In Vivo Methods to Study Protein–Protein Interactions as Key Players in Mycobacterium Tuberculosis Virulence
Abstract
1. Introduction
2. Yeast Two-Hybrid (Y2H) System
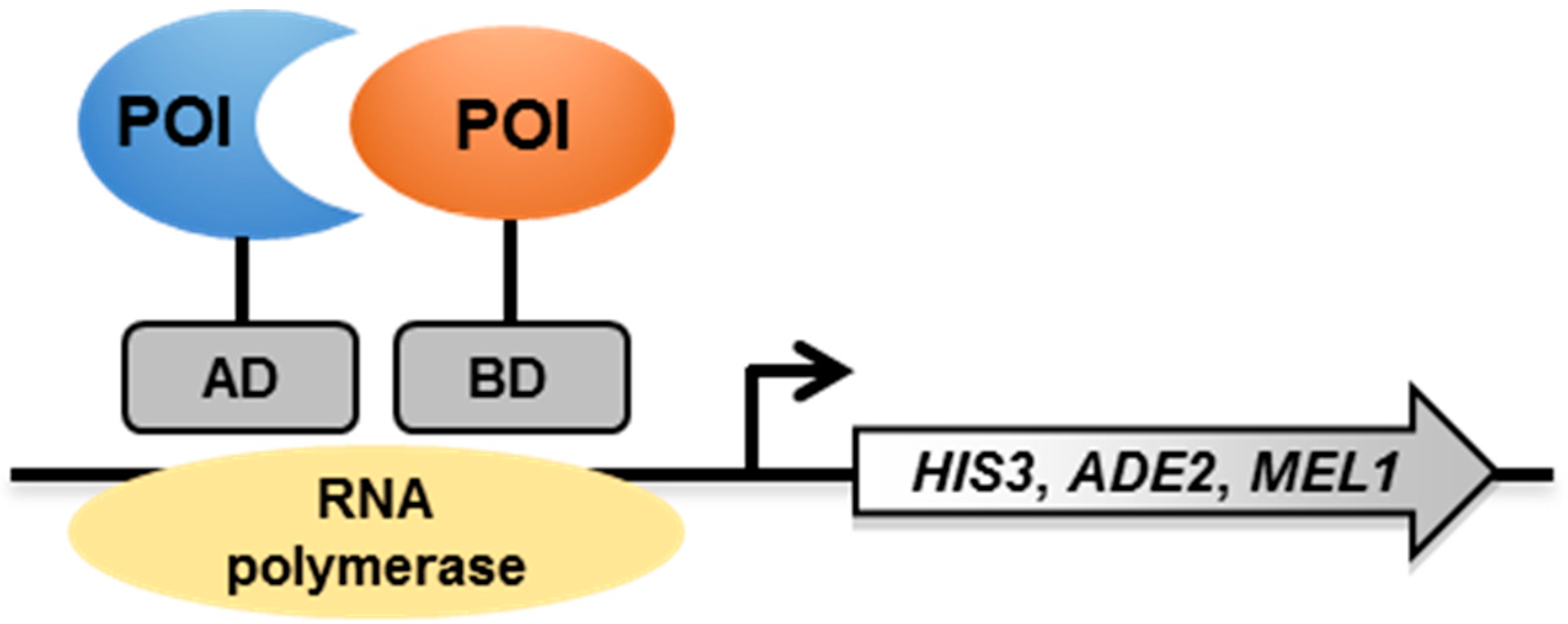
2.1. Principle
2.2. The Y2H System to Study Mycobacterial PPI
2.2.1. Signaling Pathways
2.2.2. Mtb Cell Division
2.2.3. Mtb Cell Wall Composition
2.2.4. Secretion of Mtb Virulence Factors
2.2.5. Regulation of Mycobacterial Protease Activity
2.3. Pros and Cons
3. Bacterial Adenylate Cyclase-Based Two-Hybrid (BACTH) System
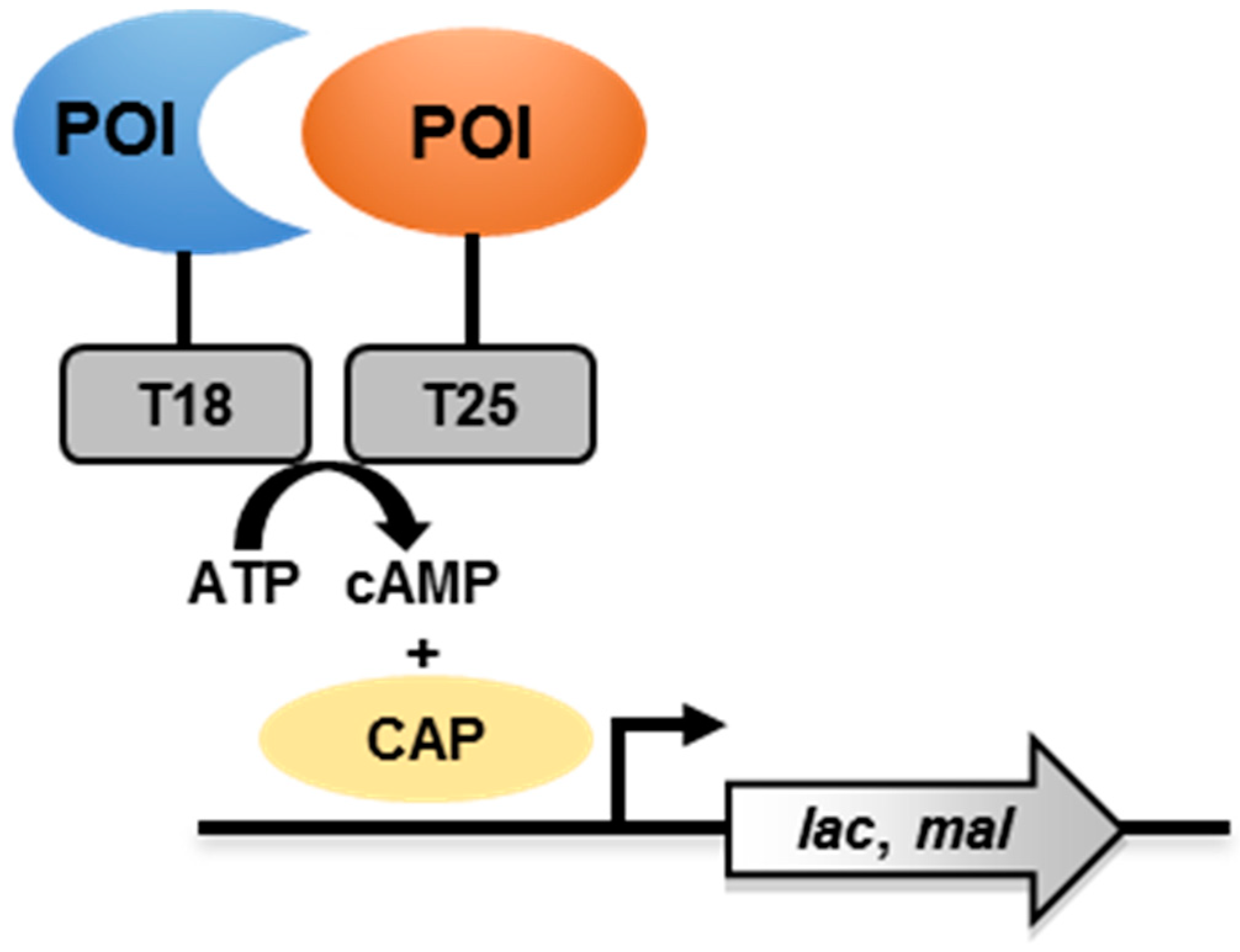
3.1. Principle
3.2. The BACTH System to Study Mycobacterial PPI
3.2.1. Signaling Pathways
3.2.2. Cell Division
3.2.3. Mtb Cell Wall Composition
3.2.4. Mtb Virulence Factors
3.2.5. High-Throughput Screening Applied to BACTH
3.3. Pros and Cons
4. Methods Developed for Use with Live Mycobacteria
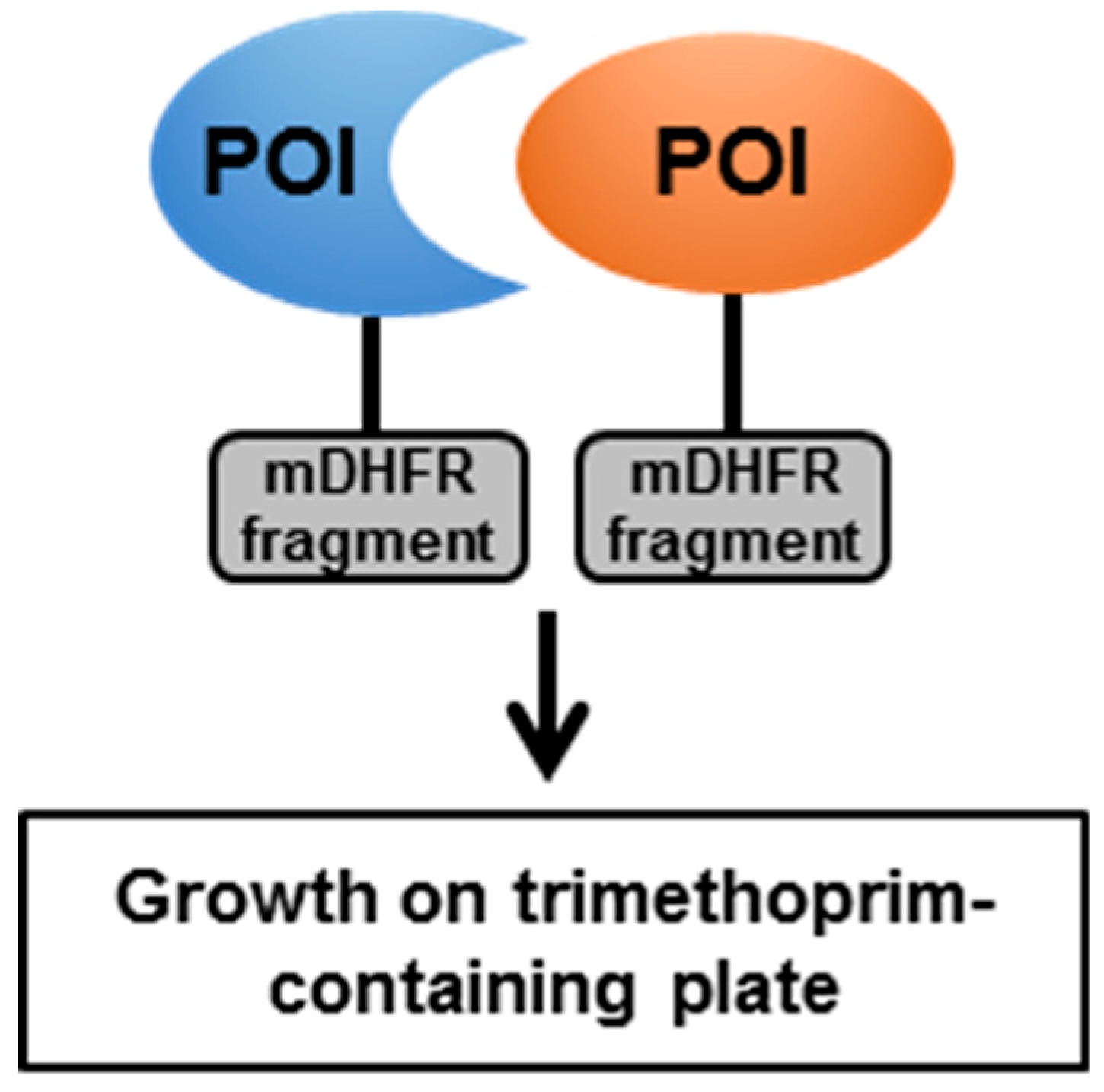
4.1. The Mycobacterial Protein Fragment Complementation (M-PFC)
4.1.1. Signaling Pathways
4.1.2. Cell Division
4.1.3. Peptidoglycan Biosynthesis
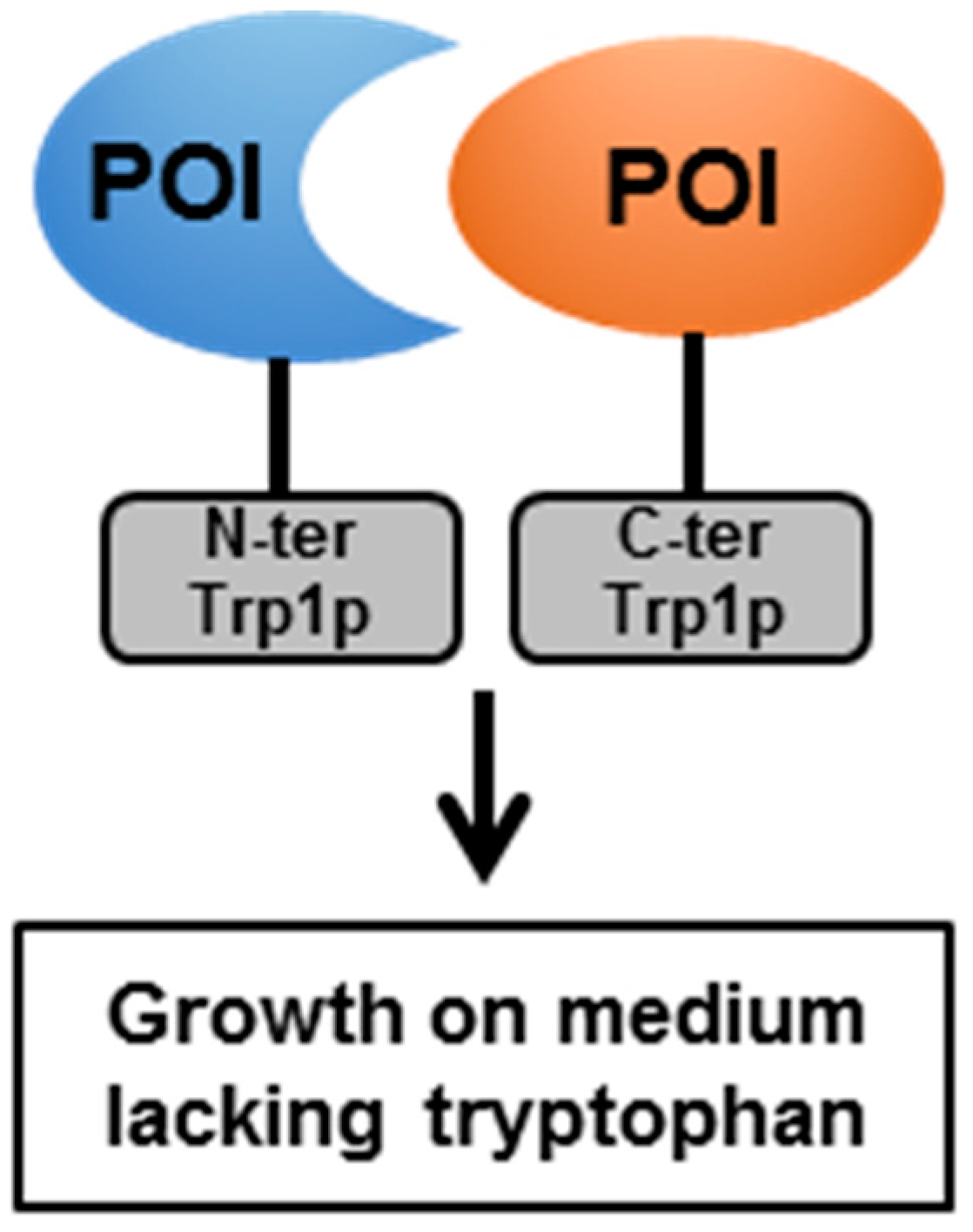
4.2. The Split-Protein Sensor (Split-Trp)
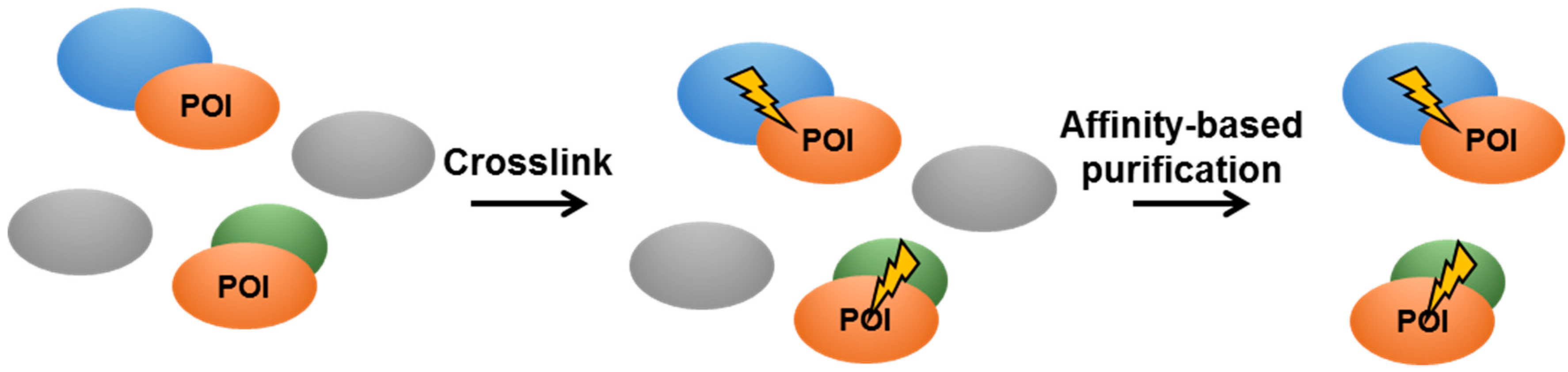
4.3. In Vivo Crosslinking in Live Mycobacteria
4.4. Pros and Cons
5. Conclusion and Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- Bellinzoni, M.; Wehenkel, A.M.; Durán, R.; Alzari, P.M. Novel mechanistic insights into physiological signaling pathways mediated by mycobacterial Ser/Thr protein kinases. Genes Immun. 2019, 20, 383–393. [Google Scholar] [CrossRef]
- Melly, G.; Purdy, G.E. MmpL Proteins in Physiology and Pathogenesis of M. tuberculosis. Microorganisms 2019, 7, 70. [Google Scholar] [CrossRef] [PubMed]
- Rameshwaram, N.R.; Singh, P.; Ghosh, S.; Mukhopadhyay, S. Lipid metabolism and intracellular bacterial virulence: Key to next-generation therapeutics. Future Microbiol. 2018, 13, 1301–1328. [Google Scholar] [CrossRef] [PubMed]
- Velasco-García, R.; Vargas-Martínez, R. The study of protein–protein interactions in bacteria. Can. J. Microbiol. 2012, 58, 1241–1257. [Google Scholar] [CrossRef] [PubMed]
- Schauer, K.; Stingl, K. ‘Guilty by Association’—Protein-Protein Interactions (PPIs) in Bacterial Pathogens. Genome Dyn. 2009, 6, 48–61. [Google Scholar] [CrossRef]
- Forrellad, M.A.; Klepp, L.I.; Gioffre, A.; Sabio y Garcia, J.; Morbidoni, H.R.; de la Paz Santangelo, M.; Cataldi, A.A.; Bigi, F. Virulence factors of the Mycobacterium tuberculosis complex. Virulence 2013, 4, 3–66. [Google Scholar] [CrossRef]
- Fu, H.; Miller, J.; Stagljar, I. Using the Yeast Two-Hybrid System to Identify Interacting Proteins. Protein Protein Interact. 2004, 261, 247–262. [Google Scholar] [CrossRef]
- Manganelli, R. Sigma Factors: Key Molecules in Mycobacterium tuberculosis Physiology and Virulence. Microbiol. Spectr. 2014, 2, 137–160. [Google Scholar] [CrossRef]
- Hurst-Hess, K.; Biswas, R.; Yang, Y.; Rudra, P.; Lasek-Nesselquist, E.; Ghosh, P. Mycobacterial SigA and SigB Cotranscribe Essential Housekeeping Genes during Exponential Growth. mBio 2019, 10, e00273-19. [Google Scholar] [CrossRef]
- Steyn, A.J.C.; Collins, D.M.; Hondalus, M.K.; Jacobs, W.R.; Kawakami, R.P.; Bloom, B.R. Mycobacterium tuberculosis WhiB3 interacts with RpoV to affect host survival but is dispensable for in vivo growth. Proc. Natl. Acad. Sci. USA 2002, 99, 3147–3152. [Google Scholar] [CrossRef]
