Identification of Antimicrobial Resistance-Associated Genes through Whole Genome Sequencing of Mycoplasma bovis Isolates with Different Antimicrobial Resistances
Abstract
1. Introduction
2. Results
2.1. MIC Testing
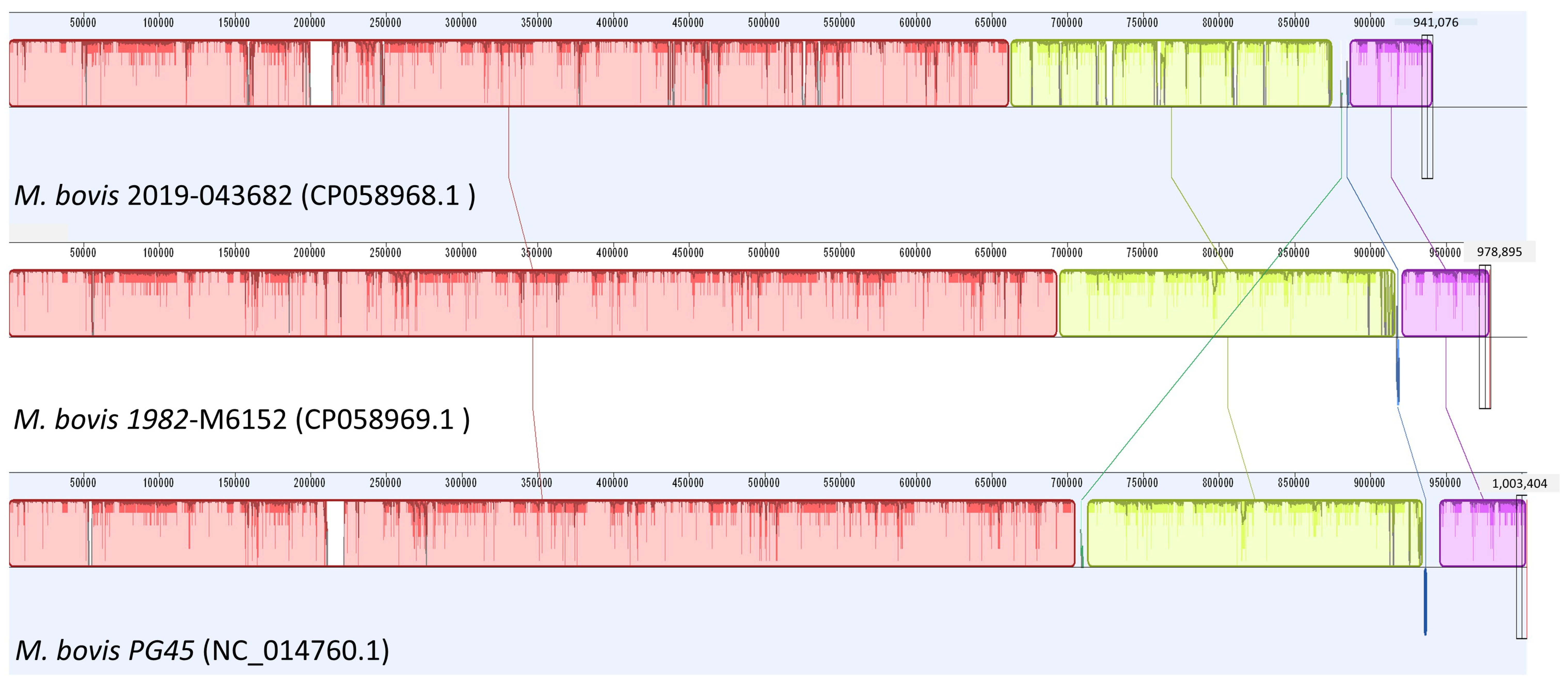
2.2. Whole-Genome Sequencing
3. Discussion
3.1. Protein Synthesis
3.1.1. Methyltransferases
3.1.2. Ribosomal Proteins
3.1.3. Aminoacyl-tRNA Synthetases
3.2. Topoisomerases
3.3. Bacterial Efflux Pumps: ABC Transporters
3.4. Future Directions
4. Materials and Methods
4.1. Culture & Isolation of Mycoplasmas
4.2. MIC Testing
4.3. Nucleic Acid Extraction
4.4. Whole Genome Sequencing & Bioinformatics
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Caswell, J.L.; Bateman, K.G.; Cai, H.Y.; Castillo-Alcala, F. Mycoplasma bovis in Respiratory Disease of Feedlot Cattle. Vet. Clin. N. Am. Food Anim. Pract. 2010, 26, 365–379. [Google Scholar] [CrossRef]
- Griffin, D.; Chengappa, M.M.; Kuszak, J.; McVey, D.S. Bacterial Pathogens of the Bovine Respiratory Disease Complex. Vet. Clin. N. Am. Food Anim. Pract. 2010, 26, 381–394. [Google Scholar] [CrossRef]
- Cai, H.Y.; McDowall, R.; Parker, L.; Kaufman, E.I.; Caswell, J.L. Changes in antimicrobial susceptibility profiles of Mycoplasma bovis over time. Can. J. Vet. Res. 2019, 83, 34–41. [Google Scholar]
- Byrne, W.J.; McCormack, R.; Egan, J.; Brice, N.; Ball, H.J.; Markey, B. Isolation of Mycoplasma bovis from bovine clinical samples in the Republic of Ireland. Vet. Rec. 2001, 148, 331–333. [Google Scholar] [CrossRef]
- Rosenbusch, R.F.; Kinyon, J.M.; Apley, M.; Funk, N.D.; Smith, S.; Hoffman, L.J. In Vitro Antimicrobial Inhibition Profiles of Mycoplasma Bovis Isolates Recovered from Various Regions of the United States from 2002 to 2003. J. Vet. Diagn. Investig. 2005, 17, 436–441. [Google Scholar] [CrossRef]
- Lysnyansky, I.; Ayling, R.D. Mycoplasma bovis: Mechanisms of Resistance and Trends in Antimicrobial Susceptibility. Front. Microbiol. 2016, 7, 595. [Google Scholar] [CrossRef]
- Baym, M.; Stone, L.K.; Kishony, R. Multidrug evolutionary strategies to reverse antibiotic resistance. Science 2016, 351, aad3292. [Google Scholar] [CrossRef] [PubMed]
- Owen, J.R.; Noyes, N.; Young, A.E.; Prince, D.J.; Blanchard, P.C.; Lehenbauer, T.W.; Aly, S.S.; Davis, J.H.; O’Rourke, S.M.; Abdo, Z.; et al. Whole-Genome Sequencing and Concordance Between Antimicrobial Susceptibility Genotypes and Phenotypes of Bacterial Isolates Associated with Bovine Respiratory Disease. G3 Genes Genomes Genet. 2017, 7, 3059–3071. [Google Scholar] [CrossRef] [PubMed]
- Sulyok, K.M.; Kreizinger, Z.; Wehmann, E.; Lysnyansky, I.; Bányai, K.; Marton, S.; Jerzsele, Á.; Rónai, Z.; Turcsányi, I.; Makrai, L.; et al. Mutations Associated with Decreased Susceptibility to Seven Antimicrobial Families in Field and Laboratory-Derived Mycoplasma bovis Strains. Antimicrob. Agents Chemother. 2017, 61. [Google Scholar] [CrossRef] [PubMed]
