Potential Biocontrol Activities of Populus Endophytes against Several Plant Pathogens Using Different Inhibitory Mechanisms
Abstract
1. Introduction
2. Results
2.1. Antimicrobial Activities of Strain WPB
2.2. Isolation and Screening of New Anti-Fungal Endophytes
2.3. Analyses for Potential Antimicrobial Mechanisms
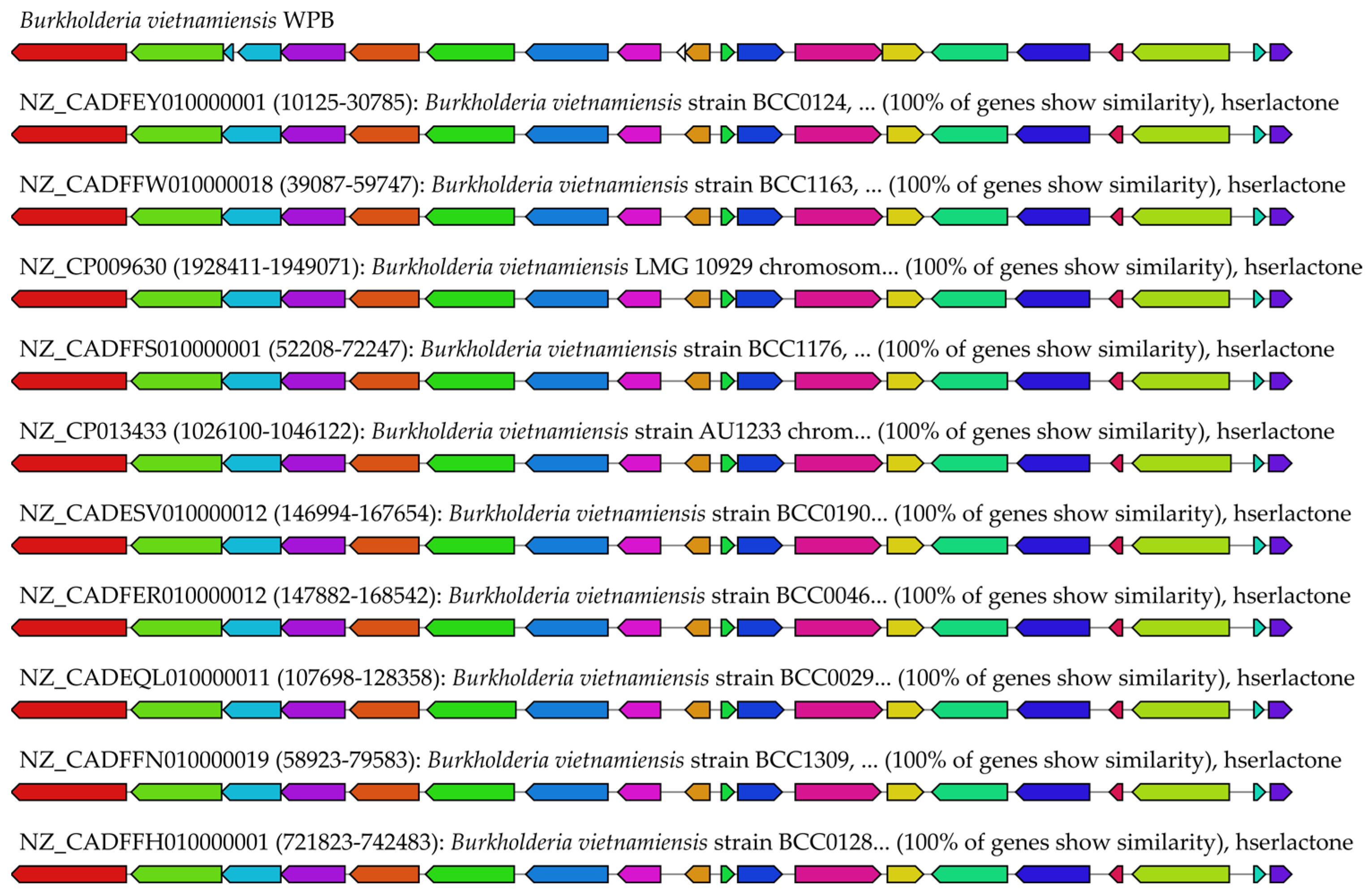
2.3.1. In silico Analysis of Antimicrobial Biosynthetic Gene Clusters in Strain WPB
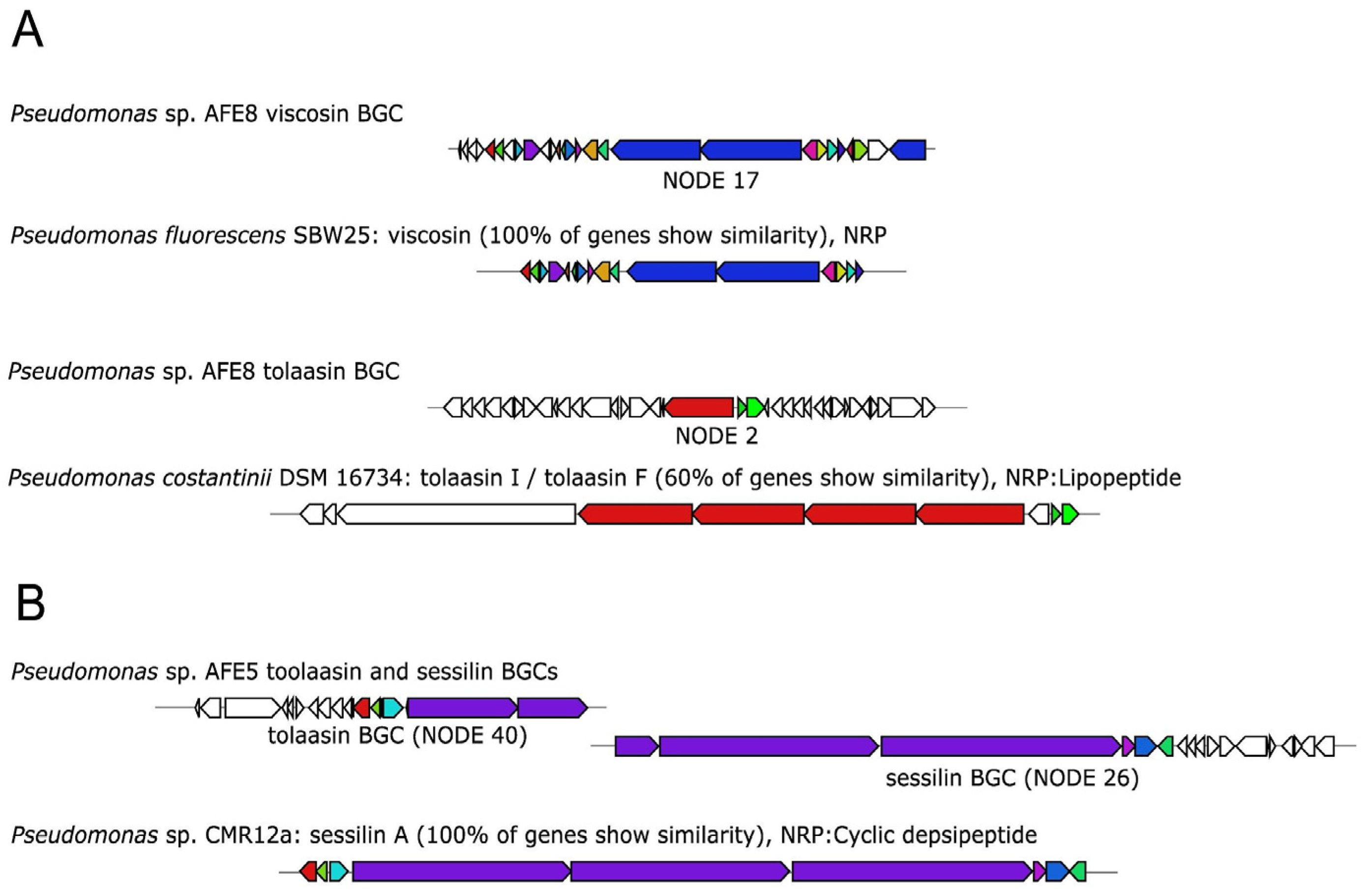
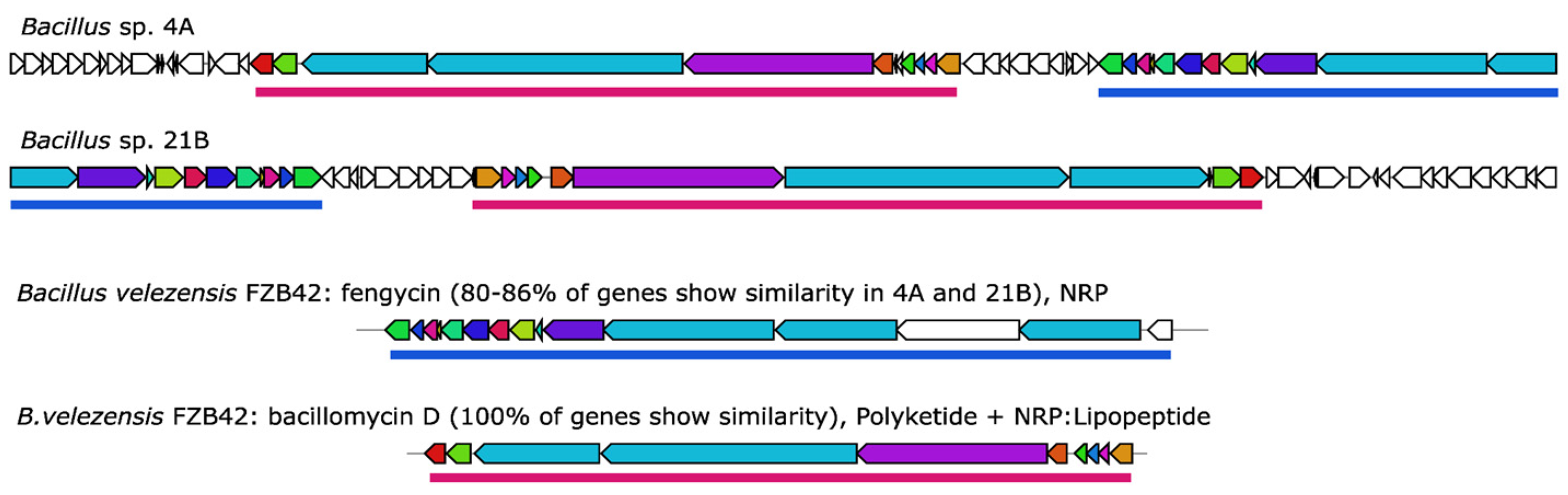
2.3.2. In Silico Analysis of Antimicrobial Biosynthetic Gene Clusters in the AFE Strains
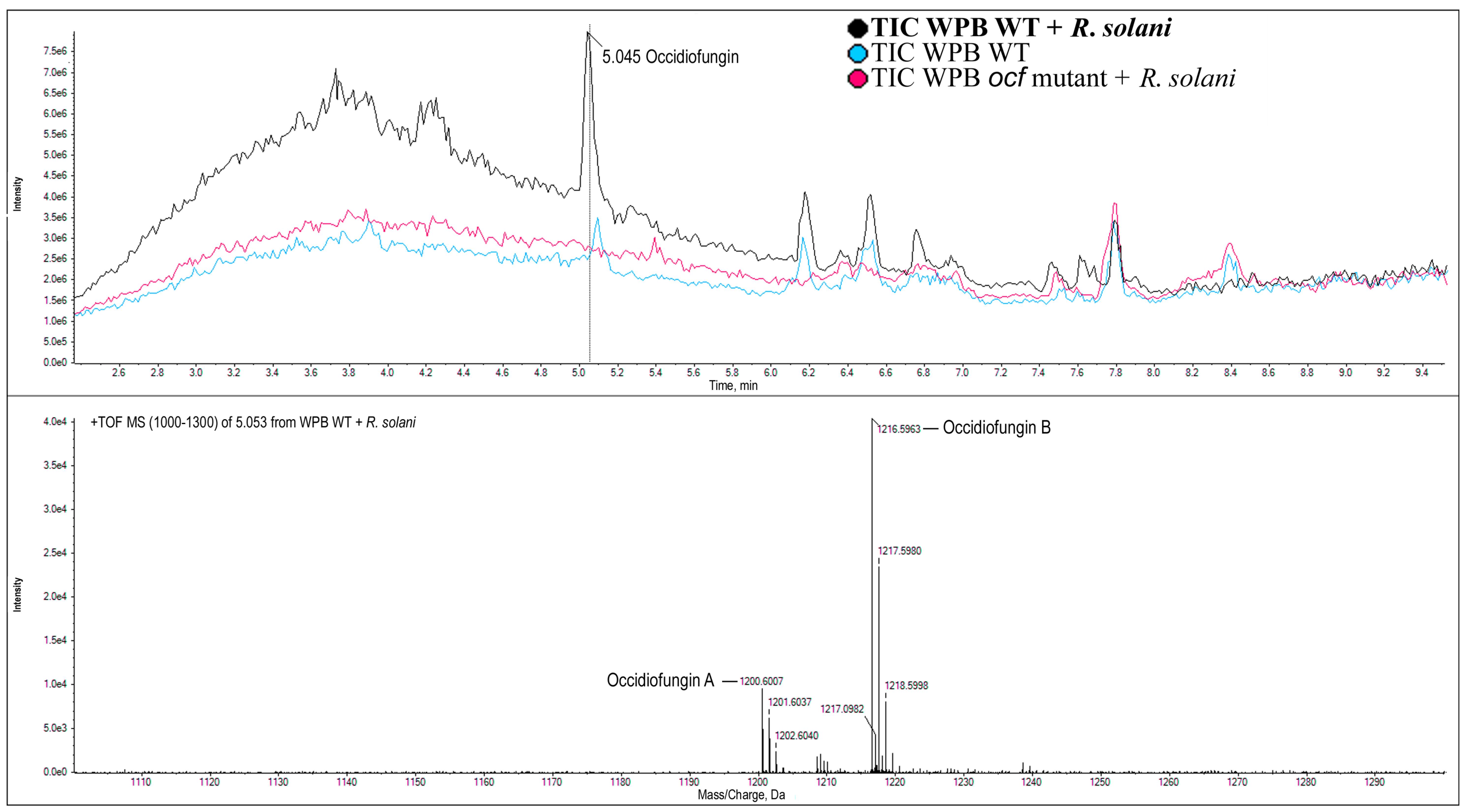
2.3.3. Occidiofungin Mutant of WPB
2.3.4. Confirmation of the Lack of Occidiofungin from AFE Strains by LC-MS-MS
2.4. Symbiotic Traits of the AFE Strains
2.5. Phylogenomic Analysis of WPB and AFE Strains for Suitability for Use in Agriculture
3. Discussion
4. Materials and Methods
4.1. Pathogen Strains
4.2. Endophyte Extraction from Plant Tissue
4.3. High-Throughput Antifungal Screen and Inhibition Assays
4.4. Endophyte Identification
4.5. Genomic Sequencing and Analysis
4.6. In Silico Analysis of Antimicrobial Biosynthetic Gene Clusters in WPB and AFE Strains
4.7. Occidiofungin Mutant Construction
4.8. Chemical Extraction and Analysis of Secreted Antimicrobials
4.9. Assays for Symbiotic Traits
5. Conclusions
6. Patents
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A

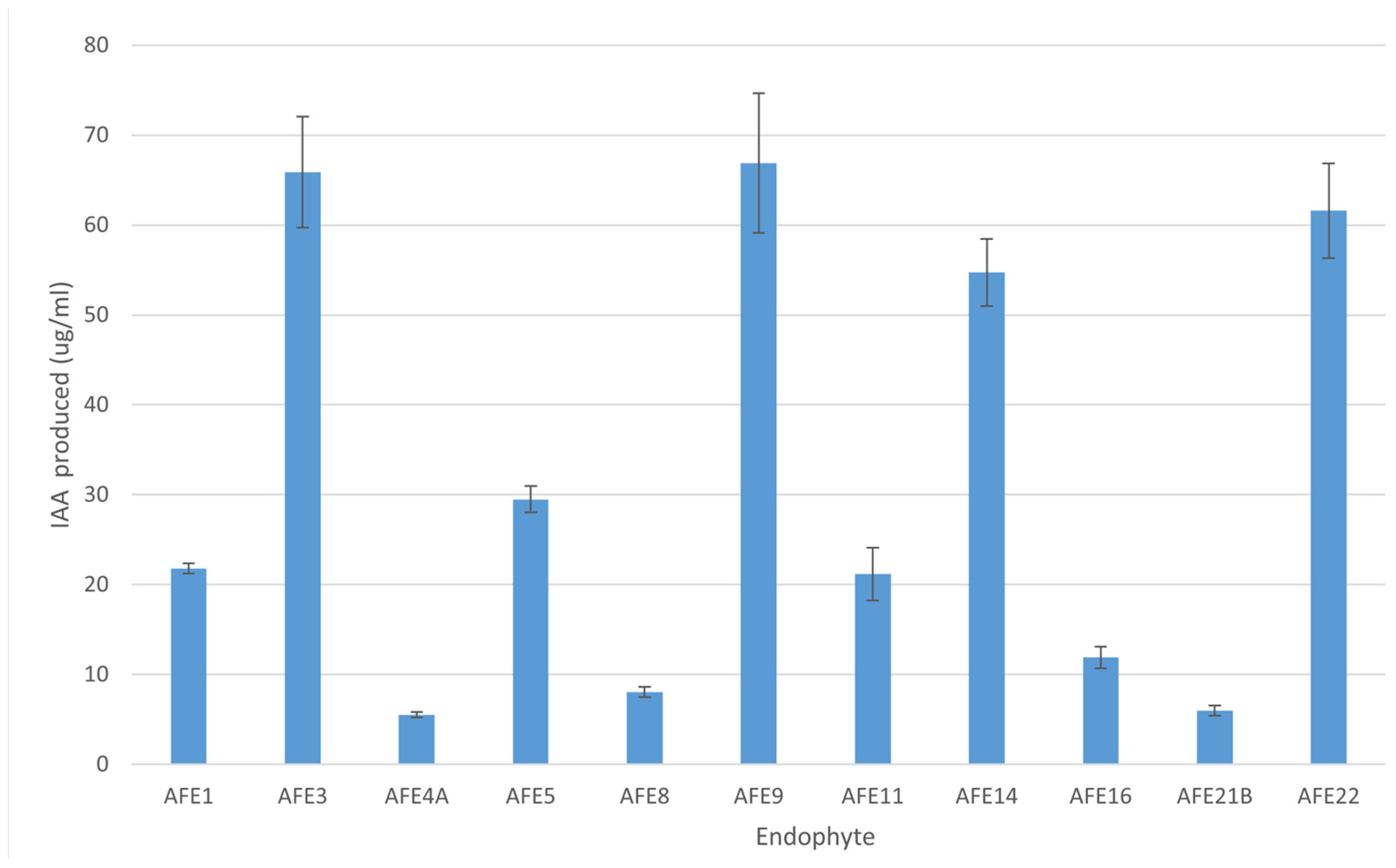
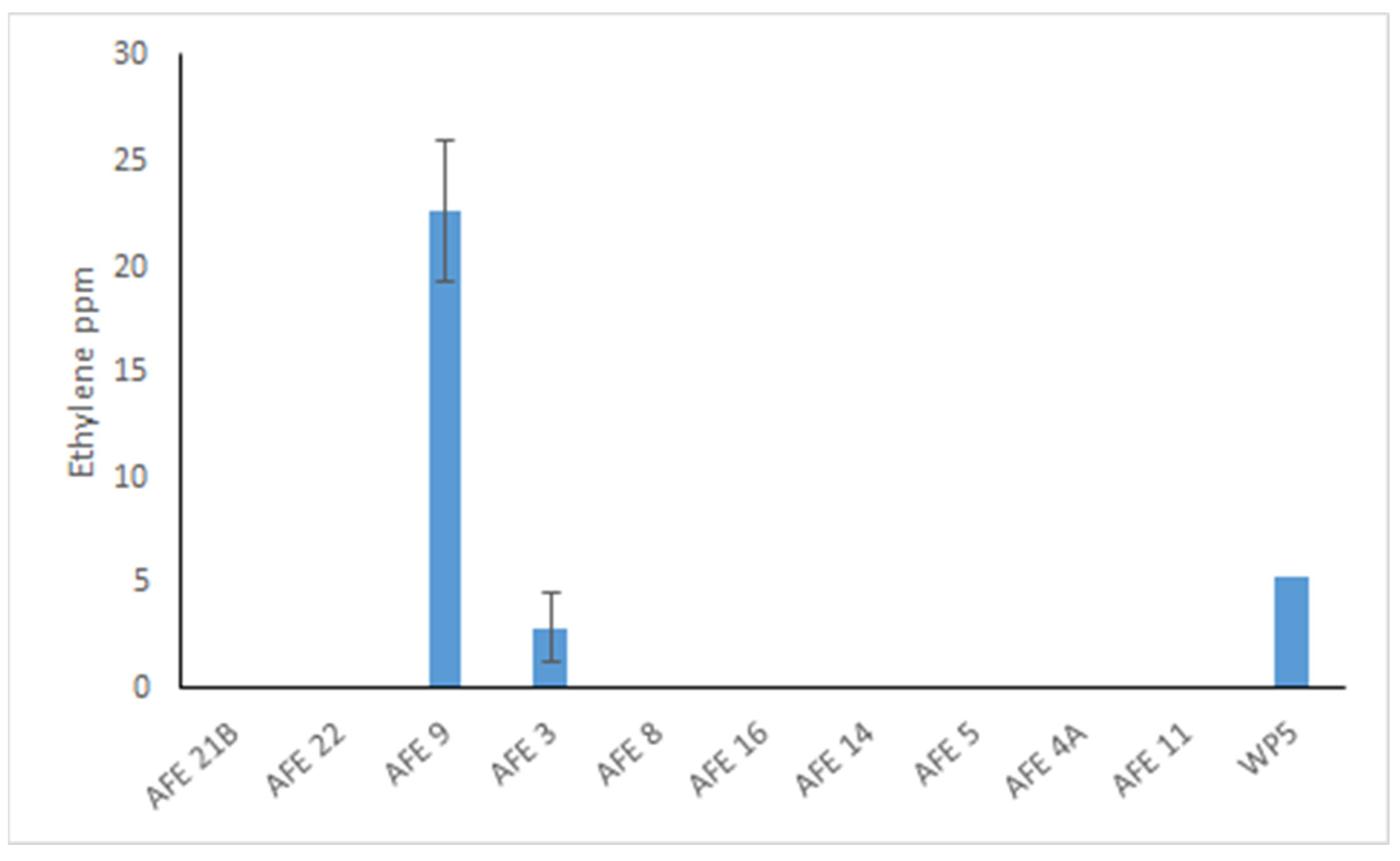
| Strain | Genus | Genome Size (Mbp) | Scaffolds | N50 | GC % | ORF | CDS |
|---|---|---|---|---|---|---|---|
| AFE1 | Erwinia | 5.30 | 19 | 708,447 | 54.9 | 5055 | 4982 |
| AFE3 | Rahnella | 5.46 | 22 | 420,146 | 53.5 | 5177 | 5099 |
| AFE4A | Bacillus | 3.96 | 25 | 422,228 | 46.4 | 4163 | 4070 |
| AFE5 | Pseudomonas | 5.91 | 68 | 154,070 | 59.5 | 5514 | 5453 |
| AFE8 | Pseudomonas | 6.14 | 30 | 506,261 | 60.5 | 5740 | 5678 |
| AFE9 | Rahnella | 5.76 | 52 | 204,001 | 52.2 | 5668 | 5589 |
| AFE16 | Pseudomonas | 5.95 | 57 | 181,015 | 60.5 | 5690 | 5627 |
| AFE21B | Bacillus | 4.12 | 22 | 585,429 | 46.2 | 4351 | 4269 |
| AFE22 | Rhanella | 5.52 | 47 | 200,569 | 52.1 | 5339 | 5262 |
| AFE Strain ID | Classification | Closest GTDB-tk Type-Strain | Is Type Strain | Closest Type-Strain NCBI Accession | Species Cluster Link |
|---|---|---|---|---|---|
| AFE1 | Erwinia billingiae | Erwinia billingiae Eb661 | YES | GCF_000196615.1 | https://gtdb.ecogenomic.org/species?id=Erwinia%20billingiae |
| AFE16 | Pseudomonas_E graminis_D | Pseudomonas graminis WP8 | NO | GCF_004364335.1 | https://gtdb.ecogenomic.org/species?id=Pseudomonas_E%20graminis_D |
| AFE21B | Bacillus velezensis | Bacillus velezensis NRRL B-41580 | YES | GCF_001461825.1 | https://gtdb.ecogenomic.org/species?id=Bacillus%20velezensis |