- Garg, S.; Alam, M.S.; Bajpai, R.; Kishan, K.R.; Agrawal, P. Redox biology of Mycobacterium tuberculosis H37Rv: Protein-protein interaction between GlgB and WhiB1 involves exchange of thiol-disulfide. BMC Biochem. 2009, 10, 1. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Karakousis, P.C.; Bishai, W.R. Roles of SigB and SigF in the Mycobacterium tuberculosis sigma factor network. J. Bacteriol. 2008, 190, 699–707. [Google Scholar] [CrossRef] [PubMed]
- Gebhard, S.; Hümpel, A.; McLellan, A.D.; Cook, G.M.; McLellan, A. The alternative sigma factor SigF of Mycobacterium smegmatis is required for survival of heat shock, acidic pH and oxidative stress. Microbiology 2008, 154, 2786–2795. [Google Scholar] [CrossRef] [PubMed]
- Parida, B.; Douglas, T.; Nino, C.; Dhandayuthapani, S. Interactions of anti-sigma factor antagonists of Mycobacterium tuberculosis in the yeast two-hybrid system. Tuberculosis 2005, 85, 347–355. [Google Scholar] [CrossRef] [PubMed]
- Kundu, M. The role of two-component systems in the physiology of Mycobacterium tuberculosis. IUBMB Life 2018, 70, 710–717. [Google Scholar] [CrossRef]
- Steyn, A.J.C.; Joseph, J.; Bloom, B.R. Interaction of the sensor module of Mycobacterium tuberculosis H37Rv KdpD with members of the Lpr family. Mol. Microbiol. 2003, 47, 1075–1089. [Google Scholar] [CrossRef]
- Lee, H.N.; Jung, K.E.; Ko, I.J.; Baik, H.S.; Oh, J.I. Protein-protein interactions between histidine kinases and response regulators of Mycobacterium tuberculosis H37Rv. J. Microbiol. 2012, 50, 270–277. [Google Scholar] [CrossRef]
- Prisic, S.; Husson, R.N. Mycobacterium tuberculosis Serine/Threonine Protein Kinases. Microbiol. Spectr. 2014, 2, 681–708. [Google Scholar] [CrossRef]
- Wu, F.L.; Liu, Y.; Jiang, H.W.; Luan, Y.Z.; Zhang, H.N.; He, X.; Xu, Z.W.; Hou, J.L.; Ji, L.Y.; Xie, Z.; et al. The Ser/Thr Protein Kinase Protein-Protein Interaction Map of M. tuberculosis. Mol. Cell. Proteom. 2017, 16, 1491–1506. [Google Scholar] [CrossRef]
- Kieser, K.J.; Rubin, E.J. How sisters grow apart: Mycobacterial growth and division. Nat. Rev. Genet. 2014, 12, 550–562. [Google Scholar] [CrossRef]
- Gola, S.; Munder, T.; Casonato, S.; Manganelli, R.; Vicente, M. The essential role of SepF in mycobacterial division. Mol. Microbiol. 2015, 97, 560–576. [Google Scholar] [CrossRef] [PubMed]
- Rosser, A.; Stover, C.; Pareek, M.; Mukamolova, G.V. Resuscitation-promoting factors are important determinants of the pathophysiology inMycobacterium tuberculosisinfection. Crit. Rev. Microbiol. 2017, 43, 621–630. [Google Scholar] [CrossRef] [PubMed]
- Hett, E.C.; Chao, M.C.; Steyn, A.J.; Fortune, S.M.; Deng, L.L.; Rubin, E.J. A partner for the resuscitation-promoting factors of Mycobacterium tuberculosis. Mol. Microbiol. 2007, 66, 658–668. [Google Scholar] [CrossRef]
- Hett, E.C.; Chao, M.C.; Rubin, E.J. Interaction and Modulation of Two Antagonistic Cell Wall Enzymes of Mycobacteria. PLoS Pathog. 2010, 6, e1001020. [Google Scholar] [CrossRef] [PubMed]
- Quémard, A. New Insights into the Mycolate-Containing Compound Biosynthesis and Transport in Mycobacteria. Trends Microbiol. 2016, 24, 725–738. [Google Scholar] [CrossRef] [PubMed]
- Veyron-Churlet, R.; Guerrini, O.; Mourey, L.; Daffé, M.; Zerbib, D.; Veyron-Churlet, R. Protein-protein interactions within the Fatty Acid Synthase-II system of Mycobacterium tuberculosis are essential for mycobacterial viability. Mol. Microbiol. 2004, 54, 1161–1172. [Google Scholar] [CrossRef] [PubMed]
