Genetic Diversity and Molecular Epidemiology of Mycobacterium tuberculosis Complex Clinical Isolates in New Brunswick, Canada—A Retrospective Chart Review
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Setting and Design
2.2. Study Samples
2.3. Identification, Drug Susceptibility Testing, and Genotyping
3. Results
3.1. Descriptive Statistics
3.2. Drug Susceptibility Testing
3.3. MIRU-VNTR Genotyping Results
4. Discussion
Limitations
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| BCG | Bacillus Calmette–Guérin |
| CAP | Capreomycin |
| CAS | Central Asian Strain |
| DNA | Deoxyribonucleic Acid |
| DST | Drug Susceptibility Testing |
| EAI | East African Indian |
| HGDI | Hunter and Gaston Discriminatory Index |
| INH | Isoniazid |
| LAM | Latin American-Mediterranean |
| MDR | Multidrug-resistant |
| MGIT | Mycobacteria Growth Indicator Tube |
| MIRU-VNTR | Mycobacterial Interspersed Repetitive Unit-Variable Number Tandem Repeat |
| MTBC | Mycobacterium tuberculosis complex |
| NB | New Brunswick |
| PAS | p-aminosalicylic acid |
| PZA | Pyrazinamide |
| TBD | Tuberculosis disease |
| TBI | Tuberculosis infection |
| TUR | Turkey |
| UPGMA | Unweighted Pair Group Method with Arithmetic Averages |
| WHO | World Health Organization |
References
- World Health Organization. Global Tuberculosis Report 2025. Available online: https://iris.who.int/server/api/core/bitstreams/e97dd6f4-b567-4396-8680-717bac6869a9/content (accessed on 2 January 2026).
- Public Health Agency of Canada. Tuberculosis Disease in Canada, 2023 (Infographic). Available online: https://www.canada.ca/en/public-health/services/publications/diseases-conditions/tuberculosis-disease-2023-infographic.html (accessed on 2 January 2026).
- Public Health Agency of Canada. Tuberculosis in Canada, Epidemiological Update 2022. Available online: https://www.canada.ca/content/dam/phac-aspc/documents/services/publications/diseases-conditions/tuberculosis-canada-epidemiological-update-2022/tuberculosis-canada-epidemiological-update-2022.pdf (accessed on 2 January 2026).
- Statistics Canada. Census Profile, 2021 Census of Population: New Brunswick. Available online: https://www12.statcan.gc.ca/census-recensement/2021/dp-pd/prof/details/page.cfm?Lang=E&SearchText=New%20Brunswick&DGUIDlist=2021A000213&GENDERlist=1,2,3&STATISTIClist=1&HEADERlist=0 (accessed on 2 January 2026).
- Mounchili, A.; Perera, R.; Lee, R.S.; Njoo, H.; Brooks, J. Chapter 1: Epidemiology of tuberculosis in Canada. Can. J. Respir. Crit. Care Sleep Med. 2022, 6, 8–21. [Google Scholar] [CrossRef]
- Public Health Agency of Canada. Tuberculosis Disease Surveillance: Summary. Available online: https://health-infobase.canada.ca/tuberculosis/ (accessed on 2 January 2026).
- Shamputa, I.C.; Nguyen, D.T.K.; Mackenzie, H.; Gaudet, D.J.; Harquail, A.; Barker, K.; Webster, D. Enhanced screening for tuberculosis infection among immigrants in southern New Brunswick: A cross-sectional pilot study. Can. Commun. Dis. Rep. 2025, 51, 167–178. [Google Scholar] [CrossRef]
- World Health Organization. Towards Tuberculosis Elimination: An Action Framework for Low-Incidence Countries. WHO/HTM/TB/2014.13. Available online: https://iris.who.int/bitstream/handle/10665/132231/9789241507707_eng.pdf?sequence=1 (accessed on 2 January 2026).
