Structure, Activity, and Function of PRMT1
Abstract
1. Introduction
2. Structural Features
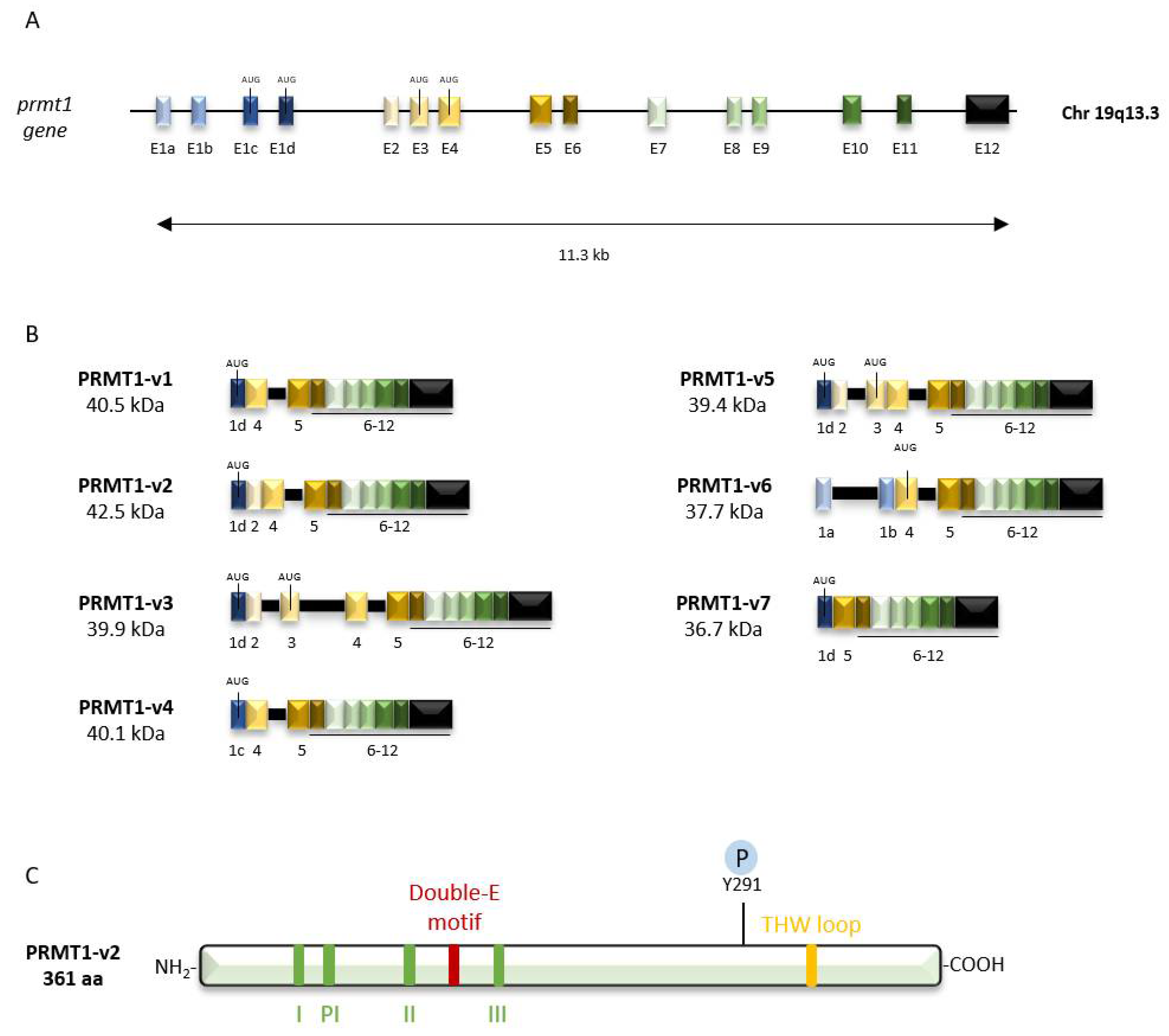
2.1. Genomic Organization
2.2. Protein Structure
2.3. PRMT1 Isoforms
3. Biochemical Features
3.1. Sequence Specificity
3.2. Product Specificity
3.3. Regulation of PRMT1 Expression and Enzymatic Activity
3.3.1. Regulation of PRMT1 Expression
3.3.2. Post-Translational Modification of PRMT1
3.3.3. PRMT1 Association with Co-Regulators
3.3.4. PTMs Influencing PRMT1 Activity
3.4. Substrates
| Biological Function | Substrate | Methylation Site | Biological Outcome | Reference |
|---|---|---|---|---|
| Transcriptional Regulation Transcriptional regulation | BRCA1 | Within the 504–802 region | Promotes BRCA1 recruitment to specific promoters | [59] |
| C/EBPα | R35, R156, R165 | Prevents C/EBPα interaction with the corepressor HDAC3 | [60] | |
| c-Myc | R299, R346 | Promotes c-Myc interaction with p300 | [61] | |
| EZH2 | R342 | Prevents EZH2 target gene expression | [62] | |
| FOXO1 | R248, R250 | Prevents FOXO1 phosphorylation by Akt | [63] | |
| FOXP3 | R48, R51 | Enhances FOXP3 transcriptional activity | [64] | |
| GLI1 | R597 | Enhances GLI1 binding to target gene promoters | [65] | |
| MyoD | R121 | Promotes MyoD DNA-binding and transcriptional activity | [66] | |
| Nrf2 | R437 | Promotes Nrf2 DNA-binding and transcriptional activity | [67] | |
| PR | R637 | Accelerates PR recycling and transcriptional activity | [68] | |
| RACO-1 | R98, R109 | Promotes c-Jun/AP1 activation | [69] | |
| RelA | R30 | Prevents RelA DNA-binding and represses NF-κB target genes | [70] | |
| RIP40 | R240, R650, R948 | Favors RIP140 nuclear export and prevents the recruitment of HDAC3 | [71] | |
| RunX1 | R206, R210 | Prevents Sin3a binding and promotes RUNX1 transcriptional activity | [72] | |
| STAT1 | R31 | Prevents STAT1 association with PIAS1 and enhances IFNα/β induced transcription | [73] | |
| TAF15 | R203 | Affects the subcellular localization of TAF-15 and enhances its transcriptional activity | [74] | |
| FUS/TLS | R216, R218, R242, R394 | Participates in the nuclear cytoplasmic shuttling of FUS/TLS and enhances its transcriptional activity | [75,76] | |
| TOP3B | R833, R835 | Promotes TOP3B interaction with TDRD3, stress granule localization and topoisomerase activity | [77] | |
| Twist1 | R34 | Regulates the nuclear import of Twist1 and represses E-cadherin expression | [78] | |
| RNA- processing | CNBP | R25, R27 | Prevents its RNA binding activity | [79] |
| G3BP1 | R435, R447 | Prevents stress granule formation during oxidative stress | [80] | |
| hnRNPA1 | R214, R226, R223, R240 | Prevents hnRNPA1 ITAF activity and RNA-binding ability | [81] | |
| HSP70 | R416, R447 | Enhances HSP70 RNA-binding and -stabilization abilities | [82] | |
| NS3 | R1493 | Affects NS3 RNA-binding and helicase activity | [46,83] | |
| RBM15 | R578 | Promotes RBM15 degradation by CNOT4 (RNA splicing) | [84] | |
| Sam68 | Within the 276–343 region | Prevents Sam68 poly(U) RNA-binding activity | [85,86] | |
| SF2/ASF | R93, R97, R109 | Affects SF2/ASF nucleocytoplasmic distribution and modulates the alternative splicing of target genes | [87,88] | |
| Translational Regulation | eIF4A1 | R362 | Prevents eIF4A1 interaction with eIF4G1 and inhibits ATPase activity | [18,89] |
| eIF4G1 | R689, R698 | Regulates eIF4G1 stability and the assembly of the translation initiation complex | [90] | |
| rpS3 | R64, R65, R67 | Promotes rpS3 import into the nucleolus and ribosome assembly | [91] | |
DNA damage repair | 53BP1 | Within the 1319–1480 region | Promotes 53BP1 recruitment to DNA-damage sites | [92] |
| APE1 | R301 | Promotes APE1 mitochondrial translocation (translocase Tom20) and protects mitochondrial DNA from oxidative damage | [93] | |
| DNA pol β | R137 | Prevents DNA pol β interaction with PCNA in BER pathway | [94] | |
| E2F-1 | R109 | Promotes E2F-1-dependent apoptosis in DNA-damaged cells | [95] | |
| FEN1 | Not determined | Stabilizes FEN1 and upregulates its DNA damage repair activities | [96] | |
| hnRNPK | R296, R299 | Prevents PKCδ-dependent apoptosis during DNA damage | [97] | |
| hnRNPUL1 | R584, R618, R620, R645, R656 | Promotes hnRNPUL1 association with NBS1 and recruitment to DNA-damage sites | [98] | |
| MRE11 | GAR domain | Promotes MRE11 recruitment to DNA-damage sites and favors its exonuclease activity | [99,100] | |
| RunX1 | R233, R237 | Confers resistance to apoptosis under stress condition and DNA damage accumulation | [101] | |
| ASK1 | R78, R80 | Prevents the stress-induced ASK1-JNK1 signaling | [102] | |
| Signal transduction | Axin | R378 | Favors Axin stability and consequently prevents Wnt/β-catenin signaling | [103] |
| BAD | R94, R96 | Prevents BAD phosphorylation by Akt and subsequent survival signaling | [104] | |
| CaMKII | R9, R275 | Prevents CaMKII-dependent signaling in cardiomyocytes | [105] | |
| CDK4 | R55, R73, R82, R163 | Prevents the formation of a CDK4/Cyc D3 complex and subsequent cell cycle progression | [106] | |
| cTnI | R146, R148 | Induces cardiac cell hypertrophy | [107] | |
| EGFR | R198, R200 | Upregulates EGFR signaling | [108] | |
| ERα | R260 | Promotes the formation of the ERα/PI3K/Src/FAK complex and subsequent activation of downstream kinase cascades | [109] | |
| INCENP | R887 | Enhances INCENP binding-affinity to AURKB and promotes cell division | [110] | |
| KCNQ | R333, R345, R353, R435 | Promotes PIP2 binding and subsequent KCNQ channel activity | [111] | |
| MYCN | R65 | Enhances MYCN stability through CDK-dependent phosphorylation | [112] | |
| NONO | R251 | Favors NONO oncogenic function | [113] | |
| p38 MAPK | R49, R149 | Promotes p38 MAPK phosphorylation by MKK3 and the subsequent activation of MAPKAK2 involved in erythroid differentiation | [114] | |
| Smad4 | R272 | Promotes Smad4 phosphorylation by GSK3 and support the activation of the canonical Wnt signaling | [115] | |
| Smad6 | R74, R81 | Participates in BMP signaling and prevents NF-κB activation | [116,117] | |
| Smad7 | R57, R67 | Facilitates TGF-β signaling | [118] | |
| TRAF6 | R88, R125 | Prevents TRAF6 ubiquitin ligase activity and regulates Toll-like receptor signaling | [119] | |