- Sato, T.; Higuchi, H.; Yokota, S.; Tamura, Y. Mycoplasma bovis isolates from dairy calves in Japan have less susceptibility than a reference strain to all approved macrolides associated with a point mutation (G748A) combined with multiple species-specific nucleotide alterations in 23S rRNA. Microbiol. Immunol. 2017, 61, 215–224. [Google Scholar] [CrossRef] [PubMed]
- Nurk, S.; Bankevich, A.; Antipov, D.; Gurevich, A.; Korobeynikov, A.; Lapidus, A.; Prjibelsky, A.; Pyshkin, A.; Sirotkin, A.; Sirotkin, Y.; et al. Assembling Genomes and Mini-metagenomes from Highly Chimeric Reads. In Proc Research Comp Molecular Bio; Deng, M., Jiang, R., Sun, F., Zhang, X., Eds.; Springer: Berlin/ Heidelberg, Germany, 2013; pp. 158–170. [Google Scholar]
- Thomsen, M.C.F.; Ahrenfeldt, J.; Cisneros, J.L.B.; Jurtz, V.; Larsen, M.V.; Hasman, H.; Aarestrup, F.M.; Lund, O. A Bacterial Analysis Platform: An Integrated System for Analysing Bacterial Whole Genome Sequencing Data for Clinical Diagnostics and Surveillance. PLoS ONE 2016, 11, e0157718. [Google Scholar] [CrossRef] [PubMed]
- Alcock, B.P.; Raphenya, A.R.; Lau, T.T.Y.; Tsang, K.K.; Bouchard, M.; Edalatmand, A.; Huynh, W.; Nguyen, A.-L.V.; Cheng, A.A.; Liu, S.; et al. CARD 2020: antibiotic resistome surveillance with the comprehensive antibiotic resistance database. Nucleic Acids Res. 2020, 48, D517–D525. [Google Scholar] [CrossRef] [PubMed]
- Perichon, B.; Tankovic, J.; Courvalin, P. Characterization of a mutation in the parE gene that confers fluoroquinolone resistance in Streptococcus pneumoniae. Antimicrob. Agents Chemother. 1997, 41, 1166–1167. [Google Scholar] [CrossRef]
- Michel, G.; Sauvé, V.; Larocque, R.; Li, Y.; Matte, A.; Cygler, M. The Structure of the RlmB 23S rRNA Methyltransferase Reveals a New Methyltransferase Fold with a Unique Knot. Structure 2002, 10, 1303–1315. [Google Scholar] [CrossRef]
- Fyfe, C.; Grossman, T.H.; Kerstein, K.; Sutcliffe, J. Resistance to Macrolide Antibiotics in Public Health Pathogens. Cold Spring Harb. Perspect. Med. 2016, 6, a025395. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Jonker, M.J.; Moustakas, I.; Brul, S.; Ter Kuile, B.H. Dynamics of Mutations during Development of Resistance by Pseudomonas aeruginosa against Five Antibiotics. Antimicrob. Agents Chemother. 2016, 60, 4229–4236. [Google Scholar] [CrossRef]
- Grossman, T.H. Tetracycline Antibiotics and Resistance. Cold Spring Harb. Perspect. Med. 2016, 6, a025387. [Google Scholar] [CrossRef]
- Wang, Z.; Kong, L.C.; Jia, B.Y.; Liu, S.M.; Jiang, X.Y.; Ma, H.X. Aminoglycoside susceptibility of Pasteurella multocida isolates from bovine respiratory infections in China and mutations in ribosomal protein S5 associated with high-level induced spectinomycin resistance. J. Vet. Med. Sci. 2017, 79, 1678–1681. [Google Scholar] [CrossRef]
