Emerging and Novel Viruses in Passerine Birds
Abstract
1. Introduction
2. Families of Viruses Affecting Passeriformes
2.1. Enveloped Double-Stranded DNA (dsDNA) Viruses
2.1.1. Family Herpesviridae
2.1.2. Family Poxviridae
2.2. Non-Enveloped dsDNA Viruses
2.2.1. Family Adenoviridae
2.2.2. Family Papillomaviridae
2.3. Single-Stranded (ss)DNA Viruses
2.3.1. Families Circoviridae, Anelloviridae and Genomoviridae
2.3.2. Family Parvoviridae
2.4. Enveloped Negative-Sense Single-Stranded RNA (ssRNA) Viruses
2.4.1. Family Orthomyxoviridae
2.4.2. Family Paramyxoviridae
2.4.3. Family Pneumoviridae
2.4.4. Family Bornaviridae
2.5. Enveloped Positive-Sense ssRNA Viruses
2.5.1. Family Flaviviridae
2.5.2. Family Togaviridae
2.5.3. Family Coronaviridae
2.6. Non-Enveloped Positive-Sense ssRNA Viruses
2.6.1. Family Picornaviridae
2.6.2. Family Astroviridae
2.6.3. Family Caliciviridae
2.6.4. Family Hepeviridae
2.7. Double-Stranded RNA (dsRNA) Viruses
Family Reoviridae
2.8. RNA Reverse Transcribing Virus
Family Retroviridae
3. General Recommendations to Reduce the Risk of Transmission of Viruses Associated with Passerines
4. Conclusions
| Viral Family | Genus | Viral Taxa | Abbreviations | Nomenclature | Bird Family | Clinical Signs | Mortality Rate | References |
|---|---|---|---|---|---|---|---|---|
| dsDNA VIRUSES | ||||||||
| Herpesviridae | Iltovirus | Gallid herpesvirus 1 | GaHV-1 | Iltovirus gallidalpha1 | Turdidae, Estrildidae, Sturnidae | Subclinical infection/ Respiratory signs | High | [7,26,34,35,37] |
| Psittacid herpesvirus 1 | PsHV-1 | Iltovirus psittacidalpha1 | Sturnidae | |||||
| passerid herpesvirus 1 | not recognized | Fringillidae | ||||||
| Mardivirus | Columbid herpesvirus 1 | CoHV-1 | Mardivirus columbidalpha1 | Corvidae, Turdidae | Subclinical infection | Probably not | [40] | |
| Poxviridae | Avipoxvirus | Fowlpox virus (Lineage A1) | FWPV | same | Fringillidae, Passeridae | Cutaneous lesions/Upper respiratory and digestive tract lesions | High | [42,43,44,45,46,47,48,49,50,51,52,53,54] |
| Pigeonpox virus (Lineage A2) | PGPV | same | Fringillidae | High/Medium | ||||
| Accipitriformespox virus (Lineage A7) | same | Lanidae | High/Medium | |||||
| Canarypox virus (Lineage B1) | CNPV | same | Fringillidae, Paridae, Corvidae (and 11 other families) | High/Medium | ||||
| Starlingpox virus (Lineage B2) | same | Sturnidae, Passeridae | High/Medium | |||||
| Lineage B3 poxvirus | not recognized | Corvidae (and eight other mostly Passeriformes families) | High | |||||
| Adenoviridae | Atadenovirus | European robin adenovirus | not recognized | Meliphagidae, Fringillidae, Ploeceidae, Estrildidae, Turdidae, Ptilonorhynchidae, Sturnidae, Zosteropidae, Maluridae, Petroicidae | Subclinical infection/ Not known | Probably not | [57,58] | |
| European greenfinch adenovirus | not recognized | |||||||
| Eurasian siskin adenovirus | not recognized | |||||||
| Eurasian bullfinch adenovirus | not recognized | |||||||
| common chaffinch adenovirus | not recognized | |||||||
| passerine adenovirus 1 | not recognized | |||||||
| white plumed honeyeater adenovirus 1, -2 | not recognized | |||||||
| vitelline masked weaver adenovirus strains 37869 and 38132 | not recognized | |||||||
| Aviadenovirus | great tit adenovirus strain 47292 | not recognized | Fringillidae, Estrildidae, Paridae, Ploceidae, Sylvidae, Falcunculidae, Petroicidae | Subclinical infection/ Not known | Probably not | |||
| european greenfinch adenovirus | not recognized | |||||||
| goldfinch adenovirus | not recognized | |||||||
| vitelline masked weaver adenovirus strain 39658 | not recognized | |||||||
| Siadenovirus | double-barred finch adenovirus | not recognized | Estrildidae, Paridae, Sylvidae, Thraupidae, Meliphagidae, Pardalotidae, Rhipiduridae, Sturnidae | Subclinical infection/ Probable renal lesions | Probably not | [56,60] | ||
| eurasian blackcap adenovirus | not recognized | |||||||
| gouldian finch adenovirus 1 | GFAdV-1 | not recognized | ||||||
| great tit adenovirus strain 47292 | GTAdV-1 | Great tit siadenovirus A | ||||||
| zebra finch adenovirus strain 47535 | not recognized | |||||||
| Papillomaviridae | Etapapillomavirus | Fringilla coelebs Papillomavirus 1 | FcPV1 | Etapapillomavirus 1 | Fringillidae | Cutaneous lesions | Low | [66,67,69,70,71,72] |
| Serinus canaria papillomavirus 1 | ScPV1 | not recognized | Fringillidae | Not known | Not known | [68] | ||
| ssDNA VIRUSES | ||||||||
| Circoviridae | Circovirus | Beak and feather disease virus | BFDV | Circovirus parrot | Artamidae, Corvidae, Estrildidae, Fringillidae, Petoicidae, Sturnidae, Thamnophilidae | Feather lesions | Probably not | [76,77,78] |
| Circovirus canary | CaCV | same | Feather diseases and immunosupression/ Subclinical infection | Yes/Not known | [25,79,80,81,82] | |||
| Circovirus finch | FiCV | same | ||||||
| Circovirus raven | RaCV | same | ||||||
| Circovirus starling | StCV | same | ||||||
| Circovirus zebrafinch | ZfiCV | same | ||||||
| Cyclovirus | Robinz virus RP_736 | RobinzV736 | Cyclovirus cervienka | Petroicidae | Not known | Not known | [17] | |
| Robinz virus RP_1170 | RobinzV1170 | Cyclovirus liepsnele | ||||||
| Robinz virus RP_493 | RobinzV493 | Cyclovirus pettirosso | ||||||
| Robinz virus RP_620 | RobinzV620 | Cyclovirus prihor | ||||||
| Robinz virus RP_526 | RobinzV526 | Cyclovirus punarinta | ||||||
| Robinz virus RP_584 | RobinzV584 | Cyclovirus rudzik | ||||||
| Robinz virus RP_259 | RobinzV259 | Cyclovirus totoi | ||||||
| Probably new genus | Gyrovirus 11 | not recognized | Thamnophilidae | Not known | Not known | [89] | ||
| Genomoviridae | Gemycirculavirus | new species | not recognized | Petroicidae, Aegithalidae, Corvidae, Emberizidae, Fringillidae, Muscicapidae, Paridae, Sylviidae, Turdidae, Zosteropidae | Not known | Not known | [88] | |
| Chickadee associated gemycircularvirus 1 | Gemycircularvirus chickad1 | Paridae | [85] | |||||
| Finch associated genomovirus 5 | Gemycircularvirus haeme1 | Fringillidae | [86] | |||||
| Finch associated genomovirus 6 | Gemycircularvirus haeme2 | [86] | ||||||
| Gemykibivirus | Blackbird associated gemykibivirus 1 | Gemykibivirus blabi1 | Turdidae | [87] | ||||
| Black robin associated gemykibivirus 1 | Gemykibivirus blaro1 | Petroicidae | ||||||
| Finch associated genomovirus 3 | Gemykibivirus haeme1 | Fringillidae | [86] | |||||
| Finch associated genomovirus 2 | Gemykibivirus haeme3 | |||||||
| Finch associated genomovirus 4 | Gemykibivirus haeme4 | |||||||
| Finch associated genomovirus 1 | Gemykibivirus haeme5 | |||||||
| Gemykroznavirus | Finch associated genomovirus 8 | Gemykronzavirus haeme1 | Fringillidae | |||||
| Gemygorvirus | Starling associated gemygorvirus 1 | Gemygorvirus stara1 | Sturnidae | [87] | ||||
| Parvoviridae | Aveparvovirus | Aveparvovirus passeriform1 | PfPV | same | Thraupidae | Not known | Not known | [95] |
| ssRNA VIRUSES | ||||||||
| Orthomyxoviridae | Alphainfluenzavirus 1 | Highly Pathogenic Avian Influenza H5N1 | FLUAV/(H5N1) | Alphainfluenzavirus influenzae | Passeridae, Turdidae, Ploceidae, Corvidae, Hirundidae, Icteridae (and many other families) | Mild/Severe disease | High | [101,102,103,104,105,106,107,108,110,116] |
| Paramyxoviridae | Orthoavulavirus | Avian paramyxovirus 1 (NDV, Newcastle Disease virus) | APMV-1 | Orthoavulavirus javaense | Passeridae, Motacillidae, Passarellidae | Not known/Subclinical disease | High/Not known | [113,123,124,126,127] |
| Metaavulavirus | Avian paramyxovirus 2 | APMV-2 | Metaavulavirus yucaipaense | Corvidae, Motacillidae, Troglodytidae, Muscicapidae, Emberizidae, Estrildidae | Mild/Severe disease | [128,129,130,131,132,133,134] | ||
| Paraavulavirus | Avian paramyxovirus 3 | APMV-3 | Paraavulavirus wisconsinense | [135,136] | ||||
| Paraavulavirus | Avian paramyxovirus 4 | APMV-4 | Paraavulavirus hongkongense | [138] | ||||
| Pneumoviridae | Metapneumovirus | Avian metapneumovirus | AMPV | Metapneumovirus avis | Passeridae, Sturnidae | Not known | Not known | [141,142] |
| Bornaviridae | Orthobornavirus | Estrildid finch bornavirus 1 | EsBV-1 | Orthobornavirus estrildidae | Estrildidae, Corvidae, Fringillidae | Not known | Not known | [146,147,148,149] |
| Canary bornavirus 1 | CnBV-1 | Orthobornavirus serini | ||||||
| Canary bornavirus 2 | CnBV-2 | |||||||
| Canary bornavirus 3 | CnBV-3 | |||||||
| Flaviviridae | Orthoflavivirus 1 | Japanese encephalitis virus | JEV | Orthoflavivirus japonicum | Passeridae, Corvidae, Fringillidae, Turdidae (and >23 more families) | Not known | High | [183] |
| Saint Louis encephalitis virus | SLEV | Orthoflavivirus louisense | [191,192,193,194] | |||||
| Murray Valley Encephalitis virus | MVEV | Orthoflavivirus murrayense | [186] | |||||
| Usutu virus | USUV | Orthoflavivirus usutuense | Severe disease | [174,175,176] | ||||
| West Nile virus | WNV | Orthoflavivirus nilense | Mild/Severe disease | [158,159,160,161,162,169] | ||||
| Togaviridae | Alphavirus 1 | Eastern equine encephalitis virus | EEEV | same | Turdidae, Hirundidae, Icteridae, Petroicidae, Estrildidae, Monarchidae | Neurological disease | Yes | [206,207,208] |
| Highlands J virus | HJV | same | Mild disease | [210] | ||||
| Sindbis virus | SINV | same | Severe disease | [215,216] | ||||
| Ross River virus | RRV | same | Not known | [219,220] | ||||
| Mayaro virus | MAYV | same | [225,226,227,228] | |||||
| Buggy Creek virus | BUCGV | Fort Morgan virus | Neurological disease | [230,231,232] | ||||
| Deltacoronavirus 2 | Bulbul coronavirus HKU11 | BulCV_HKU11 | same | Turdidae, Estrildidae, Pycnonotidae, Passeridae, Zosteropidae, Fringillidae, Emeberizidae | Severe disease | Probably high | [239,240,241] | |
| Munia coronavirus HKU13 | MunCV_HKU13 | same | ||||||
| Thrush coronavirus | not recognized | |||||||
| White-eye coronavirus HKU16 | WECoV_HKU16 | same | ||||||
| Coronavirus HKU15 | PoCoV_HKU15 | same | ||||||
| Picornaviridae | Oscivirus | Oscivirus A1 and A2 (turdivirus 2 and 3) | OsV-A1/OsV-A2 | Oscivirus A | Turdidae, Muscicapidae | Severe disease | Not known | [247,248] |
| Passerivirus | Passerivirus A and B | PasV-A/PasV-B | same | Turdidae, Estrildidae | High | |||
| Poecivirus | Poecivirus A1 | PoeV-A | Poecivirus A | Paridae | Yes | [251,252] | ||
| Probably new genus | French Guiana Picornavirus | FGPV | not recognized | Thamnophilidae | Not known | Not known | [13] | |
| Astroviridae | Avastovirus | novel avastrovirus clades 4 and 5 | not recognized | Fringillidae, Monarchidae Parulidae, Passeridae | Subclinical infectionMild disease | Not known | [258,259,260] | |
| Caliciviridae | Probably new genera | new calicivirus | not recognized | Prunellidae, Turdidae, Emberizidae, Fringillidae, Muscicapidae, Petroicidae | Subclinical infection | Not known | [7,15,275,276] | |
| Hepeviridae | Probably new genera | new hepe-like viruses | not recognized | Thamnophilidae, Turdidae, Zosteropidae | Not known | Not known | [7,13,15] | |
| dsRNA VIRUSES | ||||||||
| Reoviridae | Orthoreovirus 2 | new orthoreovirus | not recognized | Emberizidae, Turdidae, Fringillidae | Not known | High mortality | [283,284] | |
| RNA REVERSE TRANSCRIBING VIRUS | ||||||||
| Retroviridae | Alpharetrovirus | Avian leukosis virus | ALV | same | Emberizidae, Paridae, Corvidae, Muscicapidae, Thraupidae, Phylloscopidae | Subclinical infection | Possibly | [291,292,293] |
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Burrell, C.J.; Howard, C.R.; Murphy, F.A. (Eds.) Chapter 15—Emerging Virus Diseases. In Fenner and White’s Medical Virology, 5th ed.; Academic Press: London, UK, 2017; pp. 217–225. ISBN 978-0-12-375156-0. [Google Scholar]
- Daly, J.M. Zoonosis, Emerging and Re-Emerging Viral Diseases. In Encyclopedia of Virology; Bamford, D., Zuckerman, M., Eds.; Elsevier: Amsterdam, The Netherlands, 2021; pp. 569–576. ISBN 978-0-12-814516-6. [Google Scholar]
- Ertunc, F. Emerging Plant Viruses. In Emerging and Reemerging Viral Pathogens; Ennaji, M., Ed.; Elsevier: Amsterdam, The Netherlands, 2020; pp. 1041–1062. ISBN 978-0-12-819400-3. [Google Scholar]
- World Health Organization. A Brief Guide to Emerging Infectious Diseases and Zoonoses; WHO Regional Office for South-East Asia: New Delhi, India, 2014; ISBN 978-92-9022-458-7. [Google Scholar]
- World Organization for Animal Health. Terrestrial Code Online Access. Available online: https://www.woah.org/en/what-we-do/standards/codes-and-manuals/terrestrial-code-online-access/ (accessed on 19 June 2023).