- Albanna, A.S.; Reed, M.B.; Kotar, K.V.; Fallow, A.; McIntosh, F.A.; Behr, M.A.; Menzies, D. Reduced transmissibility of East African Indian strains of Mycobacterium tuberculosis. PLoS ONE 2011, 6, e25075. [Google Scholar] [CrossRef]
- Alvarez, G.G.; Zwerling, A.A.; Duncan, C.; Pease, C.; Van Dyk, D.; Behr, M.A.; Lee, R.S.; Mulpuru, S.; Pakhale, S.; Cameron, D.W.; et al. Molecular epidemiology of Mycobacterium tuberculosis to describe the transmission dynamics among Inuit residing in Iqaluit Nunavut using whole-genome sequencing. Clin. Infect. Dis. 2021, 72, 2187–2195. [Google Scholar] [CrossRef]
- Gardy, J.L.; Johnston, J.C.; Ho Sui, S.J.; Cook, V.J.; Shah, L.; Brodkin, E.; Rempel, S.; Moore, R.; Zhao, Y.; Holt, R.; et al. Whole-Genome Sequencing and Social-Network Analysis of a Tuberculosis Outbreak. N. Engl. J. Med. 2011, 364, 730–739. [Google Scholar] [CrossRef]
- Lee, R.S.; Radomski, N.; Proulx, J.F.; Manry, J.; McIntosh, F.; Desjardins, F.; Soualhine, H.; Domenech, P.; Reed, M.B.; Menzies, D.; et al. Reemergence and amplification of tuberculosis in the Canadian arctic. J. Infect. Dis. 2015, 211, 1905–1914. [Google Scholar] [CrossRef]
- Lee, R.S.; Proulx, J.F.; McIntosh, F.; Behr, M.A.; Hanage, W.P. Previously undetected super-spreading of Mycobacterium tuberculosis revealed by deep sequencing. eLife 2020, 9, e53245. [Google Scholar] [CrossRef]
- Moreau, D.; Gratrix, J.; Kunimoto, D.; Beckon, A.; Der, E.; Hansen, E.; Chui, L.; Ahmed, R. A shelter-associated tuberculosis outbreak: A novel strain introduced through foreign-born populations. Can. J. Public Health 2012, 103, e408–e412. [Google Scholar] [CrossRef]
- Bandera, A.; Gori, A.; Catozzi, L.; Degli Esposti, A.; Marchetti, G.; Molteni, C.; Ferrario, G.; Codecasa, L.; Penati, V.; Matteelli, A.; et al. Molecular epidemiology study of exogenous reinfection in an area with a low incidence of tuberculosis. J. Clin. Microbiol. 2001, 39, 2213–2218. [Google Scholar] [CrossRef]
- van Rie, A.; Warren, R.; Richardson, M.; Victor, T.C.; Gie, R.P.; Enarson, D.A.; Beyers, N.; van Helden, P.D. Exogenous reinfection as a cause of recurrent tuberculosis after curative treatment. N. Engl. J. Med. 1999, 341, 1174–1179. [Google Scholar] [CrossRef]
- Kamerbeek, J.; Schouls, L.; Kolk, A.; van Agterveld, M.; van Soolingen, D.; Kuijper, S.; Bunschoten, A.; Molhuizen, H.; Shaw, R.; Goyal, M.; et al. Simultaneous detection and strain differentiation of Mycobacterium tuberculosis for diagnosis and epidemiology. J. Clin. Microbiol. 1997, 35, 907–914. [Google Scholar] [CrossRef]
- van Embden, J.D.; Cave, M.D.; Crawford, J.T.; Dale, J.W.; Eisenach, K.D.; Gicquel, B.; Hermans, P.; Martin, C.; McAdam, R.; Shinnick, T.M. Strain identification of Mycobacterium tuberculosis by DNA fingerprinting: Recommendations for a standardized methodology. J. Clin. Microbiol. 1993, 31, 406–409. [Google Scholar] [CrossRef]
