Breeding and Genomic Approaches towards Development of Fusarium Wilt Resistance in Chickpea
Abstract
1. Introduction
2. Fusarium Wilt
3. Genetics of Resistance to Fusarium Wilt
| Fusarium Race | Name of Resistance Gene | Number and Nature of Wilt Resistance Gene | Effect of Resistance Gene on Wilting | Symptoms | References |
|---|---|---|---|---|---|
| 0 | FOC-01/FOC-01 | Monogenic or digenic | Complete resistance | Yellowing | [26] |
| FOC-02/FOC-02 | |||||
| 1A | h1 (syn FOC-1) | Trigenic | Late wilting | Wilting | [57] |
| h2 | Late wilting | ||||
| H3 | Late wilting | ||||
| 1B/C | - | - | - | Yellowing | [63] |
| 2 | FOC-2 | Monogenic | Complete resistance | Wilting | [27] |
| 3 | FOC-3/FOC-3 | Monogenic | Complete resistance | Wilting | [62] |
| 4 | FOC-4 | Monogenic recessive | Complete resistance | Wilting | [27] |
| 5 | FOC-5/FOC-5 | Monogenic | Complete resistance | Wilting | [67] |
| 6 | - | - | - | Wilting | [63] |
4. Breeding Methods Employed for Fusarium Wilt Resistance in Chickpea
5. Screening Strategies to Identify Wilt-Resistant Genotypes
5.1. Field Screening
5.2. Screening under Controlled Conditions
5.2.1. Greenhouse Screening
5.2.2. Laboratory Screening
6. Management of Fusarium Wilt in Chickpea
6.1. Utilization of Pathogen-Free Planting Material
6.2. Chemical Control
6.3. Biological Control
6.4. Cultural Control
6.5. Use of Resistant Cultivars
7. Advanced Breeding Techniques
7.1. Marker Technology
7.1.1. Molecular Markers and FW Resistance in Chickpea
7.1.2. Marker-Assisted Breeding
Variations in MAS
Marker-Assisted Backcrossing (MABC)
Marker-Assisted Gene Pyramiding (MAGP)
Marker-Assisted Recurrent Selection (MARS)
7.2. Genetic Mapping and QTL Technique
7.3. Genome Sequencing
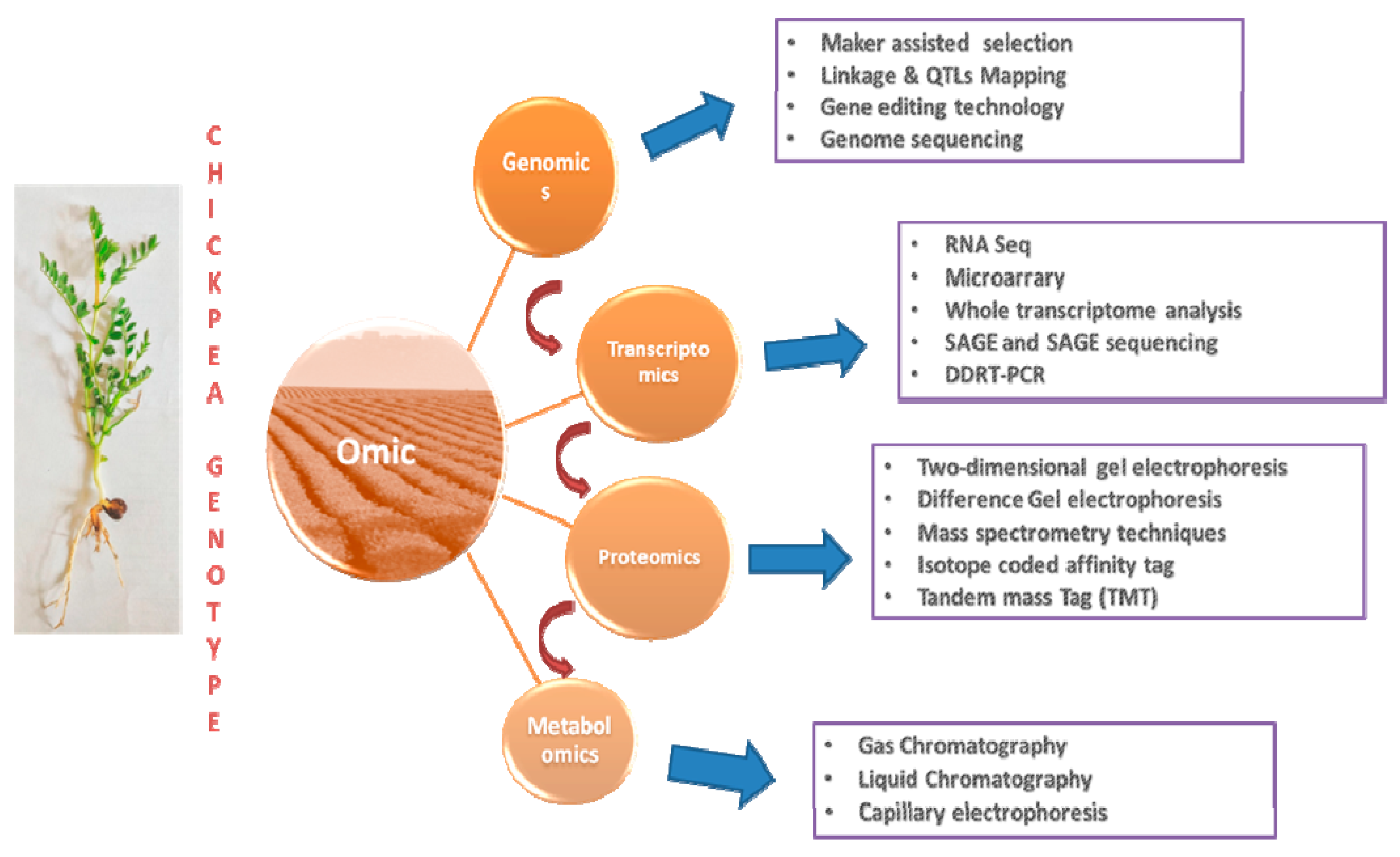
8. Multi-Omics Approaches
8.1. Transcriptomics/Gene Expression Studies
8.2. Proteomics and Metabolomics
9. Genomic Selection (GS)
10. In Vitro Selection against Fusarium Wilt Disease Tolerance/Resistance in Chickpea
11. Speed Breeding in Chickpea Improvement
12. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Varshney, R.K.; Thudi, M.; Pandey, M.K.; Tardieu, F.; Ojiewo, C.; Vadez, V.; Whit Bread, A.M.; Siddique, K.H.M.; Nguyen, H.T.; Carberry, P.S.; et al. Accelerating genetic gains in legumes for the development of prosperous smallholder agriculture: Integrating genomics, phenotyping, systems modelling and agronomy. J. Exp. Bot. 2018, 69, 3293–3312. [Google Scholar] [CrossRef] [PubMed]
- Cobos, M.J.; Rubio, J.; Fernández-Romero, M.D.; Garza, R.; Moreno, M.T.; Millán, T.; Gil, J. Genetic analysis of seed size, yield and days to flowering in a chickpea recombinant inbred line population derived from a Kabuli×Desi cross. Ann. Appl. Biol. 2007, 151, 33–42. [Google Scholar] [CrossRef]
- Singh, S.; Singh, I.; Kapoor, K.; Gaur, P.M.; Chaturvedi, S.K.; Singh, N.P.; Sandhu, J.S. Chickpea. In Broadening the Genetic Base of Grain Legumes; National Bureau of Plant Genetic Resources: New Delhi, India, 2014. [Google Scholar]
- Patil, M.G. Wilt of Chickpea with Special Reference to Characterization of Races and Variant of Fusarium oxysporum f. sp. ciceris and Parameters Associated with Resistance. Jawaharlal Nehru Krishi Vishwa Vidyalaya Jabalpur 2015. Available online: https://krishikosh.egranth.ac.in/handle/1/68318 (accessed on 15 January 2023).
- Solanki, R.S.; Babbar, A.; Tripathi, N. Genetic diversity analysis in Kabuli chickpea (Cicer arietinum L.) genotypes based on quantitative traits and molecular markers. Bangladesh J. Bot. 2022, 51, 581–587. [Google Scholar] [CrossRef]
- Kaur, R.; Grewal, S.K.; Singh, S.; Kaur, J.; Bhardwaj, R.D. Desi and kabuli chickpea cultivars had differential behavior towards salinity stress tolerance. Biol. Futur. 2020, 71, 137–146. [Google Scholar] [CrossRef]
- Zhang, C.; Chen, T.; Chen, W.; Sankaran, S. Non-invasive evaluation of Ascochyta blight disease severity in chickpea using field asymmetric ion mobility spectrometry and hyperspectral imaging techniques. Crop Prot. 2023, 165, 106163. [Google Scholar] [CrossRef]
- Kaur, K.; Grewal, S.K.; Gill, P.S.; Singh, S. Comparison of cultivated and wild chickpea genotypes for nutritional quality and antioxidant potential. J. Food Sci. Technol. 2019, 56, 1864. [Google Scholar] [CrossRef]
- Khatoon, N.; Prakash, J. Nutritional quality of microwave-cooked and pressure-cooked legumes. Int. J. Food Sci. Nutr. 2004, 55, 441. [Google Scholar] [CrossRef]
- Lukus, P.K.; Doma, K.M.; Duncan, A.M. The Role of Pulses in Cardiovascular Disease Risk for Adults with Diabetes. Am. J. Lifestyle Med. 2020, 14, 571. [Google Scholar] [CrossRef]
- Sahu, V.K.; Tiwari, S.; Gupta, N.; Tripathi, M.K.; Yasin, M. Evaluation of physiological and biochemical contents in desi and Kabuli chickpea. Legume Res. 2020, 45, 1197–1208. [Google Scholar] [CrossRef]
- Marin, J.A.; Andreu, P.; Carrasco, A.; Arbeloa, A. Determination of proline concentration, an abiotic stress marker, in root exudates of excised root cultures of fruit tree root stocks under salt stress. Rev. Region. Arid. Num. 2010, 24, 722–727. [Google Scholar]
- Wallace, T.C.; Murray, R.; Zelman, K.M. The Nutritional Value and Health Benefits of Chickpeas and Hummus. Nutrients 2016, 8, 766. [Google Scholar] [CrossRef]
- Madurapperumage, A.; Tang, L.; Thavarajah, P.; Bridges, W.; Shipe, E.; Vandemark, G.; Thavarajah, D. Chickpea (Cicer arietinum L.) as a Source of Essential Fatty Acids—A Biofortification Approach. Front. Plant Sci. 2021, 12, 734980. [Google Scholar] [CrossRef]
- Asati, R.; Tripathi, M.K.; Tiwari, S.; Yadav, R.K.; Tripathi, N. Molecular Breeding and Drought Tolerance in Chickpea. Life 2022, 12, 1846. [Google Scholar] [CrossRef]
- Belete, T. The Role of Conventional and Molecular Techniques for Chickpea (Cicer arietinum L.) Improvement. Adv. Biotechnol. Microbiol. 2018, 10, 555780. [Google Scholar]
- Gaur, P.M.; Krishnamurthy, L.; Kashiwagi, J. Improving drought-avoidance root traits in chickpea (Cicer arietinum L.)-current status of research at ICRISAT. Plant Prod Sci. 2008, 11, 3–11. [Google Scholar] [CrossRef]
- Kashiwagi, J.; Krishnamurthy, L.; Crouch, J.H.; Serraj, R. Variability of root length density and its contributions to seed yield in chickpea (Cicer arietinum L.) under terminal drought stress. Field Crop Res. 2006, 95, 171–181. [Google Scholar] [CrossRef]
- Choudhary, A.K.; Jain, S.K.; Dubey, A.K.; Kumar, J.; Sharma, M.; Gupta, K.C.; Sharma, L.D.; Prakash, V.; Kumar, S. Conventional and molecular breeding for disease resistance in chickpea: Status and strategies. Biotechnol. Genet. Eng. Rev. 2022, 12, 1–32. [Google Scholar] [CrossRef]
- Varshney, R.K.; Graner, A.; Sorrells, M.E. Genomics-assisted breeding for crop improvement. Trends Plant Sci. 2005, 10, 621–630. [Google Scholar]
- Li, H.; Rodda, M.; Gnanasambandam, A.; Aftab, M.; Redden, R.; Hobson, K.; Rosewarne, G.; Michael, M.; Sukhjiwan, K.; Anthony, S. Breeding for biotic stress resistance in chickpea: Progressand prospects. Euphytica 2015, 204, 257–288. [Google Scholar] [CrossRef]
- Ningwal, R.; Tripathi, M.K.; Tiwari, S.; Yadav, R.K.; Tripathi, N.; Solanki, R.S.; Asati, R.; Yasin, M. Assessment of genetic variability, correlation and path coefficient analysis for yield and its attributing traits in chickpea (Cicer arietinum L.). Pharma Innov J. 2023, 12, 4851–4859. [Google Scholar]
- Verma, S.; Gupta, S.; Nitesh, B.; Kumar, T.; Bharadwaj, C.; Bhatia, S. High-density linkage map construction and mapping of seed trait QTLs in chickpea (Cicer arietinum L.) using Genotyping-by Sequencing (GBS). Sci. Rep. 2015, 5, 17512. [Google Scholar] [CrossRef] [PubMed]
- Gaur, P.M.; Thudi, M.; Srinivasan, S.; Varshney, R.K. Advances in chickpea genomics. In Legumes in the Omic Era; Gupta, S., Nadarajan, N., Gupta, D.S., Eds.; Springer: New York, NY, USA, 2014; pp. 73–94. [Google Scholar]
- Sahu, V.K.; Tiwari, S.; Tripathi, M.K.; Gupta, N.; Tomar, R.S.; Yasin, M. Morpho physiological and biochemical traits analysis for Fusarium wilt disease using gene-based markers in desi and Kabuli genotypes of chickpea (Cicer arietinum L.). Indian J. Genet. 2020, 80, 163–172. [Google Scholar]
- Sharma, K.D.; Winter, P.; Kahl, G.; Muehlbauer, F.J. Molecular mapping of Fusarium oxysporum f. sp. ciceris race 3 resistance gene in chickpea. Theor. Appl. Genet. 2003, 108, 1243–1248. [Google Scholar] [CrossRef] [PubMed]
- Sharma, K.D.; Chen, W.; Muehlbauer, F.J. Genetics of Chickpea Resistance to Five Races of Fusarium Wilt and a Concise Set of Race Differentials for Fusarium oxysporum f. sp. ciceris. Plant Dis. 2005, 89, 385–390. [Google Scholar] [CrossRef]
- Bharadwaj, C.; Jorben, J.; Rao, A.; Roorkiwal, M.; Patil, B.S.; Jayalakshmi; Ahammed, S.K.; Saxena, D.R.; Yasin, M.; Jahagirdar, J.E.; et al. Development of High Yielding Fusarium Wilt Resistant Cultivar by Pyramiding of “Genes” Through Marker-Assisted Backcrossing in Chickpea (Cicer arietinum L.). Front. Genet. 2022, 13, 924287. [Google Scholar] [CrossRef]
- Wang, Y.; Dong, S. A new roadmap for the breeding of disease-resistant and high-yield crops. Stress Biol. 2021, 1, 21. [Google Scholar] [CrossRef]
- Milla´n, T.; Clarke, H.J.; Siddique, K.H.M.; Buhariwalla, H.K.; Gaur, P.M.; Kumar, J.; Gil, J.; Kahl, G.; Winter, P. Chickpea molecular breeding: New tools and concepts. Euphytica 2006, 147, 81–103. [Google Scholar] [CrossRef]
- Millan, T.; Winter, P.; Jüngling, R.; Gil, J.; Rubio, J.; Cho, S.; Cobos, M.J.; Iruela, M.; Rajesh, P.N.; Tekeoglu, M.; et al. A consensus genetic map of chickpea (Cicer ari-etinum L.) based on 10 mapping populations. Euphytica 2010, 175, 175–189. [Google Scholar] [CrossRef]