- Yadav, M.P.; Singh, R.K.; Malik, Y.S. Emerging and Transboundary Animal Viral Diseases: Perspectives and Preparedness. In Emerging and Transboundary Animal Viruses; Malik, Y.S., Singh, R.K., Yadav, M.P., Eds.; Livestock Diseases and Management; Springer: Singapore, 2020; pp. 1–25. ISBN 9789811504020. [Google Scholar]
- French, R.K.; Filion, A.; Niebuhr, C.N.; Holmes, E.C. Metatranscriptomic Comparison of Viromes in Endemic and Introduced Passerines in New Zealand. Viruses 2022, 14, 1364. [Google Scholar] [CrossRef] [PubMed]
- Manne, L.L.; Brooks, T.M.; Pimm, S.L. Relative Risk of Extinction of Passerine Birds on Continents and Islands. Nature 1999, 399, 258–261. [Google Scholar] [CrossRef]
- Jiménez-Clavero, M.Á. Animal Viral Diseases and Global Change: Bluetongue and West Nile Fever as Paradigms. Front. Genet. 2012, 3, 105. [Google Scholar] [CrossRef]
- Canuti, M.; Munro, H.J.; Robertson, G.J.; Kroyer, A.N.K.; Roul, S.; Ojkic, D.; Whitney, H.G.; Lang, A.S. New Insight into Avian Papillomavirus Ecology and Evolution from Characterization of Novel Wild Bird Papillomaviruses. Front. Microbiol. 2019, 10, 701. [Google Scholar] [CrossRef]
- Chan, J.F.-W.; To, K.K.-W.; Chen, H.; Yuen, K.-Y. Cross-Species Transmission and Emergence of Novel Viruses from Birds. Curr. Opin. Virol. 2015, 10, 63–69. [Google Scholar] [CrossRef]
- Wang, C.C.; Prather, K.A.; Sznitman, J.; Jimenez, J.L.; Lakdawala, S.S.; Tufekci, Z.; Marr, L.C. Airborne Transmission of Respiratory Viruses. Science 2021, 373, eabd9149. [Google Scholar] [CrossRef] [PubMed]
- Truchado, D.A.; Llanos-Garrido, A.; Oropesa-Olmedo, D.A.; Cerrada, B.; Cea, P.; Moens, M.A.J.; Gomez-Lucia, E.; Doménech, A.; Milá, B.; Pérez-Tris, J.; et al. Comparative Metagenomics of Palearctic and Neotropical Avian Cloacal Viromes Reveal Geographic Bias in Virus Discovery. Microorganisms 2020, 8, 1869. [Google Scholar] [CrossRef]
- Dai, Z.; Wang, H.; Wu, H.; Zhang, Q.; Ji, L.; Wang, X.; Shen, Q.; Yang, S.; Ma, X.; Shan, T.; et al. Parvovirus Dark Matter in the Cloaca of Wild Birds. GigaScience 2023, 12, giad001. [Google Scholar] [CrossRef]
- Shan, T.; Yang, S.; Wang, H.; Wang, H.; Zhang, J.; Gong, G.; Xiao, Y.; Yang, J.; Wang, X.; Lu, J.; et al. Virome in the Cloaca of Wild and Breeding Birds Revealed a Diversity of Significant Viruses. Microbiome 2022, 10, 60. [Google Scholar] [CrossRef]
- Chang, W.-S.; Eden, J.-S.; Hall, J.; Shi, M.; Rose, K.; Holmes, E.C. Metatranscriptomic Analysis of Virus Diversity in Urban Wild Birds with Paretic Disease. J. Virol. 2020, 94, e00606–e00620. [Google Scholar] [CrossRef]
- Custer, J.M.; White, R.; Taylor, H.; Schmidlin, K.; Fontenele, R.S.; Stainton, D.; Kraberger, S.; Briskie, J.V.; Varsani, A. Diverse Single-Stranded DNA Viruses Identified in New Zealand (Aotearoa) South Island Robin (Petroica australis) Fecal Samples. Virology 2022, 565, 38–51. [Google Scholar] [CrossRef]
- Woolhouse, M.; Scott, F.; Hudson, Z.; Howey, R.; Chase-Topping, M. Human Viruses: Discovery and Emergence. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2012, 367, 2864–2871. [Google Scholar] [CrossRef]
- Clements, J.; Schulenberg, T.; Iliff, M.; Billerman, S.; Fredericks, T.; Gerbracht, J.; Woods, C. Checklist of Birds of the World. Available online: https://www.birds.cornell.edu/clementschecklist/download/ (accessed on 19 July 2023).
- Callaghan, C.T.; Nakagawa, S.; Cornwell, W.K. Global Abundance Estimates for 9700 Bird Species. Proc. Natl. Acad. Sci. USA 2021, 118, e2023170118. [Google Scholar] [CrossRef] [PubMed]
- Gibb, R.; Redding, D.W.; Chin, K.Q.; Donnelly, C.A.; Blackburn, T.M.; Newbold, T.; Jones, K.E. Zoonotic Host Diversity Increases in Human-Dominated Ecosystems. Nature 2020, 584, 398–402. [Google Scholar] [CrossRef] [PubMed]
- Boseret, G.; Losson, B.; Mainil, J.G.; Thiry, E.; Saegerman, C. Zoonoses in Pet Birds: Review and Perspectives. Vet. Res. 2013, 44, 36. [Google Scholar] [CrossRef]
- Ayala, A.J.; Yabsley, M.J.; Hernandez, S.M. A Review of Pathogen Transmission at the Backyard Chicken–Wild Bird Interface. Front. Vet. Sci. 2020, 7, 539925. [Google Scholar] [CrossRef]
- Marzal, A.; Ferraguti, M.; Muriel, J.; Magallanes, S.; Ortiz, J.A.; García-Longoria, L.; Bravo-Barriga, D.; Guerrero-Carvajal, F.; Aguilera-Sepúlveda, P.; Llorente, F.; et al. Circulation of Zoonotic Flaviviruses in Wild Passerine Birds in Western Spain. Vet. Microbiol. 2022, 268, 109399. [Google Scholar] [CrossRef]
- Todd, D.; Weston, J.; Ball, N.W.; Borghmans, B.J.; Smyth, J.A.; Gelmini, L.; Lavazza, A. Nucleotide Sequence-Based Identification of a Novel Circovirus of Canaries. Avian Pathol. 2001, 30, 321–325. [Google Scholar] [CrossRef]
- Tomaszewski, E.K.; Gravendyck, M.; Kaleta, E.F.; Phalen, D.N. Genetic Characterization of a Herpesvirus Isolate from a Superb Starling (Lamprotornis superbus) as a Psittacid Herpesvirus Genotype 1. Avian Dis. 2004, 48, 212–214. [Google Scholar] [CrossRef]
- Van Riper, C., III; Forrester, D.J. Avian Pox. In Infectious Diseases of Wild Birds; Thomas, N., Hunter, D., Atkinson, C., Eds.; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2007; pp. 131–176. ISBN 978-0-470-34466-8. [Google Scholar]
- Couteaudier, M.; Denesvre, C. Marek’s Disease Virus and Skin Interactions. Vet. Res. 2014, 45, 36. [Google Scholar] [CrossRef]
- Davison, A.J.; Eberle, R.; Ehlers, B.; Hayward, G.S.; McGeoch, D.J.; Minson, A.C.; Pellett, P.E.; Roizman, B.; Studdert, M.J.; Thiry, E. The Order Herpesvirales. Arch. Virol. 2009, 154, 171–177. [Google Scholar] [CrossRef] [PubMed]
- Dhama, K.; Kumar, N.; Saminathan, M.; Tiwari, R.; Karthik, K.; Kumar, M.A.; Palanivelu, M.; Shabbir, M.Z.; Malik, Y.S.; Singh, R.K. Duck Virus Enteritis (Duck Plague)—A Comprehensive Update. Vet. Q. 2017, 37, 57–80. [Google Scholar] [CrossRef]
- Kaleta, E.F.; Redmann, T. Infectious laryngotracheitis in chickens, peacocks and pheasants and means and limitations for its control with attenuated live vaccines. Tierarztl. Prax. Ausg. G. Grosstiere Nutztiere 1997, 25, 605–610. [Google Scholar] [PubMed]
- Panigrahy, B.; Grumbles, L.C. Pacheco’s Disease in Psittacine Birds. Avian Dis. 1984, 28, 808–812. [Google Scholar] [CrossRef] [PubMed]
- Phalen, D.N.; Alvarado, C.; Grillo, V.; Mason, P.; Dobson, E.; Holz, P. Prevalence of Columbid Infection in Feral Pigeons from New South Wales and Victoria, Australia, with Spillover into a Wild Powerful Owl (Ninox struena). J. Wildl. Dis. 2017, 53, 543–551. [Google Scholar] [CrossRef]
- Žlabravec, Z.; Slavec, B.; Vrezec, A.; Kuhar, U.; Zorman Rojs, O.; Golob, Z.; Račnik, J. Detection of Herpesviruses in Wild Bird Casualties in Slovenia. Front. Vet. Sci. 2022, 9, 822212. [Google Scholar] [CrossRef] [PubMed]
- Žlabravec, Z.; Trilar, T.; Slavec, B.; Krapež, U.; Vrezec, A.; Rojs, O.Z.; Račnik, J. Detection of Herpesviruses in Passerine Birds Captured during Autumn Migration in Slovenia. J. Wildl. Dis. 2021, 57, 368–375. [Google Scholar] [CrossRef]
- Kaleta, E.F.; Docherty, D.E. Avian Herpesviruses. In Infectious Diseases of Wild Birds; Thomas, N., Hunter, D., Atkinson, C., Eds.; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2007; pp. 63–86. ISBN 978-0-470-34466-8. [Google Scholar]
- Paulman, A.; Lichtensteiger, C.A.; Kohrt, L.J. Outbreak of Herpesviral Conjunctivitis and Respiratory Disease in Gouldian Finches. Vet. Pathol. 2006, 43, 963–970. [Google Scholar] [CrossRef]
- Wellehan, J.F.X.; Gagea, M.; Smith, D.A.; Taylor, W.M.; Berhane, Y.; Bienzle, D. Characterization of a Herpesvirus Associated with Tracheitis in Gouldian Finches (Erythrura [Chloebia] gouldiae). J. Clin. Microbiol. 2003, 41, 4054–4057. [Google Scholar] [CrossRef]
- Azab, W.; Dayaram, A.; Greenwood, A.D.; Osterrieder, N. How Host Specific Are Herpesviruses? Lessons from Herpesviruses Infecting Wild and Endangered Mammals. Annu. Rev. Virol. 2018, 5, 53–68. [Google Scholar] [CrossRef]
- Woźniakowski, G.J.; Samorek-Salamonowicz, E.; Szymański, P.; Wencel, P.; Houszka, M. Phylogenetic Analysis of Columbid Herpesvirus-1 in Rock Pigeons, Birds of Prey and Non-Raptorial Birds in Poland. BMC Vet. Res. 2013, 9, 52. [Google Scholar] [CrossRef] [PubMed]
- Smadel, J.E.; Jackson, E.B.; Harman, J.W. A new virus disease of pigeons: I. Recovery of the virus. J. Exp. Med. 1945, 81, 385–398. [Google Scholar] [CrossRef]
- Lawson, B.; Lachish, S.; Colvile, K.M.; Durrant, C.; Peck, K.M.; Toms, M.P.; Sheldon, B.C.; Cunningham, A.A. Emergence of a Novel Avian Pox Disease in British Tit Species. PLoS ONE 2012, 7, e40176. [Google Scholar] [CrossRef] [PubMed]
- Williams, R.A.J.; Benitez, L. Chapter 5: Avian Poxvirus. In Ecology of Wild Bird Diseases; Fereidouni, S., Ed.; CRC Press: Boca Raton, FL, USA, in press; pp. 154–176. ISBN 978-0-8153-7945-4.
- Manuel Medina, F.; Adolfo Ramírez, G.; Hernández, A. Avian Pox in White-Tailed Laurel-Pigeons from the Canary Islands. J. Wildl. Dis. 2004, 40, 351–355. [Google Scholar] [CrossRef]
- McDonald, S.E.; Lowenstine, L.J.; Ardans, A.A. Avian Pox in Blue-Fronted Amazon Parrots. J. Am. Vet. Med. Assoc. 1981, 179, 1218–1222. [Google Scholar] [PubMed]
- Van Riper, C., III; van Riper, S.G.; Hansen, W.R. Epizootiology and Effect of Avian Pox on Hawaiian Forest Birds. Auk 2002, 119, 929–942. [Google Scholar] [CrossRef]
- Davidson, W.R.; Kellogg, F.E.; Doster, G.L. An Epornitic of Avian Pox in Wild Bobwhite Quail. J. Wildl. Dis. 1980, 16, 293–298. [Google Scholar] [CrossRef]
- Tripathy, D.N.; Schnitzlein, W.M.; Morris, P.J.; Janssen, D.L.; Zuba, J.K.; Massey, G.; Atkinson, C.T. Characterization of Poxviruses from Forest Birds in Hawaii. J. Wildl. Dis. 2000, 36, 225–230. [Google Scholar] [CrossRef]
- Tripathy, D.N.; Reed, W.M. Pox. In Diseases of Poultry; Swayne, D., Ed.; Wiley-Blackwell: Ames, IA, USA, 2013; pp. 333–349. [Google Scholar]
- Yeo, G.; Wang, Y.; Chong, S.M.; Humaidi, M.; Lim, X.F.; Mailepessov, D.; Chan, S.; How, C.B.; Lin, Y.N.; Huangfu, T.; et al. Characterization of Fowlpox Virus in Chickens and Bird-Biting Mosquitoes: A Molecular Approach to Investigating Avipoxvirus Transmission. J. Gen. Virol. 2019, 100, 838–850. [Google Scholar] [CrossRef]
- Williams, R.A.J.; Truchado, D.A.; Benitez, L. A Review on the Prevalence of Poxvirus Disease in Free-Living and Captive Wild Birds. Microbiol. Res. 2021, 12, 403–418. [Google Scholar] [CrossRef]
- Gyuranecz, M.; Foster, J.T.; Dán, Á.; Ip, H.S.; Egstad, K.F.; Parker, P.G.; Higashiguchi, J.M.; Skinner, M.A.; Höfle, U.; Kreizinger, Z.; et al. Worldwide Phylogenetic Relationship of Avian Poxviruses. J. Virol. 2013, 87, 4938–4951. [Google Scholar] [CrossRef]
- Le Loc’h, G.; Ducatez, M.F.; Camus-Bouclainville, C.; Guérin, J.-L.; Bertagnoli, S. Diversity of Avipoxviruses in Captive-Bred Houbara Bustard. Vet. Res. 2014, 45, 98. [Google Scholar] [CrossRef][Green Version]
- MacDonald, A.M.; Gibson, D.J.; Barta, J.R.; Poulson, R.; Brown, J.D.; Allison, A.B.; Nemeth, N.M. Bayesian Phylogenetic Analysis of Avipoxviruses from North American Wild Birds Demonstrates New Insights into Host Specificity and Interspecies Transmission. Avian Dis. 2019, 63, 427–432. [Google Scholar] [CrossRef] [PubMed]
- Shivaprasad, H.L.; Hill, D.; Todd, D.; Smyth, J.A. Circovirus Infection in a Gouldian Finch (Chloebia gouldiae). Avian Pathol. 2004, 33, 525–529. [Google Scholar] [CrossRef] [PubMed]
- Joseph, H.M.; Ballmann, M.Z.; Garner, M.M.; Hanley, C.S.; Berlinski, R.; Erdélyi, K.; Childress, A.L.; Fish, S.S.; Harrach, B.; Wellehan, J.F.X. A Novel Siadenovirus Detected in the Kidneys and Liver of Gouldian Finches (Erythura gouldiae). Vet. Microbiol. 2014, 172, 35–43. [Google Scholar] [CrossRef] [PubMed]
- Phalen, D.N.; Agius, J.; Vaz, F.F.; Eden, J.-S.; Setyo, L.C.; Donahoe, S. A Survey of a Mixed Species Aviary Provides New Insights into the Pathogenicity, Diversity, Evolution, Host Range, and Distribution of Psittacine and Passerine Adenoviruses. Avian Pathol. 2019, 48, 437–443. [Google Scholar] [CrossRef]
- Rinder, M.; Schmitz, A.; Baas, N.; Korbel, R. Molecular Identification of Novel and Genetically Diverse Adenoviruses in Passeriform Birds. Virus Genes. 2020, 56, 316–324. [Google Scholar] [CrossRef]
- Kovács, E.R.; Jánoska, M.; Dán, A.; Harrach, B.; Benko, M. Recognition and Partial Genome Characterization by Non-Specific DNA Amplification and PCR of a New Siadenovirus Species in a Sample Originating from Parus major, a Great Tit. J. Virol. Methods 2010, 163, 262–268. [Google Scholar] [CrossRef]
- Vaz, F.F.; Raso, T.F.; Agius, J.E.; Hunt, T.; Leishman, A.; Eden, J.-S.; Phalen, D.N. Opportunistic Sampling of Wild Native and Invasive Birds Reveals a Rich Diversity of Adenoviruses in Australia. Virus Evol. 2020, 6, veaa024. [Google Scholar] [CrossRef]
- Harrach, B.; Tarján, Z.L.; Benkő, M. Adenoviruses across the Animal Kingdom: A Walk in the Zoo. FEBS Lett. 2019, 593, 3660–3673. [Google Scholar] [CrossRef]
- Athukorala, A.; Helbig, K.J.; Mcsharry, B.P.; Forwood, J.K.; Sarker, S. Adenoviruses in Avian Hosts: Recent Discoveries Shed New Light on Adenovirus Diversity and Evolution. Viruses 2022, 14, 1767. [Google Scholar] [CrossRef] [PubMed]
- Das, S.; Fearnside, K.; Sarker, S.; Forwood, J.K.; Raidal, S.R. A Novel Pathogenic Aviadenovirus from Red-Bellied Parrots (Poicephalus rufiventris) Unveils Deep Recombination Events among Avian Host Lineages. Virology 2017, 502, 188–197. [Google Scholar] [CrossRef]
