Rocket Science: The Effect of Spaceflight on Germination Physiology, Ageing, and Transcriptome of Eruca sativa Seeds
Abstract
1. Introduction
2. Materials and Methods
2.1. Seed Material, Space Travel and Packaging
2.2. Controlled Artificial Ageing Treatment (CAAT)
2.3. Germination Assays
2.4. Conductivity Assay
2.5. RNA Extraction and Sequencing
2.6. Sequencing Quality Control, de Novo Assembly and Differentially Expressed Gene (DEG) Detection
2.7. Carotenoid Tocopherol and Chlorophyll Determinations
3. Results
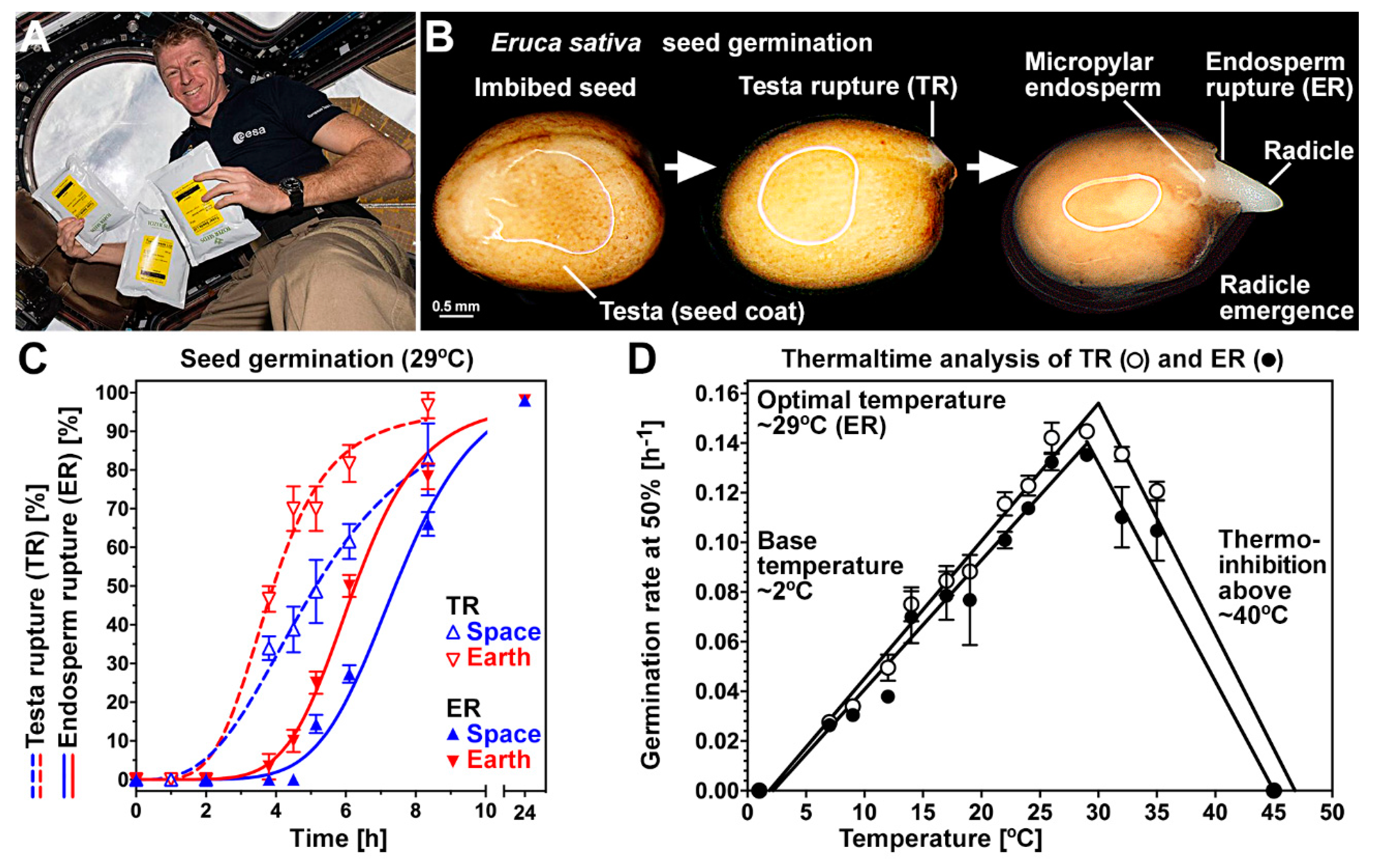
3.1. Germination of Rocket Seeds Following Spaceflight
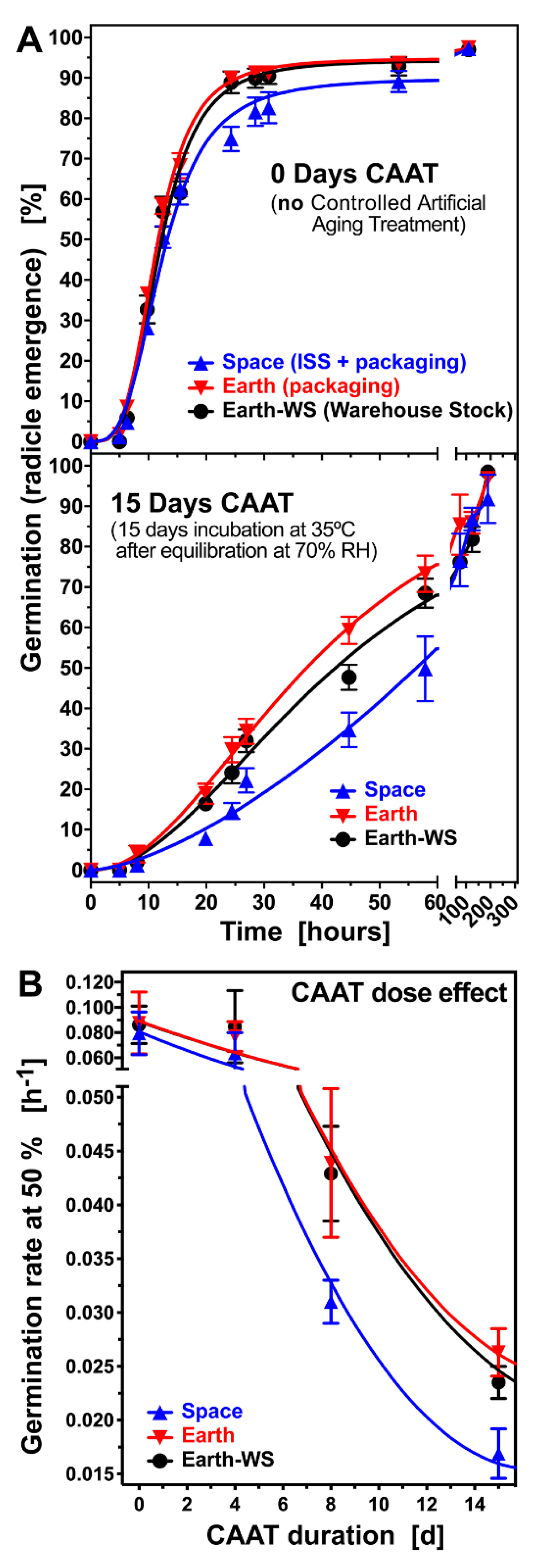
3.2. Effect of Controlled Artificial Ageing on Earth and Space Seed Germination
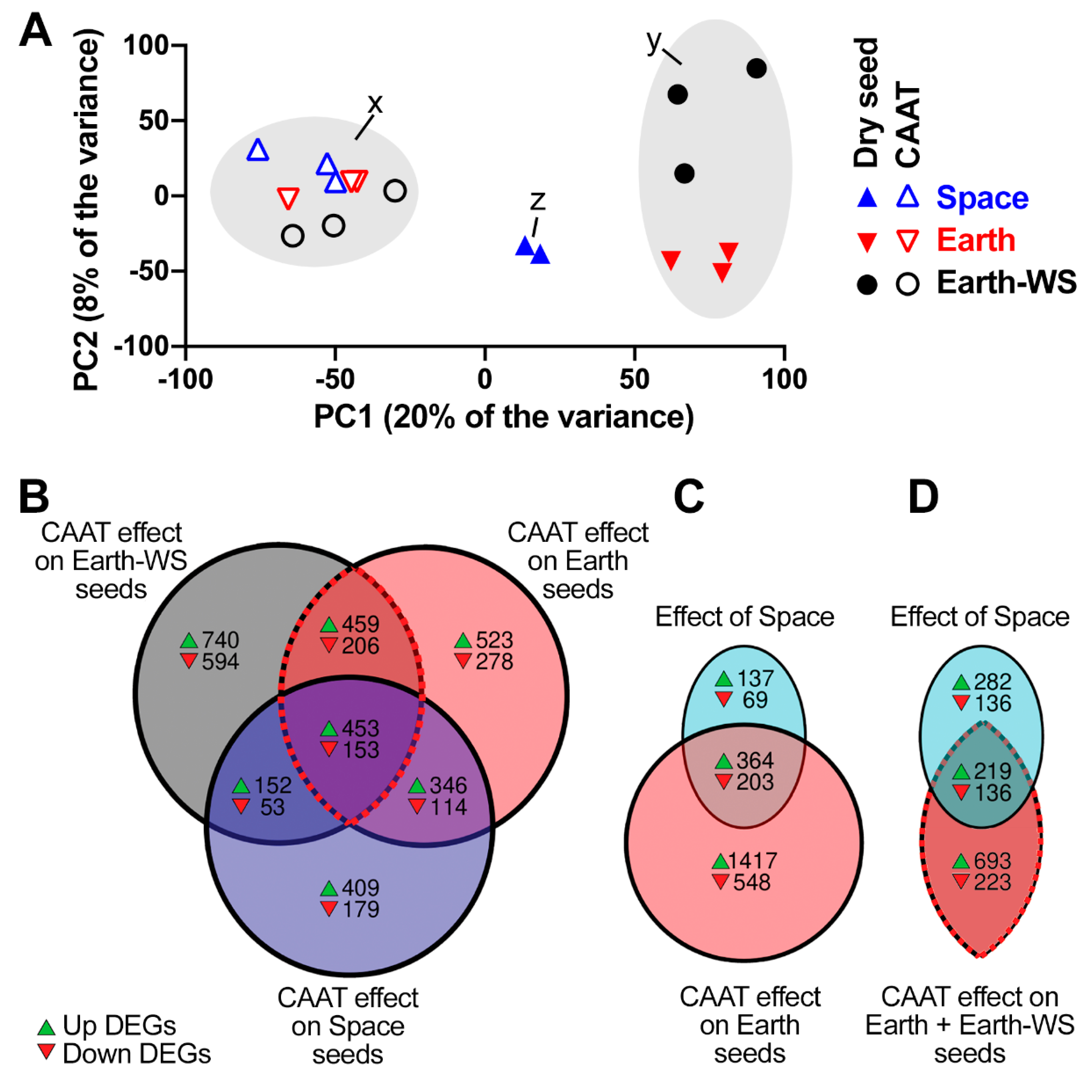
3.3. Overall Effect of Spaceflight and Controlled Accelerated Ageing on the Seed Transcriptomes
3.4. Functional Gene Groups Associated with Transcriptome Changes during Ageing and Spaceflight
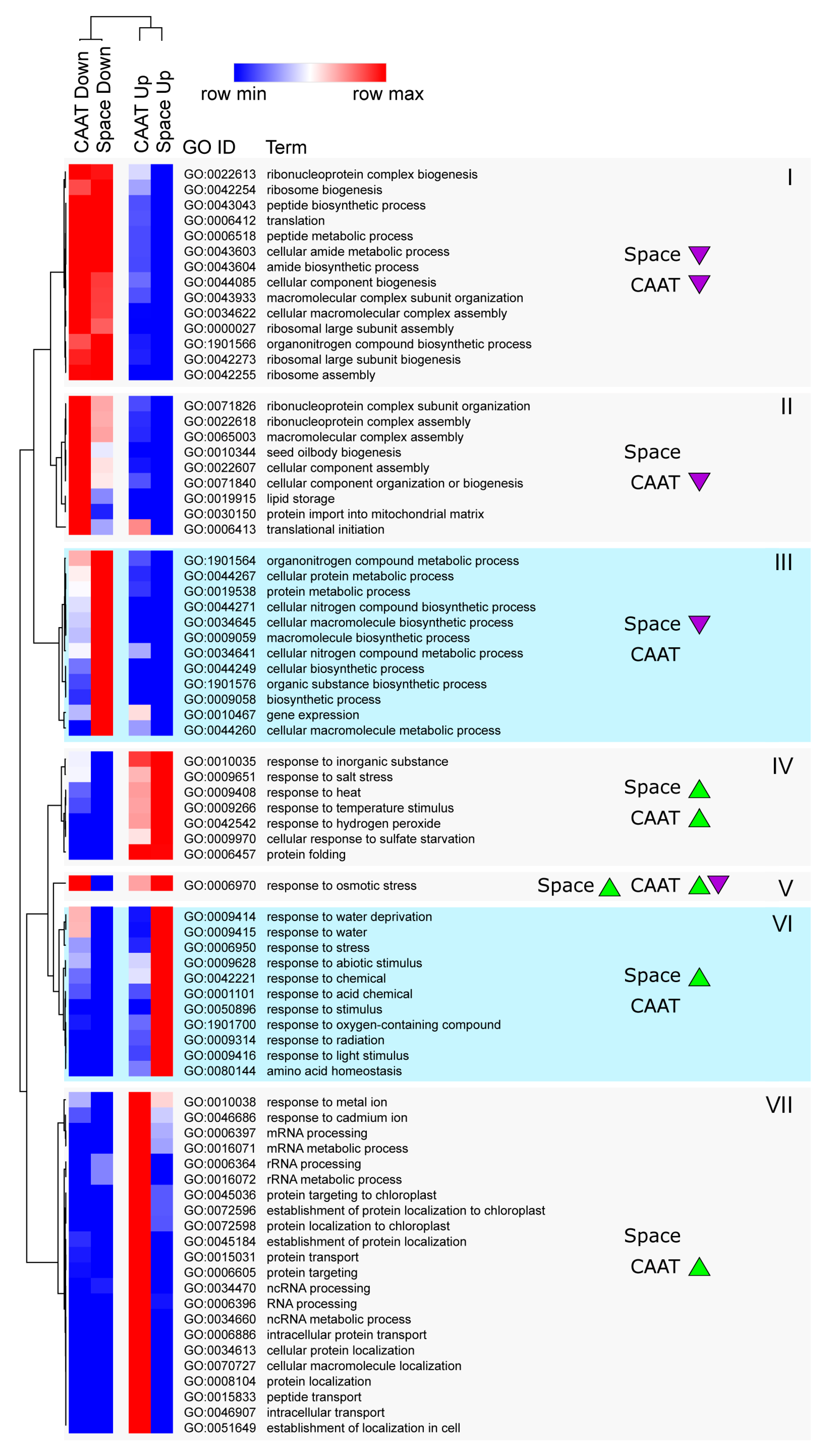
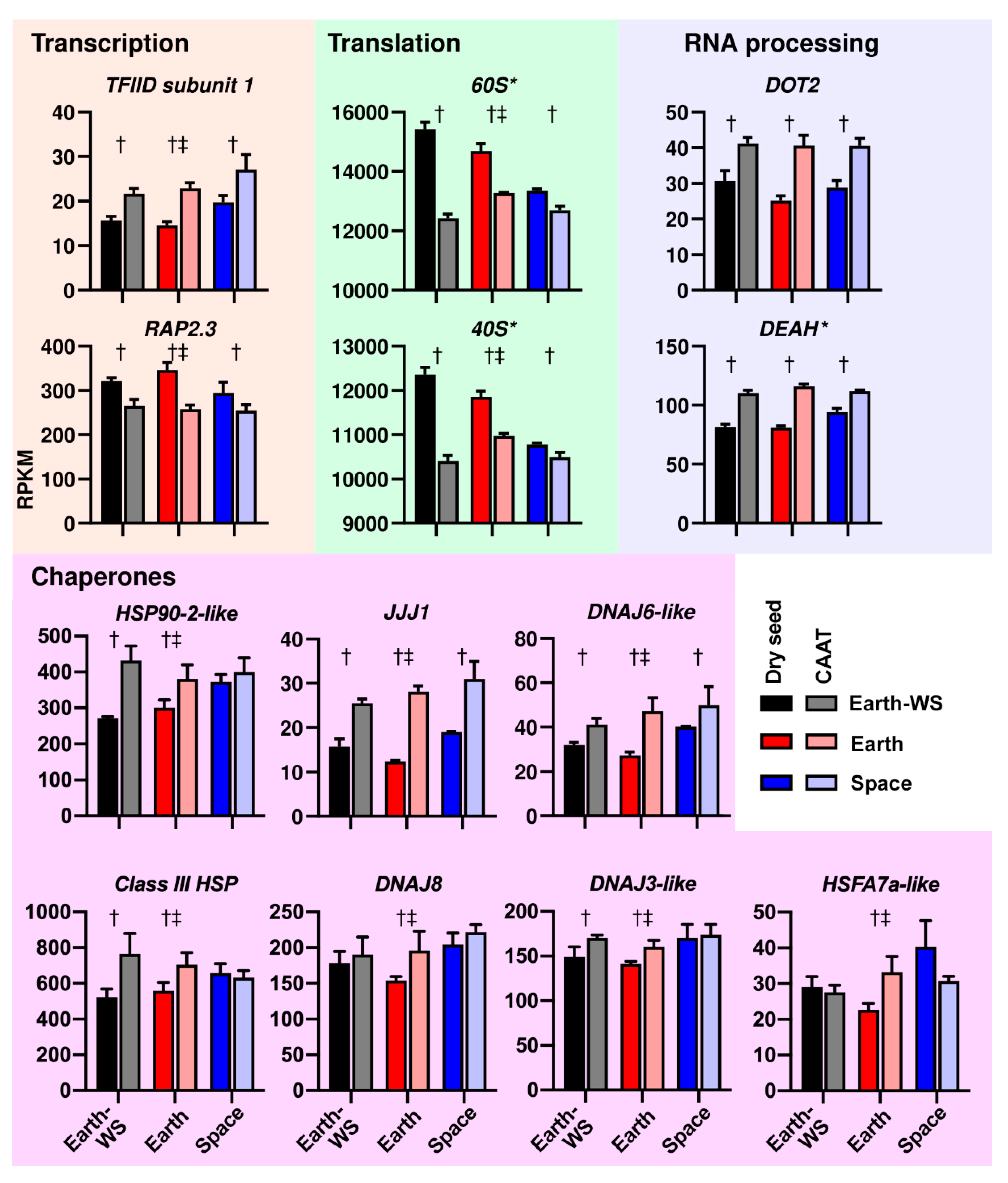
3.5. Processes Affected at the Transcript Abundance Level by Ageing and Spaceflight
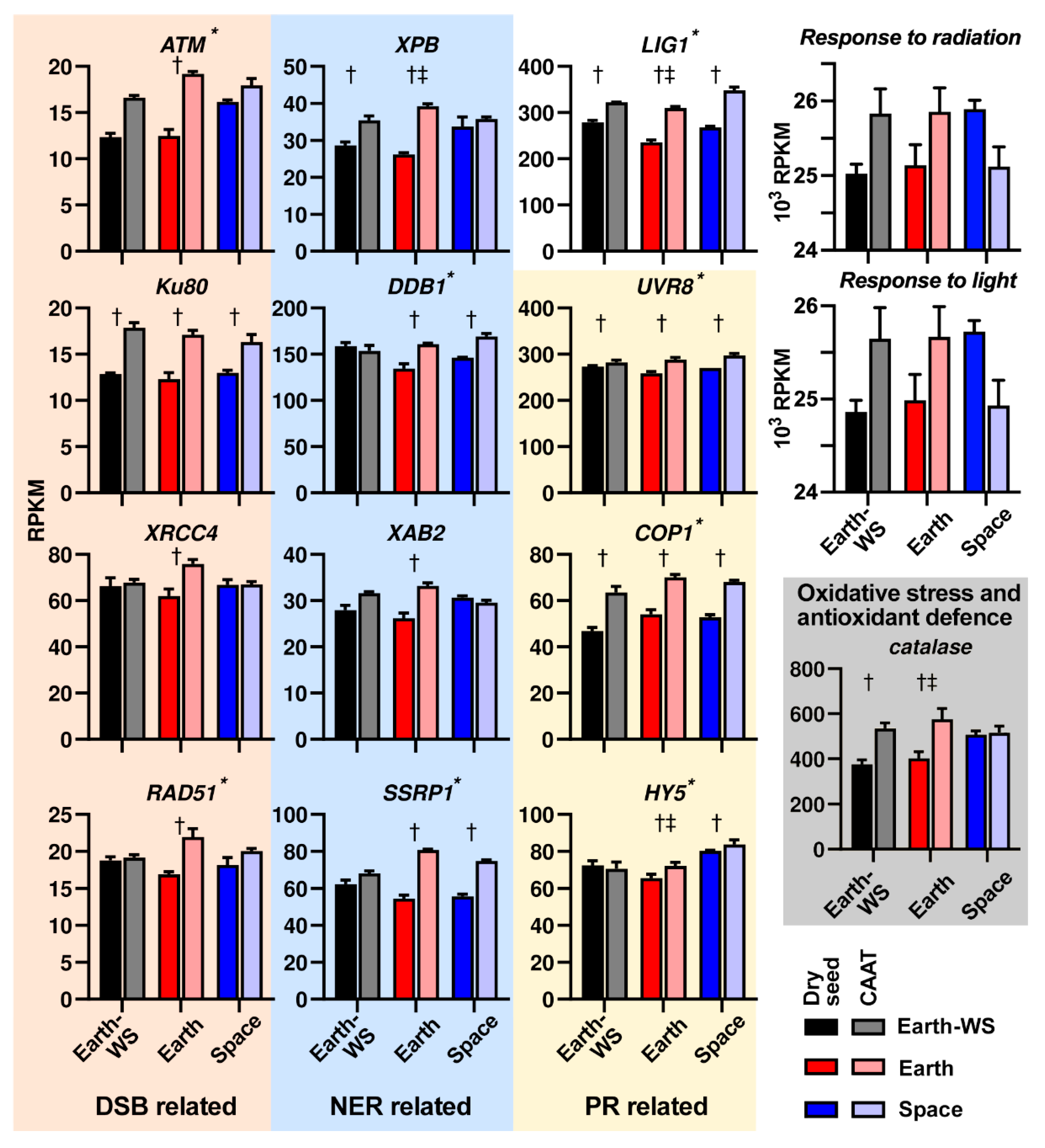
3.6. Expression of DNA Repair Related Pathways Following Ageing and Spaceflight
4. Discussion
4.1. Effects of Spaceflight Environmental Factors on Dry Stored Seeds
4.2. Effects of Spaceflight Associated Ageing on the Seed Transcriptome
5. Conclusions and Future Directions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Data Availability Statement
Abbreviations
References
- Visscher, A.M.; Seal, C.E.; Newton, R.J.; Frances, A.L.; Pritchard, H.W. Dry seeds and environmental extremes: Consequences for seed lifespan and germination. Funct. Plant Biol. 2016, 43, 656–668. [Google Scholar] [CrossRef]