| TSC2 | R1457, R1459 | Blocks the Akt-dependent phosphorylation of TSC2 and regulates mTORC1 activity | [120] |
4. Cellular Features
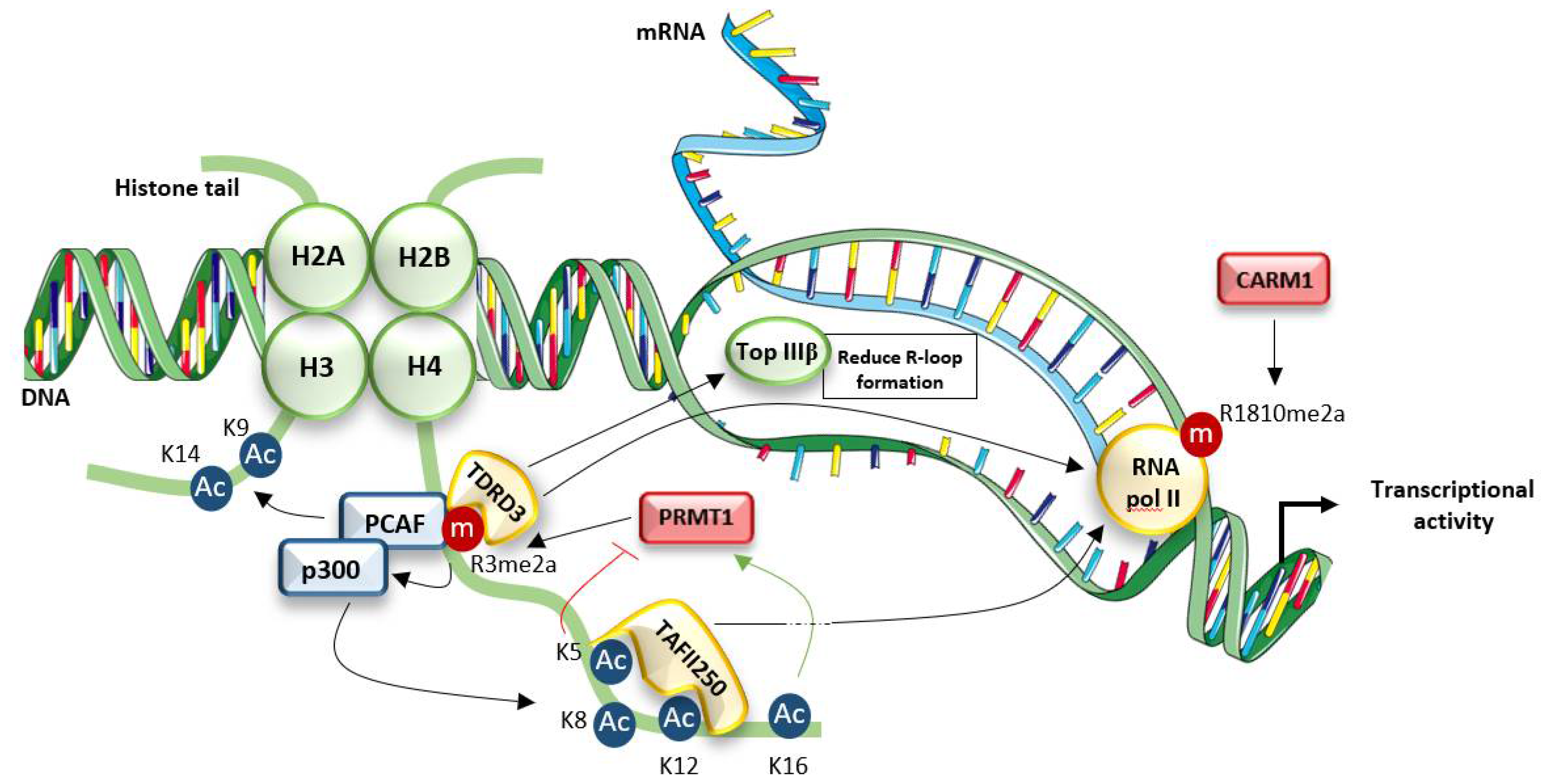
4.1. Connection with Chromatin Dynamics and Transcriptional Regulation
4.2. Connection to Cell Signaling Pathways
4.2.1. Steroid Receptors
Estrogen Receptor (ERα)
Progesterone Receptor (PR)
4.2.2. Akt Signaling Pathway
FOXO
BAD
4.2.3. NF-κB Signaling
4.2.4. Wnt Signaling
4.3. Cellular Role and Functions
4.3.1. Embryogenesis and Development
4.3.2. DNA Damage Repair
5. PRMT1 in Cancer
5.1. Breast Cancer
5.2. Colorectal Cancer
5.3. Lung Cancer
5.4. Other Cancers
5.5. PRMT1 Inhibitors
6. Outlook
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Yang, Y.; Bedford, M.T. Protein arginine methyltransferases and cancer. Nat. Rev. Cancer 2012, 13, 37–50. [Google Scholar] [CrossRef]
- Zhang, J.; Jing, L.; Li, M.; He, L.; Guo, Z. Regulation of histone arginine methylation/demethylation by methylase and demethylase. Mol. Med. Rep. 2019, 19, 3963. [Google Scholar] [CrossRef]
- Tang, J.; Frankel, A.; Cook, R.; Kim, S.; Paik, W.; Williams, K.; Clarke, S.; Herschman, H. PRMT1 is the predominant type I protein arginine methyltransferase in mammalian cells. J. Biol. Chem. 2000, 275, 7723–7730. [Google Scholar] [CrossRef]
- Scorilas, A.; Black, M.H.; Talieri, M.; Diamandis, E.P. Genomic Organization, Physical Mapping, and Expression Analysis of the Human Protein Arginine Methyltransferase 1 Gene. Biochem. Biophys. Res. Commun. 2000, 278, 349–359. [Google Scholar] [CrossRef]
- Goulet, I.; Gauvin, G.; Boisvenue, S.; Côté, J. Alternative splicing yields protein arginine methyltransferase 1 isoforms with distinct activity, substrate specificity, and subcellular localization. J. Biol. Chem. 2007, 282, 33009–33021. [Google Scholar] [CrossRef]
- Bedford, M.T.; Richard, S. Arginine methylation: An emerging regulator of protein function. Mol. Cell 2005, 18, 263–272. [Google Scholar] [CrossRef]
- Wu, H.; Min, J.; Lunin, V.V.; Antoshenko, T.; Dombrovski, L. Structural Biology of Human H3K9 Methyltransferases. PLoS ONE 2010, 5, e8570. [Google Scholar] [CrossRef]
- Tewary, S.K.; Zheng, Y.G.; Ho, M.C. Protein arginine methyltransferases: Insights into the enzyme structure and mechanism at the atomic level. Cell. Mol. Life Sci. 2019, 76, 2917–2932. [Google Scholar] [CrossRef]
- Jain, K.; Warmack, R.A.; Debler, E.W.; Hadjikyriacou, A.; Stavropoulos, P.; Clarke, S.G. Protein arginine methyltransferase product specificity is mediated by distinct active-site architectures. J. Biol. Chem. 2016, 291, 18299–18308. [Google Scholar] [CrossRef]
- Zhang, X.; Cheng, X. Structure of the predominant protein arginine methyltransferase PRMT1 and analysis of its binding to substrate peptides. Structure 2003, 11, 509–520. [Google Scholar] [CrossRef]
- Fuhrmann, J.; Clancy, K.; Thompson, P. Chemical biology of protein arginine modifications in epigenetic regulation. Chem. Rev. 2015, 115, 5413–5461. [Google Scholar] [CrossRef]
- Morales, Y.; Nitzel, D.V.; Price, O.M.; Gui, S.; Li, J.; Qu, J.; Hevel, J.M. Redox Control of Protein Arginine Methyltransferase 1 (PRMT1) Activity. J. Biol. Chem. 2015, 290, 14915–14926. [Google Scholar] [CrossRef] [PubMed]
- Weiss, V.; McBride, A.; Soriano, M.; Filman, D.; Silver, P.; Hogle, J. The structure and oligomerization of the yeast arginine methyltransferase, Hmt1. Nat. Struct. Biol. 2000, 7, 1165–1171. [Google Scholar] [CrossRef]
- Feng, Y.; Xie, N.; Jin, M.; Stahley, M.; Stivers, J.; Zheng, Y. A transient kinetic analysis of PRMT1 catalysis. Biochemistry 2011, 50, 7033–7044. [Google Scholar] [CrossRef]
- Herrmann, F.; Fackelmayer, F.O. Nucleo-cytoplasmic shuttling of protein arginine methyltransferase 1 (PRMT1) requires enzymatic activity. Genes Cells 2009, 14, 309–317. [Google Scholar] [CrossRef]
- Thandapani, P.; O’Connor, T.; Bailey, T.; Richard, S. Defining the RGG/RG motif. Mol. Cell 2013, 50, 613–623. [Google Scholar] [CrossRef]
- Morales, Y.; Cáceres, T.; May, K.; Hevel, J. Biochemistry and regulation of the protein arginine methyltransferases (PRMTs). Arch. Biochem. Biophys. 2016, 590, 138–152. [Google Scholar] [CrossRef]
- Wooderchak, W.; Zang, T.Z.; Zhou, Z.S.; Acuna, M.; Taharam, S.M.; Hevel, J. Substrate profiling of PRMT1 reveals amino acid sequences that extend beyond the “RGG” paradigm. Biochemistry 2008, 47, 9456–9466. [Google Scholar] [CrossRef] [PubMed]
- Osborne, T.C.; Obianyo, O.; Zhang, X.; Cheng, X.; Thompson, P.R. Protein Arginine Methyltransferase 1: Positively Charged Residues in Substrate Peptides Distal to the Site of Methylation Are Important for Substrate Binding and Catalysis. Biochemistry 2007, 46, 13370. [Google Scholar] [CrossRef]
- Gui, S.; Wooderchak, W.; Daly, M.; Porter, P.; Johnson, S.; Hevel, J. Investigation of the molecular origins of protein-arginine methyltransferase I (PRMT1) product specificity reveals a role for two conserved methionine residues. J. Biol. Chem. 2011, 286, 29118–29126. [Google Scholar] [CrossRef]
- Kirmizis, A.; Santos-Rosa, H.; Penkett, C.J.; Singer, M.A.; Vermeulen, M.; Mann, M.; Bähler, J.; Green, R.D.; Kouzarides, T. Arginine methylation at histone H3R2 controls deposition of H3K4 trimethylation. Nature 2007, 449, 928. [Google Scholar] [CrossRef]
- Gui, S.; Gathiaka, S.; Li, J.; Qu, J.; Acevedo, O.; Hevel, J.M. A remodeled protein arginine methyltransferase 1 (PRMT1) generates symmetric dimethylarginine. J. Biol. Chem. 2014, 289, 9320–9327. [Google Scholar] [CrossRef] [PubMed]
- Gathiaka, S.; Boykin, B.; Cáceres, T.; Hevel, J.M.; Acevedo, O. Understanding protein arginine methyltransferase 1 (PRMT1) product specificity from molecular dynamics. Bioorg. Med. Chem. 2016, 24, 4949–4960. [Google Scholar] [CrossRef]
- Gui, S.; WL, W.-D.; Zang, T.; Chen, D.; Daly, M.; Zhou, Z.; Hevel, J. Substrate-induced control of product formation by protein arginine methyltransferase 1. Biochemistry 2013, 52, 199–209. [Google Scholar] [CrossRef]
- Brown, J.I.; Koopmans, T.; van Strien, J.; Martin, N.I.; Frankel, A. Kinetic Analysis of PRMT1 Reveals Multifactorial Processivity and a Sequential Ordered Mechanism. ChemBioChem 2018, 19, 85–99. [Google Scholar] [CrossRef]