- Schachner, A.; Matos, M.; Grafl, B.; Hess, M. Fowl Adenovirus-Induced Diseases and Strategies for Their Control—A Review on the Current Global Situation. Avian Pathol. 2018, 47, 111–126. [Google Scholar] [CrossRef] [PubMed]
- Blackmore, D.K.; Keymer, I.F. Cutaneous Diseases of Wild Birds in Britain. Br. Birds 1969, 62, 316–331. [Google Scholar]
- Keymer, I.F.; Blackmore, D.K. Diseases of the Skin and Soft Parts of Wild Birds. Br. Birds 1964, 57, 175–179. [Google Scholar]
- Lina, P.H.; van Noord, M.J.; de Groot, F.G. Detection of Virus in Squamous Papillomas of the Wild Bird Species Fringilla coelebs. J. Natl. Cancer Inst. 1973, 50, 567–571. [Google Scholar] [CrossRef]
- Truchado, D.A.; Moens, M.A.J.; Callejas, S.; Pérez-Tris, J.; Benítez, L. Genomic Characterization of the First Oral Avian Papillomavirus in a Colony of Breeding Canaries (Serinus canaria). Vet. Res. Commun. 2018, 42, 111–120. [Google Scholar] [CrossRef]
- Dom, P.; Ducatelle, R.; Charlier, G.; de Groot, P. Papillomavirus-like Infections in Canaries (Serinus canarius). Avian Pathol. 1993, 22, 797–803. [Google Scholar] [CrossRef][Green Version]
- Literak, I.; Smid, B.; Dusbabek, F.; Halouzka, R.; Novotny, L. Co-Infection with Papillomavirus and Knemidokoptes Jamaicensis (Acari: Knemidokoptidae) in a Chaffinch (Fringilla coelebs) and a Case of Beak Papillomatosis in Another Chaffinch. Vet. Med. 2005, 50, 276–280. [Google Scholar] [CrossRef]
- Osterhaus, A.D.; Ellens, D.J.; Horzinek, M.C. Identification and Characterization of a Papillomavirus from Birds (Fringillidae). Intervirology 1977, 8, 351–359. [Google Scholar] [CrossRef]
- Prosperi, A.; Chiari, M.; Zanoni, M.; Gallina, L.; Casà, G.; Scagliarini, A.; Lavazza, A. Identification and Characterization of Fringilla coelebs Papillomavirus 1 (FcPV1) in Free-Living and Captive Birds in Italy. J. Wildl. Dis. 2016, 52, 756–758. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sironi, G.; Gallazzi, D. Papillomavirus Infection in Greenfinches (Carduelis chloris). J. Vet. Med. 1992, 39, 454–458. [Google Scholar] [CrossRef] [PubMed]
- Terai, M.; DeSalle, R.; Burk, R.D. Lack of Canonical E6 and E7 Open Reading Frames in Bird Papillomaviruses: Fringilla coelebs Papillomavirus and Psittacus erithacus timneh Papillomavirus. J. Virol. 2002, 76, 10020–10023. [Google Scholar] [CrossRef]
- Katoh, H.; Ogawa, H.; Ohya, K.; Fukushi, H. A Review of DNA Viral Infections in Psittacine Birds. J. Vet. Med. Sci. 2010, 72, 1099–1106. [Google Scholar] [CrossRef] [PubMed]
- Amery-Gale, J.; Marenda, M.S.; Owens, J.; Eden, P.A.; Browning, G.F.; Devlin, J.M. A High Prevalence of Beak and Feather Disease Virus in Non-Psittacine Australian Birds. J. Med. Microbiol. 2017, 66, 1005–1013. [Google Scholar] [CrossRef] [PubMed]
- Circella, E.; Legretto, M.; Pugliese, N.; Caroli, A.; Bozzo, G.; Accogli, G.; Lavazza, A.; Camarda, A. Psittacine Beak and Feather Disease-like Illness in Gouldian Finches (Chloebia gouldiae). Avian Dis. 2014, 58, 482–487. [Google Scholar] [CrossRef]
- Ahaduzzaman, M.; Nath, C.; Hossain, M.S. Evidence of Circulation of Beak and Feather Disease Virus in Captive Psittacine and Non-Psittacine Birds in Bangladesh. Arch. Virol. 2022, 167, 2567–2575. [Google Scholar] [CrossRef]
- Phenix, K.V.; Weston, J.H.; Ypelaar, I.; Lavazza, A.; Smyth, J.A.; Todd, D.; Wilcox, G.E.; Raidal, S.R. Nucleotide Sequence Analysis of a Novel Circovirus of Canaries and Its Relationship to Other Members of the Genus Circovirus of the Family Circoviridae. J. Gen. Virol. 2001, 82, 2805–2809. [Google Scholar] [CrossRef]
- Rinder, M.; Schmitz, A.; Peschel, A.; Wörle, B.; Gerlach, H.; Korbel, R. Molecular Characterization of a Recently Identified Circovirus in Zebra Finches (Taeniopygia guttata) Associated with Immunosuppression and Opportunistic Infections. Avian Pathol. 2017, 46, 106–116. [Google Scholar] [CrossRef]
- Stewart, M.E.; Perry, R.; Raidal, S.R. Identification of a Novel Circovirus in Australian Ravens (Corvus coronoides) with Feather Disease. Avian Pathol. 2006, 35, 86–92. [Google Scholar] [CrossRef]
- Todd, D.; Scott, A.N.J.; Fringuelli, E.; Shivraprasad, H.L.; Gavier-Widen, D.; Smyth, J.A. Molecular Characterization of Novel Circoviruses from Finch and Gull. Avian Pathol. 2007, 36, 75–81. [Google Scholar] [CrossRef] [PubMed]
- Goldsmith, T.L. Documentation of Passerine Circoviral Infection; Association of Avian Vets (AAV): Philadelphia, PA, USA, 1995; Volume 43, pp. 349–350. [Google Scholar]
- Rosario, K.; Duffy, S.; Breitbart, M. A Field Guide to Eukaryotic Circular Single-Stranded DNA Viruses: Insights Gained from Metagenomics. Arch. Virol. 2012, 157, 1851–1871. [Google Scholar] [CrossRef] [PubMed]
- Hanna, Z.R.; Runckel, C.; Fuchs, J.; DeRisi, J.L.; Mindell, D.P.; Van Hemert, C.; Handel, C.M.; Dumbacher, J.P. Isolation of a Complete Circular Virus Genome Sequence from an Alaskan Black-Capped Chickadee (Poecile atricapillus) Gastrointestinal Tract Sample. Genome Announc. 2015, 3, e01081-e15. [Google Scholar] [CrossRef]
- Schmidlin, K.; Sepp, T.; Khalifeh, A.; Smith, K.; Fontenele, R.S.; McGraw, K.J.; Varsani, A. Diverse Genomoviruses Representing Eight New and One Known Species Identified in Feces and Nests of House Finches (Haemorhous mexicanus). Arch. Virol. 2019, 164, 2345–2350. [Google Scholar] [CrossRef] [PubMed]
- Sikorski, A.; Massaro, M.; Kraberger, S.; Young, L.M.; Smalley, D.; Martin, D.P.; Varsani, A. Novel Myco-like DNA Viruses Discovered in the Faecal Matter of Various Animals. Virus Res. 2013, 177, 209–216. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Wu, H.; Sun, G.; Yang, S.; Shen, Q.; Wang, X.; Zhang, W. Identification of Diverse Novel Genomoviruses in Gut of Wild Birds. Biosaf. Health 2021, 3, 136–141. [Google Scholar] [CrossRef]
- Truchado, D.A.; Diaz-Piqueras, J.M.; Gomez-Lucia, E.; Doménech, A.; Milá, B.; Pérez-Tris, J.; Schmidt-Chanasit, J.; Cadar, D.; Benítez, L. A Novel and Divergent Gyrovirus with Unusual Genomic Features Detected in Wild Passerine Birds from a Remote Rainforest in French Guiana. Viruses 2019, 11, 1148. [Google Scholar] [CrossRef]
- Pénzes, J.J.; Söderlund-Venermo, M.; Canuti, M.; Eis-Hübinger, A.M.; Hughes, J.; Cotmore, S.F.; Harrach, B. Reorganizing the Family Parvoviridae: A Revised Taxonomy Independent of the Canonical Approach Based on Host Association. Arch. Virol. 2020, 165, 2133–2146. [Google Scholar] [CrossRef]
- Cotmore, S.F.; Tattersall, P. Parvovirus Diversity and DNA Damage Responses. Cold Spring Harb. Perspect. Biol. 2013, 5, a012989. [Google Scholar] [CrossRef]
- Hueffer, K.; Parrish, C.R. Parvovirus Host Range, Cell Tropism and Evolution. Curr. Opin. Microbiol. 2003, 6, 392–398. [Google Scholar] [CrossRef]
- Dong, H.V.; Tran, G.T.H.; Nguyen, H.T.T.; Nguyen, T.M.; Trinh, D.Q.; Le, V.P.; Choowongkomon, K.; Rattanasrisomporn, J. Epidemiological Analysis and Genetic Characterization of Parvovirus in Ducks in Northern Vietnam Reveal Evidence of Recombination. Animals 2022, 12, 2846. [Google Scholar] [CrossRef] [PubMed]
- Jager, M.C.; Tomlinson, J.E.; Lopez-Astacio, R.A.; Parrish, C.R.; Van de Walle, G.R. Small but Mighty: Old and New Parvoviruses of Veterinary Significance. Virol. J. 2021, 18, 210. [Google Scholar] [CrossRef] [PubMed]
- De Souza, W.M.; Dennis, T.; Fumagalli, M.J.; Araujo, J.; Sabino-Santos, G.; Maia, F.G.M.; Acrani, G.O.; Carrasco, A.D.O.T.; Romeiro, M.F.; Modha, S.; et al. Novel Parvoviruses from Wild and Domestic Animals in Brazil Provide New Insights into Parvovirus Distribution and Diversity. Viruses 2018, 10, 143. [Google Scholar] [CrossRef] [PubMed]
- Webster, R.G.; Bean, W.J.; Gorman, O.T.; Chambers, T.M.; Kawaoka, Y. Evolution and Ecology of Influenza A Viruses. Microbiol. Rev. 1992, 56, 152–179. [Google Scholar] [CrossRef]
- Krammer, F.; Smith, G.J.D.; Fouchier, R.A.M.; Peiris, M.; Kedzierska, K.; Doherty, P.C.; Palese, P.; Shaw, M.L.; Treanor, J.; Webster, R.G.; et al. Influenza. Nat. Rev. Dis. Primers 2018, 4, 3. [Google Scholar] [CrossRef]
- Alexander, D.J. An Overview of the Epidemiology of Avian Influenza. Vaccine 2007, 25, 5637–5644. [Google Scholar] [CrossRef]
- Seeger, R.M.; Hagerman, A.D.; Johnson, K.K.; Pendell, D.L.; Marsh, T.L. When Poultry Take a Sick Leave: Response Costs for the 2014–2015 Highly Pathogenic Avian Influenza Epidemic in the USA. Food Policy 2021, 102, 102068. [Google Scholar] [CrossRef]
- Bahl, J.; Pham, T.T.; Hill, N.J.; Hussein, I.T.M.; Ma, E.J.; Easterday, B.C.; Halpin, R.A.; Stockwell, T.B.; Wentworth, D.E.; Kayali, G.; et al. Ecosystem Interactions Underlie the Spread of Avian Influenza A Viruses with Pandemic Potential. PLoS Pathog. 2016, 12, e1005620. [Google Scholar] [CrossRef]
- Fouchier, R.A.; Bestebroer, T.M.; Herfst, S.; Van Der Kemp, L.; Rimmelzwaan, G.F.; Osterhaus, A.D. Detection of Influenza A Viruses from Different Species by PCR Amplification of Conserved Sequences in the Matrix Gene. J. Clin. Microbiol. 2000, 38, 4096–4101. [Google Scholar] [CrossRef]
- Morishita, T.Y.; Aye, P.P.; Ley, E.C.; Harr, B.S. Survey of Pathogens and Blood Parasites in Free-Living Passerines. Avian Dis. 1999, 43, 549–552. [Google Scholar] [CrossRef]
- Munster, V.J.; Baas, C.; Lexmond, P.; Waldenström, J.; Wallensten, A.; Fransson, T.; Rimmelzwaan, G.F.; Beyer, W.E.P.; Schutten, M.; Olsen, B.; et al. Spatial, Temporal, and Species Variation in Prevalence of Influenza A Viruses in Wild Migratory Birds. PLoS Pathog. 2007, 3, e61. [Google Scholar] [CrossRef]
- Swayne, D.E.; Suarez, D.L. Highly Pathogenic Avian Influenza. Rev. Sci. Tech. 2000, 19, 463–482. [Google Scholar] [CrossRef]
- Chang, H.; Dai, F.; Liu, Z.; Yuan, F.; Zhao, S.; Xiang, X.; Zou, F.; Zeng, B.; Fan, Y.; Duan, G. Seroprevalence Survey of Avian Influenza A (H5) in Wild Migratory Birds in Yunnan Province, Southwestern China. Virol. J. 2014, 11, 18. [Google Scholar] [CrossRef] [PubMed]
- Fuller, T.L.; Saatchi, S.S.; Curd, E.E.; Toffelmier, E.; Thomassen, H.A.; Buermann, W.; DeSante, D.F.; Nott, M.P.; Saracco, J.F.; Ralph, C.; et al. Mapping the Risk of Avian Influenza in Wild Birds in the US. BMC Infect. Dis. 2010, 10, 187. [Google Scholar] [CrossRef] [PubMed]
- Kou, Z.; Lei, F.M.; Yu, J.; Fan, Z.J.; Yin, Z.H.; Jia, C.X.; Xiong, K.J.; Sun, Y.H.; Zhang, X.W.; Wu, X.M.; et al. New Genotype of Avian Influenza H5N1 Viruses Isolated from Tree Sparrows in China. J. Virol. 2005, 79, 15460–15466. [Google Scholar] [CrossRef] [PubMed]
- Peterson, A.T.; Bush, S.E.; Spackman, E.; Swayne, D.E.; Ip, H.S. Influenza A Virus Infections in Land Birds, People’s Republic of China. Emerg. Infect. Dis. 2008, 14, 1644–1646. [Google Scholar] [CrossRef]
- Günther, A.; Pohlmann, A.; Globig, A.; Ziegler, U.; Calvelage, S.; Keller, M.; Fischer, D.; Staubach, C.; Groschup, M.H.; Harder, T.; et al. Continuous Surveillance of Potentially Zoonotic Avian Pathogens Detects Contemporaneous Occurrence of Highly Pathogenic Avian Influenza Viruses (HPAIV H5) and Flaviviruses (USUV, WNV) in Several Wild and Captive Birds. Emerg. Microbes Infect. 2023, 12, 2231561. [Google Scholar] [CrossRef]
- Shriner, S.A.; Root, J.J. A Review of Avian Influenza A Virus Associations in Synanthropic Birds. Viruses 2020, 12, 1209. [Google Scholar] [CrossRef]
- Fujimoto, Y.; Ito, H.; Shinya, K.; Yamaguchi, T.; Usui, T.; Murase, T.; Ozaki, H.; Ono, E.; Takakuwa, H.; Otsuki, K.; et al. Susceptibility of Two Species of Wild Terrestrial Birds to Infection with a Highly Pathogenic Avian Influenza Virus of H5N1 Subtype. Avian Pathol. 2010, 39, 95–98. [Google Scholar] [CrossRef]
- Root, J.J.; Bosco-Lauth, A.M.; Marlenee, N.L.; Bowen, R.A. Viral Shedding of Clade 2.3.4.4 H5 Highly Pathogenic Avian Influenza A Viruses by American Robins. Transbound. Emerg. Dis. 2018, 65, 1823–1827. [Google Scholar] [CrossRef]
- Schnebel, B.; Dierschke, V.; Rautenschlein, S.; Ryll, M.; Neumann, U. Investigations on Infection Status with H5 and H7 Avian Influenza Virus in Short-Distance and Long-Distance Migrant Birds in 2001. Avian Dis. 2007, 51, 432–433. [Google Scholar] [CrossRef]
- Caron, A.; Grosbois, V.; Etter, E.; Gaidet, N.; de Garine-Wichatitsky, M. Bridge Hosts for Avian Influenza Viruses at the Wildlife/Domestic Interface: An Eco-Epidemiological Framework Implemented in Southern Africa. Prev. Vet. Med. 2014, 117, 590–600. [Google Scholar] [CrossRef] [PubMed]
- Breithaupt, A.; Kalthoff, D.; Dale, J.; Bairlein, F.; Beer, M.; Teifke, J.P. Neurotropism in Blackcaps (Sylvia Atricapilla) and Red-Billed Queleas (Quelea quelea) after Highly Pathogenic Avian Influenza Virus H5N1 Infection. Vet. Pathol. 2011, 48, 924–932. [Google Scholar] [CrossRef] [PubMed]
- European Food Safety Authority; European Centre for Disease Prevention and Control; European Union Reference Laboratory for Avian Influenza; Adlhoch, C.; Fusaro, A.; Gonzales, J.L.; Kuiken, T.; Marangon, S.; Niqueux, É.; Staubach, C.; et al. Avian Influenza Overview June–September 2022. EFS2 2022, 20, e7597. [Google Scholar] [CrossRef]
- Animal and Plant Health Inspection Service. 2022–2023 Detections of Highly Pathogenic Avian Influenza in Wild Birds. Available online: https://www.aphis.usda.gov/aphis/ourfocus/animalhealth/animal-disease-information/avian/avian-influenza/hpai-2022/2022-hpai-wild-birds (accessed on 19 July 2023).