- Vandenbrink, J.P.; Kiss, J.Z. Space, the final frontier: A critical review of recent experiments performed in microgravity. Plant Sci. 2016, 243, 115–119. [Google Scholar] [CrossRef] [PubMed]
- Paul, A.L.; Wheeler, R.M.; Levine, H.G.; Ferl, R.J. Fundamental plant biology enabled by the space shuttle. Am. J. Bot. 2013, 100, 226–234. [Google Scholar] [CrossRef]
- Kordyum, E.; Chapman, D.; Brykova, V. Plant cell development and aging may accelerate in microgravity. Acta Astronaut. 2019, 157, 157–161. [Google Scholar] [CrossRef]
- Musgrave, M.E. Seeds in space. Seed Sci. Res. 2002, 12, 1–16. [Google Scholar] [CrossRef]
- Cannon, K.M.; Britt, D.T. Feeding one million people on Mars. New Space 2019, 7, 245–254. [Google Scholar] [CrossRef]
- Wheeler, R.M. Agriculture for space: People and places paving the way. Open Agric. 2017, 2, 14–32. [Google Scholar] [CrossRef]
- Paul, A.L.; Sng, N.J.; Zupanska, A.K.; Krishnamurthy, A.; Schultz, E.R.; Ferl, R.J. Genetic dissection of the Arabidopsis spaceflight transcriptome: Are some responses dispensable for the physiological adaptation of plants to spaceflight? PLoS ONE 2017, 12, e0180186. [Google Scholar] [CrossRef]
- Herranz, R.; Vandenbrink, J.P.; Villacampa, A.; Manzano, A.; Poehlman, W.L.; Feltus, F.A.; Kiss, J.Z.; Medina, F.J. RNAseq analysis of the response of Arabidopsis thaliana to fractional gravity under blue-light stimulation during spaceflight. Front. Plant Sci. 2019, 10, 1529. [Google Scholar] [CrossRef]
- Zhou, M.Q.; Sng, N.J.; LeFrois, C.E.; Paul, A.L.; Ferl, R.J. Epigenomics in an extraterrestrial environment: Organ-specific alteration of DNA methylation and gene expression elicited by spaceflight in Arabidopsis thaliana. BMC Genom. 2019, 20, 205. [Google Scholar] [CrossRef] [PubMed]
- Zupanska, A.K.; LeFrois, C.; Ferl, R.J.; Paul, A.L. HSFA2 functions in the physiological adaptation of undifferentiated plant cells to spaceflight. Int. J. Mol. Sci. 2019, 20, 390. [Google Scholar] [CrossRef] [PubMed]
