1. Introduction
Endophytic fungi spend the whole or a part of their life cycle colonizing the internal tissues of host plants, without causing any visible symptoms [
1,
2]. Therefore, they have potential to produce a broad range of valuable products with a large spectrum of biological activities and promote the growth and resistance of plants against biotic and abiotic stresses including drought, salinity and phytopathogenic agents [
3,
4]. Among the metabolites required for industrial and pharmaceutical usages, enzymes are currently attracting much attention [
5]. There is an increasing demand for new resources of enzymes, including proteases, with various thermo-stabilities and pH profiles suited to different industrial requirements. Proteases are among the most important commercial enzymes used in food processing, silver recovery, diagnostics, detergent industry and pharmaceutical applications [
6]. These biological catalysts account for roughly 60% to 65% of the worldwide enzyme market, which allow them to be the leader of the industrial enzymes [
7]. Acid proteases among alkaline and/or neutral proteases are widely used in various food industry processes such as cheese and food processing, animal feed, and X-ray films [
8].
Currently, several attempts have been made in exploration of new resources of proteases that possess desired criteria for diverse biotechnological applications. Nowadays, microorganisms represent preferred protease resources due to their susceptibility to genetic manipulation and their bioengineering potentials [
9]. Moreover, microbial proteases are almost extracellular enzymes which imply that they are directly secreted into the fermentation medium without intervention of complicated extraction processes compared to plant- and animal-derived proteases [
10]. Among proteases belonging to microbes, those originating from fungi are the most recommended regarding their high productivity coupled with a low materials cost. Furthermore, fungi are regarded as GRAS (Generally Recognized As Safe) organisms, with the advantage of the safety manipulation of their products, including proteases [
11]. In recent years, great number of fungi has been used to produce proteases using different substrates and different approaches. Currently, the application of response surface methodology (RSM) for optimization has increased remarkably thanks to its advantages over the traditional “one-variable-at-a-time” strategy allowing gain of time and materials as well as consideration of interaction between variables to detect the true optimum [
12]. The Plackett-Burman design is generally used for detection among a large number of factors as the significant ones that can be considered for further optimization. RSM is a collection of mathematical and technical approaches for searching optimum conditions, predicting the response and checking the adequacy of the model [
13]. The experimental design of Box-Behnken was successfully applied in some fields of biotechnology such as the production of laccases [
14], xylo-oligosaccharides [
15] and wastewater treatment [
16].
Despite the considerable data on the production of proteases, no attempt has been conducted on proteases originating from date palm-related endophytic fungi. The present study is therefore designed to optimize the production of protease from endophytic fungi using a statistical design. Using a newly isolated Penicillium bilaiae from the internal tissues of adult date palm tree roots, Plackett-Burman and Box Behnken designs were applied to optimize the conditions of production of a promising acid protease suitable for diverse industrial applications.
2. Materials and Methods
2.1. Isolation of Endophytic Fungi from Healthy Date Palm Roots
Endophytic fungi were isolated from internal tissues of roots of healthy date palms (variety Deglet-Ennour, Nefta, Tunisia), as described by Hallmann et al. [
17]. Briefly, samples were initially washed with tap water for 30 min and then surface sterilized in 70% (
v/
v) ethanol followed by 3% (
w/
v) NaClO treatment for 3 min. The roots were then rinsed several times with sterile distilled water and allowed to surface dry under aseptic conditions. Each root was cut into 0.5–1 cm pieces using a sterile scalpel. Four root pieces were, consequently, placed on plate surface containing potato dextrose agar (PDA) supplemented with 100 μg/mL streptomycin and incubated for 3 to 5 days at 30 °C [
18]. After isolation, pure fungal cultures were obtained by sub-culturing single hyphae several times on new PDA plates.
