Understanding HCMV Latency Using Unbiased Proteomic Analyses
Abstract
:1. Introduction
2. HCMV Lytic and Latent Lifecycle
2.1. Viral Gene Expression during HCMV Latency
2.2. Cellular Transcriptome Analysis of HCMV Latency
3. Studying the Proteome
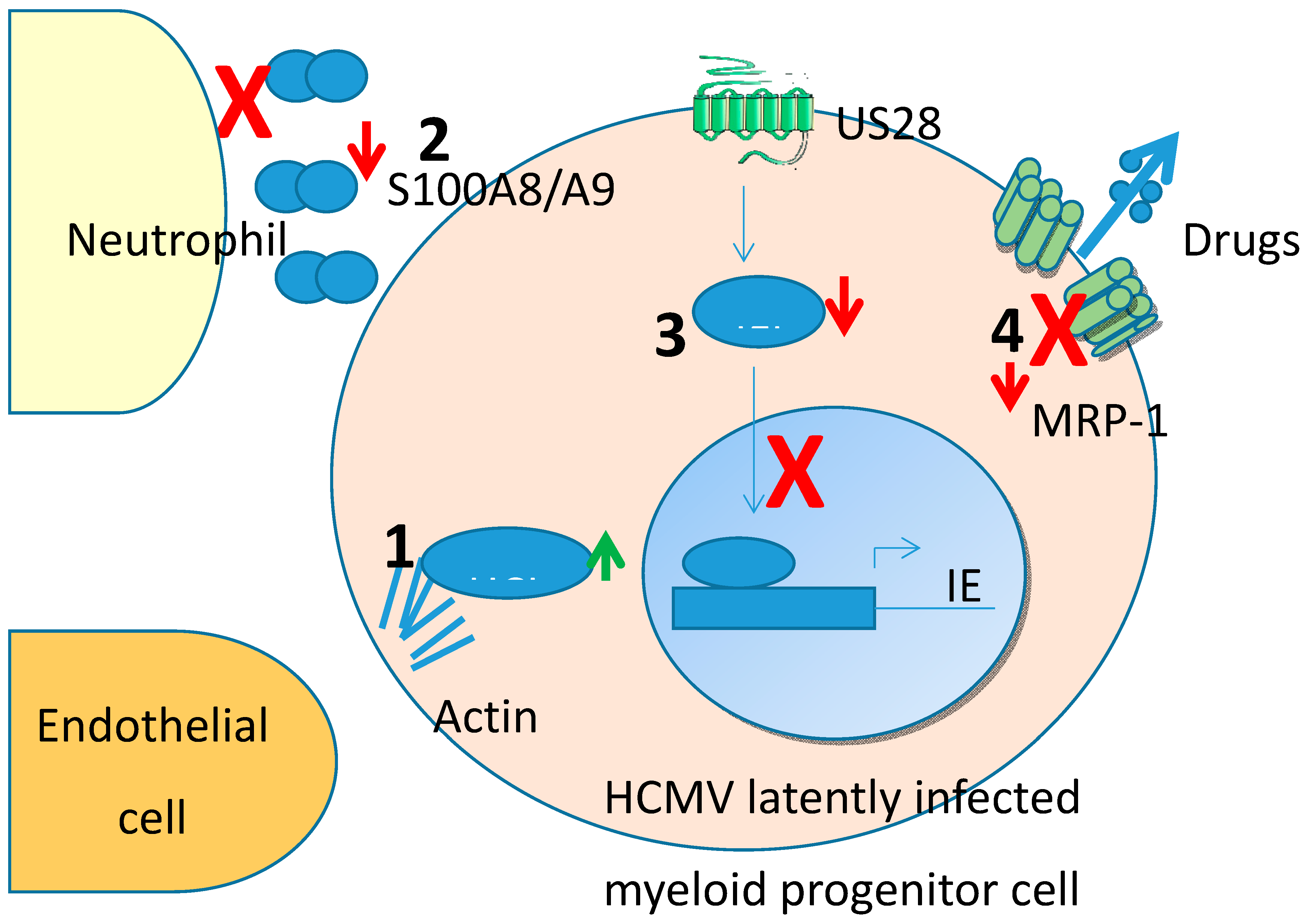
3.1. Total Proteomic Changes during HCMV Latency
3.2. HCLS1
3.3. S100A8/A9
4. Proteome Changes Induced by Expression of Latency-Associated Proteins in Isolation
4.1. US28
4.2. UL138
5. Concluding Remarks
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Cannon, M.J.; Schmid, D.S.; Hyde, T.B. Review of cytomegalovirus seroprevalence and demographic characteristics associated with infection. Rev. Med. Virol. 2010, 20, 202–213. [Google Scholar] [CrossRef]
- Ahmed, A. Antiviral treatment of cytomegalovirus infection. Infect. Disord. Drug Targets 2010, 11, 475–503. [Google Scholar] [CrossRef]
- Ehlert, K.; Groll, A.H.; Kuehn, J.; Vormoor, J. Treatment of refractory CMV-infection following hematopoietic stem cell transplantation with the combination of foscarnet and leflunomide. Klin. Padiatr. 2006, 218, 180–184. [Google Scholar] [CrossRef] [PubMed]
- Weekes, M.P.; Tomasec, P.; Huttlin, E.L.; Fielding, C.A.; Nusinow, D.; Stanton, R.J.; Wang, E.C.; Aicheler, R.; Murrell, I.; Wilkinson, G.W.; et al. Quantitative temporal viromics: An approach to investigate host-pathogen interaction. Cell 2014, 157, 1460–1472. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sinclair, J.; Poole, E. Human cytomegalovirus latency and reactivation in and beyond the myeloid lineage. Future Virol. 2014, 6, 7. [Google Scholar] [CrossRef] [Green Version]
- Goodrum, F.; Reeves, M.; Sinclair, J.; High, K.; Shenk, T. Human cytomegalovirus sequences expressed in latently infected individuals promote a latent infection in vitro. Blood 2007, 110, 937–945. [Google Scholar] [CrossRef] [Green Version]
- Beisser, P.S.; Laurent, L.; Virelizier, J.L.; Michelson, S. Human cytomegalovirus chemokine receptor gene US28 is transcribed in latently infected THP-1 monocytes. J. Virol. 2001, 75, 5949–5957. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Humby, M.S.; O’Connor, C.M. Human Cytomegalovirus US28 Is Important for Latent Infection of Hematopoietic Progenitor Cells. J. Virol. 2015, 90, 2959–2970. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Poole, E.; Walther, A.; Raven, K.; Benedict, C.A.; Mason, G.M.; Sinclair, J. The myeloid transcription factor GATA-2 regulates the viral UL144 gene during human cytomegalovirus latency in an isolate-specific manner. J. Virol. 2013, 87, 4261–4271. [Google Scholar] [CrossRef] [Green Version]
- Bego, M.; Maciejewski, J.; Khaiboullina, S.; Pari, G.; St Jeor, S. Characterization of an antisense transcript spanning the UL81-82 locus of human cytomegalovirus. J. Virol. 2005, 79, 11022–11034. [Google Scholar] [CrossRef] [Green Version]