- Perkins, L.E.L.; Swayne, D.E. Comparative Susceptibility of Selected Avian and Mammalian Species to a Hong Kong-Origin H5N1 High-Pathogenicity Avian Influenza Virus. Avian Dis. 2003, 47, 956–967. [Google Scholar] [CrossRef] [PubMed]
- Thomazelli, L.M.; Sinhorini, J.A.; Oliveira, D.B.L.; Knöbl, T.; Bosqueiro, T.C.M.; Sano, E.; Costa, G.C.V.; Monteiro, C.; Dorlass, E.G.; Utecht, N.; et al. An Outbreak in Pigeons Caused by the Subgenotype VI.2.1.2 of Newcastle Disease Virus in Brazil. Viruses 2021, 13, 2446. [Google Scholar] [CrossRef]
- Wajid, A.; Mayahi, V.; Yin, R.; Ain, Q.; Mohiuddin, A.; Khalid, F.; Rehim, A.; Manan, A.; Baksh, M. Genomic and Biological Characteristics of Avian Orthoavulavirus-1 Strains Isolated from Multiple Wild Birds and Backyard Chickens in Pakistan. Trop. Anim. Health Prod. 2021, 53, 90. [Google Scholar] [CrossRef]
- Karamendin, K.; Kydyrmanov, A. Cormorants as a Potentially Important Reservoir and Carrier of Newcastle Disease Virus on the Asian Continent. Front. Vet. Sci. 2021, 8, 648091. [Google Scholar] [CrossRef]
- Kuiken, T.; Wobeser, G.; Leighton, F.A.; Haines, D.M.; Chelack, B.; Bogdan, J.; Hassard, L.; Heckert, R.A.; Riva, J. Pathology of Newcastle Disease in Double-Crested Cormorants from Saskatchewan, with Comparison of Diagnostic Methods. J. Wildl. Dis. 1999, 35, 8–23. [Google Scholar] [CrossRef]
- Turan, N.; Ozsemir, C.; Yilmaz, A.; Cizmecigil, U.Y.; Aydin, O.; Bamac, O.E.; Gurel, A.; Kutukcu, A.; Ozsemir, K.; Tali, H.E.; et al. Identification of Newcastle Disease Virus Subgenotype VII.2 in Wild Birds in Turkey. BMC Vet. Res. 2020, 16, 277. [Google Scholar] [CrossRef]
- Silva, B.R.; Gamon, T.H.; Campos, A.C.A.; Thomazelli, L.M.; Serafini, P.P.; Chiarani, E.; Silva, T.W.; Locatelli-Dittrich, R. Molecular Diagnosis of Avian Viruses in Grassland Passerines and Captive Yellow Cardinals Gubernatrix cristata in Brazil. Pesq. Vet. Bras. 2021, 41, e06840. [Google Scholar] [CrossRef]
- Pedersen, K.; Marks, D.R.; Afonso, C.L.; Stopak, S.R.; Williams-Coplin, D.; Dimitrov, K.M.; Miller, P.J.; DeLiberto, T.J. Identification of Avian Paramyxovirus Serotype-1 in Wild Birds in the USA. J. Wildl. Dis. 2016, 52, 657–662. [Google Scholar] [CrossRef]
- Orynbayev, M.B.; Fereidouni, S.; Sansyzbai, A.R.; Seidakhmetova, B.A.; Strochkov, V.M.; Nametov, A.M.; Sadikaliyeva, S.O.; Nurgazieva, A.; Tabynov, K.K.; Rametov, N.M.; et al. Genetic Diversity of Avian Avulavirus 1 (Newcastle Disease Virus Genotypes VIg and VIIb) Circulating in Wild Birds in Kazakhstan. Arch. Virol. 2018, 163, 1949–1954. [Google Scholar] [CrossRef] [PubMed]
- Kariithi, H.M.; Christy, N.; Decanini, E.L.; Lemiere, S.; Volkening, J.D.; Afonso, C.L.; Suarez, D.L. Detection and Genome Sequence Analysis of Avian Metapneumovirus Subtype a Viruses Circulating in Commercial Chicken Flocks in Mexico. Vet. Sci. 2022, 9, 579. [Google Scholar] [CrossRef]
- Greenacre, C.B. Viral Diseases of Companion Birds. Vet. Clin. N. Am. Exot. Anim. Pract. 2005, 8, 85–105. [Google Scholar] [CrossRef]
- Fleury, H.J.; Alexander, D.J. Isolation of Twenty-Three Yucaipa-like Viruses from 616 Wild Birds in Senegal, West Africa. Avian Dis. 1979, 23, 742–744. [Google Scholar] [CrossRef]
- Mbugua, H.C.; Karstad, L.; Thorsen, J. Isolation of Avian Paramyxoviruses (Yucaipa-like) from Wild Birds in Kenya, 1980–1982. J. Wildl. Dis. 1985, 21, 52–54. [Google Scholar] [CrossRef][Green Version]
- Nymadava, P.; Konstantinow-Siebelist, I.; Schulze, P.; Starke, S. Isolation of Paramyxoviruses from Free-Flying Birds of the Order Passeriformes in the German Democratic Republic. Acta Virol. 1977, 21, 443. [Google Scholar]
- Maldonado, A.; Arenas, A.; Tarradas, M.C.; Luque, I.; Astorga, R.; Perea, J.A.; Miranda, A. Serological Survey for Avian Paramyxoviruses from Wildfowl in Aquatic Habitats in Andalusia. J. Wildl. Dis. 1995, 31, 66–69. [Google Scholar] [CrossRef]
- Goodman, B.B.; Hanson, R.P. Isolation of Avian Paramyxovirus-2 from Domestic and Wild Birds in Costa Rica. Avian Dis. 1988, 32, 713–717. [Google Scholar] [CrossRef]
- Račnik, J.; Slavec, B.; Trilar, T.; Zadravec, M.; Dovč, A.; Krapež, U.; Barlič-Maganja, D.; Zorman Rojs, O. Evidence of Avian Influenza Virus and Paramyxovirus Subtype 2 in Wild-Living Passerine Birds in Slovenia. Eur. J. Wildl. Res. 2008, 54, 529–532. [Google Scholar] [CrossRef]
- Schemera, B.; Toro, H.; Kaleta, E.F.; Herbst, W. A Paramyxovirus of Serotype 3 Isolated from African and Australian Finches. Avian Dis. 1987, 31, 921–925. [Google Scholar] [CrossRef] [PubMed]
- Shihmanter, E.; Weisman, Y.; Lublin, A.; Mahani, S.; Panshin, A.; Lipkind, M. Isolation of Avian Serotype 3 Paramyxoviruses from Imported Caged Birds in Israel. Avian Dis. 1998, 42, 829–831. [Google Scholar] [CrossRef] [PubMed]
- Yin, R.; Zhang, P.; Liu, X.; Chen, Y.; Tao, Z.; Ai, L.; Li, J.; Yang, Y.; Li, M.; Xue, C.; et al. Dispersal and Transmission of Avian Paramyxovirus Serotype 4 among Wild Birds and Domestic Poultry. Front. Cell Infect. Microbiol. 2017, 7, 212. [Google Scholar] [CrossRef]
- Muzyka, D.; Pantin-Jackwood, M.; Stegniy, B.; Rula, O.; Bolotin, V.; Stegniy, A.; Gerilovych, A.; Shutchenko, P.; Stegniy, M.; Koshelev, V.; et al. Wild Bird Surveillance for Avian Paramyxoviruses in the Azov-Black Sea Region of Ukraine (2006 to 2011) Reveals Epidemiological Connections with Europe and Africa. Appl. Environ. Microbiol. 2014, 80, 5427–5438. [Google Scholar] [CrossRef]
- Rautenschlein, S. Avian Metapneumovirus. In Diseases of Poultry; Swayne, D., Boulianne, M., Logue, C., McDougald, L., Nair, V., Suarez, D., Eds.; Wiley-Blackwell: Hoboken, NJ, USA, 2020; pp. 135–143. [Google Scholar]
- Graziosi, G.; Lupini, C.; Catelli, E. Disentangling the Role of Wild Birds in Avian Metapneumovirus (aMPV) Epidemiology: A Systematic Review and Meta-Analysis. Transbound. Emerg. Dis. 2022, 69, 3285–3299. [Google Scholar] [CrossRef]
- Bennett, R.S.; Nezworski, J.; Velayudhan, B.T.; Nagaraja, K.V.; Zeman, D.H.; Dyer, N.; Graham, T.; Lauer, D.C.; Njenga, M.K.; Halvorson, D.A. Evidence of Avian Pneumovirus Spread beyond Minnesota among Wild and Domestic Birds in Central North America. Avian Dis. 2004, 48, 902–908. [Google Scholar] [CrossRef]
- Shin, H.-J.; Njenga, M.K.; McComb, B.; Halvorson, D.A.; Nagaraja, K.V. Avian Pneumovirus (APV) RNA from Wild and Sentinel Birds in the United States Has Genetic Homology with RNA from APV Isolates from Domestic Turkeys. J. Clin. Microbiol. 2000, 38, 4282–4284. [Google Scholar] [CrossRef]
- Gharaibeh, S.; Shamoun, M. Avian Metapneumovirus Subtype B Experimental Infection and Tissue Distribution in Chickens, Sparrows, and Pigeons. Vet. Pathol. 2012, 49, 704–709. [Google Scholar] [CrossRef]
- Honkavuori, K.S.; Shivaprasad, H.L.; Williams, B.L.; Quan, P.-L.; Hornig, M.; Street, C.; Palacios, G.; Hutchison, S.K.; Franca, M.; Egholm, M.; et al. Novel Borna Virus in Psittacine Birds with Proventricular Dilatation Disease. Emerg. Infect. Dis. 2008, 14, 1883–1886. [Google Scholar] [CrossRef]
- Kistler, A.L.; Gancz, A.; Clubb, S.; Skewes-Cox, P.; Fischer, K.; Sorber, K.; Chiu, C.Y.; Lublin, A.; Mechani, S.; Farnoushi, Y.; et al. Recovery of Divergent Avian Bornaviruses from Cases of Proventricular Dilatation Disease: Identification of a Candidate Etiologic Agent. Virol. J. 2008, 5, 88. [Google Scholar] [CrossRef]
- Berg, M.; Johansson, M.; Montell, H.; Berg, A.L. Wild Birds as a Possible Natural Reservoir of Borna Disease Virus. Epidemiol. Infect. 2001, 127, 173–178. [Google Scholar] [CrossRef] [PubMed]
- Rubbenstroth, D.; Rinder, M.; Stein, M.; Höper, D.; Kaspers, B.; Brosinski, K.; Horie, M.; Schmidt, V.; Legler, M.; Korbel, R.; et al. Avian Bornaviruses Are Widely Distributed in Canary Birds (Serinus canaria f. domestica). Vet. Microbiol. 2013, 165, 287–295. [Google Scholar] [CrossRef] [PubMed]
- Rubbenstroth, D.; Schmidt, V.; Rinder, M.; Legler, M.; Corman, V.M.; Staeheli, P. Discovery of a New Avian Bornavirus Genotype in Estrildid Finches (Estrildidae) in Germany. Vet. Microbiol. 2014, 168, 318–323. [Google Scholar] [CrossRef]
- Kato, M.; Okanoya, K. Molecular Characterization of the Song Control Nucleus HVC in Bengalese Finch Brain. Brain Res. 2010, 1360, 56–76. [Google Scholar] [CrossRef] [PubMed]
- Fall, G.; Di Paola, N.; Faye, M.; Dia, M.; Freire, C.C.D.M.; Loucoubar, C.; Zanotto, P.M.D.A.; Faye, O.; Sall, A.A. Biological and Phylogenetic Characteristics of West African Lineages of West Nile Virus. PLoS Negl. Trop. Dis. 2017, 11, e0006078. [Google Scholar] [CrossRef]
- Scherret, J.H.; Poidinger, M.; Mackenzie, J.S.; Broom, A.K.; Deubel, V.; Lipkin, W.I.; Briese, T.; Gould, E.A.; Hall, R.A. The Relationships between West Nile and Kunjin Viruses. Emerg. Infect. Dis. 2001, 7, 697–705. [Google Scholar] [CrossRef]
- Komar, N.; Langevin, S.; Hinten, S.; Nemeth, N.; Edwards, E.; Hettler, D.; Davis, B.; Bowen, R.; Bunning, M. Experimental Infection of North. American Birds with the New York 1999 Strain of West Nile Virus. Emerg. Infect. Dis. 2003, 9, 311–322. [Google Scholar] [CrossRef]
- Smithburn, K.C.; Hughes, T.P.; Burke, A.W.; Paul, J.H. A Neurotropic Virus Isolated from the Blood of a Native of Uganda. Am. J. Trop. Med. Hyg. 1940, 20, 471–492. [Google Scholar] [CrossRef]
- Campbell, G.L.; Ceianu, C.S.; Savage, H.M. Epidemic West Nile Encephalitis in Romania: Waiting for History to Repeat Itself. Ann. NY Acad. Sci. 2001, 951, 94–101. [Google Scholar] [CrossRef]
- Kramer, L.D.; Ciota, A.T.; Kilpatrick, A.M. Introduction, Spread, and Establishment of West Nile Virus in the Americas. J. Med. Entomol. 2019, 56, 1448–1455. [Google Scholar] [CrossRef] [PubMed]
- Bakonyi, T.; Haussig, J.M. West Nile Virus Keeps on Moving up in Europe. Euro Surveill. 2020, 25, 2001938. [Google Scholar] [CrossRef] [PubMed]
- Zehender, G.; Veo, C.; Ebranati, E.; Carta, V.; Rovida, F.; Percivalle, E.; Moreno, A.; Lelli, D.; Calzolari, M.; Lavazza, A.; et al. Reconstructing the Recent West Nile Virus Lineage 2 Epidemic in Europe and Italy Using Discrete and Continuous Phylogeography. PLoS ONE 2017, 12, e0179679. [Google Scholar] [CrossRef] [PubMed]
