The Immobilization of Oxindole Derivatives with Use of Cube Rhombellane Homeomorphs
Abstract
1. Introduction
2. Methods
3. Results and Discussion
The Binding Affinity and Conformational Diversity
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Damascelli, B.; Patelli, G.L.; Lanocita, R.; Di Tolla, G.; Frigerio, L.F.; Marchianò, A.; Garbagnati, F.; Spreafico, C.; Tichà, V.; Gladin, C.R.; et al. A Novel Intraarterial Chemotherapy Using Paclitaxel in Albumin Nanoparticles to Treat Advanced Squamous Cell Carcinoma of the Tongue: Preliminary Findings. Am. J. Roentgenol. 2003, 181, 253–260. [Google Scholar] [CrossRef] [PubMed]
- Huang, M.; Khor, E.; Lim, L.-Y. Uptake and cytotoxicity of chitosan molecules and nanoparticles: Effects of molecular weight and degree of deacetylation. Pharm. Res. 2004, 21, 344–353. [Google Scholar] [CrossRef] [PubMed]
- Dyer, A.M.; Hinchcliffe, M.; Watts, P.; Castile, J.; Jabbal-Gill, I.; Nankervis, R.; Smith, A.; Illum, L. Nasal delivery of insulin using novel chitosan based formulations: A comparative study in two animal models between simple chitosan formulations and chitosan nanoparticles. Pharm. Res. 2002, 19, 998–1008. [Google Scholar] [CrossRef] [PubMed]
- Cascone, M.G.; Lazzeri, L.; Carmignani, C.; Zhu, Z. Gelatin nanoparticles produced by a simple W/O emulsion as delivery system for methotrexate. J. Mater. Sci. Mater. Med. 2002, 13, 523–526. [Google Scholar] [CrossRef] [PubMed]
- Paciotti, G.F.; Myer, L.; Weinreich, D.; Goia, D.; Pavel, N.; McLaughlin, R.E.; Tamarkin, L. Colloidal gold: A novel nanoparticle vector for tumor directed drug delivery. Drug Deliv. 2004, 11, 169–183. [Google Scholar] [CrossRef]
- Hainfeld, J.F.; Slatkin, D.N.; Smilowitz, H.M. The use of gold nanoparticles to enhance radiotherapy in mice. Phys. Med. Biol. 2004, 49, N309–N315. [Google Scholar] [CrossRef] [PubMed]
- Gupta, M.; Gupta, A.K. Hydrogel pullulan nanoparticles encapsulating pBUDLacZ plasmid as an efficient gene delivery carrier. J. Control. Release 2004, 99, 157–166. [Google Scholar] [CrossRef] [PubMed]
- Gupta, A.K.; Gupta, M. Synthesis and surface engineering of iron oxide nanoparticles for biomedical applications. Biomaterials 2005, 26, 3995–4021. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.Y.; Lee, Y.M.; Baik, D.J.; Kang, J.S. Toxic characteristics of methoxy poly(ethylene glycol)/poly(epsilon-caprolactone) nanospheres; in vitro and in vivo studies in the normal mice. Biomaterials 2003, 24, 55–63. [Google Scholar] [CrossRef]
- Alyautdin, R.N.; Petrov, V.E.; Langer, K.; Berthold, A.; Kharkevich, D.A.; Kreuter, J. Delivery of loperamide across the blood-brain barrier with polysorbate 80-coated poly(butylcyanoacrylate) nanoparticles. Pharm. Res. 1997, 14, 325–328. [Google Scholar] [CrossRef]
- Kreuter, J.; Ramge, P.; Petrov, V.; Hamm, S.; Gelperina, S.E.; Engelhardt, B.; Alyautdin, R.; von Briesen, H.; Begley, D.J. Direct evidence that polysorbate-80-coated poly(butylcyanoacrylate) nanoparticles deliver drugs to the CNS via specific mechanisms requiring prior binding of drug to the nanoparticles. Pharm. Res. 2003, 20, 409–416. [Google Scholar] [CrossRef] [PubMed]
- Panyam, J.; Zhou, W.Z.; Prabha, S.; Sahoo, S.K.; Labhasetwar, V. Rapid endo-lysosomal escape of poly(dl-lactide-co-glycolide) nanoparticles: Implications for drug and gene delivery. FASEB J. 2002, 16, 1217–1226. [Google Scholar] [CrossRef] [PubMed]
