.B.; Torres, C.A.; Brañdo, P.E. Assessing the economic burden of avian infectious bronchitis on poultry farms in Brazil. Rev. Sci. Tech. 2015, 34, 993–999. [Google Scholar] [CrossRef] [Green Version]
- Masters, P.S. The Molecular Biology of Coronaviruses. Adv. Virus Res. 2006, 65, 193–292. [Google Scholar]
- Denison, M.R.; Graham, R.L.; Donaldson, E.F.; Eckerle, L.D.; Baric, R.S. Coronaviruses: An RNA proofreading machine regulates replication fidelity and diversity. RNA Biol. 2011, 8. [Google Scholar] [CrossRef]
- Cavanagh, D.; Davis, P.J.; Mockett, A.P.A. Amino acids within hypervariable region 1 of avian coronavirus IBV (Massachusetts serotype) spike glycoprotein are associated with neutralization epitopes. Virus Res. 1988, 11, 141–150. [Google Scholar] [CrossRef]
- Kant, A.; Koch, G.; Van Roozelaar, D.J.; Kusters, J.G.; Poelwijk, F.A.J.; Van der Zeijst, B.A.M. Location of antigenic sites defined by neutralizing monoclonal antibodies on the S1 avian infectious bronchitis virus glycopolypeptide. J. Gen. Virol. 1992, 73, 591–596. [Google Scholar] [CrossRef] [PubMed]
- Abro, S.H.; Renström, L.H.M.; Ullman, K.; Isaksson, M.; Zohari, S.; Jansson, D.S.; Belák, S.; Baule, C. Emergence of novel strains of avian infectious bronchitis virus in Sweden. Vet. Microbiol. 2012, 155, 237–246. [Google Scholar] [CrossRef] [PubMed]
- Fellahi, S.; El Harrak, M.; Ducatez, M.; Loutfi, C.; Koraichi, S.I.S.; Kuhn, J.H.; Khayi, S.; El Houadfi, M.; Ennaji, M.M. Phylogenetic analysis of avian infectious bronchitis virus S1 glycoprotein regions reveals emergence of a new genotype in Moroccan broiler chicken flocks. Virol. J. 2015, 12. [Google Scholar] [CrossRef]
- Lee, C.W.; Jackwood, M.W. Evidence of genetic diversity generated by recombination among avian coronavirus IBV. Arch. Virol. 2000, 145, 2135–2148. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lim, T.H.; Lee, H.J.; Lee, D.H.; Lee, Y.N.; Park, J.K.; Youn, H.N.; Kim, M.S.; Lee, J.B.; Park, S.Y.; Choi, I.S.; et al. An emerging recombinant cluster of nephropathogenic strains of avian infectious bronchitis virus in Korea. Infect. Genet. Evol. 2011, 11, 678–685. [Google Scholar] [CrossRef]
- Moreno, A.; Franzo, G.; Massi, P.; Tosi, G.; Blanco, A.; Antilles, N.; Biarnes, M.; Majó, N.; Nofrarías, M.; Dolz, R.; et al. A novel variant of the infectious bronchitis virus resulting from recombination events in Italy and Spain. Avian Pathol. 2017, 46, 28–35. [Google Scholar] [CrossRef]
- Cook, J.K.A.; Jackwood, M.; Jones, R.C. The long view: 40 years of infectious bronchitis research. Avian Pathol. 2012, 41, 239–250. [Google Scholar] [CrossRef] [PubMed]
- Hidalgo, H.; Gallardo, R.; Rosende, S. Isolation of Infectious Bronchitis Virus from broiler chickens in Chile. Avian Dis. 1976, 20, 601–603. [Google Scholar] [CrossRef] [PubMed]
- de Wit, J.J.; Dijkman, R.; Guerrero, P.; Calvo, J.; Gonzalez, A.; Hidalgo, H. Variability in biological behaviour, pathogenicity, protectotype and induction of virus neutralizing antibodies by different vaccination programmes to infectious bronchitis virus genotype Q1 strains from Chile. Avian Pathol. 2017, 46, 666–675. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Valastro, V.; Holmes, E.C.; Britton, P.; Fusaro, A.; Jackwood, M.W.; Cattoli, G.; Monne, I. S1 gene-based phylogeny of infectious bronchitis virus: An attempt to harmonize virus classification. Infect. Genet. Evol. 2016, 39, 349–364. [Google Scholar] [CrossRef] [PubMed]