- Bernard, K.A.; Maffei, J.G.; Jones, S.A.; Kauffman, E.B.; Ebel, G.; Dupuis, A.P.; Ngo, K.A.; Nicholas, D.C.; Young, D.M.; Shi, P.Y.; et al. West Nile Virus Infection in Birds and Mosquitoes, New York State, 2000. Emerg. Infect. Dis. 2001, 7, 679–685. [Google Scholar] [CrossRef]
- Bakonyi, T.; Ferenczi, E.; Erdélyi, K.; Kutasi, O.; Csörgő, T.; Seidel, B.; Weissenböck, H.; Brugger, K.; Bán, E.; Nowotny, N. Explosive Spread of a Neuroinvasive Lineage 2 West Nile Virus in Central Europe, 2008/2009. Vet. Microbiol. 2013, 165, 61–70. [Google Scholar] [CrossRef]
- Jourdain, E.; Olsen, B.; Lundkvist, A.; Hubálek, Z.; Sikutová, S.; Waldenström, J.; Karlsson, M.; Wahlström, M.; Jozan, M.; Falk, K.I. Surveillance for West Nile Virus in Wild Birds from Northern Europe. Vector Borne Zoonotic Dis. 2011, 11, 77–79. [Google Scholar] [CrossRef]
- López, G.; Jiménez-Clavero, M.A.; Tejedor, C.G.; Soriguer, R.; Figuerola, J. Prevalence of West Nile Virus Neutralizing Antibodies in Spain Is Related to the Behavior of Migratory Birds. Vector Borne Zoonotic Dis. 2008, 8, 615–621. [Google Scholar] [CrossRef]
- Drennan, J.E. Northern Goshawk Food Habits and Goshawk Prey Species Habitats|Searchable Ornithological Research Archive. Stud. Avian Biol. 2006, 31, 198–218. [Google Scholar]
- Pérez-Ramírez, E.; Llorente, F.; Jiménez-Clavero, M.Á. Experimental Infections of Wild Birds with West Nile Virus. Viruses 2014, 6, 752–781. [Google Scholar] [CrossRef]
- Brault, A.C.; Langevin, S.A.; Ramey, W.N.; Fang, Y.; Beasley, D.W.C.; Barker, C.M.; Sanders, T.A.; Reisen, W.K.; Barrett, A.D.T.; Bowen, R.A. Reduced Avian Virulence and Viremia of West Nile Virus Isolates from Mexico and Texas. Am. J. Trop. Med. Hyg. 2011, 85, 758–767. [Google Scholar] [CrossRef]
- Del Amo, J.; Llorente, F.; Figuerola, J.; Soriguer, R.C.; Moreno, A.M.; Cordioli, P.; Weissenböck, H.; Jiménez-Clavero, M.Á. Experimental Infection of House Sparrows (Passer domesticus) with West Nile Virus Isolates of Euro-Mediterranean and North American Origins. Vet. Res. 2014, 45, 33. [Google Scholar] [CrossRef]
- Del Amo, J.; Llorente, F.; Pérez-Ramirez, E.; Soriguer, R.C.; Figuerola, J.; Nowotny, N.; Jiménez-Clavero, M.A. Experimental Infection of House Sparrows (Passer domesticus) with West Nile Virus Strains of Lineages 1 and 2. Vet. Microbiol. 2014, 172, 542–547. [Google Scholar] [CrossRef] [PubMed]
- Langevin, S.A.; Brault, A.C.; Panella, N.A.; Bowen, R.A.; Komar, N. Variation in Virulence of West Nile Virus Strains for House Sparrows (Passer domesticus). Am. J. Trop. Med. Hyg. 2005, 72, 99–102. [Google Scholar] [CrossRef] [PubMed]
- Work, T.H.; Hurlbut, H.S.; Taylor, R.M. Indigenous Wild Birds of the Nile Delta as Potential West Nile Virus Circulating Reservoirs. Am. J. Trop. Med. Hyg. 1955, 4, 872–888. [Google Scholar] [CrossRef]
- Centers for Disease Control and Prevention. Species of Dead Birds in Which West Nile Virus Has Been Detected, United States, 1999–2016. Available online: https://www.cdc.gov/westnile/resources/pdfs/birdspecies1999-2016.pdf (accessed on 27 June 2023).
- Williams, M.C.; Simpson, D.I.; Haddow, A.J.; Knight, E.M. The Isolation of West Nile Virus from Man and of Usutu Virus Brom He Bird-Bitting Mosquito Mansonia aurites (Theobald) in the Entebbe Area of Uganda. Ann. Trop. Med. Parasitol. 1964, 58, 367–374. [Google Scholar] [CrossRef]
- Weissenböck, H.; Kolodziejek, J.; Fragner, K.; Kuhn, R.; Pfeffer, M.; Nowotny, N. Usutu Virus Activity in Austria, 2001–2002. Microbes Infect. 2003, 5, 1132–1136. [Google Scholar] [CrossRef]
- Vilibic-Cavlek, T.; Petrovic, T.; Savic, V.; Barbic, L.; Tabain, I.; Stevanovic, V.; Klobucar, A.; Mrzljak, A.; Ilic, M.; Bogdanic, M.; et al. Epidemiology of Usutu Virus: The European Scenario. Pathogens 2020, 9, 699. [Google Scholar] [CrossRef]
- Engel, D.; Jöst, H.; Wink, M.; Börstler, J.; Bosch, S.; Garigliany, M.-M.; Jöst, A.; Czajka, C.; Lühken, R.; Ziegler, U.; et al. Reconstruction of the Evolutionary History and Dispersal of Usutu Virus, a Neglected Emerging Arbovirus in Europe and Africa. mBio 2016, 7, e01938-15. [Google Scholar] [CrossRef]
- Chevalier, V.; Marsot, M.; Molia, S.; Rasamoelina, H.; Rakotondravao, R.; Pedrono, M.; Lowenski, S.; Durand, B.; Lecollinet, S.; Beck, C. Serological Evidence of West. Nile and Usutu Viruses Circulation in Domestic and Wild Birds in Wetlands of Mali. and Madagascar in 2008. Int. J. Environ. Res. Public Health 2020, 17, 1998. [Google Scholar] [CrossRef]
- Nikolay, B.; Diallo, M.; Boye, C.S.B.; Sall, A.A. Usutu Virus in Africa. Vector Borne Zoonotic Dis. 2011, 11, 1417–1423. [Google Scholar] [CrossRef]
- Chvala, S.; Bakonyi, T.; Bukovsky, C.; Meister, T.; Brugger, K.; Rubel, F.; Nowotny, N.; Weissenböck, H. Monitoring of Usutu Virus Activity and Spread by Using Dead Bird Surveillance in Austria, 2003–2005. Vet. Microbiol. 2007, 122, 237–245. [Google Scholar] [CrossRef] [PubMed]
- Lühken, R.; Jöst, H.; Cadar, D.; Thomas, S.M.; Bosch, S.; Tannich, E.; Becker, N.; Ziegler, U.; Lachmann, L.; Schmidt-Chanasit, J. Distribution of Usutu Virus in Germany and Its Effect on Breeding Bird Populations. Emerg. Infect. Dis. 2017, 23, 1994–2001. [Google Scholar] [CrossRef] [PubMed]
- Steinmetz, H.W.; Bakonyi, T.; Weissenböck, H.; Hatt, J.-M.; Eulenberger, U.; Robert, N.; Hoop, R.; Nowotny, N. Emergence and Establishment of Usutu Virus Infection in Wild and Captive Avian Species in and around Zurich, Switzerland—Genomic and Pathologic Comparison to Other Central European Outbreaks. Vet. Microbiol. 2011, 148, 207–212. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Japanese Encephalitis. Available online: https://www.who.int/news-room/fact-sheets/detail/japanese-encephalitis (accessed on 15 July 2023).
- Mulvey, P.; Duong, V.; Boyer, S.; Burgess, G.; Williams, D.T.; Dussart, P.; Horwood, P.F. The Ecology and Evolution of Japanese Encephalitis Virus. Pathogens 2021, 10, 1534. [Google Scholar] [CrossRef]
- Buescher, E.L.; Scherer, W.F. Ecologic Studies of Japanese Encephalitis Virus in Japan. IX. Epidemiologic Correlations and Conclusions. Am. J. Trop. Med. Hyg. 1959, 8, 719–722. [Google Scholar] [CrossRef]
- Nemeth, N.; Bosco-Lauth, A.; Oesterle, P.; Kohler, D.; Bowen, R. North American Birds as Potential Amplifying Hosts of Japanese Encephalitis Virus. Am. J. Trop. Med. Hyg. 2012, 87, 760–767. [Google Scholar] [CrossRef]
- Hasegawa, T.; Takehara, Y.; Takahashi, K. Natural and Experimental Infections of Japanese Tree Sparrows with Japanese Encephalitis Virus. Arch. Virol. 1975, 49, 373–376. [Google Scholar] [CrossRef]
- Marshall, I.D.; Monath, T.P. The Arboviruses: Epidemiology and Ecology; CRC Press: Boca Raton, FL, USA, 1988; Volume 151, p. 189. [Google Scholar]
- Figueiredo, M.L.G.D.; Amarilla, A.A.; Figueiredo, G.G.D.; Alfonso, H.L.; Lippi, V.; Maia, F.G.M.; Morais, F.A.; Costa, C.A.D.; Henriques, D.A.; Durigon, E.L.; et al. Cacipacore Virus as an Emergent Mosquito-Borne Flavivirus. Rev. Soc. Bras. Med. Trop. 2017, 50, 539–542. [Google Scholar] [CrossRef]
- Curren, E.J.; Lindsey, N.P.; Fischer, M.; Hills, S.L. St. Louis Encephalitis Virus Disease in the United States, 2003–2017. Am. J. Trop. Med. Hyg. 2018, 99, 1074–1079. [Google Scholar] [CrossRef]
- Ridenour, C.L.; Cocking, J.; Poidmore, S.; Erickson, D.; Brock, B.; Valentine, M.; Roe, C.C.; Young, S.J.; Henke, J.A.; Hung, K.Y.; et al. St. Louis Encephalitis Virus in the Southwestern United States: A Phylogeographic Case for a Multi-Variant Introduction Event. Front. Genet. 2021, 12, 667895. [Google Scholar] [CrossRef]
- Kopp, A.; Gillespie, T.R.; Hobelsberger, D.; Estrada, A.; Harper, J.M.; Miller, R.A.; Eckerle, I.; Müller, M.A.; Podsiadlowski, L.; Leendertz, F.H.; et al. Provenance and Geographic Spread of St. Louis Encephalitis Virus. mBio 2013, 4, 10–1128. [Google Scholar] [CrossRef] [PubMed]
- Reisen, W.K.; Milby, M.M.; Presser, S.B.; Hardy, J.L. Ecology of Mosquitoes and St. Louis Encephalitis Virus in the Los Angeles Basin of California, 1987–1990. J. Med. Entomol. 1992, 29, 582–598. [Google Scholar] [CrossRef] [PubMed]
- Sulkin, S.E.; Allen, R.; Sims, R. Studies of Arthropod-Borne Virus Infections in Chiroptera: I. Susceptibility of Insectivorous Species to Experimental Infection with Japanese B and St. Louis Encephalitis Viruses. Am. J. Trop. Med. Hyg. 1963, 12, 800–814. [Google Scholar] [CrossRef]
- Day, J.F.; Stark, L.M. Avian Serology in a St. Louis Encephalitis Epicenter Before, During, and After a Widespread Epidemic in South Florida, USA. J. Med. Entomol. 1999, 36, 614–624. [Google Scholar] [CrossRef]
- Díaz, A.; Flores, F.S.; Quaglia, A.I.; Contigiani, M.S. Evaluation of Argentinean Bird Species as Amplifying Hosts for St. Louis Encephalitis Virus (Flavivirus, Flaviviridae). Am. J. Trop. Med. Hyg. 2018, 99, 216–221. [Google Scholar] [CrossRef] [PubMed]
- Reisen, W.K.; Chiles, R.E.; Martinez, V.M.; Fang, Y.; Green, E.N. Experimental Infection of California Birds with Western Equine Encephalomyelitis and St. Louis Encephalitis Viruses. J. Med. Entomol. 2003, 40, 968–982. [Google Scholar] [CrossRef]
- Gruwell, J.A.; Fogarty, C.L.; Bennett, S.G.; Challet, G.L.; Vanderpool, K.S.; Jozan, M.; Webb, J.P., Jr. Role of Peridomestic Birds on the Transmission of St. Louis Encephalitis Virus in Southern California. J. Wildl. Dis. 2000, 36, 13–34. [Google Scholar] [CrossRef]
- Diaz, L.A.; Quaglia, A.I.; Konigheim, B.S.; Boris, A.S.; Aguilar, J.J.; Komar, N.; Contigiani, M.S. Activity Patterns of St. Louis Encephalitis and West Nile Viruses in Free Ranging Birds during a Human Encephalitis Outbreak in Argentina. PLoS ONE 2016, 11, e0161871. [Google Scholar] [CrossRef]
- Danforth, M.E.; Snyder, R.E.; Feiszli, T.; Bullick, T.; Messenger, S.; Hanson, C.; Padgett, K.; Coffey, L.L.; Barker, C.M.; Reisen, W.K.; et al. Epidemiologic and Environmental Characterization of the Re-Emergence of St. Louis Encephalitis Virus in California, 2015–2020. PLoS Negl. Trop. Dis. 2022, 16, e0010664. [Google Scholar] [CrossRef]
- Centers for Disease Control and Prevention. Arbovirus Catalog—CDC Division of Vector-Borne Diseases (DVBD). Available online: https://wwwn.cdc.gov/arbocat/VirusBrowser.aspx (accessed on 19 July 2023).
