Activity and Function of the PRMT8 Protein Arginine Methyltransferase in Neurons
Abstract
1. Introduction
2. PRMT8 Characteristics
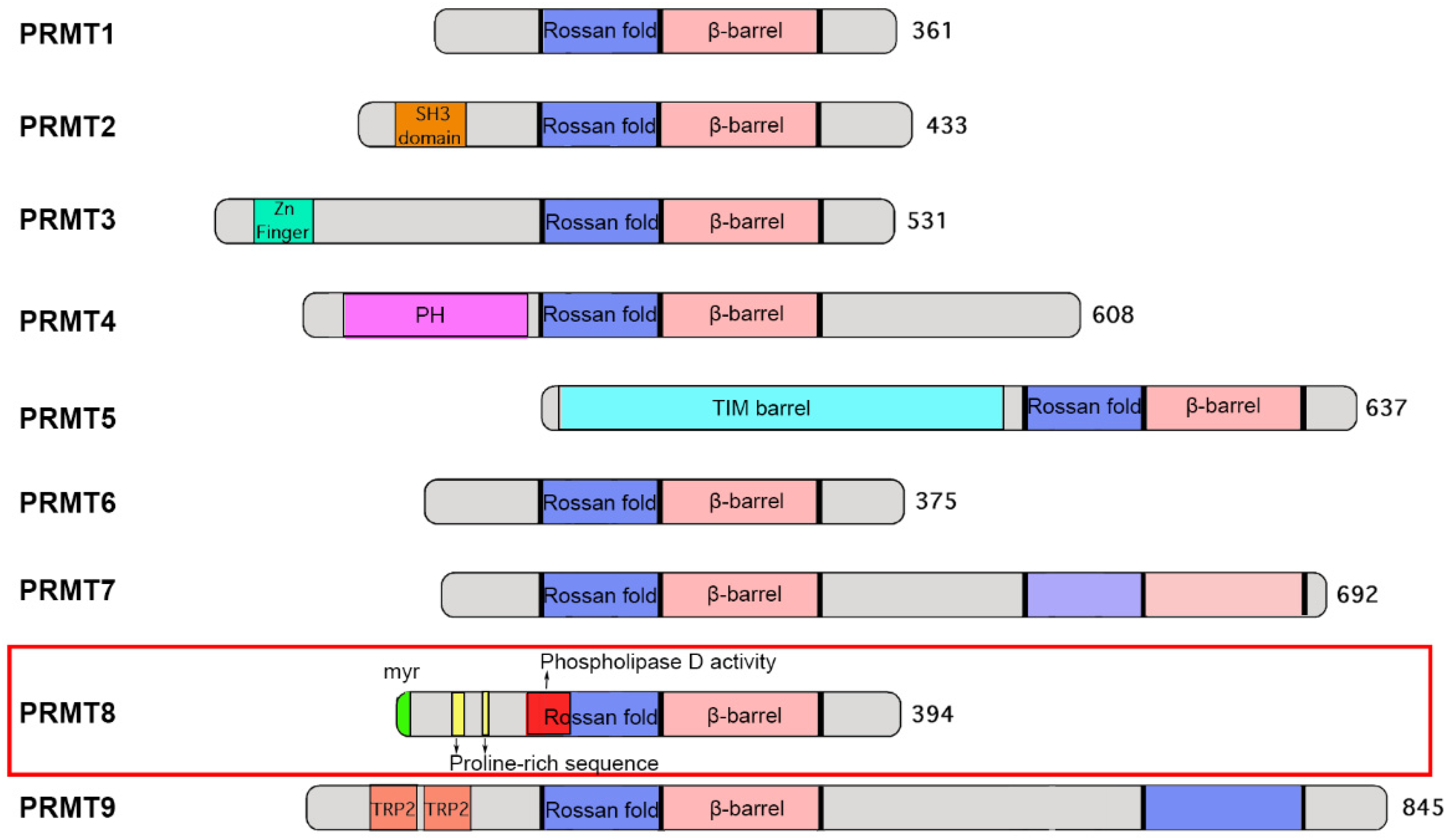
2.1. Domain and Structure
2.2. Expression Profile and Subcellular Localization
3. Enzymatic Properties of PRMT8
3.1. Regulation and Crosstalk with Other PRMTs
3.2. Substrates of PRMT8
4. Neuronal Functions of PRMT8
4.1. PRMT8 Functions as a Phospholipase to Regulate Purkinje Cell Dendritic Arborization
4.2. PRMT8 Regulates Synaptic Plasticity and Cognitive Functions
4.3. PRMT8 Regulates the Maturation of Synapse and Neural Circuit during Brain Development
4.4. PRMT8 and Neurological Diseases
5. Conclusions and Outlook
Author Contributions
Funding
Informed Consent Statement
Conflicts of Interest
References
- Paik, W.K.; Paik, D.C.; Kim, S. Historical review: The field of protein methylation. Trends Biochem. Sci. 2007, 32, 146–152. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Sayegh, J.; Daniel, J.; Clarke, S.; Bedford, M.T. PRMT8, a new membrane-bound tissue- specific member of the protein arginine methyltransferase family. J. Biol. Chem. 2005, 280, 32890–32896. [Google Scholar] [CrossRef] [PubMed]
- Hwang, J.W.; Cho, Y.; Bae, G.U.; Kim, S.N.; Kim, Y.K. Protein arginine methyltransferases: Promising targets for cancer therapy. Exp. Mol. Med. 2021, 53, 788–808. [Google Scholar] [CrossRef] [PubMed]
- Bedford, M.T.; Clarke, S.G. Protein arginine methylation in mammals: Who, what, and why. Mol. Cell 2009, 33, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Qualmann, B.; Kessels, M.M. The Role of Protein Arginine Methylation as Post-Translational Modification on Actin Cytoskeletal Components in Neuronal Structure and Function. Cells 2021, 10, 1079. [Google Scholar] [CrossRef] [PubMed]
- Swiercz, R.; Person, M.D.; Bedford, M.T. Ribosomal protein S2 is a substrate for mammalian PRMT3 (protein arginine methyltransferase 3). Biochem. J. 2005, 386, 85–91. [Google Scholar] [CrossRef]
- Frankel, A.; Clarke, S. PRMT3 is a distinct member of the protein arginine N- methyltransferase family. conferral of substrate specificity by a zinc-finger domain. J. Biol. Chem. 2000, 275, 32974–32982. [Google Scholar] [CrossRef] [PubMed]
- Antonysamy, S.; Bonday, Z.; Campbell, R.M.; Doyle, B.; Druzina, Z.; Gheyi, T.; Han, B.; Jungheim, L.N.; Qian, Y.; Rauch, C.; et al. Crystal structure of the human PRMT5:MEP50 complex. Proc. Natl. Acad. Sci. USA 2012, 109, 17960–17965. [Google Scholar] [CrossRef]
- Ho, M.-C.; Wilczek, C.; Bonanno, J.B.; Xing, L.; Seznec, J.; Matsui, T.; Carter, L.G.; Onikubo, T.; Kumar, P.R.; Chan, M.K.; et al. Structure of the arginine methyltransferase PRMT5-MEP50 reveals a mechanism for substrate specificity. PLoS ONE 2013, 8, 57008–57023. [Google Scholar] [CrossRef]