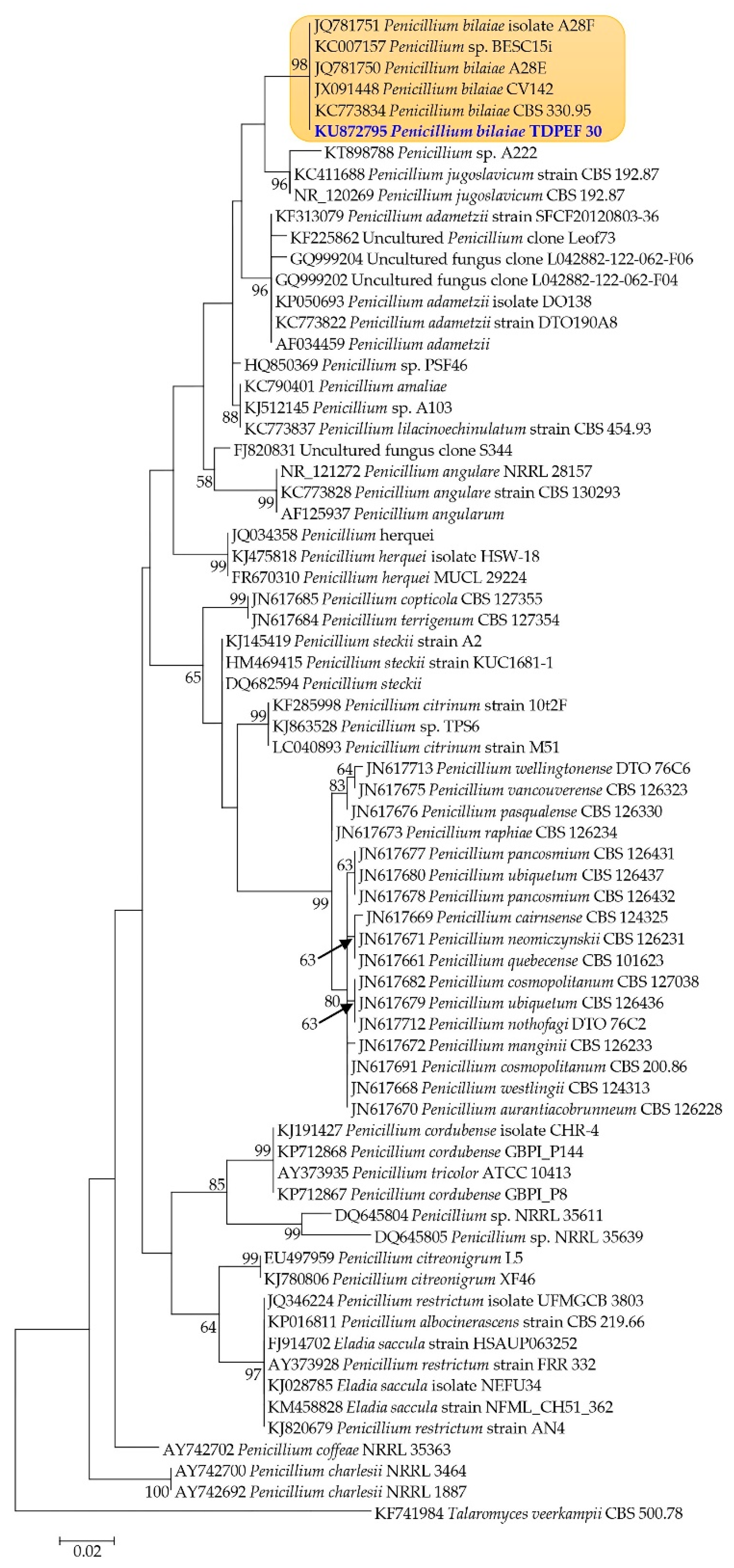
2.2. Molecular Identification and Phylogenetic Analysis of the Endophytic Fungi
Mycelial DNA was extracted from pure culture of fungi using a DNA-Easy Plant Mini kit (QIAGEN, Basel, Switzerland) according to the manufacturer’s specifications. The molecular identification was carried out using the protocol described by Belbahri et al. [
19]. The universal primers used for the PCR amplification of the ITS rRNA region were ITS1 (5′-TCCGTAGGTGAACCTGCGG-3′) and ITS4 (5′-TCCTCCGCTTATTGATATGC-3′) [
20]. Resulting PCR products were sequenced from both sides and consensus sequences were analyzed using BLASTN available from the National Center of Biotechnology Information (NCBI) web site (
http://www.ncbi.nlm.nih.gov/blast/Blast.cgi). Identity of endophytic fungi was based on the percentage of homology to sequences available in the database. CLUSTAL W was used for alignment of endophytic fungal sequences and those recovered from the GenBank [
21]. Distances were calculated and maximum likelihood (ML) trees were constructed using the PhyML program [
22].
2.3. Screening of Protease-Producing Endophytic Fungi on Solid Media
The presence of protease activity was assessed using agar plates containing Czapek Dox medium supplemented with 1% (
w/
v) skimmed milk powder [
23]. The medium was adjusted to pH 7, and poured into plates under aseptic conditions after sterilization. Mycelia from pure culture of fungi were spot inoculated on agar plates and incubated for 5 days at 30 °C. Protease-producing strains were identified by monitoring clear zone formation around them.