- Mackenzie, J.S.; Barrett, A.D.T.; Deubel, V. The Japanese Encephalitis Serological Group of Flaviviruses: A Brief Introduction to the Group. Curr. Top. Microbiol. Immunol. 2002, 267, 1–10. [Google Scholar] [CrossRef]
- Kapadia, R.K.; Chauhan, L.; Piquet, A.L.; Tyler, K.L.; Pastula, D.M. An Overview of Eastern Equine Encephalitis (EEE). Neurohospitalist 2020, 10, 161–162. [Google Scholar] [CrossRef]
- Sah, R.; Siddiq, A.; Al-Ahdal, T.; Maulud, S.Q.; Mohanty, A.; Padhi, B.K.; El-Shall, N.A.; Chandran, D.; Emran, T.B.; Hussein, N.R.; et al. The Emerging Scenario for the Eastern Equine Encephalitis Virus and Mitigation Strategies to Counteract This Deadly Mosquito-Borne Zoonotic Virus, the Cause of the Most Severe Arboviral Encephalitis in Humans—An Update. Front. Trop. Dis. 2023, 3, 1077962. [Google Scholar] [CrossRef]
- Armstrong, P.M.; Andreadis, T.G. Eastern Equine Encephalitis Virus in Mosquitoes and Their Role as Bridge Vectors. Emerg. Infect. Dis. 2010, 16, 1869–1874. [Google Scholar] [CrossRef] [PubMed]
- Arrigo, N.C.; Adams, A.P.; Watts, D.M.; Newman, P.C.; Weaver, S.C. Cotton Rats and House Sparrows as Hosts for North and South American Strains of Eastern Equine Encephalitis Virus. Emerg. Infect. Dis. 2010, 16, 1373–1380. [Google Scholar] [CrossRef] [PubMed]
- Corrin, T.; Ackford, R.; Mascarenhas, M.; Greig, J.; Waddell, L.A. Eastern Equine Encephalitis Virus: A Scoping Review of the Global Evidence. Vector Borne Zoonotic Dis. 2021, 21, 305–320. [Google Scholar] [CrossRef]
- Thompson, K.A.; Henderson, E.; Fitzgerald, S.D.; Walker, E.D.; Kiupel, M. Eastern Equine Encephalitis Virus in Mexican Wolf Pups at Zoo, Michigan, USA. Emerg. Infect. Dis. 2021, 27, 1173–1176. [Google Scholar] [CrossRef]
- White, G.; Ottendorfer, C.; Graham, S.; Unnasch, T.R. Competency of Reptiles and Amphibians for Eastern Equine Encephalitis Virus. Am. J. Trop. Med. Hyg. 2011, 85, 421–425. [Google Scholar] [CrossRef]
- Estep, L.K.; McClure, C.J.W.; Burkett-Cadena, N.D.; Hassan, H.K.; Hicks, T.L.; Unnasch, T.R.; Hill, G.E. A Multi-Year Study of Mosquito Feeding Patterns on Avian Hosts in a Southeastern Focus of Eastern Equine Encephalitis Virus. Am. J. Trop. Med. Hyg. 2011, 84, 718–726. [Google Scholar] [CrossRef]
- Molaei, G.; Armstrong, P.M.; Graham, A.C.; Kramer, L.D.; Andreadis, T.G. Insights into the Recent Emergence and Expansion of Eastern Equine Encephalitis Virus in a New Focus in the Northern New England USA. Parasit. Vectors 2015, 8, 516. [Google Scholar] [CrossRef] [PubMed]
- Molaei, G.; Thomas, M.C.; Muller, T.; Medlock, J.; Shepard, J.J.; Armstrong, P.M.; Andreadis, T.G. Dynamics of Vector-Host Interactions in Avian Communities in Four Eastern Equine Encephalitis Virus Foci in the Northeastern U.S. PLoS Negl. Trop. Dis. 2016, 10, e0004347. [Google Scholar] [CrossRef]
- Locke, L.N.; Scanlon, J.E.; Byrne, R.J.; Knisley, J.O. Occurrence of Eastern Encephalitis Virus in House Sparrows. Wilson Bull. 1962, 74, 263–266. [Google Scholar]
- Andreadis, T.G.; Anderson, J.F.; Tirrell-Peck, S.J. Multiple Isolations of Eastern Equine Encephalitis and Highlands J Viruses from Mosquitoes (Diptera: Culicidae) during a 1996 Epizootic in Southeastern Connecticut. J. Med. Entomol. 1998, 35, 296–302. [Google Scholar] [CrossRef] [PubMed]
- Kurkela, S.; Manni, T.; Myllynen, J.; Vaheri, A.; Vapalahti, O. Clinical and Laboratory Manifestations of Sindbis Virus Infection: Prospective Study, Finland, 2002–2003. J. Infect Dis. 2005, 191, 1820–1829. [Google Scholar] [CrossRef] [PubMed]
- Lwande, O.W.; Obanda, V.; Bucht, G.; Mosomtai, G.; Otieno, V.; Ahlm, C.; Evander, M. Global Emergence of Alphaviruses That Cause Arthritis in Humans. Infect. Ecol. Epidemiol. 2015, 5, 29853. [Google Scholar] [CrossRef]
- Lundström, J.O.; Turell, M.J.; Niklasson, B. Antibodies to Ockelbo Virus in Three Orders of Birds (Anseriformes, Galliformes and Passeriformes) in Sweden. J. Wildl. Dis. 1992, 28, 144–147. [Google Scholar] [CrossRef][Green Version]
- Lundström, J.O.; Turell, M.J.; Niklasson, B. Viremia in Three Orders of Birds (Anseriformes, Galliformes and Passeriformes) Inoculated with Ockelbo Virus. J. Wildl. Dis. 1993, 29, 189–195. [Google Scholar] [CrossRef]
- Hesson, J.C.; Lundström, J.O.; Tok, A.; Östman, Ö.; Lundkvist, Å. Temporal Variation in Sindbis Virus Antibody Prevalence in Bird Hosts in an Endemic Area in Sweden. PLoS ONE 2016, 11, e0162005. [Google Scholar] [CrossRef]
- Lundström, J.O.; Lindström, K.M.; Olsen, B.; Dufva, R.; Krakower, D.S. Prevalence of Sindbis Virus Neutralizing Antibodies among Swedish Passerines Indicates That Thrushes Are the Main Amplifying Hosts. J. Med. Entomol. 2001, 38, 289–297. [Google Scholar] [CrossRef]
- Lau, C.; Aubry, M.; Musso, D.; Teissier, A.; Paulous, S.; Desprès, P.; de-Lamballerie, X.; Pastorino, B.; Cao-Lormeau, V.-M.; Weinstein, P. New Evidence for Endemic Circulation of Ross River Virus in the Pacific Islands and the Potential for Emergence. Int. J. Infect. Dis. 2017, 57, 73–76. [Google Scholar] [CrossRef]
- Jacups, S.P.; Whelan, P.I.; Currie, B.J. Ross River Virus and Barmah Forest Virus Infections: A Review of History, Ecology, and Predictive Models, with Implications for Tropical Northern Australia. Vector Borne Zoonotic Dis. 2008, 8, 283–297. [Google Scholar] [CrossRef]
- Whitehead, R.H.; Doderty, R.L.; Domrow, R.; Standfast, H.A.; Wetters, E.J. Studies of the Epidemiology of Arthropod-Borne Virus Infections at Mitchell River Mission, Cape York Peninsula, North Queensland. 3. Virus Studies of Wild Birns, 1964–1967. Trans. R Soc. Trop. Med. Hyg. 1968, 62, 439–445. [Google Scholar] [CrossRef] [PubMed]
- Jones, A.; Lowry, K.; Aaskov, J.; Holmes, E.C.; Kitchen, A. Molecular Evolutionary Dynamics of Ross River Virus and Implications for Vaccine Efficacy. J. Gen. Virol. 2010, 91, 182–188. [Google Scholar] [CrossRef]
- Marshall, I.D.; Brown, B.K.; Keith, K.; Gard, G.P.; Thibos, E. Variation in Arbovirus Infection Rates in Species of Birds Sampled in a Serological Survey during an Encephalitis Epidemic in the Murray Valley of South-Eastern Australia, February 1974. Aust. J. Exp. Biol. Med. Sci. 1982, 60, 471–478. [Google Scholar] [CrossRef] [PubMed]
- Anderson, C.R.; Downs, W.G.; Wattley, G.H.; Ahin, N.W.; Reese, A.A. Mayaro Virus: A New Human Disease Agent. II. Isolation from Blood of Patients in Trinidad, B.W.I. Am. J. Trop. Med. Hyg. 1957, 6, 1012–1016. [Google Scholar] [CrossRef]
- Celone, M.; Okech, B.; Han, B.A.; Forshey, B.M.; Anyamba, A.; Dunford, J.; Rutherford, G.; Mita-Mendoza, N.K.; Estallo, E.L.; Khouri, R.; et al. A Systematic Review and Meta-Analysis of the Potential Non-Human Animal Reservoirs and Arthropod Vectors of the Mayaro Virus. PLoS Negl. Trop. Dis. 2021, 15, e0010016. [Google Scholar] [CrossRef] [PubMed]
- Talarmin, A.; Chandler, L.J.; Kazanji, M.; de Thoisy, B.; Debon, P.; Lelarge, J.; Labeau, B.; Bourreau, E.; Vié, J.C.; Shope, R.E.; et al. Mayaro Virus Fever in French Guiana: Isolation, Identification, and Seroprevalence. Am. J. Trop. Med. Hyg. 1998, 59, 452–456. [Google Scholar] [CrossRef] [PubMed]
- Degallier, N.; Rosa, A.; Vasconcelos, P.F.C.; Hervé, J.-P.; Sá Filho, G.C.; Rosa, J.F.; Rosa, E.S.; Rodrigues, S.G. Modifications of Arbovirus Transmission in Relation to Construction of Dams in Brazilian Amazonia. Ciênc. Cult. 1992, 44, 124–135. [Google Scholar]
- Hoch, A.L.; Peterson, N.E.; LeDuc, J.W.; Pinheiro, F.P. An Outbreak of Mayaro Virus Disease in Belterra, Brazil. III. Entomological and Ecological Studies. Am. J. Trop. Med. Hyg. 1981, 30, 689–698. [Google Scholar] [CrossRef] [PubMed]
- Sanmartín, C.; Mackenzie, R.B.; Trapido, H.; Barreto, P.; Mullenax, C.H.; Gutiérrez, E.; Lesmes, C. Encefalitis Equina Venezolana En Colombia. Boletín Oficina Sanit. Panam. 1973, 74, 108–137. [Google Scholar]
- Calisher, C.H.; Gutiérrez, E.; Maness, K.S.; Lord, R.D. Isolation of Mayaro Virus from a Migrating Bird Captured in Louisiana in 1967. Bull. Pan. Am. Health Organ. 1974, 8, 243–248. [Google Scholar] [PubMed]
- Pan American Health Organization. Epidemiological Alert: Mayaro Fever (1 May 2019). Available online: https://iris.paho.org/bitstream/handle/10665.2/51485/EpiUpdate1May2019_eng.pdf?sequence=1&isAllowed=y (accessed on 2 July 2023).
- Scott, T.W.; Bowen, G.S.; Monath, T.P. A Field Study on the Effects of Fort Morgan Virus, an Arbovirus Transmitted by Swallow Bugs, on the Reproductive Success of Cliff Swallows and Symbiotic House Sparrows in Morgan County, Colorado, 1976. Am. J. Trop. Med. Hyg. 1984, 33, 981–991. [Google Scholar] [CrossRef] [PubMed]