- Jenkins, C.; Abendroth, A.; Slobedman, B. A novel viral transcript with homology to human interleukin-10 is expressed during latent human cytomegalovirus infection. J. Virol. 2004, 78, 1440–1447. [Google Scholar] [CrossRef] [Green Version]
- Slobedman, B.; Cheung, A.K. Microarrays for the study of viral gene expression during human cytomegalovirus latent infection. Methods Mol. Med. 2008, 141, 153–175. [Google Scholar] [PubMed]
- Cheng, S.; Caviness, K.; Buehler, J.; Smithey, M.; Nikolich-Zugich, J.; Goodrum, F. Transcriptome-wide characterization of human cytomegalovirus in natural infection and experimental latency. Proc. Natl. Acad. Sci. USA 2017, 114, E10586–E10595. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shnayder, M.; Nachshon, A.; Krishna, B.; Poole, E.; Boshkov, A.; Binyamin, A.; Maza, I.; Sinclair, J.; Schwartz, M.; Stern-Ginossar, N. Defining the Transcriptional Landscape during Cytomegalovirus Latency with Single-Cell RNA Sequencing. MBio 2018, 9. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Poole, E.; Huang, C.J.Z.; Forbester, J.; Shnayder, M.; Nachshon, A.; Kweider, B.; Basaj, A.; Smith, D.; Jackson, S.E.; Liu, B.; et al. An iPSC-Derived Myeloid Lineage Model of Herpes Virus Latency and Reactivation. Front. Microbiol. 2019, 10, 2233. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rossetto, C.C.; Tarrant-Elorza, M.; Pari, G.S. Cis and trans acting factors involved in human cytomegalovirus experimental and natural latent infection of CD14 (+) monocytes and CD34 (+) cells. PLoS Pathog. 2013, 9, e1003366. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grey, F.; Nelson, J. Identification and function of human cytomegalovirus microRNAs. J. Clin. Virol. 2008, 41, 186–191. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lau, B.; Poole, E.; Krishna, B.; Sellart, I.; Wills, M.R.; Murphy, E.; Sinclair, J. The Expression of Human Cytomegalovirus MicroRNA MiR-UL148D during Latent Infection in Primary Myeloid Cells Inhibits Activin A-triggered Secretion of IL-6. Sci. Rep. 2016, 6, 31205. [Google Scholar] [CrossRef]
- Fu, M.; Gao, Y.; Zhou, Q.; Zhang, Q.; Peng, Y.; Tian, K.; Wang, J.; Zheng, X. Human cytomegalovirus latent infection alters the expression of cellular and viral microRNA. Gene 2014, 536, 272–278. [Google Scholar] [CrossRef]
- Shen, Z.Z.; Pan, X.; Miao, L.F.; Ye, H.Q.; Chavanas, S.; Davrinche, C.; McVoy, M.; Luo, M.H. Comprehensive analysis of human cytomegalovirus microRNA expression during lytic and quiescent infection. PLoS ONE 2014, 9, e88531. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- O’Connor, C.M.; Vanicek, J.; Murphy, E.A. Host microRNA regulation of human cytomegalovirus immediate early protein translation promotes viral latency. J. Virol. 2014, 88, 5524–5532. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Poole, E.; McGregor Dallas, S.R.; Colston, J.; Joseph, R.S.; Sinclair, J. Virally induced changes in cellular microRNAs maintain latency of human cytomegalovirus in CD34 progenitors. J. Gen. Virol. 2011, 92 (Pt 7), 1539–1549. [Google Scholar] [CrossRef]