- Veyron-Churlet, R.; Bigot, S.; Guerrini, O.; Verdoux, S.; Malaga, W.; Daffé, M.; Zerbib, D. The Biosynthesis of Mycolic Acids in Mycobacterium tuberculosis Relies on Multiple Specialized Elongation Complexes Interconnected by Specific Protein–Protein Interactions. J. Mol. Boil. 2005, 353, 847–858. [Google Scholar] [CrossRef] [PubMed]
- Cantaloube, S.; Veyron-Churlet, R.; Haddache, N.; Daffe, M.; Zerbib, D. The Mycobacterium Tuberculosis FAS-II Dehydratases and Methyltransferases Define the Specificity of the Mycolic Acid Elongation Complexes. PLoS ONE 2011, 6, e29564. [Google Scholar] [CrossRef]
- Glass, L.N.; Swapna, G.; Chavadi, S.S.; Tufariello, J.M.; Mi, K.; Drumm, J.E.; Lam, T.T.; Zhu, G.; Zhan, C.; Vilcheze, C.; et al. Mycobacterium tuberculosis universal stress protein Rv2623 interacts with the putative ATP binding cassette (ABC) transporter Rv1747 to regulate mycobacterial growth. PLoS Pathog. 2017, 13, e1006515. [Google Scholar] [CrossRef]
- Guerin, M.E.; Korduláková, J.; Alzari, P.M.; Brennan, P.J.; Jackson, M. Molecular Basis of Phosphatidyl-myo-inositol Mannoside Biosynthesis and Regulation in Mycobacteria. J. Boil. Chem. 2010, 285, 33577–33583. [Google Scholar] [CrossRef]
- Torrelles, J.B.; Schlesinger, L.S. Diversity in Mycobacterium tuberculosis mannosylated cell wall determinants impacts adaptation to the host. Tuberculosis 2010, 90, 84–93. [Google Scholar] [CrossRef] [PubMed]
- Renshaw, P.S.; Lightbody, K.L.; Veverka, V.; Muskett, F.W.; Kelly, G.; Frenkiel, T.A.; Gordon, S.V.; Hewinson, R.G.; Burke, B.; Norman, J.; et al. Structure and function of the complex formed by the tuberculosis virulence factors CFP-10 and ESAT-6. EMBO J. 2005, 24, 2491–2498. [Google Scholar] [CrossRef] [PubMed]
- Pathak, S.K.; Basu, S.; Basu, K.K.; Banerjee, A.; Pathak, S.; Bhattacharyya, A.; Kaisho, T.; Kundu, M.; Basu, J. Direct extracellular interaction between the early secreted antigen ESAT-6 of Mycobacterium tuberculosis and TLR2 inhibits TLR signaling in macrophages. Nat. Immunol. 2007, 8, 610–618. [Google Scholar] [CrossRef]
- Feltcher, M.E.; Sullivan, J.T.; Braunstein, M. Protein export systems of Mycobacterium tuberculosis: Novel targets for drug development? Future Microbiol. 2010, 5, 1581–1597. [Google Scholar] [CrossRef]
- Van Winden, V.J.C.; Houben, E.N.G.; Braunstein, M. Protein Export into and across the Atypical Diderm Cell Envelope of Mycobacteria. Microbiol. Spectr. 2019, 7. [Google Scholar] [CrossRef] [PubMed]
- Stanley, S.A.; Raghavan, S.; Hwang, W.W.; Cox, J.S. Acute infection and macrophage subversion by Mycobacterium tuberculosis require a specialized secretion system. Proc. Natl. Acad. Sci. USA 2003, 100, 13001–13006. [Google Scholar] [CrossRef]
- Champion, P.A.D.; Stanley, S.A.; Champion, M.M.; Brown, E.J.; Cox, J.S. C-Terminal Signal Sequence Promotes Virulence Factor Secretion in Mycobacterium tuberculosis. Science 2006, 313, 1632–1636. [Google Scholar] [CrossRef]
- Champion, P.A.D.; Champion, M.M.; Manzanillo, P.; Cox, J.S. ESX-1 Secreted Virulence Factors Are Recognized by Multiple Cytosolic AAA ATPases in Pathogenic Mycobacteria. Mol. Microbiol. 2009, 73, 950–962. [Google Scholar] [CrossRef]