- Supply, P.; Allix, C.; Lesjean, S.; Cardoso-Oelemann, M.; Rüsch-Gerdes, S.; Willery, E.; Savine, E.; de Haas, P.; van Deutekom, H.; Roring, S.; et al. Proposal for standardization of optimized mycobacterial interspersed repetitive unit-variable-number tandem repeat typing of Mycobacterium tuberculosis. J. Clin. Microbiol. 2006, 44, 4498–4510. [Google Scholar] [CrossRef]
- Sharma, M.K.; Janella, D.; McGurran, A.; Corbett, C.; Adam, H.; Akochy, P.M.; Haldane, D.; MacKenzie, H.; Minion, J.; Needle, R.; et al. Compilation of 10 Years of MIRU-VNTR Data: Canadian National Tuberculosis Laboratory’s Experience. Can. J. Infect. Dis. Med. Microbiol. 2022, 2022, 3505142. [Google Scholar] [CrossRef]
- Islam, M.R.; Sharma, M.K.; KhunKhun, R.; Shandro, C.; Sekirov, I.; Tyrrell, G.J.; Soualhine, H. Whole genome sequencing-based identification of human tuberculosis caused by animal-lineage Mycobacterium orygis. J. Clin. Microbiol. 2023, 61, e0026023. [Google Scholar] [CrossRef]
- Guthrie, J.L.; Delli Pizzi, A.; Roth, D.; Kong, C.; Jorgensen, D.; Rodrigues, M.; Tang, P.; Cook, V.J.; Johnston, J.; Gardy, J.L. Genotyping and Whole-genome sequencing to identify tuberculosis transmission to pediatric patients in British Columbia, Canada, 2005–2014. J. Infect. Dis. 2018, 218, 1155–1163. [Google Scholar] [CrossRef]
- Guthrie, J.L.; Strudwick, L.; Roberts, B.; Allen, M.; McFadzen, J.; Roth, D.; Jorgensen, D.; Rodrigues, M.; Tang, P.; Hanley, B.; et al. Whole genome sequencing for improved understanding of Mycobacterium tuberculosis transmission in a remote circumpolar region. Epidemiol. Infect. 2019, 147, e188. [Google Scholar] [CrossRef]
- Romanowski, K.; Sobkowiak, B.; Guthrie, J.L.; Cook, V.J.; Gardy, J.L.; Johnston, J.C. Using Whole-genome sequencing to determine the timing of secondary tuberculosis in British Columbia, Canada. Clin. Infect. Dis. 2021, 73, 535–537. [Google Scholar] [CrossRef]
- Tyler, A.D.; Randell, E.; Baikie, M.; Antonation, K.; Janella, D.; Christianson, S.; Tyrrell, G.J.; Graham, M.; Van Domselaar, G.; Sharma, M.K. Application of whole genome sequence analysis to the study of Mycobacterium tuberculosis in Nunavut, Canada. PLoS ONE 2017, 12, e0185656. [Google Scholar] [CrossRef]
- Hernández-Garduño, E.; Kunimoto, D.; Wang, L.; Rodrigues, M.; Elwood, R.K.; Black, W.; Mak, S.; FitzGerald, J.M. Predictors of clustering of tuberculosis in Greater Vancouver: A molecular epidemiologic study. CMAJ 2002, 167, 349–352. [Google Scholar]
- Hernández-Garduño, E.; Cook, V.; Kunimoto, D.; Elwood, R.K.; Black, W.A.; FitzGerald, J.M. Transmission of tuberculosis from smear negative patients: A molecular epidemiology study. Thorax 2004, 59, 286–290. [Google Scholar] [CrossRef]
