1. Introduction
Lung cancer is one of the most common cancers worldwide and the main cause of death for cancer patients. According to “Global Cancer Statistics” [
1], in 2018, there were approximately 2.1 million new cases of lung cancer worldwide and 1.77 million lung cancer-related deaths. Since lung cancer has no obvious symptoms in the early stages and is difficult to detect, it is often discovered in the middle and late stages of cancer, and the best treatment time is missed. On the one hand, studies have found that most of the lung cancer is in the form of lung nodules. The lung nodules are divided into benign and malignant. The probability of malignant lung nodules becoming cancerous are greatly increased. An accurate early identification of benign and malignant lung nodules is essential for the prevention of lung cancer. On the other hand, lung nodules are present as three-dimensional spherical shapes in images and are affected by factors such as variable shapes, different sizes, and complex surrounding tissues. The shape of pulmonary nodules is especially critical for doctors to screen. The process of using CT images to detect lung cancer is actually a process of identifying lung nodules.
The current research on lung nodules mainly focuses on two kinds of methods.
The first method is based on traditional image processing segmentation methods. Carvalh et al. [
2] combined traditional image processing methods with machine learning methods. First, the lung parenchyma was segmented from the CT image as the region of interest. Then, the candidate nodules are detected in terms of shape, texture, and extraction of expression features (such as size) through segmentation methods. Messay et al. [
3] designed an algorithm for the segmentation-based detection of nodules using candidate nodule shape, location, brightness, and gradient characteristics. Although the methods in [
2,
3] have achieved significant results, the edges of the lung nodules in the CT image are blurred due to the small size of the lung nodules (usually between 3 and 30 mm) and low contrast; additionally, the image gray scale is uneven due to adhesions. The influence of noise and artifacts also limits the accuracy of such methods in detecting and segmenting lung nodules.
The second segmentation method used to detect lung nodules is based on machine learning or deep learning. With the continuous development of deep learning, more and more deep learning-based convolutional neural networks have played a vital role in lung nodule research, mainly focusing on methods based on 2D convolutional neural networks and 3D-based methods.
- (1)
Methods based on the 2D convolutional neural network. Ding et al. [
4] borrowed from the successful application of deep convolutional neural networks (DCNNs) in natural image recognition and proposed a lung nodule detection method based on DCNNs. In the faster R-CNN neural network, a deconvolution structure is introduced for candidate detection of axis slices. Aiming at the problem that the size of lung nodules is too small, and it is easy to lose features, a deconvolution is added after the VGG16 network to restore the size of the feature map so that the network can capture features more accurately. Deng Zhonghao et al. [
5] aimed at solving the problem of low detection sensitivity of traditional algorithms and reducing the large number of false positives. The UNet network was improved to reduce the complexity of a deep neural network while maintaining its sensitivity. Although the two-dimensional detection and segmentation methods have made great progress compared with the traditional methods, a CT image is a three-dimensional image sequence, and the lung organs of the human body are not based on a two-dimensional plane, so making inferences about a three-dimensional object from single plane cuts is often not objective or specific enough. The artificial reduction of one dimension of information often results in low recall rates and high false positives. This undoubtedly brings a lot of unnecessary work to physicians. Therefore, because lung organs and lung nodules are three-dimensional objects, a three-dimensional convolutional neural network is required to further improve the detection and classification accuracy.
- (2)
Methods based on the 3D convolutional neural network. Aiming at the characteristics of lung nodules in a three-dimensional space and its variability in shape, Zhu et al. [
6] considered the three-dimensionality of lung CT data and the compactness of the dual-path network, and designed two deep three-dimensional DPNs for the nodules: Detection and classification. Specifically, a three-dimensional fast convolutional neural network area (R-CNN) is designed for nodule detection. This modified R-CNN uses a three-dimensional dual-path block and UNet encoding and decoding structure to effectively learn nodule characteristics. This method makes full use of a lung nodule’s spatial information and integrates feature maps with different abstract levels to repair the lost features, making the detection accuracy higher. Gong et al. [
7] proposed an automatic computer-aided detection scheme for lung nodules based on deep convolutional neural networks (DCNNs). A three-dimensional dynamic neural network (SE-ResNet) based on a compressed excitation network and residual network was used to detect lung nodules and reduce false positives. Specifically, by fusing the 3D-SE-ResNet module to design a three-dimensional area suggestion network, with a UNet network structure to detect candidate lung nodules, the 3D-SE-ResNet module recalibrates the residual characteristic response of the channel to enhance the network. The model uses the 3D-SE-ResNet module to effectively learn the characteristics of nodules and improve nodule detection performance. Although this method detects lung nodules in three dimensions and can make full use of the spatiality and completeness of CT image sequences, it does not fully consider the adhesion of lung nodules and surrounding tissues, resulting in inaccurate segmentation.
Considering the influence of the surrounding tissues of lung nodules on the segmentation of lung nodules, as well as the diversification of lung nodules, this paper proposes a CT image lung nodule segmentation method based on the 3D convolutional neural network. Compared with other methods, the main difference is the 3D feature extraction method. By making full use of three-dimensional spatial information, our network can learn more feature information than two-dimensional networks. We also transform the Res2Net module into a 3D-Res2Net module and integrate it into the 3D-UNet network and design a 3D-Res2UNet network for candidate nodule detection and segmentation. Therefore, the network takes CT image sequences as input, can make full use of the spatial information of lung nodules, and minimizes artificial loss of dimensionality. First of all, in the designed network, the improved Res2Net network module can make the overall network deeper, effectively solving the problems of gradient explosion and gradient disappearance that are prone to deep networks. Secondly, the 3D-UNet network acts as the basic network, which can ensure its effectiveness while restricting the size of the feature map and repairing the lost features. Experiments have shown that, compared with the existing methods, the method in this paper can detect lung nodules more accurately and segment them effectively.
This article combines the Res2Net module with 3D-UNet and has achieved good results on small targets. This structure is not only suitable for the detection of lung nodules, but also for the detection of small particles and irregular objects in other fields. Thus, this research has laid the foundation for future applied work.
2. Related Work
2.1. UNet Segmentation Network
UNet [
8] is a semantic segmentation network developed based on a fully convolutional neural network. The network has a total of 23 layers, and the number of layers is far less than the other networks while ensuring accuracy. The UNet network mainly includes two parts, down-sampling and up-sampling. Down-sampling is also called the feature extraction part, which mainly uses the convolutional and pooling layer to extract features of the input image. Up-sampling uses a deconvolution operation to up-sample the feature map. This structure of down-sampling and up-sampling is also called a decoder-encoder structure. In the down-sampling part, the input image passes through the convolutional and pooling layer to obtain feature maps of different levels. These feature maps contain image features with different levels of abstraction. In the up-sampling part, the deconvolution layer is used to gradually restore the size of the feature map, and the down-sampled feature map is merged to repair the less abstract detail information lost in the training process and improve the segmentation accuracy of the network.
However, because the lung is a three-dimensional structure, the UNet network uses two-dimensional convolution and pooling operations to extract the features of lung nodules in lung CT images, which will cause a lot of spatial information to be lost. Thus, a lot of contextual information is lost in the down-sampling process. It cannot be fully restored during up-sampling, which leads to fuzzy up-sampling results and insensitiveness to the details of the image. Combined with the above problems, it requires a three-dimensional network for further optimization.
2.2. 3D-UNet Segmentation Network
3D-UNet [
9] is an improved semantic segmentation network based on UNet, since a lot of data in the field of medical imaging are three-dimensional data. Therefore, if you directly use the UNet network to process images, you need to perform slice preprocessing on the 3D data first. That is to divide the three-dimensional image into multiple layers of two-dimensional data, and then divide the two-dimensional data. This method is not only cumbersome and inefficient, but also loses the dimensional information of the three-dimensional data. This leads to the loss of correlation between adjacent slices, making the network accuracy low. The 3D-UNet network mainly includes an encoder part and a decoder part. Additionally, in the encoder part, a 3D convolution layer and a 3D pooling layer are used to extract the expressive features of the input image, and in the decoder part, 3D deconvolution is used to restore the feature map size. Moreover, the feature maps of the decoder and the encoder are merged through the cascade operation, which brings richer semantic information to the segmentation. This makes the segmentation accuracy higher.