2.4. Protease Production from the Endophytic Fungus
Protease production was carried out in 300 mL Erlenmeyer flasks with a working volume of 50 mL of Czapek Dox medium containing (g/L): NaNO3 2.5 g, KH2PO4 1 g, MgSO4·7H2O 0.5 g, KCl 0.5 g supplemented with 1% (w/v) glucose. The medium was sterilized at 121 °C for 20 min. The flasks were inoculated with 106 spores/mL and incubated on a rotary shaker at 150 rpm. Cultures were centrifuged at 8000× g for 15 min and the supernatant was used for determination of proteolytic activity. Experiments were performed in triplicate.
2.5. Protease Assay
Protease activity was measured by the method developed by Kembhavi et al. [
24] using casein as a substrate: 0.5 mL of the enzyme, suitably diluted, was mixed with 0.5 mL of casein (1%) (
w/
v) in 100 mM sodium acetate buffer (pH 5.5). After incubation for 10 min at 50 °C, the reaction was stopped by adding 0.5 mL of TCA (trichloroacetic acid). The mixture was kept for 15 min at room temperature and then centrifuged at 10,000×
g for 15 min to remove the pellet. The absorbance of the soluble fraction was finally measured at 280 nm. A standard curve was established using tyrosine solutions (0–50 mg/L). One unit of protease activity was defined as the amount of enzyme that releases 1 μmol of tyrosine per minute according to standard curve.
2.6. Experimental Designs
2.6.1. Plackett-Burman Design
The independent variables of protease production were initial pH, temperature, MgSO
4, NaNO
3, KH
2PO
4, KCl and glucose. These variables were selected from a preliminary literature review. A Plackett-Burman design was used for multifactor rapid screening to find the most significant independent factors [
25,
26,
27,
28]. Then, the seven factors were investigated using the Plackett-Burman design with a first-order polynomial equation. Each factor was examined at low (−1) and high (+1) levels. Eleven variables (including 4 dummy variables) were screened in 15 experimental runs, as shown in