- Ribeiro-Guimarães, M.L.; Pessolani, M.C.V. Comparative genomics of mycobacterial proteases. Microb. Pathog. 2007, 43, 173–178. [Google Scholar] [CrossRef]
- Sklar, J.G.; Makinoshima, H.; Schneider, J.S.; Glickman, M.S.M. tuberculosis intramembrane protease Rip1 controls transcription through three anti-sigma factor substrates. Mol. Microbiol. 2010, 77, 605–617. [Google Scholar] [CrossRef]
- Makinoshima, H.; Glickman, M.S. Regulation of Mycobacterium tuberculosis cell envelope composition and virulence by intramembrane proteolysis. Nature 2005, 436, 406–409. [Google Scholar] [CrossRef]
- Schneider, J.S.; Reddy, S.P.; Evans, H.W.; Glickman, M.S. Site-2 protease substrate specificity and coupling in trans by a PDZ-substrate adapter protein. Proc. Natl. Acad. Sci. USA 2013, 110, 19543–19548. [Google Scholar] [CrossRef] [PubMed]
- Karimova, G.; Pidoux, J.; Ullmann, A.; Ladant, D. A bacterial two-hybrid system based on a reconstituted signal transduction pathway. Proc. Natl. Acad. Sci. USA 1998, 95, 5752–5756. [Google Scholar] [CrossRef] [PubMed]
- Ladant, D.; Ullmann, A. Bordatella pertussis adenylate cyclase: A toxin with multiple talents. Trends Microbiol. 1999, 7, 172–176. [Google Scholar] [CrossRef]
- Battesti, A.; Bouveret, E. The bacterial two-hybrid system based on adenylate cyclase reconstitution in Escherichia coli. Methods 2012, 58, 325–334. [Google Scholar] [CrossRef]
- Fontán, P.A.; Aris, V.; Alvarez, M.E.; Ghanny, S.; Cheng, J.; Soteropoulos, P.; Trevani, A.; Pine, R.; Smith, I. Mycobacterium tuberculosisSigma Factor E Regulon Modulates the Host Inflammatory Response. J. Infect. Dis. 2008, 198, 877–885. [Google Scholar] [CrossRef][Green Version]
- Casonato, S.; Provvedi, R.; Dainese, E.; Palù, G.; Manganelli, R. Mycobacterium tuberculosis Requires the ECF Sigma Factor SigE to Arrest Phagosome Maturation. PLoS ONE 2014, 9, e108893. [Google Scholar] [CrossRef]
- Barik, S.; Sureka, K.; Mukherjee, P.; Basu, J.; Kundu, M. RseA, the SigE specific anti-sigma factor of Mycobacterium tuberculosis, is inactivated by phosphorylation-dependent ClpC1P2 proteolysis. Mol. Microbiol. 2010, 75, 592–606. [Google Scholar] [CrossRef]
- Li, Y.; Zeng, J.; He, Z.G. Characterization of a functional C-terminus of the Mycobacterium tuberculosis MtrA responsible for both DNA binding and interaction with its two-component partner protein, MtrB. J. Biochem. 2010, 148, 549–556. [Google Scholar] [CrossRef]
- Zahrt, T.C.; Deretic, V. An Essential Two-Component Signal Transduction System in Mycobacterium tuberculosis. J. Bacteriol. 2000, 182, 3832–3838. [Google Scholar] [CrossRef]
- Fol, M.; Chauhan, A.; Nair, N.K.; Maloney, E.; Moomey, M.; Jagannath, C.; Madiraju, M.V.V.S.; Rajagopalan, M. Modulation of Mycobacterium tuberculosis proliferation by MtrA, an essential two-component response regulator. Mol. Microbiol. 2006, 60, 643–657. [Google Scholar] [CrossRef]
- Gannoun-Zaki, L.; Alibaud, L.; Carrère-Kremer, S.; Kremer, L.; Blanc-Potard, A.B. Overexpression of the KdpF Membrane Peptide in Mycobacterium bovis BCG Results in Reduced Intramacrophage Growth and Altered Cording Morphology. PLoS ONE 2013, 8, e60379. [Google Scholar] [CrossRef] [PubMed]
- Chalut, C. MmpL transporter-mediated export of cell-wall associated lipids and siderophores in mycobacteria. Tuberculosis 2016, 100, 32–45. [Google Scholar] [CrossRef] [PubMed]