- El-Nakhel, C.; Giordano, M.; Pannico, A.; Carillo, P.; Fusco, G.M.; De Pascale, S.; Rouphael, Y. Cultivar-specific performance and qualitative descriptors for butterhead Salanova lettuce produced in closed soilless cultivation as a candidate salad crop for human life Support in Space. Life 2019, 9, 61. [Google Scholar] [CrossRef] [PubMed]
- Wolff, S.A.; Palma, C.F.; Marcelis, L.; Kittang Jost, A.I.; van Delden, S.H. Testing new concepts for crop cultivation in space: Effects of rooting volume and nitrogen availability. Life 2018, 8, 45. [Google Scholar] [CrossRef] [PubMed]
- Odeh, R.; Guy, C.L. Gardening for therapeutic people-plant interactions during long-duration space missions. Open Agric. 2017, 2, 1–13. [Google Scholar] [CrossRef]
- Colla, G.; Battistelli, A.; Proietti, S.; Moscatello, S.; Rouphael, Y.; Cardarelli, M.; Casucci, M. Rocket seedling production on the International Space Station: Growth and Nutritional properties. Microgravity Sci. Technol. 2007, 19, 118–121. [Google Scholar] [CrossRef]
- Zabel, P.; Bamsey, M.; Schubert, D.; Tajmar, M. Review and analysis of over 40 years of space plant growth systems. Life Sci. Space Res. 2016, 10, 1–16. [Google Scholar] [CrossRef]
- Shen, Y.Z.; Guo, S.S.; Zhao, P.S.; Wang, L.J.; Wang, X.X.; Li, J.; Bian, Q. Research on lettuce growth technology onboard Chinese Tiangong II Spacelab. Acta Astronaut. 2018, 144, 97–102. [Google Scholar] [CrossRef]
- Weitbrecht, K.; Müller, K.; Leubner-Metzger, G. First off the mark: Early seed germination. J. Exp. Bot. 2011, 62, 3289–3309. [Google Scholar] [CrossRef]
- Nagel, M.; Börner, A. The longevity of crop seeds stored under ambient conditions. Seed Sci. Res. 2010, 20, 1–12. [Google Scholar] [CrossRef]
- Kranner, I.; Minibayeva, F.V.; Beckett, R.P.; Seal, C.E. What is stress? Concepts, definitions and applications in seed science. New Phytol. 2010, 188, 655–673. [Google Scholar] [CrossRef]