3. Method
Since the lung CT image is a three-dimensional tomogram, and lung nodules are small in size, have a variable morphology, and have rich semantic information around the lung nodules (such as blood vessels and bronchus), the 3D segmentation of lung nodules is an extremely difficult research problem. However, these disadvantages can be mitigated by the Res2Net module because of its ability to capture the characteristics of tiny particles. In this work, the module is converted from 2D to 3D, which optimizes the 3D-UNet network to establish a 3D-Res2UNet network. This ensures that the entire network can be more accurate to improve image segmentation of lung nodules.
3.1. 3D-Res2Net
On the one hand, with the development of deep learning, deep neural networks have made a series of achievements in image classification tasks. This relies on the ability of deep neural networks to capture features more effectively. The number of these features can be increased by stacking more layers, which shows that this method is effective to establish connections within the network through a multi-layer end-to-end approach. On the other hand, in order to make the neural network achieve better learning results, the increasing number of network layers has brought some problems to the training of the network, of which gradient disappearance and gradient explosion are most important.
A residual network (ResNet) [
10] effectively solves this problem. The residual learning unit introduces identity mapping to establish a direct correlation channel between the output and input; therefore, the parameterized layer can learn the input and output, and the residual difference between them achieves the purpose of protecting the integrity of the information and simplifies the goal and speed of learning.
For visual tasks, different scale information contains different features, so it is very important to be able to express features on multiple scales. Considering the general applicability of the 3D network in the future, this paper proposes to add a channel to the original Res2Net [
11] module to upgrade it to the 3D level. The 3D-Res2Net module is shown in
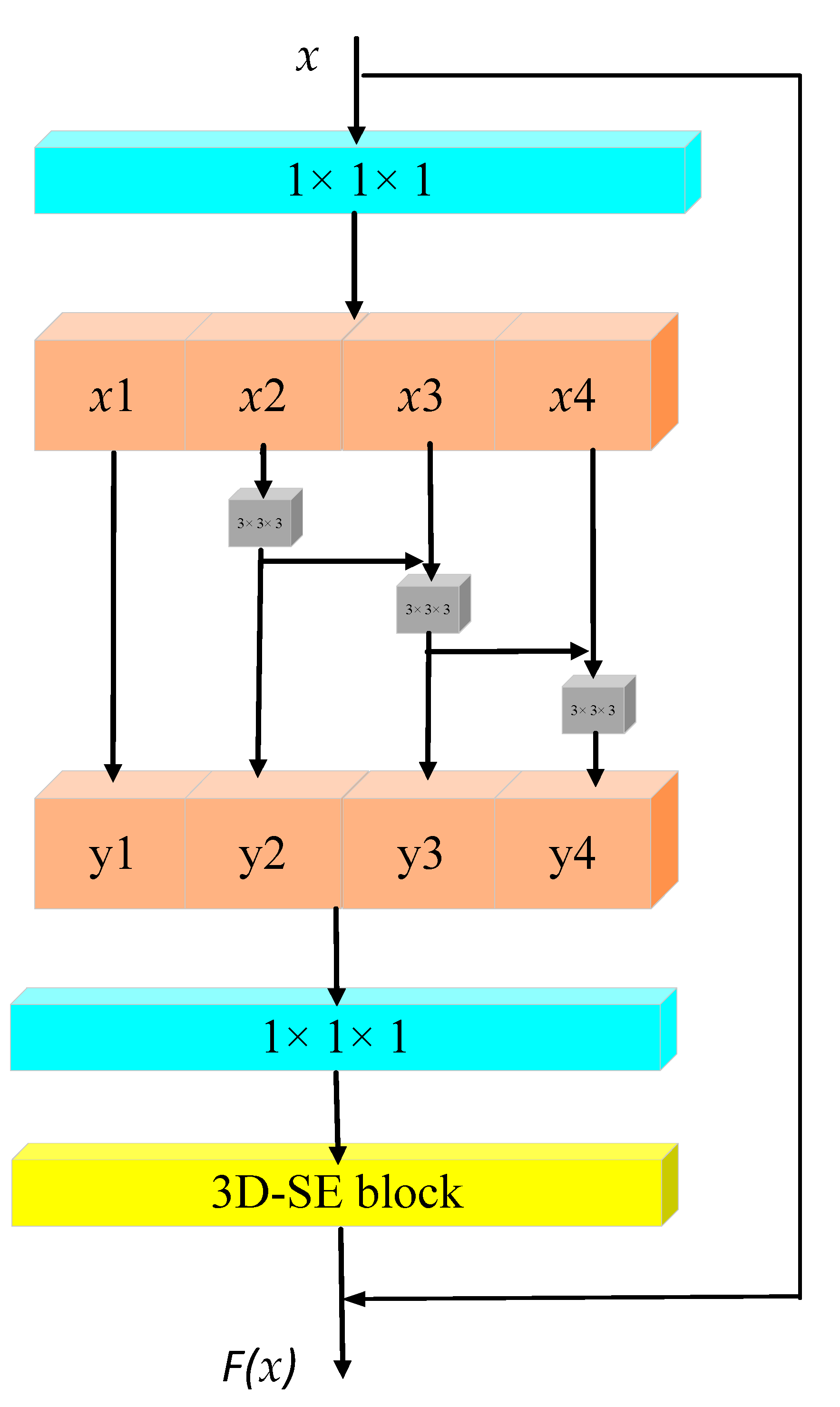
Figure 1.
3D-Res2Net changes the internal structure of the basic residual module. Compared with the basic residual unit, the new module replaces the original filter with multiple sets of 3 × 3 × 3 filters and combines different filter groups. The residual cascade is connected to construct a new layered residual connection in a single residual block. A 3D-SE block is added, after the last 1 × 1 convolution, to each channel to reallocate weight. As shown in
Figure 1, the 3D-Res2Net module uses a new size. It is the number of feature groups with scale = 4 so that the feature map sent to this structure is converted to eight channels after the 1 × 1 × 1 convolution. In addition, x_1 represents the feature map with channel numbers 1, 2, and x_2 represents the feature map with channel numbers 3 and 4; in this way, if the channels are grouped and then trained, the weight of the convolution kernel trained by each grouped channel is also different. Compared with the weight of each channel convolution kernel caused by non-group training, this technique shows a greater advantage. Although the computational load of the Res2Net module is similar to the residual network architecture, the hierarchically connected network has strong multi-scale feature extraction capabilities, which enables the network to express multi-scale features at a finer granularity, and increases the network capacity of each layer. The output formula is as follows:
Here, s is the number of groups that Res2Net integrates the 3 × 3 volumes of the original residual module into groups. Although a similar external jump connection is performed inside a bottleneck, the concat operation is performed on y_1, y_2, y_3, and y_4, so that the channel remains unchanged. The direct mapping of the first x_1 to y_1 is based on two considerations. The first is to reduce network parameters, and the second is to reuse features.
