Genetic Variability and Management in Nero di Parma Swine Breed to Preserve Local Diversity
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Avaiable, Quality Control, and Reference Populations
2.2. Genetic Variability and Population Structure
3. Results
3.1. Data Avaiable, Quality Control, and Reference Populations
3.2. Genetic Variability and Population Structure
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Muñoz, M.; Bozzi, R.; García-Casco, J.; Núñez, Y.; Ribani, A.; Franci, O.; García, F.; Škrlep, M.; Schiavo, G.; Bovo, S.; et al. Genomic diversity, linkage disequilibrium and selection signatures in European local pig breeds assessed with a high density SNP chip. Sci. Rep. 2019, 9, 13546. [Google Scholar] [CrossRef] [PubMed]
- Sabbioni, A.; Beretti, V.; Paini, V.; Superchi, P. Occurrence and inheritance of wattles and effects of genotype at wattle locus on Gompertz growth curve parameters in “Nero di Parma” pigs. Livest. Sci. 2011, 137, 226–230. [Google Scholar] [CrossRef]
- Rozzi, U. L’allevamento suino in provincia di Parma; Tipografia Fratelli Godi, Ed.; Godi: Parma, Italy, 1934. [Google Scholar]
- Ablondi, M.; Vasini, M.; Beretti, V.; Superchi, P.; Sabbioni, A. Exploring genetic diversity in an Italian horse native breed to develop strategies for preservation and management. J. Anim. Breed. Genet. 2018, 135, 450–459. [Google Scholar] [CrossRef] [PubMed]
- Fabbri, M.C.; de Rezende, M.P.G.; Dadousis, C.; Biffani, S.; Negrini, R.; Carneiro, P.L.S.; Bozzi, R. Population structure and genetic diversity of Italian beef breeds as a tool for planning conservation and selection strategies. Animals 2019, 9, 880. [Google Scholar] [CrossRef]
- Hammami, H.; Croquet, C.; Stoll, J.; Rekik, B.; Gengler, N. Genetic Diversity and Joint-Pedigree Analysis of Two Importing Holstein Populations. J. Dairy Sci. 2007, 90, 3530–3541. [Google Scholar] [CrossRef]
- Caballero, A.; Toro, M.A. Interrelations between effective population size and other pedigree tools for the management of conserved populations. Genet. Res. 2000, 75, 331–343. [Google Scholar] [CrossRef]
