An Asymmetric Bimodal Double Regression Model
Abstract
:1. Introduction
2. The GSC Regression Model
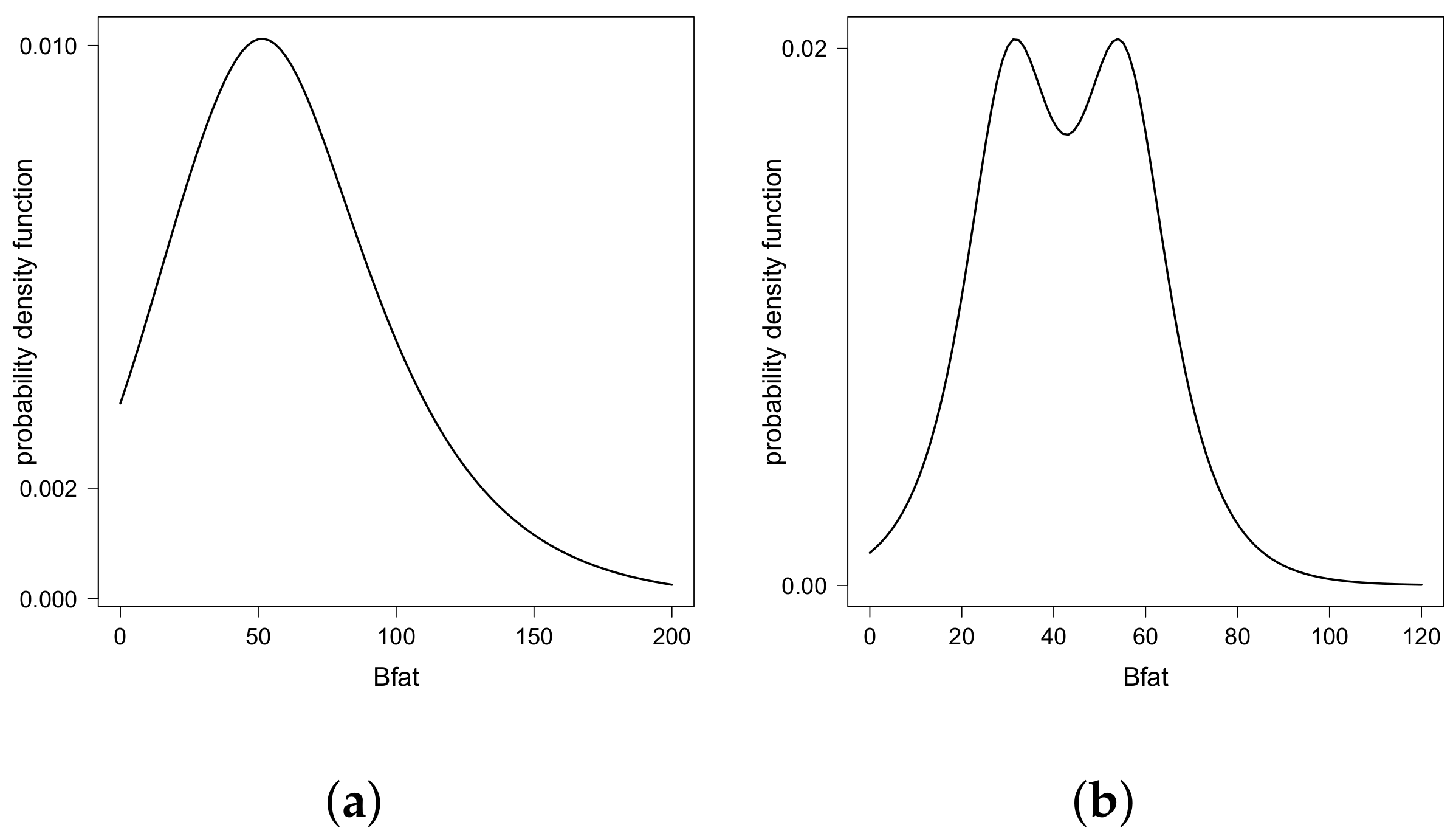