- Sano, N.; Rajjou, L.; North, H.M.; Debeaujon, I.; Marion-Poll, A.; Seo, M. Staying alive: Molecular aspects of seed longevity. Plant Cell Physiol. 2016, 57, 660–674. [Google Scholar] [CrossRef] [PubMed]
- Bradford, K.J.; Dahal, P.; van Asbrouck, J.; Kunusoth, K.; Bello, P.; Thompson, J.; Wu, F. The dry chain: Reducing postharvest losses and improving food safety in humid climates. Trends Food Sci. Technol. 2018, 71, 84–93. [Google Scholar] [CrossRef]
- Fleming, M.B.; Patterson, E.L.; Reeves, P.A.; Richards, C.M.; Gaines, T.A.; Walters, C. Exploring the fate of mRNA in aging seeds: Protection, destruction, or slow decay? J. Exp. Bot. 2018, 69, 4309–4321. [Google Scholar] [CrossRef] [PubMed]
- Fleming, M.B.; Hill, L.M.; Walters, C. The kinetics of ageing in dry-stored seeds: A comparison of viability loss and RNA degradation in unique legacy seed collections. Ann. Bot. 2019, 123, 1133–1146. [Google Scholar] [CrossRef]
- Waterworth, W.M.; Bray, C.M.; West, C.E. The importance of safeguarding genome integrity in germination and seed longevity. J. Exp. Bot. 2015, 66, 3549–3558. [Google Scholar] [CrossRef]
- Mene-Saffrane, L.; Jones, A.D.; DellaPenna, D. Plastochromanol-8 and tocopherols are essential lipid-soluble antioxidants during seed desiccation and quiescence in Arabidopsis. Proc. Natl. Acad. Sci. USA 2010, 107, 17815–17820. [Google Scholar] [CrossRef]
- Steinbrecher, T.; Leubner-Metzger, G. The biomechanics of seed germination. J. Exp. Bot. 2017, 68, 765–783. [Google Scholar] [CrossRef]
- Nicholson, W.L.; Ricco, A.J. Nanosatellites for biology in space: In situ measurement of Bacillus subtilis spore germination and growth after 6 months in Low Earth Orbit on the O/OREOS mission. Life 2019, 10, 1. [Google Scholar] [CrossRef]