- Slobedman, B.; Stern, J.L.; Cunningham, A.L.; Abendroth, A.; Abate, D.A.; Mocarski, E.S. Impact of human cytomegalovirus latent infection on myeloid progenitor cell gene expression. J. Virol. 2004, 78, 4054–4062. [Google Scholar] [CrossRef] [Green Version]
- Chan, G.; Bivins-Smith, E.R.; Smith, M.S.; Smith, P.M.; Yurochko, A.D. Transcriptome analysis reveals human cytomegalovirus reprograms monocyte differentiation toward an M1 macrophage. J. Immunol. 2008, 181, 698–711. [Google Scholar] [CrossRef] [PubMed]
- Zhu, D.; Pan, C.; Sheng, J.; Liang, H.; Bian, Z.; Liu, Y.; Trang, P.; Wu, J.; Liu, F.; Zhang, C.Y.; et al. Human cytomegalovirus reprogrammes haematopoietic progenitor cells into immunosuppressive monocytes to achieve latency. Nat. Microbiol. 2018, 3, 503–513. [Google Scholar] [CrossRef] [PubMed]
- Roche, K.L.; Nukui, M.; Krishna, B.A.; O’Connor, C.M.; Murphy, E.A. Selective 4-Thiouracil Labeling of RNA Transcripts within Latently Infected Cells after Infection with Human Cytomegalovirus Expressing Functional Uracil Phosphoribosyltransferase. J. Virol. 2018, 92. [Google Scholar] [CrossRef] [Green Version]
- Galinato, M.; Shimoda, K.; Aguiar, A.; Hennig, F.; Boffelli, D.; McVoy, M.A.; Hertel, L. Single-Cell Transcriptome Analysis of CD34(+) Stem Cell-Derived Myeloid Cells Infected With Human Cytomegalovirus. Front. Microbiol. 2019, 10, 577. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shnayder, M.; Nachshon, A.; Rozman, B.; Bernshtein, B.; Lavi, M.; Fein, N.; Poole, E.; Avdic, S.; Blyth, E.; Gottlieb, D.; et al. Single cell analysis reveals human cytomegalovirus drives latently infected cells towards an anergic-like monocyte state. Elife 2020, 9, e52168. [Google Scholar] [CrossRef]
- Poole, E.; Avdic, S.; Hodkinson, J.; Jackson, S.; Wills, M.; Slobedman, B.; Sinclair, J. Latency-associated viral interleukin-10 (IL-10) encoded by human cytomegalovirus modulates cellular IL-10 and CCL8 Secretion during latent infection through changes in the cellular microRNA hsa-miR-92a. J. Virol. 2014, 88, 13947–13955. [Google Scholar] [CrossRef] [Green Version]
- Mason, G.M.; Poole, E.; Sissons, J.G.; Wills, M.R.; Sinclair, J.H. Human cytomegalovirus latency alters the cellular secretome, inducing cluster of differentiation (CD)4+ T-cell migration and suppression of effector function. Proc. Natl. Acad. Sci. USA 2012, 109, 14538–14543. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stevenson, E.V.; Collins-McMillen, D.; Kim, J.H.; Cieply, S.J.; Bentz, G.L.; Yurochko, A.D. HCMV reprogramming of infected monocyte survival and differentiation: A Goldilocks phenomenon. Viruses 2014, 6, 782–807. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Poole, E.; Lau, J.C.; Sinclair, J. Latent infection of myeloid progenitors by human cytomegalovirus protects cells from FAS-mediated apoptosis through the cellular IL-10/PEA-15 pathway. J. Gen Virol. 2015, 96, 2355–2359. [Google Scholar] [CrossRef] [PubMed]