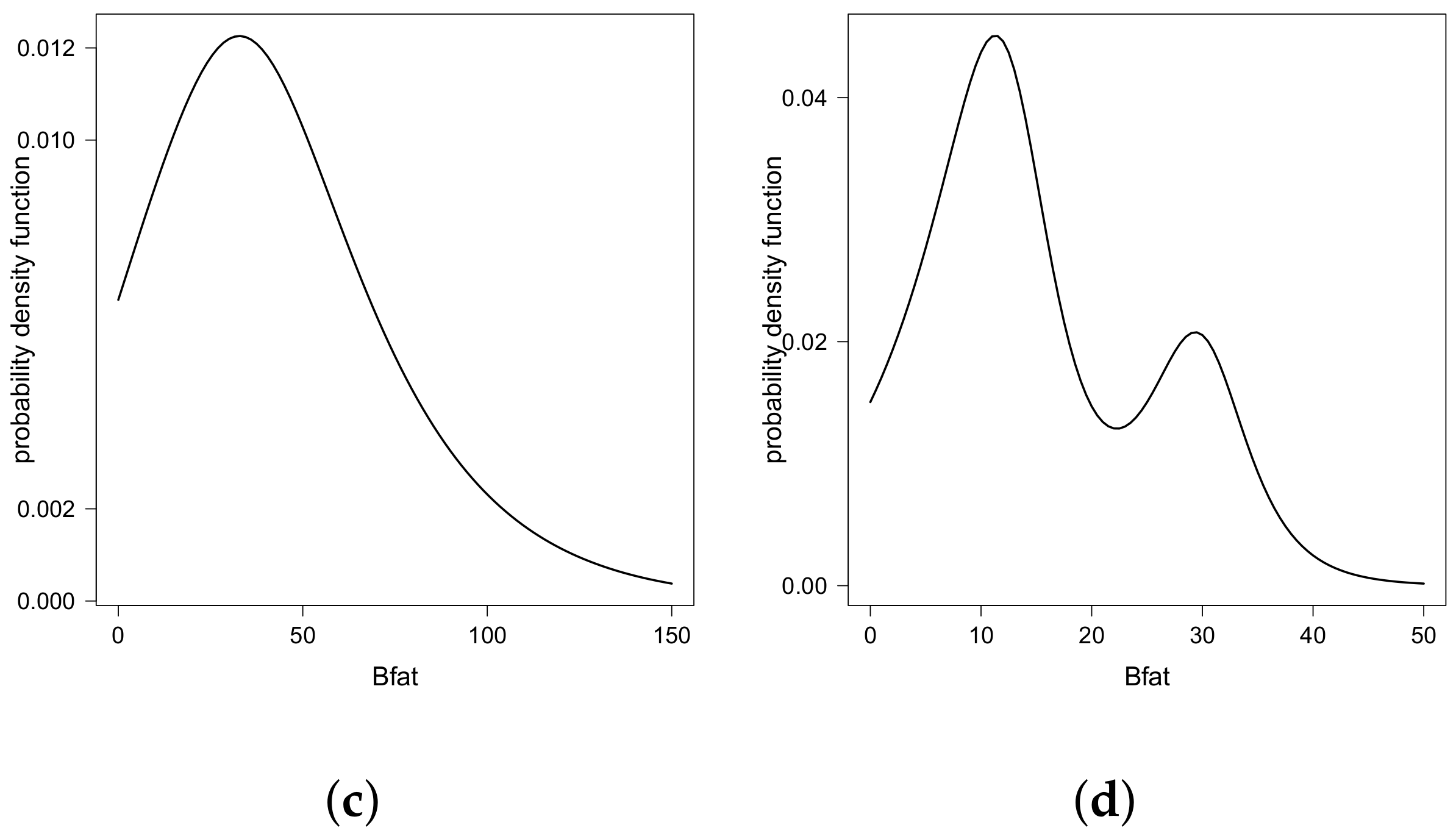
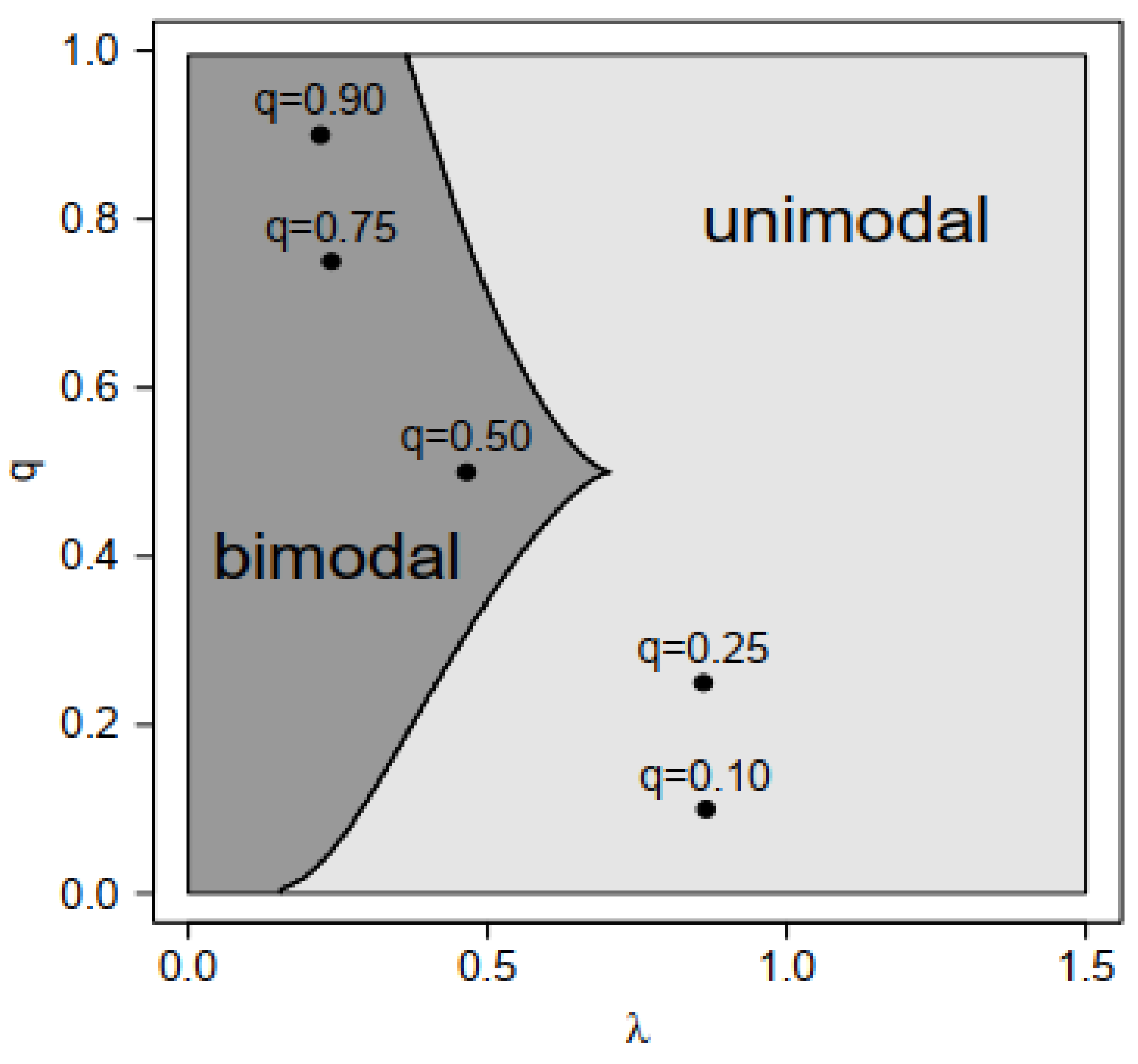
- For , the equation has three solutions. In this case, , and , . Then, are the two modes of the distribution.
- For , the equation has one solution. In this case, , and , . Then, is the only mode of the distribution.