- Kölbel, K.; Ihling, C.; Bellmann-Sickert, K.; Neundorf, I.; Beck-Sickinger, A.G.; Sinz, A.; Kühn, U.; Wahle, E. Type I arginine methyltransferases PRMT1 and PRMT-3 act distributively. J. Biol. Chem. 2009, 284, 8274–8282. [Google Scholar] [CrossRef]
- Lakowski, T.M.; Frankel, A. Kinetic analysis of human protein arginine N-methyltransferase 2: Formation of monomethyl- and asymmetric dimethyl-arginine residues on histone H4. Biochem. J. 2009, 421, 253–261. [Google Scholar] [CrossRef]
- Hu, H.; Luo, C.; Zheng, Y.G. Transient kinetics define a complete kinetic model for protein arginine methyltransferase 1. J. Biol. Chem. 2016, 291, 26722–26738. [Google Scholar] [CrossRef]
- Obianyo, O.; Osborne, T.C.; Thompson, P.R. Kinetic mechanism of protein arginine methyltransferase. Biochemistry 2008, 47, 10420–10427. [Google Scholar] [CrossRef]
- Obianyo, O.; Causey, C.P.; Jones, J.E.; Thompson, P.R. Activity-Based Protein Profiling of Protein Arginine Methyltransferase 1. ACS Chem. Biol. 2011, 6, 1127–1135. [Google Scholar] [CrossRef]
- Wang, Y.; Wysocka, J.; Sayegh, J.; Lee, Y.; Perlin, J.; Leonelli, L.; Sonbuchner, L.; McDonald, C.; Cook, R.; Dou, Y.; et al. Human PAD4 regulates histone arginine methylation levels via demethylimination. Science 2004, 306, 279–283. [Google Scholar] [CrossRef]
- Chang, B.; Chen, Y.; Zhao, Y.; Bruick, R. JMJD6 is a histone arginine demethylase. Science 2007, 318, 444–447. [Google Scholar] [CrossRef]
- Li, S.; Ali, S.; Duan, X.; Liu, S.; Du, J.; Liu, C.; Dai, H.; Zhou, M.; Zhou, L.; Yang, L.; et al. JMJD1B Demethylates H4R3me2s and H3K9me2 to Facilitate Gene Expression for Development of Hematopoietic Stem and Progenitor Cells. Cell Rep. 2018, 23, 389–403. [Google Scholar] [CrossRef]
- Zhang, X.; Li, L.; Li, Y.; Li, Z.; Zhai, W.; Sun, Q.; Yang, X.; Roth, M.; Lu, S. mTOR regulates PRMT1 expression and mitochondrial mass through STAT1 phosphorylation in hepatic cell. Biochim. Biophys. Acta Mol. Cell Res. 2021, 1868, 119017. [Google Scholar] [CrossRef]
- Li, B.; Liu, L.; Li, X.; Wu, L. miR-503 suppresses metastasis of hepatocellular carcinoma cell by targeting PRMT1. Biochem. Biophys. Res. Commun. 2015, 464, 982–987. [Google Scholar] [CrossRef]
- Rust, H.L.; Subramanian, V.; West, G.M.; Young, D.D.; Schultz, P.G.; Thompson, P.R. Using unnatural amino acid mutagenesis to probe the regulation of PRMT1. ACS Chem. Biol. 2014, 9, 649–655. [Google Scholar] [CrossRef]
- Bao, X.; Siprashvili, Z.; Zarnegar, B.J.; Shenoy, R.M.; Rios, E.J.; Nady, N.; Qu, K.; Mah, A.; Webster, D.E.; Rubin, A.J.; et al. CSNK1a1 Regulates PRMT1 to Maintain the Progenitor State in Self-Renewing Somatic Tissue. Dev. Cell 2017, 43, 227–239.e5. [Google Scholar] [CrossRef]
- Musiani, D.; Giambruno, R.; Massignani, E.; Ippolito, M.R.; Maniaci, M.; Jammula, S.; Manganaro, D.; Cuomo, A.; Nicosia, L.; Pasini, D.; et al. PRMT1 Is Recruited via DNA-PK to Chromatin Where It Sustains the Senescence-Associated Secretory Phenotype in Response to Cisplatin. Cell Rep. 2020, 30, 1208–1222.e9. [Google Scholar] [CrossRef]
- Hirata, Y.; Katagiri, K.; Nagaoka, K.; Morishita, T.; Kudoh, Y.; Hatta, T.; Naguro, I.; Kano, K.; Udagawa, T.; Natsume, T.; et al. TRIM48 Promotes ASK1 Activation and Cell Death through Ubiquitination-Dependent Degradation of the ASK1-Negative Regulator PRMT1. Cell Rep. 2017, 21, 2447–2457. [Google Scholar] [CrossRef]
- Bhuripanyo, K.; Wang, Y.; Liu, X.; Zhou, L.; Liu, R.; Duong, D.; Zhao, B.; Bi, Y.; Zhou, H.; Chen, G.; et al. Identifying the substrate proteins of U-box E3s E4B and CHIP by orthogonal ubiquitin transfer. Sci. Adv. 2018, 4, e1701393. [Google Scholar] [CrossRef]
- Nie, M.; Wang, Y.; Guo, C.; Li, X.; Wang, Y.; Deng, Y.; Yao, B.; Gui, T.; Ma, C.; Liu, M.; et al. CARM1-mediated methylation of protein arginine methyltransferase 5 represses human γ-globin gene expression in erythroleukemia cells. J. Biol. Chem. 2018, 293, 17454–17463. [Google Scholar] [CrossRef]
- Sakabe, K.; Hart, G.W. O-GlcNAc Transferase Regulates Mitotic Chromatin Dynamics. J. Biol. Chem. 2010, 285, 34460–34468. [Google Scholar] [CrossRef]
- Lin, W.J.; Gary, J.D.; Yang, M.C.; Clarke, S.; Herschman, H.R. The mammalian immediate-early TIS21 protein and the leukemia-associated BTG1 protein interact with a protein-arginine N-methyltransferase. J. Biol. Chem. 1996, 271, 15034–15044. [Google Scholar] [CrossRef]
- Robin-Lespinasse, Y.; Sentis, S.; Kolytcheff, C.; Rostan, M.C.; Corbo, L.; Le Romancer, M. hCAF1, a new regulator of PRMT1-dependent arginine methylation. J. Cell Sci. 2007, 120, 638–647. [Google Scholar] [CrossRef]
- Pak, M.L.; Lakowski, T.M.; Thomas, D.; Vhuiyan, M.I.; Hüsecken, K.; Frankel, A. A protein arginine N-methyltransferase 1 (PRMT1) and 2 heteromeric interaction increases PRMT1 enzymatic activity. Biochemistry 2011, 50, 8226–8240. [Google Scholar] [CrossRef]
- Duong, F.H.T.; Christen, V.; Berke, J.M.; Penna, S.H.; Moradpour, D.; Heim, M.H. Upregulation of protein phosphatase 2Ac by hepatitis C virus modulates NS3 helicase activity through inhibition of protein arginine methyltransferase 1. J. Virol. 2005, 79, 15342–15350. [Google Scholar] [CrossRef]
- Gasperini, L.; Rossi, A.; Cornella, N.; Peroni, D.; Zuccotti, P.; Potrich, V.; Quattrone, A.; Macchi, P. The hnRNP RALY regulates PRMT1 expression and interacts with the ALS-linked protein FUS: Implication for reciprocal cellular localization. Mol. Biol. Cell 2018, 29, 3067–3081. [Google Scholar] [CrossRef]
- Lei, N.; Zhang, X.; Chen, H.; Wang, Y.; Zhan, Y.; Zheng, Z.; Shen, Y.; Wu, Q. A feedback regulatory loop between methyltransferase PRMT1 and orphan receptor TR3. Nucleic Acids Res. 2009, 37, 832–848. [Google Scholar] [CrossRef]
- Sun, Q.; Liu, L.; Mandal, J.; Molino, A.; Stolz, D.; Tamm, M.; Lu, S.; Roth, M. PDGF-BB induces PRMT1 expression through ERK1/2 dependent STAT1 activation and regulates remodeling in primary human lung fibroblasts. Cell. Signal. 2016, 28, 307–315. [Google Scholar] [CrossRef]
- Vadnais, C.; Chen, R.; Fraszczak, J.; Yu, Z.; Boulais, J.; Pinder, J.; Frank, D.; Khandanpour, C.; Hébert, J.; Dellaire, G.; et al. GFI1 facilitates efficient DNA repair by regulating PRMT1 dependent methylation of MRE11 and 53BP1. Nat. Commun. 2018, 9, 1418. [Google Scholar] [CrossRef]