Table 1. The fitted first-order model is:
Y is the predicted response, β0 and βi are constant coefficients and xi is the coded independent factors.
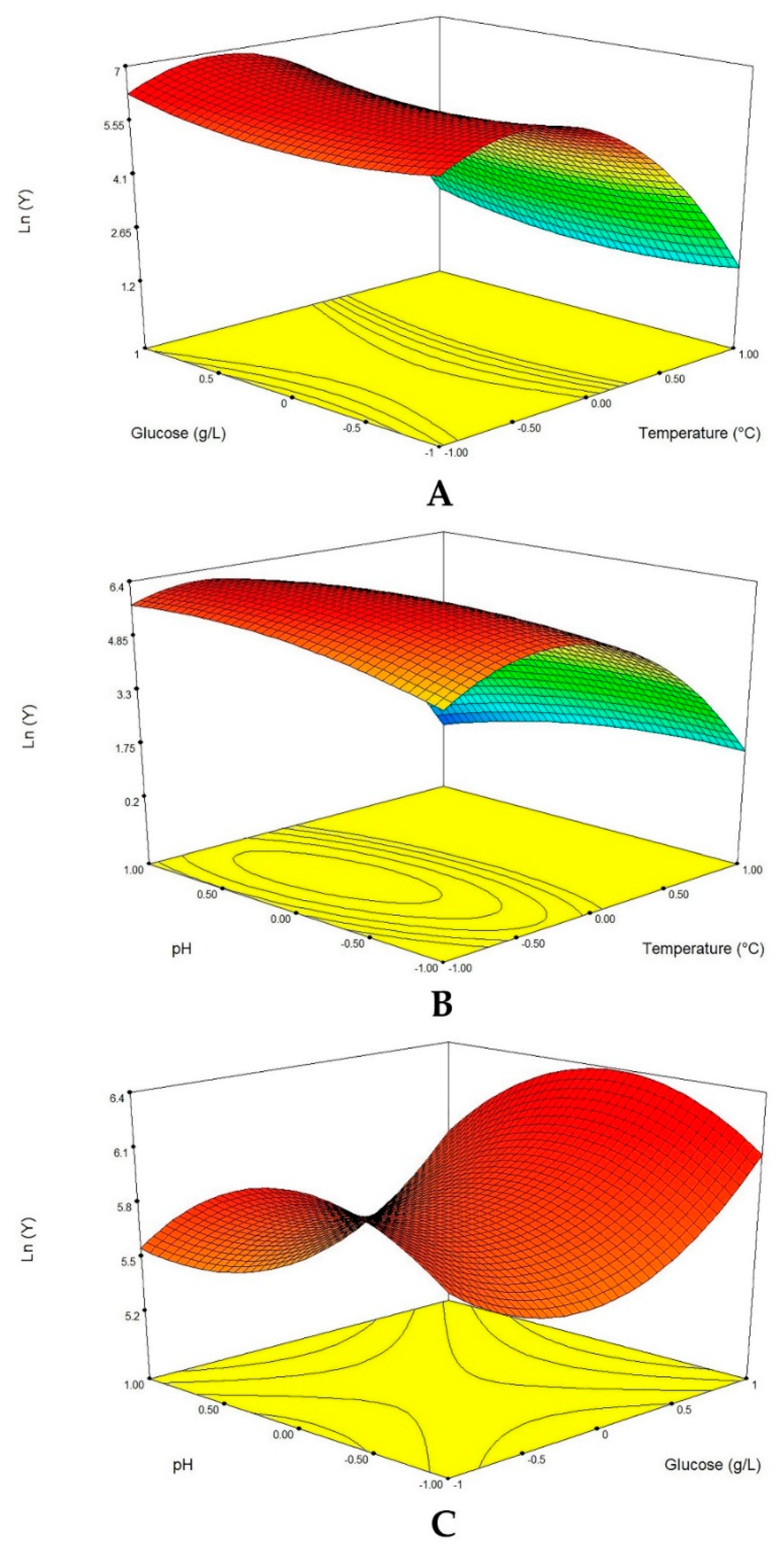
2.6.2. Box-Behnken Design and Response Surface Analysis
The Box-Behnken design (BBD) of RSM was employed to optimize the three most significant factors (temperature, glucose concentration and initial pH) for enhancing protease production from
Penicillium bilaiae, screened by Plackett-Burman design [
29]. The three independent factors were investigated at three levels (−1, 0, 1) and the experimental design used for study is shown in
Table 2. The protease production was fitted using a second order polynomial equation and multiple regression of the data was carried out for obtaining an empirical model related to the most significant factors. The general form of the second-order polynomial equation is:
where
Y is the predicted response,
xi and
xj are independent factors,
β0 is the model intercept,
βi is the linear coefficient,
βii is the quadratic coefficient and
βij is the interaction coefficient.
2.7. Data Analysis and Software
Design Expert 7.0.0 (trial version, Stat Ease Inc., Minneapolis, MN, USA) was used for experimental design and regression analysis of the experimental data. Analysis of variance (ANOVA) was employed to estimate the statistical parameters.
2.8. Effect of Carbon and Nitrogen Sources on Proteolytic Activity
Different sources of carbon and nitrogen were examined for their effect on the production of protease from the endophytic fungus. In total, eight sources of carbon including starch, galactose, sucrose, dextrose, fructose, mannose, xylose and maltose and seven of simple and complex nitrogen sources including casein peptone, ammonium sulfate, urea, casein, yeast extract, KNO3 and malt extract were investigated on the basis of the C/N ratio. The medium amended with glucose and NaNO3 as sources of carbon and nitrogen, respectively, was considered as control.