3. Simulation Study
4. Application
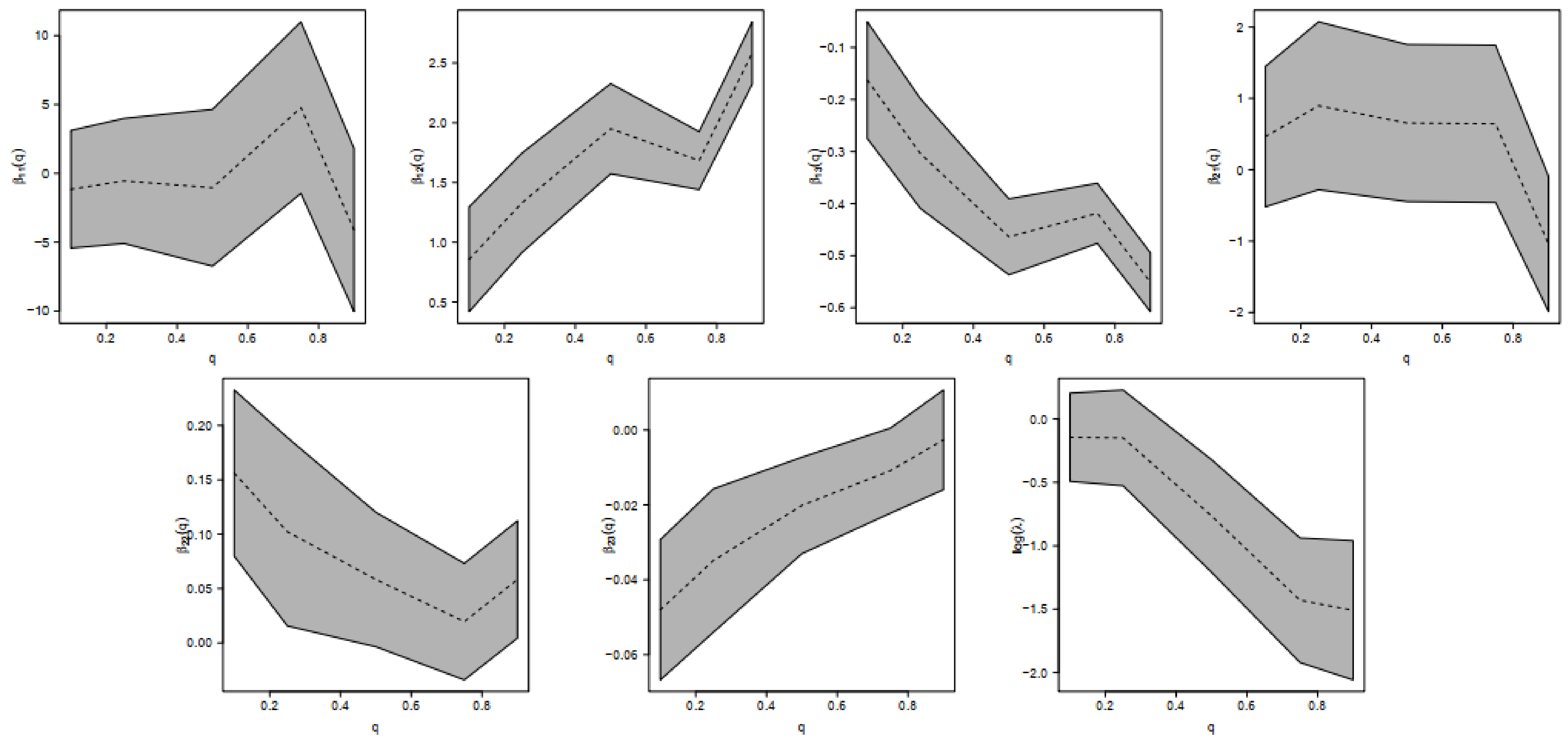
- (Interpreting ) For a fixed lbm, for athletes in the lowest 10% of Bfat, the Bfat is increased by 0.8572 units (95% confidence interval 0.4191; 1.2954) for each unit increase in bmi, and for athletes in the highest 90% of Bfat, the Bfat is increased by 2.5834 units (95% confidence interval 2.3227; 2.8442) for each unit increase in bmi.
- (Interpreting ) For a fixed bmi, for athletes in the lowest 10% of Bfat, the Bfat is decreased by 0.3039 units (95% confidence interval −0.4093; −0.1984) for each unit increase in lbm, and for athletes in the highest 10% of Bfat, the Bfat is decreased by 0.5511 units (95% confidence interval −0.6077; −0.4945) for each unit increase in lbm.