- Kim, J.D.; Park, K.E.; Ishida, J.; Kako, K.; Hamada, J.; Kani, S.; Takeuchi, M.; Namiki, K.; Fukui, H.; Fukuhara, S.; et al. PRMT8 as a phospholipase regulates Purkinje cell dendritic arborization and motor coordination. Sci. Adv. 2015, 1, e1500615. [Google Scholar] [CrossRef]
- Sayegh, J.; Webb, K.; Cheng, D.; Bedford, M.T.; Clarke, S.G. Regulation of protein arginine methyltransferase 8 (PRMT8) activity by its N-terminal domain. J. Biol. Chem. 2007, 282, 36444–36453. [Google Scholar] [CrossRef] [PubMed]
- Kousaka, A.; Mori, Y.; Koyama, Y.; Taneda, T.; Miyata, S.; Tohyama, M. The distribution and characterization of endogenous protein arginine N-methyltransferase 8 in mouse CNS. Neuroscience 2009, 163, 1146–1157. [Google Scholar] [CrossRef] [PubMed]
- Park, S.W.; Jun, Y.W.; Choi, H.E.; Lee, J.A.; Jang, D.J. Deciphering the molecular mechanisms underlying the plasma membrane targeting of PRMT8. BMB Rep. 2019, 52, 601–606. [Google Scholar] [CrossRef]
- Penney, J.; Seo, J.; Kritskiy, O. Loss of Protein Arginine Methyltransferase 8 Alters Synapse Composition and Function, Resulting in Behavioral Defects. J. Neurosci. 2017, 37, 8655–8666. [Google Scholar] [CrossRef] [PubMed]
- Lo, L.H.-Y.; Dong, R.; Lyu, Q.; Lai, K.O. The Protein Arginine Methyltransferase PRMT8 and Substrate G3BP1 Control Rac1-PAK1 Signaling and Actin Cytoskeleton for Dendritic Spine Maturation. Cell Rep. 2020, 31, 107744. [Google Scholar] [CrossRef]
- Samuel, S.F.; Barry, A.; Greenman, J.; Beltran-Alvarez, P. Arginine methylation: The promise of a ‘silver bullet’ for brain tumours? Amino Acids 2021, 53, 489–506. [Google Scholar] [CrossRef]
- Simandi, Z.; Czipa, E.; Horvath, A.; Koszeghy, A.; Bordas, C.; Póliska, S.; Juhász, I.; Imre, L.; Szabó, G.; Dezso, B.; et al. PRMT1 and PRMT8 regulate retinoic acid-dependent neuronal differentiation with implications to neuropathology. Stem Cells 2015, 33, 726–741. [Google Scholar] [CrossRef]
- Yang, Y.; Bedford, M.T. Protein arginine methyltransferases and cancer. Nat. Rev. Cancer 2013, 13, 37–50. [Google Scholar] [CrossRef] [PubMed]
- Hernandez, S.J.; Dolivo, D.M.; Dominko, T. PRMT8 demonstrates variant-specific expression in cancer cells and correlates with patient survival in breast, ovarian and gastric cancer. Oncol. Lett. 2017, 13, 1983–1989. [Google Scholar] [CrossRef]
- Lin, H.; Wang, B.; Yu, J.; Wang, J.; Li, Q.; Cao, B. Protein arginine methyltransferase 8 gene enhances the colon cancer stem cell (CSC) function by upregulating the pluripotency transcription factor. J. Cancer 2018, 9, 1394–1402. [Google Scholar] [CrossRef]