- Hasan, N.; Choudhary, S.; Naaz, N.; Sharma, N.; Laskar, R.A. Recent advancements in molecular marker-assisted selection and applications in plant breeding programmes. J. Genet. Eng. Biotechnol. 2021, 19, 1. [Google Scholar] [CrossRef]
- Metzker, M.L. Sequencing technologies—The next generation. Nat. Rev. Genet. 2009, 11, 31–46. [Google Scholar] [CrossRef]
- Llaca, V. Sequencing technologies and their use in plant biotechnology and breeding. In DNA Sequencing–Methods and Applications; Munshi, A., Ed.; InTech: Rijeka, Croatia, 2012; pp. 35–60. [Google Scholar]
- Wang, Z.; Gerstein, M.; Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 2009, 10, 57. [Google Scholar] [CrossRef]
- Jia, J.; Zhao, S.; Kong, X.; Li, Y.; Zhao, G.; He, W.; Appels, R.; Pfeifer, M.; Tao, Y.; Zhang, X.; et al. Aegilops tauschii draft genome sequence reveals a gene repertoire for wheat adaptation. Nature 2013, 496, 91–95. [Google Scholar] [CrossRef]
- Sierro, N.; Battey, J.N.; Ouadi, S.; Bakaher, N.; Bovet, L.; Willig, A.; Goepfert, S.; Peitsch, M.C.; Ivanov, N.V. The tobacco genome sequence and its comparison with those of tomato and potato. Nat. Commun. 2014, 5, 3833. [Google Scholar] [CrossRef]
- Gaur, R.; Azam, S.; Jeena, G.; Khan, A.W.; Choudhary, S.; Jain, M.; Yadav, G.; Tyagi, A.K.; Chattopadhyay, D.; Bhatia, S. High-Throughput SNP Discovery and Genotyping for Constructing a Saturated Linkage Map of Chickpea (Cicer arietinum L.). DNA Res. 2012, 19, 357–373. [Google Scholar] [CrossRef]
- Azam, S.; Thakur, V.; Ruperao, P.; Shah, T.; Balaji, J.; Amindala, B.; Farmer, A.D.; Studholme, D.J.; May, G.D.; Edwards, D.; et al. Coverage-Based Consensus Calling (CBCC) of Short Sequence Reads and Comparison of CBCC Results to Identify SNPs In Chickpea (Cicer arietinum; Fabaceae), A Crop Species Without A Reference Genome. Am. J. Bot. 2012, 99, 1–7. [Google Scholar] [CrossRef]
- Kudapa, H.; Azam, S.; Sharpe, A.G.; Taran, B.; Li, R.; Deonovic, B.; Cameron, C.; Farmer, A.D.; Cannon, S.B.; Varshney, R.K. Comprehensive transcriptome assembly of Chickpea (Cicer arietinum L.) using sanger and next generation sequencing platforms: Development and applications. PLoS ONE 2014, 9, e86039. [Google Scholar] [CrossRef]
- Basandrai, A.S.; Basandrai, D.; Duraimurugan, P.; Srinivasan, T. Breeding for biotic stress. In Biology and Breeding of Food Legumes; Pratap, A., Kumar, J., Eds.; CAB International: Wallingford, UK, 2011; pp. 220–240. [Google Scholar]
- Ahmad, F.; Gaur, P.; Croser, J. Chickpea (Cicer arietinum L.). In Genetic Resources, Chromosome Engineering and Crop Improvement—Grain Legumes; Singh, R., Jauhar, P.P., Eds.; CRC Press: Boca Raton, FL, USA, 2005; pp. 187–217. [Google Scholar]
- Knights, E.J.; Southwell, R.J.; Schwinghamer, M.W.; Harden, S. Resistance to Phytophthora medicaginis Hansen and Maxwell in wild Cicer species and its use in breeding root rot resistant chickpea (Cicer arietinum L.). Aust. J. Agric. Res. 2008, 59, 383–387. [Google Scholar] [CrossRef]
- Singh, R.; Sharma, P.; Varshney, R.K.; Sharma, S.K.; Singh, N.K. Chickpea improvement: Role of wild species and genetic markers. Biotechnol. Genet. Eng. Rev. 2008, 25, 267–314. [Google Scholar] [CrossRef]
- Fahmy, F.I.; Taha, O.; Eiashry, A.N. First Genome Analysis and Molecular Characterization of Chickpea Chlorotic DwarfVirus Egyptian Isolate Infecting Squash. Virus Dis. 2015, 2, 33–41. [Google Scholar] [CrossRef]
- Madrid, E.; Seoane, P.; Claros, M.G.; Barro, F.; Rubio, J.; Gil, J.; Millan, T. Genetic and physical mapping of the QTLAR3 controlling blight resistance in chickpea (Cicer arietinum L). Euphytica 2014, 198, 69–78. [Google Scholar] [CrossRef]
- Singh, P.K.; Singh, M.; Agnihotri, V.K.; Vyas, D. Arbuscular Mycorrhizal Fungi: Biocontrol against Fusarium wilt of Chickpea. Int. J. Sci. Res. Publ. 2013, 3, 1–5. [Google Scholar]
- Cunnington, J.; Lindbeck, K.; Jones, R.H. National Diagnostic Protocol for the Detection of Fusarium Wilt of Chick-pea (Fusarium Oxysporum f. sp. ciceris); Plant Health Australia: Camberra, Australia, 2007. [Google Scholar]
- Sharma, M.; Nagavardhini, A.; Thudi, M.; Ghosh, R.; Pande, S.; Varshney, R.K. Development of DArT markers and assessment of diversity in Fusarium oxysporum f. sp. ciceris, wilt pathogen of chickpea (Cicer arietinum L.). BMC Genom. 2014, 15, 454. [Google Scholar] [CrossRef] [PubMed]
- Achari, S.R.; Mann, R.C.; Sharma, M.; Edwards, J. Diagnosis of Fusarium oxysporum f. sp. ciceris causing Fusarium wilt of chickpea using loop-mediated isothermal amplification (LAMP) and conventional end-point PCR. Sci. Rep. 2023, 13, 2640. [Google Scholar] [CrossRef]
- Jiménez-díaz, R.M.; Jiménez-gasco, M.M. Integrated Management of Fusarium Wilt Diseases. In Control of Fusarium Diseases; Transworld Research Network: Kerala, India, 2011; pp. 177–215. [Google Scholar]
- Singh, U.M.; Sareen, P.; Sengar, R.; Kumar, A. Plant ionomics: A newer approach to study mineral transport and its regulation. Acta Physiol. Plant. 2013, 35, 2641. [Google Scholar] [CrossRef]
- Chand, H.; Khirbat, S.K.; L, C.C. Chickpea Wilt and Its Management—A Review. Agric Rev. 2009, 30, 1–12. [Google Scholar]
- Sampaio, A.M.; Araújo, S.D.S.; Rubiales, D.; Patto, M.C.V. Fusarium Wilt Management in Legume Crops. Agronomy 2020, 10, 1073. [Google Scholar] [CrossRef]
- Jiménez-Fernández, D.; Landa, B.B.; Kang, S.; Jiménez-Díaz, R.M.; Navas-Cortés, J.A. Quantitative and Microscopic Assessment of Compatible and Incompatible Interactions between Chickpea Cultivars and Fusarium oxysporum f. sp. ciceris Races. PLoS ONE 2013, 8, E61360. [Google Scholar] [CrossRef]
- Jimenez-Gasco, D.M.; Jimenez-Diaz, R.M. Development of a specific polymerase chain reaction-based assay for the identification of Fusarium oxysporum f. sp. ciceris and its pathogenic races 0, 1A, 5, and 6. Phytopathology 2003, 93, 200–209. [Google Scholar] [CrossRef]
- Sharma, K.D.; Muehlbauer, F.J. Fusarium wilt of chickpea: Physiological specialization, genetics of resistance and resistance gene tagging. Euphytica 2007, 157, 1–14. [Google Scholar] [CrossRef]
- Jendoubi, W.; Bouhadida, M.; Boukteb, A.; Béji, M.; Kharrat, M. Fusarium Wilt Affecting Chickpea Crop. Agriculture 2017, 7, 23. [Google Scholar] [CrossRef]
- Castro, P.; Rubio, J.; Millán, T.; Gil, J.; Cobos, M.J. Fusarium wilt in chickpea: General aspect and molecular Breed-ing. In Fusarium: Epidemiology, Environmental Sources and Prevention; Rios, T.F., Ortega, E.R., Eds.; Nova Science Publishers: New York, NY, USA, 2012; pp. 101–122. [Google Scholar]
- Sharma, M.; Telangre, R.; Ghosh, R.; Pande, S. Multi-environment field testing to identify broad, stable resistance to sterility mosaic disease of pigeonpea. J. Gen. Plant Pathol. 2015, 81, 249–259. [Google Scholar] [CrossRef]
- Gowda, M.; Radhika, P.; Kadoo, N.; Mhase, L.B.; Gupta, V.S. Molecular mapping of wilt resistance genes in chickpea. Mol. Breed. 2009, 24, 177–183. [Google Scholar] [CrossRef]
- Tekeoglu, M.; Tullu, A.; Kaiser, W.J.; Muehlbauer, F.J. Inheritance and linkage of two genes that confer resistance to Fusarium wilt in chickpea. Crop Sci. 2000, 40, 1247–1251. [Google Scholar] [CrossRef]
- Raj, J.D.; Nelson, J.A.; Rao, K.S.P. A Study on the Effects of Some Reinforcers to Improve Performance of Employees in a Retail Industry. Behav. Modif. 2006, 90, 365–375. [Google Scholar] [CrossRef]
- Al-taae, A.K.; Hadwan, H.A.; Al-jobory, S.A.E. Physiological Races of Fusarium oxysporum f. sp. Ciceris in Iraq. J. Life Sci. 2013, 7, 1070–1075. [Google Scholar]
- Shehabu, M.; Ahmed, S.; Sakhuja, P.K. Pathogenic variability in Ethiopian isolates of Fusarium oxysporum f. sp. ciceris and reaction of chickpea improved varieties to the isolates. Int. J. Pest Manag. 2008, 54, 143–149. [Google Scholar] [CrossRef]
- Jimenez-Gasco, M.D.M.; Pérez-Artés, E.; Jiménez-Diaz, R.M. Pathogenic variability in Ethiopian isolates of Fusarium oxysporum f. sp. ciceris and reaction of chickpea improved varieties to the isolates. Eur. J. Plant Pathol. 2001, 107, 237–248. [Google Scholar] [CrossRef]
- Tullu, A.; Muehlbauer, F.; Simon, C.; Mayer, M.; Kumar, J.; Kaiser, W.; Kraft, J. Inheritance and linkage of a gene for resistance to race 4 of fusarium wilt and RAPD markers in chickpea. Euphytica 1998, 102, 227–232. [Google Scholar] [CrossRef]
- Srivastava, A.K.; Saxena, D.R.; Saabale, P.R.; Raghuvanshi, K.S.; Anandani, V.P.; Singh, R.K.; Sharma, O.P.; Wasinikar, A.R.; Sahni, S.; Varshney, R.K. Delineation of genotype-by-environment interactions for identification and validation of resistant genotypes in chickpea to fusarium wilt using GGE biplot. Crop Prot. 2021, 144, 105571. [Google Scholar] [CrossRef]
- Sharma, M.; Tarafdar, A.; Ghosh, R.; Gopalakrishanan, S. Biological control as a tool for eco-friendly management of plant pathogens. In Advances in Soil Microbiology: Recent Trends and Future Prospects; Adhya, T., Mishra, B., Annapurna, K., Verma, D., Kumar, U., Eds.; Springer Press: Singapore, 2017; pp. 153–188. [Google Scholar] [CrossRef]
- Pande, S.; Kishore, G.K.; Upadhyaya, H.D.; Rao, J.N. Identification of Sources of Multiple Disease Resistance in Mini-core Collection of Chickpea. Plant Dis. 2006, 90, 1214–1218. [Google Scholar] [CrossRef] [PubMed]
- Mirzapour, S.; Darvishnia, M.; Bazgir, E.; Goodarzi, D. Identification of resistant sources in chickpea against Fusarium wilt under greenhouse condition. Intl. J. Farm. Alli. Sci. 2014, 3, 772–776. [Google Scholar]
