Morphology and Multi-Gene Phylogeny Reveal Pestalotiopsis pinicola sp. nov. and a New Host Record of Cladosporium anthropophilum from Edible Pine (Pinus armandii) Seeds in Yunnan Province, China
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection and Specimen Examination
Isolation
2.2. DNA Extraction, PCR Amplification, and DNA Sequencing
2.3. Phylogenetic Analyses
3. Results
3.1. Phylogenetic Analysis of Combined Sequence Data
3.2. Taxonomy
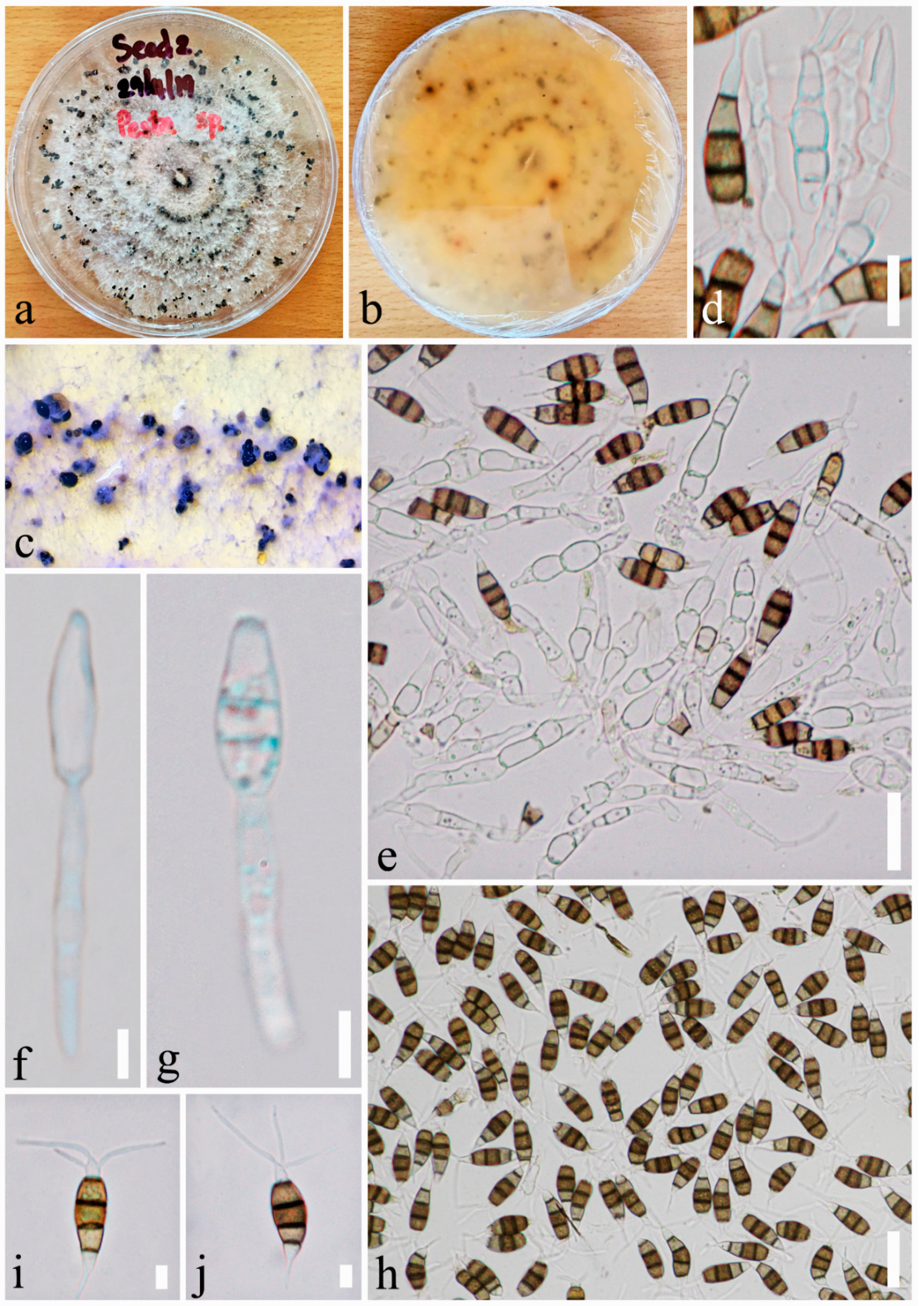
- Index Fungorum number: IF556765, Facesoffungi number: FoF 06276, Figure 5.
- Etymology: named after the host genus, Pinus.
- Holotype: MFLU19-2363.