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ashman, K.M.; Bird, C.M.; Zepf, S.E. Detecting bimodality in astronomical datasets. Astron. J. 1994, 108, 2348–2361. [Google Scholar] [CrossRef] [Green Version]
- De Michele, C.; Accatino, F. Tree cover bimodality in savannas and forests emerging from the switching between two fire dynamics. PLoS ONE 2014, 9, e91195. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.; Wen, S.; Symmans, W.F.; Pusztai, L.; Coombes, K.R. The bimodality index: A criterion for discovering and ranking bimodal signatures from cancer gene expression profiling data. Cancer Inform. 2009, 7, 199–216. [Google Scholar] [CrossRef] [Green Version]
- Freeman, J.B.; Dale, R. Assessing bimodality to detect the presence of a dual cognitive process. Behav. Res. Methods 2013, 45, 83–97. [Google Scholar] [CrossRef] [Green Version]
- Sambrook-Smith, G.H.; Nicholas, A.P.; Ferguson, R.I. Measuring and defining bimodal sediments: Problems and implications. Water Resour. Res. 1997, 33, 1179–1195. [Google Scholar] [CrossRef]
- Sturrock, P.A. Analysis of bimodality in histograms formed from GALLEX and GNO solar neutrino data. Sol. Phys. 2008, 249, 1–10. [Google Scholar] [CrossRef] [Green Version]
- Gómez, H.W.; Elal-Olivero, D.; Salinas, H.S.; Bolfarine, H. Bimodal extension based on the skew-normal distribution with application to pollen data. Environmetrics 2018, 22, 50–62. [Google Scholar] [CrossRef]
- Venegas, O.; Salinas, H.S.; Gallardo, D.I.; Bolfarine, H.; Gómez, H.W. Bimodality based on the generalized skew-normal distribution. J. Stat. Comput. Simul. 2018, 88, 156–181. [Google Scholar] [CrossRef]
- Butt, N.S.; Khalil, M.G. A new bimodal distribution for modeling asymmetric bimodal heavy-tail real lifetime data. Symmetry 2020, 12, 2058. [Google Scholar] [CrossRef]
- Iriarte, Y.A.; de Castro, M.; Gómez, H.W. A Unimodal/Bimodal Skew/Symmetric Distribution Generated from Lambert’s Transformation. Symmetry 2021, 13, 269. [Google Scholar] [CrossRef]
- Reyes, J.; Gómez-Déniz, E.; Gómez, H.W.; Calderín-Ojeda, E. A bimodal extension of the exponential distribution with applications in risk theory. Symmetry 2021, 13, 679. [Google Scholar] [CrossRef]
- Reyes, J.; Arrué, J.; Leiva, V.; Martín-Barreiro, C. A new Birnbaum–Saunders distribution and its mathematical features applied to bimodal real-world data from environment and medicine. Mathematics 2021, 9, 1891. [Google Scholar] [CrossRef]
- Gómez, Y.; Gómez-Déniz, E.; Venegas, O.; Gallardo, D.I.; Gómez, H.W. An asymmetric bimodal distribution with application to quantile regression. Symmetry 2019, 11, 899. [Google Scholar] [CrossRef] [Green Version]
- Bayes, C.L.; García, C. A new robust regression model for proportions. Bayesian Anal. 2012, 7, 841–866. [Google Scholar] [CrossRef]
- Bayes, C.L.; Bazán, J.L.; de Castro, M. A quantile parametric mixed regression model for bounded response variables. Stat. Interface 2017, 10, 483–493. [Google Scholar] [CrossRef]
- Sánchez, L.; Leiva, V.; Galea, M.; Saulo, H. Birnbaum-Saunders quantile regression and its diagnostics with application to economic data. Appl. Stoch. Model. Bus. Ind. 2021, 37, 53–73. [Google Scholar] [CrossRef]