- Frankel, A.; Yadav, N.; Lee, J.; Branscombe, T.L.; Clarke, S.; Bedford, M.T. The novel human protein arginine N-methyltransferase PRMT6 is a nuclear enzyme displaying unique substrate specificity. J. Biol. Chem. 2002, 277, 3537–3543. [Google Scholar] [CrossRef] [PubMed]
- Dillon, M.B.; Rust, H.L.; Thompson, P.R.; Mowen, K.A. Automethylation of protein arginine methyltransferase 8 (PRMT8) regulates activity by impeding S-adenosylmethionine sensitivity. J. Biol. Chem. 2013, 288, 27872–27880. [Google Scholar] [CrossRef] [PubMed]
- Lin, W.J.; Gary, J.D.; Yang, M.C.; Clarke, S.; Herschman, H.R. The mammalian immediate-early TIS21 protein and the leukemia-associated BTG1 protein interact with a protein-arginine N- methyltransferase. J. Biol. Chem. 1996, 271, 15034–15044. [Google Scholar] [CrossRef] [PubMed]
- Singh, V.; Miranda, T.B.; Jiang, W.; Frankel, A.; Roemer, M.E.; Robb, V.A.; Gutmann, D.H.; Herschman, H.R.; Clarke, S.; Newsham, I.F. DAL-1/4.1B tumor suppressor interacts with protein arginine N-methyltransferase 3 (PRMT3) and inhibits its ability to methylate substrates in vitro and in vivo. Oncogene 2004, 23, 7761–7771. [Google Scholar] [CrossRef]
- Hu, H.; Qian, K.; Ho, M.C.; Zheng, Y.G. Small Molecule Inhibitors of Protein Arginine Methyltransferases. Expert Opin. Investig. Drugs 2016, 25, 335–358. [Google Scholar] [CrossRef]
- Kim, J.D.; Kako, K.; Kakiuchi, M.; Park, G.G.; Fukamizu, A. EWS is a substrate of type I protein arginine methyltransferase, PRMT8. Int. J. Mol. Med. 2008, 22, 309–315. [Google Scholar] [CrossRef]
- Burd, C.G.; Dreyfuss, G. Conserved structures and diversity of functions of RNA-binding proteins. Science 1994, 265, 615–621. [Google Scholar] [CrossRef]
- Najbauer, J.; Johnson, B.A.; Young, A.L.; Aswad, D.W. Peptides with sequences similar to glycine, arginine-rich motifs in proteins interacting with RNA are efficiently recognized by methyltransferase(s) modifying arginine in numerous proteins. J. Biol. Chem. 1993, 268, 10501–10509. [Google Scholar] [CrossRef]
- Gary, J.D.; Clarke, S. RNA and protein interactions modulated by protein arginine methylation. Prog. Nucl. Acid Res. Mol. Biol. 1998, 61, 65–131. [Google Scholar]
- Lee, W.C.; Lin, W.L.; Matsui, T.; Chen, E.S.; Wei, T.Y.; Lin, W.H.; Hu, H.; Zheng, Y.G.; Tsai, M.D.; Ho, M.C. Protein Arginine Methyltransferase 8: Tetrameric Structure and Protein Substrate Specificity. Biochemistry 2015, 54, 7514–7523. [Google Scholar] [CrossRef]
- Baek, J.H.; Rubinstein, M.; Scheuer, T.; Trimmer, J.S. Reciprocal changes in phosphorylation and methylation of mammalian brain sodium channels in response to seizures. J. Biol. Chem. 2014, 289, 15363–15373. [Google Scholar] [CrossRef]
- Tsai, W.C.; Gayatri, S.; Reineke, L.C.; Sbardella, G.; Bedford, M.T.; Lloyd, R.E. Arginine Demethylation of G3BP1 Promotes Stress Granule Assembly. J. Biol. Chem. 2016, 291, 22671–22685. [Google Scholar] [CrossRef]