3.2. Network Design
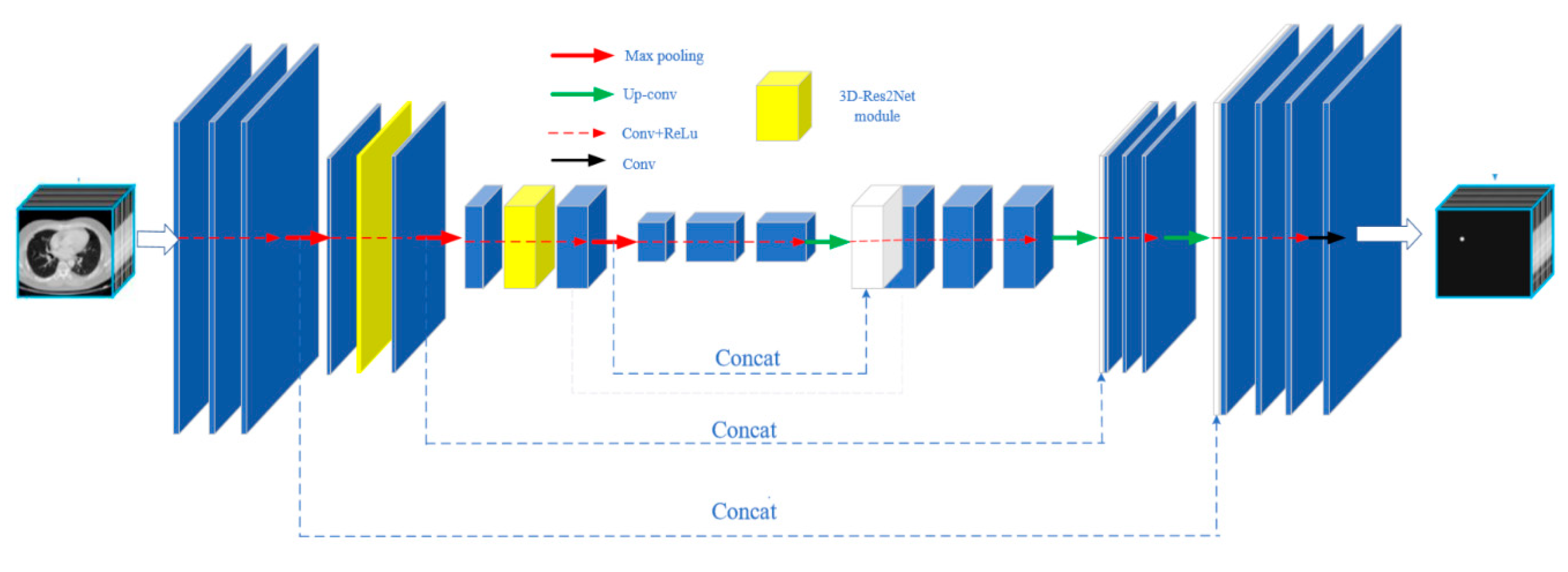
This article combines the 3D-Res2Net module with the 3D-UNet network to form a new network structure, namely 3D-Res2UNet. The network structure diagram is shown in
Figure 2.
Down-sampling is the most important process of extracting features in a network, and it can extract feature details well, which has an important significance for subsequent segmentation operations. Compared with the 3D-UNet convolutional neural network, the 3D-Res2UNet convolutional neural network changes the original CBR module in the two down-samplings, namely Conv + BN + ReLU, and replaces the 3D basic convolution module with the 3D-Res2Net module as a whole. Therefore, the method of integrating the residual network into the entire network can make the network deeper, and because of the strong feature extraction ability of the 3D-Res2Net module and the phased update of the convolution kernel weights of each channel, the overall network is able to capture very small features. With an improved performance, the connection method where x_1 is directly mapped to y_1 in the module effectively reduces network parameters and the resource consumption. Therefore, the network will not affect the network speed due to excessive redundancy.
4. Experience and Results
4.1. Dataset
This method is validated on the public dataset of LUNA16 [
12]. The dataset contains a total of 888 sets of CT images with a total of 1186 lung nodules. During the experiment, the data was randomly divided into three parts, 70% of the data was used for training, 20% of the data was used for testing, and 10% of the data was used for verification. The criterion for judging a nodule in the LUNA16 dataset is that at least three of the four radiologists determine that the radius of the nodule is greater than 3 mm. Therefore, in the annotations of the dataset, non-nodules, nodules with a radius less than 3 mm, and nodules with a radius greater than 3 mm are considered by one or two radiologists as unrelated findings.
4.2. Data Preprocessing
4.2.1. Data Format Conversion
In the dataset, each chest CT sample mainly contains two parts: (1) The raw data of chest CT, including two files with the suffix zraw and mhd. The zraw file saves the original CT data and the mhd file saves the header file information of the CT data. The most important information is establishing an axis origin and pixel spacing. The origin represents the coordinates of the origin of the CT data in the world coordinate system, while the spacing subdivides the CT data to a length of a pixel in the world coordinates. (2) A CSV (Comma Separated Values) file with nodule annotation information. The content of the file is shown in
Table 1.
Since the CT data are expressed in a natural coordinate system, the world coordinates of the lung nodules need to be converted to the natural coordinates of the CT voxels to correspond to the CT data before the actual operation. The conversion formulas are as follows:
The terms voxelcoord and voxeldiam, respectively represent the coordinates and diameter of the lung nodule in the CT pixel coordinate system, the terms coord and diameter_mm, respectively represent the coordinates and diameter of the nodule in the world coordinate system.
4.2.2. Lung Parenchymal Segmentation
Since the lung tissue is relatively complex, there are a large number of lung trachea, pulmonary blood vessels, tissue mucosa, and other structures around the lung nodules. As a result, there are some inevitable errors in the direct use of the original slice image to segment the lung nodules. If the data are preprocessed first, the lung parenchyma are segmented before the detection step. The lung nodules are detected in the lung parenchyma, which can avoid the interference of the external tissues and organs of the lung parenchyma on the detection task, thereby improving the detection accuracy. The lung CT image is shown in
Figure 3.
In this paper, the method proposed by Mansoor et al. [
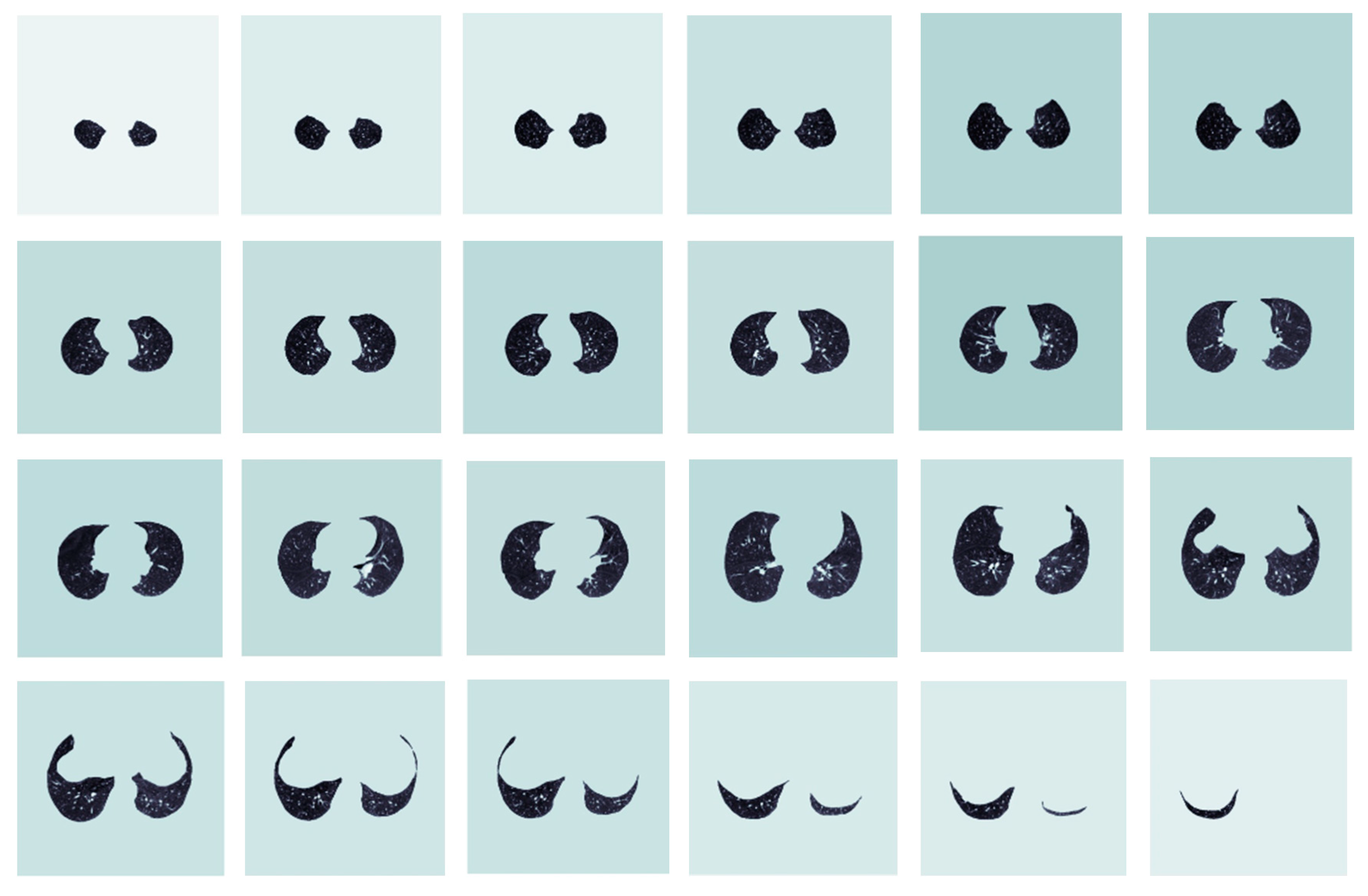
13] is used to segment the lung parenchyma. After the lung is segmented, the lung parenchyma is accurately segmented to remove the influence of surrounding tissues. This has a positive effect on lung nodule segmentation. Since the lung is a 3D model, the segmentation is divided into multiple horizontal layers, and the overall segmentation result is shown in
Figure 4.