- Bernardi, M.; Bottone, M.; Petrella, L. Bayesian quantile regression using the skew exponential power distribution. Comput. Stat. Data Anal. 2018, 126, 92–111. [Google Scholar] [CrossRef] [Green Version]
- Korkmaz, M.Ç.; Chesneau, C.; Korkmaz, Z.S. On the arcsecant hyperbolic normal distribution. Properties, quantile regression modeling and applications. Symmetry 2021, 13, 117. [Google Scholar] [CrossRef]
- Cade, B.S.; Noon, B.R. A gentle introduction to quantile regression for ecologists. Front. Ecol. Environ. 2003, 1, 412–420. [Google Scholar] [CrossRef]
- Xiao, Z.; Guo, H.; Lam, M.S. Quantile regression and value at risk. In Handbook of Financial Econometrics and Statistics; Lee, C.F., Lee, J., Eds.; Springer: New York, NY, USA, 2015; pp. 1143–1167. [Google Scholar]
- Alencar, A.P.; Santos, B.R. Association of pollution with quantiles and expectations of the hospitalization rate of elderly people by respiratory diseases in the city of São Paulo, Brazil. Environmetrics 2014, 25, 165–171. [Google Scholar] [CrossRef]
- Wei, Y.; Pere, A.; Koenker, R.; He, X. Quantile regression methods for reference growth charts. Stat. Med. 2005, 25, 1369–1382. [Google Scholar] [CrossRef] [PubMed]
- Bourguignon, M.; Gallardo, D.I.; Medeiros, R.M.R. A simple and useful regression model for underdispersed count data based on Bernoulli–Poisson convolution. Stat. Pap. 2021, in press. [Google Scholar] [CrossRef]
- Bourguignon, M.; Santos-Neto, M.; de Castro, M. A new regression model for positive random variables with skewed and long tail. METRON 2021, 79, 33–55. [Google Scholar] [CrossRef]
- Bourguignon, M.; Leão, J.; Gallardo, D.I. Parametric modal regression with varying precision. Biom. J. 2020, 62, 2002–2020. [Google Scholar] [CrossRef]
- Mittelhammer, R.C.; Jodge, G.G.; Miller, D.J. Econometric Foundations; Cambridge University Press: New York, NY, USA, 2000. [Google Scholar]
- Core Team, R. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2021. [Google Scholar]
- Azzalini, A. The R Package sn: The Skew-Normal and Related Distributions Such as the Skew-t and the SUN (Version 2.0.0); Università di Padova: Padova, Italy, 2021. [Google Scholar]
- Galarza, C.E.; Lachos, V.H.; Barbosa, C.; Castro, L.M. Robust quantile regression using a generalized class of skewed distributions. Stat 2017, 6, 113–130. [Google Scholar]
- Akaike, H. A new look at the statistical model identification. IEEE Trans. Autom. Control 1974, 19, 379–397. [Google Scholar] [CrossRef]
- Dunn, P.K.; Smyth, G.K. Randomized Quantile Residuals. J. Comput. Graph. Stat. 1996, 5, 236–244. [Google Scholar]

| q | Parameter | Bias | SE | RMSE | CP | Bias | SE | RMSE | CP | Bias | SE | RMSE | CP | |||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0.10 | (1, −1, 0.5) | (−1, 1.6, −0.5) | −1.39 | 0.0146 | 0.0514 | 0.0648 | 0.8694 | 0.0066 | 0.0329 | 0.0363 | 0.9198 | 0.0025 | 0.0200 | 0.0208 | 0.9394 | |
| 0.0060 | 0.0275 | 0.0340 | 0.8786 | 0.0029 | 0.0177 | 0.0191 | 0.9246 | 0.0010 | 0.0106 | 0.0110 | 0.9398 | |||||