- Malo, A.; Orbell, S.; Huggins, M.; Woods, M.; Cook, J. Cross protection studies after the use of live- attenuated IBV vaccines Nobilis® IB 4-91 and Nobilis® IB Ma5 (Massachusetts type). Intervet VSD Newsl. 1998, 5, 1–6. [Google Scholar]
- Taylor, P.; Cook, J.K.A.; Orbell, S.J.; Woods, M.A.; Michael, B. Breadth of protection of the respiratory tract provided by different live-attenuated infectious bronchitis vaccines against challenge with infectious bronchitis viruses of heter. Avian Pathol. 1999, 28, 477–485. [Google Scholar]
- Terregino, C.; Toffan, A.; Serena Beato, M.; De Nardi, R.; Vascellari, M.; Meini, A.; Ortali, G.; Mancin, M.; Capua, I. Pathogenicity of a QX strain of infectious bronchitis virus in specific pathogen free and commercial broiler chickens, and evaluation of protection induced by a vaccination programme based on the Ma5 and 4/91 serotypes. Avian Pathol. 2008, 37, 487–493. [Google Scholar] [CrossRef] [PubMed]
- De Wit, J.J.; Brandao, P.; Torres, C.A.; Koopman, R.; Villarreal, L.Y. Increased level of protection of respiratory tract and kidney by combining different infectious bronchitis virus vaccines against challenge with nephropathogenic Brazilian genotype subcluster 4 strains. Avian Pathol. 2015, 44, 352–357. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Belkasmi, S.F.Z.; Fellahi, S.; Umar, S.; Delpont, M.; Delverdier, M.; Lucas, M.-N.; Bleuart, C.; Kichou, F.; Nassik, S.; Guerin, J.-L.; et al. Efficacy of Massachusetts and 793B Vaccines Against Infectious Bronchitis Moroccan-Italy 02 Virus in Specific-Pathogen-Free Chickens and Commercial Broilers. Avian Dis. 2017, 61, 466–471. [Google Scholar] [CrossRef]
- Polson, A. Isolation of IgY from the Yolks of Eggs by a Chloroform Polyethylene Glycol Procedure. Inmunol. Investig. 1990, 19, 253–258. [Google Scholar]
- Gelb, J. patente seq RNA.pdf. In A laboratory Manual for the Isolation and Identification of Avian Pathogens; Swayne, D.E., Glisson, J.R., Pearson, J.E., Reed, W.M., Jackwood, M.W., Woolcock, P.R., Eds.; American Association of Avian Pathologists: Athens, Greece, 2008; pp. 146–149. [Google Scholar]
- Jackwood, M.W.; de Wit, J.J. Infectious Bronchitis. In Disease of Poultry; Swayne, D.E., McDougald, L.R., Nolan, L.K., Suarez, D.L., Nair, V.L., Eds.; Wiley-Blackwell: Marblehead, MA, USA, 2013; pp. 117–135. [Google Scholar]
- Zhang, Z.; Zhou, Y.; Wang, H.; Zeng, F.; Yang, X.; Zhang, Y.; Zhang, A. Molecular detection and Smoothing spline clustering of the IBV strains detected in China during 2011–2012. Virus Res. 2016, 211, 145–150. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Fungiflora, O.; Gascuel, O. A simple, fast and accurate method to estimate large phylogenies by maximum-likelihood. Syst. Biol. 2003, 52, 696–704. [Google Scholar]