- Carter, J.M.; Demizieux, L.; Campenot, R.B. Phosphatidylcholine Biosynthesis via CTP: Phosphocholine Cytidylyltransferase β2 Facilitates Neurite Outgrowth and Branching. J. Biol. Chem. 2008, 283, 202–212. [Google Scholar] [CrossRef]
- Gould, R.M.; Connell, F.; Spivack, W. Phospholipid Metabolism in Mouse Sciatic Nerve In Vivo. J. Neurochem. 1987, 48, 853–859. [Google Scholar] [CrossRef] [PubMed]
- Kanaho, Y.; Funakoshi, Y.; Hasegawa, H. Phospholipase D signalling and its involvement in neurite outgrowth. Biochim. Biophys. Acta BBA Mol. Cell Biol. Lipids 2009, 1791, 898–904. [Google Scholar] [CrossRef] [PubMed]
- Vance, J.E.; de Chaves, E.P.; Campenot, R.B.; Vance, D.E. Role of axons in membrane phospholipid synthesis in rat sympathetic neurons. Neurobiol. Aging 1995, 16, 493–498. [Google Scholar] [CrossRef]
- Picciotto, M.R.; Higley, M.J.; Mineur, Y.S. Acetylcholine as a neuromodulator: Cholinergic signaling shapes nervous system function and behavior. Neuron 2012, 76, 116–129. [Google Scholar] [CrossRef] [PubMed]
- Ammar, M.-R.; Kassas, N.; Bader, M.-F.; Vitale, N. Phosphatidic acid in neuronal development: A node for membrane and cytoskeleton rearrangements. Biochimie 2014, 107, 51–57. [Google Scholar] [CrossRef]
- Selvy, P.E.; Lavieri, R.R.; Lindsley, C.W.; Brown, H.A. Phospholipase D—Enzymology, functionality, and chemical modulation. Chem. Rev. 2011, 111, 6064–6119. [Google Scholar] [CrossRef] [PubMed]
- Ponting, C.P.; Kerr, I.D. A novel family of phospholipase D homologues that includes phospholipid synthases and putative endonucleases: Identification of duplicated repeats and potential active site residues. Protein Sci. 1996, 5, 914–922. [Google Scholar] [CrossRef]
- Park, K.-E.; Kim, J.-D.; Nagashima, Y.; Kako, K.; Daitoku, H.; Matsui, M.; Park, G.G.; Fukamizu, A. Detection of choline and phosphatidic acid (PA) catalyzed by phospholipase D (PLD) using MALDI-QIT-TOF/MS with 9-aminoacridine matrix. Biosci. Biotechnol. Biochem. 2014, 78, 981–988. [Google Scholar] [CrossRef]
- Lee, K.J.; Jung, J.G.; Arii, T.; Imoto, K.; Rhyu, I.J. Morphological changes in dendritic spines of Purkinje cells associated with motor learning. Neurobiol. Learn. Mem. 2007, 88, 445–450. [Google Scholar] [CrossRef] [PubMed]
- Cerminara, N.L.; Lang, E.J.; Sillitoe, R.V.; Apps, R. Redefining the cerebellar cortex as an assembly of non-uniform Purkinje cell microcircuits. Nat. Rev. Neurosci. 2005, 16, 79–93. [Google Scholar] [CrossRef] [PubMed]
- Jan, Y.-N.; Jan, L.Y. The Control of Dendrite Development. Neuron 2003, 40, 229–242. [Google Scholar] [CrossRef]