- Poole, E.; Sinclair, J. Sleepless latency of human cytomegalovirus. Med. Microbiol. Immunol. 2015, 204, 421–429. [Google Scholar] [CrossRef] [Green Version]
- Jenkins, C.; Garcia, W.; Godwin, M.J.; Spencer, J.V.; Stern, J.L.; Abendroth, A.; Slobedman, B. Immunomodulatory properties of a viral homolog of human interleukin-10 expressed by human cytomegalovirus during the latent phase of infection. J. Virol. 2008, 82, 3736–3750. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Elder, E.G.; Krishna, B.A.; Williamson, J.; Lim, E.Y.; Poole, E.; Sedikides, G.X.; Wills, M.; O’Connor, C.M.; Lehner, P.J.; Sinclair, J. Interferon-Responsive Genes Are Targeted during the Establishment of Human Cytomegalovirus Latency. MBio 2019, 10. [Google Scholar] [CrossRef] [Green Version]
- Dumortier, J.; Streblow, D.N.; Moses, A.V.; Jacobs, J.M.; Kreklywich, C.N.; Camp, D.; Smith, R.D.; Orloff, S.L.; Nelson, J.A. Human cytomegalovirus secretome contains factors that induce angiogenesis and wound healing. J. Virol. 2008, 82, 6524–6535. [Google Scholar] [CrossRef] [Green Version]
- Lee, C.H.; Grey, F. Systems Virology and Human Cytomegalovirus: Using High Throughput Approaches to Identify Novel Host-Virus Interactions During Lytic Infection. Front. Cell Infect Microbiol. 2020, 10, 280. [Google Scholar] [CrossRef]
- Arend, K.C.; Lenarcic, E.M.; Vincent, H.A.; Rashid, N.; Lazear, E.; McDonald, I.M.; Gilbert, T.S.; East, M.P.; Herring, L.E.; Johnson, G.L.; et al. Kinome Profiling Identifies Druggable Targets for Novel Human Cytomegalovirus (HCMV) Antivirals. Mol. Cell Proteom. 2017, 16 (Suppl. 1), S263–S276. [Google Scholar] [CrossRef] [Green Version]
- Cohen, Y.; Stern-Ginossar, N. Manipulation of host pathways by human cytomegalovirus: Insights from genome-wide studies. Semin. Immunopathol. 2014, 36, 651–658. [Google Scholar] [CrossRef]
- Aslam, Y.; Williamson, J.; Romashova, V.; Elder, E.; Krishna, B.; Wills, M.; Lehner, P.; Sinclair, J.; Poole, E. Human Cytomegalovirus Upregulates Expression of HCLS1 Resulting in Increased Cell Motility and Transendothelial Migration during Latency. iScience 2019, 20, 60–72. [Google Scholar] [CrossRef] [Green Version]
- Elder, E.; Krishna, B.; Williamson, J.; Aslam, Y.; Farahi, N.; Wood, A.; Romashova, V.; Roche, K.; Murphy, E.; Chilvers, E.; et al. Monocytes latently infected with human cytomegalovirus evade neutrophil killing. Iscience 2019, 20, 60–72. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Skokowa, J.; Klimiankou, M.; Klimenkova, O.; Lan, D.; Gupta, K.; Hussein, K.; Carrizosa, E.; Kusnetsova, I.; Li, Z.; Sustmann, C.; et al. Interactions among HCLS1, HAX1 and LEF-1 proteins are essential for G-CSF-triggered granulopoiesis. Nature Med. 2012, 18, 1550–1559. [Google Scholar] [CrossRef] [PubMed]