- Weronika, E.; Lukasz, K. Tardigrades in space research-past and future. Orig. Life Evol. B 2017, 47, 545–553. [Google Scholar] [CrossRef]
- Tepfer, D.; Leach, S. Survival and DNA damage in plant seeds exposed for 558 and 682 days outside the International Space Station. Astrobiology 2017, 17, 205–215. [Google Scholar] [CrossRef]
- Tepfer, D.; Zalar, A.; Leach, S. Survival of plant seeds, their UV screens, and nptII DNA for 18 months outside the International Space Station. Astrobiology 2012, 12, 517–528. [Google Scholar] [CrossRef]
- Zimmermann, M.W.; Gartenbach, K.E.; Kranz, A.R. First radiobiological results of Ldef-1 experiment-A0015 with Arabidopsis seed embryos and Sordaria fungus spores. Adv. Space Res. 1994, 14, 47–51. [Google Scholar] [CrossRef]
- Sugimoto, M.; Oono, Y.; Kawahara, Y.; Gusev, O.; Maekawa, M.; Matsumoto, T.; Levinskikh, M.; Sychev, V.; Novikova, N.; Grigoriev, A. Gene expression of rice seeds surviving 13- and 20-month exposure to space environment. Life Sci. Space Res. 2016, 11, 10–17. [Google Scholar] [CrossRef] [PubMed]
- Cui, J.; Xia, W.Y.; Wei, S.Y.; Zhang, M.; Wang, W.; Zeng, D.Y.; Liu, M.Y.; Sun, Y.Q.; Lu, W.H. Photosynthetic performance of rice seedlings originated from seeds exposed to spaceflight conditions. Photochem. Photobiol. 2019, 95, 1205–1212. [Google Scholar] [CrossRef] [PubMed]
- Xiong, H.C.; Guo, H.J.; Xie, Y.D.; Zhao, L.S.; Gu, J.Y.; Zhao, S.R.; Li, J.H.; Liu, L.X. RNAseq analysis reveals pathways and candidate genes associated with salinity tolerance in a spaceflight-induced wheat mutant. Sci. Rep. 2017, 7, 2731. [Google Scholar] [CrossRef] [PubMed]
- RHS. Rocket Science: Our Voyage to Discovery; Royal Horticultural Society: London, UK, 2016; Available online: https://schoolgardening.rhs.org.uk/getmedia/a3385b8e-0eaf-4953-8d90-bc163ff0f982/Final-Rocket-Science-Report-Low-Res (accessed on 31 March 2020).