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Anon. Dendrology (Southern Edition); China Forest Press: Beijing, China, 1994. (In Chinese) [Google Scholar]
- Yan, K.; He, X.Y.; Wei, C.; Tao, L.; Zhang, G. Variation of Antioxidant System in Pinus armandii under Elevated O3 in an Entire Growth Season. Acta Hydrochim. Hydrobiol. 2013, 41, 5–10. [Google Scholar]
- Destaillats, F.; Cruzhernandez, C.; Giuffrida, F.; Dionisi, F. Identification of the botanical origin of pine nuts found in food products by gas-liquid chromatography analysis of fatty acid profile. J. Agric. Food Chem. 2010, 58, 2082–2087. [Google Scholar] [CrossRef] [PubMed]
- CCCFNA. Chinese pine nuts. In China Chamber of Commerce of Imports/Exports of Foodstuffs, Native Produce, and Animal by-Products; The International Tea Committee, International Nut and Dried Fruit Council: Reus, Spain, 2011. [Google Scholar]
- Joshi, V.; Mukerji, K.G. Seed-borne mycoflora of two under–exploited legumes: Vigna umbellata and psophocarpus tetragonolobus from Northeastern parts of India. In From Ethnomycology to Fungal Biotechnology; Springer: Boston, MA, USA, 1999; pp. 257–268. [Google Scholar]
- Cram, M.M.; Fraedrich, S.W. Seed diseases and seedborne pathogens of North America. Tree Plant. Notes. 2010, 53, 35–44. [Google Scholar]
- Sutherland, J.R.; Miller, T.; Quinard, R.S. Cone and seed diseases of North American conifers. N. Am. For. Comm. Publ. Number 1 1987, 1, 1–70. [Google Scholar]
- Promputtha, I.; Lumyong, S.; Dhanasekaran, V.; Mckenzie, E.H.C.; Hyde, K.D.; Jeewon, R. A phylogenetic evaluation of whether endophytes become saprotrophs at host senescence. Microb. Ecol. 2007, 53, 579–590. [Google Scholar] [CrossRef]
- Tibpromma, S.; Hyde, K.D.; Bhat, J.D.; Mortimer, P.E.; Xu, J.C.; Promputtha, I.; Doilom, M.; Yang, J.-B.; Tang, A.M.C.; Karunarathna, S.C. Identification of endophytic fungi from leaves of Pandanaceae based on their morphotypes and DNA sequence data from southern Thailand. MycoKeys 2018, 33, 25–67. [Google Scholar] [CrossRef]
- Mittal, R.K.; Anderson, R.L.; Mathur, S.B. Microorganisms associated with tree seeds: World checklist Petawawa National Forestry Institute, Chalk River, Ontario. Available online: https://cfs.nrcan.gc.ca/publications?id=10573 (accessed on 29 November 2019).
- Pittayakhajonwut, P.; Theerasilp, M.; Kongsaeree, P.; Rungrod, A.; Tanticharoen, M.; Thebtaranonth, Y.; Pughiinin, A. A sesquiterpene from the fungus Kionochaeta pughii BCC 3878. Planta Med. 2002, 68, 1017–1019. [Google Scholar] [CrossRef]
- Chinworrungsee, M.; Kittakoop, P.; Isaka, M.; Maithip, P.; Supothina, S.; Thebtaranonth, Y. Isolation and structure elucidation of a novel antimalarial macrocyclic polylactone, menisporopsin A, from the fungus Menisporopsis theobromae. J. Nat. Prod. 2004, 67, 689–692. [Google Scholar] [CrossRef]
- Link, H.F. Observationes in ordines plantarum naturales. Observationes in ordines plantarum naturales. 2. Mag. Gesell. Naturf. Freunde Berlin 1816, 8, 25–45. [Google Scholar]
- Bensch, K.; Braun, U.; Groenewald, J.Z.; Crous, P.W. The genus Cladosporium. Stud. Mycol. 2012, 72, 1–401. [Google Scholar] [CrossRef]
- Crous, P.W.; Shivas, R.G.; Quaedvlieg, W.; van der Bank, M.; Zhang, Y.; Summerell, B.A.; Guarro, J.; Wingfield, M.J.; Wood, A.R.; Alfenas, A.C.; et al. Fungal planet description sheets: 214–280. Persoonia 2013, 31, 188. [Google Scholar] [CrossRef] [PubMed]
- Sandoval-Denis, M.; Genã, J.; Sutton, D.A.; Wiederhold, N.P.; Cano-Lira, J.F.; Guarro, J. New species of Cladosporium associated with human and animal infections. Persoonia 2016, 36, 281–298. [Google Scholar] [CrossRef] [PubMed]
- Ma, R.; Chen, Q.; Fan, Y.; Wang, Q.; Chen, S.F.; Liu, X.Z.; Cai, L.; Yao, B. Six new soil–inhabiting Cladosporium species from plateaus in China. Mycologia 2017, 109, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Tibpromma, S.; Hyde, K.D.; Mckenzie, E.H.C.; Bhat, D.J.; Phillips, A.J.L.; Wanasinghe, D.N.; Samarakoon, M.C.; Jayawardena, R.S.; Dissanayake, A.J.; Tennakoon, D.S.; et al. Fungal diversity notes 840–928: Micro-fungi associated with Pandanaceae. Fungal Divers. 2018, 72, 1–160. [Google Scholar] [CrossRef]
- Fradkin, A.; Tarlo, S.M.; Tobin, R.S.; Tucic–Porretta, M.; Malloch, D. Species identification of airborne molds and its significance for the detection of indoor pollution. Air Repair. 1987, 37, 51–53. [Google Scholar] [CrossRef]