- Hu, M.; Nandi, S.; Davies, C.; Nicholas, R.A. High-Level Chromosomally Mediated Tetracycline Resistance in Neisseria gonorrhoeae Results from a Point Mutation in the rpsJ Gene Encoding Ribosomal Protein S10 in Combination with the mtrR and penB Resistance Determinants. Antimicrob. Agents Chemother. 2005, 49, 4327–4334. [Google Scholar] [CrossRef]
- Long, K.S.; Hansen, L.H.; Jakobsen, L.; Vester, B. Interaction of Pleuromutilin Derivatives with the Ribosomal Peptidyl Transferase Center. Antimicrob. Agents Chemother. 2006, 50, 1458–1462. [Google Scholar] [CrossRef]
- Hölzel, C.S.; Harms, K.S.; Schwaiger, K.; Bauer, J. Resistance to Linezolid in a Porcine Clostridium perfringens Strain Carrying a Mutation in the rplD Gene Encoding the Ribosomal Protein L4. Antimicrob. Agents Chemother. 2010, 54, 1351–1353. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Cagliero, C.; Mouline, C.; Cloeckaert, A.; Payot, S. Synergy between Efflux Pump CmeABC and Modifications in Ribosomal Proteins L4 and L22 in Conferring Macrolide Resistance in Campylobacter jejuni and Campylobacter coli. Antimicrob. Agents Chemother. 2006, 50, 3893–3896. [Google Scholar] [CrossRef] [PubMed]
- Liu, A.; Tran, L.; Becket, E.; Lee, K.; Chinn, L.; Park, E.; Tran, K.; Miller, J.H. Antibiotic Sensitivity Profiles Determined with an Escherichia coli Gene Knockout Collection: Generating an Antibiotic Bar Code. Antimicrob. Agents Chemother. 2010, 54, 1393–1403. [Google Scholar] [CrossRef] [PubMed]
- Milija, J.; Lilic, M.; Janjusevic, R.; Jovanovic, G.; Savic, D.J. tRNA Synthetase Mutants of Escherichia coli K-12 Are Resistant to the Gyrase Inhibitor Novobiocin. J. Bacteriol. 1999, 181, 2979–2983. [Google Scholar] [CrossRef]
- Magalhães, S.; Aroso, M.; Roxo, I.; Ferreira, S.; Cerveira, F.; Ramalheira, E.; Ferreira, R.; Vitorino, R. Proteomic profile of susceptible and multidrug-resistant clinical isolates of Escherichia coli and Klebsiella pneumoniae using label-free and immunoproteomic strategies. Res. Microbiol. 2017, 168, 222–233. [Google Scholar] [CrossRef] [PubMed]
- Yanagisawa, T.; Lee, J.T.; Wu, H.C.; Kawakami, M. Relationship of protein structure of isoleucyl-tRNA synthetase with pseudomonic acid resistance of Escherichia coli. A proposed mod of action of pseudomonic acid as an inhibitor of isoleucyl-tRNA synthetase. J. Biol. Chem. 1994, 269, 24304–24309. [Google Scholar]
- Dordel, J.; Kim, C.; Chung, M.; Pardos de la Gándara, M.; Holden, M.T.J.; Parkhill, J.; de Lencastre, H.; Bentley, S.D.; Tomasz, A. Novel Determinants of Antibiotic Resistance: Identification of Mutated Loci in Highly Methicillin-Resistant Subpopulations of Methicillin-Resistant Staphylococcus aureus. MBio 2014, 5. [Google Scholar] [CrossRef]