- Chobe, D.R.; Gupta, O.; Pawar, M. Radiation induced mutation for resistance against races/pathotypes of Fusarium oxysporum f. sp ciceris in chickpea (Cicer arietinum L.). Indian Phytopath 2016, 69, 260–265. [Google Scholar]
- Sharma, M.; Babu, T.K.; Gaur, P.; Ghosh, R.; Rameshwar, T.; Chaudhary, R.; Upadhyay, J.; Gupta, O.; Saxena, D.; Kaur, L.; et al. Identification and multi-environment validation of resistance to Fusarium oxysporum f. sp. ciceris in chickpea. Field Crop. Res. 2012, 135, 82–88. [Google Scholar] [CrossRef]
- Fikre, A.; Korbu, L.; Eshete, M.; Bekele, D.; Girma, N.; Mohamed, R.; Assefe, S.; Admasu, D.; Tilahun, G.; Tesfaye, T. A decade of research progress in chickpea and lentil breeding and genetics. Ethiop. J. Crop Sci. 2018, 6, 101–113. [Google Scholar]
- Moore, R.; Casale, F.P.; Bonder, M.J.; Horta, D.; BIOS Consortium; Franke, L.; Barroso, I.; Stegle, O. A linear mixed-model approach to study multivariate gene–environment interactions. Nat. Genet. 2018, 51, 180–186. [Google Scholar] [CrossRef]
- Yan, W. GGE biplot–a windows application for graphical analysis of multi-environment trial data and other types of two-way data. Agronomy 2001, 93, 1111–1118. [Google Scholar] [CrossRef]
- Yan, W.; Hunt, L.A.; Sheng, Q.L.; Szlavnics, Z. Cultivar evaluation and mega-environment investiga-tion based on the GGE biplot. Crop Sci. 2000, 40, 597–605. [Google Scholar] [CrossRef]
- Sharma, M.; Babu, T.K.; Ghosh, R.; Telangre, R.; Rathore, A.; Kaur, L.; Kushwaha, K.; Das, R.; Pande, S. Multi-environment field testing for identification and validation of genetic resistance to Botrytis cinerea causing Botrytis grey mold in chickpea (Cicer arietinum L.). Crop. Prot. 2013, 54, 106–113. [Google Scholar] [CrossRef]
- Gaur, P.M.; Jukanti, A.K.; Srinivasan, S.; Gowda, C.L.L. Chickpea (Cicer arietinum L.). In Breeding of Field Crops; Bharadwaj, D.N., Ed.; Agrobios: Jodhpur, India, 2012; pp. 165–194. [Google Scholar]
- Millan, T.; Madrid, E.; JI, C.; Amri, M.; Castro, P.; Rubio, J. Chickpea. InGrain Legumes; De Ron Antonio, M., Ed.; Grain legumes: New York, NY, USA, 2015. [Google Scholar]
- Pande, S.; Gaur, P.M.; Sharma, M.; Rao, J.N.; Rao, B.V.; Krishna Kishore, G. Identification of single and multiple disease resistance in desi chickpea genotypes to Ascochyta blight, Botrytis gray mold and Fusarium wilt. SAT J. 2007, 3, 1–3. [Google Scholar]
- Berrada, A.F.; Shivakumar, B.G.; Yaburaju, N.T. Chickpea in cropping systems. In Chickpea Breeding and Man-Agement; Yadav, S.S.S., Redden, R., Chen, W., Sharma, B., Eds.; CABI Publishing: Wallingford, UK, 2007; pp. 193–212. [Google Scholar]
- Kaiser, W.J.; Alcala-Jimenez, A.R.; Hervas-Vargas, A.; Trapero-Casas, J.L.; Jiménez-Díaz, R.M. Screening of wild Cicer species for resistance to races 0 and 5 of Fusarium oxysporum f. sp. ciceris. Plant Dis. 1994, 78, 962–967. [Google Scholar] [CrossRef]
- Ali, M.; Inanaga, S.; Sugimoto, Y. Sources of resistance to Fusarium wilt of chickpea in Sudan. Phytopathol. Mediterr. 2002, 41, 163–169. [Google Scholar]
- Sharma, M.; Ghosh, R.; Tarafdar, A.; Rathore, A.; Chobe, D.R.; Kumar, A.V.; Gaur, P.M.; Samineni, S.; Gupta, O.; Singh, N.P.; et al. Exploring the Genetic Cipher of Chickpea (Cicer arietinum L.) Through Identification and Multi-environment Validation of Resistant Sources Against Fusarium Wilt (Fusarium oxysporum f. sp. ciceris). Front. Sustain. Food Syst. 2019, 3, 78. [Google Scholar] [CrossRef]
- Bohra, A.; Jha, U.C.; Kavi Kishor, P.B.; Pandey, S.; Singh, N.P. Genomics and molecular breeding in lesser ex-plored pulse crops: Current trends and future opportunities. Biotechnol. Adv. 2014, 32, 1410–1428. [Google Scholar] [CrossRef]
- Bohra, A.; Pandey, M.K.; Jha, U.C.; Singh, B.; Singh, I.P.; Datta, D.; Chaturvedi, S.K.; Nadarajan, N.; Varshney, R.K. Genomics-assisted breeding in four major pulse crops of developing countries: Present status and prospects. Theor. Appl. Genet. 2014, 127, 1263–1291. [Google Scholar] [CrossRef]
- Gaur, P.M.; Gowda, C.L.L.; Knights, E.J.; Warkentin, T.; Acikoz, N.; Yadav, S.S.; Kumar, J. Breeding Achievements. In Chickpea Breeding; Centre for Agriculture and Bioscience International (CABI): Wallingford, UK, 2007. [Google Scholar]
- Deshmukh, R.B.; Patil, J.V.; Mhase, L.B.; Kamble, M.S.; Deshmukh, D.V.; Jamadagni, B.M. Digvijay—An High Yielding, Wilt Resistant Chickpea Variety for Maharashtra State. J. Maharashtra agric. Univ. 2010, 35, 367–370. [Google Scholar]
- Chaudhary, R.G. Chickpea diseases. In 25 Years of Research at IIPR; Indian Institute of Pulses Research: Kanpur, India, 2009; pp. 85–115. [Google Scholar]
- Roorkiwal, M.; Bharadwaj, C.; Barmukh, R.; Dixit, G.P.; Thudi, M.; Gaur, P.M.; Chaturvedi, S.K.; Fikre, A.; Hamwieh, A.; Kumar, S.; et al. Integrating genomics for chickpea improvement: Achievements and opportunities. Theor. Appl. Genet. 2020, 133, 1703–1720. [Google Scholar] [CrossRef]
- Pande, S.; Sharma, M.; Nagavardhini, A.; Rameshwar, T. High throughput phenotyping of chickpea diseases: Stepwise identification of host plant resistance. In Information Bulletin No. 92; ICRISAT: Patancheru, Andhra Pradesh, India, 2012. [Google Scholar]
- Infantino, A.; Kharrat, M.; Riccioni, L.; Coyne, C.J.; McPhee, K.E.; Grunwald, N.J. Screening techniques and sources of resistance to root diseases of food legumes. Euphytica 2006, 147, 201–221. [Google Scholar] [CrossRef]
- Nene, Y.L.; Haware, M.P. Screening chickpea for resistance to wilt. Pl. Dis. 1980, 64, 379–380. [Google Scholar] [CrossRef]
- Sharma, K.D.; Muehlbauer, F.J. Genetic mapping of Fusarium oxysporum f. sp. ciceris race-speciWc resistance genes in chickpea (Cicer arietinum L.). In Abstract of the International Food Legume Research Conference—IV; Indian Agricultural Research Institute: New Delhi, India, 2005; pp. 18–22. [Google Scholar]
- Ratna Babu, D.; Ravikumar, R.L. Genetic evidence for resistance to Fusarium wilt of pollen grains in chickpea (Cicer arietinum L.). Curr. Sci. 2010, 96, 811–815. [Google Scholar]
- Landa, B.B.; Navas-Cortés, J.A.; Jiménez-Díaz, R.M. Integrated management of fusarium wilt of chickpea with sowing date, host resistance, and biological control. Phytopathology 2004, 94, 946–960. [Google Scholar] [CrossRef] [PubMed]
- Jiménez-Díaz, R.M.; Castillo, P.; Jiménez-Gasco, M.D.M.; Landa, B.B.; Navas-Cortés, J.A. Fusarium wilt of chickpeas: Biology, ecology and management. Crop Prot. 2015, 73, 16–27. [Google Scholar] [CrossRef]
- Jime´nez-Dı´az, R.M.; Jime´nez-Gasco, M.M.; Landa, B.B.; Castillo, P.; Navas-Corte´s, J.A. Fusarium wilt of chickpea. In Compendium of Chickpea and Lentil Diseases; Chen, W., Sharma, H.C., Muehlbauer, F.J., Eds.; APS Press: Paul, MN, USA, 2011; pp. 16–20. ISBN 978-0-89054-384-9. [Google Scholar]
- Zacharia, S.; Jaiswal, K.K.; Pandey, P. Integrated Management of Chickpea Wilt Incited by Fusarium oxysporum f. sp. ciceris. Int. J. Agric. Res. 2012, 7, 284–290. [Google Scholar]
- Jamil, A.; Ashraf, S. Utilization of chemical fungicides in managing the wilt disease of chickpea caused by Fusarium oxysporum f. sp. ciceri. Arch. Phytopathol. Plant Prot. 2020, 53, 876. [Google Scholar] [CrossRef]
- Singh, P.K.; Vyas, D. Biocontrol of plant diseases and sustainable agriculture. Proc. Natl. Acad. Sci. India–Sect. B Biol. Sci. 2009, 79, 110–128. [Google Scholar]
- Shukla, A.; Dehariya, K.; Vyas, D.; Jha, A. Interactions between arbuscular mycorrhizae and Fusarium oxysporum f. sp. ciceris: Effects on fungal development, seedling growth and wilt disease suppression in Cicer arietinum L. Arch. Phytopathol. Plant Prot. 2014, 48, 240–252. [Google Scholar] [CrossRef]
- Dehariya, K.; Shukla, A.; Sheikh, I.A.; Vyas, D. Trichoderma and arbuscular mycorrhizal fungi based biocontrol of fusariumudum butler and their growth promotion effects on pigeon pea. J. Agric. Sci. Technol. 2015, 17, 505–517. [Google Scholar]
- Dubey, S.C.; Suresh, M.; Singh, B. Evaluation of Trichoderma species against Fusarium oxysporum f. sp. ciceris for integrated management of chickpea wilt. Biol. Control 2007, 40, 118–127. [Google Scholar] [CrossRef]
- Fravel, D.; Olivain, C.; Alabouvette, C. Fusarium oxysporum and its biocontrol. New Phytol. 2003, 157, 493–502. [Google Scholar] [CrossRef]
- Landa, B.B.; Hervás, A.; Bettiol, W.; Jiménez-Díaz, R.M. Antagonistic activity of Bacteria from the chickpea rhizo-sphere against Fusarium Oxysporum f. sp. ciceris. Phytoparasitica 1997, 25, 305–318. [Google Scholar] [CrossRef]
- Larkin, R.P.; Fravel, D.R. Efficacy of Various Fungal and Bacterial Biocontrol Organisms for Control of Fusarium Wilt of Tomato. Plant Dis. 2007, 82, 1022–1028. [Google Scholar] [CrossRef]
- Chérif, M.; Arfaoui, A.; Rhaiem, A. Phenolic compounds and their role in bio-control and resistance of chickpea to fungal pathogenic attacks. Tunis. J. Plant Prot. 2007, 2, 7–21. [Google Scholar]
- Tamietti, G.; Valentino, D. Soil solarization as an ecological method for the control of Fusarium wilt of melon in Italy. Crop. Prot. 2006, 25, 389. [Google Scholar] [CrossRef]
- Landa, B.B.; Navas-Cortés, J.A.; Jiménez-Gasco, M.M.; Katan, J.; Retig, B.; Jiménez-Díaz, R.M. Temperature response of chickpea cultivars to races of Fusarium oxysporum f. sp. ciceris, causal agent of Fusarium wilt. Phytopathology 2006, 90, 365–374. [Google Scholar] [CrossRef]
- Strange, R.N. Introduction to Plant Pathology; John Wiley & Sons Ltd.: London, UK, 2003. [Google Scholar]
- Orr, R.; Nelson, P.L. Impacts of soil abiotic attributes on Fusarium wilt, focusing on bananas. Appl. Soil Ecol. 2018, 132, 20–33. [Google Scholar] [CrossRef]
- Bayraktar, H.; Dolar, F.S. Pathogenic variability of Fusarium oxysporum f. sp. ciceris isolates from chickpea in Turkey. Pakistan J. Bot. 2012, 44, 821–823. [Google Scholar]