- Flannigan, B.; Samson, R.A.; Miller, J.D. Microorganisms in Home and Indoor Work Environments; CRC Press: Boca Raton, FL, USA, 2011; Volume 3, pp. 55–58. [Google Scholar]
- Bullerman, L.B. SPOILAGE|Fungi in food—An overview. Encycl. Food Sci. Nutr. 2003, 1, 5511–5522. [Google Scholar]
- Horner, W.E.; Worthan, A.G.; Morey, P.R. Air-and dustborne mycoflora in houses free of water damage and fungal growth. Appl. Environ. Microbiol. 2004, 70, 6394–6400. [Google Scholar] [CrossRef]
- Horner, W.E.; Helbling, A.; Salvaggio, J.E.; Lehrer, S.B. Fungal allergens. Clin. Microbiol. Rev. 1995, 8, 161–179. [Google Scholar] [CrossRef]
- Sundell, J. On the history of indoor air quality and health. Indoor Air. 2004, 14, 8–51. [Google Scholar] [CrossRef]
- Matheson, M.C.; Abramson, M.J.; Dharmage, S.C.; Forbes, A.B.; Raven, J.M.; Thien, F.C.; Walters, E.H. Changes in indoor allergen and fungal levels predict changes in asthma activity among young adults. Clin. Exp. Allergy 2010, 35, 907–913. [Google Scholar] [CrossRef] [PubMed]
- Denning, D.W.; O’Driscoll, B.R.; Hogaboam, C.M.; Bowyer, P.; Niven, R.M. The link between fungi and severe asthma: A summary of the evidence. Eur. Respir. J. 2006, 27, 615–626. [Google Scholar] [CrossRef] [PubMed]
- Simonnobbe, B.; Denk, U.; Pöll, V.; Rid, R.; Breitenbach, M. The spectrum of fungal allergy. Int. Arch. Allergy Immunol. 2008, 145, 58–86. [Google Scholar] [CrossRef] [PubMed]
- Mari, A.; Rasi, C.; Palazzo, P.; Scala, E. Allergen databases: Current status and perspectives. Curr. Allergy Asthma Rep. 2009, 9, 376–384. [Google Scholar] [CrossRef] [PubMed]
- Reboux, G.; Bellanger, A.P.; Roussel, S.; Grenouillet, F.; Millon, L. Moulds in dwellings: Health risks and involved species. Rev. Mal. Respir. 2010, 27, 169–179. [Google Scholar] [CrossRef] [PubMed]
- Heinzerling, L.; Mari, A.; Bergmann, K.C.; Bresciani, M.; Burbach, G.; Darsow, U.; Durham, S.; Fokkens, W.; Gjomarkaj, M.; Haahtela, T.; et al. The skin prick test—European standards. Clin. Transl. Allergy 2013, 3, 1–10. [Google Scholar] [CrossRef]
- Segers, F.J.J.; Meijer, M.; Houbraken, J.; Samson, R.A.; Han, A.B.W.; Dijksterhuis, J. Xerotolerant Cladosporium sphaerospermum are predominant on indoor surfaces compared to other Cladosporium species. PLoS ONE 2015, 10, 145–150. [Google Scholar] [CrossRef]
- Jayasiri, S.C.; Hyde, K.D.; Jones, E.B.G.; McKenzie, E.H.C.; Jeewon, R.; Phillips, A.J.L.; Bhat, D.J.; Wanasinghe, D.N.; Liu, J.K.; Lu, Y.Z.; et al. Diversity, morphology and molecular phylogeny of Dothideomycetes on decaying wild seed pods and fruits. Mycosphere 2019, 10, 1–186. [Google Scholar] [CrossRef]
- Chen, M.M. Forest Fungi Phytogeography: Forest Fungi Phytogeography of China, North America, and Siberia and International Quarantine of Tree Pathogens; Pacific Mushroom Research and Education Center: Sacramento, CA, USA, 2002. [Google Scholar]
- Farr, D.F.; Rossman, A.Y. Fungal Databases, US National Fungus Collections, ARS, USDA. Available online: https://nt.ars–grin.gov/fungaldatabases (accessed on 1 August 2019).
- Steyaert, R.L. Contribution a l’etude monographique de Pestalotia de Not. et Monochaetia Sacc. (Truncatella gen. nov. et Pestalotiopsis gen. nov.). Available online: https://www.jstor.org/stable/3666710?seq=1 (accessed on 1 August 2019).
- Jeewon, R.; Liew, E.C.Y.; Hyde, K.D. Phylogenetic relationships of Pestalotiopsis and allied genera inferred from ribosomal DNA sequences and morphological characters. Mol. Phylogenet. Evol. 2002, 25, 378–392. [Google Scholar] [CrossRef]
- Jeewon, R.; Liew, E.C.Y.; Simpson, J.A.; Hodgkiss, I.J.; Hyde, K.D. Phylogenetic significance of morphological characters in the taxonomy of Pestalotiopsis species. Mol. Phylogenet. Evol. 2013, 27, 372–383. [Google Scholar] [CrossRef]
- Hu, H.; Jeewon, R.; Zhou, D.; Zhou, T.; Hyde, K.D. Phylogenetic diversity of endophytic Pestalotiopsis species in Pinus armandii and Ribes spp.: Evidence from rDNA and β–tubulin gene phylogenies. Fungal Divers. 2007, 45–89. [Google Scholar]
- Maharachchikumbura, S.S.N.; Guo, L.D.; Chukeatirote, E.; Bahkali, A.H.; Hyde, K.D. Pestalotiopsis-morphology, phylogeny, biochemistry and diversity. Fungal Divers. 2011, 50, 167–187. [Google Scholar] [CrossRef]