- Liu, M.; Douthwaite, S. Resistance to the macrolide antibiotic tylosin is conferred by single methylations at 23S rRNA nucleotides G748 and A2058 acting in synergy. Proc. Natl. Acad. Sci. USA 2002, 99, 14658–14663. [Google Scholar] [CrossRef]
- Doi, Y.; Arakawa, Y. 16S Ribosomal RNA Methylation: Emerging Resistance Mechanism against Aminoglycosides. Clin. Infect. Dis. 2007, 45, 88–94. [Google Scholar] [CrossRef]
- Masuda, I.; Matsubara, R.; Christian, T.; Rojas, E.R.; Yadavalli, S.S.; Zhang, L.; Goulian, M.; Foster, L.J.; Huang, K.C.; Hou, Y.-M. tRNA Methylation Is a Global Determinant of Bacterial Multi-drug Resistance. Cell Syst. 2019, 8, 302–314. [Google Scholar] [CrossRef]
- Giguère, S. Lincosamides, Pleuromutilins, and Streptogramins. In Antimicrobial Therapy in Veterinary Medicine; John Wiley & Sons Ltd: Hoboken, NJ, USA, 2013; pp. 199–210. ISBN 978-1-118-67501-4. [Google Scholar]
- Hurdle, J.G.; O’Neill, A.J.; Chopra, I. Prospects for Aminoacyl-tRNA Synthetase Inhibitors as New Antimicrobial Agents. Antimicrob. Agents Chemother. 2005, 49, 4821–4833. [Google Scholar] [CrossRef] [PubMed]
- Tse-Dinh, Y.-C. Bacterial topoisomerase I as a target for discovery of antibacterial compounds. Nucleic Acids Res. 2009, 37, 731–737. [Google Scholar] [CrossRef] [PubMed]
- Bansal, S.; Tawar, U.; Singh, M.; Nikravesh, A.; Good, L.; Tandon, V. Old class but new dimethoxy analogue of benzimidazole: A bacterial topoisomerase I inhibitor. Int. J. Antimicrob. Agents 2010, 35, 186–190. [Google Scholar] [CrossRef] [PubMed]
- Nagaraja, V.; Godbole, A.A.; Henderson, S.R.; Maxwell, A. DNA topoisomerase I and DNA gyrase as targets for TB therapy. Drug Discov. Today 2017, 22, 510–518. [Google Scholar] [CrossRef]
- Webber, M.A.; Piddock, L.J.V. The importance of efflux pumps in bacterial antibiotic resistance. J Antimicrob. Chemother. 2003, 51, 9–11. [Google Scholar] [CrossRef]
- Marquez, B. Bacterial efflux systems and efflux pumps inhibitors. Biochimie 2005, 87, 1137–1147. [Google Scholar] [CrossRef]
- Ruhnke, H.L.; Rosendal, S. Useful Protocols for Diagnosis of Animal Mycoplasmas. In Mycoplasmosis in Animals: Laboratory Diagnosis; Whitford, H.W., Rosenbusch, R.F., Lauerman, L.H., Eds.; Iowa State University Press: Ames, IA, USA, 1994; pp. 141–166. [Google Scholar]
| Antimicrobial | PG45 | 2019-043682 | 1982-M6152 |
|---|---|---|---|
| Neomycin | >32 | >32 | >32 |
| Spectinomycin | <8 | 16 | <8 |
| Trimethoprim/Sulfa | >2/38 | >2/38 | >2/38 |
| Danofloxacin | 0.5 | >1 | 0.5 |
| Enrofloxacin | 0.5 | 2 | 0.5 |
| Clindamycin | <0.25 | >16 | <0.25 |
| Tilmicosin | <4 | >64 | <4 |
| Tulathromycin | 8 | >64 | 8 |
| Tylosin Tartrate | 1 | >32 | 1 |
| Tiamulin | 1 | 8 | 1 |