- Burkhardt, U.; Stegner, D.; Hattingen, E.; Beyer, S.; Nieswandt, B.; Klein, J. Impaired brain development and reduced cognitive function in phospholipase D-deficient mice. Neurosci. Lett. 2014, 572, 48–52. [Google Scholar] [CrossRef]
- Cai, D.; Zhong, M.; Wang, R.; Netzer, W.J.; Shields, D.; Zheng, H.; Sisodia, S.S.; Foster, D.A.; Gorelick, F.S.; Xu, H.; et al. Phospholipase D1 corrects impaired βAPP trafficking and neurite outgrowth in familial Alzheimer’s disease-linked presenilin-1 mutant neurons. Proc. Natl. Acad. Sci. USA 2006, 103, 1936–1940. [Google Scholar] [CrossRef]
- Lin, Y.; Tsai, Y.-J.; Liu, Y.-F.; Cheng, Y.-C.; Hung, C.-M.; Lee, Y.-J.; Pan, H.; Li, C. The Critical Role of Protein Arginine Methyltransferase prmt8 in Zebrafish Embryonic and Neural Development Is Non-Redundant with Its Paralogue prmt1. PLoS ONE 2013, 8, e55221. [Google Scholar] [CrossRef]
- Morris, R.G.M.; Anderson, E.; Lynch, G.S.; Baudry, M. Selective impairment of learning and blockade of long-term potentiation by an N-methyl-D-aspartate receptor antagonist, AP5. Nature 1986, 319, 774–776. [Google Scholar] [CrossRef]
- Tsien, J.Z.; Huerta, P.T.; Tonegawa, S. The Essential Role of Hippocampal CA1 NMDA Receptor–Dependent Synaptic Plasticity in Spatial Memory. Cell 1996, 87, 1327–1338. [Google Scholar] [CrossRef]
- Akazawa, C.; Shigemoto, R.; Bessho, Y.; Nakanishi, S.; Mizuno, N. Differential expression of five N-methyl-D-aspartate receptor subunit mRNAs in the cerebellum of developing and adult rats. J. Comp. Neurol. 1994, 347, 150–160. [Google Scholar] [CrossRef]
- Monyer, H.; Burnashev, N.; Laurie, D.J.; Sakmann, B.; Seeburg, P.H. Developmental and regional expression in the rat brain and functional properties of four NMDA receptors. Neuron 1994, 12, 529–540. [Google Scholar] [CrossRef]
- Paoletti, P.; Bellone, C.; Zhou, Q. NMDA receptor subunit diversity: Impact on receptor properties, synaptic plasticity and disease. Nat. Rev. Neurosci. 2013, 14, 383–400. [Google Scholar] [CrossRef] [PubMed]
- Baez, M.V.; Oberholzer, M.V.; Cercato, M.C.; Snitcofsky, M.; Aguirre, A.I.; Jerusalinsky, D.A. NMDA receptor subunits in the adult rat hippocampus undergo similar changes after 5 minutes in an open field and after LTP induction. PLoS ONE 2013, 8, e55244. [Google Scholar] [CrossRef] [PubMed]
- Bannerman, D.M.; Niewoehner, B.; Lyon, L.; Romberg, C.; Schmitt, W.B.; Taylor, A.; Sanderson, D.J.; Cottam, J.; Sprengel, R.; Seeburg, P.H.; et al. NMDA receptor subunit NR2A is required for rapidly acquired spatial working memory but not incremental spatial reference memory. J. Neurosci. 2008, 28, 3623–3630. [Google Scholar] [CrossRef]