- Mira, S.; Estrelles, E.; Gonzalez-Benito, M.E. Effect of water content and temperature on seed longevity of seven Brassicaceae species after 5 years of storage. Plant Biol 2015, 17, 153–162. [Google Scholar] [CrossRef] [PubMed]
- Matthews, S.; Powell, A. Electrical conductivity vigour test: Physiological basis and use. Seed Test. Int. 2006, 131, 32–35. [Google Scholar]
- Graeber, K.; Linkies, A.; Wood, A.T.; Leubner-Metzger, G. A guideline to family-wide comparative state-of-the-art quantitative RT-PCR analysis exemplified with a Brassicaceae cross-species seed germination case study. Plant Cell 2011, 23, 2045–2063. [Google Scholar] [CrossRef] [PubMed]
- Andrews, S. Bioinformatics tools of the Babraham Institute. 2019. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 10 September 2019).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics (Oxford, England) 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Wu, T.D.; Nacu, S. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics (Oxford, England) 2010, 26, 873–881. [Google Scholar] [CrossRef]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 2015, 12, 59–60. [Google Scholar] [CrossRef] [PubMed]
- Huson, D.H.; Auch, A.F.; Qi, J.; Schuster, S.C. MEGAN analysis of metagenomic data. Genome Res. 2007, 17, 377–386. [Google Scholar] [CrossRef]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef]
- Simao, F.A.; Waterhouse, R.M.; Ioannidis, P.; Kriventseva, E.V.; Zdobnov, E.M. BUSCO: Assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics (Oxford, England) 2015, 31, 3210–3212. [Google Scholar] [CrossRef] [PubMed]
- TimeLogic Tera-BLAST, Active Motif Inc., Carlsbad (CA, USA). 2019. Available online: http://www.timelogic.com/catalog/757/tera-blast (accessed on 10 September 2019).
- Conesa, A.; Gotz, S.; Garcia-Gomez, J.M.; Terol, J.; Talon, M.; Robles, M. Blast2GO: A universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics (Oxford, England) 2005, 21, 3674–3676. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. FeatureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics (Oxford, England) 2014, 30, 923–930. [Google Scholar] [CrossRef]
- Tarazona, S.; Furio-Tari, P.; Turra, D.; Di Pietro, A.; Nueda, M.J.; Ferrer, A.; Conesa, A. Data quality aware analysis of differential expression in RNA-seq with NOISeq R/Bioc package. Nucleic Acids Res. 2015, 43. [Google Scholar] [CrossRef]
- Chamberlain, S.; Szocs, E.; Boettiger, C.; Ram, K.; Bartomeus, I.; Baumgartner, J.; Foster, Z.; O’Donnell, J.; Oksanen, J. Taxize: Taxonomic Information from around the Web. R package Version 0.9.0. 2017. Available online: https://github.com/ropensci/taxize (accessed on 10 September 2019).
- Alexa, A.; Rahnenfuhrer, J. topGO: Enrichment Analysis for Gene Ontology. R package version 2.38.1. 2019. Available online: https://www.mpi-inf.mpg.de/home/ (accessed on 20 December 2019).
- Morpheus Heat Map Tool. 2019. Available online: https://software.broadinstitute.org/morpheus/ (accessed on 20 December 2019).