| Gentamicin | 8 | 16 | 8 |
| Florfenicol | 4 | >8 | 4 |
| Sulphadimethoxine | >256 | >256 | >256 |
| Chlortetracycline | <0.5 | >8 | <0.5 |
| Oxytetracycline | <0.5 | >8 | 1 |
| Ceftiofur | >8 | >8 | >8 |
| Genetic Events | 1982-M6152 | 2019-043682 | |
|---|---|---|---|
| Variation types | Identical | 696 | 183 |
| Deletion | 8 | 18 | |
| Deleted_end_3prime | 3 | 3 | |
| Deleted_end_5prime | 1 | 3 | |
| Indel | 3 | 4 | |
| Insertion | 22 | 22 | |
| Not_Mapped | 39 | 173 | |
| Substitution | 105 | 471 | |
| TOTAL | 877 | 877 | |
| Unmapped features | Excised (<5% coverage) | 24 | 81 |
| Truncated (5–95% coverage) | 11 | 48 | |
| Highly variable (>95% coverage) | 4 | 44 | |
| TOTAL | 39 | 173 | |
| Functional Role: | Gene: | 1982-M6152 | 2019-043682 | Associated AMR: | Reference: | |
|---|---|---|---|---|---|---|
| Topoisomerases | gyrA | 0 | 8 | fluoroquinolones | [9] | |
| gyrB | 0 | 6 | fluoroquinolones | [9] | ||
| parC | 1 * | 19 * | fluoroquinolones | [9] | ||
| parE | 0 | 4 | fluoroquinolones | [14] | ||
| topA | 0 | 1 ^ | ||||
| Protein Synthesis: | ||||||
| Methyltransferases: | MBOVPG45_RS00465 | 0 | 2 | |||
| MBOVPG45_RS00470 | 0 | 7 | ||||
| MBOVPG45_RS02280 | 0 | 4 | ||||
| rlmB | 0 | 5 | Predicted AMR | [15] | ||
| rlmD | 0 | 11 | ||||
| rlmH | 0 | 1 | ||||
| rsmA | 0 | 6 | aminoglycosides | [16] | ||
| rsmD | 0 | 1 | aminoglycosides | [16] | ||
| rsmH | 0 | 5 | aminoglycosides | [16] | ||
| rsmI | 0 | 5 | aminoglycosides | [16] | ||
| trmB | 0 | 4 ^ | ||||
| 30S Ribosomal Proteins | rpsB | 0 | 3 | aminoglycosides | [17] | |
| rpsC | 2 | 1 | tetracyclines | [18] | ||
| rpsD | 0 | 2 | ||||
| rpsE | 1 | 2 | aminoglycosides | [19] | ||
| rpsH | 0 | 1 | ||||
| rpsJ | 1 | 0 | tetracyclines | [20] | ||
| rpsP | 0 | 2 | ||||
| rpsS | 0 | 1 | ||||
| rbfA | 0 | 1 | ||||
| 50S Ribosomal Proteins | MBOVPG45_RS00445 | 0 | 1 | |||
| rplB | 0 | 1 | ||||
| rplC | 0 | 1 | pleuromutilins | [21] | ||
| rplD | 0 | 10 | linezolid | [22] | ||
| MBOVPG45_RS03525 | 0 | 2 | ||||
| rplV | 0 | 1 | macrolides | [23] | ||
| MBOVPG45_RS01360 | 0 | 1 | ||||
| rpmE | 0 | 1 | MDR | [24] | ||
| tRNA ligases | alaS | 1 | 8 ^ | novobiocin | [25] | |
| MBOVPG45_RS01640 | 0 | 5 | ||||
| asnS | 0 | 1 | multi-drug resistance | [26] | ||
| MBOVPG45_RS00205 | 0 | 9 | ||||
| MBOVPG45_RS01150 | 0 | 6 ^ | ||||
| MBOVPG45_RS02730 | 0 | 2 | ||||
| MBOVPG45_RS02640 | 0 | 1 | ||||
| ileS | 1 | 5 ^ | pseudomonic acid | [27] | ||
| MBOVPG45_RS03145 | 0 | 1 | ||||
| MBOVPG45_RS02255 | 0 | 16 | ||||
| lysS | 0 | 3 | methicillin | [28] | ||
| MBOVPG45_RS03150 | 0 | 10 | ||||
| pheS | 0 | 2 | MDR | [26] | ||
| MBOVPG45_RS00380 | 0 | 18 | ||||
| serS | 0 | 1 | ||||
| tRNA ligases | MBOVPG45_RS02170 | 1 | 8 ^ | |||
| trpS | 0 | 1 | ||||
| MBOVPG45_RS04210 | 0 | 9 ^ | ||||
| MBOVPG45_RS00740 | 0 | 7 ^ | ||||