- Guindon, S.; Dufayard, J.F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New algorithms and methods to estimate maximum-likelihood phylogenies: Assessing the performance of PhyML 3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef] [PubMed]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Reed, L.J.; Muench, H. A simple method of estimating fifty per cent endpoints. Am. J. Epidemiol. 1938, 27, 493–497. [Google Scholar] [CrossRef]
- Archetti, I.; Horsfall, F.L. Persistent antigenic variation of Influenza A viruses after incomplete neutralization in ovo with heterologous immune serum. J. Exp. Med. 1950, 92, 441–462. [Google Scholar] [CrossRef] [PubMed]
- Brooksby, J.B. Variants and immunity in foot and mouth disease. Symp. Ser. Immunobiol. Stand. 1968, 8, 1–10. [Google Scholar]
- Marandino, A.; Pereda, A.; Tomás, G.; Hernández, M.; Iraola, G.; Craig, M.I.; Hernaández, D.; Banda, A.; Villegas, P.; Panzera, Y.; et al. Phylodynamic analysis of avian infectious bronchitis virus in South America. J. Gen. Virol. 2015, 96, 1340–1346. [Google Scholar] [CrossRef]
- Tataje-Lavanda, L.; Izquierdo-Lara, R.; Ormeño-Vásquez, P.; Huamán-Gutiérrez, K.; Zimic-Peralta, M.; Fernández-Díaz, M. Near-Complete Genome Sequence of Infectious Bronchitis Virus Strain VFAR-047 (GI-16 Lineage), Isolated in Peru. Microbiol. Resour. Announc. 2019, 8, e01555-18. [Google Scholar] [CrossRef] [Green Version]
- Nabavi, H.; Karimi, V.; Ghalyanchilangeroudi, A.; Shateri, S.; Seger, W.; Najafi, H. Phylogenetic analysis and full-length characterization of S1 gene of IS-1494 (Variant 2) like infectious bronchitis virus isolates, Iran, 2015. Virol. Rep. 2016, 6, 18–24. [Google Scholar] [CrossRef] [Green Version]
- Shimazaki, Y.; Watanabe, Y.; Harada, M.; Seki, Y.; Kuroda, Y.; Fukuda, M.; Honda, E.; Suzuki, S.; Nakamura, S. Genetic analysis of the S1 gene of 4/91 type infectious bronchitis virus isolated in Japan. J. Vet. Med. Sci. 2009, 71, 583–588. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Han, Z.; Jiang, L.; Sun, J.; Zhao, Y.; Liu, S. Genetic diversity of avian infectious bronchitis virus in China in recent years. Infect. Genet. Evol. 2018, 66, 82–94. [Google Scholar] [CrossRef] [PubMed]
- De Wit, J.J. Detection of infectious bronchitis virus. Avian Pathol. 2000, 29, 71–93. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alexander, D.; Gough, R. Isolation of avian infectious bronchitis virus from experimentally infected chickens. Res. Vet. Sci. 1977, 23, 344–347. [Google Scholar] [CrossRef]
- Alexander, D.; Gough, R. A long-term study of the pathogenesis of infection of fowls with three strains of avian infectious bronchitis virus. Res. Vet. Sci. 1978, 24, 228–233. [Google Scholar] [CrossRef]
- Cavanagh, D.; Picault, J.P.; Gough, R.E.; Hess, M.; Mawditt, K.; Britton, P. Variation in the spike protein of the 793/B type of infectious bronchitis virus, in the field and during alternate passage in chickens and embryonated eggs. Avian Pathol. 2005, 34, 20–25. [Google Scholar] [CrossRef]
- SAG Informe Sanidad Animal año 2012. Available online: https://www.sag.gob.cl/sites/default/files/situacion_sanitaria_animal_2012_0.pdf (accessed on 1 June 2019).