- Drapal, M.; Rossel, G.; Heider, B.; Fraser, P.D. Metabolic diversity in sweet potato (Ipomoea batatas, Lam.) leaves and storage roots. Hortic. Res. Engl. 2019, 6, 2. [Google Scholar] [CrossRef]
- Bligh, E.G.; Dyer, W.J. A rapid method of total lipid extraction and purification. Canadian J. Biochem. Physiol. 1959, 37, 911–917. [Google Scholar] [CrossRef]
- Fraser, P.D.; Pinto, M.E.; Holloway, D.E.; Bramley, P.M. Technical advance: Application of high-performance liquid chromatography with photodiode array detection to the metabolic profiling of plant isoprenoids. Plant J. 2000, 24, 551–558. [Google Scholar] [CrossRef] [PubMed]
- Leubner-Metzger, G. ß-1,3-Glucanase gene expression in low-hydrated seeds as a mechanism for dormancy release during tobacco after-ripening. Plant J. 2005, 41, 133–145. [Google Scholar] [CrossRef] [PubMed]
- Finch-Savage, W.E.; Bassel, G.W. Seed vigour and crop establishment: Extending performance beyond adaptation. J. Exp. Bot. 2016, 67, 567–591. [Google Scholar] [CrossRef]
- Ohtani, M.; Demura, T.; Sugiyama, M. Arabidopsis ROOT INITIATION DEFECTIVE1, a DEAH-box RNA helicase involved in pre-mRNA splicing, is essential for plant development. Plant Cell 2013, 25, 2056–2069. [Google Scholar] [CrossRef]
- Kozeko, L.; Talalaiev, O.; Neimash, V.; Povarchuk, V. A protective role of HSP90 chaperone in gamma-irradiated Arabidopsis thaliana seeds. Life Sci. Space Res. 2015, 6, 51–58. [Google Scholar] [CrossRef] [PubMed]
- Meyer, A.E.; Hoover, L.A.; Craig, E.A. The cytosolic J-protein, Jjj1, and Rei1 function in the removal of the pre-60 S subunit factor Arx1. J. Boil. Chem. 2010, 285, 961–968. [Google Scholar] [CrossRef] [PubMed]
- Waterworth, W.M.; Footitt, S.; Bray, C.M.; Finch-Savage, W.E.; West, C.E. DNA damage checkpoint kinase ATM regulates germination and maintains genome stability in seeds. Proc. Natl. Acad. Sci. USA 2016, 113, 9647–9652. [Google Scholar] [CrossRef] [PubMed]
- Waterworth, W.M.; Bray, C.M.; West, C.E. Seeds and the art of genome maintenance. Front. Plant Sci. 2019, 10, 706. [Google Scholar] [CrossRef]
- Friesner, J.; Britt, A.B. Ku80- and DNA ligase IV-deficient plants are sensitive to ionizing radiation and defective in T-DNA integration. Plant J. 2003, 34, 427–440. [Google Scholar] [CrossRef]
- Costa, R.M.A.; Morgante, P.G.; Berra, C.M.; Nakabashi, M.; Bruneau, D.; Bouchez, D.; Sweder, K.S.; Van Sluys, M.A.; Menck, C.F.M. The participation of AtXPB1, the XPB/RAD25 homologue gene from Arabidopsis thaliana, in DNA repair and plant development. Plant J. 2001, 28, 385–395. [Google Scholar] [CrossRef]
- Manova, V.; Gruszka, D. DNA damage and repair in plants-from models to crops. Front. Plant Sci. 2015, 6, 885. [Google Scholar] [CrossRef] [PubMed]
- Waterworth, W.M.; Masnavi, G.; Bhardwaj, R.M.; Jiang, Q.; Bray, C.M.; West, C.E. A plant DNA ligase is an important determinant of seed longevity. Plant J. 2010, 63, 848–860. [Google Scholar] [CrossRef] [PubMed]
- Heijde, M.; Ulm, R. UV-B photoreceptor-mediated signalling in plants. Trends Plant Sci. 2012, 17, 230–237. [Google Scholar] [CrossRef]
- Tossi, V.E.; Regalado, J.J.; Iannicelli, J.; Laino, L.E.; Burrieza, H.P.; Escandon, A.S.; Pitta-Alvarez, S.I. Beyond Arabidopsis: Differential UV-B response mediated by UVR8 in diverse species. Front. Plant Sci. 2019, 10, 780. [Google Scholar] [CrossRef] [PubMed]
- Gangappa, S.N.; Botto, J.F. The multifaceted roles of HY5 in plant gowth and development. Mol. Plant 2016, 9, 1353–1365. [Google Scholar] [CrossRef]
- Bailly, C. Active oxygen species and antioxidants in seed biology. Seed Sci. Res. 2004, 14, 93–107. [Google Scholar] [CrossRef]
- Chen, H.; Osuna, D.; Colville, L.; Lorenzo, O.; Graeber, K.; Kuster, H.; Leubner-Metzger, G.; Kranner, I. Transcriptome-wide mapping of pea seed ageing reveals a pivotal role for genes related to oxidative stress and programmed cell death. PLoS ONE 2013, 8, e78471. [Google Scholar] [CrossRef]
- Kibinza, S.; Bazin, J.; Bailly, C.; Farrant, J.M.; Corbineau, F.; El-Maarouf-Bouteau, H. Catalase is a key enzyme in seed recovery from ageing during priming. Plant Sci. 2011, 181, 309–315. [Google Scholar] [CrossRef]
- Benton, E.R.; Benton, E. Space radiation dosimetry in low-Earth orbit and beyond. Nucl. Instruments Methods Phys. Res. Sect. B 2001, 184, 255–294. [Google Scholar] [CrossRef]
- Kerr, R.A. Radiation will make astronauts’ trip to Mars even riskier. Science 2013, 340, 1031. [Google Scholar] [CrossRef]
- Bell, L.; Wagstaff, C. Rocket science: A review of phytochemical & health-related research in Eruca & Diplotaxis species. Food Chem. 2019, 1, 100002. [Google Scholar] [CrossRef]
- Gu, R.L.; Huang, R.; Jia, G.Y.; Yuan, Z.P.; Ren, L.S.; Li, L.; Wang, J.H. Effect of mechanical threshing on damage and vigor of maize seed threshed at different moisture contents. J. Integr. Agric. 2019, 18, 1571–1578. [Google Scholar] [CrossRef]
- Uchida, A.; Yamamoto, K.T. Effects of mechanical vibration on seed germination of Arabidopsis thaliana (L.) Heynh. Plant Cell Physiol. 2002, 43, 647–651. [Google Scholar] [CrossRef]
- Chancellor, J.C.; Scott, G.B.; Sutton, J.P. Space radiation: The number one risk to astronaut health beyond Low Earth Orbit. Life 2014, 4, 491–510. [Google Scholar] [CrossRef]
- Akopova, A.B.; Manaseryan, M.M.; Melkonyan, A.A.; Tatikyan, S.S.; Potapov, Y. Radiation measurement on the International Space Station. Radiat. Meas. 2005, 39, 225–228. [Google Scholar] [CrossRef]