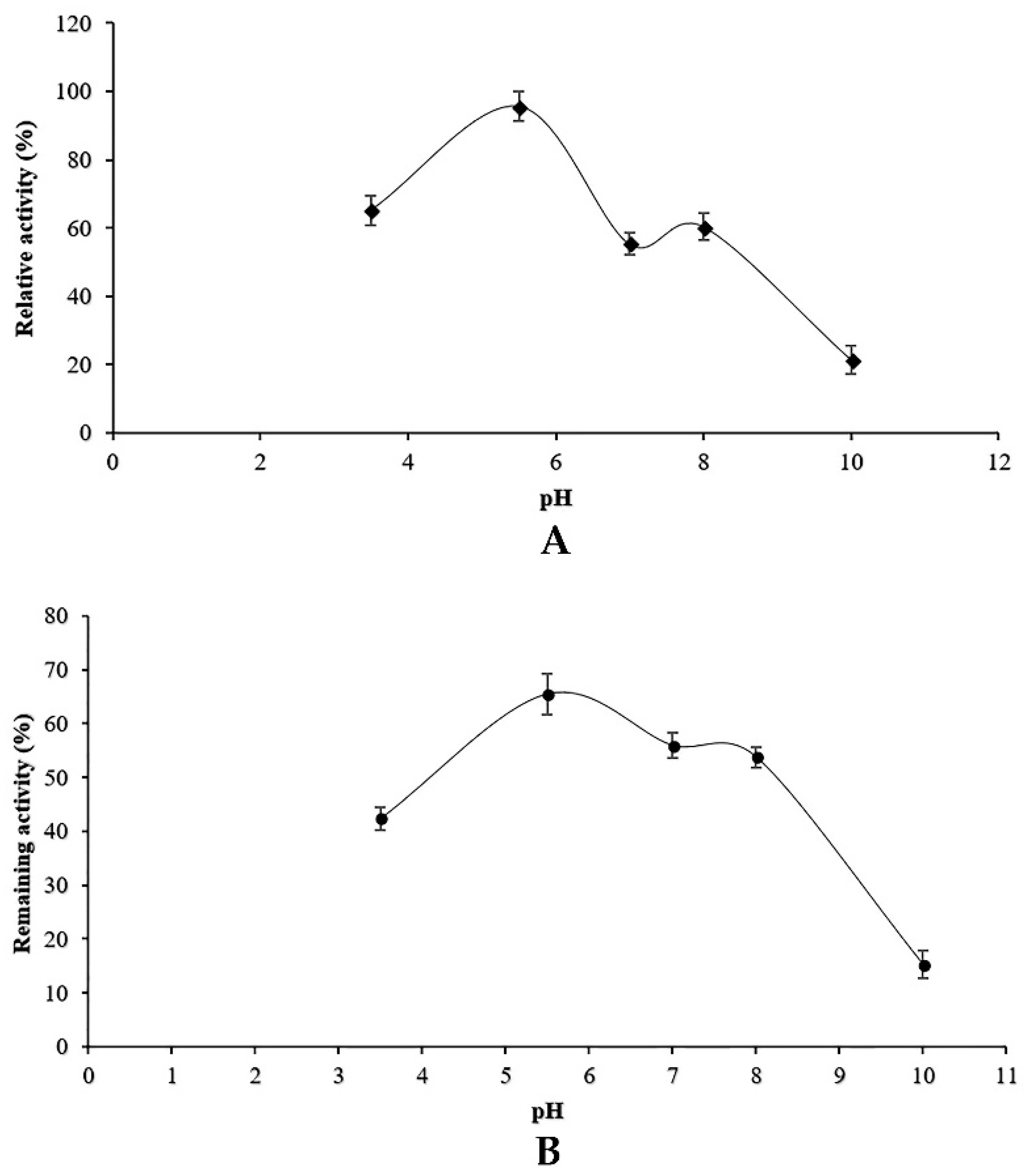
2.9. Effect of pH and Temperature on Enzyme Activity and Stability
Both optimal pH and stability of proteolytic activity at different pH values were studied. For the determination of optimum pH, the enzyme preparation was experimented at various pH values (3–9) using casein (1%) as a substrate. The pH stability experiments were conducted after pre-incubation of protease containing supernatant in different buffers for 1 h at optimum temperature (25 °C). The following buffers were used: 100 mM glycine-HCl for pH 3, 100 mM sodium acetate for pH 4–6, Tris-HCl for pH 6–8, glycine-NaOH for pH 9. The remaining activity tests were assessed under standard assay conditions.
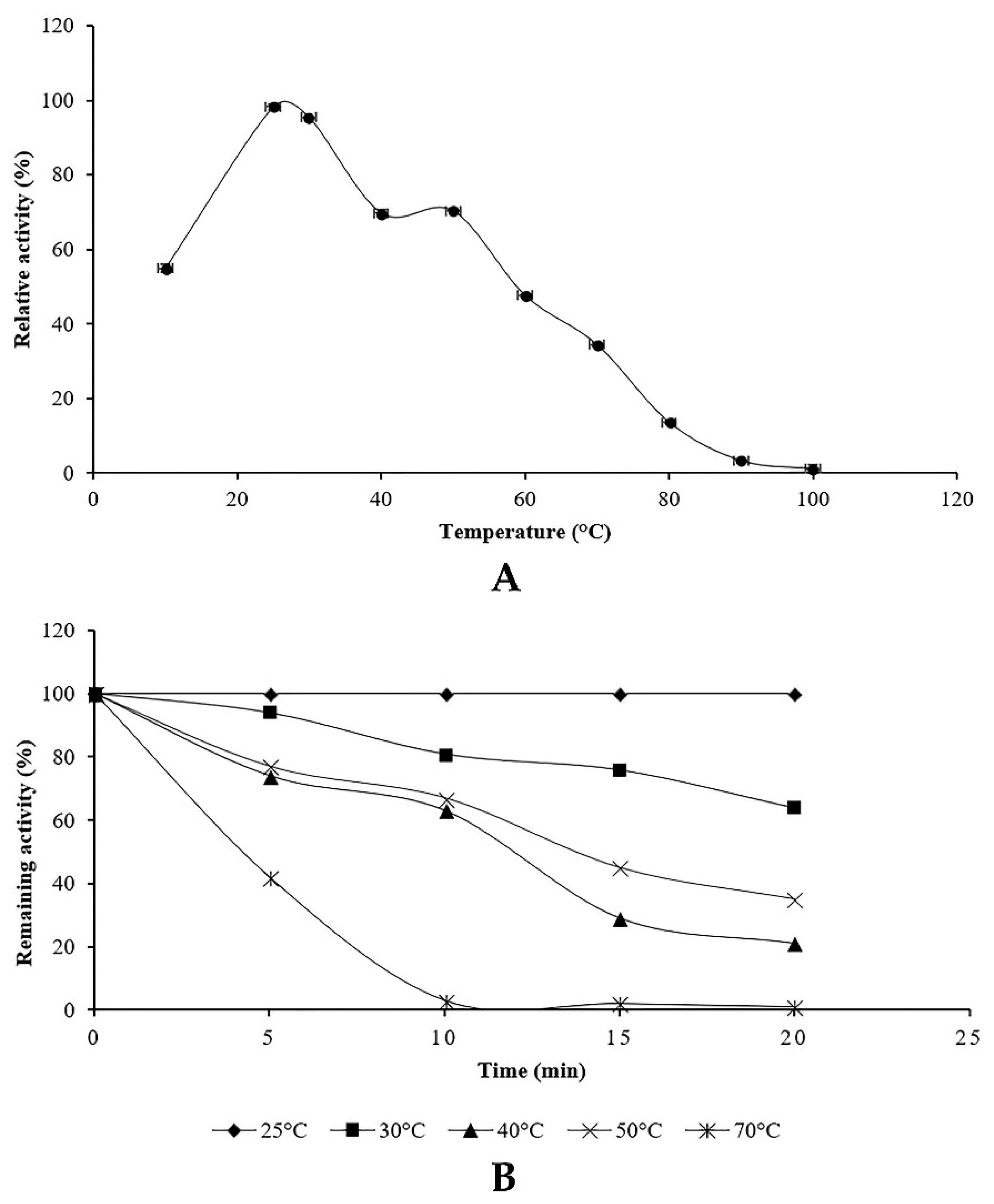
The optimum temperature for proteolytic activity was evaluated at pH 6 using protease assay at different temperatures values (10–100 °C). The effect of temperature on the stability of the protease was assessed by pre-incubating the aliquots at different temperatures for a fixed time. Afterwards, the residual proteolytic activity was determined using casein as a substrate. The unheated crude enzyme was considered as control.
2.10. Effect of Metal Ions, Detergents and Enzyme Inhibitors on Proteolytic Activity
The effect of metal ions was investigated by measuring the residual activity after addition of 5 mM of CuSO4, CaCl2, FeSO4, ZnSO4, MnCl2, HgCl2, H3BO4 and NaCl. In addition, triton X-100, tween 80 and 3-[(3-cholamidopropyl) dimethylammonio]-1-propane sulfonate (CHAPS) were added separately to the production