- Viljoen, A.; Dubois, V.; Blaise, M.; Kremer, L.; Girard-Misguich, F.; Girard-Misguich, F.; Herrmann, J.L.; Girard-Misguich, F.; Herrmann, J. The diverse family of MmpL transporters in mycobacteria: From regulation to antimicrobial developments. Mol. Microbiol. 2017, 104, 889–904. [Google Scholar] [CrossRef] [PubMed]
- Olvera, M.R.; Vivès, E.; Molle, V.; Blanc-Potard, A.B.; Gannoun-Zaki, L. Endogenous and Exogenous KdpF Peptide Increases Susceptibility of Mycobacterium bovis BCG to Nitrosative Stress and Reduces Intramacrophage Replication. Front. Microbiol. 2017, 7, 5. [Google Scholar] [CrossRef]
- Datta, P.; Dasgupta, A.; Singh, A.K.; Mukherjee, P.; Kundu, M.; Basu, J. Interaction between FtsW and penicillin-binding protein 3 (PBP3) directs PBP3 to mid-cell, controls cell septation and mediates the formation of a trimeric complex involving FtsZ, FtsW and PBP3 in mycobacteria. Mol. Microbiol. 2006, 62, 1655–1673. [Google Scholar] [CrossRef]
- Dziedzic, R.; Kiran, M.; Plocinski, P.; Ziolkiewicz, M.; Brzostek, A.; Moomey, M.; Vadrevu, I.S.; Dziadek, J.; Madiraju, M.; Rajagopalan, M. Mycobacterium tuberculosis ClpX Interacts with FtsZ and Interferes with FtsZ Assembly. PLoS ONE 2010, 5, e11058. [Google Scholar] [CrossRef]
- Plocinski, P.; Ziolkiewicz, M.; Kiran, M.; Vadrevu, S.I.; Nguyen, H.B.; Hugonnet, J.; Veckerle, C.; Arthur, M.; Dziadek, J.; Cross, T.A.; et al. Characterization of CrgA, a New Partner of the Mycobacterium tuberculosis Peptidoglycan Polymerization Complexes. J. Bacteriol. 2011, 193, 3246–3256. [Google Scholar] [CrossRef]
- Plocinski, P.; Arora, N.; Sarva, K.; Blaszczyk, E.; Qin, H.; Das, N.; Plocinska, R.; Ziolkiewicz, M.; Dziadek, J.; Kiran, M.; et al. Mycobacterium tuberculosis CwsA Interacts with CrgA and Wag31, and the CrgA-CwsA Complex Is Involved in Peptidoglycan Synthesis and Cell Shape Determination. J. Bacteriol. 2012, 194, 6398–6409. [Google Scholar] [CrossRef]
- Donovan, C.; Bramkamp, M. Cell division in Corynebacterineae. Front. Microbiol. 2014, 5, 132. [Google Scholar] [CrossRef]
- Kruh, N.A.; Borgaro, J.G.; Ruzsicska, B.P.; Xu, H.; Tonge, P.J. A novel interaction linking the FAS-II and phthiocerol dimycocerosate (PDIM) biosynthetic pathways. J. Boil. Chem. 2008, 283, 31719–31725. [Google Scholar] [CrossRef]
- Cox, J.S.; Chen, B.; McNeil, M.; Jacobs, W.R. Complex lipid determines tissue-specific replication of Mycobacterium tuberculosis in mice. Nature 1999, 402, 79–83. [Google Scholar] [CrossRef] [PubMed]
- Trivedi, O.A.; Arora, P.; Vats, A.; Ansari, M.Z.; Tickoo, R.; Sridharan, V.; Mohanty, D.; Gokhale, R.S. Dissecting the Mechanism and Assembly of a Complex Virulence Mycobacterial Lipid. Mol. Cell 2005, 17, 631–643. [Google Scholar] [CrossRef] [PubMed]
- Joshi, S.A.; Ball, D.A.; Sun, M.G.; Carlsson, F.; Watkins, B.Y.; Aggarwal, N.; McCracken, J.M.; Huynh, K.K.; Brown, E.J. EccA1, a Component of the Mycobacterium marinum ESX-1 Protein Virulence Factor Secretion Pathway, Regulates Mycolic Acid Lipid Synthesis. Chem. Boil. 2012, 19, 372–380. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.A.G.; Abrahams, K.A.; Alemparte, C.; Ghidelli-Disse, S.; Rullas, J.; Angulo-Barturen, I.; Singh, A.; Gurcha, S.S.; Nataraj, V.; Bethell, S.; et al. THPP target assignment reveals EchA6 as an essential fatty acid shuttle in mycobacteria. Nat. Microbiol. 2016, 1, 15006. [Google Scholar] [CrossRef]