- Maharachchikumbura, S.S.N.; Guo, L.D.; Chukeatirote, E.; Wu, W.P.; Sun, X.; Crous, P.W.; Bhat, D.J.; McKenzie, E.H.C.; Bahkali, A.H.; Hyde, H.D. A multi-locus backbone tree for Pestalotiopsis, with a polyphasic characterization of 14 new species. Fungal Divers. 2012, 56, 95–129. [Google Scholar] [CrossRef]
- Maharachchikumbura, S.S.N.; Guo, L.D.; Chukeatirote, E.; Hyde, K.D. Improving the backbone tree for the genus Pestalotiopsis; addition of P. steyaertii and P. magna sp. nov. Mycol. Prog. 2013, 13, 617–624. [Google Scholar] [CrossRef]
- Maharachchikumbura, S.S.N.; Hyde, K.D.; Groenewald, J.Z.; Xu, J.; Crous, P.W. Pestalotiopsis revisited. Stud. Mycol. 2014, 79, 121–186. [Google Scholar] [CrossRef] [PubMed]
- Maharachchikumbura, S.S.N.; Guo, L.D.; Liu, Z.Y.; Hyde, K.D. Pseudopestalotiopsis ignota and Ps. camelliae spp. nov. associated with grey blight disease of tea in China. Mycol. Prog. 2016, 15, 22–58. [Google Scholar] [CrossRef]
- Maharachchikumbura, S.S.N.; Hyde, K.D.; Jones, E.B.G.; Mckenzie, E.H.C.; Bhat, J.D.; Dayarathne, M.C.; Huang, C.-K.; Norphanphoun, C.; Senanayake, I.C.; Perera, R.H.; et al. Families of Sordariomycetes. Fungal Divers. 2016, 79, 1–317. [Google Scholar] [CrossRef]
- Sutton, D.A. Coelomycetous fungi in human disease. A review: Clinical entities, pathogenesis, identification and therapy. Rev. Iberoam. Micol. 1999, 16, 171–179. [Google Scholar]
- Monden, Y.; Yamamoto, S.; Sunada, A.; Asari, S.; Makimura, K.; Inoue, Y. First case of fungal keratitis caused by Pestalotiopsis clavispora. Clin. Ophthalmol. 2013, 7, 2261–2264. [Google Scholar] [CrossRef]
- Zhuang, W.Y. Fungi of Northwestern China. Mycotaxon, Ltd.: Ithaca, NY, USA, 2005. [Google Scholar]
- Figueras, M.J.; Guarro, J. A scanning electron microscopic study of ascoma development in Chaetomium Malays. Mycologia 1988, 80, 298–306. [Google Scholar] [CrossRef]
- Index Fungorum. Available online: http://www.indexfungorumorg/Names/Namesasp (accessed on 15 July 2019).
- Jayasiri, S.C.; Hyde, K.D.; Ariyawansa, H.A.; Bhat, J.; Buyck, B.; Cai, L.; Dai, Y.C.; Abd-Elsalam, K.A.; Ertz, D.; Hidayat, I.; et al. The Faces of Fungi database: Fungal names linked with morphology, phylogeny and human impacts. Fungal Divers. 2015, 74, 3–18. [Google Scholar] [CrossRef]
- Carbone, I.; Kohn, L.M. A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 1999, 91, 553–556. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: New York, NY, USA, 1990; pp. 315–322. [Google Scholar]
- Rehner, S.A. Primers for Elongation Factor 1–alpha (EF1–alpha). 2001. Available online: http://www.aftol.org/pdfs/EF1primer.pdf (accessed on 15 July 2019).
- O’Donnell, K.; Cigelnik, E. Two divergent intragenomic rDNA ITS2 types within a monophyletic lineage of the fungusfusariumare nonorthologous. Mol. Phylogenet. Evol. 1997, 7, 103–116. [Google Scholar] [CrossRef] [PubMed]
- Ariyawansa, H.A.; Hyde, K.D. Additions to Pestalotiopsis in Taiwan. Mycosphere 2018, 9, 999–1013. [Google Scholar] [CrossRef]
- Bensch, K.; Groenewald, J.Z.; Meijer, M.; Dijksterhuis, J.; Jurjevic, Z.; Houbraken, J.; Crous, P.W.; Samson, R.A. Cladosporium species in indoor environments. Stud. Mycol. 2018, 89, 177–301. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. A simple method to control over-alignment in the MAFFT multiple sequence alignment program. Bioinformatics 2016, 32, 1933–1942. Available online: http://mafft.cbrc.jp/alignment/server/index.html (accessed on 15 September 2019). [CrossRef]
- Hall, T. Bioedit Version 6.0.7; Department of Microbiology, North Carolina State University: Raleigh, NC, USA, 2004. [Google Scholar]
- Glez-Peña, D.; Gómez-Blanco, D.; Reboiro-Jato, M.; Fdez-Riverola, F.; Posada, D. ALTER: Program-Oriented Format Conversion of DNA and Protein Alignments. Nucleic Acids Research. Web Server Issue. 2010. Available online: http://www.sing-group.org/ALTER/ (accessed on 15 September 2019).
- Miller, M.A.; Pfeiffer, W.; Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In Proceedings of the Gateway Computing Environments Workshop (GCE), New Orleans, LA, USA, 14 November 2010; Institute of Electrical and Electronics Engineers: New Orleans, LA, USA, 2010. Available online: http://www.phylo.org/ (accessed on 15 September 2019).