4.2.3. Voxel Value Normalization
X-rays are used to scan the lungs when collecting data in CT. Since different tissues of the human body have different X-ray absorption characteristics, different ray attenuations will occur when X-rays pass by, and the remaining X-rays after attenuation will be converted into a digital signal to acquire the desired CT value. The CT values of different human tissues are shown in
Table 2.
Table 2 shows that the CT value of the lung is at −500, so the HU value of the CT sample is intercepted as [−1000, +400]. The [−1000, +400] is normalized to a [0, 1] range. This helps reduce the influence of other scalings and enhances the ability of neural networks to capture features.
4.2.4. Data Enhancement
Medical image data are essential for training the model. The severe imbalance of positive and negative samples in the dataset will affect the network performance. As a result, the network weight cannot reach the optimal value. Therefore, the AugGAN network [
14] proposed by Huang SW et al. is used for data enhancement. It can solve the problem of insufficient positive samples to a certain extent. This also helps slow down the occurrence of model overfitting. In addition, the detection accuracy of lung nodules can also be improved. The resulting lung nodules are shown in
Figure 5.
4.3. Evaluation Standard
This article evaluates the article method from two perspectives, namely the dice index recall rate (Recall) and the average number of false positives per sample (FP/scan).
The dice index [
15] refers to the degree of fit between the original target and the segmented target. The more the two objects fit, the higher the dice value and the lower the loss function value, indicating that the network segmentation of the lung nodules is more complete. The dice coefficient formula is as follows:
The relationship between the dice coefficient and loss coefficient is as follows:
Here, Ntruepositive represents the area where the lung nodule exists and is correctly segmented, Nfalsepositive represents the area where the lung nodule exists but is not correctly segmented, and Nfalsenegative represents the area where the lung nodule does not exist and is not segmented. When the invalid dice coefficient is close to 1, the loss function loss is infinitely close to 0. At this time, the model segmentation result matches the real result.
The Recall indicator refers to the sensitivity of the network to lung nodules, describing the ability to segment lung nodules. The larger the value, the more the lung nodules found through the network are complete. Among them,
N’real is the number of real nodules detected by the network, and
Nnodule is the number of real nodules in the sample. The formula is as follows:
The average number of false positives (FP/scan) describes the network’s ability to judge lung nodules, whether it can effectively avoid vascular tomography or lung tissue, and accurately distinguish lung nodules, where
N’no is the network detected. The number of non-nodules,
Nsample is the total number of training samples, and the formula is as follows:
4.4. Experimental Results
The method in this paper is tested on the LUNA16 public dataset. There are many types of lung nodules in the sample, which are divided into three categories according to the nodule’s density, e.g., solid nodules, mixed nodules, and ground glass nodules. The variable density structure can fully detect the ability of network segmentation and avoid the occurrence of contingency.
Figure 6 shows the results of the detection of lung nodules by the 3D-Res2UNet neural network and the overall effect of segmentation, and
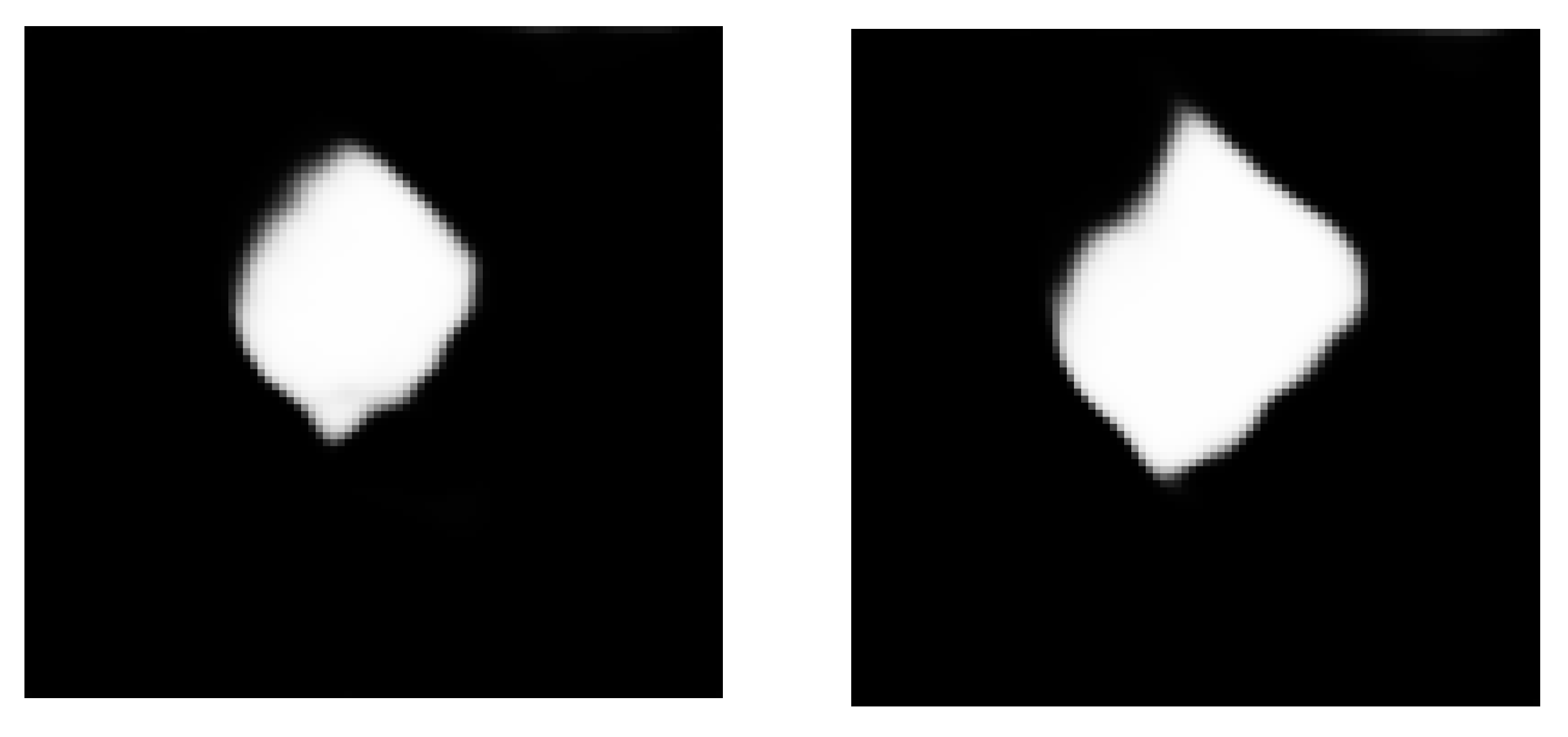
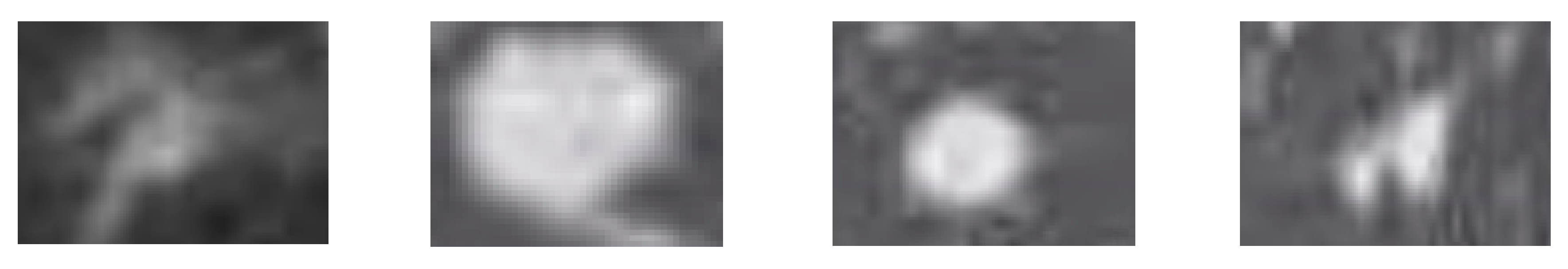
Figure 7 shows the local effect of lung nodule segmentation. In the local renderings, it can be found that the lung nodules are completely segmented. Either ellipse-like smooth edges or prominent feature edges can be accurately segmented by the 3D-Res2UNet neural network. This is not only helpful to assist doctors in the diagnosis and treatment, but also lays the foundation for the follow-up study of false positive detection of lung nodules.