| AFE22 | Rahnella variigena | Rahnella variigena CIP 105588 | YES | GCF_003610915.1 | https://gtdb.ecogenomic.org/species?id=Rahnella%20variigena |
| AFE3 | Rahnella victoriana | Rahnella victoriana DSM 27397 | YES | GCF_004330295.1 | https://gtdb.ecogenomic.org/species?id=Rahnella%20victoriana |
| AFE4A | Bacillus velezensis | Bacillus velezensis NRRL B-41580 | YES | GCF_001461825.1 | https://gtdb.ecogenomic.org/species?id=Bacillus%20velezensis |
| AFE5 | Pseudomonas kitaguniensis | Pseudomonas kitaguniensis MAFF 212408 | YES | GCF_009296165.1 | https://gtdb.ecogenomic.org/species?id=Pseudomonas_E%20kitaguniensis |
| AFE8 | Pseudomonas lurida | Pseudomonas lurida LMG 21995 | YES | GCF_002563895.1 | https://gtdb.ecogenomic.org/species?id=Pseudomonas_E%20lurida |
| AFE9 | Rahnella aceris | Rahnella aceris SAP-19 | YES | GCF_011684115.1 | https://gtdb.ecogenomic.org/species?id=Rahnella%20aceris |
References
- Fisher, M.C.; Henk, D.A.; Briggs, C.J.; Brownstein, J.S.; Madoff, L.C.; McCraw, S.L.; Gurr, S.J. Emerging fungal threats to animal, plant and ecosystem health. Nature 2012, 484, 186–194. [Google Scholar] [CrossRef] [PubMed]
- Moore, D.; Robson, G.D.; Trinci, A.P.J. 21st Century Guidebook to Fungi; Cambridge University Press: Cambridge, UK, 2020. [Google Scholar]
- Hane, J.K.; Anderson, J.P.; Williams, A.H.; Sperschneider, J.; Singh, K.B. Genome sequencing and comparative genomics of the broad host-range pathogen Rhizoctonia solani AG8. PLoS Genet. 2014, 10, e1004281. [Google Scholar] [CrossRef] [PubMed]
- Paulitz, T.C. Low input no-till cereal production in the Pacific Northwest of the U.S.: The challenges of root diseases. Eur. J. Plant Pathol. 2006, 115, 271–281. [Google Scholar] [CrossRef]
- Syed, A.; Rahman, S.F.; Singh, E.; Pieterse, C.M.J.; Schenk, P.M. Emerging microbial biocontrol strategies for plant pathogens. Plant Sci. 2018, 267, 102–111. [Google Scholar] [CrossRef]
- Mavrodi, O.V.; Walter, N.; Elateek, S.; Taylor, C.G.; Okubara, P.A. Suppression of Rhizoctonia and Pythium root rot of wheat by new strains of Pseudomonas. Biol. Control 2012, 62, 93–102. [Google Scholar] [CrossRef]
- Palmieri, D.; Ianiri, G.; Del Grosso, C.; Barone, G.; De Curtis, F.; Castoria, R.; Lima, G. Advances and perspectives in the use of biocontrol agents against fungal plant diseases. Horticulturae 2022, 8, 577. [Google Scholar] [CrossRef]
- Velivelli, S.L.; De Vos, P.; Kromann, P.; Declerck, S.; Prestwich, B.D. Biological control agents: From field to market, problems and challenges. Trends Biotechnol. 2014, 32, 493–496. [Google Scholar] [CrossRef]
- Berg, G. Plant-microbe interactions promoting plant growth and health: Perspectives for controlled use of microorganisms in agriculture. Appl. Microbiol. Biotechnol 2009, 84, 11–18. [Google Scholar] [CrossRef]