- Larrouy-Maumus, G.; Škovierová, H.; Dhouib, R.; Angala, S.K.; Zuberogoitia, S.; Pham, H.; Villela, A.D.; Mikušová, K.; Noguera, A.; Gilleron, M.; et al. A Small Multidrug Resistance-like Transporter Involved in the Arabinosylation of Arabinogalactan and Lipoarabinomannan in Mycobacteria*. J. Boil. Chem. 2012, 287, 39933–39941. [Google Scholar] [CrossRef]
- Belardinelli, J.M.; Stevens, C.M.; Li, W.; Tan, Y.Z.; Jones, V.; Mancia, F.; Zgurskaya, H.I.; Jackson, M. The MmpL3 interactome reveals a complex crosstalk between cell envelope biosynthesis and cell elongation and division in mycobacteria. Sci. Rep. 2019, 9, 10728. [Google Scholar] [CrossRef]
- Berthet, F. Attenuation of Virulence by Disruption of the Mycobacterium tuberculosis erp Gene. Science 1998, 282, 759–762. [Google Scholar] [CrossRef]
- Klepp, L.I.; Soria, M.; Blanco, F.C.; Bianco, M.V.; Santangelo, M.P.; Cataldi, A.A.; Bigi, F. Identification of two proteins that interact with the Erp virulence factor from Mycobacterium tuberculosis by using the bacterial two-hybrid system. BMC Mol. Boil. 2009, 10, 3. [Google Scholar] [CrossRef]
- Belon, C.; Gannoun-Zaki, L.; Lutfalla, G.; Kremer, L.; Blanc-Potard, A.B. Mycobacterium marinum MgtC Plays a Role in Phagocytosis but is Dispensable for Intracellular Multiplication. PLoS ONE 2014, 9, e116052. [Google Scholar] [CrossRef]
- Belon, C.; Olvera, M.R.; Vives, E.; Kremer, L.; Gannoun-Zaki, L.; Blanc-Potard, A.B. Use of the Salmonella MgtR peptide as an antagonist of the Mycobacterium MgtC virulence factor. Future Microbiol. 2016, 11, 215–225. [Google Scholar] [CrossRef]
- Alix, E.; Blanc-Potard, A.B. Peptide-assisted degradation of the Salmonella MgtC virulence factor. EMBO J. 2008, 27, 546–557. [Google Scholar] [CrossRef] [PubMed]
- Pethe, K.; Alonso, S.; Biet, F.; Delogu, G.; Brennan, M.J.; Locht, C.; Menozzi, F.D. The heparin-binding haemagglutinin of M. tuberculosis is required for extrapulmonary dissemination. Nature 2001, 412, 190–194. [Google Scholar] [CrossRef] [PubMed]
- Raze, D.; Verwaerde, C.; Deloison, G.; Werkmeister, E.; Coupin, B.; Loyens, M.; Brodin, P.; Rouanet, C.; Locht, C. Heparin-Binding Hemagglutinin Adhesin (HBHA) Is Involved in Intracytosolic Lipid Inclusions Formation in Mycobacteria. Front. Microbiol. 2018, 9, 2258. [Google Scholar] [CrossRef] [PubMed]
- Veyron-Churlet, R.; Dupres, V.; Saliou, J.M.; Lafont, F.; Raze, D.; Locht, C. Rv0613c/MSMEG_1285 Interacts with HBHA and Mediates Its Proper Cell-Surface Exposure in Mycobacteria. Int. J. Mol. Sci. 2018, 19, 1673. [Google Scholar] [CrossRef]
- Tharad, M.; Samuchiwal, S.K.; Bhalla, K.; Ghosh, A.; Kumar, K.; Kumar, S.; Ranganathan, A. A Three-Hybrid System to Probe In Vivo Protein-Protein Interactions: Application to the Essential Proteins of the RD1 Complex of M. tuberculosis. PLoS ONE 2011, 6, e27503. [Google Scholar] [CrossRef]
- Wang, Y.; Cui, T.; Zhang, C.; Yang, M.; Huang, Y.; Li, W.; Zhang, L.; Gao, C.H.; He, Y.; Li, Y.; et al. Global Protein−Protein Interaction Network in the Human PathogenMycobacterium tuberculosisH37Rv. J. Proteome Res. 2010, 9, 6665–6677. [Google Scholar] [CrossRef]
- Singh, A.; Mai, D.; Kumar, A.; Steyn, A.J.C. Dissecting virulence pathways of Mycobacterium tuberculosis through protein–protein association. Proc. Natl. Acad. Sci. USA 2006, 103, 11346–11351. [Google Scholar] [CrossRef]