In addition,
Figure 8 lists some images whose segmentation is not very accurate. This is because the shape of such lung nodules is different from most lung nodules. Its edges are rough and jagged. Such lung nodules require a deeper network structure when segmenting the edges. However, a deeper network will inevitably lead to an increase in neural network training time and waste of resources. In fact, the ultimate purpose of segmentation of lung nodules is to assist doctors in the diagnosis, but the false positive probability of such nodules is relatively high. Therefore, the practical value of deepening the network for a more precise segmentation of such lung nodules is very small.
Model Comparison
The experiment first compared the ability of 3D-Res2UNet and the original network to segment and fit lung nodules. The comparison parameters are shown in
Table 3.
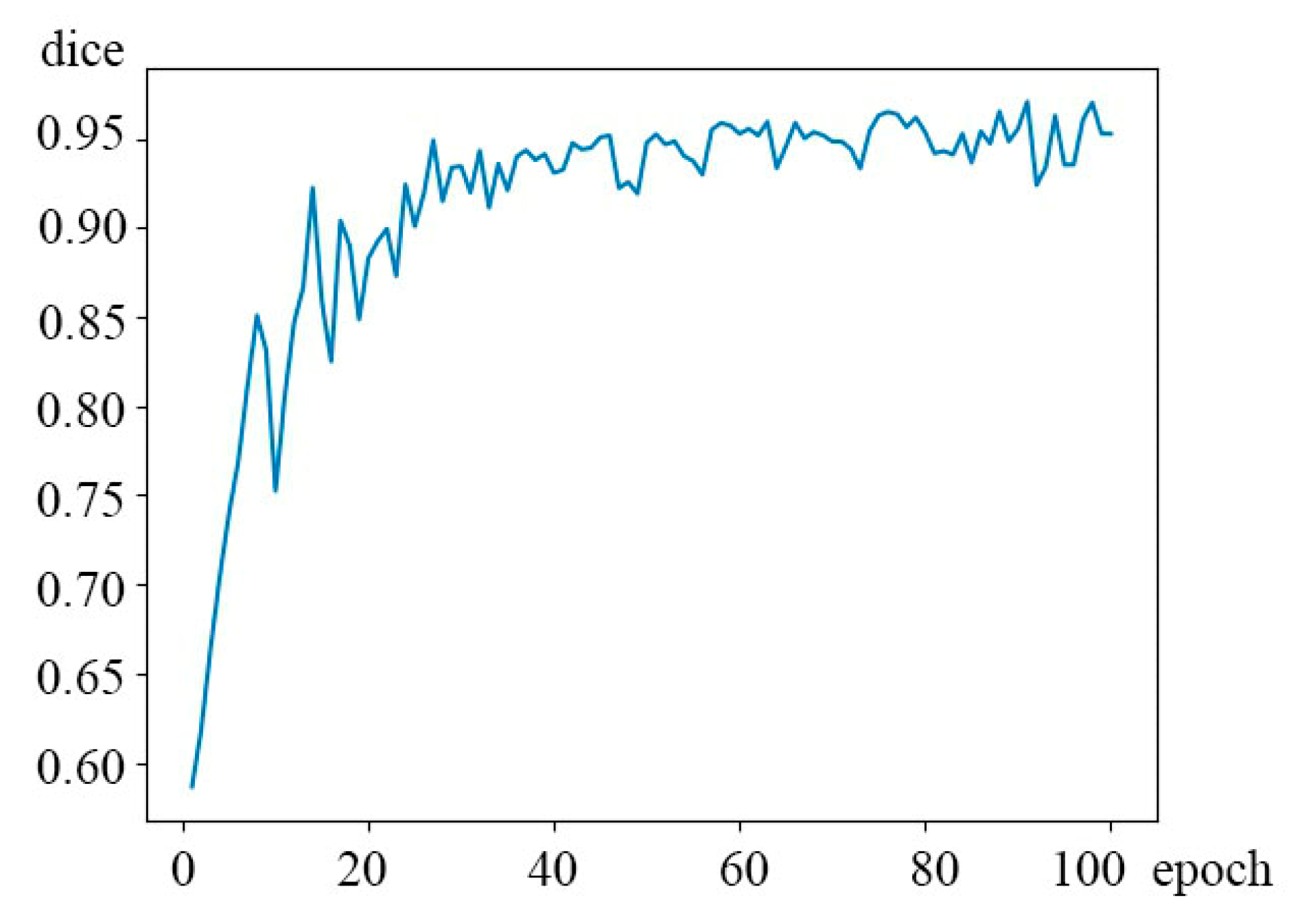
Due to the three-dimensional CT tomogram of the lungs, as shown in the table, the 3D network has obvious advantages in capturing lung nodules compared with a 2D network. Since the 3D-Res2net module fused in this paper uses multiple sets of 3 × 3 filters, and different filter sets are connected by the residual cascade, a new structure is constructed in a single residual block. The hierarchical residual class is connected and finally a 3D-SE block is added to re-assign a weight to each channel. Such hierarchical filtering and gradual fusion of details make the 3D-Res2Unet network more sensitive to the small edges often found in lung nodules; thus, this method can more accurately restore the original shape of lung nodules during segmentation. The final dice coefficient is significantly better than the other networks by 95.30%, as shown in
Figure 9, where the abscissa is the number of epochs and the ordinate is the dice coefficient.
Secondly, in terms of recall rate and the average number of false positive lung nodules, this article compares traditional and existing methods, as shown in
Table 4.
In
Table 4, ISICAD [
17] is a traditional image processing method that uses artificially designed features to detect lung nodules and processes them according to the edge shape of lung nodules. This method mainly focuses on the main characteristics of large nodules. The eature extraction of mixed nodules and ground glass nodules is insufficient, so the overall recall rate is relatively low. LUNA16_V1 [
20], as the official method, is representative of traditional image processing methods. It integrates the advantages of all previous traditional methods. This method has a high sensitivity to high-density nodules. The method works best on solid nodules and other large nodules. LUNA16_V2 [
20] is an upgraded version based on LUNA16_V1. In addition to focusing on high-density large nodules, the network also has targeted designs for low-density mixed nodules and ground glass nodules. However, the impact on other lung tissues and the indiscriminate detection of vascular cross-sections has led to a sharp increase in the number of false positive lung nodules, which has caused great trouble for doctors when reading the film. A three-dimensional fully convolutional neural network designed by Dou et al. [
21] can effectively detect the lung nodule memory. However, because the network does not have an up-sampling process, it cannot effectively repair the degree of abstraction lost in the training process due to the low detail information. The method in this paper can effectively restore the lost information. At the same time, this method can also extract detailed information many times to significantly improve the recall rate.