- Inoue, H.; Hanawa, N.; Katsumata, S.-I.; Aizawa, Y.; Katsumata-Tsuboi, R.; Tanaka, M.; Takahashi, N.; Uehara, M. Iron deficiency negatively regulates protein methylation via the downregulation of protein arginine methyltransferase. Heliyon 2020, 6, e05059. [Google Scholar] [CrossRef]
- Wagner, S.; Weber, S.; Kleinschmidt, M.A.; Nagata, K.; Bauer, U.-M. SET-mediated promoter hypoacetylation is a prerequisite for coactivation of the estrogen-responsive pS2 gene by PRMT1. J. Biol. Chem. 2006, 281, 27242–27250. [Google Scholar] [CrossRef] [PubMed]
- Fulton, M.D.; Dang, T.; Brown, T.; Zheng, Y.G. Effects of substrate modifications on the arginine dimethylation activities of PRMT1 and PRMT5. Epigenetics 2020, 31, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Strahl, B.D.; Briggs, S.D.; Brame, C.J.; Caldwell, J.A.; Koh, S.S.; Ma, H.; Cook, R.G.; Shabanowitz, J.; Hunt, D.F.; Stallcup, M.R.; et al. Methylation of histone H4 at arginine 3 occurs in vivo and is mediated by the nuclear receptor coactivator PRMT1. Curr. Biol. 2001, 11, 996–1000. [Google Scholar] [CrossRef]
- Stallcup, M.R. Role of protein methylation in chromatin remodeling and transcriptional regulation. Oncogene 2001, 20, 3014–3020. [Google Scholar] [CrossRef]
- Wang, H.; Huang, Z.; Xia, L.; Feng, Q.; Erdjument-Bromage, H.; Strahl, B.; Briggs, S.; Allis, C.; Wong, J.; Tempst, P.; et al. Methylation of histone H4 at arginine 3 facilitating transcriptional activation by nuclear hormone receptor. Science 2001, 293, 853–857. [Google Scholar] [CrossRef] [PubMed]
- Waldmann, T.; Izzo, A.; Kamieniarz, K.; Richter, F.; Vogler, C.; Sarg, B.; Lindner, H.; Young, N.L.; Mittler, G.; Garcia, B.A.; et al. Methylation of H2AR29 is a novel repressive PRMT6 target. Epigenetics Chromatin 2011, 4, 11. [Google Scholar] [CrossRef]
- Dhar, S.; Vemulapalli, V.; Patananan, A.N.; Huang, G.L.; Di Lorenzo, A.; Richard, S.; Comb, M.J.; Guo, A.; Clarke, S.G.; Bedford, M.T. Loss of the major type i arginine methyltransferase PRMT1 causes substrate scavenging by other PRMTs. Sci. Rep. 2013, 3, 138–152. [Google Scholar] [CrossRef]
- Guendel, I.; Carpio, L.; Pedati, C.; Schwartz, A.; Teal, C.; Kashanchi, F.; Kehn-Hall, K. Methylation of the Tumor Suppressor Protein, BRCA1, Influences Its Transcriptional Cofactor Function. PLoS ONE 2010, 5, e11379. [Google Scholar] [CrossRef]
- Liu, L.-M.; Sun, W.-Z.; Fan, X.-Z.; Xu, Y.-L.; Cheng, M.-B.; Zhang, Y. Molecular Cell Biology Methylation of C/EBPa by PRMT1 Inhibits Its Tumor-Suppressive Function in Breast Cancer. Cancer Res. 2019, 79, 2865–2877. [Google Scholar] [CrossRef]
- Tikhanovich, I.; Zhao, J.; Bridges, B.; Kumer, S.; Roberts, B.; Weinman, S.A. Arginine methylation regulates c-Myc–dependent transcription by altering promoter recruitment of the acetyltransferase p300. J. Biol. Chem. 2017, 292, 13333–13344. [Google Scholar] [CrossRef]
- Li, Z.; Wang, D.; Lu, J.; Huang, B.; Wang, Y.; Dong, M.; Fan, D.; Li, H.; Gao, Y.; Hou, P.; et al. Methylation of EZH2 by PRMT1 regulates its stability and promotes breast cancer metastasis. Cell Death Differ. 2020, 27, 3226–3242. [Google Scholar] [CrossRef]
- Yamagata, K.; Daitoku, H.; Takahashi, Y.; Namiki, K.; Hisatake, K.; Kako, K.; Mukai, H.; Kasuya, Y.; Fukamizu, A. Arginine methylation of FOXO transcription factors inhibits their phosphorylation by Akt. Mol. Cell 2008, 32, 221–231. [Google Scholar] [CrossRef] [PubMed]
- Kagoya, Y.; Saijo, H.; Matsunaga, Y.; Guo, T.; Saso, K.; Anczurowski, M.; Wang, C.H.; Sugata, K.; Murata, K.; Butler, M.O.; et al. Arginine methylation of FOXP3 is crucial for the suppressive function of regulatory T cells. J. Autoimmun. 2019, 97, 10–21. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Hsu, J.-M.; Kang, Y.; Wei, Y.; Lee, P.-C.; Chang, S.-J.; Hsu, Y.-H.; Hsu, J.L.; Wang, H.-L.; Chang, W.-C.; et al. Oncogenic functions of Gli in pancreatic adenocarcinoma are supported by its PRMT1-mediated methylation. Cancer Res. 2016, 76, 7049. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Zhang, X.; Cheng, M.; Zhang, Y. PRMT1 activates myogenin transcription via MyoD arginine methylation at R121. Biochim. Biophys. Acta Gene Regul. Mech. 2019, 1862, 194442. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Li, H.; Liu, L.; Lu, Y.; Gao, Y.; Geng, P.; Li, X.; Huang, B.; Zhang, Y.; Lu, J. Methylation of arginine by PRMT1 regulates Nrf2 transcriptional activity during the antioxidative response. Biochim. Biophys. Acta 2016, 1863, 2093–2103. [Google Scholar] [CrossRef]
- Malbeteau, L.; Poulard, C.; Languilaire, C.; Mikaelian, I.; Flamant, F.; Le Romancer, M.; Corbo, L. PRMT1 Is Critical for the Transcriptional Activity and the Stability of the Progesterone Receptor. IScience 2020, 23, 101236. [Google Scholar] [CrossRef]
- Davies, C.C.; Chakraborty, A.; Diefenbacher, M.E.; Skehel, M.; Behrens, A. Arginine methylation of the c-Jun coactivator RACO-1 is required for c-Jun/AP-1 activation. EMBO J. 2013, 32, 1556. [Google Scholar] [CrossRef]
- Reintjes, A.; Fuchs, J.E.; Kremser, L.; Lindner, H.H.; Liedl, K.R.; Huber, L.A.; Valovka, T. Asymmetric arginine dimethylation of RelA provides a repressive mark to modulate TNFα/NF-κB response. Proc. Natl. Acad. Sci. USA 2016, 113, 4326–4331. [Google Scholar] [CrossRef]
- Huq, M.D.M.; Gupta, P.; Tsai, N.-P.; White, R.; Parker, M.G.; Wei, L.-N. Suppression of receptor interacting protein 140 repressive activity by protein arginine methylation. EMBO J. 2006, 25, 5094–5104. [Google Scholar] [CrossRef]
- Zhao, X.; Jankovic, V.; Gural, A.; Huang, G.; Pardanani, A.; Menendez, S.; Zhang, J.; Dunne, R.; Xiao, A.; Erdjument-Bromage, H.; et al. Methylation of RUNX1 by PRMT1 abrogates SIN3A binding and potentiates its transcriptional activity. Genes Dev. 2008, 22, 640. [Google Scholar] [CrossRef]
- Mowen, K.A.; Tang, J.; Zhu, W.; Schurter, B.T.; Shuai, K.; Herschman, H.R.; David, M. Arginine Methylation of STAT1 Modulates IFNα/β-Induced Transcription. Cell 2001, 104, 731–741. [Google Scholar] [CrossRef]
- Jobert, L.; Argentini, M.; Tora, L. PRMT1 mediated methylation of TAF15 is required for its positive gene regulatory function. Exp. Cell Res. 2009, 315, 1273–1286. [Google Scholar] [CrossRef] [PubMed]
- Tradewell, M.; Yu, Z.; Tibshirani, M.; Boulanger, M.; Durham, H.; Richard, S. Arginine methylation by PRMT1 regulates nuclear-cytoplasmic localization and toxicity of FUS/TLS harbouring ALS-linked mutations. Hum. Mol. Genet. 2012, 21, 136–149. [Google Scholar] [CrossRef]