- Uruno, T.; Zhang, P.; Liu, J.; Hao, J.J.; Zhan, X. Haematopoietic lineage cell-specific protein 1 (HS1) promotes actin-related protein (Arp) 2/3 complex-mediated actin polymerization. Biochem. J. 2003, 371 (Pt 2), 485–493. [Google Scholar] [CrossRef] [Green Version]
- Mukherjee, S.; Kim, J.; Mooren, O.L.; Shahan, S.T.; Cohan, M.; Cooper, J.A. Role of cortactin homolog HS1 in transendothelial migration of natural killer cells. PLoS ONE 2015, 10, e0118153. [Google Scholar] [CrossRef] [PubMed]
- Ryckman, C.; Vandal, K.; Rouleau, P.; Talbot, M.; Tessier, P.A. Proinflammatory activities of S100: Proteins S100A8, S100A9, and S100A8/A9 induce neutrophil chemotaxis and adhesion. J. Immunol. 2003, 170, 3233–3242. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lau, B.; Poole, E.; Van Damme, E.; Bunkens, L.; Sowash, M.; King, H.; Murphy, E.; Wills, M.; Van Loock, M.; Sinclair, J. Human cytomegalovirus miR-UL112-1 promotes the down-regulation of viral immediate early-gene expression during latency to prevent T-cell recognition of latently infected cells. J. Gen. Virol. 2016, 97, 2387–2398. [Google Scholar] [CrossRef]
- Poole, E.L.; Kew, V.G.; Lau, J.C.H.; Murray, M.J.; Stamminger, T.; Sinclair, J.H.; Reeves, M.B. A Virally Encoded DeSUMOylase Activity Is Required for Cytomegalovirus Reactivation from Latency. Cell Rep. 2018, 24, 594–606. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Krishna, B.A.; Poole, E.L.; Jackson, S.E.; Smit, M.J.; Wills, M.R.; Sinclair, J.H. Latency-Associated Expression of Human Cytomegalovirus US28 Attenuates Cell Signaling Pathways To Maintain Latent Infection. MBio 2017, 8. [Google Scholar] [CrossRef] [Green Version]
- Krishna, B.A.; Spiess, K.; Poole, E.L.; Lau, B.; Voigt, S.; Kledal, T.N.; Rosenkilde, M.M.; Sinclair, J.H. Targeting the latent cytomegalovirus reservoir with an antiviral fusion toxin protein. Nat. Commun. 2017, 8, 14321. [Google Scholar] [CrossRef] [Green Version]
- Crawford, L.B.; Caposio, P.; Kreklywich, C.; Pham, A.H.; Hancock, M.H.; Jones, T.A.; Smith, P.P.; Yurochko, A.D.; Nelson, J.A.; Streblow, D.N. Human Cytomegalovirus US28 Ligand Binding Activity Is Required for Latency in CD34(+) Hematopoietic Progenitor Cells and Humanized NSG Mice. MBio 2019, 10, e01889-19. [Google Scholar] [CrossRef] [Green Version]
- Krishna, B.A.; Wass, A.B.; Sridharan, R.; O’Connor, C.M. The Requirement for US28 During Cytomegalovirus Latency Is Independent of US27 and US29 Gene Expression. Front. Cell Infect. Microbiol. 2020, 10, 186. [Google Scholar] [CrossRef] [PubMed]