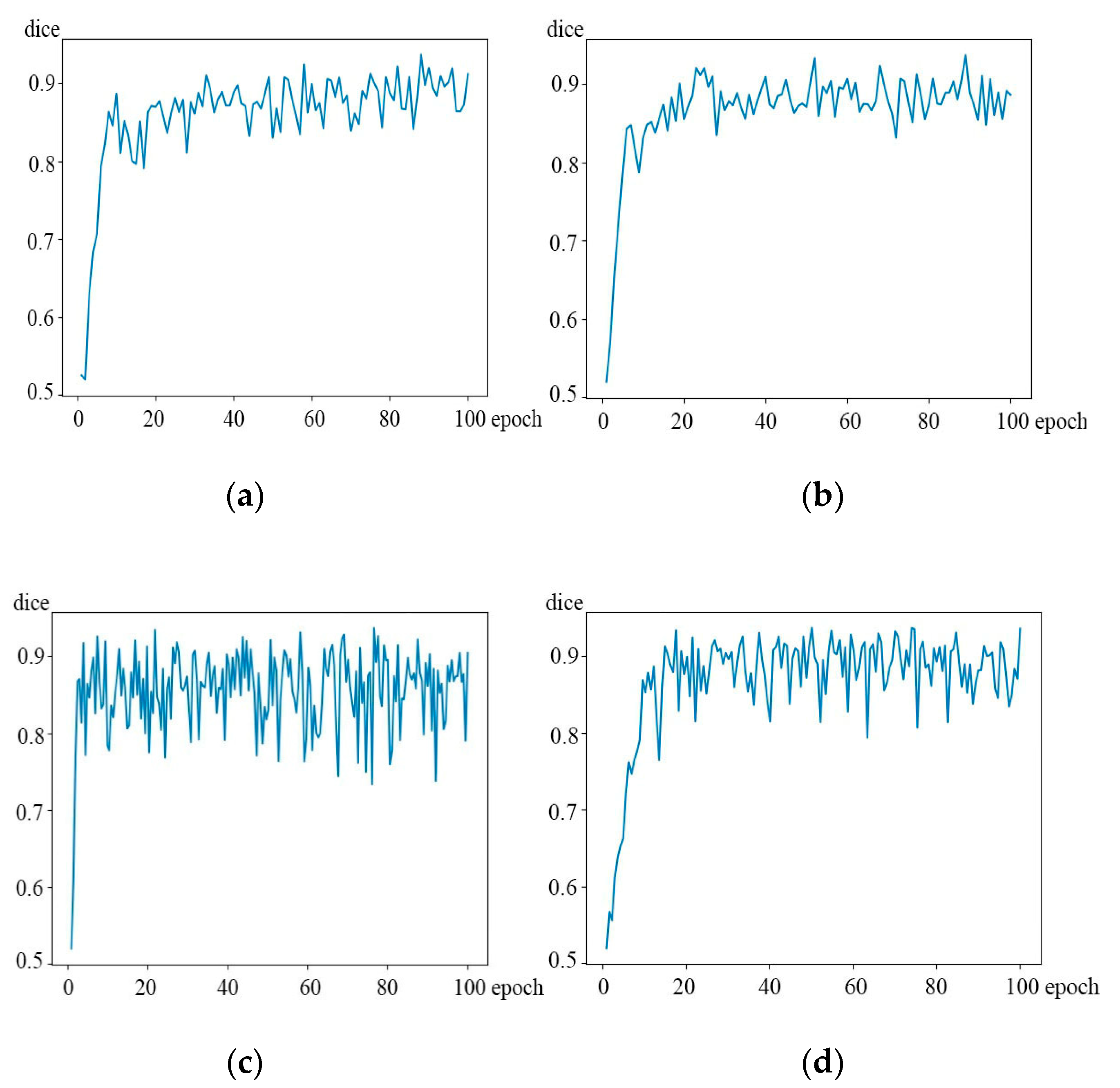
Finally, since the down-sampling process is a gradual amplification process, this article adds the 3D-Res2Net module to the second and third down-sampling of the 3D-UNet network to form symmetry in order to capture more target information. This effectively prevents the network from not capturing enough detail because the target is too large in the first down-sampling. What is more, it also avoids the target area of being too limited in the last down-sampling. In order to prove its rationality, the following comparative experiments were done. The comparison of experimental correlation coefficients is shown in
Table 5. What is more, the dice coefficient variation curve is shown in
Figure 10. Among them, the upper corner is marked as the position where the 3D-Res2Net module was added to the down-sampling.
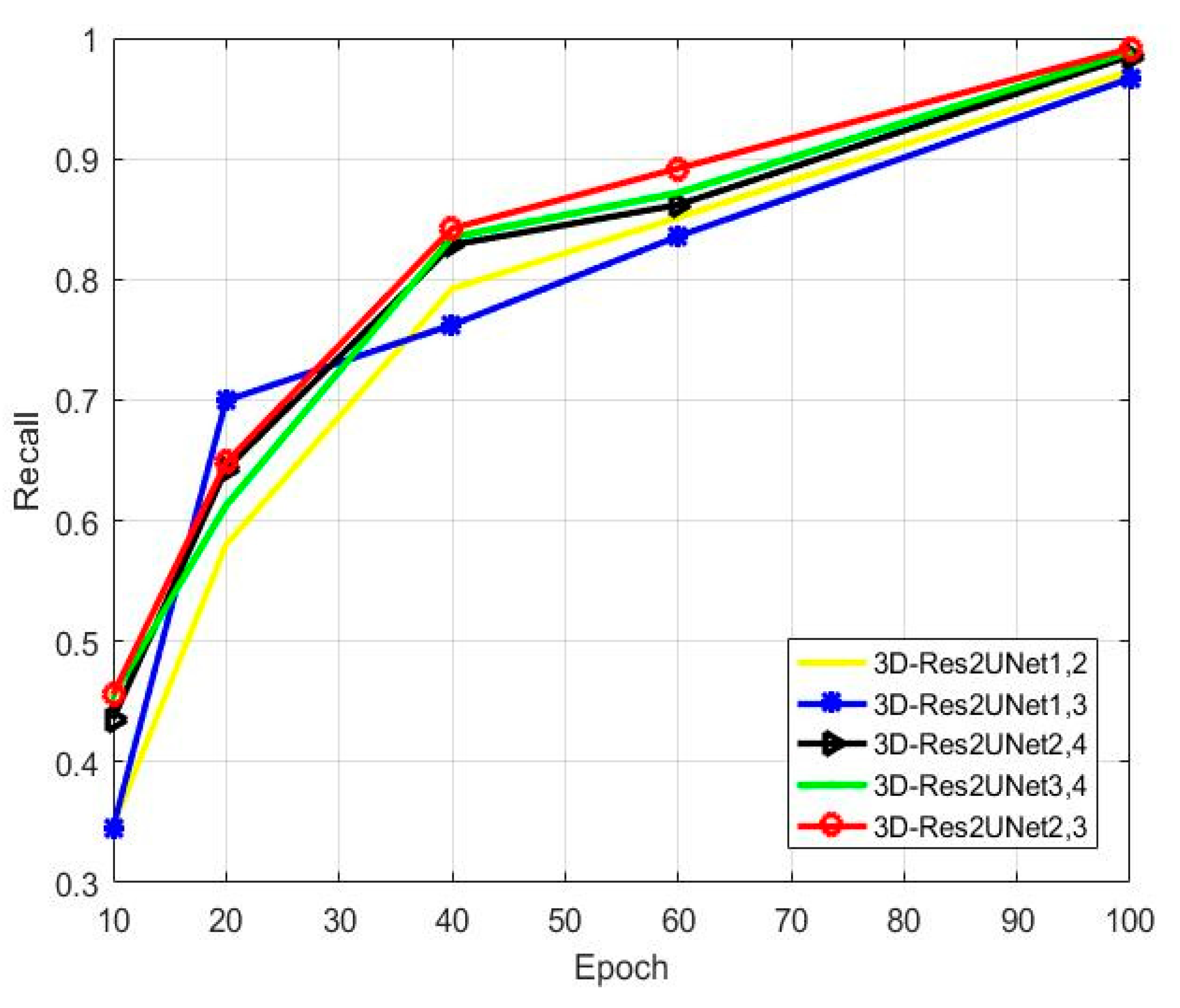
Figure 11 shows the changes in the recall rate caused by the 3D-Res2Net module that is being added to different positions. Among them, the recall rate is the best when the module is added to the second and third down-sampling.
The method proposed in this paper has a dice coefficient of 95.30%, a recall rate of 99.1%, and a false positive nodule: 276.3 false positive nodules appear on each CT sample. These results are significantly better than the other design methods in terms of segmentation fit and recall.