- Stamatakis, A. RAxML-VI-HPC: Maximum likelihood–based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 2006, 22, 2688–2690. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post–analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef]
- Nylander, J.A.A. MrModeltest 20 Program Distributed by the Author. Uppsala University, Evolutionary Biology Centre: Uppsala, Sweden, 2004. [Google Scholar]
- Rannala, B.; Yang, Z.H. Probability distribution of molecular evolutionary trees: A new method of phylogenetic inference. J. Mol. Evol. 1996, 43, 304–311. [Google Scholar] [CrossRef]
- Liu, J.K.; Doilom, M.; Wikee, S.; Li, Y.M.; Ariyawansha, H.; Boonmee, S.; Chomnunti, P.; Dai, D.Q.; Bhat, J.D.; Romero, A.I.; et al. Towards a natural classification of Botryosphaeriales. Fungal Divers. 2012, 57, 149–210. [Google Scholar] [CrossRef]
- Cai, L.; Jeewon, R.; Hyde, K.D. Phylogenetic investigations of Sordariaceae based on multiple gene sequences and morphology. Mycol. Res. 2006, 110, 137–150. [Google Scholar] [CrossRef] [PubMed]
- Zhaxybayeva, O.; Gogarten, J.P. Bootstrap, Bayesian probability and maximum likelihood mapping: Exploring new tools for comparative genome analyses. Genomics 2002, 3, 4. [Google Scholar] [PubMed]
- Ariyawansa, H.A.; Thambugala, K.M.; Manamgoda, D.S.; Jayawardena, R.; Camporesi, E.; Boonmee, S.; Wanasinghe, D.N.; Phookamsak, R.; Hongsanan, S.; Singtripop, C.; et al. Towards a natural classification and backbone tree for Pleosporaceae. Fungal Divers. 2015, 71, 85–139. [Google Scholar] [CrossRef]
- Rambaut, A.; Drummond, A. FigTree: Tree figure drawing tool, version 1.2.2. Institute of Evolutionary Biology; University of Edinburgh: Edinburgh, Scotland, 2008. [Google Scholar]
- Sandoval-Denis, M.; Sutton, D.A.; Martin-Vicente, A.; Cano-Lira, J.F.; Wiederhold, N.; Guarro, J.; Gené, J. Cladosporium species recovered from clinical samples in the United States. J. Clin. Microbiol. 2015, 53, 2990–3000. [Google Scholar] [CrossRef] [PubMed]
- Jeewon, R.; Hyde, K.D. Establishing species boundaries and new taxa among fungi: Recommendations to resolve taxonomic ambiguities. Mycosphere 2016, 7, 1669–1677. [Google Scholar] [CrossRef]
- Lee, J.C.; Yang, X.S.; Schwartz, M.; Strobel, G.; Clardy, J. The relationship between an endangered North American tree and an endophytic fungus. Chem. Biol. 1995, 2, 721. [Google Scholar] [CrossRef][Green Version]
- Xu, J.; Ebada, S.S.; Proksch, P. Pestalotiopsis a highly creative genus: Chemistry and bioactivity of secondary metabolites. Fungal Divers. 2010, 44, 15–31. [Google Scholar] [CrossRef]
- Yang, X.L.; Zhang, J.Z.; Luo, D.Q. The taxonomy, biology and chemistry of the fungal Pestalotiopsis genus. Nat. Prod. Rep. 2012, 29, 622–641. [Google Scholar] [CrossRef]



| Genes | Primers (Forward/Reverse) | References |
|---|---|---|
| Cladosporium | ||
| ACT | 512F/783R | [51] |
| ITS | ITS5/ITS4 | [52] |
| TEF1 | 728F/986R | [51] |
| Pestalotiopsis | ||
| ITS | ITS5/ITS4 | [52] |
| TEF1 | 526F/1567R | [53] |
| TUB2 | T1/T2 | [54] |
| Species | Culture Collection Number | GenBank Accession Numbers | ||
|---|---|---|---|---|
| ITS | TEF1 | ACT | ||
| Cercospora beticola | CBS 116456 | NR_121315 | AY840494 | AY840458 |
| Cladosporium acalyphae | CBS 125982 | HM147994 | HM148235 | HM148481 |
| C. alboflavescens | CBS 140690 | LN834420 | LN834516 | LN834604 |
| C. angustisporum | CBS 125983 | HM147995 | HM148236 | HM148482 |
| C. angustisporum | UTHSC-DI-13-240 | LN834356 | LN834452 | LN834540 |
| C. anthropophilum | CBS 117483 | HM148007 | HM148248 | HM148494 |
| C. anthropophilum | CBS 140685 | LN834437 | LN834533 | LN834621 |
| C. anthropophilum | KUMCC 19-0182 | MN412638 | MN417513 | MN417511 |
| C. anthropophilum | KUMCC 19-0202 | MN412639 | MN417514 | MN417512 |
| C. anthropophilum | UTHSC-DI-13-168 | LN834407 | LN834503 | LN834591 |
| C. anthropophilum | UTHSC-DI-13-169 | LN834408 | LN834504 | LN834592 |
| C. anthropophilum | UTHSC-DI-13-178 | LN834410 | LN834506 | LN834594 |
| C. anthropophilum | UTHSC-DI-13-179 | LN834411 | LN834507 | LN834595 |
| C. anthropophilum | UTHSC-DI-13-207 | LN834413 | LN834509 | LN834597 |
| C. anthropophilum | UTHSC-DI-13-226 | LN834421 | LN834517 | LN834605 |
| C. anthropophilum | UTHSC-DI-13-228 | LN834423 | LN834519 | LN834607 |
