Clustering of Bacterial Growth Dynamics in Response to Growth Media by Dynamic Time Warping
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Growth Data
2.2. Growth Data Processing
2.3. Similarity Comparison of the Growth Curves
2.4. Hierarchical Clustering of the Growth Curves
2.5. Evaluation of the Goodness of Clustering
3. Results
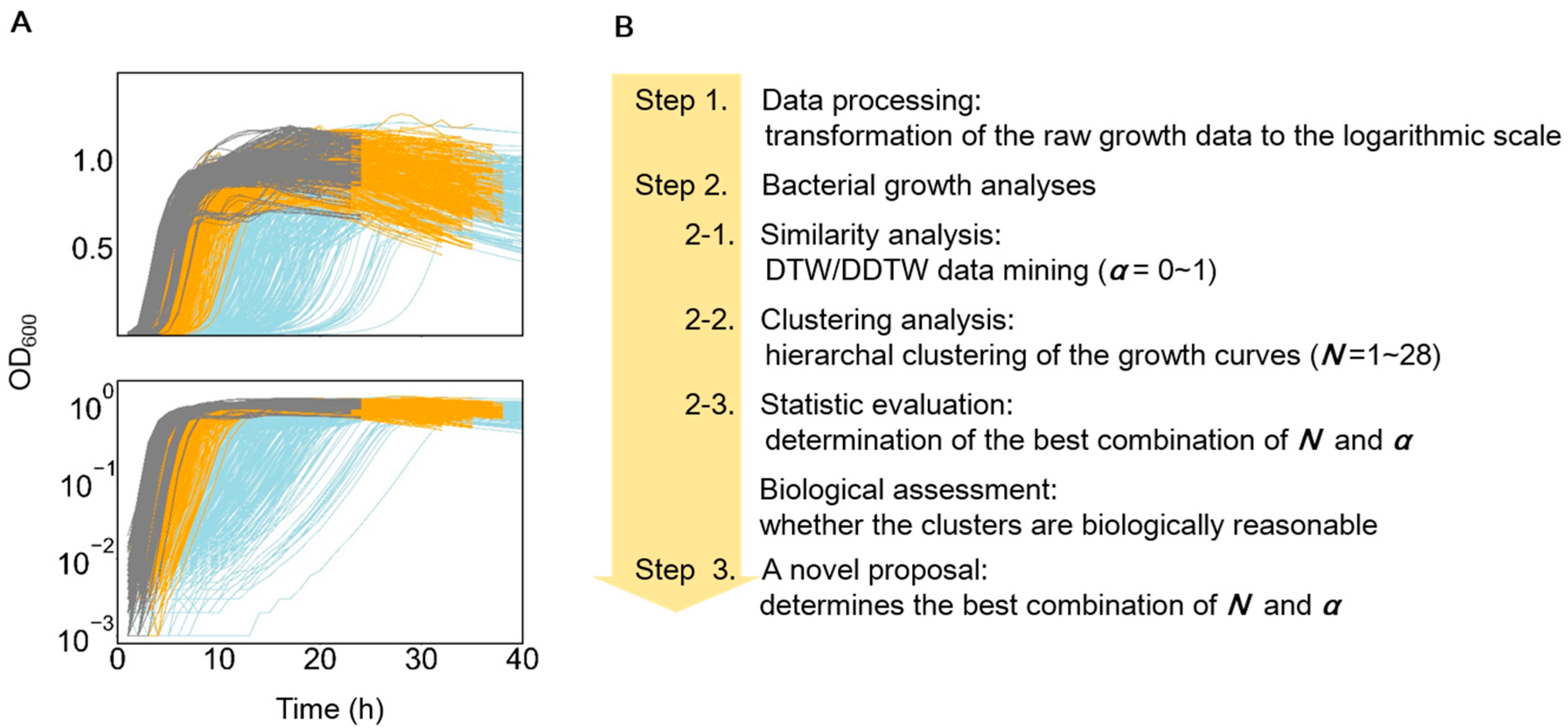
3.1. Features of the Growth Curves and Analytical Approaches
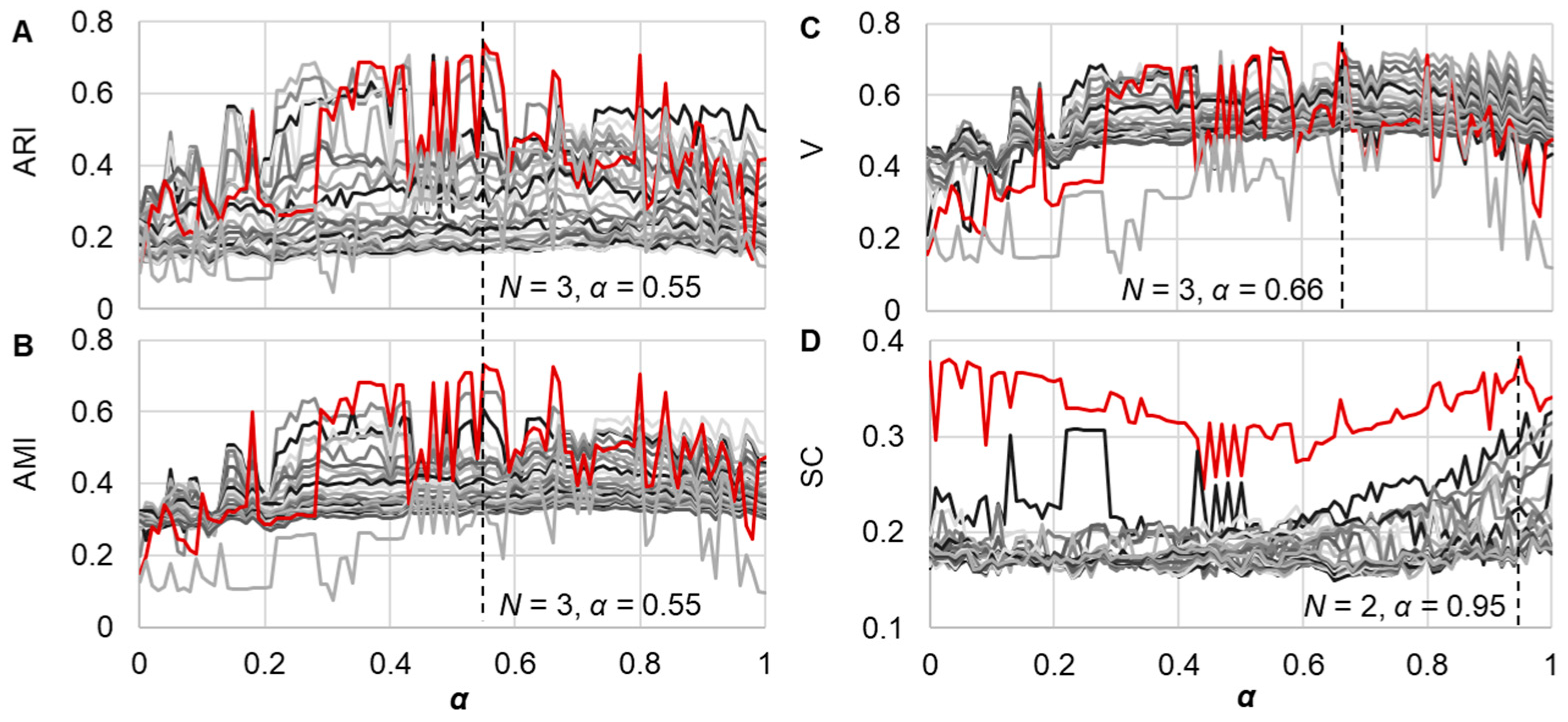
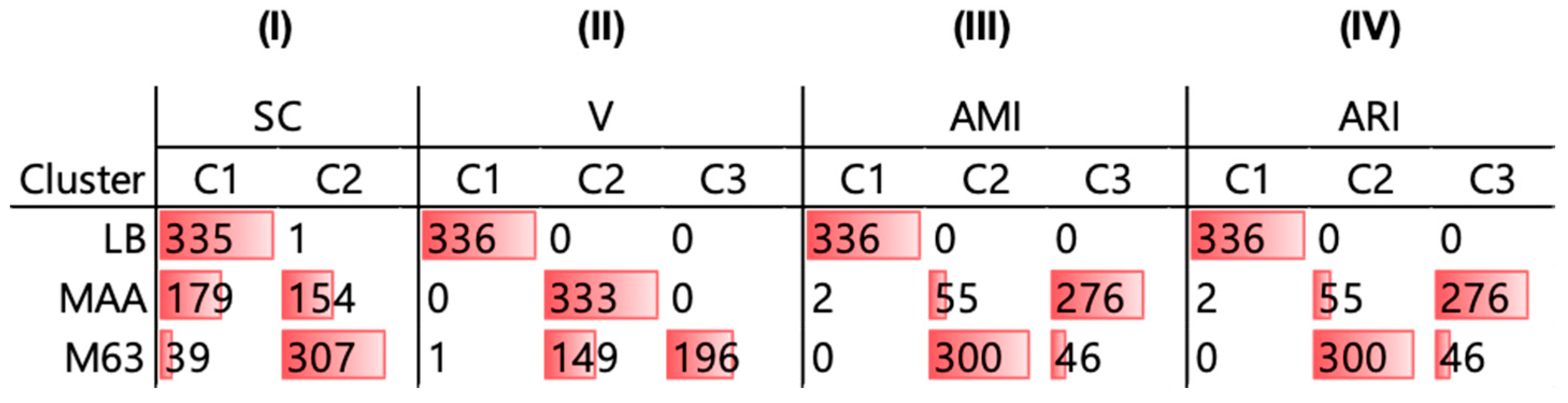
3.2. Similarity Evaluation and Hierarchical Clustering
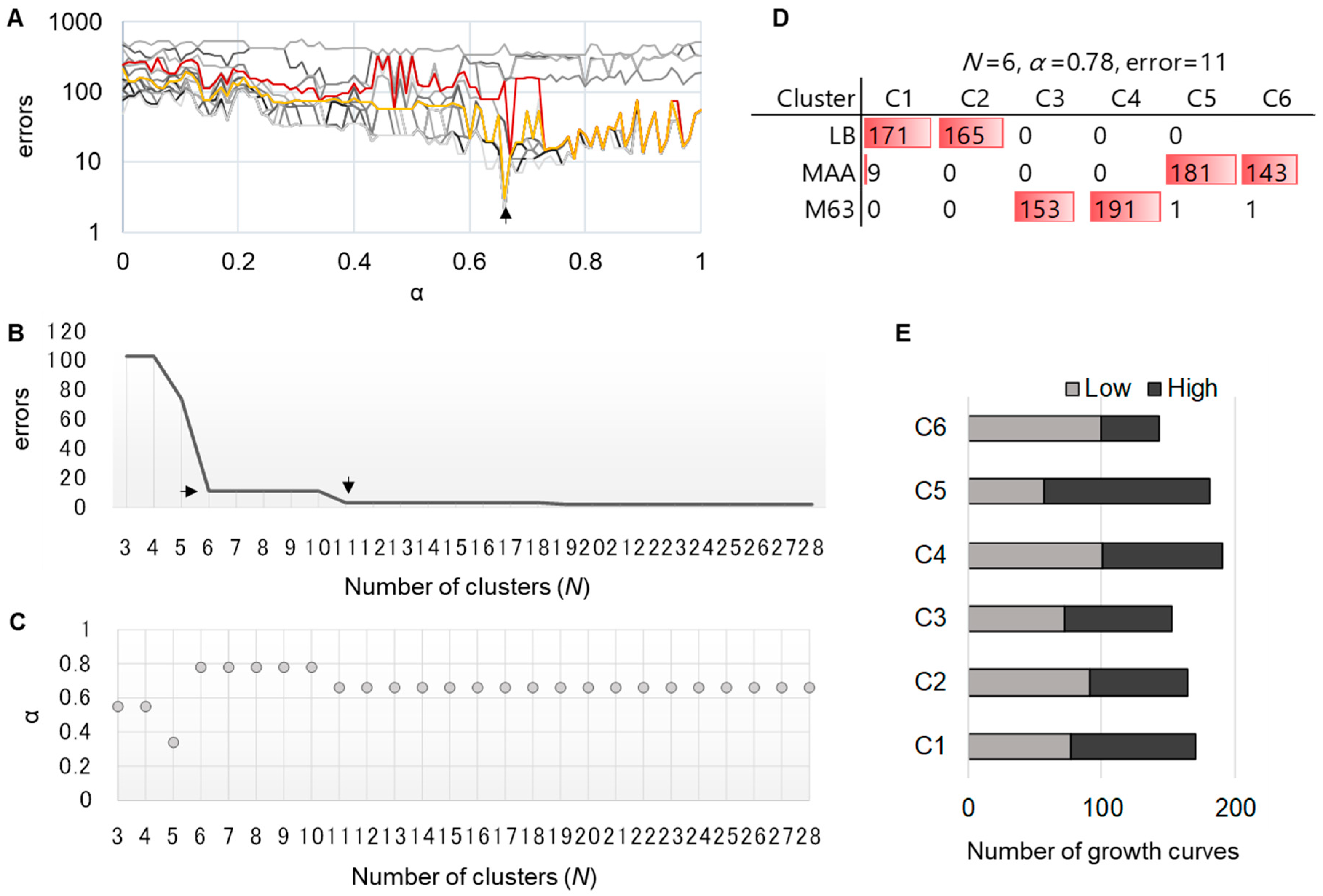
3.3. A novel Algorithm for Improved Clustering
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Swinnen, I. Predictive modelling of the microbial lag phase: A review. Int. J. Food Microbiol. 2004, 94, 137–159. [Google Scholar] [CrossRef]
- Maier, R.M.; Pepper, I.L. Bacterial Growth. In Environmental Microbiology, 3rd ed.; Academic Press: London, UK, 2015; pp. 37–56. [Google Scholar]
- Gibson, A.M.; Bratchell, N.; Roberts, T.A. Predicting microbial growth: Growth responses of salmonellae in a laboratory medium as affected by pH, sodium chloride and storage temperature. Int. J. Food Microbiol. 1988, 6, 155–178. [Google Scholar] [CrossRef]
- Baranyi, J.; Roberts, T.A. Mathematics of predictive food microbiology. Int. J. Food Microbiol. 1995, 26, 199–218. [Google Scholar] [CrossRef]
- Ponciano, J.M.; Vandecasteele, F.P.J.; Hess, T.F.; Forney, L.J.; Crawford, R.L.; Joyce, P. Use of Stochastic Models To Assess the Effect of Environmental Factors on Microbial Growth. Appl. Environ. Microbiol. 2005, 71, 2355–2364. [Google Scholar] [CrossRef] [PubMed]