5. Discussion
This article quantitatively analyzes the performance of the network through experiments. Firstly, for the situation of pulmonary nodules with variable shapes and different sizes, we chose the Res2Net module for improvement. The module is upgraded from two- to three-dimensional. This is because the 3D-Res2UNet network uses the means of decomposing and then fusing the input to effectively perform the multi-layer feature extraction. It helps improve the segmentation accuracy of the network, and it can more accurately distinguish the edges of lung nodules from other irrelevant tissues. Secondly, adding the 3D-Res2Net module twice during down-sampling is to be able to complement information at multiple levels. With the continuous enlargement of the feature map, the multi-layer detection network can progressively capture the missing target points in the previous layer to achieve better results.
The network proposed in this paper has achieved good results in the segmentation of lung nodules, and has made great progress in the segmentation of small nodules. This method can be effectively applied to other fields related to small detection and segmentation tasks. However, this method also has limitations, including high training costs and high false positives. This is also the direction of continued efforts in the future. It can be expected that 3D detection and segmentation methods will have better development in the future.
6. Conclusions
This paper proposes a CT image lung nodule segmentation method based on 3D-UNet and Res2Net. The 3D-Res2UNet neural network improves the training speed of the model while making the segmentation method more complete. Before the experiment, the preparation of and preprocessing methodology applied to the experimental data are introduced. After the experiment, the experimental results of the method in this paper are shown. The experimental results are compared with an original basic network, an existing same-task network, and the method presented in this paper. After testing, the method in this paper demonstrated better results than the other methods in terms of the dice coefficient and recall rate. The method in this paper also has a good overall performance in lung nodule segmentation.
Author Contributions
Conceptualization, Z.X.; validation, Z.X. and B.L.; investigation, L.G.; resources, F.Z.; writing—original draft preparation, B.L.; writing—review and editing, Y.L.; supervision, Y.L.; project administration, F.Z. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by Tianjin Science and Technology Major Projects and Engineering (No. 17ZXSCSY00060, No. 17ZXHLSY00040) and the Program for Innovative Research Team in the University of Tianjin (No. TD13-5034).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2019. CA Cancer J. Clin. 2019, 69, 7–34. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, A.O.; Sampaio, W.B.; Silva, A.C.; de Paiva, A.C.; Nunes, R.A.; Gattass, M. Automatic detection of solitary lung nodules using quality threshold clustering, genetic algorithm and diversity index. Artif. Intell. Med. 2014, 60, 165–177. [Google Scholar] [CrossRef] [PubMed]
- Messay, T.; Hardie, R.C.; Rogers, S.K. A new computationally efficient CAD system for pulmonary nodule detection in CT imagery. Med Image Anal. 2010, 14, 390–406. [Google Scholar] [CrossRef] [PubMed]
- Ding, J.; Li, A.; Hu, Z. Accurate pulmonary nodule detection in computed tomography images using deep convolutional neural networks. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI 2017), Quebec City, QC, Canda, 10–14 September 2017; pp. 559–567. [Google Scholar]
- Deng, Z.; Chen, X. Pulmonary nodule detection algorithm based on deep convolutional neural network. J. Comput. Appl. 2019, 39, 2019–2115. [Google Scholar]
- Zhu, W.; Liu, C.; Fan, W.; Xie, X. Deeplung: Deep 3D dual path nets for automated pulmonary nodule detection and classification. In Proceedings of the 18th IEEE Winter Conference on Applications of Computer Vision, Lake Tahoe, Stateline, NV, USA; 2017; pp. 673–681. [Google Scholar]
- Gong, L.; Jiang, S.; Yang, Z.Y.; Zhang, G.; Wang, L. Automated pulmonary nodule detection in CT images using 3D deep squeeze-and-excitation networks. Int. J. Comput. Assist. Radiol. Surg. 2019, 14, 1969–1979. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Munich, Germany, 5–9 October 2015; 9351, pp. 234–241. Available online: https://arxiv.org/abs/1505.04597v1 (accessed on 11 September 2020).
- Çiçek, Ö.; Abdulkadir, A.; Lienkamp, S.S.; Brox, T.; Ronneberger, O. 3D U-Net: Learning Dense Volumetric Segmentation from Sparse Annotation. In Proceedings of the International Conference on Medical Image Computing and Computer Assisted Intervention, Athens, Greece, 17–21 October 2016; pp. 424–432. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition; IEEE: New York, NY, USA, 2016; pp. 770–778. Available online: http://arXiv.org/pdf/1512.03385 (accessed on 15 September 2020).
- Gao, S.; Cheng, M.M.; Zhao, K.; Zhang, X.Y.; Yang, M.H.; Torr, P.H. Res2Net: A New Multi-scale Backbone Architecture. In IEEE Transactions on Pattern Analysis and Machine Intelligence; IEEE: New York, NY, USA, 2019. [Google Scholar] [CrossRef]
- Armato, S.G.; Mclennan, G.; Bidautl, L.; McNitt-Gray, M.F.; Meyer, C.R.; Reeves, A.P.; Zhao, B.; Aberle, D.R.; Henschke, C.I.; Hoffman, E.A.; et al. The lung image database consortium (LIDC) and image database resource initiative (IDRI): A completed reference database of lung nodules on CT scans. Med. Phys. 2011, 38, 915–931. [Google Scholar] [CrossRef] [PubMed]
- Mansoor, A.; Bagci, U.; Xu, Z.; Foster, B.; Olivier, K.N.; Elinoff, J.M.; Suffredini, A.F.; Jayaram, K.U.; Mollura, D.J. A generic approach to pathological lung segmentation. IEEE Trans. Med. Imaging 2014, 33, 2293–2310. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.W.; Lin, C.T.; Chen, S.P.; Wu, Y.-Y.; Hsu, P.-H.; Lai, S.-H. AugGAN: Cross Domain Adaptation with GAN-based Data Augmentation. In ECCV 2018: European Conference on Computer Vision; Springer: Cham, UK, 2018; pp. 731–744. [Google Scholar] [CrossRef]
- Dice, L.R.; Dice, L.R. Measures of the Amount of Ecologic Association Between Species. Ecology 1944, 26, 297–302. [Google Scholar] [CrossRef]
- Hou, T.X.; Zhao, J.J.; Qiang, Y. Pulmonary nodules segmentation based on CRF 3D-UNet structure. Comput. Eng. Des. 2020, 41, 1663–1669. [Google Scholar]
- Murphy, K.; Ginneken, B.V.; Schilham, A.M.R.; de Hoop, B.J.; Gietema, H.A.; Prokop, M. A large-scale evaluation of automatic pulmonary nodule detection in chest CT using local image features and k-nearest-neighbour classification. Med. Image Anal. 2009, 13, 757–770. [Google Scholar] [CrossRef] [PubMed]
- Tan, M.; Deklerck, R.; Jansen, B.; Bister, M.; Corneli, J. A novel computer-aided lung nodule detection system for CT images. Med. Phys. 2011, 38, 5630–5645. [Google Scholar] [CrossRef] [PubMed]
- Setio, A.A.A.; Ciompi, F.; Litjens, G.; Gerke, P.; Jacobs, C.; van Riel, S.J.; Winkler, M.M.; Naqibullah, M.; Sánchez, C.I.; van Ginneken, B. Pulmonary nodule detection in CT images: False positive reduction using multi-view convolutional networks. IEEE Trans. Med. Imaging 2016, 35, 1160–1169. [Google Scholar] [CrossRef] [PubMed]
- Lung Nodule Analysis 2016. Available online: https://luna16.grand-challenge.org (accessed on 18 January 2016).
- Dou, Q.; Chen, H.; Jin, Y. Automated pulmonary nodule detection via 3D convnets with online sample filtering and hybrid-loss residual learning. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI 2017), Quebec City, QC, Canada, 10–14 September 2017; pp. 630–638. [Google Scholar]
Figure 1.
Three-dimensional residual network (3D-Res2Net) module.
Figure 1.
Three-dimensional residual network (3D-Res2Net) module.
Figure 2.
Three-dimensional residual UNet (3D-Res2UNet) network structure diagram.
Figure 2.