- Thambugala, K.M.; Daranagama, D.A.; Phillips, A.J.L.; Kannangara, S.D.; Promputtha, I. Fungi vs. Fungi in Biocontrol: An Overview of Fungal Antagonists Applied Against Fungal Plant Pathogens. Front. Cell. Infect. Microbiol. 2020, 10, 604923. [Google Scholar] [CrossRef]
- Okubara, P.A.; Kornoely, J.P.; Landa, B.B. Rhizosphere colonization of hexaploid wheat by Pseudomonas fluorescens strains Q8r1-96 and Q2-87 is cultivar-variable and associated with changes in gross root morphology. Biol. Control 2004, 30, 392–403. [Google Scholar] [CrossRef]
- Nevada, S.; Lupien, S.; Watson, B.; Okubara, P.A. Growth inhibition of Botrytis cinerea by native vineyard yeasts from Puget Sound, Washington State, USA. J. Biol. Nat. 2021, 13, 42–53. [Google Scholar]
- Bonanomi, G.; Lorito, M.; Vinale, F.; Woo, S.L. Organic Amendments, Beneficial Microbes, and Soil Microbiota: Toward a Unified Framework for Disease Suppression. Annu. Rev. Phytopathol. 2018, 56, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Kandel, S.L.; Firrincieli, A.; Joubert, P.M.; Okubara, P.A.; Leston, N.D.; McGeorge, K.M.; Mugnozza, G.S.; Harfouche, A.; Kim, S.-H.; Doty, S.L. An in vitro study of bio-control and plant growth promotion potential of Salicaceae endophytes. Front. Microbiol. 2017, 8, 386. [Google Scholar] [CrossRef] [PubMed]
- Le, C.K.; Gurr, S.J.; Hirsch, P.R.; Mauchline, T.H. Exploitation of endophytes for sustainable agricultural intensification. Mol. Plant Pathol. 2017, 18, 469–473. [Google Scholar]
- Doty, S.L. Functional importance of the plant microbiome: Implications for agriculture, forestry and bioenergy. In Functional Importance of the Plant Microbiome: Implications for Agriculture, Forestry and Bioenergy; Springer: Cham, Switzerland, 2017; pp. 1–6. [Google Scholar]
- Santoyo, G.; Moreno-Hagelsieb, G.; del Carmen Orozco-Mosqueda, M.; Glick, B.R. Plant growth-promoting bacterial endophytes. Microbiol. Res. 2016, 183, 92–99. [Google Scholar] [CrossRef]
- Compant, S.; Clement, C.; Sessitsch, A. Plant growth-promoting bacteria in the rhizo- and endosphere of plants: Their role, colonization, mechanisms involved and prospects for utilization. Soil Biol. Biochem. 2010, 42, 669–678. [Google Scholar] [CrossRef]
- Kandel, S.L.; Joubert, P.M.; Doty, S.L. Bacterial endophyte colonization and distribution in plants. Microorganisms 2017, 5, 77. [Google Scholar] [CrossRef]
- Chiarini, L.; Bevivino, A.; Dalmastri, C.; Tabacchioni, S.; Visca, P. Burkholderia cepacia complex species: Health hazards and biotechnological potential. Trends Microbiol. 2006, 14, 277–286. [Google Scholar] [CrossRef]