- Zhang, X.-M.; Yan, X.-Y.; Zhang, B.; Yang, Q.; Ye, M.; Cao, W.; Qiang, W.-B.; Zhu, L.-J.; Du, Y.-L.; Xu, X.-X.; et al. Activity-induced synaptic delivery of the GluN2A-containing NMDA receptor is dependent on endoplasmic reticulum chaperone Bip and involved in fear memory. Cell Res. 2015, 25, 818–836. [Google Scholar] [CrossRef]
- Lee, P.K.M.; Goh, W.W.B.; Sng, J.C.G. Network-based characterization of the synaptic proteome reveals that removal of epigenetic regulator Prmt8 restricts proteins associated with synaptic maturation. J. Neurochem. 2017, 140, 613–628. [Google Scholar] [CrossRef]
- Bedford, M.T.; Richard, S. Arginine methylation an emerging regulator of protein function. Mol. Cell 2005, 18, 263–272. [Google Scholar] [CrossRef]
- Dong, X.; Weng, Z. The correlation between histone modifications and gene expression. Epigenomics 2013, 5, 113–116. [Google Scholar] [CrossRef]
- Poulard, C.; Corbo, L.; Le Romancer, M. Protein arginine methylation/demethylation and cancer. Oncotarget 2016, 7, 67532–67550. [Google Scholar] [CrossRef]
- Cajigas, I.J.; Tushev, G.; Will, T.J.; Dieck, S.T.; Fuerst, N.; Schuman, E.M. The local transcriptome in the synaptic neuropil revealed by deep sequencing and high-resolution imaging. Neuron 2012, 74, 453–466. [Google Scholar] [CrossRef]
- Fiala, J.C.; Feinberg, M.; Popov, V.; Harris, K.M. Synaptogenesis Via Dendritic Filopodia in Developing Hippocampal Area CA1. J. Neurosci. 1998, 18, 8900–8911. [Google Scholar] [CrossRef] [PubMed]
- Bourne, J.N.; Harris, K.M. Balancing structure and function at hippocampal dendritic spines. Annu. Rev. Neurosci. 2008, 31, 47–67. [Google Scholar] [CrossRef] [PubMed]
- Dansie, L.; Phommahaxay, K.; Okusanya, A.; Uwadia, J.; Huang, M.; Rotschafer, S.; Razak, K.; Ethell, D.; Ethell, I. Long-lasting effects of minocycline on behavior in young but not adult Fragile X mice. Neuroscience 2013, 246, 186–198. [Google Scholar] [CrossRef] [PubMed]
- Hutsler, J.J.; Zhang, H. Increased dendritic spine densities on cortical projection neurons in autism spectrum disorders. Brain Res. 2010, 1309, 83–94. [Google Scholar] [CrossRef]
- Wu, H.; Cottingham, C.; Chen, L.; Wang, H.; Che, P.; Liu, K.; Wang, Q. Age-dependent differential regulation of anxiety- and depression-related behaviors by neurabin and spinophilin. PLoS ONE 2017, 12, e0180638. [Google Scholar] [CrossRef] [PubMed]
- Cheadle, L.; Biederer, T. The novel synaptogenic protein Farp1 links postsynaptic cytoskeletal dynamics and transsynaptic organization. J. Cell Biol. 2012, 199, 985–1001. [Google Scholar] [CrossRef] [PubMed]
- Hotulainen, P.; Llano, O.; Smirnov, S.; Tanhuanpää, K.; Faix, J.; Rivera, C.; Lappalainen, P. Defining mechanisms of actin polymerization and depolymerization during dendritic spine morphogenesis. J Cell Biol. 2009, 185, 323–339. [Google Scholar] [CrossRef]