Three-dimensional residual UNet (3D-Res2UNet) network structure diagram.
Figure 3.
CT image of lung.
Figure 3.
CT image of lung.
Figure 4.
Results of lung parenchymal segmentation.
Figure 4.
Results of lung parenchymal segmentation.
Figure 5.
Lung nodules generated by the adversarial network.
Figure 5.
Lung nodules generated by the adversarial network.
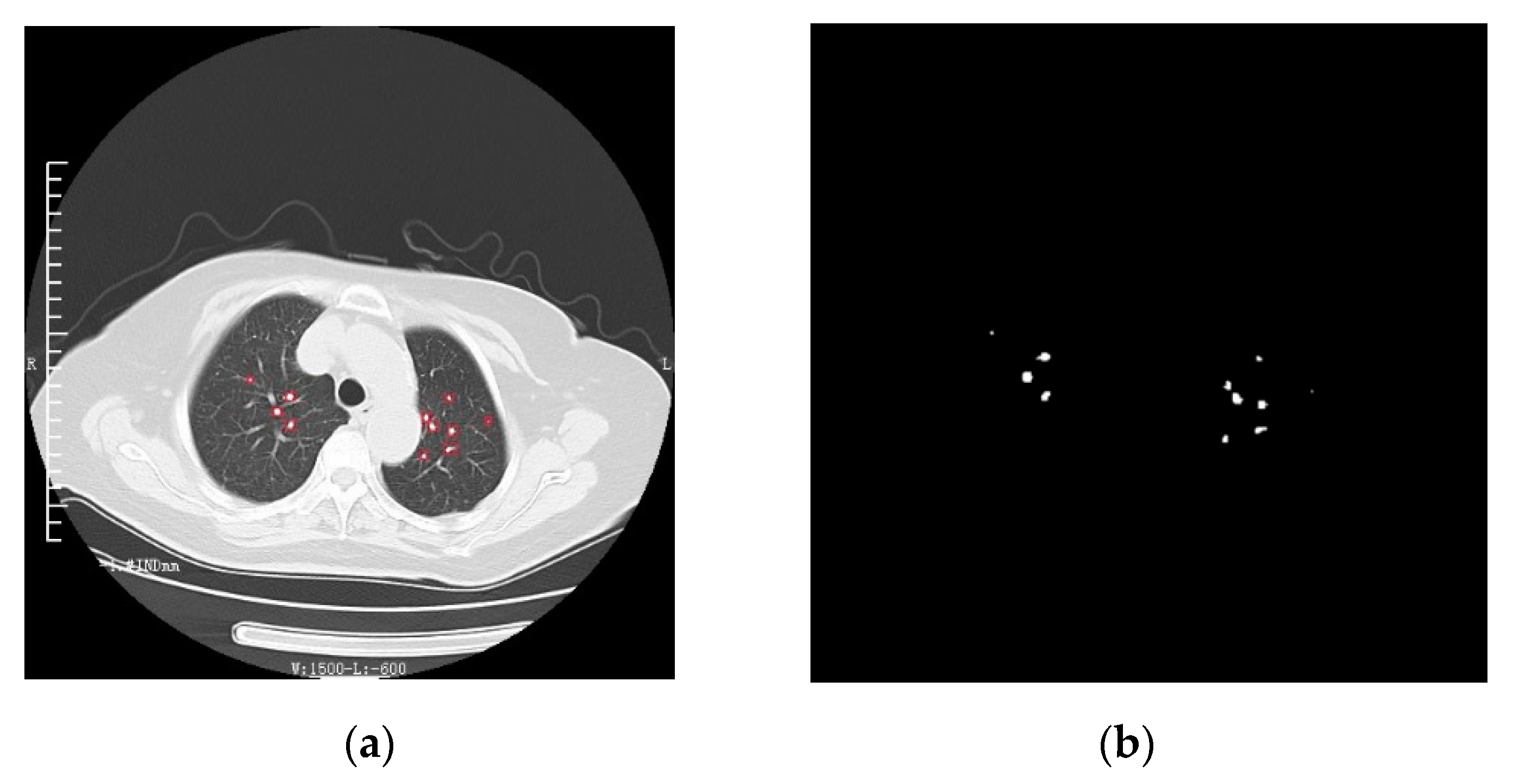
Figure 6.
The detection results of lung nodules. (a) Results of lung nodule detection. (b) The overall effect of lung nodule segmentation.
Figure 6.
The detection results of lung nodules. (a) Results of lung nodule detection. (b) The overall effect of lung nodule segmentation.
Figure 7.
Local effect diagram of lung nodule segmentation.
Figure 7.
Local effect diagram of lung nodule segmentation.
Figure 8.
Image of lung nodules whose segmentation effect is not very accurate.
Figure 8.
Image of lung nodules whose segmentation effect is not very accurate.
Figure 9.
3D-Res2UNet network dice coefficient.
Figure 9.
3D-Res2UNet network dice coefficient.
Figure 10.
3D-Res2UNet contrast network dice coefficient. (a) Describe the dice coefficient of 3D-Res2UNet 1,2, (b) describe the dice coefficient of 3D-Res2UNet 1,3, (c) describe the dice coefficient of 3D-Res2UNet 2,4, (d) describe the dice coefficient of 3D-Res2UNet 3,4.
Figure 10.
3D-Res2UNet contrast network dice coefficient. (a) Describe the dice coefficient of 3D-Res2UNet 1,2, (b) describe the dice coefficient of 3D-Res2UNet 1,3, (c) describe the dice coefficient of 3D-Res2UNet 2,4, (d) describe the dice coefficient of 3D-Res2UNet 3,4.
Figure 11.
Comparison of recall rates of the 3D-Res2Net module added to different positions.
Figure 11.
Comparison of recall rates of the 3D-Res2Net module added to different positions.
Table 1.
Labeling information of pulmonary nodules in a CSV file.
Table 1.
Labeling information of pulmonary nodules in a CSV file.
| Seriesuid | CoordX | CoordY | CoordZ | Diameter (mm) |
|---|
| LKDS-00539 | −70.7218023 | −87.5886319 | 36.5 | 9.4028638 |
| LKDS-00540 | −50.2048319 | −90.0324701 | 50.5 | 7.9106295 |
Table 2.
CT values of different human tissues.
Table 2.
CT values of different human tissues.
| Name of Organization | HU Values |
|---|
| Air | −1000 |
| Lung | −500 |
| Fat | −100~−50 |
| Water | 0 |
| Aorta | 35~50 |
| Kidney | 40~60 |
| Bones | 150~3000 |
Table 3.
The dice coefficient of different networks.
Table 3.
The dice coefficient of different networks.
| Network Name | Dice (%) |
|---|
| UNet | 81.32 |
| 3D-UNet | 89.12 |
| 3D-UNet+fully CRF [16] | 93.25 |
| 3D-Res2UNet (Ours) | 95.30 |
Table 4.
Comparison of detection algorithms.
Table 4.
Comparison of detection algorithms.
| Algorithm | Recall (%) | Number of False Positive Lung Nodules/CT |
|---|
| ISICAD [17] | 85.7 | 329.3 |
| ETROCAD [18] | 92.2 | 333.0 |
| DIAG_CONVENT [19] | 93.3 | 269.0 |
| LUNA16_V1 [20] | 94.4 | 622.0 |
| Dou [21] | 97.1 | 219.1 |
| LUNA16_V2 [20] | 98.3 | 850.2 |
| 3D-Res2UNet (Ours) | 99.1 | 276.3 |
Table 5.
Comparison of the parameters of the 3D-Res2Net module in different positions.
Table 5.
Comparison of the parameters of the 3D-Res2Net module in different positions.
| Name | Dice | Recall | Number of False Positive Lung Nodules/CT |
|---|
| 3D-Res2UNet 1,2 | 92.4 | 97.2 | 320.7 |
| 3D-Res2UNet 1,3 | 92.7 | 96.6 | 351.5 |
| 3D-Res2UNet 2,4 | 93.50 | 98.5 | 300.1 |
| 3D-Res2UNet 3,4 | 94.52 | 98.8 | 330.4 |
| 3D-Res2UNet 2,3 | 95.30 | 99.1 | 276.3 |
| Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).