- Heather, J.M.; Chain, B. The sequence of sequencers: The history of sequencing DNA. Genomics 2015, 107, 1–8. [Google Scholar] [CrossRef]
- Gasperskaja, E.; Kučinskas, V. The most common technologies and tools for functional genome analysis. Acta Med. Lituanica. 2017, 24, 1–11. [Google Scholar] [CrossRef]
- Bunnik, E.M.; Le Roch, K.G. An Introduction to Functional Genomics and Systems Biology. Adv. Wound Care 2013, 2, 490–498. [Google Scholar] [CrossRef]
- Collard, B.C.Y.; Jahufer, M.Z.Z.; Brouwer, J.B.; Pang, E.C.K. An introduction to markers, quantitative trait loci (QTL) mapping and marker-assisted selection for crop improvement: The basic concepts. Euphytica 2005, 142, 169–196. [Google Scholar] [CrossRef]
- Platten, J.D.; Cobb, J.N.; Zantua, R.E. Criteria for evaluating molecular markers: Comprehensive quality metrics to improve marker-assisted selection. PLoS ONE 2019, 14, e0210529. [Google Scholar] [CrossRef] [PubMed]
- Dheer, P.; Rautela, I.; Sharma, V.; Dhiman, M.; Sharma, A.; Sharma, N.; Sharma, M.D. Evolution in crop improvement approaches and future prospects of molecular markers to CRISPR/Cas9 system. Gene 2020, 753, 144795. [Google Scholar] [CrossRef] [PubMed]
- Mohammadi, M.; Maibody, S.A.; Golkar, P. Application of DNA Molecular Markers in Plant Breeding. J. Plant Genet. Res. 2019, 6, 1–30. [Google Scholar]
- Ahmar, S.; Gill, R.A.; Jung, K.-H.; Faheem, A.; Qasim, M.U.; Mubeen, M.; Zhou, W. Conventional and Molecular Techniques from Simple Breeding to Speed Breeding in Crop Plants: Recent Advances and Future Outlook. Int. J. Mol. Sci. 2020, 21, 2590. [Google Scholar] [CrossRef]
- Mallikarjuna, B.P.; Samineni, S.; Thudi, M.; Sajja, S.B.; Khan, A.W.; Patil, A.; Viswanatha, K.P.; Varshney, R.K.; Gaur, P.M. Molecular Mapping of Flowering Time Major Genes and QTLs in Chickpea (Cicer arietinum L.). Front. Plant Sci. 2017, 8, 1140. [Google Scholar] [CrossRef]
- Koul, B.; Sharma, K.; Sehgal, V.; Yadav, D.; Mishra, M.; Bharadwaj, C. Chickpea (Cicer arietinum L.) Biology and Biotechnology: From Domestication to Biofortification and Biopharming. Plants 2022, 11, 2926. [Google Scholar] [CrossRef]
- Cobos, M.; Fernandez, M.; Rubio, J.; Kharrat, M.; Moreno, M.; Gill, J.; Millan, T. A linkage map of chickpea based on popu-lations from Kabuli x Desi crosses: Location of genes for resistance to Fusarium wilt race 0. Theor. Appl. Genet. 2005, 110, 1347–1353. [Google Scholar] [CrossRef]
- Hajibarat, Z.; Saidi, A.; Hajibarat, Z.; Talebi, R. Characterization of genetic diversity in chickpea using SSR markers, Start Codon Targeted Polymorphism (SCoT) and Conserved DNA-Derived Polymorphism (CDDP). Physiol. Mol. Biol. Plants 2015, 21, 365–373. [Google Scholar] [CrossRef]
- Garg, R.; Patel, R.K.; Jhanwar, S.; Priya, P.; Bhattacharjee, A.; Yadav, G.; Bhatia, S.; Chattopadhyay, D.; Tyagi, A.K.; Jain, M. Gene discovery and tissue-specific transcriptome analysis in chickpea with massively parallel pyrosequencing and web resource development. Plant Physiol. 2011, 156, 661–678. [Google Scholar] [CrossRef]
- Hiremath, P.J.; Farmer, A.; Cannon, S.B.; Woodward, J.; Kudapa, H.; Tuteja, R.; Kumar, A.; Bhanuprakash, A.; Mulaosmanovic, B.; Gujaria, N.; et al. Largescale transcriptome analysis in chickpea (Cicer arietinum L.), an orphan legume crop of the semiarid tropics of Asia and Africa. Plant Biotechnol. J. 2011, 9, 922–931. [Google Scholar] [CrossRef]
- Hiremath, P.J.; Kumar, A.; Penmetsa, R.V.; Farmer, A.; Schlueter, J.; Chamarthi, S.K.; Whaley, A.M.; Carrasquilla-Garcia, N.; Gaur, P.; Upadhyaya, H.D.; et al. Large-scale development of cost-effective SNP marker assays for diversity assessment and genetic mapping in chickpea and comparative mapping in legumes. Plant Biotechnol. J. 2012, 10, 716–732. [Google Scholar] [CrossRef]
- Thudi, M.; Bohra, A.; Nayak, S.N.; Varghese, N.; Shah, T.; Penmetsa, R.V.; Thirunavukkarasu, N.; Gudipati, S.; Gaur, P.; Kulwal, P.L.; et al. Novel SSR Markers from BAC-End Sequences, DArT Arrays and a Comprehensive Genetic Map with 1291 Marker Loci for Chickpea (Cicer arietinum L.). PLoS ONE 2011, 6, e27275. [Google Scholar] [CrossRef]
- Ali, L.; Azam, S.; Rubio, J.; Kudapa, H.; Madrid, E.; Varshney, R.K.; Castro, P.; Chen, W.; Gil, J.; Millan, T. Detection of a new QTL/gene for growth habit in chickpea CaLG1 using wide and narrow crosses. Euphytica 2015, 204, 473–485. [Google Scholar] [CrossRef]
- Kumar, J.; Choudhary, A.K.; Solanki, R.K.; Pratap, A. Towards marker-assisted selection in pulses: A review. Plant Breed. 2011, 130, 297–313. [Google Scholar] [CrossRef]
- Torres, A. Application of molecular markers for breeding disease resistant varieties in crop plants. In Molecular Techniques in Crop Improvement; Jain, S.M., Brar, D.S., Eds.; Springer Science Business Media B.V.: Berlin/Heidelberg, Germany, 2010. [Google Scholar] [CrossRef]
- Varshney, R.K.; Mohan, S.M.; Gaur, P.M.; Chamarthi, S.K.; Singh, V.K.; Srinivasan, S.; Swapna, N.; Sharma, M.; Singh, S.; Kaur, L.; et al. Marker-assisted backcrossing to introgress resistance to Fusarium wilt (FW) race 1 and Ascochyta blight (AB) in C 214, an elite cultivar of chickpea. Plant Genome 2014, 7. [Google Scholar] [CrossRef]
- Pratap, A.; Chaturvedi, S.K.; Tomar, R.; Rajan, N.; Malviya, N.; Thudi, M.; Saabale, P.R.; Prajapati, U.; Varshney, R.K.; Singh, N.P. Marker-assisted introgression of resistance to fusarium wilt race 2 in Pusa 256, an elite cultivar of desi chickpea. Mol. Genet. Genom. 2017, 292, 1237–1245. [Google Scholar] [CrossRef]
- Castro, P.; Pistón, F.; Madrid, E.; Millán, T.; Gil, J.; Rubio, J. Development of chickpea near-isogenic lines for fusari-um wilt. Theor. Appl. Genet. 2010, 121, 1519–1526. [Google Scholar] [CrossRef]
- Radhika, P.; Gowda, S.J.; Kadoo, N.Y.; Mhase, L.B.; Jamadagni, B.M.; Sainani, M.N.; Chandra, S.; Gupta, V. Development of an 461 integrated intraspecific map of chickpea (Cicer ari-etinum 462 L.) using two recombinant inbred line populations. Theor. 463 Appl.Genet. 2007, 115, 209–216. [Google Scholar] [CrossRef]
- Winter, P.; Benko-Iseppon, A.-M.; Hüttel, B.; Ratnaparkhe, M.; Tullu, A.; Sonnante, G.; Pfaff, T.; Tekeoglu, M.; Santra, D.; Sant, V.J.; et al. A linkage map of the chickpea (Cicer arietinum L.) genome based on recombinant inbred lines from a C. arietinum×C. reticulatum cross: Localization of resistance genes for fusarium wilt races 4 and 5. Theor. Appl. Genet. 2000, 101, 1155–1167. [Google Scholar] [CrossRef]
- Mayer, M.S.; Tullu, A.; Simon, C.J.; Kumar, J.; Kaiser, W.J.; Kraft, J.M.; Muehlbauer, F.J. Development of a DNA Marker for Fusarium Wilt Resistance in Chickpea. Crop. Sci. 1997, 37, 1625–1629. [Google Scholar] [CrossRef]
- Ratnaparkhe, M.B.; Santra, D.K.; Tullu, A.; Muehlbauer, F.J. Inheritance of inter-simple-sequence-repeat polymorphisms and linkage with a fusarium wilt resistance gene in chickpea. Theor. Appl. Genet. 1998, 96, 348–353. [Google Scholar] [CrossRef] [PubMed]
- Ratnaparkhe, M.B.; Tekeoglu, M.; Muehlbauer, F.J. Inter-simple-sequence-repeat (ISSR) polymorphisms are useful for finding markers associated with disease resistance gene clusters. Theor. Appl. Genet. 1998, 97, 515–519. [Google Scholar] [CrossRef]
- Gupta, P.; Varshney, R. The development and use of microsatellite markers for genetic analysis and plant breeding with emphasis on bread wheat. Euphytica 2000, 113, 163–185. [Google Scholar] [CrossRef]
- Sethy, N.K.; Shokeen, B.; Edwards, K.J.; Bhatia, S. Development of microsatellite markers and analysis of intraspe-cific genetic variability in chickpea (Cicer arietinum L.). Theor. Appl. Genet. 2006, 112, 1416–1428. [Google Scholar] [CrossRef] [PubMed]
- Nayak, S.N.; Zhu, H.; Varghese, N.; Datta, S.; Choi, H.-K.; Horres, R.; Jüngling, R.; Singh, J.; Kishor, P.B.K.; Sivaramakrishnan, S.; et al. Integration of novel SSR and gene-based SNP marker loci in the chickpea genetic map and establishment of new anchor points with Medicago truncatula genome. Theor. Appl. Genet. 2010, 120, 1415–1441. [Google Scholar] [CrossRef]
- Varshney, R.K.; Graner, A.; Sorrells, M.E. Genic microsatellite markers in plants: Features and applications. Trends Biotechnol. 2005, 23, 48–55. [Google Scholar] [CrossRef]
- Gujaria, N.; Kumar, A.; Dauthal, P.; Dubey, A.; Hiremath, P.; Prakash, A.B.; Farmer, A.; Bhide, M.; Shah, T.; Gaur, P.M.; et al. Development and use of genic molecular markers (GMMs) for construction of a transcript map of chickpea (Cicer arietinum L.). Theor. Appl. Genet. 2011, 122, 1577–1589. [Google Scholar] [CrossRef]
- Varshney, R.K.; Hiremath, P.J.; Lekha, P.; Kashiwagi, J.; Balaji, J.; A Deokar, A.; Vadez, V.; Xiao, Y.; Srinivasan, R.; Gaur, P.M.; et al. A comprehensive resource of drought- and salinity- responsive ESTs for gene discovery and marker development in chickpea (Cicer arietinum L.). BMC Genom. 2009, 10, 523. [Google Scholar] [CrossRef]
- Ravikumar, R.L.; Babu, D.R. In vitro screenings of chickpea genotypes for Fusarium wilt resistance through root feeding of pathotoxin. Curr. Sci. 2007, 93, 20–21. [Google Scholar]
- Tiwari, S.; Tripathi, N.; Tsuji, K.; Tantwai, K. Genetic diversity and population structure of Indian soybean [Glycine max (L.) Merr.] as revealed by microsatellite markers. Physiol. Mol. Biol. Plants 2019, 25, 953–964. [Google Scholar] [CrossRef]
- Soregaon, C.D.; Ravikumar, R.L. Marker assisted characterization of wilt resistance in productive chickpea genotypes. Electron. J. Plant Breed. 2021, 1, 1159–1163. [Google Scholar]