- Hill, W.G.; Mackay, T.F.C. D. S. Falconer and introduction to quantitative genetics. Genetics 2004, 167, 1529–1536. [Google Scholar]
- Maignel, L.; Boichard, D.; Verrier, E. Genetic variability of French dairy breeds estimated from pedigree information. Interbull Bull. 1996, 14, 49–53. [Google Scholar]
- Lacy, R.C. Analysis of founder representation in pedigrees: Founder equivalents and founder genome equivalents. Zoo Biol. 1989, 8, 111–123. [Google Scholar] [CrossRef]
- Boichard, D.; Maignel, L.; Verrier, É. The value of using probabilities of gene origin to measure genetic variability in a population. Genet. Sel. Evol. 1997, 29, 5. [Google Scholar] [CrossRef]
- Paixão, G.; Esteves, A.; Payan-Carreira, R.; Carolino, N. Demographic structure and genetic diversity of the endangered Bísaro pig: Evolution and current status. Czech J. Anim. Sci. 2018, 63, 452–461. [Google Scholar] [CrossRef]
- Meuwissen, T.; Luo, Z. Computing inbreeding coefficients in large populations. Genet. Sel. Evol. 1992, 24, 305. [Google Scholar] [CrossRef]
- Gutiérrez, J.P.; Cervantes, I.; Molina, A.; Valera, M.; Goyache, F. Individual increase in inbreeding allows estimating effective sizes from pedigrees. Genet. Sel. Evol. 2008, 40, 359–378. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, J.P.; Cervantes, I.; Goyache, F. Improving the estimation of realized effective population sizes in farm animals. J. Anim. Breed. Genet. 2009, 126, 327–332. [Google Scholar] [CrossRef] [PubMed]
- Cervantes, I.; Goyache, F.; Molina, A.; Valera, M.; Gutiérrez, J.P. Application of individual increase in inbreeding to estimate realized effective sizes from real pedigrees. J. Anim. Breed. Genet. 2008, 125, 301–310. [Google Scholar] [CrossRef]
- Cervantes, I.; Gutiérrez, J.P.; Meuwissen, T.H.E. Response to selection while maximizing genetic variance in small populations. Genet. Sel. Evol. 2016, 48, 69. [Google Scholar] [CrossRef]
- Gutierrez, J.P.; Goyache, F. A note on ENDOG: A computer program for analysing pedigree information. J. Anim. Breed. Genet. 2005, 122, 172–176. [Google Scholar] [CrossRef]