- Fassbinder-Orth, C.A.; Killpack, T.L.; Goto, D.S.; Rainwater, E.L.; Shearn-Bochsler, V.I. High Costs of Infection: Alphavirus Infection Reduces Digestive Function and Bone and Feather Growth in Nestling House Sparrows (Passer domesticus). PLoS ONE 2018, 13, e0195467. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, V.A.; Meteyer, C.U.; Ip, H.S.; Long, R.R.; Brown, C.R. Pathology and Virus Detection in Tissues of Nestling House Sparrows Naturally Infected with Buggy Creek Virus (Togaviridae). J. Wildl. Dis. 2010, 46, 23–32. [Google Scholar] [CrossRef]
- Forrester, N.L.; Kenney, J.L.; Deardorff, E.; Wang, E.; Weaver, S.C. Western Equine Encephalitis Submergence: Lack of Evidence for a Decline in Virus Virulence. Virology 2008, 380, 170–172. [Google Scholar] [CrossRef] [PubMed]
- Ludwig, A.; Zheng, H.; Vrbova, L.; Drebot, M.; Iranpour, M.; Lindsay, L. Increased Risk of Endemic Mosquito-Borne Diseases in Canada Due to Climate Change. Can. Commun. Dis. Rep. 2019, 45, 91–97. [Google Scholar] [CrossRef] [PubMed]
- Alluwaimi, A.M.; Alshubaith, I.H.; Al-Ali, A.M.; Abohelaika, S. The Coronaviruses of Animals and Birds: Their Zoonosis, Vaccines, and Models for SARS-CoV and SARS-CoV2. Front. Vet. Sci. 2020, 7, 582287. [Google Scholar] [CrossRef]
- Woo, P.C.Y.; de Groot, R.J.; Haagmans, B.; Lau, S.K.P.; Neuman, B.W.; Perlman, S.; Sola, I.; van der Hoek, L.; Wong, A.C.P.; Yeh, S.-H. ICTV Virus Taxonomy Profile: Coronaviridae 2023. J. Gen. Virol. 2023, 104, 001843. [Google Scholar] [CrossRef]
- Cavanagh, D.; Mawditt, K.; Sharma, M.; Drury, S.E.; Ainsworth, H.L.; Britton, P.; Gough, R.E. Detection of a Coronavirus from Turkey Poults in Europe Genetically Related to Infectious Bronchitis Virus of Chickens. Avian Pathol. 2001, 30, 355–368. [Google Scholar] [CrossRef]
- Chacón, J.L.; Chacón, R.D.; Sánchez-Llatas, C.J.; Morín, J.G.; Astolfi, C.S.; Piantino, A.J. Antigenic and Molecular Characterization of Isolates of the Brazilian Genotype BR-I (GI-11) of Infectious Bronchitis Virus Supports Its Recognition as BR-I Serotype. Avian Pathol. 2023, 52, 1–16. [Google Scholar] [CrossRef]
- Miłek, J.; Blicharz-Domańska, K. Coronaviruses in Avian Species—Review with Focus on Epidemiology and Diagnosis in Wild Birds. J. Vet. Res. 2018, 62, 249–255. [Google Scholar] [CrossRef]
- Woo, P.C.Y.; Lau, S.K.P.; Lam, C.S.F.; Lai, K.K.Y.; Huang, Y.; Lee, P.; Luk, G.S.M.; Dyrting, K.C.; Chan, K.-H.; Yuen, K.-Y. Comparative Analysis of Complete Genome Sequences of Three Avian Coronaviruses Reveals a Novel Group 3c Coronavirus. J. Virol. 2009, 83, 908–917. [Google Scholar] [CrossRef]
- Woo, P.C.Y.; Lau, S.K.P.; Lam, C.S.F.; Lau, C.C.Y.; Tsang, A.K.L.; Lau, J.H.N.; Bai, R.; Teng, J.L.L.; Tsang, C.C.C.; Wang, M.; et al. Discovery of Seven Novel Mammalian and Avian Coronaviruses in the Genus Deltacoronavirus Supports Bat Coronaviruses as the Gene Source of Alphacoronavirus and Betacoronavirus and Avian Coronaviruses as the Gene Source of Gammacoronavirus and Deltacoronavirus. J. Virol. 2012, 86, 3995–4008. [Google Scholar] [CrossRef] [PubMed]
- Domańska-Blicharz, K.; Miłek-Krupa, J.; Pikuła, A. Diversity of Coronaviruses in Wild Representatives of the Aves Class in Poland. Viruses 2021, 13, 1497. [Google Scholar] [CrossRef] [PubMed]
- Zhigailov, A.V.; Maltseva, E.R.; Perfilyeva, Y.V.; Ostapchuk, Y.O.; Naizabayeva, D.A.; Berdygulova, Z.A.; Kuatbekova, S.A.; Nizkorodova, A.S.; Mashzhan, A.; Gavrilov, A.E.; et al. Prevalence and Genetic Diversity of Coronaviruses, Astroviruses and Paramyxoviruses in Wild Birds in Southeastern Kazakhstan. Heliyon 2022, 8, e11324. [Google Scholar] [CrossRef] [PubMed]
- Franzo, G.; Massi, P.; Tucciarone, C.M.; Barbieri, I.; Tosi, G.; Fiorentini, L.; Ciccozzi, M.; Lavazza, A.; Cecchinato, M.; Moreno, A. Think Globally, Act Locally: Phylodynamic Reconstruction of Infectious Bronchitis Virus (IBV) QX Genotype (GI-19 Lineage) Reveals Different Population Dynamics and Spreading Patterns When Evaluated on Different Epidemiological Scales. PLoS ONE 2017, 12, e0184401. [Google Scholar] [CrossRef]
- Rahman, M.M.; Talukder, A.; Chowdhury, M.M.H.; Talukder, R.; Akter, R. Coronaviruses in Wild Birds—A Potential and Suitable Vector for Global Distribution. Vet. Med. Sci. 2021, 7, 264–272. [Google Scholar] [CrossRef] [PubMed]
- Tannock, G.A.; Shafren, D.R. Avian Encephalomyelitis: A Review. Avian Pathol. 1994, 23, 603–620. [Google Scholar] [CrossRef]
- Woo, P.C.Y.; Lau, S.K.P.; Huang, Y.; Lam, C.S.F.; Poon, R.W.S.; Tsoi, H.-W.; Lee, P.; Tse, H.; Chan, A.S.L.; Luk, G.; et al. Comparative Analysis of Six Genome Sequences of Three Novel Picornaviruses, Turdiviruses 1, 2 and 3, in Dead Wild Birds, and Proposal of Two Novel Genera, Orthoturdivirus and Paraturdivirus, in the Family Picornaviridae. J. Gen. Virol. 2010, 91, 2433–2448. [Google Scholar] [CrossRef]
- Pankovics, P.; Boros, Á.; Phan, T.G.; Delwart, E.; Reuter, G. A Novel Passerivirus (Family Picornaviridae) in an Outbreak of Enteritis with High Mortality in Estrildid Finches (Uraeginthus sp.). Arch. Virol. 2018, 163, 1063–1071. [Google Scholar] [CrossRef]
- Handel, C.M.; Pajot, L.M.; Matsuoka, S.M.; Hemert, C.V.; Terenzi, J.; Talbot, S.L.; Mulcahy, D.M.; Meteyer, C.U.; Trust, K.A. Epizootic of Beak Deformities Among Wild Birds in Alaska: An Emerging Disease in North America? Auk 2010, 127, 882–898. [Google Scholar] [CrossRef]
- Hemert, C.V.; Handel, C.M. Beak Deformities in Northwestern Crows: Evidence of a Multispecies Epizootic. Auk 2010, 127, 746–751. [Google Scholar] [CrossRef]
- Zylberberg, M.; Van Hemert, C.; Handel, C.M.; DeRisi, J.L. Avian Keratin Disorder of Alaska Black-Capped Chickadees Is Associated with Poecivirus Infection. Virol. J. 2018, 15, 100. [Google Scholar] [CrossRef]
- Zylberberg, M.; Van Hemert, C.; Handel, C.M.; Liu, R.M.; DeRisi, J.L. Poecivirus Is Present in Individuals with Beak Deformities in Seven Species of North American Birds. J. Wildl. Dis. 2021, 57, 273–281. [Google Scholar] [CrossRef] [PubMed]
- De Benedictis, P.; Schultz-Cherry, S.; Burnham, A.; Cattoli, G. Astrovirus Infections in Humans and Animals—Molecular Biology, Genetic Diversity, and Interspecies Transmissions. Infect. Genet. Evol. 2011, 11, 1529–1544. [Google Scholar] [CrossRef]
- Pantin-Jackwood, M.; Todd, D.; Koci, M.D. Avian Astroviruses. Astrovirus Res. 2012, 1, 151–180. [Google Scholar] [CrossRef]
- Yamaguchi, S.; Imada, T.; Kawamura, H. Characterization of a Picornavirus Isolated from Broiler Chicks. Avian Dis. 1979, 23, 571–581. [Google Scholar] [CrossRef]
- Schultz-Cherry, S.; Kapczynski, D.R.; Simmons, V.M.; Koci, M.D.; Brown, C.; Barnes, H.J. Identifying Agent(s) Associated with Poult Enteritis Mortality Syndrome: Importance of the Thymus. Avian Dis. 2000, 44, 256–265. [Google Scholar] [CrossRef]
- Li, L.; Sun, M.; Zhang, Y.; Liao, M. A Review of the Emerging Poultry Visceral Gout Disease Linked to Avian Astrovirus Infection. Int. J. Mol. Sci. 2022, 23, 10429. [Google Scholar] [CrossRef]
- Fernández-Correa, I.; Truchado, D.A.; Gomez-Lucia, E.; Doménech, A.; Pérez-Tris, J.; Schmidt-Chanasit, J.; Cadar, D.; Benítez, L. A Novel Group of Avian Astroviruses from Neotropical Passerine Birds Broaden the Diversity and Host Range of Astroviridae. Sci. Rep. 2019, 9, 9513. [Google Scholar] [CrossRef]
- Mendenhall, I.H.; Smith, G.J.D.; Vijaykrishna, D. Ecological Drivers of Virus Evolution: Astrovirus as a Case Study. J. Virol. 2015, 89, 6978–6981. [Google Scholar] [CrossRef]
- Mendenhall, I.H.; Yaung, K.N.; Joyner, P.H.; Keatts, L.; Borthwick, S.; Neves, E.S.; San, S.; Gilbert, M.; Smith, G.J. Detection of a Novel Astrovirus from a Black-Naped Monarch (Hypothymis azurea) in Cambodia. Virol. J. 2015, 12, 182. [Google Scholar] [CrossRef]
- Cook, J.K. Avian Astroviruses. In Diseases of Poultry; Swayne, D., Boulianne, M., Logue, C., McDougald, L., Nair, V., Suarez, D., Eds.; Wiley-Blackwell: Hoboken, NJ, USA, 2020; ISBN 978-1-119-37116-8. [Google Scholar]
- Koci, M.D.; Schultz-Cherry, S. Avian Astroviruses. Avian Pathol. 2002, 31, 213–227. [Google Scholar] [CrossRef]
- Chu, D.K.W.; Poon, L.L.M.; Guan, Y.; Peiris, J.S.M. Novel Astroviruses in Insectivorous Bats. J. Virol. 2008, 82, 9107–9114. [Google Scholar] [CrossRef] [PubMed]
- De Battisti, C.; Salviato, A.; Jonassen, C.M.; Toffan, A.; Capua, I.; Cattoli, G. Genetic Characterization of Astroviruses Detected in Guinea Fowl (Numida meleagris) Reveals a Distinct Genotype and Suggests Cross-Species Transmission between Turkey and Guinea Fowl. Arch. Virol. 2012, 157, 1329–1337. [Google Scholar] [CrossRef] [PubMed]