- An, H.B.; Li, F.; Zhao, M.L.; Liu, Y.J. Optimized Spectral Indices Based Estimation of Forage Grass Biomass. Guang Pu Xue Yu Guang Pu Fen Xi 2015, 35, 3155–3160. [Google Scholar] [PubMed]
- López, S.; Prieto, M.; Dijkstra, J.; Dhanoa, M.S.; France, J. Statistical evaluation of mathematical models for microbial growth. Int. J. Food Microbiol. 2004, 96, 289–300. [Google Scholar] [CrossRef] [PubMed]
- Peleg, M.; Corradini, M.G. Microbial growth curves: What the models tell us and what they cannot. Crit. Rev. Food Sci. Nutr. 2011, 51, 917–945. [Google Scholar] [CrossRef] [PubMed]
- Zwietering, M.H.; Jongenburger, I.; Rombouts, F.M.; Van’t Riet, K. Modeling of the bacterial growth curve. Appl. Environ. Microbiol. 1990, 56, 1875–1881. [Google Scholar] [CrossRef]
- Merks, R.M.H.; Tjørve, K.M.C.; Tjørve, E. The use of Gompertz models in growth analyses, and new Gompertz-model approach: An addition to the Unified-Richards family. PLoS ONE 2017, 12, e0178691. [Google Scholar]
- Yates, G.T.; Smotzer, T. On the lag phase and initial decline of microbial growth curves. J. Theor. Biol. 2007, 244, 511–517. [Google Scholar] [CrossRef]
- Tsuchiya, K.; Cao, Y.-Y.; Kurokawa, M.; Ashino, K.; Yomo, T.; Ying, B.-W. A decay effect of the growth rate associated with genome reduction in Escherichia coli. BMC Microbiol. 2018, 18, 101. [Google Scholar] [CrossRef] [PubMed]
- Fujikawa, H.; Kai, A.; Morozumi, S. A new logistic model for Escherichia coli growth at constant and dynamic temperatures. Food Microbiol. 2004, 21, 501–509. [Google Scholar] [CrossRef]