- Shaheen, T.S.; Kausar, H.; Mahmood-ur-Rahman, A.; Mahmud, T. Breeding the conventional and mo-lecular methods for identification of resistance against fusarium wilt in chickpea germplasm. J. Anim. Plant Sci. 2017, 27, 559–566. [Google Scholar]
- Udupa, S.M.; Baum, M. Genetic dissection of pathotype-specific resistance to ascochyta blight disease in chickpea (Cicer arietinum L.) using microsatellite markers. Theor. Appl. Genet. 2002, 106, 1196–1202. [Google Scholar] [CrossRef]
- Lichtenzveig, J.; Bonfil, D.J.; Zhang, H.-B.; Shtienberg, D.; Abbo, S. Mapping quantitative trait loci in chickpea asso-ciated with time to flowering and resistance to Didymellarabiei the causal agent of Ascochyta blight. Theor. Appl. Genet. 2006, 113, 1357–1369. [Google Scholar] [CrossRef]
- Gaur, P.M.; Jukanti, A.K.; Varshney, R.K. Impact of Genomic Technologies on Chickpea Breeding Strategies. Agronomy 2012, 2, 199–221. [Google Scholar] [CrossRef]
- Temesgen, B.; Begna, T.; Yesuf, H.; Abdurezake, M.; Eshetu, G. Genetic mapping in crop plants. Open J. Plant Sci. 2021, 6, 19–26. [Google Scholar] [CrossRef]
- Collard, B.C.Y.; Mackill, D.J. Marker-assisted selection: An approach for precision plant breeding in the twenty-first century Marker-assisted selection: An approach for precision plant breeding in the twenty-first century. Phil. Trans. R. Soc. B 2008, 363, 557–572. [Google Scholar] [CrossRef]
- Allahverdipoor, K.H.; Bahramnejad, B.; Amini, J. Selection of molecular markers associated with resistance to Fusarium wilt disease in chickpea (Cicer arietinum L.) using multivariate statistical techniques. AJCS 2011, 5, 1801–1809. [Google Scholar]
- Choudhary, K.; Choudhary, O.P.; Shekhawat, N.S. Marker assisted selection: A novel approach for crop improvement. Am. Eura. J. Sci. Res. 2008, 1, 26–30. [Google Scholar]
- Sato, S.; Isobe, S.; Tabata, S. Structural analyses of the genomes in legumes. Curr. Opin. Plant Biol. 2010, 13, 146–152. [Google Scholar] [CrossRef]
- Nadeem, M.A.; Nawaz, M.A.; Shahid, M.Q.; Doğan, Y.; Comertpay, G.; Yıldız, M.; Hatipoğlu, R.; Ahmad, F.; Alsaleh, A.; Labhane, N.; et al. DNA molecular markers in plant breeding: Current status and recent advancements in ge-nomic selection and genome editing. Biotechnol. Biotechnol. Equip. 2018, 32, 261–285. [Google Scholar] [CrossRef]
- Holland, J.B. Implementation of molecular markers for quantitative traits in breeding programs—challenges and opportunities. In New Directions for a Diverse Planet: Proceedings for the 4th International Crop Science Congress; Regional Institute: Gosford, Australia, 2004; Available online: https://www.cropscience.org.au/icsc2004 (accessed on 15 January 2023).
- Ribaut, J.M.; Sawkins, M.C.; Bänziger, M.; Vargas, M.; Huerta, E.; Martinez, C.; Moreno, M. Marker-assisted selection in tropical maize based on consensus map, perspectives, and limitations. Resilient Crops Water Ltd. Environ. 2004, 267–268. [Google Scholar]
- Mannur, D.M.; Babbar, A.; Thudi, M.; Sabbavarapu, M.M.; Roorkiwal, M.; Yeri, S.B.; Bansal, V.P.; Jayalakshmi, S.K.; Singh Yadav, S.; Rathore, A.; et al. Super An-nigeri 1 and improved JG 74: Two Fusarium wilt-resistant introgression lines developed using marker-assisted backcrossing approach in chickpea (Cicer arietinum L.). Mol. Breed. 2019, 39, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Varshney, R.K. Exciting journey of 10 years from genomes to fields and markets: Some success stories of ge-nomics-assisted breeding in chickpea, pigeonpea and groundnut. Plant Sci. 2016, 242, 98–107. [Google Scholar] [CrossRef] [PubMed]
- Rana, M.; Sood, A.; Hussain, W.; Kaldate, R.; Sharma, T.R.; Gill, R.K.; Kumar, S.; Singh, S. Chapter 6—Gene Pyramiding and Multiple Character Breeding. In Lentils: Potential Resources for Enhancing Genetic Gains, 1st ed.; Singh, M., Ed.; Academic Press: London, UK, 2019; pp. 83–124. [Google Scholar]
- Qi, L.; Ma, G. Marker-Assisted Gene Pyramiding and the Reliability of Using SNP Markers Located in the Recombination Suppressed Regions of Sunflower (Helianthus annuus L.). Genes 2019, 11, 10. [Google Scholar] [CrossRef] [PubMed]
- Dormatey, R.; Sun, C.; Ali, K.; Coulter, J.; Bi, Z.; Bai, J. Gene Pyramiding for Sustainable Crop Improvement against Biotic and Abiotic Stresses. Agronomy 2020, 10, 1255. [Google Scholar] [CrossRef]
- Jaganathan, D.; Bohra, A.; Thudi, M.; Varshney, R.K. Fine mapping and gene cloning in the post-NGS era: Advances and prospects. Theor. Appl. Genet. 2020, 133, 1791–1810. [Google Scholar] [CrossRef] [PubMed]
- Yadava, D.K. Fundamentals of Field Crop Breeding; Springer Nature: Berlin/Heidelberg, Germany, 2022. [Google Scholar] [CrossRef]
- Jha, U.C.; Sharma, K.D.; Nayyar, H.; Parida, S.K.; Siddique, K.H.M. Breeding and Genomics Interventions for Developing Ascochyta Blight Resistant Grain Legumes. Int. J. Mol. Sci. 2022, 23, 2217. [Google Scholar] [CrossRef]
- Chagné, D.; Brown, G.; Lalanne, C.; Madur, D.; Pot, D.; Neale, D.; Plomion, C. Comparative genome and QTL mapping between maritime and loblolly pines. Mol. Breed. 2003, 12, 185–195. [Google Scholar] [CrossRef]
- Halila, I.; Cobos, M.J.; Rubio, J.; Millán, T.; Kharrat, M.; Marrakchi, M.; Gil, J. Tagging and mapping a second resistance gene for Fusarium wilt race 0 in chickpea. Eur. J. Plant Pathol. 2008, 124, 87–92. [Google Scholar] [CrossRef]
- Jendoubi, W.; Bouhadida, M.; Millan, T.; Kharrat, M.; Gil, J.; Rubio, J.; Madrid, E. Identification of the target region including the Foc0 1 /foc0 1 gene and development of near isogenic lines for resistance to Fusarium Wilt race 0 in chickpea. Euphytica 2016, 210, 119–133. [Google Scholar] [CrossRef]
- Sabbavarapu, M.M.; Sharma, M.; Chamarthi, S.K.; Swapna, N.; Rathore, A.; Thudi, M.; Gaur, P.M.; Pande, S.; Singh, S.; Kaur, L.; et al. Molecular mapping of QTLs for resistance to Fusarium wilt (race 1) and Ascochyta blight in chickpea (Cicer arietinum L.). Euphytica 2013, 193, 121–133. [Google Scholar] [CrossRef]
- Cobos, M.; Winter, P.; Kharrat, M.; Cubero, J.; Gil, J.; Millan, T.; Rubio, J. Genetic analysis of agronomic traits in a wide cross of chickpea. Field Crop. Res. 2009, 111, 130–136. [Google Scholar] [CrossRef]
- Rajesh, P.N.; Tullu, A.; Gil, J.; Gupta, V.; Ranjekar, P.; Muehlbauer, F. Identification of an STMS marker for the double-podding gene in chickpea. Theor. Appl. Genet. 2002, 105, 604–607. [Google Scholar] [CrossRef]
- Jingade, P.; Ravikumar, R.L. Development of molecular map and identification of QTLs linked to Fusarium wilt resistance in chickpea. J. Genet. 2015, 94, 723–729. [Google Scholar] [CrossRef]
- Garg, T.; Mallikarjuna, B.P.; Thudi, M.; Samineni, S.; Singh, S.; Sandhu, J.S.; Kaur, L.; Singh, I.; Sirari, A.; Basandrai, A.K.; et al. Identification of QTLs for resistance to Fusarium wilt and Ascochyta blight in a recombinant inbred population of chickpea (Cicer arietinum L.). Euphytica 2018, 214, 45. [Google Scholar] [CrossRef]
- Caballo, C.; Castro, P.; Gil, J.; Millan, T.; Rubio, J.; Die, J.V. Candidate genes expression profiling during wilting in chickpea caused by Fusarium oxysporum f. sp. ciceris race 5. PLoS ONE 2019, 14, e0224212. [Google Scholar] [CrossRef]
- Benko-Iseppon, A.M.; Winter, P.; Huettel, B.; Staginnus, C.; Muehlbauer, F.J.; Kahl, G. Molecular markers closely linked to Fusarium resistance genes in chickpea show signiWcant alignments to pathogenesis-related genes located on Ara-bidopsis chromosomes 1 and 5. Theoret. Appl. Genet. 2003, 107, 379–386. [Google Scholar] [CrossRef]
- Ali, H.; Haq, M.A.U.; Shah, T.M.; Rahman, M.U.; Chen, W. Validation of molecular markers for resistance among Pakistani chickpea germplasm to races of Fusarium oxysporum f. sp. ciceris. Eur. J. Plant Pathol. 2011, 132, 237–244. [Google Scholar] [CrossRef]
- Nimbalkar, S.B.; Harsulkar, A.M.; Giri, A.P.; Sainani, M.N.; Franceschi, V.; Gupta, V.S. Differentially expressed gene transcripts in roots of resistant and susceptible chickpea plant (Cicer arietinum L.) upon Fusarium oxysporum infection. Physiol. Mol. Plant Pathol. 2006, 68, 176–188. [Google Scholar] [CrossRef]
- Lam, H.Y.; Clark, M.J.; Chen, R.; Chen, R.; Natsoulis, G.; O’huallachain, M.; Dewey, F.E.; Habegger, L.; Ashley, E.A.; Gerstein, M.B. Performance comparison of whole-genome sequencing platforms. Nat. Biotechnol. 2012, 30, 78–83. [Google Scholar] [CrossRef] [PubMed]
- Leo, V.; Morgan, N.; Bem, D.; Jones, M.; Lowe, G.; Lordkipanidzé, M.; Drake, S.; Simpson, M.; Gissen, P.; Mumford, A.; et al. Use of next-generation sequencing and candidate gene analysis to identify underlying defects in patients with inherited platelet function disorders. J. Thromb. Haemost. 2015, 13, 643–650. [Google Scholar] [CrossRef] [PubMed]
- Kulski, J.K.; Suzuki, S.; Ozaki, Y.; Mitsunaga, S.; Inoko, H.; Shiina, T. In Phase HLA Genotyping by Next Generation Sequencing—A Comparison Between Two Massively Parallel Sequencing Bench-Top Systems, the Roche GS Junior and Ion Torrent PGM. InTech 2014, 141–181. [Google Scholar] [CrossRef]
- Rabbani, B.; Tekin, M.; Mahdieh, N. The promise of whole-exome sequencing in medical genetics. J. Hum. Genet. 2013, 59, 5–15. [Google Scholar] [CrossRef] [PubMed]
- Ashraf, N.; Ghai, D.; Barman, P.; Basu, S.; Gangisetty, N.; Mandal, M.K.; Chakraborty, N.; Datta, A.; Chakraborty, S. Comparative analyses of genotype dependent expressed sequence tags and stress-responsive transcriptome of chickpea wilt illustrate predicted and unexpected genes and novel regulators of plant immunity. BMC Genom. 2009, 10, 415. [Google Scholar] [CrossRef] [PubMed]