- Cheng, J.M.; Hiscoe, L.; Pollock, S.L.; Hasselback, P.; Gardy, J.L.; Parker, R. A clonal outbreak of tuberculosis in a homeless population in the interior of British Columbia, Canada, 2008–2015. Epidemiol. Infect. 2015, 143, 3220–3226. [Google Scholar] [CrossRef]
- Martin, M.A.; Lee, R.S.; Cowley, L.A.; Gardy, J.L.; Hanage, W.P. Within-host Mycobacterium tuberculosis diversity and its utility for inferences of transmission. Microb. Genom. 2018, 4, e000217. [Google Scholar] [CrossRef]
- Guthrie, J.L.; Kong, C.; Roth, D.; Jorgensen, D.; Rodrigues, M.; Hoang, L.; Tang, P.; Cook, V.; Johnston, J.; Gardy, J.L. Molecular epidemiology of tuberculosis in British Columbia, Canada: A 10-Year Retrospective Study. Clin. Infect. Dis. 2018, 66, 849–856. [Google Scholar] [CrossRef]
- Guthrie, J.L.; Kong, C.; Roth, D.; Jorgensen, D.; Rodrigues, M.; Tang, P.; Thejoe, M.; Elwood, K.; Cook, V.J.; Johnston, J.; et al. Universal genotyping for tuberculosis prevention programs: A 5-year comparison with on-request genotyping. J. Clin. Microbiol. 2018, 56, e01778-17. [Google Scholar] [CrossRef]
- Khan, K.; Hirji, M.M.; Miniota, J.; Hu, W.; Wang, J.; Gardam, M.; Rawal, S.; Ellis, E.; Chan, A.; Creatore, M.I.; et al. Domestic impact of tuberculosis screening among new immigrants to Ontario, Canada. CMAJ 2015, 187, E473–E481. [Google Scholar] [CrossRef]
- Alexander, D.C.; Guthrie, J.L.; Pyskir, D.; Maki, A.; Kurepina, N.; Kreiswirth, B.N.; Chedore, P.; Drews, S.J.; Jamieson, F. Mycobacterium tuberculosis in Ontario, Canada: Insights from IS6110 restriction fragment length polymorphism and mycobacterial interspersed repetitive-unit-variable-number tandem-repeat genotyping. J. Clin. Microbiol. 2009, 47, 2651–2654. [Google Scholar] [CrossRef]
- Kulaga, S.; Behr, M.; Musana, K.; Brinkman, J.; Menzies, D.; Brassard, P.; Kunimoto, D.; Tannenbaum, T.N.; Thibert, L.; Joseph, L.; et al. Molecular epidemiology of tuberculosis in Montreal. CMAJ 2002, 167, 353–354. [Google Scholar]
- Kulaga, S.; Behr, M.; Nguyen, D.; Brinkman, J.; Westley, J.; Menzies, D.; Brassard, P.; Tannenbaum, T.; Thibert, L.; Boivin, J.F.; et al. Diversity of Mycobacterium tuberculosis isolates in an immigrant population: Evidence against a founder effect. Am. J. Epidemiol. 2004, 159, 507–513. [Google Scholar] [CrossRef]
- Nguyen, D.; Proulx, J.F.; Westley, J.; Thibert, L.; Dery, S.; Behr, M.A. Tuberculosis in the Inuit community of Quebec, Canada. Am. J. Respir. Crit. Care Med. 2003, 168, 1353–1357. [Google Scholar] [CrossRef]
- Blackwood, K.S.; Al-Azem, A.; Elliott, L.J.; Hershfield, E.S.; Kabani, A.M. Conventional and molecular epidemiology of tuberculosis in Manitoba. BMC Infect. Dis. 2003, 3, 18. [Google Scholar] [CrossRef]