- Reeves, M.B.; Breidenstein, A.; Compton, T. Human cytomegalovirus activation of ERK and myeloid cell leukemia-1 protein correlates with survival of latently infected cells. Proc. Natl. Acad. Sci. USA 2012, 109, 588–593. [Google Scholar] [CrossRef] [Green Version]
- Dutta, D.; Dutta, S.; Veettil, M.V.; Roy, A.; Ansari, M.A.; Iqbal, J.; Chikoti, L.; Kumar, B.; Johnson, K.E.; Chandran, B. BRCA1 Regulates IFI16 Mediated Nuclear Innate Sensing of Herpes Viral DNA and Subsequent Induction of the Innate Inflammasome and Interferon-beta Responses. PLoS Pathog. 2015, 11, e1005030. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kerur, N.; Veettil, M.V.; Sharma-Walia, N.; Bottero, V.; Sadagopan, S.; Otageri, P.; Chandran, B. IFI16 acts as a nuclear pathogen sensor to induce the inflammasome in response to Kaposi Sarcoma-associated herpesvirus infection. Cell Host Microbe 2011, 9, 363–375. [Google Scholar] [CrossRef] [Green Version]
- Li, T.; Chen, J.; Cristea, I.M. Human cytomegalovirus tegument protein pUL83 inhibits IFI16-mediated DNA sensing for immune evasion. Cell Host Microbe 2013, 14, 591–599. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cristea, I.M.; Moorman, N.J.; Terhune, S.S.; Cuevas, C.D.; O’Keefe, E.S.; Rout, M.P.; Chait, B.T.; Shenk, T. Human cytomegalovirus pUL83 stimulates activity of the viral immediate-early promoter through its interaction with the cellular IFI16 protein. J. Virol. 2010, 84, 7803–7814. [Google Scholar] [CrossRef] [Green Version]
- Biolatti, M.; Dell’Oste, V.; Pautasso, S.; von Einem, J.; Marschall, M.; Plachter, B.; Gariglio, M.; De Andrea, M.; Landolfo, S. Regulatory Interaction between the Cellular Restriction Factor IFI16 and Viral pp65 (pUL83) Modulates Viral Gene Expression and IFI16 Protein Stability. J. Virol. 2016, 90, 8238–8250. [Google Scholar] [CrossRef] [Green Version]
- Merkl, P.E.; Orzalli, M.H.; Knipe, D.M. Mechanisms of Host IFI16, PML, and Daxx Protein Restriction of Herpes Simplex Virus 1 Replication. J. Virol. 2018, 92. [Google Scholar] [CrossRef] [Green Version]
- Cai, H.; Yan, L.; Liu, N.; Xu, M.; Cai, H. IFI16 promotes cervical cancer progression by upregulating PD-L1 in immunomicroenvironment through STING-TBK1-NF-kB pathway. Biomed. Pharmacother. 2020, 123, 109790. [Google Scholar] [CrossRef]
- Dunphy, G.; Flannery, S.M.; Almine, J.F.; Connolly, D.J.; Paulus, C.; Jonsson, K.L.; Jakobsen, M.R.; Nevels, M.M.; Bowie, A.G.; Unterholzner, L. Non-canonical Activation of the DNA Sensing Adaptor STING by ATM and IFI16 Mediates NF-kappaB Signaling after Nuclear DNA Damage. Mol. Cell 2018, 71, 745–760.e5. [Google Scholar] [CrossRef] [Green Version]
- Bawadekar, M.; De Andrea, M.; Lo Cigno, I.; Baldanzi, G.; Caneparo, V.; Graziani, A.; Landolfo, S.; Gariglio, M. The Extracellular IFI16 Protein Propagates Inflammation in Endothelial Cells Via p38 MAPK and NF-kappaB p65 Activation. J. Interferon Cytokine Res. 2015, 35, 441–453. [Google Scholar] [CrossRef]