- Matsuki, H.; Takahashi, M.; Higuchi, M.; Makokha, G.N.; Oie, M.; Fujii, M. Both G3BP1 and G3BP2 contribute to stress granule formation. Genes Cells 2013, 18, 135–146. [Google Scholar] [CrossRef]
- Martin, S.; Bellora, N.; González-Vallinas, J.; Irimia, M.; Chebli, K.; de Toledo, M.; Raabe, M.; Eyras, E.; Urlaub, H.; Blencowe, B.J.; et al. Preferential binding of a stable G3BP ribonucleoprotein complex to intron-retaining transcripts in mouse brain and modulation of their expression in the cerebellum. J. Neurochem. 2016, 139, 349–368. [Google Scholar] [CrossRef]
- Santini, E.; Huynh, T.N.; Longo, F.; Koo, S.Y.; Mojica, E.; D’Andrea, L.; Bagni, C.; Klann, E. Reducing eIF4E-eIF4G interactions restores the balance between protein synthesis and actin dynamics in fragile X syndrome model mice. Sci. Signal. 2017, 10. [Google Scholar] [CrossRef]
- Guo, A.; Gu, H.; Zhou, J.; Mulhern, D.; Wang, Y.; Lee, K.A.; Yang, V.; Aguiar, M.; Kornhauser, J.; Jia, X.; et al. Immunoaffinity Enrichment and Mass Spectrometry Analysis of Protein Methylation. Mol. Cell. Proteom. 2014, 13, 372–387. [Google Scholar] [CrossRef] [PubMed]
- Lim, C.S.; Alkon, D.L. Inhibition of coactivator-associated arginine methyltransferase 1 modulates dendritic arborization and spine maturation of cultured hippocampal neurons. J. Biol. Chem. 2017, 292, 6402–6413. [Google Scholar] [CrossRef] [PubMed]
- Miyata, S.; Mori, Y.; Tohyama, M. PRMT3 is essential for dendritic spine maturation in rat hippocampal neurons. Brain Res. 2010, 1352, 11–20. [Google Scholar] [CrossRef]
- Hou, W.; Nemitz, S.; Schopper, S.; Nielsen, M.; Kessels, M.M.; Qualmann, B. Arginine Methylation by PRMT2 Controls the Functions of the Actin Nucleator Cobl. Dev. Cell 2018, 45, 262–275. [Google Scholar] [CrossRef]
- Zhou, H.; Mangelsdorf, M.; Liu, J.; Zhu, L.; Wu, J.Y. RNA-binding proteins in neurological diseases. Sci. China Life Sci. 2014, 57, 432–444. [Google Scholar] [CrossRef]
- Kabashi, E.; Valdmanis, P.; Dion, P.; Spiegelman, D.; McConkey, B.J.; Velde, C.V.; Bouchard, J.-P.; Lacomblez, L.; Pochigaeva, K.; Salachas, F.; et al. TARDBP mutations in individuals with sporadic and familial amyotrophic lateral sclerosis. Nat. Genet. 2008, 40, 572–574. [Google Scholar] [CrossRef]
- Vance, C.; Rogelj, B.; Hortobágyi, T.; De Vos, K.J.; Nishimura, A.L.; Sreedharan, J.; Hu, X.; Smith, B.; Ruddy, D.; Wright, P.; et al. Mutations in FUS, an RNA Processing Protein, Cause Familial Amyotrophic Lateral Sclerosis Type 6. Science 2009, 323, 1208–1211. [Google Scholar] [CrossRef] [PubMed]
- Scaramuzzino, C.; Monaghan, J.; Milioto, C.; Lanson, N.A., Jr.; Maltare, A.; Aggarwal, T.; Casci, I.; Fackelmayer, F.O.; Pennuto, M.; Pandey, U.B. Protein arginine methyltransferase 1 and 8 interact with FUS to modify its sub-cellular distribution and toxicity in vitro and in vivo. PLoS ONE 2013, 8, e61576. [Google Scholar] [CrossRef]