- Varshney, R.K.; Roorkiwal, M.; Sun, S.; Bajaj, P.; Chitikineni, A.; Thudi, M.; Singh, N.P.; Du, X.; Upadhyaya, H.D.; Khan, A.W.; et al. A chickpea genetic variation map based on the sequencing of 3366 genomes. Nature 2021, 599, 622–627. [Google Scholar] [CrossRef]
- Williams, A.H.; Sharma, M.; Thatcher, L.F.; Azam, S.; Hane, J.K.; Sperschneider, J.; Kidd, B.N.; Anderson, J.P.; Ghosh, R.; Garg, G.; et al. Comparative genomics and prediction of conditionally dispensable sequences in legume-infecting Fusarium ox- ysporum formaes-peciales facilitates identification of candidate effectors. BMC Genom. 2016, 17, 191. [Google Scholar] [CrossRef]
- Srivastava, A.K.; Kashyap, P.L.; Chakdar, H.; Kumar, M.; Srivastava, A.K.; Yadav, J.; Jamali, H.; Srivastava, R.; Sharma, A.; Ti- wari, P.; et al. First de novo draft genome sequence of the pathogenic fungus Fusarium udum F02845, associated with pigeonpea (Cajanus cajan L. Millspaugh) wilt. Microbiol. Resour. Announc. 2018, 7, e1001–e1018. [Google Scholar] [CrossRef]
- Muthamilarasan, M.; Singh, N.K.; Prasad, M. Multi-omics approaches for strategic improvement of stress tolerance in underuti- lized crop species: A climate change perspective. Adv. Genet. 2019, 103, 1–38. [Google Scholar] [CrossRef]
- Salt, D.E.; Baxter, I.; Lahner, B. Ionomics and the Study of the Plant Ionome. Annu. Rev. Plant Biol. 2008, 59, 709–733. [Google Scholar] [CrossRef]
- Houle, D.; Govindaraju, D.R.; Omholt, S. Phenomics: The next challenge. Nat. Rev. Genet. 2010, 11, 855–866. [Google Scholar] [CrossRef]
- Talukdar, D.; Sinjushin, A. Cytogenomics and mutagenomics in plant functional biology and breeding. In PlantOmics: The Omics of Plant Science, 1st ed.; Barh, D., Khan, M., Davies, E., Eds.; Springer: New Delhi, India, 2015; pp. 113–156. [Google Scholar] [CrossRef]
- Wu, S.; Ning, F.; Zhang, Q.; Wu, X.; Wang, W. Enhancing Omics Research of Crop Responses to Drought under Field Conditions. Front. Plant Sci. 2017, 8, 174. [Google Scholar] [CrossRef]
- Saabale, P.R.; Dubey, S.C.; Priyanka, K.; Sharma, T.R. Analysis of differential transcript expression in chickpea during compatible and incompatible interactions with Fusarium oxysporum f. sp. ciceris Race 4. 3 Biotech 2018, 8, 111. [Google Scholar] [CrossRef]
- Ashraf, N.; Basu, S.; Narula, K.; Ghosh, S.; Tayal, R.; Gangisetty, N.; Biswas, S.; Aggarwal, P.R.; Chakraborty, N.; Chakraborty, S. Integrative network analyses of wilt transcriptome in chickpea reveal genotype dependent regulatory hubs in immunity and susceptibility. Sci. Rep. 2018, 8, 6528. [Google Scholar] [CrossRef]
- Gupta, S.; Bhar, A.; Chatterjee, M.; Das, S. Fusarium oxysporum f.sp. ciceri Race 1 Induced Redox State Alterations Are Coupled to Downstream Defense Signaling in Root Tissues of Chickpea (Cicer arietinum L.). PLoS ONE 2013, 8, e73163. [Google Scholar] [CrossRef]
- Priyadarshini, P.; Sahu, S.; Kalwan, G.; Yadava, Y.K.; Nagar, R.; Rai, V.; Bharadwaj, C.; Gaikwad, K.; Jain, P.K. Unravelling the mechanism of Fusarium wilt resistance in chickpea seedlings using biochemical studies and expression analysis of NBS-LRR and WRKY genes. Physiol. Mol. Plant Pathol. 2023, 124, 101958. [Google Scholar] [CrossRef]
- Kumar, Y.; Zhang, L.; Panigrahi, P.; Dholakia, B.B.; Dewangan, V.; Chavan, S.G.; Kunjir, S.M.; Wu, X.; Li, N.; Rajmohanan, P.R.; et al. Fusarium oxysporum mediates systems metabolic reprogramming of chickpea roots as revealed by a combination of proteomics and metabolomics. Plant Biotechnol. J. 2016, 14, 1589–1603. [Google Scholar] [CrossRef]
- Upasani, M.L.; Gurjar, G.S.; Kadoo, N.Y.; Gupta, V.S. Dynamics of Colonization and Expression of Pathogenicity Related Genes in Fusarium oxysporum f.sp. ciceri during Chickpea Vascular Wilt Disease Progression. PLoS ONE 2016, 11, e0156490. [Google Scholar] [CrossRef]
- Kohli, D.; Joshi, G.; Deokar, A.A.; Bhardwaj, A.R.; Agarwal, M.; Katiyar-Agarwal, S.; Srinivasan, R.; Jain, P.K. Identification and Characterization of Wilt and Salt Stress-Responsive MicroRNAs in Chickpea through High-Throughput Sequencing. PLoS ONE 2014, 9, e108851. [Google Scholar] [CrossRef]
- Dandale, S.N.; Mane, S.S.; Ingle, S.T.; Patil, A.N.; Nandanwal, R.S.; Jadhav, P.V. Candidate gene expression pro-filing during wilting in chickpea caused by Fusarium oxysporum f. sp. ciceri in race 2 and race. Pharma—Novation J. 2022, 11, 223–228. [Google Scholar]
- Pelizzola, M.; Ecker, J.R. The DNA methylome. FEBS Lett. 2011, 585, 1994–2000. [Google Scholar] [CrossRef] [PubMed]
- Jain, S.; Weeden, N.F.; Kumar, A.; Chittem, K.; McPhee, K. Functional Codominant Marker for Selecting the Fw Gene Conferring Resistance to Fusarium Wilt Race 1 in Pea. Crop. Sci. 2015, 55, 2639–2646. [Google Scholar] [CrossRef]
- Kole, C. Genomic Designing for Biotic Stress Resistant Pulse Crops; Springer Nature: Berlin/Heidelberg, Germany, 2022. [Google Scholar] [CrossRef]
- Pratap, A.; Das, A.; Kumar, S.; Gupta, S. Current Perspectives on Introgression Breeding in Food Legumes. Front. Plant Sci. 2021, 11, 589189. [Google Scholar] [CrossRef] [PubMed]
- Großkinsky, D.K.; Syaifullah, S.J.; Roitsch, T. Integration of multi-omics techniques and physiological phenotyping within a holistic phenomics approach to study senescence in model and crop plants. J. Exp. Bot. 2017, 69, 825–844. [Google Scholar] [CrossRef]
- Raza, A.; Tabassum, J.; Kudapa, H.; Varshney, R.K. Can omics deliver temperature resilient ready-to-grow crops? Crit. Rev. Biotechnol. 2021, 41, 1209. [Google Scholar] [CrossRef]
- Vahdati, K. Abiotic Stress—Plant Responses and Applications in Agriculture; BoD–Books on Demand: Paris, France, 2013; pp. 49–102. [Google Scholar] [CrossRef]
- El-Metwally, S.; Ouda, O.M.; Helmy, M. Next Generation Sequencing Technologies and Challenges in Sequence Assembly, 1st ed.; Springer: New York, NY, USA, 2014. [Google Scholar] [CrossRef]
- Nataraja, K.N.; Madhura, B.G.; Parvathi, S.M. Omics: Modern tools for precise understanding of drought adaptation in plants. In Plant OMICS and Crop Breeding; Zargar, S.M., Rai, V., Eds.; Apple Academic Press: Palm Bay, FL, USA, 2017; pp. 289–320. [Google Scholar]
- Kawahara, Y.; Oono, Y.; Kanamori, H.; Matsumoto, T.; Itoh, T.; Minami, E. Simultaneous RNA-Seq Analysis of a Mixed Tran- scriptome of Rice and Blast Fungus Interaction. PLoS ONE 2012, 7, e49423. [Google Scholar] [CrossRef]
- De Cremer, K.; Mathys, J.; Vos, C.; Froenicke, L.; Michelmore, R.W.; Cammue, B.P.A.; DE Coninck, B. RNA seq-based transcrip- tome analysis of Lactuca sativa, infected by the fungal necrotrophy. Botrytis Cinerea. Plant Cell Environ. 2013, 36, 1992–2007. [Google Scholar] [CrossRef]
- Ke, R.; Mignardi, M.; Pacureanu, A.; Svedlund, J.; Botling, J.; Wählby, C.; Nilsson, M. In situ sequencing for RNA analysis in preserved tissue and cells. Nat. Methods 2013, 10, 857–860. [Google Scholar] [CrossRef]
- Burgess, D.J. Putting transcriptomics in its place. Nat. Rev. Genet. 2015, 16, 319. [Google Scholar] [CrossRef]
- Wise, R.P.; Moscou, M.J.; Bogdanove, A.J.; Whitham, S.A. Transcript Profiling in Host–Pathogen Interactions. Annu. Rev. Phyto- pathol. 2007, 45, 329–369. [Google Scholar] [CrossRef]
- Xue, R.; Wu, J.; Zhu, Z.; Wang, L.; Wang, X.; Wang, S.; Blair, M.W. Differentially Expressed Genes in Resistant and Susceptible Common Bean (Phaseolus vulgaris L.) Genotypes in Response to Fusarium oxysporum f. sp. phaseoli. PLoS ONE 2015, 10, e0127698. [Google Scholar] [CrossRef]
- Gupta, S.; Chakraborti, D.; Rangi, R.K.; Basu, D.; Das, S. A Molecular Insight into the Early Events of Chickpea (Cicer arietinum) and Fusarium oxysporum f. sp. ciceri (Race 1) Interaction Through cDNA-AFLP Analysis. Phytopathology 2009, 99, 1245–1257. [Google Scholar] [CrossRef]
- Li, C.Y.; Deng, G.M.; Yang, J.; Viljoen, A.; Jin, Y.; Kuang, R.-B.; Zuo, C.W.; Lv, Z.C.; Yang, Q.S.; Sheng, O.; et al. Transcriptome profiling of resistant and susceptible Cavendish banana roots following inoculation with Fusarium oxysporum f. sp. cubense tropical race. BMC Genom. 2012, 13, 374. [Google Scholar] [CrossRef]
- Caballo, C.; Madrid, E.; Gil, J.; Chen, W.; Rubio, J.; Millan, T. Saturation of genomic region implicated in resis- tance to Fusarium oxysporum f. sp. ciceris race 5 in chickpea. Mol. Breed. 2019, 39, 16. [Google Scholar] [CrossRef]
- Mosa, K.A.; Ismail, A.; Helmy, M. Omics and system biology approaches in plant stress research. In Plant Stress Tolerance: An Integrated Omics Approach; Mosa, K.A., Ismail, A., Helmy, M., Eds.; Springer: Cham, Germany, 2017; pp. 21–34. [Google Scholar] [CrossRef]
- Aizat, W.M.; Hassan, M. Proteomics in systems biology. In Omics Applications for Systems Biology. Advances in Experimental Med- icine and Biology; Aizat, W., Goh, H.H., Baharum, S., Eds.; Springer: Cham, Germany, 2018; pp. 31–49. [Google Scholar] [CrossRef]
- Castillejo, M.; Bani, M.; Rubiales, D. Understanding pea resistance mechanisms in response to Fusarium oxysporum through proteomic analysis. Phytochemistry 2015, 115, 44–58. [Google Scholar] [CrossRef]
- Rep, M.; Dekker, H.L.; Vossen, J.H.; de Boer, A.D.; Houterman, P.M.; Speijer, D.; Back, J.; de Koster, C.G.; Cornelissen, B.J. Mass Spectrometric Identification of Isoforms of PR Proteins in Xylem Sap of Fungus-Infected Tomato. Plant Physiol. 2002, 130, 904–917. [Google Scholar] [CrossRef]
- Berrocal-Lobo, M.; Molina, A. Arabidopsis defense response against Fusarium oxysporum. Trends Plant Sci. 2008, 13, 145–150. [Google Scholar] [CrossRef]
- Castillejo, M.; Curto, M.; Fondevilla, S.; Rubiales, D.; Jorrín, J.V. Two-Dimensional Electrophoresis Based Proteomic Analysis of the Pea (Pisum sativum) in Response to Mycosphaerella pinodes. J. Agric. Food Chem. 2010, 58, 12822–12832. [Google Scholar] [CrossRef]