- Doty, S.L.; Oakley, B.; Xin, G.; Kang, J.W.; Singleton, G.; Khan, Z.; Vajzovic, A.; Staley, J.T. Diazotrophic endophytes of native black cottonwood and willow. Symbiosis 2009, 47, 23–33. [Google Scholar] [CrossRef]
- Xin, G.; Zhang, G.; Kang, J.W.; Staley, J.T.; Doty, S.L. A diazotrophic, indole-3-acetic acid-producing endophyte from wild cottonwood. Biol. Fertil. Soils 2009, 45, 669–674. [Google Scholar] [CrossRef]
- Mullins, A.J.; Murray, J.A.; Bull, M.J.; Jenner, M.; Jones, C.; Webster, G.; Green, A.; Neill, D.R.; Connor, T.R.; Parkhill, J.; et al. Genome mining identifies cepacin as a plant-protective metabolite of the biopesticidal bacterium Burkholderia ambifaria. Nat. Microbiol. 2019, 4, 996–1005. [Google Scholar] [CrossRef] [PubMed]
- Lu, S.E.; Novak, J.; Austin, F.W.; Gu, G.; Ellis, D.; Kirk, M.; Wilson-Stanford, S.; Tonelli, M.; Smith, L. Occidiofungin, a unique antifungal glycopeptide produced by a strain of Burkholderia contaminans. Biochemistry 2009, 48, 8312–8321. [Google Scholar] [CrossRef]
- Wang, X.; Liu, A.; Guerrero, A.; Liu, J.; Yu, X.; Deng, P.; Ma, L.; Baird, S.; Smith, L.; Li, X.; et al. Occidiofungin is an important component responsible for the antifungal activity of Burkholderia pyrrocinia strain Lyc2. J. Appl. Microbiol. 2016, 120, 607–618. [Google Scholar] [CrossRef] [PubMed]
- Deng, P.; Foxfire, A.; Xu, J.; Baird, S.M.; Jia, J.; Delgado, K.H.; Shin, R.; Smith, L.; Lu, S.-E. The Siderophore Product Ornibactin Is Required for the Bactericidal Activity of Burkholderia contaminans MS14. Appl. Environ. Microbiol. 2017, 83, e00051-17. [Google Scholar] [CrossRef] [PubMed]
- Thomas, M.S. Iron acquisition mechanisms of the Burkholderia cepacia complex. Biometals 2007, 20, 431–452. [Google Scholar] [CrossRef]
- Butt, A.T.; Thomas, M.S. Iron Acquisition Mechanisms and Their Role in the Virulence of Burkholderia Species. Front. Cell. Infect. Microbiol. 2017, 7, 460. [Google Scholar] [CrossRef]
- Hoskisson, P.A.; Seipke, R.F. Cryptic or Silent? The Known Unknowns, Unknown Knowns, and Unknown Unknowns of Secondary Metabolism. MBio 2020, 11, e02642-20. [Google Scholar] [CrossRef]
- Hermenau, R.; Kugel, S.; Komor, A.J.; Hertweck, C. Helper bacteria halt and disarm mushroom pathogens by linearizing structurally diverse cyclolipopeptides. Proc. Natl. Acad. Sci. USA 2020, 117, 23802–23806. [Google Scholar] [CrossRef]
- Ferrarini, E.; De, R.V.; Geudens, N.; Martins, J.C.; Hofte, M. Altering in vivo membrane sterol composition affects the activity of the cyclic lipopeptides tolaasin and sessilin against Pythium. Biochim. Biophys. Acta Biomembr. 2022, 1864, 184008. [Google Scholar] [CrossRef]
- Xiao, J.; Guo, X.; Qiao, X.; Zhang, X.; Chen, X.; Zhang, D. Activity of Fengycin and Iturin A Isolated From Bacillus subtilis Z-14 on Gaeumannomyces graminis Var. tritici and Soil Microbial Diversity. Front. Microbiol. 2021, 12, 682437. [Google Scholar]