| C. anthropophilum | UTHSC-DI-13-244 | LN834428 | LN834524 | LN834612 |
| C. anthropophilum | UTHSC-DI-13-246 | LN834430 | LN834526 | LN834614 |
| C. anthropophilum | UTHSC-DI-13-269 | LN834437 | LN834533 | LN834621 |
| C. anthropophilum | UTHSC-DI-13-271 | LN834439 | LN834535 | LN834623 |
| C. asperulatum | CBS 126339 | HM147997 | HM148238 | HM148484 |
| C. asperulatum | CBS 126340 | HM147998 | HM148239 | HM148485 |
| C. australiense | CBS 125984 | HM147999 | HM148240 | HM148486 |
| C. austroafricanum | CPC 16763 | KT600381 | KT600478 | KT600577 |
| C. chalastosporoides | CBS 125985 | HM148001 | HM148242 | HM148488 |
| C. chubutense | CBS 124457 | FJ936158 | FJ936161 | FJ936165 |
| C. cladosporioides | CBS 112388 | HM148003 | HM148244 | HM148490 |
| C. cladosporioides | CBS 113738 | HM148004 | HM148245 | HM148491 |
| C. cladosporioides | CPC 14292 | HM148046 | HM148287 | HM148533 |
| C. cladosporioides | UTHSC-DI-13-215 | LN834360 | LN834456 | LN834544 |
| C. colocasiae | CBS 119542 | HM148066 | HM148309 | HM148554 |
| C. colocasiae | CBS 386.64 | HM148067 | HM148310 | HM148555 |
| C. colombiae | CBS 274.80B | FJ936159 | FJ936163 | FJ936166 |
| C. crousii | UTHSC-DI-13-247 | LN834431 | LN834527 | LN834615 |
| C. cucumerinum | CBS 171.52 | HM148072 | HM148316 | HM148561 |
| C. cucumerinum | CBS 173.54 | HM148074 | HM148318 | HM148563 |
| C. delicatulum | CBS 126342 | HM148079 | HM148323 | HM148568 |
| C. delicatulum | CBS 126344 | HM148081 | HM148325 | HM148570 |
| C. exile | CBS 125987 | HM148091 | HM148335 | HM148580 |
| C. flabelliforme | CBS 126345 | HM148092 | HM148336 | HM148581 |
| C. flabelliforme | UTHSC-DI-13-267 | LN834361 | LN834457 | LN834545 |
| C. flavovirens | UTHSC-DI-13-273 | LN834440 | LN834536 | LN834624 |
| C. funiculosum | CBS 122128 | HM148093 | HM148337 | HM148582 |
| C. funiculosum | CBS 122129 | HM148094 | HM148338 | HM148583 |
| C. gamsianum | CBS 125989 | HM148095 | HM148339 | HM148584 |
| C. globisporum | CBS 812.96 | HM148096 | HM148340 | HM148585 |
| C. grevilleae | CBS 114271 | JF770450 | JF770472 | JF770473 |
| C. hillianum | CBS 125988 | HM148097 | HM148341 | HM148586 |
| C. inversicolor | CBS 143.65 | HM148100 | HM148344 | HM148589 |
| C. ipereniae | CBS 140483 | KT600394 | KT600491 | KT600589 |
| C. ipereniae | CPC 16855 | KT600395 | KT600492 | KT600590 |
| C. iranicum | CBS 126346 | HM148110 | HM148354 | HM148599 |
| C. limoniforme | CBS 113737 | KT600396 | KT600493 | KT600591 |
| C. longicatenatum | CPC 17189 | KT600403 | KT600500 | KT600598 |
| C. lycoperdinum | CBS 126347 | HM148112 | HM148356 | HM148601 |
| C. lycoperdinum | CBS 574.78C | HM148115 | HM148359 | HM148604 |
| C. montecillanum | CPC 15605 | KT600407 | KT600505 | KT600603 |
| C. montecillanum | CPC 17953 | KT600406 | KT600504 | KT600602 |
| C. myrtacearum | CBS 126350 | HM148117 | HM148361 | HM148606 |
| C. oxysporum | CBS 125991 | HM148118 | HM148362 | HM148607 |
| C. oxysporum | CBS 126351 | HM148119 | HM148363 | HM148608 |
| C. paracladosporioides | CBS 171.54 | HM148120 | HM148364 | HM148609 |
| C. parapenidielloides | CPC 17193 | KT600410 | KT600508 | KT600606 |
| C. phaenocomae | CBS 128769 | JF499837 | JF499875 | JF499881 |
| C. phyllactiniicola | CBS 126353 | HM148151 | HM148395 | HM148640 |
| C. phyllactiniicola | CBS 126355 | HM148153 | HM148397 | HM148642 |
| C. phyllophilum | CBS 125992 | HM148154 | HM148398 | HM148643 |
| C. phyllophilum | CBS 125990 | HM148111 | HM148355 | HM148600 |
| C. pini-ponderosae | CBS 124456 | FJ936160 | FJ936164 | FJ936167 |
| C. pseudochalastosporoides | CPC 17823 | KT600415 | KT600513 | KT600611 |
| C. pseudocladosporioides | CBS 125993 | HM148158 | HM148402 | HM148647 |
| C. pseudocladosporioides | CBS 667.80 | HM148165 | HM148409 | HM148654 |
| C. pseudocladosporioides | CPC 13683 | HM148173 | HM148417 | HM148662 |
| C. pseudocladosporioides | CPC 14020 | HM148185 | HM148429 | HM148674 |
| C. pseudocladosporioides | CPC 14295 | HM148188 | HM148432 | HM148677 |
| C. pseudocladosporioides | UTHSC-DI-13-165 | LN834406 | LN834502 | LN834590 |
| C. pseudocladosporioides | UTHSC-DI-13-190 | LN834412 | LN834508 | LN834596 |
| C. pseudocladosporioides | UTHSC-DI-13-210 | LN834414 | LN834510 | LN834598 |