- De Micco, V.; Paradiso, R.; Aronne, G.; De Pascale, S.; Quarto, M.; Arena, C. Leaf anatomy and photochemical behaviour of Solanum lycopersicum L. plants from seeds irradiated with low-LET ionising radiation. Sci. World J. 2014, 2014, 428141. [Google Scholar] [CrossRef]
- Norfadzrin, F.; Ahmed, O.H.; Shaharudin, S.; Abdul Rahman, D. A preliminary study on gamma radiosensitivity of tomato (Lycopersicon esculentum) and okra (Abelmoschus esculentus). Int. J. Agric. Res. 2007, 2, 620–625. [Google Scholar] [CrossRef]
- Marcu, D.; Damian, G.; Cosma, C.; Cristea, V. Gamma radiation effects on seed germination, growth and pigment content, and ESR study of induced free radicals in maize (Zea mays). J. Boil. Phys. 2013, 39, 625–634. [Google Scholar] [CrossRef] [PubMed]
- Kahn, B.A.; Stoffella, P.J. No evidence of adverse effects on germination, emergence, and fruit yield due to space exposure of tomato seeds. J. Am. Soc. Hortic. Sci. 1996, 121, 414–418. [Google Scholar] [CrossRef]
- Mei, M.; Qiu, Y.; He, Y.; Bucker, H.; Yang, C.H. Mutational effects of space-flight on Zea mays seeds. Adv. Space Res. 1994, 14, 33–39. [Google Scholar] [CrossRef]
- Nechitailo, G.S.; Lu, J.Y.; Xue, H.A.; Pan, Y.; Tang, C.Q.; Liu, M. Influence of long term exposure to space flight on tomato seeds. Adv. Space Res. 2005, 36, 1329–1333. [Google Scholar] [CrossRef]
- Iglesias-Fernandez, R.; Rodriguez-Gacio, M.D.; Matilla, A.J. Progress in research on dry afterripening. Seed Sci. Res. 2011, 21, 69–80. [Google Scholar] [CrossRef]
- Bazin, J.; Langlade, N.; Vincourt, P.; Arribat, S.; Balzergue, S.; El-Maarouf-Bouteau, H.; Bailly, C. Targeted mRNA oxidation regulates sunflower seed dormancy alleviation during dry after-ripening. Plant Cell 2011, 23, 2196–2208. [Google Scholar] [CrossRef] [PubMed]
- Nelson, S.K.; Ariizumi, T.; Steber, C.M. Biology in the dry seed: Transcriptome changes associated with dry seed dormancy and dormancy loss in the Arabidopsis GA-insensitive sleepy1-2 mutant. Front. Plant Sci. 2017, 8, 2158. [Google Scholar] [CrossRef] [PubMed]
- Mei, T.; Yang, G.; Quan, Y.; Wang, W.K.; Zhang, W.M.; Xue, J.M.; Wu, L.J.; Gu, H.Y.; Schettino, G.; Wang, Y.G. Oxidative metabolism involved in non-targeted effects induced by proton radiation in intact arabidopsis seeds. J. Radiat. Res. 2011, 52, 159–167. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kim, S.H.; Song, M.; Lee, K.J.; Hwang, S.G.; Jang, C.S.; Kim, J.B.; Kim, S.H.; Ha, B.K.; Kang, S.Y.; Kim, D.S. Genome-wide transcriptome profiling of ROS scavenging and signal transduction pathways in rice (Oryza sativa L.) in response to different types of ionizing radiation. Mol. Boil. Rep. 2012, 39, 11231–11248. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.Y.; Hou, L.T.; Jian, H.J.; Di, F.F.; Li, J.N.; Liu, L.Z. Combined QTL mapping, physiological and transcriptomic analyses to identify candidate genes involved in Brassica napus seed aging. Mol. Genet. Genom. 2018, 293, 1421–1435. [Google Scholar] [CrossRef]
- Rajjou, L.; Lovigny, Y.; Groot, S.P.; Belghazi, M.; Job, C.; Job, D. Proteome-wide characterization of seed aging in Arabidopsis: A comparison between artificial and natural aging protocols. Plant Physiol. 2008, 148, 620–641. [Google Scholar] [CrossRef]

| Group | Comparison | A | B | Down in A vs. B | Up in A vs. B |
|---|---|---|---|---|---|
| Dry seed vs. CAAT | CAAT effect on Earth-WS seeds | Earth-WS CAAT | Earth-WS | 1006 | 1804 |
| CAAT effect on Earth seeds | Earth CAAT | Earth | 751 | 1781 | |
| CAAT effect on Space seeds | Space CAAT | Space | 499 | 1360 | |
| Dry seed | Effect of Packaging and Space | Space | Earth-WS | 655 | 658 |
| Effect of Space | Space | Earth | 272 | 501 | |
| Effect of Packaging | Earth | Earth-WS | 603 | 339 | |
| CAAT | Effect of Packaging and Space | Space CAAT | Earth-WS CAAT | 45 | 343 |
| Effect of Space | Space CAAT | Earth CAAT | 32 | 12 | |
| Effect of Packaging | Earth CAAT | Earth-WS CAAT | 49 | 300 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chandler, J.O.; Haas, F.B.; Khan, S.; Bowden, L.; Ignatz, M.; Enfissi, E.M.A.; Gawthrop, F.; Griffiths, A.; Fraser, P.D.; Rensing, S.A.; et al. Rocket Science: The Effect of Spaceflight on Germination Physiology, Ageing, and Transcriptome of Eruca sativa Seeds. Life 2020, 10, 49. https://doi.org/10.3390/life10040049
Chandler JO, Haas FB, Khan S, Bowden L, Ignatz M, Enfissi EMA, Gawthrop F, Griffiths A, Fraser PD, Rensing SA, et al. Rocket Science: The Effect of Spaceflight on Germination Physiology, Ageing, and Transcriptome of Eruca sativa Seeds. Life. 2020; 10(4):49. https://doi.org/10.3390/life10040049
Chicago/Turabian StyleChandler, Jake O., Fabian B. Haas, Safina Khan, Laura Bowden, Michael Ignatz, Eugenia M. A. Enfissi, Frances Gawthrop, Alistair Griffiths, Paul D. Fraser, Stefan A. Rensing, and et al. 2020. "Rocket Science: The Effect of Spaceflight on Germination Physiology, Ageing, and Transcriptome of Eruca sativa Seeds" Life 10, no. 4: 49. https://doi.org/10.3390/life10040049
APA StyleChandler, J. O., Haas, F. B., Khan, S., Bowden, L., Ignatz, M., Enfissi, E. M. A., Gawthrop, F., Griffiths, A., Fraser, P. D., Rensing, S. A., & Leubner-Metzger, G. (2020). Rocket Science: The Effect of Spaceflight on Germination Physiology, Ageing, and Transcriptome of Eruca sativa Seeds. Life, 10(4), 49. https://doi.org/10.3390/life10040049