medium at a rate of 0.1% to evaluate their effect on enzyme activity. In order to study the effect of inhibitors such as β-mercaptoethanol, ethylene diamine tetra-acetic acid (EDTA), phenylmethylsulfonyl fluoride (PMSF) and dithiothreitol (DTT) at a rate of 1 mM, the enzyme preparations were pre-incubated with each inhibitor for 1 h at 4 °C before determination of proteolytic activity. The aliquots without any additive were considered as the 100% control.
4. Discussion
The improvement of enzyme production of endophytic microorganisms is the purpose of several investigations owing to their abilities to produce many valuables metabolites with potential bioactivity [
31]. In the present study, endophytic fungi were isolated from the root internal tissues of adult date palm trees. These isolates were evaluated for their ability to produce proteases. Plating on skim milk agar medium, as described by Saran et al. [
32], is an easy and simple technique to screen for protease activity, although it cannot be recommended for quantitative analysis. The wide use of proteases in biotechnological and industrial processes has attracted researchers to obtain proteolytic enzymes that exhibit various properties suitable for different applications. However, each microbial strain has its specific conditions to maximize protease production. According to our results, the endophytic fungus
Penicillium bilaiae TDPEF30 was found to be the best producer of proteases and was selected for further assays of protease production in liquid medium. Few studies have been done to induce protease production from endophytic fungi using a statistical design. Using Czapek Dox as fermentation medium, the TDPEF30 endophytic strain displayed the best level of protease production with 203 ± 2.1 U/mL. Czapek Dox is an easy to prepare medium with simple components [
33].