- Cheung, A.K.; Abendroth, A.; Cunningham, A.L.; Slobedman, B. Viral gene expression during the establishment of human cytomegalovirus latent infection in myeloid progenitor cells. Blood 2006, 108, 3691–3699. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.H.; Albright, E.R.; Lee, J.H.; Jacobs, D.; Kalejta, R.F. Cellular defense against latent colonization foiled by human cytomegalovirus UL138 protein. Sci. Adv. 2015, 1, e1501164. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Qi, Y.; Ma, Y.; He, R.; Ji, Y.; Sun, Z.; Ruan, Q. Down-regulation of human cytomegalovirus UL138, a novel latency-associated determinant, by hcmv-miR-UL36. J. Biosci. 2013, 38, 479–485. [Google Scholar] [CrossRef] [PubMed]
- Han, L.; Ma, Y.; Liu, Z.; Liu, C.; Lu, Y.; Qi, Y.; Huang, Y.; Sun, Z.; Ruan, Q. Transcriptional regulation and influence on replication of the human cytomegalovirus UL138 1.4 kb transcript. Mol. Med. Rep. 2017, 16, 5649–5658. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.H.; Caviness, K.; Albright, E.R.; Lee, J.H.; Gelbmann, C.B.; Rak, M.; Goodrum, F.; Kalejta, R.F. Long and Short Isoforms of the Human Cytomegalovirus UL138 Protein Silence IE Transcription and Promote Latency. J. Virol. 2016, 90, 9483–9494. [Google Scholar] [CrossRef] [Green Version]
- Gelbmann, C.B.; Kalejta, R.F. The Golgi sorting motifs of human cytomegalovirus UL138 are not required for latency maintenance. Virus Res. 2019, 270, 197646. [Google Scholar] [CrossRef]
- Gelbmann, C.B.; Kalejta, R.F. The Membrane-Spanning Peptide and Acidic Cluster Dileucine Sorting Motif of UL138 Are Required To Downregulate MRP1 Drug Transporter Function in Human Cytomegalovirus-Infected Cells. J. Virol. 2019, 93. [Google Scholar] [CrossRef] [Green Version]
- Weekes, M.P.; Tan, S.Y.; Poole, E.; Talbot, S.; Antrobus, R.; Smith, D.L.; Montag, C.; Gygi, S.P.; Sinclair, J.H.; Lehner, P.J. Latency-associated degradation of the MRP1 drug transporter during latent human cytomegalovirus infection. Science 2013, 340, 199–202. [Google Scholar] [CrossRef] [Green Version]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Poole, E.; Sinclair, J. Understanding HCMV Latency Using Unbiased Proteomic Analyses. Pathogens 2020, 9, 590. https://doi.org/10.3390/pathogens9070590
Poole E, Sinclair J. Understanding HCMV Latency Using Unbiased Proteomic Analyses. Pathogens. 2020; 9(7):590. https://doi.org/10.3390/pathogens9070590
Chicago/Turabian StylePoole, Emma, and John Sinclair. 2020. "Understanding HCMV Latency Using Unbiased Proteomic Analyses" Pathogens 9, no. 7: 590. https://doi.org/10.3390/pathogens9070590
APA StylePoole, E., & Sinclair, J. (2020). Understanding HCMV Latency Using Unbiased Proteomic Analyses. Pathogens, 9(7), 590. https://doi.org/10.3390/pathogens9070590