- Palomares-Rius, J.E.; Castillo, P.; Navas-Cortés, J.A.; Jiménez-Díaz, R.M.; Tena, M. A proteomic study of in-root interactions between chickpea pathogens: The root-knot nematode Meloidogyne artiellia and the soil-borne fungus Fusarium oxysporum f. sp. ciceris race. J. Proteom. 2011, 74, 2034–2051. [Google Scholar] [CrossRef]
- Yang, Y.; Shah, J.; Klessig, D.F. Signal perception and transduction in plant defense responses. Genes Dev. 1997, 11, 1621–1639. [Google Scholar] [CrossRef]
- De Ascensao, A.R.; Dubery, I.A. Panama disease: Cell wall reinforcement in banana roots in response to elic-itors from Fusarium oxysporum f. sp. cubense race four. Phytopathology 2000, 90, 1173–1180. [Google Scholar] [CrossRef] [PubMed]
- Sebastiani, M.S.; Bagnaresi, P.; Sestili, S.; Biselli, C.; Zechini, A.; Orrù, L.; Cattivelli, L.; Ficcadenti, N. Transcriptome Analysis of the Melon-Fusarium oxysporum f. sp. melonis Race 1.2 Pathosystem in Susceptible and Resistant Plants. Front. Plant Sci. 2017, 8, 362. [Google Scholar] [CrossRef] [PubMed]
- Fiehn, O. Metabolomics—the link between genotypes and phenotypes. Plant Mol. Biol. 2002, 48, 155–171. [Google Scholar] [CrossRef]
- Baharum, S.N.; Azizan, K.A. Metabolomics in systems biology. Adv. Exp. Med. Biol. 2018, 1102, 51–68. [Google Scholar] [CrossRef] [PubMed]
- Weckwerth, W. Unpredictability of metabolism—The key role of metabolomics science in combination with next-generation genome sequencing. Anal. Bioanal. Chem. 2011, 400, 1967–1978. [Google Scholar] [CrossRef]
- Pandey, M.K.; Roorkiwal, M.; Singh, V.K.; Ramalingam, A.; Kudapa, H.; Thudi, M.; Chitikineni, A.; Rathore, A.; Varshney, R.K. Emerging Genomic Tools for Legume Breeding: Current Status and Future Prospects. Front. Plant Sci. 2016, 7, 455. [Google Scholar] [CrossRef]
- Morkunas, I.; Ratajczak, L. The role of sugar signaling in plant defense responses against fungal pathogens. Acta Physiol. Plant. 2014, 36, 1607–1619. [Google Scholar] [CrossRef]
- Behmand, T.; Berger, J.; Elekcioğlu, H. Identification of resistance to Pratylenchus thornei, P. neglectus, P. penetrans and Ditylenchus dipsaci in Turkish accessions of five wild chickpea species (Cicer bijugum, C. echinospermum, C. pinnatifidum, C. reticulatum and C. turcicum). Phytoparasitica 2023, 51, 337–351. [Google Scholar] [CrossRef]
- Bani, M.; Cimmino, A.; Evidente, A.; Rubiales, D.; Rispai, N. Pisatin involvement in the variation of inhibition of Fusarium ox- ysporum f. sp. pisi spore germination by root exudates of Pisum spp. germplasm. Plant Pathol. 2018, 67, 1046–1054. [Google Scholar] [CrossRef]
- Verbruggen, N.; Hanikenne, M.; Clemens, S. A more complete picture of metal hyperaccumulation through next-generation sequencing technologies. Front. Plant Sci. 2013, 4, 199. [Google Scholar] [CrossRef]
- Shyam, C.; Tripathi, M.K.; Tiwari, S.; Tripathi, N.; Solanki, R.S.; Sapre, S.; Ahuja, A.; Tiwari, S. In vitro production of somaclones with decreased erucic acid content in Indian mustard [Brassica juncea (Linn.) Czern&Coss]. Plants 2021, 10, 1297. [Google Scholar] [CrossRef]
- Upadhyay, S.; Singh, A.K.; Tripathi, M.K.; Tiwari, S.; Tripathi, N.; Patel, R.P. In vitro selection for resistance against char-coal rot disease of soybean [Glycine max (L.) Merrill] caused by Macrophomina phaseolina (Tassi) Gold. Leg. Res. Int. J. 2020. [Google Scholar] [CrossRef]
- Eathington, S.R.; Crosbie, T.M.; Edwards, M.D.; Reiter, R.S.; Bull, J.K. Molecular markers in a commercial breeding program. Crop Sci. 2007, 47, S-154. [Google Scholar] [CrossRef]
- Eagles, H.A.; Bariana, H.S.; Ogbonnaya, F.C.; Rebetzke, G.J.; Hollamby, G.J.; Henry, R.J.; Henschke, P.H.; Carter, M. Implemen- tation of markers in Australian wheat breeding. Aust. J. Agric. Res. 2001, 52, 1349–1356. [Google Scholar] [CrossRef]
- Hayes, B.J.; Goddard, M.E. Prediction of total genetic value using genome-wide dense marker maps. Genetics 2001, 1, 1819–1829. [Google Scholar]
- Miedaner, T.; Boeven, A.L.G.-C.; Gaikpa, D.S.; Kistner, M.B.; Grote, C.P. Genomics-Assisted Breeding for Quantitative Disease Resistances in Small-Grain Cereals and Maize. Int. J. Mol. Sci. 2020, 21, 9717. [Google Scholar] [CrossRef]
- Tripathi, M.; Tripathi, N.; Tiwari, S.; Tiwari, G.; Mishra, N.; Bele, D.; Patel, R.; Sapre, S.; Tiwari, S. Optimization of Different Factors for Initiation of Somatic Embryogenesis in Suspension Cultures in Sandalwood (Santalum album L.). Horticulturae 2021, 7, 118. [Google Scholar] [CrossRef]
- Jhankare, A.; Tiwari, G.; Tripathi, M.K.; Pandey, G.N.; Patel, R.P.; Tiwari, S.; Baghel, B.S. Development of resistant lines against leaf blight disease of [(Withania somnifera (L.) Dunal.)] caused by Alternaria alternata through In vitro selection. Plant Cell Bio- technol. Mol. Biol. 2011, 12, 21–30. [Google Scholar]
- Tripathi, M.K.; Tiwari, S.; Khare, U.K. In vitro selection for resistance against purple blotch disease of on-ion (Allium cepa L.) caused by Alternaria porri. Biotechnology 2008, 7, 80–86. [Google Scholar]
- Mishra, N.; Tripathi, M.K.; Tiwari, S.; Tripathi, N.; Sapre, S.; Ahuja, A.; Tiwari, S. Cell suspension cul-ture and in vitro screening for drought tolerance in soybean using poly-ethylene glycol. Plants 2021, 10, 517. [Google Scholar] [CrossRef]
- Rai, M.K.; Kalia, R.K.; Singh, R.; Gangola, M.P.; Dhawan, A. Developing stress tolerant plants through in vitro selection—An overview of the recent progress. Environ. Exp. Bot. 2011, 71, 89–98. [Google Scholar] [CrossRef]
- Matsumoto, K.; Barbosa, M.L.; Souza, L.A.C.; Teixeira, J.B. Race 1 fusarium wilt tolerance on banana plants selected by fusaric acid. Euphytica 1995, 84, 67–71. [Google Scholar] [CrossRef]
- Gayatri, M.C.; Roopa Darshini, V.; Kavyashree, R. Selection of turmeric callus for tolerant to cul-ture filtrate of Pythium gram- inicolum and regeneration of plants. Plant Cell Tissue Organ Cult. 2005, 83, 33–40. [Google Scholar] [CrossRef]
- Ahuja, A.; Tripathi, M.K.; Tiwari, S.; Tripathi, N.; Tiwari, G.; Mishra, N.; Bhargav, S.; Tiwari, S. Recent Advancements on Callus and Cell Suspension Cultures: An Effectual Reserve for the Production of Pharmaceutically Significant Metabolites. Curr. Asp. Pharm. Res. 2021, 6, 96–111. [Google Scholar] [CrossRef]
- Tripathi, M.K.; Tiwari, S.; Tripathi, N.; Tiwari, G.; Bhatt, D.; Vibhute, M.; Gupta, N.; Mishra, N.; Parihar, P.; Singh, P.; et al. Plant Tissue Culture Techniques for Conservation of Biodiversity of Some Plants Appropriate for Propgation in Degraded and Tem- perate Areas. In Current Topics in Agricultural Sciences; B.P. International Publisher: Bhanjipur, India, 2021. [Google Scholar] [CrossRef]
- Singh, R.; Sindhu, A.; Singal, H.R. Biochemical basis of resistance in chickpea (Cicer anetinum L) against Fusarium wilt. Acta. Phytopathol. Entomol. Hung 2003, 38, 13–19. [Google Scholar] [CrossRef]
- Rao, S.; Padmaja, M. Selection of chickpea coll lines resistant to culture filtrate of Fusarium oxysporum fpcionri. Phytomorphology 2000, 50, 41–46. [Google Scholar]
- Hamid, K.; Strange, R.N. Phytotoicity of solanapyrones A and B produced by the chickpea pathogen Ascochyta rabiel (paas) L’abr, and the apparent metabolism of solanapyrone A by chickpea tissues. Physiol. Mol. Plant Pathol. 2000, 56, 235–244. [Google Scholar] [CrossRef]
- Tripathi, N.; Tripathi, M.K.; Tiwari, S.; Payasi, D.K. Molecular Breeding to Overcome Biotic Stresses in Soybean: Update. Plants 2022, 11, 1967. [Google Scholar] [CrossRef]
- Rockström, J.; Williams, J.; Daily, G.; Noble, A.; Matthews, N.; Gordon, L.; Wetterstrand, H.; DeClerck, F.; Shah, M.; Steduto, P.; et al. Sustainable intensification of agriculture for human prosperity and global sustainability. AMBIO 2016, 46, 4–17. [Google Scholar] [CrossRef]
- Ray, D.K.; Mueller, N.D.; West, P.C.; Foley, J.A. Yield trends are insufficient to double global crop production. PLoS ONE 2013, 8, e66428. [Google Scholar] [CrossRef]
- Lin, Z.; Cogan, N.O.I.; Pembleton, L.W.; Spangenberg, G.C.; Forster, J.W.; Hayes, B.J.; Daetwyler, H.D. Genetic gain and in- breeding from genomic selection in a simulated commercial breeding program for perennial ryegrass. Plant Genome 2016, 9, 1–12. [Google Scholar] [CrossRef]
- Samantara, K.; Bohra, A.; Mohapatra, S.R.; Prihatini, R.; Asibe, F.; Singh, L.; Reyes, V.P.; Tiwari, A.; Maurya, A.K.; Croser, J.S.; et al. Breeding more crops in less time: A perspective on speed breeding. Biology 2022, 11, 275. [Google Scholar] [CrossRef]
- Pandey, S.; Singh, A.; Parida, S.K.; Prasad, M. Combining speed breeding with traditional and genomics-assisted breeding for crop improvement. Plant Breed. 2022, 141, 301–313. [Google Scholar] [CrossRef]
- Lamichhane, S.; Thapa, S. Advances from Conventional to Modern Plant Breeding Methodologies. Plant Breed. Biotechnol. 2022, 10, 1–14. [Google Scholar] [CrossRef]
- Ren, J.; Wu, P.; Trampe, B.; Tian, X.; Lübberstedt, T.; Chen, S. Novel technologies in doubled haploid line development. Plant Biotechnol. J. 2017, 15, 1361–1370. [Google Scholar] [CrossRef]
- Wang, K.; Frame, B.; Ishida, Y.; Komari, T. Maize transformation. In Handbook of maize. Genetics and genomics; Bennetzen, K., Hake, S., Eds.; Springer Science + Business Media: New York, NY, USA, 2009; pp. 609–639. [Google Scholar]
- Rizal, G.; Karki, S.; Alcasid, M.; Montecillo, F.; Acebron, K.; Larazo, N.; Garcia, R.; Slamet-Loedin, I.H.; Quick, W.P. Shortening the breeding cycle of sorghum, a model crop for research. Crop Sci. 2019, 54, 520–529. [Google Scholar] [CrossRef]