- Kurokawa, M.; Seno, S.; Matsuda, H.; Ying, B.-W. Correlation between genome reduction and bacterial growth. DNA Res. 2016, 23, 517–525. [Google Scholar] [CrossRef] [PubMed]
- Kurokawa, M.; Ying, B.W.J.M. Experimental Challenges for Reduced Genomes: The Cell Model Escherichia coli. Microorganisms 2020, 8, 3. [Google Scholar] [CrossRef] [PubMed]
- Ashino, K.; Sugano, K.; Amagasa, T.; Ying, B.W. Predicting the decision making chemicals used for bacterial growth. Sci. Rep. 2019, 9, 7251. [Google Scholar] [CrossRef]
- Aghabozorgi, S.; Shirkhorshidi, A.S.; Wah, T.Y. Time-series clustering—A decade review. Inf. Syst. 2015, 53, 16–38. [Google Scholar] [CrossRef]
- Senin, P.J.I.; Computer Science Department University of Hawaii at Manoa Honolulu, U. Dynamic time warping algorithm review. Int. J. Res. Electr. Comput. Eng. 2008, 855, 40. [Google Scholar]
- Sakoe, H.; Chiba, S. Dynamic programming algorithm optimization for spoken word recognition. Inf. Comput. Sci. Dep. Univ. Hawaii Manoa Honol. 1978, 26, 43–49. [Google Scholar] [CrossRef]
- Górecki, T.; Łuczak, M. First and Second Derivatives in Time Series Classification Using DTW. Commun. Stat. Simul. Comput. 2014, 43, 2081–2092. [Google Scholar]
- Górecki, T.; Łuczak, M. Using derivatives in time series classification. Data Min. Knowl. Discov. 2012, 26, 310–331. [Google Scholar] [CrossRef]
- Keogh, E.J.; Pazzani, M.J. Derivative dynamic time warping. In Proceedings of the 2001 SIAM international conference on data mining, Chicago, IL, USA, 5–7 April 2001; pp. 1–11. [Google Scholar]
- Łuczak, M. Hierarchical clustering of time series data with parametric derivative dynamic time warping. Expert Syst. Appl. 2016, 62, 116–130. [Google Scholar] [CrossRef]