- Chao, J.D.; Papavinasasundaram, K.G.; Zheng, X.; Chávez-Steenbock, A.; Wang, X.; Lee, G.Q.; Av-Gay, Y. Convergence of Ser/Thr and Two-component Signaling to Coordinate Expression of the Dormancy Regulon in Mycobacterium tuberculosis*. J. Boil. Chem. 2010, 285, 29239–29246. [Google Scholar] [CrossRef]
- Gupta, S.; Banerjee, S.K.; Chatterjee, A.; Sharma, A.K.; Kundu, M.; Basu, J. Essential protein SepF of mycobacteria interacts with FtsZ and MurG to regulate cell growth and division. Microbiology 2015, 161, 1627–1638. [Google Scholar] [CrossRef]
- Angala, S.K.; Belardinelli, J.M.; Huc-Claustre, E.; Wheat, W.H.; Jackson, M. The cell envelope glycoconjugates of Mycobacterium tuberculosis. Crit. Rev. Biochem. Mol. Boil. 2014, 49, 361–399. [Google Scholar] [CrossRef]
- Munshi, T.; Gupta, A.; Evangelopoulos, D.; Guzmán, J.D.; Gibbons, S.; Keep, N.H.; Bhakta, S. Characterisation of ATP-Dependent Mur Ligases Involved in the Biogenesis of Cell Wall Peptidoglycan in Mycobacterium tuberculosis. PLoS ONE 2013, 8, e60143. [Google Scholar] [CrossRef] [PubMed]
- O’Hare, H.; Juillerat, A.; Dianišková, P.; Johnsson, K. A split-protein sensor for studying protein–protein interaction in mycobacteria. J. Microbiol. Methods 2008, 73, 79–84. [Google Scholar] [CrossRef] [PubMed]
- Lougheed, K.E.; Bennett, M.H.; Williams, H.D. An in vivo crosslinking system for identifying mycobacterial protein-protein interactions. J. Microbiol. Methods 2014, 105, 67–71. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Robbins, S.; Guo, J.; Shen, W.; Schultz, P.G. Genetic Incorporation of Unnatural Amino Acids into Proteins in Mycobacterium tuberculosis. PLoS ONE 2010, 5, e9354. [Google Scholar] [CrossRef] [PubMed]
- Touchette, M.H.; Van Vlack, E.R.; Bai, L.; Kim, J.; Cognetta, A.B., III; Previti, M.L.; Backus, K.M.; Martin, D.W.; Cravatt, B.F.; Seeliger, J.C. A Screen for Protein-Protein Interactions in Live Mycobacteria Reveals a Functional Link between the Virulence-Associated Lipid Transporter LprG and the Mycolyltransferase Antigen 85A. ACS Infect. Dis. 2017, 3, 336–348. [Google Scholar] [CrossRef]
- Shukla, S.; Richardson, E.T.; Athman, J.J.; Shi, L.; Wearsch, P.A.; McDonald, D.; Banaei, N.; Boom, W.H.; Jackson, M.; Harding, C.V. Mycobacterium tuberculosis Lipoprotein LprG Binds Lipoarabinomannan and Determines Its Cell Envelope Localization to Control Phagolysosomal Fusion. PLoS Pathog. 2014, 10, e1004471. [Google Scholar] [CrossRef]
- Gaur, R.L.; Ren, K.; Blumenthal, A.; Bhamidi, S.; Gibbs, S.; Jackson, M.; Zare, R.N.; Ehrt, S.; Ernst, J.D.; Banaei, N. LprG-Mediated Surface Expression of Lipoarabinomannan Is Essential for Virulence of Mycobacterium tuberculosis. PLoS Pathog. 2014, 10, e1004376. [Google Scholar] [CrossRef]
- Martinot, A.J.; Farrow, M.; Bai, L.; Layre, E.; Cheng, T.Y.; Tsai, J.H.; Iqbal, J.; Annand, J.W.; Sullivan, Z.A.; Hussain, M.M.; et al. Mycobacterial Metabolic Syndrome: LprG and Rv1410 Regulate Triacylglyceride Levels, Growth Rate and Virulence in Mycobacterium tuberculosis. PLoS Pathog. 2016, 12, e1005351. [Google Scholar] [CrossRef]
- Robinson, K.E.; Reardon, P.N.; Spicer, L.D. In-cell NMR spectroscopy in Escherichia coli. Methods Mol. Biol. 2012, 831, 261–277. [Google Scholar] [CrossRef]
- Sciolino, N.; Burz, D.S.; Shekhtman, A. In-Cell NMR Spectroscopy of Intrinsically Disordered Proteins. Proteomics 2019, 19, 1800055. [Google Scholar] [CrossRef]