- Gaur, P.M.; Srinivasan, S.; Gowda, C.L.L.; Bao, B.V. Rapid generation advancement in chickpea. J. SAT Agric. Res. 2007, 3, 1–3. [Google Scholar]
- Watson, A.; Ghosh, S.; Williams, M.J.; Cuddy, W.S.; Simmonds, J.; Rey, M.D.; Asyraf-Md-Hatta, M.; Hinchliffe, A.; Steed, A.; Reynolds, D.; et al. Speed breeding is a powerful tool to accelerate crop research and breeding. Nat. Plants 2018, 4, 23–29. [Google Scholar] [CrossRef]
- Hickey, L.T.; Hafeez, A.N.; Robinson, H.; Jackson, S.A.; Leal-Bertioli, S.C.M.; Tester, M.; Gao, C.; Godwin, I.D.; Hayes, B.J.; Wulff, B.B.H. Breeding crops to feed 10 billion. Nat. Biotechnol. 2019, 37, 744–754. [Google Scholar] [CrossRef]
- Saxena, K.; Saxena, R.K.; Varshney, R.K. Use of immature seed germination and single seed descent for rapid genetic gains in pigeonpea. Plant Breed. 2017, 136, 954–957. [Google Scholar] [CrossRef]
- Samineni, S.; Sen, M.; Sajja, S.B.; Gaur, P.M. Rapid generation advance (RGA) in chickpea to produce up to seven generations per year and enable speed breeding. Crop J. 2019, 8, 164–169. [Google Scholar] [CrossRef]
- Fikre, A.; Tulu, D. A Guide to Accelerated Breeding Cycle in Chickpea to Enhance Rate of Gain A Guide to Accelerated Breeding Cycle in Chickpea to Enhance Rate of Gain. Guide Man. 2019, 1–35. [Google Scholar] [CrossRef]
- Fikre, A.; Tulu, D.; Tesfaye, G.; Thudi, M.; Gaur, P.; Ojiewo, C.; Hickey, L.; Varshney, R.K. Rapid Generation Advance in Chickpea for Accelerated Breeding Gain in Ethiopia: What Speed Breeding Imply? Ethiop. J. Agric. Sci. 2021, 31, 1–10. [Google Scholar]


| Important Varieties/Donors | Country | Reference |
|---|---|---|
| Surutato-77, Sonora-80, UC-15, UC-27, and Gavilan | Mexico | [27] |
| BG-312, ICCVs 98505, 07105, 07111, 07305, 08113, and 93706, ICCVs 08123, 08125, 96858, 07118, 08124, 04514, 08323, and08117(moderately resistant) | India | [85] |
| WR 315, JG 315, CPS 1, JG 74, Avrodhi, and Phule G | India | [84] |
| ICCV 2,3,4,5 and ICC 11322, 14424, and 14433 (against race I) | India | [88] |
| Digvijay | India | [89] |
| ICC 14194, ICC 17109, and WR 315 | India | [90] |
| Three lines derived from MABC-based C 214 and WR 315 cross | India | [91] |
| ICCV 09118, ICCV 09113, ICCV 09115, ICCV 09308, ICCV 09314, ICCV 05527, ICCV 05528, and ICCV 96818 | India | [73] |
| Super Annigeri and improved JG74 (resistant against FOC4) | India | [92] |
| ICC 7537 resistant to all races (except race 4) | Ethiopia | [27] |
| FLIP 84-43C (against race 0), ILC-5411, FLIP 85-20C (against race 5), FLIP 85-29C, FLIP 85-30C, ILC-127 (against race 0), ILC-219 (against race 0), ILC-237, ILC-267, and ILC-513 (against race 0) | Santaella, Córdoba, Spain | [93] |
| Annigeri | India | [27] |
| ICC-7520 | Iran | [27] |
| Andom1 and Ayala | - | [63] |
| Rating | Wilt/Mortality (%) | Field Observation |
|---|---|---|
| 1 | 0% | No lesions visible |
| 2 | <10% | Few scattered lesions, usually seen after careful examination |
| 3 | 11–20% | Lesions and defoliation on some plants; little damage |
| 4 | 21–50% | Lesions very common and damaging; 25% plants killed |
| 5 | 51–80% | All plants with extensive lesions, causing defoliation and drying of branches; 50% plants killed |
| 6 | >81% | Lesions extensive on all plants; defoliation and drying of branches; more than 75% plants killed |
| Fusarium Race | Name of Population | QTLs | Marker Identified | Linkage Group | References |
|---|---|---|---|---|---|
| Race 1 Race 4 | C-104 × WR-315 | - | CS-27700, UBC-170550 (RAPD) | - | [140] |
| Race 3 | WR-315 × C-104 | FOC-3 | TA96 and TA27, TA196 (STMS) | - | [26] |
| Race 1 Race 4 | - | FOC-1 (syn. h (1)) and FOC-4 | CS27A (STS/SCAR) TA194 (STMS) | - | [61,138] |
| Race 5 | - | FOC-5 | TA59 and TA96 (SSR) | - | [174] |
| Race 2 | - | FOC-2 | TA96 and H3A12 (STMS) | - | |
| Race 4 Race 5 | C. arietinum × C. reticulatum | - | STM S and a SCAR | - | [138] |
| Race 1 | F9 | FOC-1 | H3A12, TA110 (STMS) | - | [61] |
| Race 0 | CA 2139 × JG 62 | FOC01/FOC01 | OPJ20(600) (RAPD) TR59 (STMS) | LG3 | [138] |
| Race 0 | CA 2139 × JG 62 | FOC02/FOC02 | TA59 (STMS) | LG2 | [174] |
| Race 1A | C 214 × WR 315 | FW-Q-APR-6-1 (FOC-1) and FW-Q-APR-6-2 (FOC-1) | CaM1402 and CaM1101 (flanking) CaM1125-TA22 | LG6 | [176] |
| Race 5 | - | FOC-5 | TA59 (STMS) | LG2 | [59] |
| Race 1 | JG 62 × WR 315 | - | TA27-TA59 (STMS) | LG2 | [4] |
| Race 1 Race 3 | C 214 × WR 315 | FOC-1 and FOC-3 | GA16, TA110, and TS82 | LG2 | [134] |
| Race 1 | JG 62 × ICC V05530 | 3QTL (race 1), FW-Q-APR-2-1 FW-Q-APR-4-1 FW-Q-APR-6-1 | TR19 and H2B061, TA132 and TA46 (STMS) | CaLG02, CaLG04, and CaLG06 | [178] |
| Race 3 | JG 62 × ICC V05530 | 2QTLs (race 3) FW-Q-APR-2-1 and FW-Q-APR-4-1 | CKAM1256 and TS72 | CaLG02 and CaLG04 | [178] |
| Race 0 | CA 2156 × JG 62 | FOC01/FOC01 | H2I20 and TS43 (STMS) | LG5 | [58] |
| Race 5 | WR 315 × ILC 3279 | FOC-5 | TA59, CaGM07922, and SNPs | LG2 | [179] |
| Race 4 | Annigeri1 × WR-315 | FOC-4 | TA59, TA96, TR19, and TA27 | LG2 | [164] |
| Race 4 | JG 74 × WR 315 | FOC-4 | GA16andTA96 | [164] | |
| Race 5 | - | FOC-5/FOC-5 | TA27 and TA59 TA96 CS27700 (RAPD) UBC170550 (RAPD) | LG2 | [57,138] |
| Race 5 | - | FOC-5/FOC-5 | ECAMCTA07 OP-M20-21045 OP-M20-31103 | LG2 | [164] |
| Genotype Used in Study | Platform/Technology | Differentially Expressed Genes (DEGs)/Candidate Gene | Study Based on | References |
|---|---|---|---|---|
| WR 315 and JG 62 | cDNA-RAPD and cDNA-AFLP | 273 DEGs related to stress response, gamma-glutamyl-cysteine synthetase, and NBS-LRR | Race 1 | [182] |
| RNA blot analysis | 6272 ESTs belonged to stress-responsive genes and cell signaling, transcription, RNA processing, modification, cellular transport, homeostasis, and hormone response-related genes | Race 1 | [197] | |
| Suppression subtractive hybridization | 162 ESTs belonged to genes responsible for defense signaling pathways, energy metabolism, cell rescue, and superoxide dismutase | Race 4 | [196] | |
| qPCR, Microarray analysis | Stress-responsive and other defense-associated genes, including aquaporin, ATP synthase, immunity-associated genes, cystatin and DnaJ, pectinesterase and xyloglucosyl transferase, actin- and profilin-like genes, cytochrome P450, and peroxidase | Race 1 | [197] | |
| qPCR | Transporter gene, transporter like gene, redox regulatory respiratory burst oxidase homolog F (RBOHF), thioredoxin 3 (TRX3), cationic peroxidase 3 (OCP3), flavodoxin-like quinone reductase 1 (FQR1), iron superoxide dismutase 1, NADH cytochrome b5 reductase (CBR), Fe (II) oxidoreductase 7 (FRO7), genes related to intracellular transportation ABC transporter-like gene, polyol transporter gene, translocase, heavy metal transporter (detoxifying protein) (FRS6), bZIP, homeodomain leucine zipper, MYB, helix loop helix, zinc finger (CCHC type), heat shock family protein, sucrose synthase (SUS4), b-amylase (BAM1), serine threonine kinase (CDKB1.1), and vacuolar ATPase (TUF) | [198] | ||
| Expression analysis | NBS-LRR and WRKY genes | [199] | ||
| Digvijay and JG 62 | qPCR | Stress-responsive genes | [200] | |
| qRT-PCR and LongSAGE | 3816 DEGs and G protein b subunit gene lignification, hormonal homeostasis, plant defense signaling, ROS homeostasis, and R-gene mediated defense | [201,202] | ||
| qRT-PCR | 5 DEGs related to stress-responsive category, glycosyltransferase gene, GroEs2, 60srp, and Betvi E | Races 1, 2, and 4 | [203] | |
| ICC4958 | Illumina (NGS) and Poly(A)-based qRT-PCR | 122 conserved miRNAs, 59 novel miRNAs, and defense gene encoding Toll/Interleukin-1 receptor–nucleotide binding site leucine-rich repeats miR2111 targets a Kelch repeat-containing F-box protein | [204] | |
| NILs—RIP8-94-5/RIP8-94-11 | qPCR | 22 potential defense-related genes encoding a MADS-box transcription factor, and TMV resistance protein | Race 5 | [205] |
| WR315 and BG256 | Sequencing (Roche 454 GS FLX system) | 202 DEGs related to polyubiquitin, chlorophyll a-b binding protein, ferredoxin-NADP, translation factor sui1, carbonic anhydrase, ribulose bisphosphate carboxylase, oxygen evolving enhancer, elongation factor 1-alpha, and post-translational modification genes | [206] | |
| JG 62, WR 315 and JAKI9218 | qRT-PCR | 6 DEGs, including transcription factors such as extracellular calcium-sensing receptor, Nitric oxide reductase, growth hormone-releasing hormone receptor, Cytochrome C oxidase Cbb-3 type subunit I, Hydroxynitrite lyase, Tir chaperone, and ionotropic glutamate receptor | Races 2 and 4 | [207] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yadav, R.K.; Tripathi, M.K.; Tiwari, S.; Tripathi, N.; Asati, R.; Patel, V.; Sikarwar, R.S.; Payasi, D.K. Breeding and Genomic Approaches towards Development of Fusarium Wilt Resistance in Chickpea. Life 2023, 13, 988. https://doi.org/10.3390/life13040988
Yadav RK, Tripathi MK, Tiwari S, Tripathi N, Asati R, Patel V, Sikarwar RS, Payasi DK. Breeding and Genomic Approaches towards Development of Fusarium Wilt Resistance in Chickpea. Life. 2023; 13(4):988. https://doi.org/10.3390/life13040988
Chicago/Turabian StyleYadav, Rakesh Kumar, Manoj Kumar Tripathi, Sushma Tiwari, Niraj Tripathi, Ruchi Asati, Vinod Patel, R. S. Sikarwar, and Devendra K. Payasi. 2023. "Breeding and Genomic Approaches towards Development of Fusarium Wilt Resistance in Chickpea" Life 13, no. 4: 988. https://doi.org/10.3390/life13040988
APA StyleYadav, R. K., Tripathi, M. K., Tiwari, S., Tripathi, N., Asati, R., Patel, V., Sikarwar, R. S., & Payasi, D. K. (2023). Breeding and Genomic Approaches towards Development of Fusarium Wilt Resistance in Chickpea. Life, 13(4), 988. https://doi.org/10.3390/life13040988