RSM was carried out to optimize culture medium nutrients and conditions for the production of protease by the endophytic fungus
Penicillium bilaiae TDPEF30. At the beginning, the significant variables that influence the protease production were selected using Plackett-Burman Design. Rodriguez et al. [
34] as well as Hajji et al. [
35] reported that proteases from
Penicillium and
Aspergillus fungi were affected by temperature and pH. In line with these studies, it was demonstrated in the present study that glucose concentration, pH and temperature had significant effects on protease synthesis by the endophytic fungus
Penicillium bilaiae TDPEF30. Then, the BBD was adopted to find out the optimum levels of each significant variable and their interactive effects on protease yield.
Nowadays, application of RSM in biotechnology is increasingly used to statistically determine the best condition for enzyme production, owing to its easy applicability, validity and reliability [
36]. In our study, protease production by the endophytic fungus of adult Tunisian date palm was enhanced. However, maximum protease production was achieved at temperature 24.46 °C, pH 6.26 and 13.75 g/L glucose while the other components were used at the lowest level 0.2 g/L MgSO
4, 1.5 g/L NaNO
3, 0.5 g/L KH
2PO
4 and 0.5 g/L KCl. The RSM process resulted in a proteolytic activity of 1086.95 U/mL, with a 5.8 increase compared to 186.21 U/mL obtained before optimization. Furthermore, the high degree of similarity between experimental and predicted values reflected the accuracy and the validity of the experimental design applied.
It is well known that microbial protease production is dependent on the availability of both carbon and nitrogen sources, which play a regulatory role in enzyme synthesis [
37]. In the current study, various sources of carbon and nitrogen were investigated in order to find the maximum protease production by the endophytic fungus
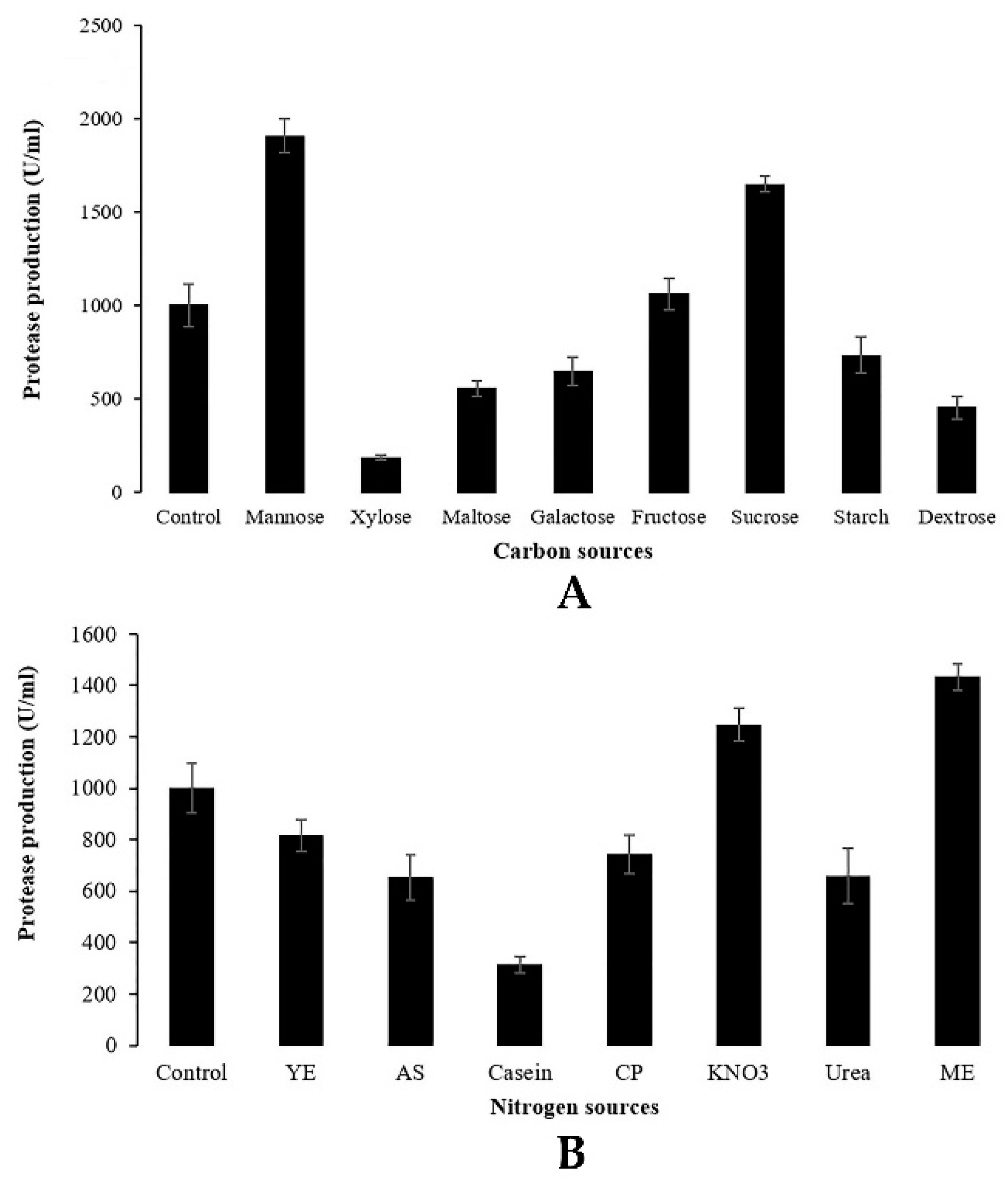
Penicillium bilaiae TDPEF30. The significant effect of sources of carbon and nitrogen either on protease activity or on enzyme production is shown in
Figure 3. Concerning carbon source influences, the highest level of protease activity was obtained in presence of the simple carbohydrate mannose with 1907.28 U/mL followed by complex sources of carbon, namely, sucrose and fructose, with 1649.12 and 1060.15 U/mL, respectively. These different carbon sources have the benefits of their commercial availability. Nonetheless, other carbon sources such as xylose, dextrose, maltose, galactose and starch decreased the protease production by the endophytic fungus TDPEF30 compared to the control glucose-based medium. In general, both organic and inorganic sources of nitrogen were evaluated for their influence on protease production. In the present study, only malt extract and KNO
3 increased the proteolytic enzyme yield compared to the control with NaNO
3. Our findings are in agreement with the results of Hameed et al. [
38], who reported that the addition of mannose induced significant proteolytic activity in etiolated wheat leaves. In addition, Sumantha et al. [