- Riopel, N.D.; Long, R.; Heffernan, C.; Tyrrell, G.J.; Shandro, C.; Li, V.; Islam, M.R.; Stobart, M.; Sharma, M.K.; Soualhine, H.; et al. Characterization of Mycobacterium orygis, Mycobacterium bovis, and Mycobacterium caprae Infections in humans in Western Canada. J. Infect. Dis. 2024, 230, e789–e797. [Google Scholar] [CrossRef]
- Kunimoto, D.; Chedore, P.; Allen, R.; Kasatiya, S. Investigation of tuberculosis transmission in Canadian Arctic Inuit communities using DNA fingerprinting. Int. J. Tuberc. Lung Dis. 2001, 5, 642–647. [Google Scholar]
- Case, C.; Kandola, K.; Chui, L.; Li, V.; Nix, N.; Johnson, R. Examining DNA fingerprinting as an epidemiology tool in the tuberculosis program in the Northwest Territories, Canada. Int. J. Circumpolar Health 2013, 72, 20067. [Google Scholar] [CrossRef]
- Christianson, S.; Wolfe, J.; Orr, P.; Karlowsky, J.; Levett, P.N.; Horsman, G.B.; Thibert, L.; Tang, P.; Sharma, M.K. Evaluation of 24 locus MIRU-VNTR genotyping of Mycobacterium tuberculosis isolates in Canada. Tuberculosis 2010, 90, 31–38. [Google Scholar] [CrossRef]
- FitzGerald, J.M.; Fanning, A.; Hoepnner, V.; Hershfield, E.; Kunimoto, D.; Canadian Molecular Epidemiology of TB Study Group. The molecular epidemiology of tuberculosis in western Canada. Int. J. Tuberc. Lung Dis. 2003, 7, 132–138. [Google Scholar]
- Patel, S.; Paulsen, C.; Heffernan, C.; Saunders, D.; Sharma, M.; King, M.; Hoeppner, V.; Orr, P.; Kunimoto, D.; Menzies, D.; et al. Tuberculosis transmission in the Indigenous peoples of the Canadian prairies. PLoS ONE 2017, 12, e0188189. [Google Scholar] [CrossRef]
- Statistics Canada. Health Regions: Boundaries and Correspondence with Census Geography—Map 4 New Brunswick Health Regions, 2022 (Catalogue No. 82-402-X); Statistics Canada: Ottawa, ON, Canada, 2022; Available online: https://www150.statcan.gc.ca/n1/pub/82-402-x/2023001/rm-cr/rm-cr04-eng.htm (accessed on 2 January 2026).
- Labbé, G.; Kruczkiewicz, P.; Robertson, J.; Mabon, P.; Schonfeld, J.; Kein, D.; Rankin, M.A.; Gopez, M.; Hole, D.; Son, D.; et al. Rapid and accurate SNP genotyping of clonal bacterial pathogens with BioHansel. Microb. Genom. 2021, 7, 000651. [Google Scholar] [CrossRef]
- Sharma, M.K.; Stobart, M.; Akochy, P.M.; Adam, H.; Janella, D.; Rabb, M.; Alawa, M.; Sekirov, I.; Tyrrell, G.J.; Soualhine, H. Evaluation of whole genome sequencing-based predictions of antimicrobial resistance to TB first line agents: A lesson from 5 years of data. Int. J. Mol. Sci. 2024, 25, 6245. [Google Scholar] [CrossRef]
- Allix-Béguec, C.; Harmsen, D.; Weniger, T.; Supply, P.; Niemann, S. Evaluation and strategy for use of MIRU-VNTRplus, a multifunctional database for online analysis of genotyping data and phylogenetic identification of Mycobacterium tuberculosis complex isolates. J. Clin. Microbiol. 2008, 46, 2692–2699. [Google Scholar] [CrossRef]