| tilS | 0 | 8 | ||||
| thiI | 0 | 4 | ||||
| mnmA | 0 | 5 | ||||
| Gene: | 1982-M6152 | 2019-043682 | Description: |
|---|---|---|---|
| MBOVPG45_RS00090 | 0 | 1 | ABC transporter ATP-binding protein |
| MBOVPG45_RS00180 | 0 | 2 | ABC transporter permease |
| MBOVPG45_RS00555 | 0 | 2 | ABC transporter permease |
| MBOVPG45_RS00570 | 0 | 4 | ATP-binding cassette domain-containing protein |
| MBOVPG45_RS00600 | 0 | 1 | ATP-binding cassette domain-containing protein |
| MBOVPG45_RS01485 | 0 | 2 | energy-coupling factor transporter transmembrane protein EcfT |
| MBOVPG45_RS01540 | 0 | 1 | sugar ABC transporter permease |
| MBOVPG45_RS01545 | 0 | 4 | ATP-binding cassette domain-containing protein |
| MBOVPG45_RS01720 | 0 | 1 | ABC transporter permease subunit |
| MBOVPG45_RS01770 | 0 | 1 | ABC transporter ATP-binding protein |
| MBOVPG45_RS01775 | 1 | 7 | ABC transporter permease |
| MBOVPG45_RS02005 | 0 | 5 | ABC transporter ATP-binding protein |
| MBOVPG45_RS02710 | 0 | 1 | ABC transporter permease subunit |
| MBOVPG45_RS02715 | 0 | 2 | ATP-binding cassette domain-containing protein |
| MBOVPG45_RS02905 | 1 | 1 | ABC transporter permease subunit |
| MBOVPG45_RS03425 | 0 | 1 | ATP-binding cassette domain-containing protein |
| MBOVPG45_RS03465 | 0 | 4 | ABC transporter ATP-binding protein |
| MBOVPG45_RS03470 | 0 | 6 | ABC transporter ATP-binding protein |
| MBOVPG45_RS03705 | 0 | 6 ^ | carbohydrate ABC transporter permease |
| MBOVPG45_RS03710 | 0 | 1 | sugar ABC transporter permease |
| MBOVPG45_RS04310 | 0 | 5 | ABC transporter ATP-binding protein |
| MBOVPG45_RS04315 | 1 | 89 | ABC transporter permease |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ledger, L.; Eidt, J.; Cai, H.Y. Identification of Antimicrobial Resistance-Associated Genes through Whole Genome Sequencing of Mycoplasma bovis Isolates with Different Antimicrobial Resistances. Pathogens 2020, 9, 588. https://doi.org/10.3390/pathogens9070588
Ledger L, Eidt J, Cai HY. Identification of Antimicrobial Resistance-Associated Genes through Whole Genome Sequencing of Mycoplasma bovis Isolates with Different Antimicrobial Resistances. Pathogens. 2020; 9(7):588. https://doi.org/10.3390/pathogens9070588
Chicago/Turabian StyleLedger, Lisa, Jason Eidt, and Hugh Y. Cai. 2020. "Identification of Antimicrobial Resistance-Associated Genes through Whole Genome Sequencing of Mycoplasma bovis Isolates with Different Antimicrobial Resistances" Pathogens 9, no. 7: 588. https://doi.org/10.3390/pathogens9070588
APA StyleLedger, L., Eidt, J., & Cai, H. Y. (2020). Identification of Antimicrobial Resistance-Associated Genes through Whole Genome Sequencing of Mycoplasma bovis Isolates with Different Antimicrobial Resistances. Pathogens, 9(7), 588. https://doi.org/10.3390/pathogens9070588