- Kaufman, L.; Rousseeuw, P.J. Finding Groups in Data: An Introduction to Cluster Analysis, 2nd ed.; John Wiley & Sons: New York, NY, USA, 2009; pp. 1–162. [Google Scholar]
- Rousseeuw, P.J. Silhouettes: A graphical aid to the interpretation and validation of cluster analysis. J. Comput. Appl. Math. 1987, 20, 53–65. [Google Scholar] [CrossRef]
- Vinh, N.X.; Epps, J.; Bailey, J. Information theoretic measures for clusterings comparison: Variants, properties, normalization and correction for chance. J. Mach. Learn. Res. 2010, 11, 2837–2854. [Google Scholar]
- Vinh, N.X.; Epps, J.; Bailey, J. Information Theoretic Measures for Clusterings Comparison: Is a Correction for Chance Necessary? In Proceedings of the 26th Annual International Conference on Machine Learning, Montreal, QC, Canada, 14–18 June 2009; pp. 1073–1080. [Google Scholar]
- Rosenberg, A.; Hirschberg, J. V-Measure: A Conditional Entropy-Based External Cluster Evaluation Measure. In Proceedings of the 2007 Joint Conference on Empirical Methods in Natural Language Processing and Computational Natural Language Learning (EMNLP-CoNLL), Prague, Czech Republic, 28–30 June 2007; pp. 410–420. [Google Scholar]
- Myers, J.A.; Curtis, B.S.; Curtis, W.R. Improving accuracy of cell and chromophore concentration measurements using optical density. BMC Biophys. 2013, 6, 4. [Google Scholar] [CrossRef] [PubMed]
- Mytilinaios, I.; Salih, M.; Schofield, H.K.; Lambert, R.J.W. Growth curve prediction from optical density data. Int. J. Food Microbiol. 2012, 154, 169–176. [Google Scholar] [CrossRef]
- Deza, M.M.; Deza, E. Encyclopedia of Distances. In Encyclopedia of Distances, 2nd ed.; Springer: Berlin/Heidelberg, Germany, 2009; pp. 1–583. [Google Scholar]