| −0.0011 | 0.0078 | 0.0099 | 0.8676 | −0.0003 | 0.0058 | 0.0066 | 0.9086 | −0.0002 | 0.0038 | 0.0040 | 0.9410 | |||||
| −0.0544 | 0.1004 | 0.1184 | 0.9006 | −0.0257 | 0.0699 | 0.0750 | 0.9314 | −0.0085 | 0.0438 | 0.0449 | 0.9424 | |||||
| 0.0194 | 0.0505 | 0.0598 | 0.9040 | 0.0083 | 0.0357 | 0.0385 | 0.9314 | 0.0033 | 0.0224 | 0.0236 | 0.9390 | |||||
| −0.0089 | 0.0457 | 0.0503 | 0.9222 | −0.0051 | 0.0332 | 0.0359 | 0.9268 | −0.0009 | 0.0206 | 0.0210 | 0.9462 | |||||
| −0.0983 | 0.3151 | 0.3520 | 0.9332 | −0.0498 | 0.2163 | 0.2273 | 0.9460 | −0.0162 | 0.1340 | 0.1360 | 0.9498 | |||||
| 1.61 | 0.0003 | 0.0449 | 0.0536 | 0.8920 | −0.0002 | 0.0287 | 0.0310 | 0.9310 | 0.0002 | 0.0175 | 0.0179 | 0.9428 | ||||
| 0.0003 | 0.0255 | 0.0311 | 0.8866 | −0.0001 | 0.0164 | 0.0181 | 0.9222 | 0.0001 | 0.0099 | 0.0102 | 0.9410 | |||||
| −0.0001 | 0.0069 | 0.0098 | 0.8308 | 0.0001 | 0.0053 | 0.0064 | 0.8950 | 0.0000 | 0.0036 | 0.0039 | 0.9334 | |||||
| −0.0627 | 0.1116 | 0.1311 | 0.8904 | −0.0315 | 0.0791 | 0.0863 | 0.9162 | −0.0113 | 0.0501 | 0.0514 | 0.9398 | |||||
| 0.0113 | 0.0460 | 0.0504 | 0.9264 | 0.0047 | 0.0337 | 0.0352 | 0.9386 | 0.0014 | 0.0215 | 0.0216 | 0.9472 | |||||
| −0.0058 | 0.0467 | 0.0503 | 0.9280 | −0.0023 | 0.0336 | 0.0350 | 0.9388 | −0.0012 | 0.0211 | 0.0215 | 0.9464 | |||||
| −0.1492 | 0.3012 | 0.3461 | 0.9266 | −0.0777 | 0.2073 | 0.2277 | 0.9312 | −0.0274 | 0.1288 | 0.1333 | 0.9446 | |||||
| 0.50 | (−0.5, −2, 1) | (−1, 1.6, −0.5) | −1.39 | 0.0126 | 0.0512 | 0.0636 | 0.8744 | 0.0073 | 0.0331 | 0.0368 | 0.9182 | 0.0023 | 0.0200 | 0.0204 | 0.9398 | |
| 0.0050 | 0.0275 | 0.0338 | 0.8812 | 0.0032 | 0.0178 | 0.0195 | 0.9230 | 0.0009 | 0.0106 | 0.0108 | 0.9422 | |||||
| −0.0011 | 0.0077 | 0.0098 | 0.8628 | −0.0005 | 0.0058 | 0.0067 | 0.9118 | −0.0003 | 0.0039 | 0.0041 | 0.9350 | |||||
| −0.0576 | 0.1001 | 0.1197 | 0.8924 | −0.0239 | 0.0700 | 0.0748 | 0.9288 | −0.0087 | 0.0438 | 0.0446 | 0.9446 | |||||
| 0.0186 | 0.0499 | 0.0586 | 0.9074 | 0.0080 | 0.0358 | 0.0381 | 0.9342 | 0.0036 | 0.0224 | 0.0236 | 0.9376 | |||||
| −0.0084 | 0.0455 | 0.0489 | 0.9274 | −0.0044 | 0.0333 | 0.0352 | 0.9346 | −0.0011 | 0.0207 | 0.0207 | 0.9492 | |||||
| −0.1142 | 0.3149 | 0.3507 | 0.9382 | −0.0432 | 0.2162 | 0.2242 | 0.9448 | −0.0154 | 0.1340 | 0.1391 | 0.9422 | |||||
| (1, 0.7, −0.3) | −1.39 | −0.0068 | 0.0529 | 0.0668 | 0.8808 | −0.0020 | 0.0346 | 0.0384 | 0.9198 | −0.0008 | 0.0210 | 0.0220 | 0.9428 | |||
| −0.0025 | 0.0297 | 0.0382 | 0.8720 | −0.0005 | 0.0196 | 0.0223 | 0.9164 | −0.0003 | 0.0118 | 0.0124 | 0.9428 | |||||
| 0.0002 | 0.0082 | 0.0121 | 0.8276 | 0.0001 | 0.0063 | 0.0079 | 0.8782 | 0.0000 | 0.0043 | 0.0047 | 0.9342 | |||||
| −0.0667 | 0.1081 | 0.1297 | 0.8862 | −0.0275 | 0.0765 | 0.0813 | 0.9306 | −0.0102 | 0.0482 | 0.0497 | 0.9402 | |||||
| 0.0154 | 0.0511 | 0.0574 | 0.9182 | 0.0048 | 0.0375 | 0.0393 | 0.9352 | 0.0023 | 0.0238 | 0.0245 | 0.9430 | |||||
| −0.0070 | 0.0512 | 0.0560 | 0.9208 | −0.0047 | 0.0371 | 0.0386 | 0.9426 | −0.0010 | 0.0232 | 0.0234 | 0.9510 | |||||
| −0.1579 | 0.3161 | 0.3625 | 0.9240 | −0.0694 | 0.2164 | 0.2292 | 0.9398 | −0.0227 | 0.1342 | 0.1375 | 0.9440 | |||||