| C. pseudocladosporioides | UTHSC-DI-13-218 | LN834418 | LN834514 | LN834602 |
| C. pseudocladosporioides | UTHSC-DI-13-227 | LN834422 | LN834518 | LN834606 |
| C. pseudocladosporioides | UTHSC-DI-13-234 | LN834424 | LN834520 | LN834608 |
| C. pseudocladosporioides | UTHSC-DI-13-238 | LN834426 | LN834522 | LN834610 |
| C. pseudocladosporioides | UTHSC-DI-13-241 | LN834427 | LN834523 | LN834611 |
| C. pseudocladosporioides | UTHSC-DI-13-245 | LN834429 | LN834525 | LN834613 |
| C. pseudocladosporioides | UTHSC-DI-13-251 | LN834432 | LN834528 | LN834616 |
| C. pseudocladosporioides | UTHSC-DI-13-261 | LN834384 | LN834480 | LN834568 |
| C. pseudocladosporioides | UTHSC-DI-13-265 | LN834435 | LN834531 | LN834619 |
| C. pseudocladosporioides | UTHSC-DI-13-268 | LN834436 | LN834532 | LN834620 |
| C. pseudocladosporioides | UTHSC-DI-13-270 | LN834438 | LN834534 | LN834622 |
| C. rectoides | CBS 125994 | HM148193 | HM148438 | HM148683 |
| C. ruguloflabelliforme | CPC 19707 | KT600458 | KT600557 | KT600655 |
| C. scabrellum | CBS 126358 | HM148195 | HM148440 | HM148685 |
| C. silenes | CBS 109082 | EF679354 | EF679429 | EF679506 |
| C. subinflatum | CBS 121630 | EF679389 | EF679467 | EF679543 |
| C. subinflatum | CBS 121630 | EF679389 | EF679467 | EF679543 |
| C. subuliforme | CBS 126500 | HM148196 | HM148441 | HM148686 |
| C. tenuissimum | CPC 13222 | HM148210 | HM148455 | HM148700 |
| C. tenuissimum | CPC 14250 | HM148211 | HM148456 | HM148701 |
| C. tenuissimum | UTHSC-DI-13-258 | LN834404 | LN834500 | LN834588 |
| C. varians | CBS 126362 | HM148224 | HM148470 | HM148715 |
| C. verrucocladosporioides | CBS 126363 | HM148226 | HM148472 | HM148717 |
| C. versiforme | CPC 19053 | KT600417 | KT600515 | KT600613 |
| C. xantochromaticum | CBS 140691 | LN834415 | LN834511 | LN834599 |
| C. xylophilum | CBS 125997 | HM148230 | HM148476 | HM148721 |
| Species | Culture Collection Number | GenBank Accession Numbers | ||
|---|---|---|---|---|
| ITS | TUB2 | TEF1 | ||
| Neopestalotiopsis clavispora | CBS 447.73 | KM199374 | KM199443 | KM199539 |
| N. formicarum | CBS 362.72 | KM199358 | KM199455 | KM199517 |
| Pestalotiopsis adusta | MFLUCC 10-0146 | JX399006 | JX399037 | JX399070 |
| P. aggestorum | LC8186 | KY464140 | KY464160 | KY464150 |
| P. anacardiacearum | IFRDCC 2397 | KC247154 | KC247155 | KC247156 |
| P. arceuthobii | CBS 434.65 | KM199341 | KM199427 | KM199516 |
| P. arengae | CBS 331.92 | KM199340 | KM199426 | KM199515 |
| P. australasiae | CBS 114126 | KM199297 | KM199409 | KM199499 |
| P. australis | CBS 114193 | KM199332 | KM199383 | KM199475 |
| P. biciliata | CBS 124463 | KM199308 | KM199399 | KM199505 |
| P. biciliata | CBS 790.68 | KM199305 | KM199400 | KM199507 |
| P. biciliata | MFLUCC 12-0598 | KX816920 | KX816948 | KX816890 |
| P. brachiata | LC2988 | KX894933 | KX895265 | KX895150 |
| P. brassicae | CBS 170.26 | KM199379 | - | KM199558 |
| P. camelliae | MFLUCC 12-0277 | JX399010 | JX399041 | JX399074 |
| P. chamaeropis | CBS 186.71 | KM199326 | KM199391 | KM199473 |
| P. clavata | MFLUCC 12-0268 | JX398990 | JX399025 | JX399056 |
| P. colombiensis | CBS 118553 | KM199307 | KM199421 | KM199488 |
| P. digitalis | ICMP 5434 | KP781879 | KP781883 | - |
| P. dilucida | LC3232 | KX894961 | KX895293 | KX895178 |
| P. diploclisiae | CBS 115587 | KM199320 | KM199419 | KM199486 |
| P. distincta | LC8185 | KY464139 | KY464159 | KY464149 |
| P. diversiseta | MFLUCC 12-0287 | JX399009 | JX399040 | JX399073 |
| P. dracontomelon | MFUCC 10-0149 | KP781877 | - | KP781880 |
| P. ericacearum | IFRDCC 2439 | KC537807 | KC537821 | KC537814 |
| P. ericacearum | OP023 | KC537807 | KC537821 | KC537814 |
| P. formosana | NTUCC 17-010 | MH809382 | MH809386 | MH809390 |
| P. furcata | MFLUCC 12-0054 | JQ683724 | JQ683708 | JQ683740 |
| P. gaultheria | IFRD 411-014 | KC537805 | KC537819 | KC537812 |
| P. gaultheria | OP137 | KC537805 | KC537819 | KC537812 |
| P. grevilleae | CBS 114127 | KM199300 | KM199407 | KM199504 |
| P. hawaiiensis | CBS 114491 | KM199339 | KM199428 | KM199514 |
| P. hispanica | CBS 115391 | MH553981 | MH554640 | MH554399 |
| P. hollandica | CBS 265.33 | KM199328 | KM199388 | KM199481 |
| P. humus | CBS 336.97 | KM199317 | KM199420 | KM199484 |
| P. inflexa | MFLUCC 12-0270 | JX399008 | JX399039 | JX399072 |