- Wright, S. Evolution and the Genetics of Populations. Volume 4: Variability within and among Natural Populations; University of Chicago Press: Chicago, IL, USA, 1978. [Google Scholar]
- R Development Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018. [Google Scholar]
- FAO. The State of the World’s Biodiversity for Food and Agriculture; FAO: Rome, Italy, 2019; ISBN 9789251312704. [Google Scholar]
- Candek-Potokar, M.; Nieto Linan, R. (Eds.) European Local Pig Breeds—Diversity and Performance. A Study of project TREASURE; IntechOpen: Rijeka, Croatia, 2019; ISBN 978-1-78985-407-7. [Google Scholar]
- Gandini, G.; Orlandini, P.; Franci, O.; Campodoni, G. Policies to decrease inbreeding in the Cinta Senese pig. CIHEAM Tradit. Innov. Mediterr. Pig Prod. 2000, 35–37. [Google Scholar]
- Commission on Genetic Resources for Food and Agriculture Food and Agriculture Organization of the United Nations. The Second Report on the State of the World’s Animal Genetic Resources for Food and Agriculture; FAO: Rome, Italy, 2007; ISBN 978-92-5-108820-3. [Google Scholar]
- Tang, G.Q.; Xuea, J.; Lian, M.J.; Yang, R.F.; Liu, T.F.; Zeng, Z.Y.; Jiang, A.A.; Jiang, Y.Z.; Zhu, L.; Bai, L.; et al. Inbreeding and genetic diversity in three imported swine breeds in China using pedigree data. Asian-Australas. J. Anim. Sci. 2013, 26, 755–765. [Google Scholar] [CrossRef]
- Krupa, E.; Žáková, E.; Krupová, Z. Evaluation of inbreeding and genetic variability of five pig breeds in Czech Republic. Asian-Australas. J. Anim. Sci. 2015, 28, 25–36. [Google Scholar] [CrossRef]
- Melka, M.G.; Schenkel, F. Analysis of genetic diversity in four Canadian swine breeds using pedigree data. Can. J. Anim. Sci. 2010, 90, 331–340. [Google Scholar] [CrossRef]
- Čandek-Potokar, M.; Batorek Lukač, N.; Tomažin, U.; Škrlep, M.; Nieto, R. Analytical Review of Productive Performance of Local Pig Breeds. In European Local Pig Breeds—Diversity and Performance. A Study of Project TREASURE; IntechOpen: Rijeka, Croatia, 2019. [Google Scholar]