- Karpinski, T.M. Marine Macrolides with Antibacterial and/or Antifungal Activity. Mar. Drugs 2019, 17, 241. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Sun, C. Fengycins, Cyclic Lipopeptides from Marine Bacillus subtilis Strains, Kill the Plant-Pathogenic Fungus Magnaporthe grisea by Inducing Reactive Oxygen Species Production and Chromatin Condensation. Appl. Environ. Microbiol. 2018, 84, e00445-18. [Google Scholar] [CrossRef] [PubMed]
- Gu, G.; Smith, L.; Wang, N.; Wang, H.; Lu, S.E. Biosynthesis of an antifungal oligopeptide in Burkholderia contaminans strain MS14. Biochem. Biophys. Res. Commun. 2009, 380, 328–332. [Google Scholar] [CrossRef] [PubMed]
- Bashan, Y.; Kamnev, A.A.; de-Bashan, L.E. Tricalcium phosphate is inappropriate as a universal selection factor for isolating and testing phosphate-solubilizing bacteria that enhance plant growth: A proposal for an alternative procedure. Biol. Fertil. Soils 2013, 49, 465–479. [Google Scholar] [CrossRef]
- Varga, T.; Hixson, K.K.; Ahkami, A.H.; Sher, A.W.; Barnes, M.E.; Chu, R.K.; Battu, A.K.; Nicora, C.D.; Winkler, T.E.; Reno, L.R.; et al. Endophyte-Promoted Phosphorus Solubilization in Populus. Front. Plant Sci. 2020, 11, 567918. [Google Scholar] [CrossRef]
- Brady, C.; Hunter, G.; Kirk, S.; Arnold, D.; Denman, S. Rahnella victoriana sp. nov., Rahnella bruchi sp. nov., Rahnella woolbedingensis sp. nov., classification of Rahnella genomospecies 2 and 3 as Rahnella variigena sp. nov. and Rahnella inusitata sp. nov., respectively and emended description of the genus Rahnella. Syst. Appl. Microbiol. 2014, 37, 545–552. [Google Scholar]
- Lee, S.D.; Jeon, D.; Kim, I.S.; Choe, H.; Kim, J.S. Rahnella aceris sp. nov., isolated from sap drawn from Acer pictum. Arch. Microbiol. 2020, 202, 2411–2417. [Google Scholar] [CrossRef]
- Kube, M.; Migdoll, A.M.; Gehring, I.; Heitmann, K.; Mayer, Y.; Kuhl, H.; Knaust, F.; Geider, K.; Reinhardt, R. Genome comparison of the epiphytic bacteria Erwinia billingiae and E. tasmaniensis with the pear pathogen E. pyrifoliae. BMC Genom. 2010, 11, 393. [Google Scholar] [CrossRef]
- Llop, P. Genetic islands in pome fruit pathogenic and non-pathogenic Erwinia species and related plasmids. Front. Microbiol. 2015, 6, 874. [Google Scholar] [CrossRef]
- United States Environmental Protection Agency. EPA Proposes Registration of Products Containing Bacillus velezensis Strain RTI301 and Bacillus subtilis Strain RTI477, New Microbial Active Ingredients; 6-25-0021; United States Environmental Protection Agency: Washington, DC, USA, 2021. Available online: https://www.epa.gov/pesticides/epa-proposes-registration-products-containing-bacillus-velezensis-strain-rti301-and (accessed on 20 September 2022).