- Du, K.; Arai, S.; Kawamura, T.; Matsushita, A.; Kurokawa, R. TLS and PRMT1 synergistically coactivate transcription at the survivin promoter through TLS arginine methylation. Biochem. Biophys. Res. Commun. 2011, 404, 991–996. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Wang, Z.; Narayanan, N.; Yang, Y. Arginine methylation of the C-terminus RGG motif promotes TOP3B topoisomerase activity and stress granule localization. Nucleic Acids Res. 2018, 46, 3061–3074. [Google Scholar] [CrossRef]
- Avasarala, S.; Van Scoyk, M.; Kumar, M.; Rathinam, K.; Zerayesus, S.; Zhao, X.; Zhang, W.; Pergande, M.R.; Borgia, J.A.; Degregori, J.; et al. PRMT1 Is a Novel Regulator of Epithelial-Mesenchymal-Transition in Non-small Cell Lung Cancer. J. Biol. Chem. 2015, 290, 13479–13489. [Google Scholar] [CrossRef]
- Wei, H.-M.; Hu, H.-H.; Chang, G.-Y.; Lee, Y.-J.; Li, Y.-C.; Chang, H.-H.; Li, C. Arginine methylation of the cellular nucleic acid binding protein does not affect its subcellular localization but impedes RNA binding. FEBS Lett. 2014, 588, 1542–1548. [Google Scholar] [CrossRef] [PubMed]
- Tsai, W.-C.; Gayatri, S.; Reineke, L.C.; Sbardella, G.; Bedford, M.T.; Lloyd, R.E. Arginine Demethylation of G3BP1 Promotes Stress Granule Assembly. J. Biol. Chem. 2016, 291, 22671. [Google Scholar] [CrossRef]
- Wall, M.L.; Lewis, S.M. Methylarginines within the RGG-Motif Region of hnRNP A1 Affect Its IRES Trans-Acting Factor Activity and Are Required for hnRNP A1 Stress Granule Localization and Formation. J. Mol. Biol. 2017, 429, 295–307. [Google Scholar] [CrossRef]
- Wang, L.; Jia, Z.; Xie, D.; Zhao, T.; Tan, Z.; Zhang, S.; Kong, F.; Wei, D.; Xie, K. Methylation of HSP70 orchestrates its binding to and stabilization of BCL2 mRNA and renders pancreatic cancer cells resistant to therapeutics. Cancer Res. 2021, 80, 4500–4513. [Google Scholar] [CrossRef] [PubMed]
- Rho, J.; Choi, S.; Seong, Y.R.; Choi, J.; Im, D.-S. The Arginine-1493 Residue in QRRGRTGR1493G Motif IV of the Hepatitis C Virus NS3 Helicase Domain Is Essential for NS3 Protein Methylation by the Protein Arginine Methyltransferase 1. J. Virol. 2001, 75, 8031. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhang, L.; Tran, N.-T.; Su, H.; Wang, R.; Lu, Y.; Tang, H.; Aoyagi, S.; Guo, A.; Khodadadi-Jamayran, A.; Zhou, D.; et al. Cross-talk between PRMT1-mediated methylation and ubiquitylation on RBM15 controls RNA splicing. eLife 2015, 4, e60742. [Google Scholar] [CrossRef] [PubMed]
- Côté, J.; Boisvert, F.-M.; Boulanger, M.-C.; Bedford, M.T.; Richard, S. Sam68 RNA Binding Protein Is an In Vivo Substrate for Protein Arginine N-Methyltransferase 1. Mol. Biol. Cell 2003, 14, 274. [Google Scholar] [CrossRef]
- Rho, J.; Choi, S.; Jung, C.R.; Im, D.S. Arginine methylation of Sam68 and SLM proteins negatively regulates their poly(U) RNA binding activity. Arch. Biochem. Biophys. 2007, 466, 49–57. [Google Scholar] [CrossRef]
- Sinha, R.; Allemand, E.; Zhang, Z.; Karni, R.; Myers, M.P.; Krainer, A.R. Arginine Methylation Controls the Subcellular Localization and Functions of the Oncoprotein Splicing Factor SF2/ASF. Mol. Cell. Biol. 2010, 30, 2762–2774. [Google Scholar] [CrossRef]
- Jia, H.; Du, C.H.; Bao, S.L.; Zheng, H.Y. Protein arginine methyltransferase 1 methylates SF2/ASF at arginine. Chin. J. Cancer Biother. 2009, 16, 216–220. [Google Scholar] [CrossRef]
- Tahara, S.M.; Acuna, M. Discrimination of eIF4A isoforms by protein arginine methyltransferase 1 (PRMT1). FASEB J. 2006, 20, A109–A110. [Google Scholar] [CrossRef]
- Hsu, J.H.-R.; Hubbell-Engler, B.; Adelmant, G.; Huang, J.; Joyce, C.E.; Vazquez, F.; Weir, B.A.; Montgomery, P.; Tsherniak, A.; Giacomelli, A.O.; et al. Prmt1-mediated translation regulation is a crucial vulnerability of cancer. Cancer Res. 2017, 77, 4613. [Google Scholar] [CrossRef]
- Shin, H.-S.; Jang, C.-Y.; Kim, H.D.; Kim, T.-S.; Kim, S.; Kim, J. Arginine methylation of ribosomal protein S3 affects ribosome assembly. Biochem. Biophys. Res. Commun. 2009, 385, 273–278. [Google Scholar] [CrossRef]
- Boisvert, F.-M.; Rhie, A.; Richard, S.; Doherty, A.J. The GAR Motif of 53BP1 is Arginine Methylated by PRMT1 and is Necessary for 53BP1 DNA Binding Activity. Cell Cycle 2005, 4, 1834–1841. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, Q.; Li, L.L.; Mu, D.; Hua, K.; Ci, S.; Shen, L.; Zheng, L.; Shen, B.; Guo, Z. Arginine methylation of APE1 promotes its mitochondrial translocation to protect cells from oxidative damage. Free Radic. Biol. Med. 2020, 158, 60–73. [Google Scholar] [CrossRef]
- El-Andaloussi, N.; Valovka, T.; Toueille, M.; Hassa, P.O.; Gehrig, P.; Covic, M.; Hübscher, U.; Hottiger, M.O. Methylation of DNA polymerase ß by protein arginine methyltransferase 1 regulates its binding to proliferating cell nuclear antigen. FASEB J. 2007, 21, 26–34. [Google Scholar] [CrossRef] [PubMed]
- Zheng, S.; Moehlenbrink, J.; Lu, Y.; Zalmas, L.; Sagum, C.; Carr, S.; McGouran, J.; Alexander, L.; Fedorov, O.; Munro, S.; et al. Arginine methylation-dependent reader-writer interplay governs growth control by E2F-1. Mol. Cell 2013, 52, 37–51. [Google Scholar] [CrossRef] [PubMed]
- He, L.; Hu, Z.; Sun, Y.; Zhang, M.; Zhu, H.; Jiang, L.; Zhang, Q.; Mu, D.; Zhang, J.; Gu, L.; et al. PRMT1 is critical to FEN1 expression and drug resistance in lung cancer cells. DNA Repair 2020, 95, 102953. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Chiou, Y.; Fu, S.; Shih, I.; Weng, T.; Lin, W.; Lin, C. Arginine methylation of hnRNPK negatively modulates apoptosis upon DNA damage through local regulation of phosphorylation. Nucleic Acids Res. 2014, 42, 9908–9924. [Google Scholar] [CrossRef]
- Gurunathan, G.; Yu, Z.; Coulombe, Y.; Masson, J.-Y.; Richard, S. Arginine methylation of hnRNPUL1 regulates interaction with NBS1 and recruitment to sites of DNA damage. Sci. Rep. 2015, 5, 1–9. [Google Scholar] [CrossRef]
- Boisvert, F.M.; Déry, U.; Masson, J.Y.; Richard, S. Arginine methylation of MRE11 by PRMT1 is required for DNA damage checkpoint control. Genes Dev. 2005, 19, 671–676. [Google Scholar] [CrossRef]