39] and Zaferanloo et al. [
33] found that protease production by
Rhizopus microsporus NRRL 3671 and
Alternaria alternata, respectively, was significantly influenced by specific sources of carbon and nitrogen, which could be explained by the specificity of protease enzyme.
Along with carbon and nitrogen sources, the environmental conditions such as pH medium and temperature could play an important role in the induction or repression of protease activity. Our results indicated that
Penicillium bilaiae produced protease at pH range 5–6.5 with an optimum at six. In addition, maximum protease activity was obtained at 25 °C. In this study, it was found that protease activity increased slightly at pH 8 and 50 °C. These findings could be explained by the presence of isoenzymes produced by the endophytic fungus TDPEF30 during fermentation. This suggests the potentiality of endophytic fungus TDPEF30 to produce a wide range of valuable enzymes with different properties and even in the same class. Enzyme inactivation represents one of the major limitations in the use of proteases in biotechnological processes. The protease produced by the endophytic fungus TDPEF30 lost its activity at 70 °C and pH 10. Most fungi have an optimum of temperature at the range of 28–30 °C (Hajji et al., 2008; Germano et al., 2003) [
35,
40]. However, earlier studies reported that production of proteases from
Penicillium species like
P. citrinum,
P. perpurogerum and
P. funculosum was obtained at temperature lower than 30 °C [
41].
The effect of metal ions on protease production by the endophytic fungus
Penicillium bilaiae TDPEF30 at a concentration of 5 mM was evaluated under optimal conditions of pH and temperature. Results indicated that the addition of both Mn
2+ and B
3+ enhanced the fungal protease activity. In contrast, Cu
2+, Ca
2+, Fe
2+ and Hg
2+ significantly decreased protease activity. Similar effects of Mn
2+ on proteolytic enzyme production were obtained with
Penicillium sp. [
42]. This finding might be attributed to manganese ion involvement in stabilization of the enzyme molecular structure. In contrast, our findings are in disagreement with the study of Negi & Banerjee [
37], who found that Ca
2+ and Hg
2+ simulated protease production by
Aspergillus awamori.
The influence of detergents on protease activity was well studied with bacteria, but with fungi, few studies have been reported. Among the detergents, only CHAPS decreased protease activity. Our results are in agreement with the study of Guleria et al. [
43], which revealed that the alkaline protease from
Bacillus amyloliquefaciens SP1 increased in the presence of triton X-100. Furthermore, Guleria et al. [
43] reported the high level of proteolytic activity obtained after incubation of enzyme solution in tween 80.
The effect of various enzyme inhibitors (used at a concentration of 5 mM) was determined to evaluate their influence on protease activity. Only PMSF, a well-known as serine protease inhibitor, inhibited protease activity from the endophytic fungus of adult date palm, TDPEF30. We speculate that the protease optimized during the current study can be classified as serine protease.