- European Food Safety Authority. Safety and Efficacy of a Feed Additive Consisting of Bacillus velezensis ATCC PTA-6737 (Bacillus velezensis PB6) for Turkeys for Fattening, Turkeys Reared for Breeding, Laying Hens, Minor Poultry Species for Laying, Piglets (Weaned), Weaned Minor Porcine Species and Sows (Kemin Europe, N.V.); 4-13-0022; European Food Safety Authority: Washington, DC, USA, 2021; Available online: https://www.efsa.europa.eu/en/efsajournal/pub/7244 (accessed on 20 September 2022).
- Scales, B.S.; Erb-Downward, J.R.; Huffnagle, I.M.; LiPuma, J.J.; Huffnagle, G.B. Comparative genomics of Pseudomonas fluorescens subclade III strains from human lungs. BMC Genom. 2015, 16, 1032. [Google Scholar] [CrossRef]
- Scales, B.S.; Dickson, R.P.; LiPuma, J.J.; Huffnagle, G.B. Microbiology, genomics, and clinical significance of the Pseudomonas fluorescens species complex, an unappreciated colonizer of humans. Clin. Microbiol. Rev. 2014, 27, 927–948. [Google Scholar] [CrossRef] [PubMed]
- Dickson, R.P.; Erb-Downward, R.; Freeman, C.M.; Walker, N.; Scales, B.S.; Beck, J.M.; Martinez, F.J.; Curtis, J.L.; Lama, V.N.; Huffnagle, G.B. Changes in the lung microbiome following lung transplantation include the emergence of two distinct Pseudomonas species with distinct clinical associations. PLoS ONE 2014, 9, e97214. [Google Scholar] [CrossRef]
- Sessitsch, A.; Coenye, T.; Sturz, A.V.; Vandamme, P.; Barka, E.A.; Salles, J.F.; Van Elsas, J.D.; Faure, D.; Reiter, B.; Glick, B.R.; et al. Burkholderia phytofirmans sp. nov., a novel plant-associated bacterium with plant-beneficial properties. Int. J. Syst. Evol. Microbiol. 2005, 55, 1187–1192. [Google Scholar] [CrossRef] [PubMed]
- Ait, B.E.; Nowak, J.; Clement, C. Enhancement of chilling resistance of inoculated grapevine plantlets with a plant growth-promoting rhizobacterium, Burkholderia phytofirmans strain PsJN. Appl. Environ. Microbiol. 2006, 72, 7246–7252. [Google Scholar]
- Andreolli, M.; Lampis, S.; Poli, M.; Gullner, G.; Biro, B.; Vallini, G. Endophytic Burkholderia fungorum DBT1 can improve phytoremediation efficiency of polycyclic aromatic hydrocarbons. Chemosphere 2013, 92, 688–694. [Google Scholar] [CrossRef]
- Elliott, G.N.; Chen, W.M.; Chou, J.H.; Wang, H.C.; Sheu, S.Y.; Perin, L.; Reis, V.M.; Moulin, L.; Simon, M.F.; Bontemps, C.; et al. Burkholderia phymatum is a highly effective nitrogen-fixing symbiont of Mimosa spp. and fixes nitrogen ex planta. New Phytol. 2007, 173, 168–180. [Google Scholar] [CrossRef]
- Govindarajan, M.; Balandreau, J.; Kwon, S.W.; Weon, H.Y.; Lakshminarasimhan, C. Effects of the inoculation of Burkholderia vietnamensis and related endophytic diazotrophic bacteria on grain yield of rice. Microb. Ecol. 2008, 55, 21–37. [Google Scholar] [CrossRef]
- Sawana, A.; Adeolu, M.; Gupta, R.S. Molecular signatures and phylogenomic analysis of the genus Burkholderia: Proposal for division of this genus into the emended genus Burkholderia containing pathogenic organisms and a new genus Paraburkholderia gen. nov. harboring environmental species. Front. Genet. 2014, 5, 429. [Google Scholar] [CrossRef]
- Angus, A.A.; Agapakis, C.M.; Fong, S.; Yerrapragada, S.; Santos, P.E.D.L.; Yang, P.; Song, N.; Kano, S.; Caballero-Mellado, J.; De Faria, S.M.; et al. Plant-associated symbiotic Burkholderia species lack hallmark strategies required in mammalian pathogenesis. PLoS ONE 2014, 9, e83779. [Google Scholar] [CrossRef]
- Dobritsa, A.P.; Samadpour, M. Transfer of eleven species of the genus Burkholderia to the genus Paraburkholderia and proposal of Caballeronia gen. nov. to accommodate twelve species of the genera Burkholderia and Paraburkholderia. Int. J. Syst. Evol. Microbiol. 2016, 66, 2836–2846. [Google Scholar] [CrossRef]
- Firrincieli, A.; Khorasani, M.; Frank, A.C.; Doty, S.L. Influences of climate on phyllosphere endophytic bacterial communities of wild poplar. Frontiers in Plant Science Front. Plant Sci. 2020, 11, 203. [Google Scholar] [CrossRef] [PubMed]
- Speare, L.; Woo, M.; Bultman, K.M.; Mandel, M.J.; Wollenberg, M.S.; Septer, A.N. Host-Like Conditions Are Required for T6SS-Mediated Competition among Vibrio fischeri Light Organ Symbionts. mSphere 2021, 6, e0128820. [Google Scholar] [CrossRef] [PubMed]