- Franci, O.; Pugliese, C. Italian autochthonous pigs: Progress report and research perspectives. Ital. J. Anim. Sci. 2007, 6, 663–671. [Google Scholar] [CrossRef]
- Álvarez, I.; Royo, L.J.; Pérez-Pardal, L.; Fernández, I.; Payeras, L.; Goyache, F. Assessing losses of genetic variability in the endangered Mallorquí horse. Czech J. Anim. Sci. 2010, 55, 456–462. [Google Scholar]
- Álvarez, I.; Royo, L.J.; Gutiérrez, J.P.; Fernández, I.; Arranz, J.J.; Goyache, F. Relationship between genealogical and microsatellite information characterizing losses of genetic variability: Empirical evidence from the rare Xalda sheep breed. Livest. Sci. 2008, 115, 80–88. [Google Scholar] [CrossRef]
- Menendez, J.; Alvarez, I.; Fernandez, I.; Goyache, F. Genealogical analysis of the Gochu Asturcelta pig breed: Insights for conservation. Czech J. Anim. Sci. 2016, 61, 140–149. [Google Scholar] [CrossRef]
- Biermann, A.D.M.; Pimentel, E.C.G.; Tietze, M.; Pinent, T.; König, S. Implementation of genetic evaluation and mating designs for the endangered local pig breed “Bunte Bentheimer”. J. Anim. Breed. Genet. 2014, 131, 36–45. [Google Scholar] [CrossRef]
- Welsh, C.S.; Stewart, T.S.; Schwab, C.; Blackburn, H.D. Pedigree analysis of 5 swine breeds in the United States and the implications for genetic conservation. J. Anim. Sci. 2010, 88, 1610–1618. [Google Scholar] [CrossRef]
- Fernandes, S.; Malovrh, Š.; Kovac, M.; Cadavez, V. Study of genetic diversity of Bisaro pigs breed by pedigree analysis. Lucr. Ştiinţifice. Ser. Zooteh. 2010, 326–330. [Google Scholar]
- Fontanesi, L. Genetic authentication and traceability of food products of animal origin: New developments and perspectives. Ital. J. Anim. Sci. 2009, 8, 9–18. [Google Scholar] [CrossRef]
- Russo, V.; Fontanesi, L.; Scotti, E.; Tazzoli, M.; Dall’Olio, S.; Davoli, R. Analysis of melanocortin 1 receptor (MC1R) gene polymorphisms in some cattle breeds: Their usefulness and application for breed traceability and authentication of Parmigiano Reggiano cheese. Ital. J. Anim. Sci. 2007, 6, 257–272. [Google Scholar] [CrossRef]