- Darwin, K.H. Prokaryotic ubiquitin-like protein (Pup), proteasomes and pathogenesis. Nat. Rev. Genet. 2009, 7, 485–491. [Google Scholar] [CrossRef] [PubMed]
- Maldonado, A.Y.; Burz, D.S.; Reverdatto, S.; Shekhtman, A. Fate of Pup inside the Mycobacterium Proteasome Studied by in-Cell NMR. PLoS ONE 2013, 8, e74576. [Google Scholar] [CrossRef] [PubMed]
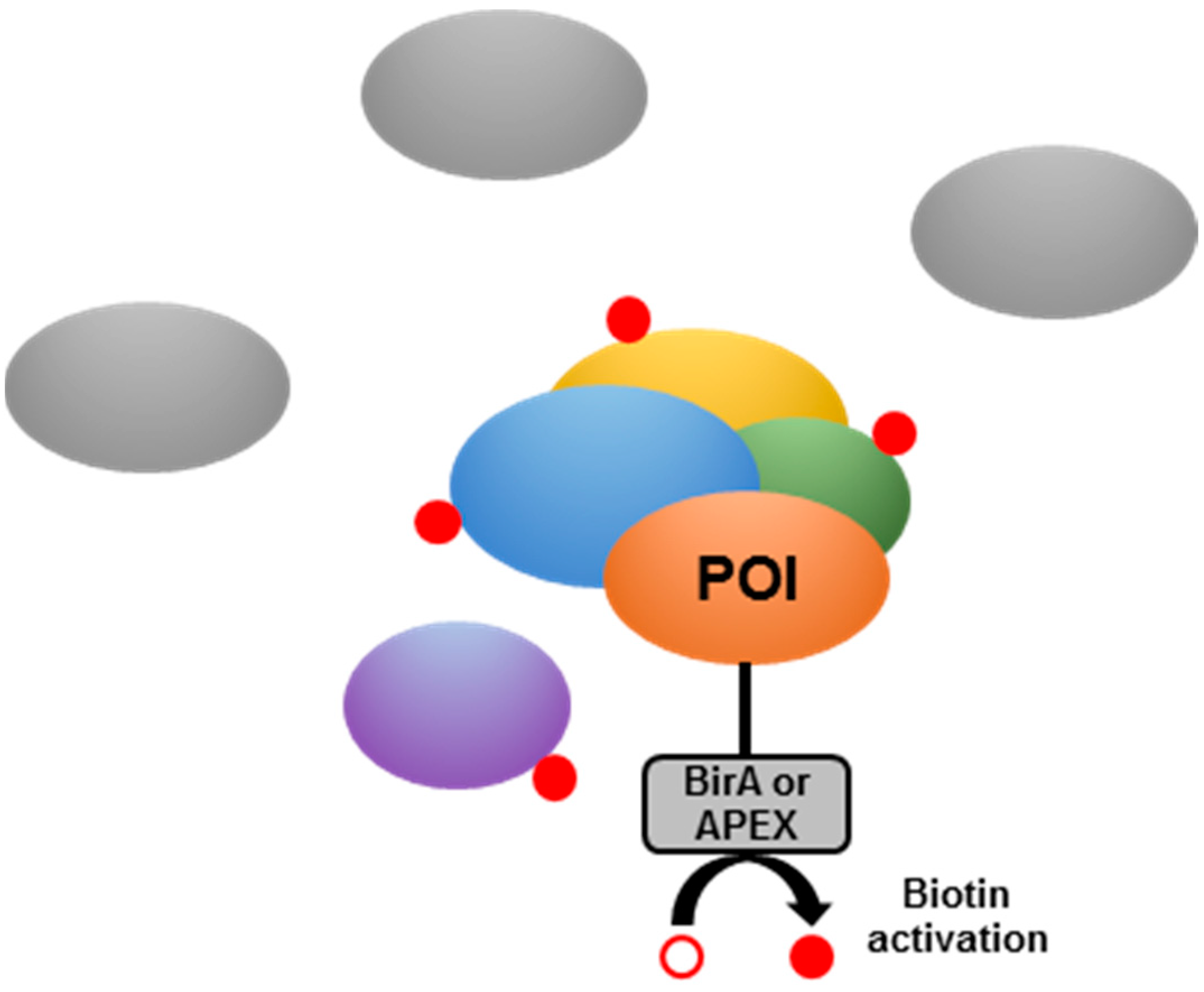
- Chen, C.L.; Perrimon, N. Proximity-dependent labeling methods for proteomic profiling in living cells. Wiley Interdiscip. Rev. Dev. Boil. 2017, 6, e272. [Google Scholar] [CrossRef] [PubMed]
- Roux, K.J.; Kim, D.I.; Raida, M.; Burke, B. A promiscuous biotin ligase fusion protein identifies proximal and interacting proteins in mammalian cells. J. Cell Boil. 2012, 196, 801–810. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.I.; Jensen, S.C.; Noble, K.A.; KC, B.; Roux, K.H.; Motamedchaboki, K.; Roux, K.J. An improved smaller biotin ligase for BioID proximity labeling. Mol. Boil. Cell 2016, 27, 1188–1196. [Google Scholar] [CrossRef]
- Rhee, H.W.; Zou, P.; Udeshi, N.D.; Martell, J.D.; Mootha, V.K.; Carr, S.A.; Ting, A.Y. Proteomic mapping of mitochondria in living cells via spatially restricted enzymatic tagging. Science 2013, 339, 1328–1331. [Google Scholar] [CrossRef]
- Lobingier, B.T.; Hüttenhain, R.; Eichel, K.; Miller, K.B.; Ting, A.Y.; Von Zastrow, M.; Krogan, N.J. An Approach to Spatiotemporally Resolve Protein Interaction Networks in Living Cells. Cell 2017, 169, 350–360. [Google Scholar] [CrossRef]
- Salaemae, W.; Polyak, S.W.; Booker, G.W. The Role of Biotin in Bacterial Physiology and Virulence: A Novel Antibiotic Target for Mycobacterium tuberculosis. Virulence Mechanisms of Bacterial Pathogens, 5th ed.; ASM Press: Washington, DC, USA, 2016; Volume 4, pp. 797–822. [Google Scholar] [CrossRef]
| Methods | Contact | Membrane Proteins | Nature of the Interaction | Cellular Context | PTM and Cofactors | HTS |
|---|---|---|---|---|---|---|
| Y2H | direct | no | binary | no | no | yes |
| BACTH | direct | yes | binary | yes/no | yes/no | yes |
| M-PFC | direct | yes | binary | yes | yes | yes |
| Split-Trp | direct | yes | binary | yes | yes | yes |
| Crosslinking | direct | yes | complex | yes | yes | no |
| In-cell NMR | proximity | yes | complex | yes/no | yes/no | no |
| Biotinylation | proximity | yes | complex | yes | yes | yes |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Veyron-Churlet, R.; Locht, C. In Vivo Methods to Study Protein–Protein Interactions as Key Players in Mycobacterium Tuberculosis Virulence. Pathogens 2019, 8, 173. https://doi.org/10.3390/pathogens8040173
Veyron-Churlet R, Locht C. In Vivo Methods to Study Protein–Protein Interactions as Key Players in Mycobacterium Tuberculosis Virulence. Pathogens. 2019; 8(4):173. https://doi.org/10.3390/pathogens8040173
Chicago/Turabian StyleVeyron-Churlet, Romain, and Camille Locht. 2019. "In Vivo Methods to Study Protein–Protein Interactions as Key Players in Mycobacterium Tuberculosis Virulence" Pathogens 8, no. 4: 173. https://doi.org/10.3390/pathogens8040173
APA StyleVeyron-Churlet, R., & Locht, C. (2019). In Vivo Methods to Study Protein–Protein Interactions as Key Players in Mycobacterium Tuberculosis Virulence. Pathogens, 8(4), 173. https://doi.org/10.3390/pathogens8040173