- Khan, N.; Maymon, M.; Hirsch, A.M. Combating Fusarium Infection Using Bacillus-Based Antimicrobials. Microorganisms 2017, 5, 75. [Google Scholar] [CrossRef] [PubMed]
- Geudens, N.; Nasir, M.N.; Crowet, J.-M.; Raaijmakers, J.; Fehér, K.; Coenye, T.; Martins, J.C.; Lins, L.; Sinnaeve, D.; Deleu, M. Membrane Interactions of Natural Cyclic Lipodepsipeptides of the Viscosin Group. Biochim. Biophys. Acta Biomembr. 2017, 1859, 331–339. [Google Scholar] [CrossRef]
- Cho, K.H.; Kim, Y.K. Two types of ion channel formation of tolaasin, a Pseudomonas peptide toxin. FEMS Microbiol. Lett. 2003, 221, 221–226. [Google Scholar] [CrossRef]
- Andrić, S.; Meyer, T.; Rigolet, A.; Prigent-Combaret, C.; Höfte, M.; Balleux, G.; Steels, S.; Hoff, G.; De Mot, R.; McCann, A.; et al. Lipopeptide Interplay Mediates Molecular Interactions between Soil Bacilli and Pseudomonads. Microbiol. Spectr. 2021, 9, e0203821. [Google Scholar] [CrossRef]
- Barbosa, J.; Caetano, T.; Mendo, S. Class I and Class II Lanthipeptides Produced by Bacillus spp. J. Nat. Prod. 2015, 78, 2850–2866. [Google Scholar] [CrossRef]
- Mohr, K.I.; Volz, C.; Jansen, R.; Wray, V.; Hoffmann, J.; Bernecker, S.; Wink, J.; Gerth, K.; Stadler, M.; Müller, R. Pinensins: The first antifungal lantibiotics. Angew. Chem. Int. Ed. Engl. 2015, 54, 11254–11258. [Google Scholar] [CrossRef]
- Agirman, B.; Erten, H. Biocontrol ability and action mechanisms of Aureobasidium pullulans GE17 and Meyerozyma guilliermondii KL3 against Penicillium digitatum DSM2750 and Penicillium expansum DSM62841 causing postharvest diseases. Yeast 2020, 37, 437–448. [Google Scholar] [CrossRef]
- Mullins, A.J.; Jones, C.; Bull, M.J.; Webster, G.; Parkhill, J.; Connor, T.R.; Murray, J.A.; Challis, G.L.; Mahenthiralingam, E. Genomic Assemblies of Members of Burkholderia and Related Genera as a Resource for Natural Product Discovery. Microbiol. Resour. Announc. 2020, 9, e00485-20. [Google Scholar] [CrossRef]
- Overbeek, R.; Olson, R.; Pusch, G.D.; Olsen, G.J.; Davis, J.J.; Disz, T.; Edwards, R.A.; Gerdes, S.; Parrello, B.; Shukla, M.; et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res. 2014, 42, D206–D214. [Google Scholar] [CrossRef] [PubMed]
- Aramaki, T.; Blanc-Mathieu, R.; Endo, H.; Ohkubo, K.; Kanehisa, M.; Goto, S.; Ogata, H. KofamKOALA: KEGG Ortholog assignment based on profile HMM and adaptive score threshold. Bioinformatics 2020, 36, 2251–2252. [Google Scholar] [CrossRef] [PubMed]
- Mavromatis, K.; Ivanova, N.N.; Chen, I.M.; Szeto, E.; Markowitz, V.M.; Kyrpides, N.C. The DOE-JGI Standard Operating Procedure for the Annotations of Microbial Genomes. Stand. Genom. Sci. 2009, 1, 63–67. [Google Scholar] [CrossRef] [PubMed]
- Meier-Kolthoff, J.P.; Goker, M. TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat. Commun. 2019, 10, 2182. [Google Scholar] [CrossRef] [PubMed]
- Cosentino, S.; Voldby, L.M.; Moller, A.F.; Lund, O. PathogenFinder--distinguishing friend from foe using bacterial whole genome sequence data. PLoS ONE 2013, 8, e77302. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Aziz, R.K.; Bartels, D.; Best, A.A.; DeJongh, M.; Disz, T.; Edwards, R.A.; Formsma, K.; Gerdes, S.; Glass, E.M.; Kubal, M.; et al. The RAST Server: Rapid annotations using subsystems technology. BMC Genom. 2008, 9, 75. [Google Scholar] [CrossRef]
- Parks, D.H.; Chuvochina, M.; Rinke, C.; Mussig, A.J.; Chaumeil, P.A.; Hugenholtz, P. GTDB: An ongoing census of bacterial and archaeal diversity through a phylogenetically consistent, rank normalized and complete genome-based taxonomy. Nucleic Acids Res. 2022, 50, D785–D794. [Google Scholar] [CrossRef]
- Blin, K.; Shaw, S.; Kloosterman, A.M.; Charlop-Powers, Z.; van Wezel, G.P.; Medema, M.H.; Weber, T. antiSMASH 6.0: Improving cluster detection and comparison capabilities. Nucleic Acids Res. 2021, 49, W29–W35. [Google Scholar] [CrossRef]
- Zúñiga, A.; Poupin, M.J.; Donoso, R.; Ledger, T.; Guiliani, N.; Gutiérrez, R.A.; González, B. Quorum sensing and indole-3-acetic acid degradation play a role in colonization and plant growth promotion of Arabidopsis thaliana by Burkholderia phytofirmans PsJN. Mol. Plant Microbe Interact. 2013, 26, 546–553. [Google Scholar] [CrossRef]