| 0.75 | (1, −1, 0.5) | (1, 0.7, −0.3) | 1.61 | −0.0061 | 0.5813 | 0.6422 | 0.9130 | −0.0004 | 0.4112 | 0.4314 | 0.9352 | −0.0020 | 0.2552 | 0.2604 | 0.9442 | |
| 0.0032 | 0.3709 | 0.4267 | 0.9004 | 0.0015 | 0.2696 | 0.2875 | 0.9296 | 0.0000 | 0.1632 | 0.1668 | 0.9426 | |||||
| 0.0059 | 0.2288 | 0.2930 | 0.8690 | 0.0061 | 0.1650 | 0.1863 | 0.9152 | −0.0001 | 0.1074 | 0.1115 | 0.9414 | |||||
| −0.0629 | 0.1118 | 0.1297 | 0.8912 | −0.0292 | 0.0792 | 0.0866 | 0.9180 | −0.0127 | 0.0500 | 0.0510 | 0.9422 | |||||
| 0.0079 | 0.0458 | 0.0497 | 0.9274 | 0.0033 | 0.0338 | 0.0351 | 0.9400 | 0.0019 | 0.0214 | 0.0219 | 0.9490 | |||||
| −0.0037 | 0.0467 | 0.0503 | 0.9292 | −0.0012 | 0.0337 | 0.0354 | 0.9382 | −0.0007 | 0.0211 | 0.0213 | 0.9462 | |||||
| −0.1481 | 0.3016 | 0.3408 | 0.9286 | −0.0727 | 0.2073 | 0.2253 | 0.9362 | −0.0313 | 0.1289 | 0.1305 | 0.9508 | |||||
| (−0.5, −2, 1) | (1, 0.7, −0.3) | 1.61 | −0.0043 | 0.1204 | 0.1302 | 0.9226 | −0.0013 | 0.0834 | 0.0886 | 0.9322 | −0.0007 | 0.0516 | 0.0519 | 0.9474 | ||
| 0.0026 | 0.0772 | 0.0880 | 0.9102 | 0.0009 | 0.0547 | 0.0594 | 0.9242 | 0.0005 | 0.0330 | 0.0333 | 0.9462 | |||||
| −0.0013 | 0.0494 | 0.0627 | 0.8624 | −0.0006 | 0.0338 | 0.0366 | 0.9208 | 0.0001 | 0.0217 | 0.0224 | 0.9370 | |||||
| −0.0547 | 0.2266 | 0.2380 | 0.9246 | −0.0307 | 0.1588 | 0.1659 | 0.9306 | −0.0099 | 0.1000 | 0.0997 | 0.9500 | |||||
| 0.0264 | 0.1123 | 0.1207 | 0.9282 | 0.0098 | 0.0805 | 0.0833 | 0.9414 | 0.0045 | 0.0504 | 0.0517 | 0.9428 | |||||
| −0.0142 | 0.1147 | 0.1221 | 0.9342 | −0.0080 | 0.0803 | 0.0843 | 0.9382 | −0.0014 | 0.0496 | 0.0508 | 0.9470 | |||||
| −0.0064 | 0.2947 | 0.2999 | 0.9452 | −0.0107 | 0.2041 | 0.2084 | 0.9432 | −0.0028 | 0.1282 | 0.1277 | 0.9488 | |||||
| AIC | log-Likelihood | LRT | KS | |||||
|---|---|---|---|---|---|---|---|---|
| GSC | GSC | SKL | GSC | GSC | Statistical | p-Value | p-Value | |
| 0.10 | 1168.54 | 1154.64 | 1194.28 | −574.27 | −563.32 | 21.90 | <0.0001 | 0.988 |
| 0.25 | 1172.72 | 1164.55 | 1172.70 | −576.36 | −568.27 | 16.17 | 0.0003 | 0.646 |
| 0.50 | 1174.74 | 1171.83 | 1182.66 | −577.37 | −571.91 | 10.91 | 0.0043 | 0.180 |
| 0.75 | 1171.50 | 1176.51 | 1221.65 | −575.75 | −574.25 | 3.00 | 0.2235 | 0.839 |
| 0.90 | 1223.71 | 1229.75 | 1280.45 | −601.85 | −600.88 | 1.95 | 0.3768 | 0.004 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gómez, Y.M.; Gallardo, D.I.; Venegas, O.; Magalhães, T.M. An Asymmetric Bimodal Double Regression Model. Symmetry 2021, 13, 2279. https://doi.org/10.3390/sym13122279
Gómez YM, Gallardo DI, Venegas O, Magalhães TM. An Asymmetric Bimodal Double Regression Model. Symmetry. 2021; 13(12):2279. https://doi.org/10.3390/sym13122279
Chicago/Turabian StyleGómez, Yolanda M., Diego I. Gallardo, Osvaldo Venegas, and Tiago M. Magalhães. 2021. "An Asymmetric Bimodal Double Regression Model" Symmetry 13, no. 12: 2279. https://doi.org/10.3390/sym13122279
APA StyleGómez, Y. M., Gallardo, D. I., Venegas, O., & Magalhães, T. M. (2021). An Asymmetric Bimodal Double Regression Model. Symmetry, 13(12), 2279. https://doi.org/10.3390/sym13122279