- Weniger, T.; Krawczyk, J.; Supply, P.; Niemann, S.; Harmsen, D. MIRU-VNTRplus: A web tool for polyphasic genotyping of Mycobacterium tuberculosis complex bacteria. Nucleic Acids Res. 2010, 38, W326–W331. [Google Scholar] [CrossRef]
- Hunter, P.R.; Gaston, M.A. Numerical index of the discriminatory ability of typing systems: An application of Simpson’s index of diversity. J. Clin. Microbiol. 1988, 26, 2465–2466. [Google Scholar] [CrossRef] [PubMed]
- Sola, C.; Filliol, I.; Legrand, E.; Lesjean, S.; Locht, C.; Supply, P.; Rastogi, N. Genotyping of the Mycobacterium tuberculosis complex using MIRUs: Association with VNTR and spoligotyping for molecular epidemiology and evolutionary genetics. Infect. Genet. Evol. 2003, 3, 125–133. [Google Scholar] [CrossRef]
- Meehan, C.J.; Moris, P.; Kohl, T.A.; Pečerska, J.; Akter, S.; Merker, M.; Utpatel, C.; Beckert, P.; Gehre, F.; Lempens, P.; et al. The relationship between transmission time and clustering methods in Mycobacterium tuberculosis epidemiology. EBioMedicine 2018, 37, 410–416. [Google Scholar] [CrossRef] [PubMed]
- Small, P.M.; Hopewell, P.C.; Singh, S.P.; Paz, A.; Parsonnet, J.; Ruston, D.C.; Schecter, G.F.; Daley, C.L.; Schoolnik, G.K. The epidemiology of tuberculosis in San Francisco. A population-based study using conventional and molecular methods. N. Engl. J. Med. 1994, 330, 1703–1709. [Google Scholar] [CrossRef] [PubMed]
- Public Health Agency of Canada. Tuberculosis in Canada: 2012–2021 Expanded Report. Available online: https://www.canada.ca/content/dam/phac-aspc/documents/services/publications/diseases-conditions/tuberculosis-canada-expanded-report-2012-2021/tuberculosis-canada-expanded-report-2012-2021.pdf (accessed on 2 January 2026).
- Chen, Y.Y.; Chang, J.R.; Huang, W.F.; Hsu, C.H.; Cheng, H.Y.; Sun, J.R.; Kuo, S.C.; Su, I.J.; Lin, M.S.; Chen, W.; et al. Genetic diversity of the Mycobacterium tuberculosis East African-Indian family in three tropical Asian countries. J. Microbiol. Immunol. Infect. 2017, 50, 886–892. [Google Scholar] [CrossRef]
- Greenaway, C.; Diefenbach-Elstob, T.; Schwartzman, K.; Cook, V.J.; Giovinazzo, G.; Njoo, H.; Mounchili, A.; Brooks, J. Chapter 13: Tuberculosis surveillance and tuberculosis infection testing and treatment in migrants. Can. J. Respir. Crit. Care Sleep Med. 2022, 6, 194–204. [Google Scholar] [CrossRef]
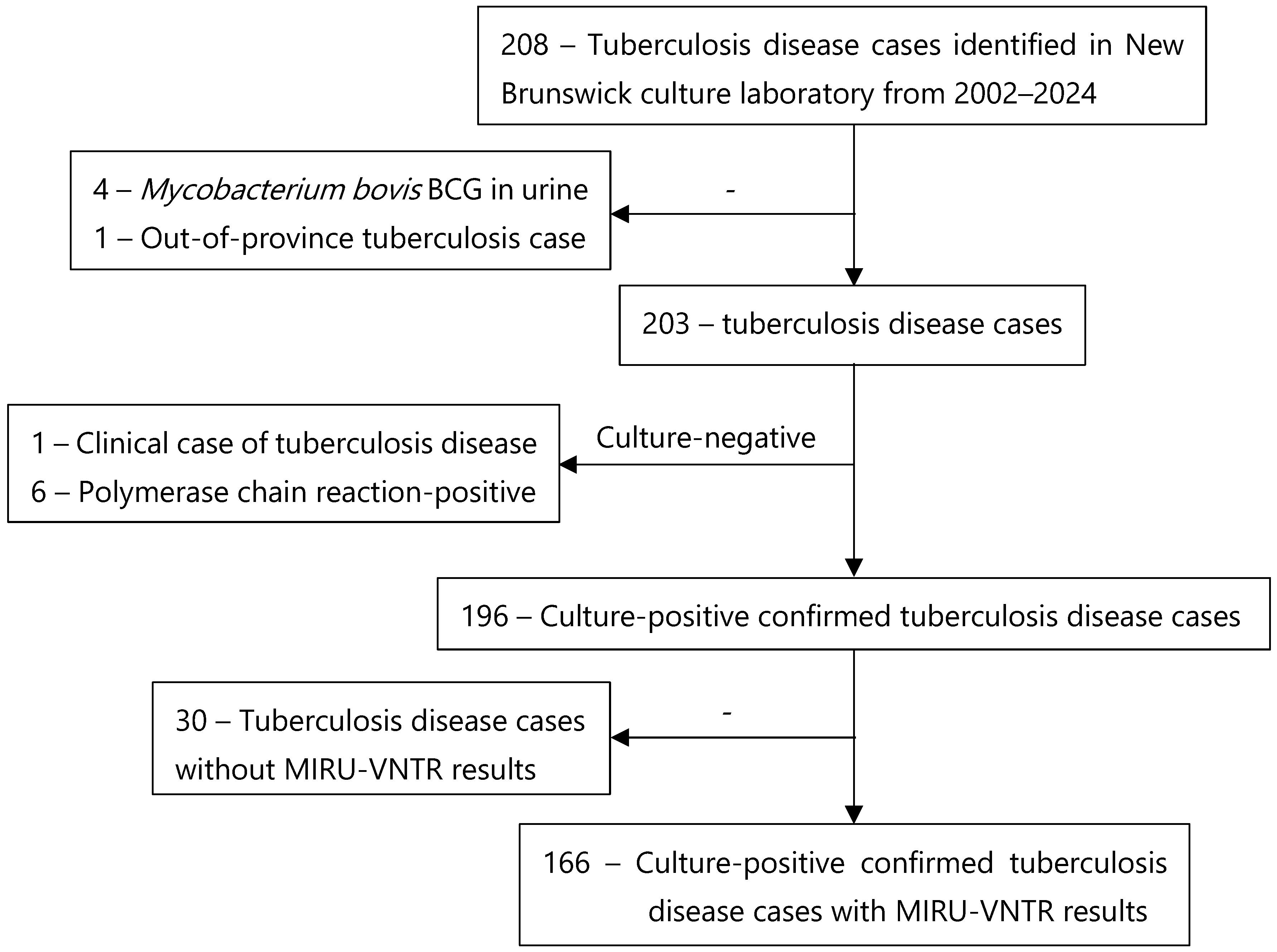


| Characteristics | Biological Sex | Total (Percentage) | |
|---|---|---|---|
| M | F | ||
| Age | 80 | 86 | 166 (100) |
| Age Group | |||
| <15 | 0 | 2 | 2 (1.2) |
| 15–24 | 17 | 17 | 34 (20.5) |
| 25–34 | 17 | 17 | 34 (20.5) |
| 35–44 | 3 | 12 | 15 (9.0) |
| 45–54 | 14 | 9 | 23 (13.9) |
| 55–64 | 7 | 7 | 14 (8.4) |