- Déry, U.; Coulombe, Y.; Rodrigue, A.; Stasiak, A.; Richard, S.; Masson, J.-Y. A Glycine-Arginine Domain in Control of the Human MRE11 DNA Repair Protein. Mol. Cell. Biol. 2008, 28, 3058. [Google Scholar] [CrossRef]
- Matsumura, T.; Nakamura-Ishizu, A.; Anurag Muddineni, S.S.N.; Tan, D.Q.; Wang, C.Q.; Tokunaga, K.; Tirado-Magallanes, R.; Sian, S.; Benoukraf, T.; Okuda, T.; et al. Hematopoietic stem cells acquire survival advantage by loss of RUNX1 methylation identified in familial leukemia. Blood 2020, 136, 1919–1932. [Google Scholar] [CrossRef]
- Cho, J.-H.; Lee, M.-K.; Yoon, K.W.; Lee, J.; Cho, S.-G.; Choi, E.-J. Arginine methylation-dependent regulation of ASK1 signaling by PRMT1. Cell Death Differ. 2012, 19, 859. [Google Scholar] [CrossRef]
- Cha, B.; Kim, W.; Kim, Y.K.; Hwang, B.N.; Park, S.Y.; Yoon, J.W.; Park, W.S.; Cho, J.W.; Bedford, M.T.; Jho, E.H. Methylation by protein arginine methyltransferase 1 increases stability of Axin, a negative regulator of Wnt signaling. Oncogene 2011, 30, 2379–2389. [Google Scholar] [CrossRef]
- Sakamaki, J.; Daitoku, H.; Ueno, K.; Hagiwara, A.; Yamagata, K.; Fukamizu, A. Arginine methylation of BCL-2 antagonist of cell death (BAD) counteracts its phosphorylation and inactivation by Akt. Proc. Natl. Acad. Sci. USA 2011, 108, 6085–6090. [Google Scholar] [CrossRef] [PubMed]
- Pyun, J.; Kim, H.; Jeong, M.; Ahn, B.; Vuong, T.; Lee, D.; Choi, S.; Koo, S.; Cho, H.; Kang, J. Cardiac specific PRMT1 ablation causes heart failure through CaMKII dysregulation. Nat. Commun. 2018, 9, 1–15. [Google Scholar] [CrossRef]
- Dolezal, E.; Infantino, S.; Drepper, F.; Börsig, T.; Singh, A.; Wossning, T.; Fiala, G.J.; Minguet, S.; Warscheid, B.; Tarlinton, D.M.; et al. The BTG2-PRMT1 module limits pre-B cell expansion by regulating the CDK4-Cyclin-D3 complex. Nat. Immunol. 2017, 18, 911–920. [Google Scholar] [CrossRef] [PubMed]
- Onwuli, D.O.; Samuel, S.F.; Sfyri, P.; Welham, K.; Goddard, M.; Abu-Omar, Y.; Loubani, M.; Rivero, F.; Matsakas, A.; Benoit, D.M.; et al. The inhibitory subunit of cardiac troponin (cTnI) is modified by arginine methylation in the human heart. Int. J. Cardiol. 2019, 282, 76–80. [Google Scholar] [CrossRef]
- Liao, H.-W.; Hsu, J.-M.; Xia, W.; Wang, H.-L.; Wang, Y.-N.; Chang, W.-C.; Arold, S.T.; Chou, C.-K.; Tsou, P.-H.; Yamaguchi, H.; et al. PRMT1-mediated methylation of the EGF receptor regulates signaling and cetuximab response. J. Clin. Investig. 2015, 125, 4529–4543. [Google Scholar] [CrossRef] [PubMed]
- Le Romancer, M.; Treilleux, I.; Leconte, N.; Robin-Lespinasse, Y.; Sentis, S.; Bouchekioua-Bouzaghou, K.; Goddard, S.; Gobert-Gosse, S.; Corbo, L. Regulation of Estrogen Rapid Signaling through Arginine Methylation by PRMT1. Mol. Cell 2008, 31, 212–221. [Google Scholar] [CrossRef]
- Deng, X.; Keudell, G. Von Suzuki, T.; Dohmae, N.; Nakakido, M.; Piao, L.; Yoshioka, Y.; Nakamura, Y.; Hamamoto, R. PRMT1 promotes mitosis of cancer cells through arginine methylation of INCENP. Oncotarget 2015, 6, 35173. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.J.; Jeong, M.H.; Kim, K.R.; Jung, C.Y.; Lee, S.Y.; Kim, H.; Koh, J.; Vuong, T.A.; Jung, S.; Yang, H.; et al. Protein arginine methylation facilitates KCNQ channel-PIP2 interaction leading to seizure suppression. eLife 2016, 5, e17159. [Google Scholar] [CrossRef] [PubMed]
- Eberhardt, A.; Hansen, J.N.; Koster, J.; Lotta, L.T., Jr.; Wang, S.; Livingstone, E.; Qian, K.; Valentijn, L.J.; Zheng, Y.G.; Schor, N.F.; et al. Protein arginine methyltransferase 1 is a novel regulator of MYCN in neuroblastoma. Oncotarget 2016, 7, 63629. [Google Scholar] [CrossRef] [PubMed]
- Yin, X.-K.; Wang, Y.-L.; Wang, F.; Feng, W.-X.; Bai, S.-M.; Zhao, W.-W.; Feng, L.-L.; Wei, M.-B.; Qin, C.-L.; Wang, F.; et al. PRMT1 enhances oncogenic arginine methylation of NONO in colorectal cancer. Oncogene 2021, 40, 1375. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Hua, W.; Chen, C.; Lin, W. The MKK-Dependent Phosphorylation of p38α Is Augmented by Arginine Methylation on Arg49/Arg149 during Erythroid Differentiation. Int. J. Mol. Sci. 2020, 21, 3546. [Google Scholar] [CrossRef]
- Albrecht, L.V.; Ploper, D.; Tejeda-Muñoz, N.; De Robertis, E.M. Arginine methylation is required for canonical Wnt signaling and endolysosomal trafficking. Proc. Natl. Acad. Sci. USA 2018, 115, E5317–E5325. [Google Scholar] [CrossRef]
- Xu, J.; Wang, A.H.; Oses-Prieto, J.; Makhijani, K.; Katsuno, Y.; Pei, M.; Yan, L.; Zheng, Y.G.; Burlingame, A.; Brückner, K.; et al. Arginine Methylation Initiates BMP-Induced Smad Signaling. Mol. Cell 2013, 51, 5–19. [Google Scholar] [CrossRef]
- Zhang, T.; Wu, J.; Ungvijanpunya, N.; Jackson-Weaver, O.; Gou, Y.; Feng, J.; Ho, T.; Shen, Y.; Liu, J.; Richard, S.; et al. Smad6 Methylation Represses NFκB Activation and Periodontal Inflammation. J. Dent. Res. 2018, 97, 810–819. [Google Scholar] [CrossRef]
- Katsuno, Y.; Qin, J.; Oses-Prieto, J.; Wang, H.; Jackson-Weaver, O.; Zhang, T.; Lamouille, S.; Wu, J.; Burlingame, A.; Xu, J.; et al. Arginine methylation of SMAD7 by PRMT1 in TGF-β-induced epithelial-mesenchymal transition and epithelial stem-cell generation. J. Biol. Chem. 2018, 293, 13059–137072. [Google Scholar] [CrossRef]
- Tikhanovich, I.; Kuravi, S.; Artigues, A.; Villar, M.T.; Dorko, K.; Nawabi, A.; Roberts, B.; Weinman, S.A. Dynamic Arginine Methylation of Tumor Necrosis Factor (TNF) Receptor-associated Factor 6 Regulates Toll-like Receptor Signaling. J. Biol. Chem. 2015, 290, 22236–22249. [Google Scholar] [CrossRef]
- Gen, S.; Matsumoto, Y.; Kobayashi, K.-I.; Suzuki, T.; Inoue, J.; Yamamoto, Y. Stability of tuberous sclerosis complex 2 is controlled by methylation at R1457 and R1459. Sci. Rep. 2020, 10, 1–9. [Google Scholar] [CrossRef]
- Yang, Y.; Lu, Y.; Espejo, A.; Wu, J.; Xu, W.; Liang, S.; Bedford, M.T. TDRD3 is an Effector Molecule for Arginine Methylated Histone Marks. Mol. Cell 2010, 40, 1016. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; McBride, K.M.; Hensley, S.; Lu, Y.; Chedin, F.; Bedford, M.T. Arginine methylation facilitates the recruitment of TOP3B to chromatin to prevent R-loop accumulation. Mol. Cell 2014, 53, 484. [Google Scholar] [CrossRef]
- Sims, R.J., III; Rojas, L.A.; Beck, D.B.; Bonasio, R.; Schüller, R.; Drury, W.J., III; Eick, D.; Reinberg, D. The C-Terminal Domain of RNA Polymerase II Is Modified by Site-Specific Methylation. Science 2011, 332, 99. [Google Scholar] [CrossRef]