- D’Alessandro, E.; Fontanesi, L.; Liotta, L.; Davoli, R.; Chiofalo, V.; Russo, V. Analysis of the MC1R gene in the nero siciliano pig breed and usefulness of this locus for breed traceability. Vet. Res. Commun. 2007, 31, 389–392. [Google Scholar] [CrossRef] [PubMed]
- Fontanesi, L.; Bozzi, R.; Tazzoli, M.; Crovetti, A.; Davoli, R.; Franci, O.; Russo, V. Genetic Characterization of Cinta Senese Pig Breed: Analysis of Polymorphisms in Four Genes Affecting Performance and Phenotypic Traits. In Proceedings of the Role of Biotechnology, Turin, Italy, 5–7 March 2005; pp. 173–174. [Google Scholar]

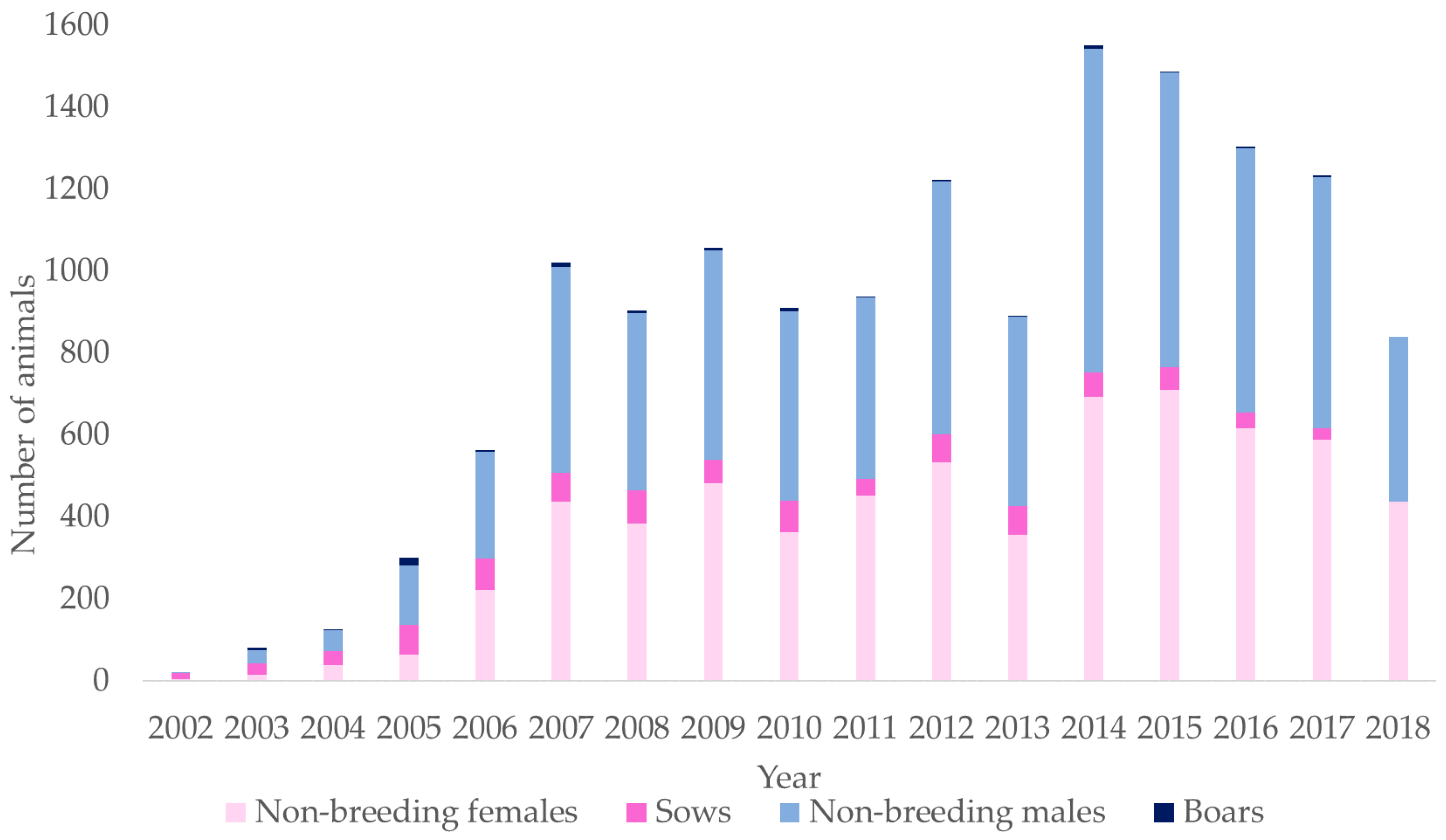
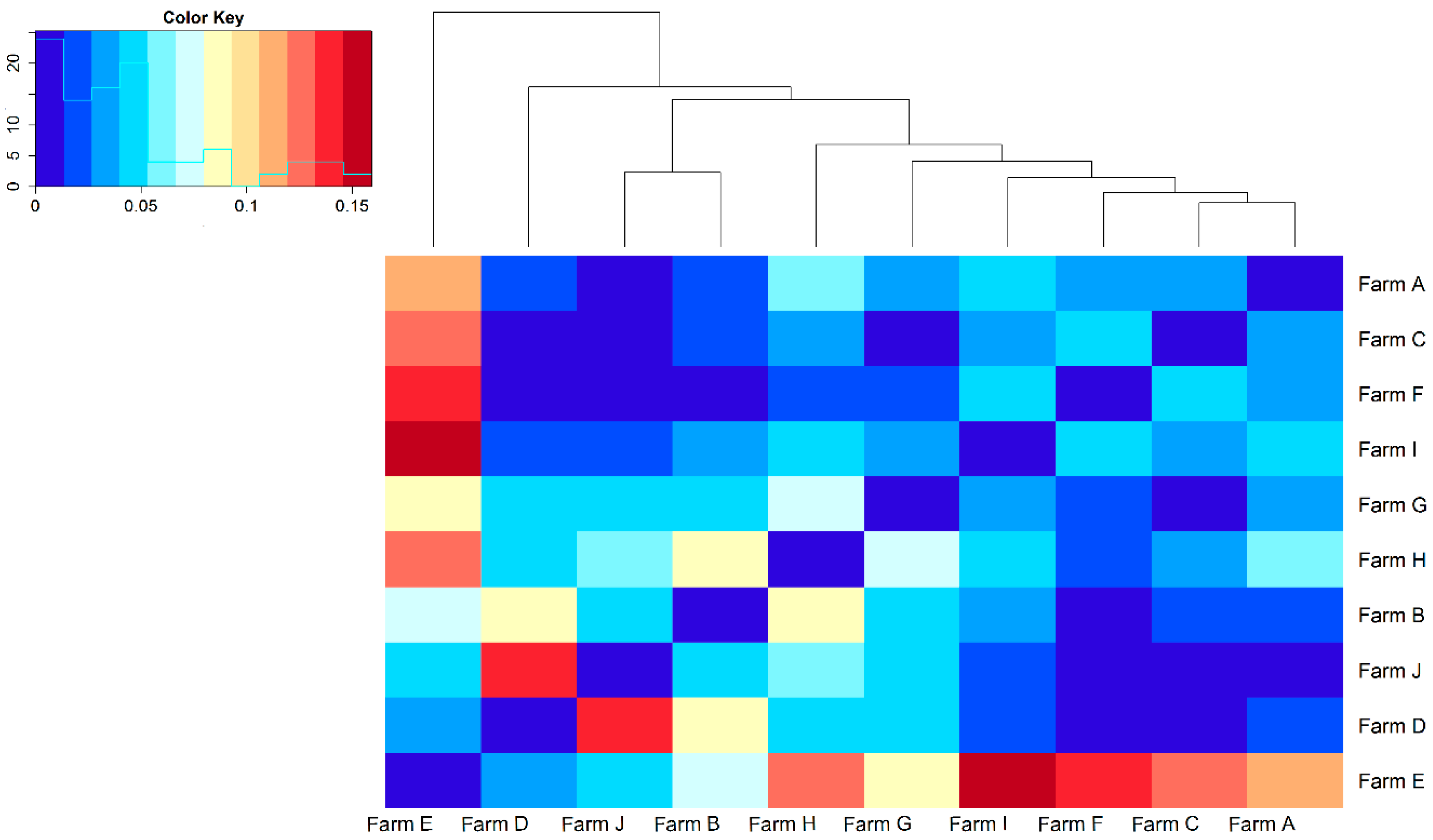
| Parameters | TP 1 | RP 2 | BP 3 |
|---|---|---|---|
| Numbers of pigs | 14,485 | 1603 | 336 |
| Numbers of males | 7197 | 641 | 21 |
| Numbers of females | 7288 | 962 | 315 |
| Numbers of pigs with no progeny | 13,507 | 1267 | 0 |
| Pathway | N1 | GI | SD 2 |
|---|---|---|---|
| Father-Son | 96 | 2.65 | 1.71 |
| Father-Daughter | 876 | 2.69 | 1.32 |
| Mother-Son | 94 | 2.26 | 1.08 |
| Mother-Daughter | 870 | 2.24 | 0.98 |
| Total | 1936 | 2.46 | 1.20 |
| Genealogical Parameter | TP | RP | BP |
|---|---|---|---|
| Population size | 14,485 | 1,603 | 336 |
| Number of Founders (f) | 32.0 | 19.0 | 11.0 |
| Number of Ancestors | 16.0 | 13.0 | 11.0 |
| Equivalent complete generations (CGE) | 6.47 | 7.94 | 7.22 |
| Average inbreeding (F) | 0.28 | 0.32 | 0.27 |
| Average Relationship (AR) | 0.50 | 0.51 | 0.50 |
| Effective Number of Founders (fe) | 3.42 | 3.00 | 3.00 |
| Effective Number of Ancestors (fa) | 3.00 | 3.00 | 3.00 |
| Founder genome equivalents (fg) | 2.01 | 1.81 | 1.91 |
| Effective Number of non-founders (nfe) | 4.88 | 4.58 | 5.29 |
| fe/f | 0.11 | 0.16 | 0.27 |
| fa/fe | 0.88 | 1.00 | 1.00 |
| fg/fe | 0.59 | 0.60 | 0.64 |
| Individual increase in inbreeding (ΔFi) | 0.07 | 0.06 | 0.05 |
| Realized effective population size (Ne) | 7.68 | 8.72 | 9.66 |