| >65 | 22 | 22 | 44 (26.5) |
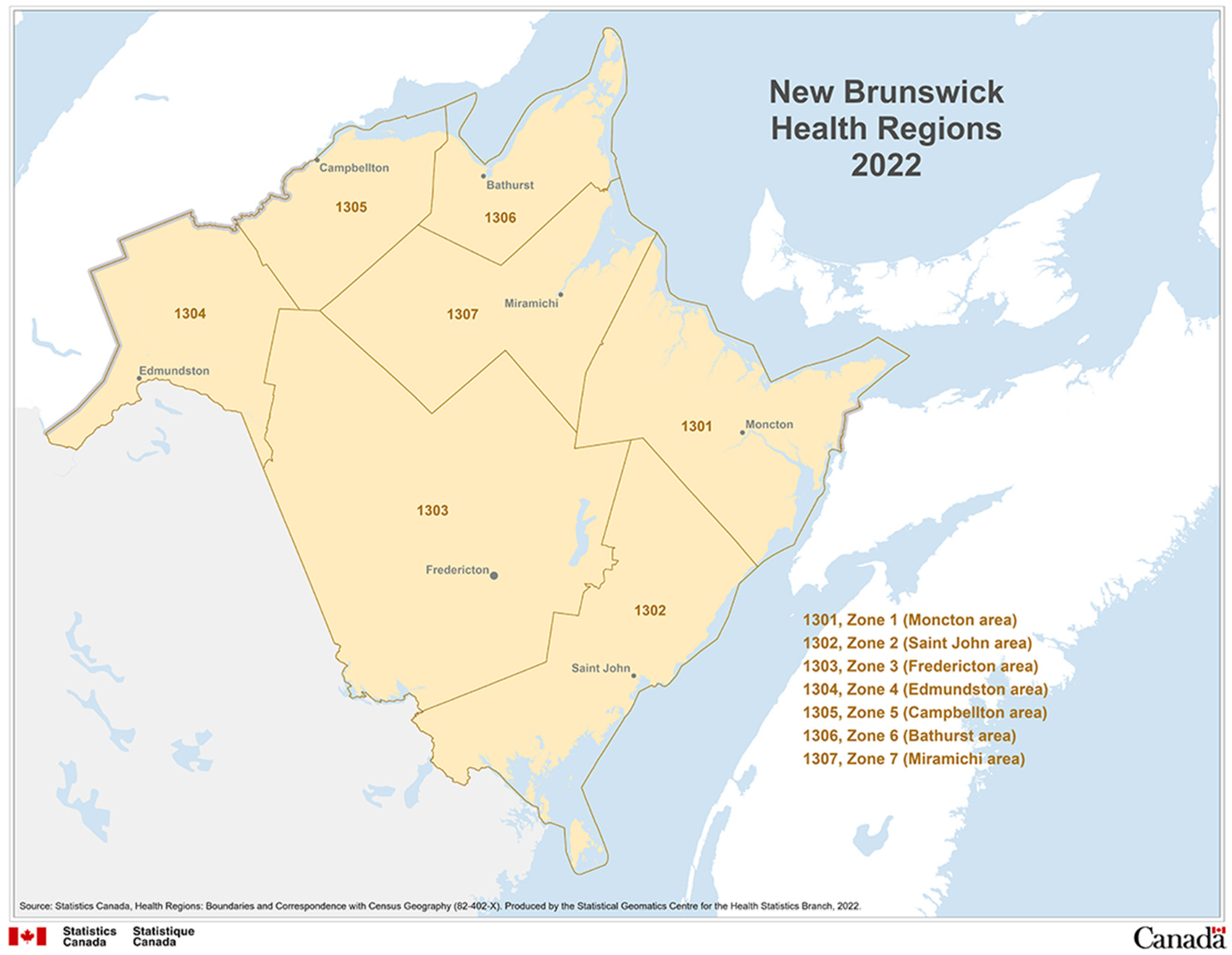
| Number of isolates per healthcare zone | |||
| Zone 1 (Moncton area) | 40 | 37 | 77 (46.4) |
| Zone 2 (Saint John area) | 10 | 20 | 30 (18.1) |
| Zone 3 (Fredericton area) | 14 | 20 | 34 (30.5) |
| Zone 4 (Edmunston area) | 3 | 3 | 6 (3.6) |
| Zone 5 (Campbellton area) | 5 | 3 | 8 (4.8) |
| Zone 6 (Bathurst area) | 6 | 2 | 8 (4.8) |
| Zone 7 (Miramichi) | 2 | 1 | 3 (1.8) |
| Anatomical isolation site: | |||
| Respiratory | 55 | 63 | 118 (71.1) |
| Lymphatic | 9 | 13 | 22 (13.3) |
| Musculoskeletal | 9 | 2 | 11 (6.6) |
| Genitourinary | 2 | 2 | 4 (2.4) |
| Gastrointestinal | 2 | 2 | 4 (2.4) |
| Central nervous system | 0 | 1 | 1 (0.6) |
| Respiratory, Lymphatic | 1 | 0 | 1 (0.6) |
| Respiratory, Gastrointestinal | 1 | 0 | 1 (0.6) |
| Respiratory, Musculoskeletal | 0 | 1 | 1 (0.6) |
| Respiratory, Musculoskeletal, Genitourinary | 0 | 1 | 1 (0.6) |
| Respiratory, Genitourinary, Central nervous system | 1 | 0 | 1 (0.6) |
| Locus | Number of Isolates with Identified MIRU-VNTR Alleles | Allelic Diversity | Designation | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 16 | 19 | NA | |||
| 4052 | 11 | 4 | 10 | 37 | 24 | 38 | 26 | 6 | 3 | 2 | 1 | 4 | 0.8394 | High | ||||
| 2163b | 2 | 31 | 36 | 35 | 26 | 16 | 2 | 6 | 5 | 7 | 0.8279 | High | ||||||
| 1955 | 13 | 31 | 50 | 38 | 9 | 4 | 4 | 11 | 1 | 1 | 4 | 0.8026 | High | |||||
| 802 | 27 | 33 | 65 | 26 | 5 | 8 | 2 | 0.7574 | High | |||||||||
| 424 | 23 | 68 | 30 | 36 | 7 | 1 | 1 | 0.7327 | High | |||||||||
| 2996 | 6 | 23 | 5 | 20 | 80 | 11 | 18 | 3 | 0.7197 | High | ||||||||
| 960 | 2 | 5 | 71 | 55 | 19 | 9 | 3 | 1 | 1 | 0.6902 | High | |||||||
| 3690 | 3 | 43 | 77 | 19 | 8 | 5 | 1 | 1 | 1 | 3 | 1 | 4 | 0.6899 | High | ||||
| 3192 | 24 | 81 | 24 | 34 | 3 | 0.6819 | High | |||||||||||
| 2165 | 2 | 30 | 51 | 72 | 5 | 2 | 1 | 3 | 0.676 | High | ||||||||
| 4156 | 2 | 35 | 83 | 33 | 12 | 1 | 0.6605 | High | ||||||||||
| 2401 | 21 | 81 | 1 | 62 | 1 | 0.61 | High | |||||||||||
| 2531 | 7 | 10 | 98 | 47 | 2 | 2 | 0.569 | Moderate | ||||||||||
| 577 | 17 | 36 | 103 | 8 | 1 | 1 | 0.5531 | Moderate | ||||||||||
| 1644 | 12 | 26 | 107 | 20 | 1 | 0.5435 | Moderate | |||||||||||
| 4348 | 5 | 107 | 52 | 1 | 1 | 0.4821 | Moderate | |||||||||||
| 580 | 1 | 126 | 16 | 3 | 19 | 1 | 0.4035 | Moderate | ||||||||||
| 2461 | 16 | 128 | 2 | 3 | 4 | 11 | 1 | 1 | 0.393 | Moderate | ||||||||
| 2347 | 1 | 8 | 31 | 126 | 0.389 | Moderate | ||||||||||||
| 2687 | 138 | 27 | 1 | 0.2841 | Poor | |||||||||||||
| 2059 | 23 | 143 | 0.2402 | Poor | ||||||||||||||
| 3171 | 2 | 6 | 149 | 2 | 7 | 0.1921 | Poor | |||||||||||
| 3007 | 4 | 6 | 154 | 2 | 0.1382 | Poor | ||||||||||||
| 154 | 5 | 157 | 4 | 0.1046 | Poor | |||||||||||||