- Huang, S.; Litt, M.; Felsenfeld, G. Methylation of histone H4 by arginine methyltransferase PRMT1 is essential in vivo for many subsequent histone modifications. Genes Dev. 2005, 19, 1885–1893. [Google Scholar] [CrossRef]
- Li, X.; Hu, X.; Patel, B.; Zhou, Z.; Liang, S.; Ybarra, R.; Qiu, Y.; Felsenfeld, G.; Bungert, J.; Huang, S. H4R3 methylation facilitates β-globin transcription by regulating histone acetyltransferase binding and H3 acetylation. Blood 2010, 115, 2028–2037. [Google Scholar] [CrossRef]
- Fulton, M.D.; Zhang, J.; He, M.; Ho, M.-C.; Zheng, Y.G. Intricate Effects of α-Amino and Lysine Modifications on Arginine Methylation of the N-Terminal Tail of Histone H4. Biochemistry 2017, 56, 3539–3548. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Wang, J.; Asher, S.; Hoang, L.; Guardiani, C.; Ivanov, I.; Zheng, Y.G. Histone H4 acetylation differentially modulates arginine methylation by an in cis mechanism. J. Biol. Chem. 2011, 286, 20323–20334. [Google Scholar] [CrossRef]
- Thiebaut, C.; Vlaeminck-Guillem, V.; Trédan, O.; Poulard, C.; Le Romancer, M. Non-genomic signaling of steroid receptors in cancer. Mol. Cell. Endocrinol. 2021, 538, 111453. [Google Scholar] [CrossRef]
- Choucair, A.; Pham, T.H.; Omarjee, S.; Jacquemetton, J.; Kassem, L.; Trédan, O.; Rambaud, J.; Marangoni, E.; Corbo, L.; Treilleux, I.; et al. The arginine methyltransferase PRMT1 regulates IGF-1 signaling in breast cancer. Oncogene 2019, 38, 4015–4027. [Google Scholar] [CrossRef]
- Poulard, C.; Rambaud, J.; Hussein, N.; Corbo, L.; Le Romancer, M. JMJD6 regulates ERα methylation on arginine. PLoS ONE 2014, 9, e87982. [Google Scholar] [CrossRef] [PubMed]
- Greer, E.L.; Brunet, A. FOXO transcription factors at the interface between longevity and tumor suppression. Oncogene 2005, 24, 7410–7425. [Google Scholar] [CrossRef] [PubMed]
- Brunet, A.; Bonni, A.; Zigmond, M.J.; Lin, M.Z.; Juo, P.; Hu, L.S.; Anderson, M.J.; Arden, K.C.; Blenis, J.; Greenberg, M.E. Akt Promotes Cell Survival by Phosphorylating and Inhibiting a Forkhead Transcription Factor. Cell 1999, 96, 857–868. [Google Scholar] [CrossRef]
- Kops, G.; Burgering, B. Forkhead transcription factors: New insights into protein kinase B (c-akt) signaling. J. Mol. Med. 1999, 77, 656–665. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Tindall, D.J. Regulation of FoxO protein stability via ubiquitination and proteasome degradation. Biochim. Biophys. Acta 2011, 1813, 1961. [Google Scholar] [CrossRef] [PubMed]
- Matsuzaki, H.; Daitoku, H.; Hatta, M.; Tanaka, K.; Fukamizu, A. Insulin-induced phosphorylation of FKHR (Foxo1) targets to proteasomal degradation. Proc. Natl. Acad. Sci. USA 2003, 100, 11285–11290. [Google Scholar] [CrossRef]
- Hassa, P.O.; Covic, M.; Bedford, M.T.; Hottiger, M.O. Protein Arginine Methyltransferase 1 Coactivates NF-κB-Dependent Gene Expression Synergistically with CARM1 and PARP1. J. Mol. Biol. 2008, 377, 668–678. [Google Scholar] [CrossRef]
- Pawlak, M.R.; Scherer, C.A.; Chen, J.; Roshon, M.J.; Ruley, H.E. Arginine N-Methyltransferase 1 Is Required for Early Postimplantation Mouse Development, but Cells Deficient in the Enzyme Are Viable. Mol. Cell. Biol. 2000, 20, 4859. [Google Scholar] [CrossRef]
- Tsai, Y.; Pan, H.; Hung, C.; Hou, P.; Li, Y.; Lee, Y.; Shen, Y.; Wu, T.; Li, C. The predominant protein arginine methyltransferase PRMT1 is critical for zebrafish convergence and extension during gastrulation. FEBS J. 2011, 278, 905–917. [Google Scholar] [CrossRef]
- Shibata, Y.; Okada, M.; Miller, T.C.; Shi, Y.-B. Knocking out histone methyltransferase PRMT1 leads to stalled tadpole development and lethality in Xenopus tropicalis. Biochim. Biophys. Acta Gen. Subj. 2020, 1864, 129482. [Google Scholar] [CrossRef]
- Batut, J.; Vandel, L.; Leclerc, C.; Daguzan, C.; Moreau, M.; Néant, I. The Ca2+-induced methyltransferase xPRMT1b controls neural fate in amphibian embryo. Proc. Natl. Acad. Sci. USA 2005, 102, 15128–15133. [Google Scholar] [CrossRef]
- Sato, A.; Kim, J.D.; Mizukami, H.; Nakashima, M.; Kako, K.; Ishida, J.; Itakura, A.; Takeda, S.; Fukamizu, A. Gestational changes in PRMT1 expression of murine placentas. Placenta 2018, 65, 47–54. [Google Scholar] [CrossRef]
- Yu, Z.; Chen, T.; Hébert, J.; Li, E.; Richard, S. A Mouse PRMT1 Null Allele Defines an Essential Role for Arginine Methylation in Genome Maintenance and Cell Proliferation. Mol. Cell. Biol. 2009, 29, 2982. [Google Scholar] [CrossRef] [PubMed]
- Yu, Z.; Vogel, G.; Coulombe, Y.; Dubeau, D.; Spehalski, E.; Hébert, J.; Ferguson, D.O.; Masson, J.Y.; Richard, S. The MRE11 GAR motif regulates DNA double-strand break processing and ATR activation. Cell Res. 2012, 22, 305. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, T.R.H.; Glenfield, K.; Jeyanthan, K.; Zhu, X.-D. Arginine Methylation Regulates Telomere Length and Stability. Mol. Cell. Biol. 2009, 29, 4918–4934. [Google Scholar] [CrossRef]
- Zhao, F.; Kim, W.; Kloeber, J.A.; Lou, Z. DNA end resection and its role in DNA replication and DSB repair choice in mammalian cells. Exp. Mol. Med. 2020, 52, 1705–1714. [Google Scholar] [CrossRef]
- Adams, M.M.; Wang, B.; Xia, Z.; Morales, J.C.; Lu, X.; Donehower, L.A.; Bochar, D.A.; Elledge, S.J.; Carpenter, P.B. 53BP1 Oligomerization is Independent of its Methylation by PRMT1. Cell Cycle 2005, 4, 12. [Google Scholar] [CrossRef] [PubMed]
- Guo, Z.; Zheng, L.; Xu, H.; Dai, H.; Zhou, M.; Pascua, M.; Chen, Q.; Shen, B. Methylation of FEN1 suppresses nearby phosphorylation and facilitates PCNA binding. Nat. Chem. Biol. 2010, 6, 766–773. [Google Scholar] [CrossRef]
- Mathioudaki, K.; Scorilas, A.; Ardavanis, A.; Lymberi, P.; Tsiambas, E.; Devetzi, M.; Apostolaki, A.; Talieri, M. Clinical evaluation of PRMT1 gene expression in breast cancer. Tumor Biol. 2011, 32, 575–582. [Google Scholar] [CrossRef]
- Poulard, C.; Treilleux, I.; Lavergne, E.; Bouchekioua-Bouzaghou, K.; Goddard-Léon, S.; Chabaud, S.; Trédan, O.; Corbo, L.; Le Romancer, M. Activation of rapid oestrogen signalling in aggressive human breast cancers. EMBO Mol. Med. 2012, 4, 1200–1213. [Google Scholar] [CrossRef]