| Birth Year Cohort | Total Population (TP) | Breeding Animal within TP | ||||||
|---|---|---|---|---|---|---|---|---|
| N1 | F 2 | SD 3 | Highest F | N | F | SD | Highest F | |
| 2001–2003 | 130 | 0.18 | 0.16 | 0.50 | 72 | 0.18 | 0.16 | 0.59 |
| 2004–2006 | 991 | 0.31 | 0.13 | 0.61 | 208 | 0.29 | 0.13 | 0.62 |
| 2007–2009 * | 2983 | 0.29 | 0.12 | 0.66 | 233 | 0.28 | 0.15 | 0.66 |
| 2010–2012 | 3072 | 0.28 | 0.10 | 0.61 | 200 | 0.28 | 0.10 | 0.47 |
| 2013–2015 | 3930 | 0.26 | 0.07 | 0.53 | 192 | 0.27 | 0.10 | 0.53 |
| 2016–2018 ** | 3379 | 0.31 | 0.10 | 0.62 | 73 | 0.29 | 0.09 | 0.55 |
| Grand Total | 14,485 | 0.28 | 0.10 | 0.66 | 978 | 0.27 | 0.13 | 0.66 |
| Herd Class 1 | Farms 2 | N3 | CGE | Own Father (%) 4 | F | AR | ||||
|---|---|---|---|---|---|---|---|---|---|---|
| Mean | SD 5 | Highest 6 | Mean | SD 5 | Highest 6 | |||||
| 1 | 11 | 15.0 | 6.6 | 9.31 | 0.32 | 0.15 | 0.57 | 0.50 | 0.10 | 0.61 |
| 2 | 11 | 42 | 5.9 | 11.69 | 0.30 | 0.10 | 0.58 | 0.52 | 0.05 | 0.64 |
| 3 | 11 | 130 | 6.7 | 11.75 | 0.31 | 0.11 | 0.61 | 0.49 | 0.07 | 0.60 |
| 4 | 11 | 1124 | 6.5 | 66.70 | 0.28 | 0.10 | 0.66 | 0.50 | 0.06 | 0.63 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mariani, E.; Summer, A.; Ablondi, M.; Sabbioni, A. Genetic Variability and Management in Nero di Parma Swine Breed to Preserve Local Diversity. Animals 2020, 10, 538. https://doi.org/10.3390/ani10030538
Mariani E, Summer A, Ablondi M, Sabbioni A. Genetic Variability and Management in Nero di Parma Swine Breed to Preserve Local Diversity. Animals. 2020; 10(3):538. https://doi.org/10.3390/ani10030538
Chicago/Turabian StyleMariani, Elena, Andrea Summer, Michela Ablondi, and Alberto Sabbioni. 2020. "Genetic Variability and Management in Nero di Parma Swine Breed to Preserve Local Diversity" Animals 10, no. 3: 538. https://doi.org/10.3390/ani10030538
APA StyleMariani, E., Summer, A., Ablondi, M., & Sabbioni, A. (2020). Genetic Variability and Management in Nero di Parma Swine Breed to Preserve Local Diversity. Animals, 10(3), 538. https://doi.org/10.3390/ani10030538