- Moore, K.M.; Jackwood, M.W.; Hilt, D.A. Identification of amino acids involved in a serotype and neutralization specific epitope within the s1 subunit of avian infectious bronchitis virus. Arch. Virol. 1997, 142, 2249–2256. [Google Scholar] [CrossRef]
- Fellahi, S.; El Harrak, M.; Kuhn, J.H.; Sebbar, G.; Bouaiti, E.A.; Khataby, K.; Fihri, O.F.; El Houadfi, M.; Ennaji, M.M. Comparison of SYBR green i real-time RT-PCR with conventional agarose gel-based RT-PCR for the diagnosis of infectious bronchitis virus infection in chickens in Morocco. BMC Res. Notes 2016, 9, 231. [Google Scholar] [CrossRef]
- Yu, L.; Jiang, Y.; Low, S.; Wang, Z.; Nam, S.J.; Liu, W.; Kwang, J. Characterization of Three Infectious Bronchitis Virus Isolates from China Associated with Proventriculus in Vaccinated Chickens. Avian Dis. 2001, 45, 416–424. [Google Scholar] [CrossRef]
- de Wit, J.J.; Malo, A.; Cook, J.K. Induction of IBV strain-specific neutralizing antibodies and broad spectrum protection in layer pullets primed with IBV Massachusetts (Mass) and 793B vaccines prior to injection of inactivated vaccine containing Mass antigen. Avian Pathol. 2019, 48, 135–147. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Zhang, X.; Wang, Y.; Li, C.; Han, Z.; Shao, Y.; Li, H.; Kong, X. Molecular characterization and pathogenicity of infectious bronchitis coronaviruses: Complicated evolution and epidemiology in China caused by cocirculation of multiple types of infectious bronchitis coronaviruses. Intervirology 2009, 52, 223–234. [Google Scholar] [CrossRef] [PubMed]
- Vaezirad, M.M.; Koene, M.G.; Wagenaar, J.A.; van Putten, J.P.M. Chicken immune response following in ovo delivery of bacterial flagellin. Vaccine 2018, 36, 2139–2146. [Google Scholar] [CrossRef] [PubMed]
- Channappanavar, R.; Zhao, J.; Perlman, S. T cell-mediated immune response to respiratory coronaviruses. Immunol. Res. 2014, 59, 118–128. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Awad, F.; Forrester, A.; Baylis, M.; Lemiere, S.; Jones, R.; Ganapathy, K. Immune responses and interactions following simultaneous application of live Newcastle disease, infectious bronchitis and avian metapneumovirus vaccines in specific-pathogen-free chicks. Res. Vet. Sci. 2015, 98, 127–133. [Google Scholar] [CrossRef] [PubMed]
- De Wit, J.J.S.; Cook, J.K.A.; van der Heijden, H.M.J.F. Infectious bronchitis virus variants: A review of the history, current situation and control measures. Avian Pathol. 2011, 40, 223–235. [Google Scholar] [CrossRef] [PubMed]
- Oade, M.; Keep, S.; Freimanis, G.; Orton, R.; Britton, P.; Hammond, J.; Bickerton, E. Attenuation of infectious bronchitis virus in eggs results in different patterns of genomic variation across multiple replicates. J. Virol. 2019. [Google Scholar] [CrossRef] [PubMed]
- McKinley, E.T.; Jackwood, M.W.; Hilt, D.A.; Kissinger, J.C.; Robertson, J.S.; Lemke, C.; Paterson, A.H. Attenuated live vaccine usage affects accurate measures of virus diversity and mutation rates in avian coronavirus infectious bronchitis virus. Virus Res. 2011, 158, 225–234. [Google Scholar] [CrossRef] [PubMed]
- Durães-Carvalho, R.; Caserta, L.C.; Barnabé, A.C.S.; Martini, M.C.; Simas, P.V.M.; Santos, M.M.B.; Salemi, M.; Arns, C.W. Phylogenetic and phylogeographic mapping of the avian coronavirus spike protein-encoding gene in wild and synanthropic birds. Virus Res. 2015, 201, 101–112. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Han, Z.; Xu, Q.; Wang, Q.; Gao, M.; Wu, W.; Shao, Y.; Li, H.; Kong, X.; Liu, S. Serotype shift of a 793/B genotype infectious bronchitis coronavirus by natural recombination. Infect. Genet. Evol. 2015, 32, 377–387. [Google Scholar] [CrossRef]