- Lee, R.J.; Nicewander, W.A. Thirteen Ways to Look at the Correlation Coefficient. Am. Stat. 1988, 42, 59–66. [Google Scholar]
- Huttenlocher, D.P.; Klanderman, G.A.; Rucklidge, W.J. Comparing images using the Hausdorff distance. IEEE Trans. Pattern Anal. Mach. Intell. 1993, 15, 850–863. [Google Scholar] [CrossRef]
- Rai, P.; Singh, S. A survey of clustering techniques. Int. J. Comput. Appl. 2010, 7, 1–5. [Google Scholar] [CrossRef]
- Aghabozorgi, S.; Teh, Y.W. Stock market co-movement assessment using a three-phase clustering method. Expert Syst. Appl. 2014, 41, 1301–1314. [Google Scholar] [CrossRef]
- Elangasinghe, M.A.; Singhal, N.; Dirks, K.N.; Salmond, J.A.; Samarasinghe, S. Complex time series analysis of PM10 and PM2.5 for a coastal site using artificial neural network modelling and k-means clustering. Atmos. Environ. 2014, 94, 106–116. [Google Scholar] [CrossRef]
- Ernst, J.; Nau, G.J.; Bar-Joseph, Z. Clustering short time series gene expression data. Bioinformatics 2005, 21, i159–i168. [Google Scholar] [CrossRef] [PubMed]
- Steinbach, M.; Tan, P.-N.; Kumar, V.; Klooster, S.; Potter, C. Discovery of climate indices using clustering. Proc. Proc. Ninth Acm Sigkdd Int. Conf. Knowl. Discov. Data Min. 2003, 3, 446–455. [Google Scholar]
- Wismüller, A.; Lange, O.; Dersch, D.R.; Leinsinger, G.L.; Hahn, K.; Pütz, B.; Auer, D. Cluster analysis of biomedical image time-series. Int. J. Comput. Vis. 2002, 46, 103–128. [Google Scholar] [CrossRef]
- Bishopric, N.H.; Pyatnitskiy, M.; Mazo, I.; Shkrob, M.; Schwartz, E.; Kotelnikova, E. Clustering Gene Expression Regulators: New Approach to Disease Subtyping. PLoS ONE 2014, 9, e84955. [Google Scholar]