- Rennie, R.J. A single medium for the isolation of acetylene-reducing (dinitrogen-fixing) bacteria from soils. Can. J. Microbiol. 1981, 27, 8–14. [Google Scholar] [CrossRef] [PubMed]
- Doty, S.L.; Sher, A.W.; Fleck, N.D.; Khorasani, M.; Bumgarner, R.E.; Khan, Z.; Ko, A.W.K.; Kim, S.-H.; DeLuca, T.H. Variable nitrogen fixation in wild Populus. PLoS ONE 2016, 11, e0155979. [Google Scholar] [CrossRef] [PubMed]
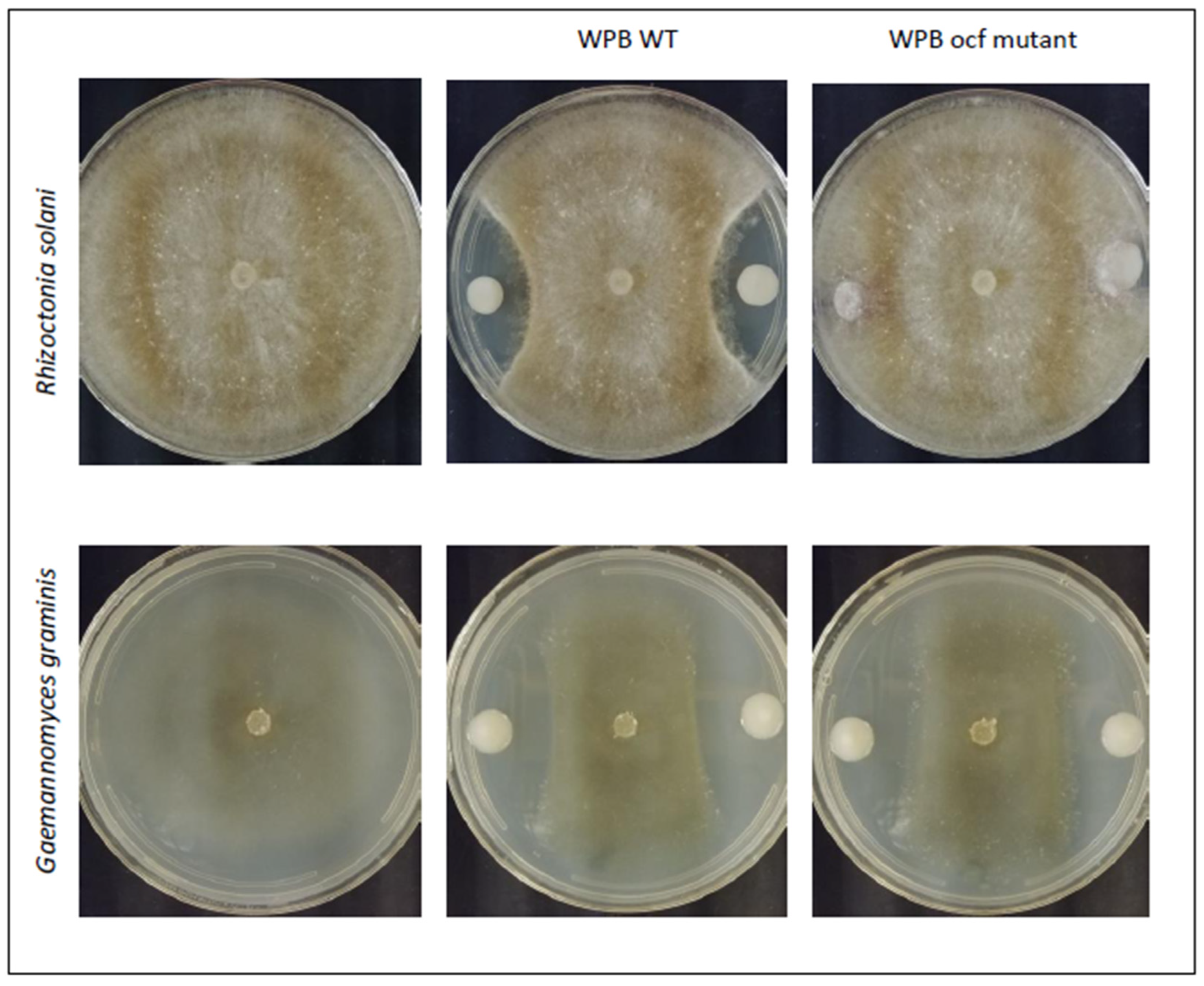

| Strain | R. solani | Ggt | F. culmorum | P. ultimum |
|---|---|---|---|---|
| WPB | 5.03 ± 0.64 | 11.40 ± 1.25 | 4.44 ± 0.87 | 2.17 ± 0.98 |
| WPB ocf | 0.00 ± 0.00 | 5.46 ± 0.61 | 0.00 ± 0.00 | 0.00 ± 0.00 |
| Strain | Species | Source | R. solani | Ggt | F. culmorum | P. ultimum |
|---|---|---|---|---|---|---|
| AFE 1 | Rahnella sp. | Snoqualmie River wild poplar 4 | ++ | ++++ | ++ | − |
| AFE 3 | Rahnella sp. | Snoqualmie River wild poplar 4 | − | ++++ | − | + |
| AFE 4A | Bacillus velezensis | Snoqualmie River wild poplar 4 | ++++ | ++++ | +++ | + |
| AFE 5 | Pseudomonas sp. | Snoqualmie River wild poplar 4 | +++ | ++ | + | + |
| AFE 8 | Pseudomonas sp. | Yakima River wild poplar 12 | +++ | ++ | ++ | − |
| AFE 9 | Rahnella aquatilis | Yakima River wild poplar 12 | ++ | +++ | − | ++ |
| AFE 11 | Aureobasidium pullulans | Skykomish River wild poplar | + | ++ | − | − |
| AFE 14 | Pantoea agglomerans | Skykomish River wild poplar | − | ++ | − | − |
| AFE 16 | Pseudomonas graminis | Skykomish River wild poplar | + | +++ | − | − |
| AFE 21B | Bacillus velezensis | Skykomish River wild poplar | ++ | ++++ | ++ | − |
| AFE 22 | Rahnella sp. | Skykomish River wild poplar | − | +++ | − | + |
| Strain | Total # of BGC | BGC’s with Known Antimicrobial Activity 1 | Activity |
|---|---|---|---|
| WPB | 12 | cepacin A (62%) | antibiotic/antioomycetes |
| ornibactin (93%) | antibiotic | ||
| occidiofungin (94%) | antifungal | ||
| AFE 1 | 7 | - | |
| AFE 3 | 6 | - | |
| AFE 4A | 21 | difficidin 2 (100%) | antibacterial |
| macrolactin (100%) | antibacterial/antifungal | ||
| bacillaene (100%) | antibacterial | ||
| fengycin (86%) | antibacterial | ||
| bacilysin (100%) | antibacterial/antifungal | ||
| bacillibactin (100%) | antibacterial | ||
| lanthipeptide class II | antibacterial | ||
| AFE 5 | 18 | sessilin 2 (100%) | antifungal |
| AFE 8 | 14 | viscosine (100%) | antifungal |
| tolaasin (60%) | antibacterial/antifungal | ||
| AFE 9 | 5 | - | |
| AFE 16 | 5 | - | |
| AFE 21B | 21 | difficidin (100%) | antibacterial |
| macrolactin (100%) | antibacterial/antifungal | ||
| bacillaene (100%) | antibacterial | ||
| fengycin (80%) | antibacterial | ||
| bacilysin (100%) | antibacterial/antifungal | ||
| bacillibactin (100%) | antibacterial | ||
| surfactin (52%) | antibacterial | ||
| AFE 22 | 7 |
| Strain | Level of Siderophore Production |
|---|---|
| AFE 1 | ++ |
| AFE 3 | ++ |
| AFE 4A | − |
| AFE 5 | +++ |
| AFE 8 | +++ |
| AFE 9 | ++ |
| AFE 11 | − |
| AFE 14 | ++ |
| AFE 16 | + |
| AFE 21B | − |
| AFE 22 | +++ |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Doty, S.L.; Joubert, P.M.; Firrincieli, A.; Sher, A.W.; Tournay, R.; Kill, C.; Parikh, S.S.; Okubara, P. Potential Biocontrol Activities of Populus Endophytes against Several Plant Pathogens Using Different Inhibitory Mechanisms. Pathogens 2023, 12, 13. https://doi.org/10.3390/pathogens12010013
Doty SL, Joubert PM, Firrincieli A, Sher AW, Tournay R, Kill C, Parikh SS, Okubara P. Potential Biocontrol Activities of Populus Endophytes against Several Plant Pathogens Using Different Inhibitory Mechanisms. Pathogens. 2023; 12(1):13. https://doi.org/10.3390/pathogens12010013
Chicago/Turabian StyleDoty, Sharon L., Pierre M. Joubert, Andrea Firrincieli, Andrew W. Sher, Robert Tournay, Carina Kill, Shruti S. Parikh, and Patricia Okubara. 2023. "Potential Biocontrol Activities of Populus Endophytes against Several Plant Pathogens Using Different Inhibitory Mechanisms" Pathogens 12, no. 1: 13. https://doi.org/10.3390/pathogens12010013
APA StyleDoty, S. L., Joubert, P. M., Firrincieli, A., Sher, A. W., Tournay, R., Kill, C., Parikh, S. S., & Okubara, P. (2023). Potential Biocontrol Activities of Populus Endophytes against Several Plant Pathogens Using Different Inhibitory Mechanisms. Pathogens, 12(1), 13. https://doi.org/10.3390/pathogens12010013