| P. intermedia | MFLUCC 12-0259 | JX398993 | JX399028 | JX399059 |
| P. italiana | MFLUCC 12-0657 | KP781878 | KP781882 | KP781881 |
| P. jesteri | CBS109350 | KM199380 | KM199468 | KM199554 |
| P. jiangxiensis | LC4399 | KX895009 | KX895341 | KX895227 |
| P. jinchanghensis | LC8191 | KY464145 | KY464165 | KY464155 |
| P. kenyana | CBS 442.67 | KM199302 | KM199395 | KM199502 |
| P. knightiae | CBS 114138 | KM199310 | KM199408 | KM199497 |
| P. krabiensis | MFLUCC 16-0260 | MH388360 | MH412722 | MH388395 |
| P. leucadendri | CBS 121417 | MH553987 | MH554654 | MH554412 |
| P. licualacola | HGUP 4057 | KC492509 | KC481683 | KC481684 |
| P. linearis | MFLUCC 12-0271 | JX398992 | JX399027 | JX399058 |
| P. longiappendiculata | LC3013 | KX894939 | KX895271 | KX895156 |
| P. lushanensis | LC8183 | KY464137 | KY464157 | KY464147 |
| P. lushanensis | OP086 | KC537804 | KC537818 | KC537811 |
| P. macadamiae | BRIP 63738 | KX186588 | KX186680 | KX186621 |
| P. malayana | CBS 102220 | KM199306 | KM199411 | KM199482 |
| P. monochaeta | CBS 144.97 | KM199327 | KM199386 | KM199479 |
| P. montelica | MFLUCC 12-0279 | JX399012 | JX399043 | JX399076 |
| P. neolitseae | NTUCC 17-012 | MH809384 | MH809388 | MH809392 |
| P. novae-hollandiae | CBS 130973 | KM199337 | KM199425 | KM199511 |
| P. oryzae | CBS 353.69 | KM199299 | KM199398 | KM199496 |
| P. pandanicola | MFLUCC 16-0255 | MH388361 | MH412723 | MH388396 |
| P. papuana | CBS 331.96 | KM199321 | KM199413 | KM199491 |
| P. parva | CBS 265.37 | KM199312 | KM199404 | KM199508 |
| P. pinicola | KUMCC 19-0183 | MN412636 | MN417507 | MN417509 |
| P. pinicola | KUMCC 19-0203 | MN412637 | MN417508 | MN417510 |
| P. portugalica | CBS 393.48 | KM199335 | KM199422 | KM199510 |
| P. rhododendri | IFRDCC 2399 | KC537804 | KC537818 | KC537811 |
| P. rhodomyrtus | HGUP 4230 | KF412648 | KC537818 | KF412645 |
| P. rosea | CL0441 | KY228790 | - | - |
| P. rosea | MFLUCC 12-0258 | JX399005 | JX399036 | JX399069 |
| P. scoparia | CBS 176.25 | KM199330 | KM199393 | KM199478 |
| P. sequoia | MFLUCC 13-0399 | KX572339 | - | - |
| P. shorea | MFLUCC 12-0314 | KJ503811 | KJ503814 | KJ503817 |
| P. spathulata | CBS 356.86 | KM199338 | KM199423 | KM199513 |
| P. spathuliappendiculata | CBS 144035 | MH554172 | MH554845 | MH554607 |
| P. telopeae | CBS 114161 | KM199296 | KM199403 | KM199500 |
| P. terricola | CBS 141.69 | MH554004 | MH554680 | MH554438 |
| P. trachicarpicola | IFRDCC 2240 | NR_120109 | - | - |
| P. trachicarpicola | OP143 | JQ845947 | JQ845945 | JQ845946 |
| P. unicolor | MFLUCC 12-0276 | JX398999 | JX399030 | JX399063 |
| P. verruculosa | MFLUCC 12-0274 | JX398996 | - | JX399061 |
| P. yanglingensis | LC3375 | KX894975 | KX895307 | KX895192 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tibpromma, S.; Mortimer, P.E.; Karunarathna, S.C.; Zhan, F.; Xu, J.; Promputtha, I.; Yan, K. Morphology and Multi-Gene Phylogeny Reveal Pestalotiopsis pinicola sp. nov. and a New Host Record of Cladosporium anthropophilum from Edible Pine (Pinus armandii) Seeds in Yunnan Province, China. Pathogens 2019, 8, 285. https://doi.org/10.3390/pathogens8040285
Tibpromma S, Mortimer PE, Karunarathna SC, Zhan F, Xu J, Promputtha I, Yan K. Morphology and Multi-Gene Phylogeny Reveal Pestalotiopsis pinicola sp. nov. and a New Host Record of Cladosporium anthropophilum from Edible Pine (Pinus armandii) Seeds in Yunnan Province, China. Pathogens. 2019; 8(4):285. https://doi.org/10.3390/pathogens8040285
Chicago/Turabian StyleTibpromma, Saowaluck, Peter E. Mortimer, Samantha C. Karunarathna, Fangdong Zhan, Jianchu Xu, Itthayakorn Promputtha, and Kai Yan. 2019. "Morphology and Multi-Gene Phylogeny Reveal Pestalotiopsis pinicola sp. nov. and a New Host Record of Cladosporium anthropophilum from Edible Pine (Pinus armandii) Seeds in Yunnan Province, China" Pathogens 8, no. 4: 285. https://doi.org/10.3390/pathogens8040285
APA StyleTibpromma, S., Mortimer, P. E., Karunarathna, S. C., Zhan, F., Xu, J., Promputtha, I., & Yan, K. (2019). Morphology and Multi-Gene Phylogeny Reveal Pestalotiopsis pinicola sp. nov. and a New Host Record of Cladosporium anthropophilum from Edible Pine (Pinus armandii) Seeds in Yunnan Province, China. Pathogens, 8(4), 285. https://doi.org/10.3390/pathogens8040285