- Poulard, C.; Jacquemetton, J.; Trédan, O.; Cohen, P.A.; Vendrell, J.; Ghayad, S.E.; Treilleux, I.; Marangoni, E.; Le Romancer, M. Oestrogen non-genomic signalling is activated in tamoxifen-resistant breast cancer. Int. J. Mol. Sci. 2019, 20, 2773. [Google Scholar] [CrossRef] [PubMed]
- Jacquemetton, J.; Kassem, L.; Poulard, C.; Dahmani, A.; De Plater, L.; Montaudon, E.; Sourd, L.; Morisset, L.; El Botty, R.; Chateau-Joubert, S.; et al. Analysis of genomic and non-genomic signaling of estrogen receptor in PDX models of breast cancer treated with a combination of the PI3K inhibitor Alpelisib (BYL719) and fulvestrant. Breast Cancer Res. 2021; in press. [Google Scholar]
- Nakai, K.; Xia, W.; Liao, H.; Saito, M.; Hung, M.; Yamaguchi, H. The role of PRMT1 in EGFR methylation and signaling in MDA-MB-468 triple-negative breast cancer cells. Breast Cancer 2018, 25, 74–80. [Google Scholar] [CrossRef]
- Gao, Y.; Zhao, Y.; Zhang, J.; Lu, Y.; Liu, X.; Geng, P.; Huang, B.; Zhang, Y.; Lu, J. The dual function of PRMT1 in modulating epithelial-mesenchymal transition and cellular senescence in breast cancer cells through regulation of ZEB1. Sci. Rep. 2016, 6, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Papadokostopoulou, A.; Mathioudaki, K.; Scorilas, A.; Xynopoulos, D.; Ardavanis, A.; Kouroumalis, E.; Talieri, M. Colon cancer and protein arginine methyltransferase 1 gene expression. Anticancer Res. 2009, 29, 1361–1366. [Google Scholar] [PubMed]
- Mathioudaki, K.; Papadokostopoulou, A.; Scorilas, A.; Xynopoulos, D.; Agnanti, N.; Talieri, M. The PRMT1 gene expression pattern in colon cancer. Br. J. Cancer 2008, 99, 2094. [Google Scholar] [CrossRef]
- Yao, B.; Gui, T.; Zeng, X.; Deng, Y.; Wang, Z.; Wang, Y.; Yang, D.; Li, Q.; Xu, P.; Hu, R.; et al. PRMT1-mediated H4R3me2a recruits SMARCA4 to promote colorectal cancer progression by enhancing EGFR signaling. Genome Med. 2021, 13, 1–21. [Google Scholar] [CrossRef]
- Yoshimatsu, M.; Toyokawa, G.; Hayami, S.; Unoki, M.; Tsunoda, T.; Field, H.I.; Kelly, J.D.; Neal, D.E.; Maehara, Y.; Ponder, B.A.J.; et al. Dysregulation of PRMT1 and PRMT6, Type I arginine methyltransferases, is involved in various types of human cancers. Int. J. Cancer 2011, 128, 562–573. [Google Scholar] [CrossRef]
- Zhao, Y.; Lu, Q.; Li, C.; Wang, X.; Jiang, L.; Huang, L.; Wang, C.; Chen, H. PRMT1 regulates the tumour-initiating properties of esophageal squamous cell carcinoma through histone H4 arginine methylation coupled with transcriptional activation. Cell Death Dis. 2019, 10, 1–17. [Google Scholar] [CrossRef]
- He, X.; Zhu, Y.; Lin, Y.; Li, M.; Du, J.; Dong, H.; Sun, J.; Zhu, L.; Wang, H.; Ding, Z.; et al. PRMT1-mediated FLT3 arginine methylation promotes maintenance of FLT3-ITD + acute myeloid leukemia. Blood 2019, 134, 548–560. [Google Scholar] [CrossRef]
- Matsubara, H.; Fukuda, T.; Awazu, Y.; Nanno, S.; Shimomura, M.; Inoue, Y.; Yamauchi, M.; Yasui, T.; Sumi, T. PRMT1 expression predicts sensitivity to platinum-based chemotherapy in patients with ovarian serous carcinoma. Oncol. Lett. 2021, 21, 1. [Google Scholar] [CrossRef]
- Seligson, D.B.; Horvath, S.; Shi, T.; Yu, H.; Tze, S.; Grunstein, M.; Kurdistani, S.K. Global histone modification patterns predict risk of prostate cancer recurrence. Nature 2005, 435, 1262–1266. [Google Scholar] [CrossRef]
- Cheng, D.; Yadav, N.; King, R.W.; Swanson, M.S.; Weinstein, E.J.; Bedford, M.T. Small Molecule Regulators of Protein Arginine Methyltransferases. J. Biol. Chem. 2004, 279, 23892–23899. [Google Scholar] [CrossRef]
- Eram, M.S.; Shen, Y.; Szewczyk, M.; Wu, H.; Senisterra, G.; Li, F.; Butler, K.V.; Kaniskan, H.Ü.; Speed, B.A.; Seña, C.; et al. A Potent, Selective and Cell-active Inhibitor of Human Type I Protein Arginine Methyltransferases. ACS Chem. Biol. 2016, 11, 772. [Google Scholar] [CrossRef]
- Fong, J.Y.; Pignata, L.; Goy, P.-A.; Kawabata, K.C.; Lee, S.C.-W.; Koh, C.M.; Musiani, D.; Massignani, E.; Kotini, A.G.; Penson, A.; et al. Therapeutic Targeting of RNA Splicing Catalysis through Inhibition of Protein Arginine Methylation. Cancer Cell 2019, 36, 194. [Google Scholar] [CrossRef] [PubMed]
- Fedoriw, A.; Rajapurkar, S.R.; O’Brien, S.; Gerhart, S.V.; Mitchell, L.H.; Adams, N.D.; Rioux, N.; Lingaraj, T.; Ribich, S.A.; Pappalardi, M.B.; et al. Anti-tumor Activity of the Type I PRMT Inhibitor, GSK3368715, Synergizes with PRMT5 Inhibition through MTAP Loss. Cancer Cell 2019, 36, 100–114.e25. [Google Scholar] [CrossRef]
- Zhang, P.; Tao, H.; Yu, L.; Zhou, L.; Zhu, C. Developing protein arginine methyltransferase 1 (PRMT1) inhibitor TC-E-5003 as an antitumor drug using INEI drug delivery systems. Drug Deliv. 2020, 27, 491–501. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.; Qian, K.; Ho, M.-C.; Zheng, Y.G. Small Molecule Inhibitors of Protein Arginine Methyltransferases. Expert Opin. Investig. Drugs 2016, 25, 335. [Google Scholar] [CrossRef] [PubMed]
| Name | Mechanism of Action | Target(s) | IC50 | Reference |
|---|---|---|---|---|
| AMI-1 | Substrate competitive SAM uncompetitive | PRMT1 | 8.81 µM | [162] |
| MS023 | Substrate competitive SAM uncompetitive | PRMT1 | 30 nM | [163] |
| PRMT3 | 119 nM | |||
| PRMT4/CARM1 | 83 nM | |||
| PRMT6 | 4 nM | |||
| PRMT8 | 5 nM | |||
| GSK3368715 | Substrate competitive SAM uncompetitive Reversible | PRMT1 | 33.1 nM | [165] |
| PRMT3 | 162 nM | |||
| PRMT4/CARM1 | 38 nM | |||
| PRMT6 | 4.7 nM | |||
| PRMT8 | 3.9 nM | |||
| TC-E-5003 | ND | PRMT1 | 1.5 µM | [166] |
| C7280948 | Interaction with the substrate-binding pocket | PRMT1 | 12.8 µM | [113] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Thiebaut, C.; Eve, L.; Poulard, C.; Le Romancer, M. Structure, Activity, and Function of PRMT1. Life 2021, 11, 1147. https://doi.org/10.3390/life11111147
Thiebaut C, Eve L, Poulard C, Le Romancer M. Structure, Activity, and Function of PRMT1. Life. 2021; 11(11):1147. https://doi.org/10.3390/life11111147
Chicago/Turabian StyleThiebaut, Charlène, Louisane Eve, Coralie Poulard, and Muriel Le Romancer. 2021. "Structure, Activity, and Function of PRMT1" Life 11, no. 11: 1147. https://doi.org/10.3390/life11111147
APA StyleThiebaut, C., Eve, L., Poulard, C., & Le Romancer, M. (2021). Structure, Activity, and Function of PRMT1. Life, 11(11), 1147. https://doi.org/10.3390/life11111147