- Möller-Levet, C.S.; Klawonn, F.; Cho, K.H.; Wolkenhauer, O. Fuzzy clustering of short time-series and unevenly distributed sampling points. In International Symposium on Intelligent Data Analysis; Springer: Berlin/Heidelberg, Germany, 2003; pp. 330–340. [Google Scholar]
- Subhani, N.; Rueda, L.; Ngom, A.; Burden, C.J. Multiple gene expression profile alignment for microarray time-series data clustering. Bioinformatics 2010, 26, 2281–2288. [Google Scholar] [CrossRef]
- Fujita, A.; Severino, P.; Kojima, K.; Sato, J.R.; Patriota, A.G.; Miyano, S. Functional clustering of time series gene expression data by Granger causality. BMC Syst. Biol. 2012, 6, 137. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cao, Y.-Y.; Yomo, T.; Ying, B.-W. Clustering of Bacterial Growth Dynamics in Response to Growth Media by Dynamic Time Warping. Microorganisms 2020, 8, 331. https://doi.org/10.3390/microorganisms8030331
Cao Y-Y, Yomo T, Ying B-W. Clustering of Bacterial Growth Dynamics in Response to Growth Media by Dynamic Time Warping. Microorganisms. 2020; 8(3):331. https://doi.org/10.3390/microorganisms8030331
Chicago/Turabian StyleCao, Yang-Yang, Tetsuya Yomo, and Bei-Wen Ying. 2020. "Clustering of Bacterial Growth Dynamics in Response to Growth Media by Dynamic Time Warping" Microorganisms 8, no. 3: 331. https://doi.org/10.3390/microorganisms8030331
APA StyleCao, Y.-Y., Yomo, T., & Ying, B.-W. (2020). Clustering of Bacterial Growth Dynamics in Response to Growth Media by Dynamic Time Warping. Microorganisms, 8(3), 331. https://doi.org/10.3390/microorganisms8030331