- Abozeid, H.H.; Paldurai, A.; Khattar, S.K.; Afifi, M.A.; El-Kady, M.F.; El-Deeb, A.H.; Samal, S.K. Complete genome sequences of two avian infectious bronchitis viruses isolated in Egypt: Evidence for genetic drift and genetic recombination in the circulating viruses. Infect. Genet. Evol. 2017, 53, 7–14. [Google Scholar] [CrossRef] [PubMed]


| Strain | OD Values | S/P Ratios a |
|---|---|---|
| Negative control | 0.044 | |
| Negative control | 0.046 | |
| Positive control | 0.225 | |
| Positive control | 0.227 | |
| Mass | 0.192 | 0.875 |
| 0.125 | 0.476 | |
| 0.132 | 0.518 | |
| 0.155 | 0.655 | |
| 0.137 | 0.548 | |
| 4/91 | 0.100 | 0.306 |
| 0.103 | 0.324 | |
| 0.106 | 0.335 | |
| 0.89 | 0.243 | |
| 0.85 | 0.220 | |
| 12101SP-09 | 0.148 | 0.598 |
| 0.211 | 0.982 | |
| 0.197 | 0.880 | |
| 0.237 | 1.14 | |
| 0.256 | 1.256 | |
| 13347SP-15 | 0.194 | 0.864 |
| 0.221 | 1.023 | |
| 0.192 | 0.875 | |
| 0.236 | 1.112 | |
| 0.205 | 0.929 | |
| 13885R-17 | 0.144 | 0.573 |
| 0.148 | 0.598 | |
| 0.208 | 0.963 | |
| 0.220 | 1.037 | |
| 0.288 | 1.451 |
| Virus Strain | Antiserum | ||||
|---|---|---|---|---|---|
| Mass | 4/91 | 12101SP-09 | 13347SP-15 | 13885R-17 | |
| Mass | 100 | 32 | 41 | 42 | 38 |
| 4/91 | - | 100 | 34 | 33 | 73 |
| 12101SP-09 | - | - | 100 | 53 | 62 |
| 13347SP-15 | - | - | - | 100 | 57 |
| 13885R-17 | - | - | - | - | 100 |
| Virus Challenged | Clinical Signs and Lesions at Necropsy a | |||
|---|---|---|---|---|
| Inflamed Infraorbital Sinuses | Tracheal Rales | Tracheal Exudate | Palatine Exudate | |
| 12101SP-09 | ++ | +++ | +++ | +++ |
| 13347SP-15 | ++ | +++ | +++ | +++ |
| 13885R-17 | - | - | + | + |
| Strain | Ct |
|---|---|
| 12101SP-09 | 32.53 |
| 25 | |
| 28.68 | |
| 29.19 | |
| 33.45 | |
| 31.96 | |
| 30.4 | |
| 31.85 | |
| 35.61 | |
| 31.01 | |
| 26.47 | |
| 29.83 | |
| 29.77 | |
| 27.18 | |
| 32.53 | |
| 13347SP-15 | 29.63 |
| 30.11 | |
| 30.06 | |
| 30.85 | |
| 31.66 | |
| 26.32 | |
| 31.77 | |
| 31.19 | |
| 34.43 | |
| 32.13 | |
| 29.92 | |
| 28.26 | |
| 30.39 | |
| 29.63 | |
| 13885R-17 | 36.59 |
| 34.34 | |
| 32.07 | |
| 32.36 | |
| 32.67 | |
| 32.77 | |
| 33.48 | |
| 3467 | |
| 32.58 | |
| 33.82 | |
| 34.58 | |
| 34.99 | |
| 32.02 | |
| 34.73 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guzmán, M.; Sáenz, L.; Hidalgo, H. Molecular and Antigenic Characterization of GI-13 and GI-16 Avian Infectious Bronchitis Virus Isolated in Chile from 2009 to 2017 Regarding 4/91 Vaccine Introduction. Animals 2019, 9, 656. https://doi.org/10.3390/ani9090656
Guzmán M, Sáenz L, Hidalgo H. Molecular and Antigenic Characterization of GI-13 and GI-16 Avian Infectious Bronchitis Virus Isolated in Chile from 2009 to 2017 Regarding 4/91 Vaccine Introduction. Animals. 2019; 9(9):656. https://doi.org/10.3390/ani9090656
Chicago/Turabian StyleGuzmán, Miguel, Leonardo Sáenz, and Héctor Hidalgo. 2019. "Molecular and Antigenic Characterization of GI-13 and GI-16 Avian Infectious Bronchitis Virus Isolated in Chile from 2009 to 2017 Regarding 4/91 Vaccine Introduction" Animals 9, no. 9: 656. https://doi.org/10.3390/ani9090656
APA StyleGuzmán, M., Sáenz, L., & Hidalgo, H. (2019). Molecular and Antigenic Characterization of GI-13 and GI-16 Avian Infectious Bronchitis Virus Isolated in Chile from 2009 to 2017 Regarding 4/91 Vaccine Introduction. Animals, 9(9), 656. https://doi.org/10.3390/ani9090656