| Cluster | Patient ID | Isolation Year | Age | Sex | Anatomical Site | Region of Isolation |
|---|---|---|---|---|---|---|
| A | 61 | 2003 | 66 | F | Respiratory | 3 |
| 152 | 2016 | 67 | F | Respiratory | 6 | |
| B | 78 | 2017 | 25 | F | Respiratory | 1 |
| 80 | 2017 | 65 | F | Respiratory | 1 | |
| 149 | 2019 | 64 | M | Respiratory | 1 | |
| 101 | 2023 | 27 | F | Respiratory | 2 | |
| C | 84 | 2016 | 74 | M | Respiratory | 2 |
| 66 | 2016 | 76 | M | Respiratory | 7 | |
| D | 98 | 2022 | 24 | F | Lymphatic | 4 |
| 131 | 2024 | 46 | F | Respiratory | 1 | |
| E | 161 | 2020 | 26 | F | Respiratory | 5 |
| 139 | 2021 | 50 | F | Respiratory | 5 | |
| F | 93 | 2021 | 36 | F | Lymphatic | 1 |
| 109 | 2022 | 25 | M | Musculoskeletal | 1 | |
| 126 | 2023 | 19 | F | Respiratory | 1 | |
| G | 39 | 2010 | 88 | M | Respiratory | 3 |
| 40 | 2010 | 50 | M | Respiratory | 1 | |
| 150 | 2020 | 69 | M | Respiratory | 1 | |
| H | 116 | 2022 | 26 | F | Respiratory | 1 |
| 79 | 2024 | 30 | F | Respiratory | 1 | |
| I | 1 | 2004 | 77 | F | Respiratory | 4 |
| 49 | 2013 | 85 | F | Musculoskeletal | 1 | |
| J | 108 | 2009 | 70 | F | Respiratory | 2 |
| 4 | 2009 | 61 | F | Respiratory | 2 | |
| 33 | 2010 | 63 | F | Respiratory | 1 | |
| K | 48 | 2022 | 51 | M | Respiratory | 2 |
| 134 | 2024 | 56 | M | Respiratory | 1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Shamputa, I.C.; Gaudet, D.J.; McKinney, J.; Barker, K.; Soualhine, H.; Yoshida, C.; Sharma, M.K.; Webster, D. Genetic Diversity and Molecular Epidemiology of Mycobacterium tuberculosis Complex Clinical Isolates in New Brunswick, Canada—A Retrospective Chart Review. Pathogens 2026, 15, 115. https://doi.org/10.3390/pathogens15010115
Shamputa IC, Gaudet DJ, McKinney J, Barker K, Soualhine H, Yoshida C, Sharma MK, Webster D. Genetic Diversity and Molecular Epidemiology of Mycobacterium tuberculosis Complex Clinical Isolates in New Brunswick, Canada—A Retrospective Chart Review. Pathogens. 2026; 15(1):115. https://doi.org/10.3390/pathogens15010115
Chicago/Turabian StyleShamputa, Isdore Chola, Derek J. Gaudet, Jason McKinney, Kim Barker, Hafid Soualhine, Catherine Yoshida, Meenu Kaushal Sharma, and Duncan Webster. 2026. "Genetic Diversity and Molecular Epidemiology of Mycobacterium tuberculosis Complex Clinical Isolates in New Brunswick, Canada—A Retrospective Chart Review" Pathogens 15, no. 1: 115. https://doi.org/10.3390/pathogens15010115
APA StyleShamputa, I. C., Gaudet, D. J., McKinney, J., Barker, K., Soualhine, H., Yoshida, C., Sharma, M. K., & Webster, D. (2026). Genetic Diversity and Molecular Epidemiology of Mycobacterium tuberculosis Complex Clinical Isolates in New Brunswick, Canada—A Retrospective Chart Review. Pathogens, 15(1), 115. https://doi.org/10.3390/pathogens15010115