- Simandi, Z.; Pajer, K.; Karolyi, K.; Sieler, T.; Jiang, L.-L.; Kolostyak, Z.; Sari, Z.; Fekecs, Z.; Pap, A.; Patsalos, A.; et al. Arginine Methyltransferase PRMT8 Provides Cellular Stress Tolerance in Aging Motoneurons. J. Neurosci. 2018, 38, 7683–7700. [Google Scholar] [CrossRef] [PubMed]
- Shaw, P.J. Molecular and cellular pathways of neurodegeneration in motor neurone disease. J. Neurol. Neurosurg. Psychiatry 2005, 76, 1046–1057. [Google Scholar] [CrossRef] [PubMed]
- Rincic, M.; Rados, M.; Kopic, J.; Krsnik, Z.; Liehr, T. 7p21.3 Together With a 12p13.32 Deletion in a Patient with Microcephaly—Does 12p13.32 Locus Possibly Comprises a Candidate Gene Region for Microcephaly? Front. Mol. Neurosci. 2021, 14, 2. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Fok, A.H.K.; Fan, R.; Kwan, P.Y.; Chan, H.L.; Lo, L.H.; Chan, Y.S.; Yung, W.H.; Huang, J.; Lai, C.S.W.; et al. Specific depletion of the motor protein KIF5B leads to deficits in dendritic transport, synaptic plasticity and memory. eLife 2020, 9, e53456. [Google Scholar] [CrossRef] [PubMed]
- Migazzi, A.; Scaramuzzino, C.; Anderson, E.N. Huntingtin-mediated axonal transport requires arginine methylation by PRMT6. Cell Rep. 2021, 35, 108980. [Google Scholar] [CrossRef] [PubMed]
- Darnell, J.C.; Richter, J.D. Cytoplasmic RNA-Binding Proteins and the Control of Complex Brain Function. Cold Spring Harb. Perspect. Biol. 2012, 4, a012344. [Google Scholar] [CrossRef] [PubMed]
- Dolzhanskaya, N.; Merz, G.; Aletta, J.M.; Denman, R.B. Methylation regulates the intracellular protein-protein and protein-RNA interactions of FMRP. J. Cell Sci. 2006, 119, 1933–1946. [Google Scholar] [CrossRef] [PubMed]
- Mackenzie, I.R.; Rademakers, R.; Neumann, M. TDP-43 and FUS in amyotrophic lateral sclerosis and frontotemporal dementia. Lancet Neurol. 2010, 9, 995–1007. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dong, R.; Li, X.; Lai, K.-O. Activity and Function of the PRMT8 Protein Arginine Methyltransferase in Neurons. Life 2021, 11, 1132. https://doi.org/10.3390/life11111132
Dong R, Li X, Lai K-O. Activity and Function of the PRMT8 Protein Arginine Methyltransferase in Neurons. Life. 2021; 11(11):1132. https://doi.org/10.3390/life11111132
Chicago/Turabian StyleDong, Rui, Xuejun Li, and Kwok-On Lai. 2021. "Activity and Function of the PRMT8 Protein Arginine Methyltransferase in Neurons" Life 11, no. 11: 1132. https://doi.org/10.3390/life11111132
APA StyleDong, R., Li, X., & Lai, K.-O. (2021). Activity and Function of the PRMT8 Protein Arginine Methyltransferase in Neurons. Life, 11(11), 1132. https://doi.org/10.3390/life11111132