- De Souza, W.M.; Fumagalli, M.J.; de Araujo, J.; Ometto, T.; Modha, S.; Thomazelli, L.M.; Durigon, E.L.; Murcia, P.R.; Figueiredo, L.T.M. Discovery of Novel Astrovirus and Calicivirus Identified in Ruddy Turnstones in Brazil. Sci. Rep. 2019, 9, 5556. [Google Scholar] [CrossRef] [PubMed]
- van Hemert, F.J.; Berkhout, B.; Lukashov, V.V. Host-Related Nucleotide Composition and Codon Usage as Driving Forces in the Recent Evolution of the Astroviridae. Virology 2007, 361, 447–454. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.-R.; Kwon, Y.-K.; Jang, I.; Bae, Y.-C. Viral Metagenomic Analysis of Chickens with Runting-Stunting Syndrome in the Republic of Korea. Virol. J. 2020, 17, 53. [Google Scholar] [CrossRef]
- Lima, D.A.; Cibulski, S.P.; Finkler, F.; Teixeira, T.F.; Varela, A.P.M.; Cerva, C.; Loiko, M.R.; Scheffer, C.M.; Dos Santos, H.F.; Mayer, F.Q.; et al. Faecal Virome of Healthy Chickens Reveals a Large Diversity of the Eukaryote Viral Community, Including Novel Circular ssDNA Viruses. J. Gen. Virol. 2017, 98, 690–703. [Google Scholar] [CrossRef]
- Lima, D.A.; Cibulski, S.P.; Tochetto, C.; Varela, A.P.M.; Finkler, F.; Teixeira, T.F.; Loiko, M.R.; Cerva, C.; Junqueira, D.M.; Mayer, F.Q.; et al. The Intestinal Virome of Malabsorption Syndrome-Affected and Unaffected Broilers through Shotgun Metagenomics. Virus Res. 2019, 261, 9–20. [Google Scholar] [CrossRef]
- Wolf, S.; Reetz, J.; Hoffmann, K.; Gründel, A.; Schwarz, B.-A.; Hänel, I.; Otto, P.H. Discovery and Genetic Characterization of Novel Caliciviruses in German and Dutch Poultry. Arch. Virol. 2012, 157, 1499–1507. [Google Scholar] [CrossRef]
- Phan, T.G.; Vo, N.P.; Boros, Á.; Pankovics, P.; Reuter, G.; Li, O.T.W.; Wang, C.; Deng, X.; Poon, L.L.M.; Delwart, E. The Viruses of Wild Pigeon Droppings. PLoS ONE 2013, 8, e72787. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Wang, M.; Dong, Y.; Zhang, B.; Zhang, D. Genetic Characterization of a Novel Calicivirus from a Goose. Arch. Virol. 2017, 162, 2115–2118. [Google Scholar] [CrossRef] [PubMed]
- Wille, M.; Shi, M.; Klaassen, M.; Hurt, A.C.; Holmes, E.C. Virome Heterogeneity and Connectivity in Waterfowl and Shorebird Communities. ISME J. 2019, 13, 2603–2616. [Google Scholar] [CrossRef] [PubMed]
- Devaney, R.; Trudgett, J.; Trudgett, A.; Meharg, C.; Smyth, V. A Metagenomic Comparison of Endemic Viruses from Broiler Chickens with Runting-Stunting Syndrome and from Normal Birds. Avian Pathol. 2016, 45, 616–629. [Google Scholar] [CrossRef]
- French, R.K.; Stone, Z.L.; Parker, K.A.; Holmes, E.C. Novel Viral and Microbial Species in a Translocated Toutouwai (Petroica longipes) Population from Aotearoa/New Zealand. One Health Outlook 2022, 4, 16. [Google Scholar] [CrossRef] [PubMed]
- Chang, W.-S.; Rose, K.; Holmes, E.C. Meta-Transcriptomic Analysis of the Virome and Microbiome of the Invasive Indian Myna (Acridotheres tristis) in Australia. One Health 2021, 13, 100360. [Google Scholar] [CrossRef]
- Cong, W.; Meng, Q.-F.; Shan, X.-F.; Sun, W.-W.; Qin, S.-Y.; Zhang, X.-X.; Huang, S.-Y.; Qian, A.-D. Seroprevalence and Risk Factors Associated with Hepatitis E Virus Infection in Three Species of Pet Birds in Northwest China. Sci. World J. 2014, 2014, 296285. [Google Scholar] [CrossRef]
- Estes, M.K.; Cohen, J. Rotavirus Gene Structure and Function. Microbiol. Rev. 1989, 53, 410–449. [Google Scholar] [CrossRef]
- Sun, X.; Li, D.; Duan, Z. Structural Basis of Glycan Recognition of Rotavirus. Front. Mol. Biosci. 2021, 8, 658029. [Google Scholar] [CrossRef]
- Kattoor, J.J.; Malik, Y.S.; Sharma, K.; Kumar, N.; Batra, M.; Jindal, N.; Yadav, A.S. Molecular Evidence of Group D Rotavirus in Commercial Broiler Chicks in India. Avian Biol. Res. 2013, 6, 313–316. [Google Scholar] [CrossRef]
- Otto, P.; Liebler-Tenorio, E.M.; Elschner, M.; Reetz, J.; Löhren, U.; Diller, R. Detection of Rotaviruses and Intestinal Lesions in Broiler Chicks from Flocks with Runting and Stunting Syndrome (RSS). Avian Dis. 2006, 50, 411–418. [Google Scholar] [CrossRef] [PubMed]
- McNulty, M.S.; Allan, G.M.; Stuart, J.C. Rotavirus Infection in Avian Species. Vet. Rec. 1978, 103, 319–320. [Google Scholar] [CrossRef] [PubMed]
- Ursu, K.; Papp, H.; Kisfali, P.; Rigó, D.; Melegh, B.; Martella, V.; Bányai, K. Monitoring of Group A Rotaviruses in Wild-Living Birds in Hungary. Avian Dis. 2011, 55, 123–127. [Google Scholar] [CrossRef]
- Duarte Júnior, J.W.B.; Chagas, E.H.N.; Serra, A.C.S.; Souto, L.C.D.S.; da Penha Júnior, E.T.; Bandeira, R.D.S.; e Guimarães, R.J.D.P.S.; Oliveira, H.G.D.S.; Sousa, T.K.S.; Lopes, C.T.D.A.; et al. Ocurrence of Rotavirus and Picobirnavirus in Wild and Exotic Avian from Amazon Forest. PLoS Negl. Trop. Dis. 2021, 15, e0008792. [Google Scholar] [CrossRef] [PubMed]
- Luchs, A.; Timenetsky, M. do C.S.T. Gastroenterite por rotavírus do grupo A: Era pós-vacinal, genótipos e transmissão zoonótica. Einstein 2016, 14, 278–287. [Google Scholar] [CrossRef] [PubMed]
- Barros, B.D.C.V.D.; Chagas, E.N.; Bezerra, L.W.; Ribeiro, L.G.; Duarte Júnior, J.W.B.; Pereira, D.; Penha Junior, E.T.D.; Silva, J.R.; Bezerra, D.A.M.; Bandeira, R.S.; et al. Rotavirus A in Wild and Domestic Animals from Areas with Environmental Degradation in the Brazilian Amazon. PLoS ONE 2018, 13, e0209005. [Google Scholar] [CrossRef] [PubMed]
- Pavlin, B.I.; Schloegel, L.M.; Daszak, P. Risk of Importing Zoonotic Diseases through Wildlife Trade, United States. Emerg. Infect. Dis. 2009, 15, 1721–1726. [Google Scholar] [CrossRef] [PubMed]
- McCowan, C.; Crameri, S.; Kocak, A.; Shan, S.; Fegan, M.; Forshaw, D.; Rubbenstroth, D.; Chen, H.; Holmes, C.; Harper, J.; et al. A Novel Group A Rotavirus Associated with Acute Illness and Hepatic Necrosis in Pigeons (Columba livia), in Australia. PLoS ONE 2018, 13, e0203853. [Google Scholar] [CrossRef]
- Shen, Y.; Cai, L.; Wang, Y.; Wei, R.; He, M.; Wang, S.; Wang, G.; Cheng, Z. Genetic Mutations of Avian Leukosis Virus Subgroup J Strains Extended Their Host Range. J. Gen. Virol. 2014, 95, 691–699. [Google Scholar] [CrossRef]
- Reinišová, M.; Plachý, J.; Kučerová, D.; Šenigl, F.; Vinkler, M.; Hejnar, J. Genetic Diversity of NHE1, Receptor for Subgroup J Avian Leukosis Virus, in Domestic Chicken and Wild Anseriform Species. PLoS ONE 2016, 11, e0150589. [Google Scholar] [CrossRef]
- Varejka, F.; Tomsik, F. The Role of House Sparrow (Passer domesticus L.) in the Spread of Leukosis Viruses in Poultry. I. Determination of Neutralizing Antibodies. Acta Vet. Brno 1974, 43, 367–370. [Google Scholar]
- Jiang, L.; Zeng, X.; Hua, Y.; Gao, Q.; Fan, Z.; Chai, H.; Wang, Q.; Qi, X.; Wang, Y.; Gao, H.; et al. Genetic Diversity and Phylogenetic Analysis of Glycoprotein Gp85 of Avian Leukosis Virus Subgroup J Wild-Bird Isolates from Northeast China. Arch. Virol. 2014, 159, 1821–1826. [Google Scholar] [CrossRef]
- Han, C.; Hao, R.; Liu, L.; Zeng, X. Molecular Characterization of 3’UTRs of J Subgroup Avian Leukosis Virus in Passerine Birds in China. Arch. Virol. 2015, 160, 845–849. [Google Scholar] [CrossRef] [PubMed]
- Walker, P.J.; Siddell, S.G.; Lefkowitz, E.J.; Mushegian, A.R.; Adriaenssens, E.M.; Alfenas-Zerbini, P.; Davison, A.J.; Dempsey, D.M.; Dutilh, B.E.; García, M.L.; et al. Changes to Virus Taxonomy and to the International Code of Virus Classification and Nomenclature Ratified by the International Committee on Taxonomy of Viruses (2021). Arch. Virol. 2021, 166, 2633–2648. [Google Scholar] [CrossRef] [PubMed]
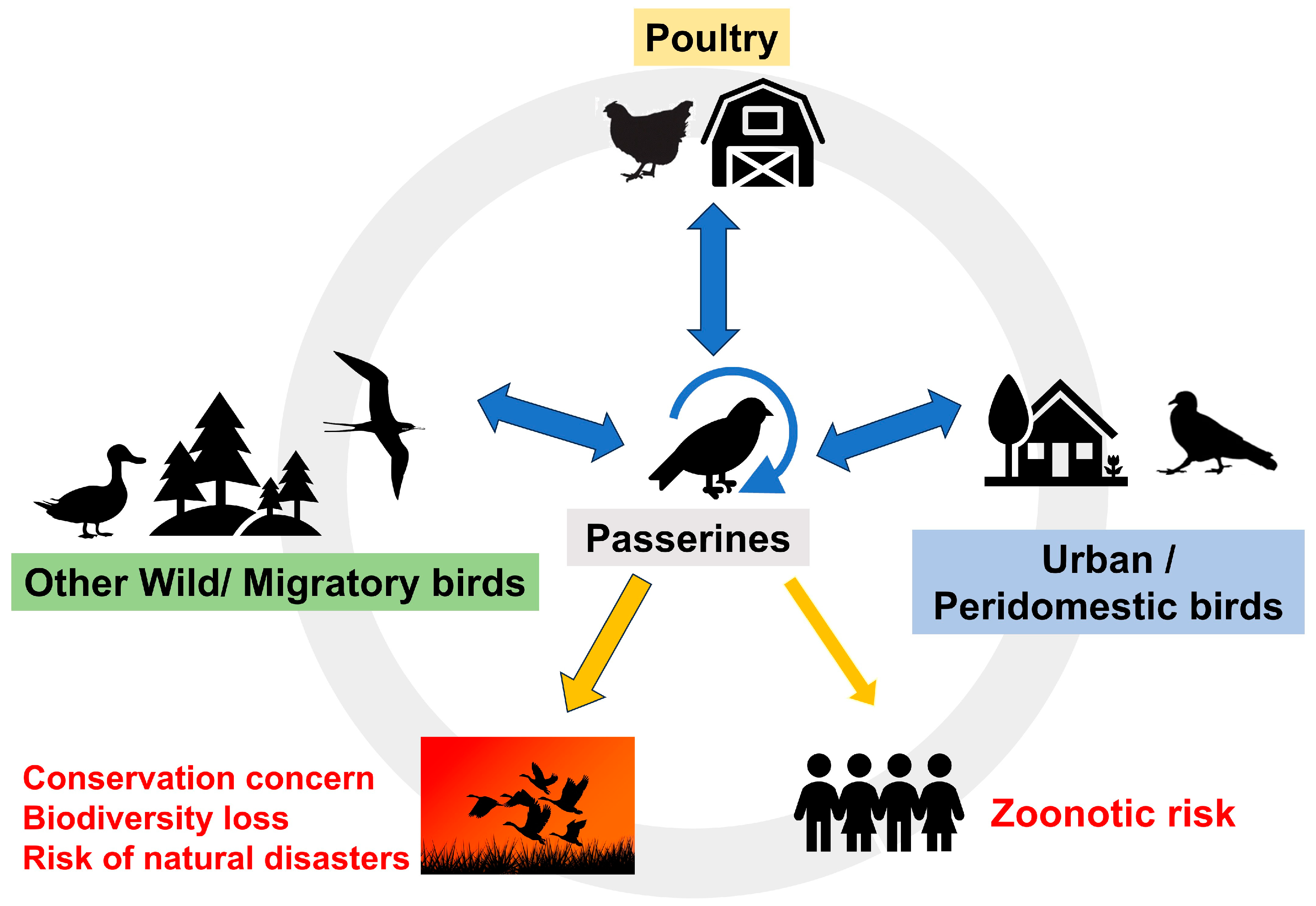
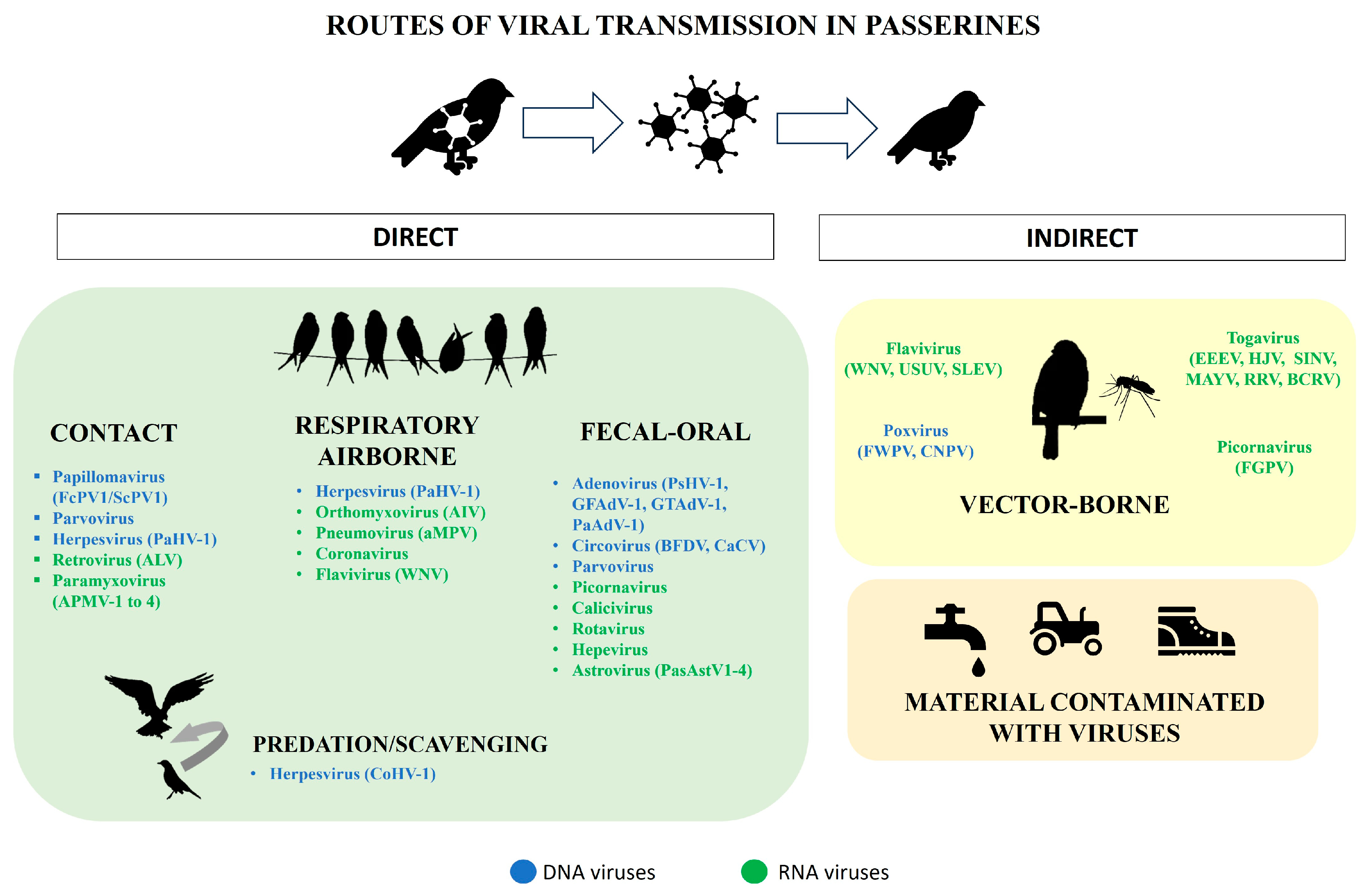
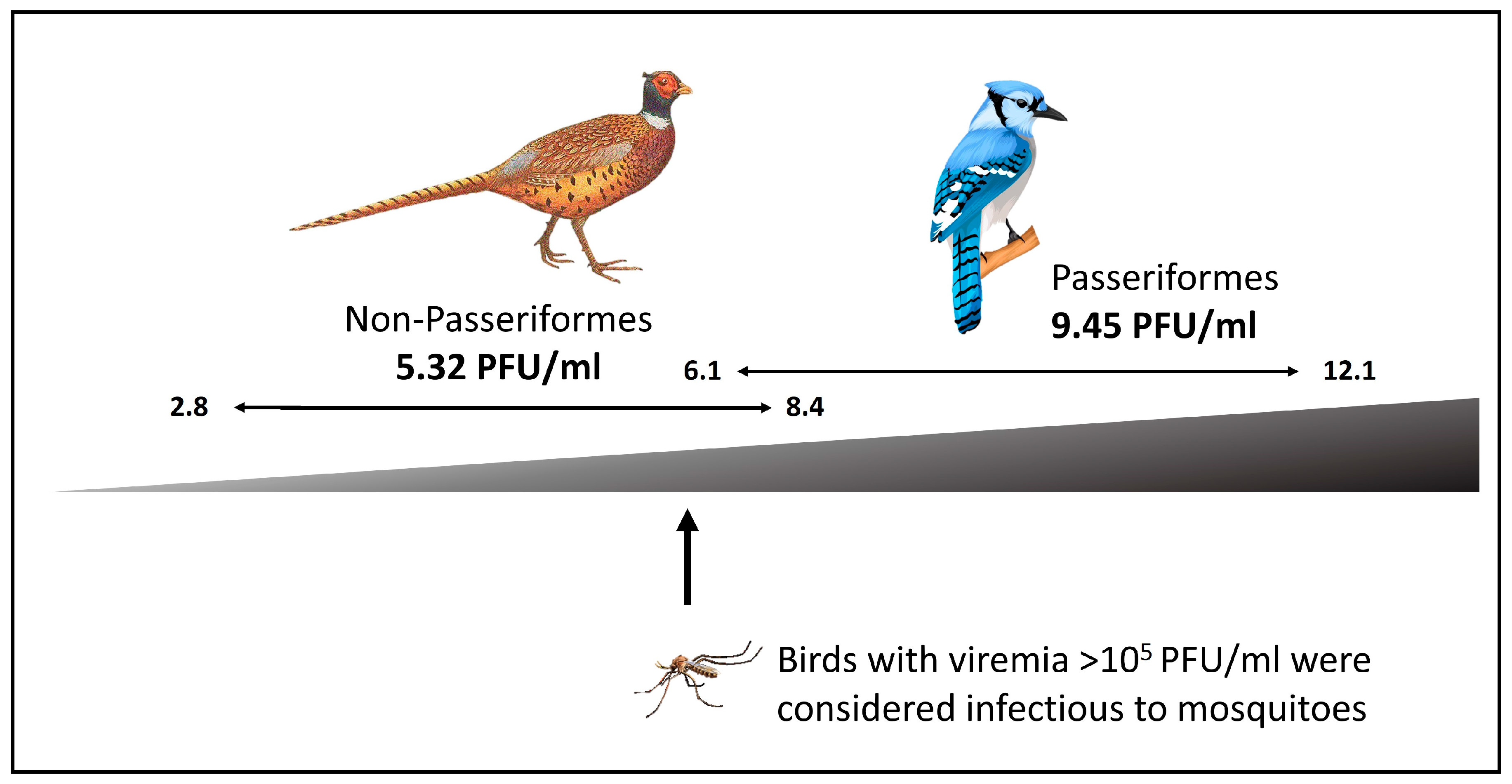
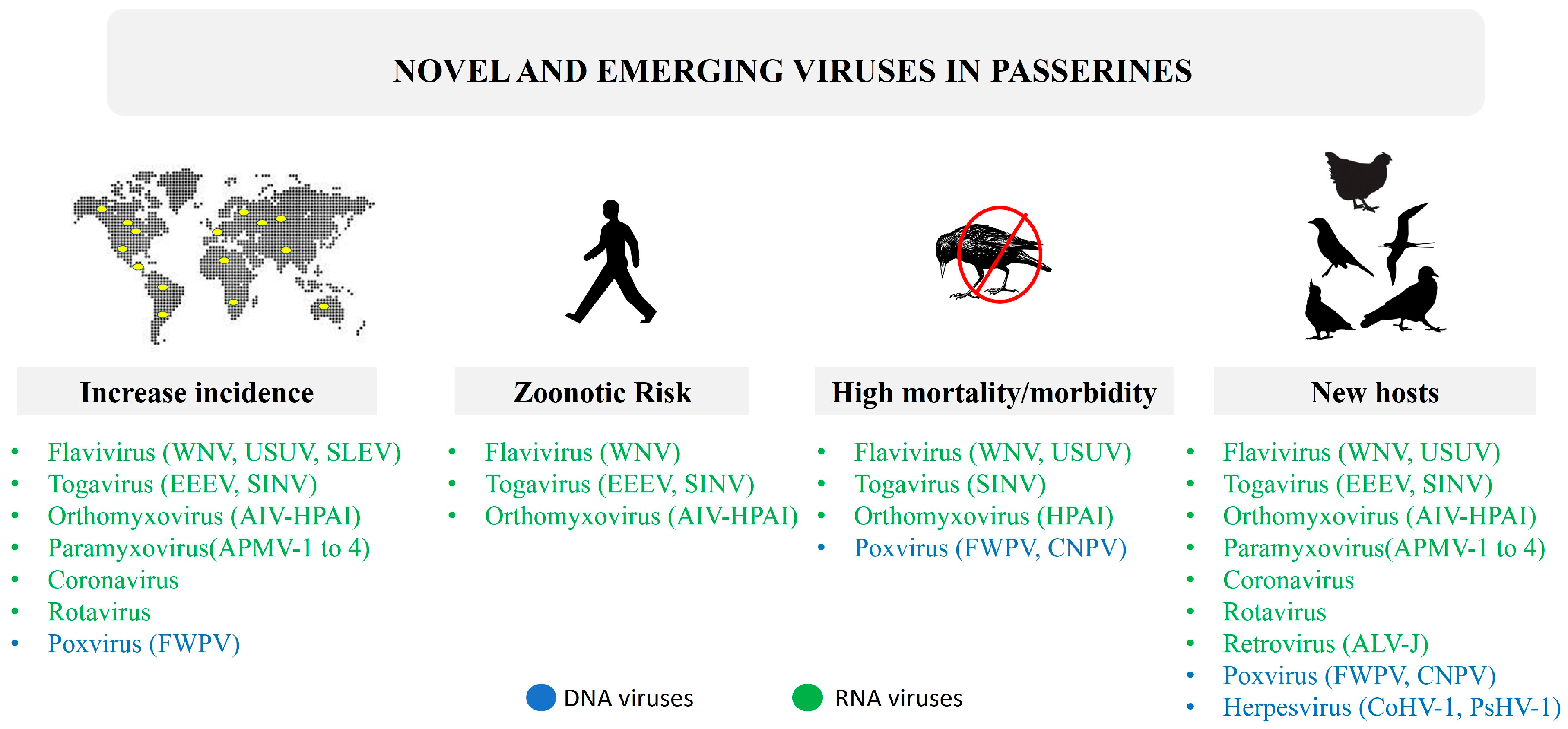
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Williams, R.A.J.; Sánchez-Llatas, C.J.; Doménech, A.; Madrid, R.; Fandiño, S.; Cea-Callejo, P.; Gomez-Lucia, E.; Benítez, L. Emerging and Novel Viruses in Passerine Birds. Microorganisms 2023, 11, 2355. https://doi.org/10.3390/microorganisms11092355
Williams RAJ, Sánchez-Llatas CJ, Doménech A, Madrid R, Fandiño S, Cea-Callejo P, Gomez-Lucia E, Benítez L. Emerging and Novel Viruses in Passerine Birds. Microorganisms. 2023; 11(9):2355. https://doi.org/10.3390/microorganisms11092355
Chicago/Turabian StyleWilliams, Richard A. J., Christian J. Sánchez-Llatas, Ana Doménech, Ricardo Madrid, Sergio Fandiño, Pablo Cea-Callejo, Esperanza Gomez-Lucia, and Laura Benítez. 2023. "Emerging and Novel Viruses in Passerine Birds" Microorganisms 11, no. 9: 2355. https://doi.org/10.3390/microorganisms11092355
APA StyleWilliams, R. A. J., Sánchez-Llatas, C. J., Doménech, A., Madrid, R., Fandiño, S., Cea-Callejo, P., Gomez-Lucia, E., & Benítez, L. (2023). Emerging and Novel Viruses in Passerine Birds. Microorganisms, 11(9), 2355. https://doi.org/10.3390/microorganisms11092355