- Weissenböck, A.; Wirth, M.; Gabor, F. WGA-grafted PLGA-nanospheres: Preparation and association with Caco-2 single cells. J. Control. Release 2004, 99, 383–392. [Google Scholar] [CrossRef] [PubMed]
- Cataldo, F.; Da Ros, T. Medicinal Chemistry and Pharmacological Potential of Fullerenes and Carbon Nanotubes; Springer: Berlin, Germany, 2008; ISBN 9781402068454. [Google Scholar]
- De Jong, W.H.; Borm, P.J.A. Drug delivery and nanoparticles: Applications and hazards. Int. J. Nanomed. 2008, 3, 133–149. [Google Scholar] [CrossRef]
- Andrievsky, G.; Klochkov, V.; Derevyanchenko, L. Is the C60 Fullerene Molecule Toxic?! Fuller. Nanotub. Carbon Nanostructures 2005, 13, 363–376. [Google Scholar] [CrossRef]
- Szefler, B. Nanotechnology, from quantum mechanical calculations up to drug delivery. Int. J. Nanomed. 2018, 13, 6143–6176. [Google Scholar] [CrossRef] [PubMed]
- Panchuk, R.R.; Prylutska, S.V.; Chumakl, V.V.; Skorokhyd, N.R.; Lehka, L.V.; Evstigneev, M.P.; Prylutskyy, Y.I.; Berger, W.; Heffeter, P.; Scharff, P.; et al. Application of C60 Fullerene-Doxorubicin Complex for Tumor Cell Treatment In Vitro and In Vivo. J. Biomed. Nanotechnol. 2015, 11, 1139–1152. [Google Scholar] [CrossRef]
- Morgen, M.; Bloom, C.; Beyerinck, R.; Bello, A.; Song, W.; Wilkinson, K.; Steenwyk, R.; Shamblin, S. Polymeric Nanoparticles for Increased Oral Bioavailability and Rapid Absorption Using Celecoxib as a Model of a Low-Solubility, High-Permeability Drug. Pharm. Res. 2012, 29, 427–440. [Google Scholar] [CrossRef]
- Gao, Z.; Zhang, L.; Sun, Y. Nanotechnology applied to overcome tumor drug resistance. J. Control. Release 2012, 162, 45–55. [Google Scholar] [CrossRef]
- Turov, V.V.; Chehun, V.F.; Barvinchenko, V.N.; Krupskaya, T.V.; Prylutskyy, Y.I.; Scharff, P.; Ritter, U. Low-temperature 1H-NMR spectroscopic study of doxorubicin influence on the hydrated properties of nanosilica modified by DNA. J. Mater. Sci. Mater. Med. 2011, 22, 525–532. [Google Scholar] [CrossRef]
- Schuetze, C.; Ritter, U.; Scharff, P.; Fernekorn, U.; Prylutska, S.; Bychko, A.; Rybalchenko, V.; Prylutskyy, Y. Interaction of N-fluorescein-5-isothiocyanate pyrrolidine-C60 with a bimolecular lipid model membrane. Mater. Sci. Eng. C 2011, 31, 1148–1150. [Google Scholar] [CrossRef]
- Qiao, R.; Roberts, A.P.; Mount, A.S.; Klaine, S.J.; Ke, P.C. Translocation of C60 and Its Derivatives Across a Lipid Bilayer. Nano Lett. 2007, 7, 614–619. [Google Scholar] [CrossRef] [PubMed]
- Prylutska, S.; Bilyy, R.; Overchuk, M.; Bychko, A.; Andreichenko, K.; Stoika, R.; Rybalchenko, V.; Prylutskyy, Y.; Tsierkezos, N.G.; Ritter, U. Water-soluble pristine fullerenes C60 increase the specific conductivity and capacity of lipid model membrane and form the channels in cellular plasma membrane. J. Biomed. Nanotechnol. 2012, 8, 522–527. [Google Scholar] [CrossRef] [PubMed]
- Prylutska, S.V.; Grynyuk, I.I.; Grebinyk, S.M.; Matyshevska, O.P.; Prylutskyy, Y.I.; Ritter, U.; Siegmund, C.; Scharff, P. Comparative study of biological action of fullerenes C60 and carbon nanotubes in thymus cells. Materwiss. Werksttech. 2009, 40, 238–241. [Google Scholar] [CrossRef]
- Johnston, H.J.; Hutchison, G.R.; Christensen, F.M.; Aschberger, K.; Stone, V. The Biological Mechanisms and Physicochemical Characteristics Responsible for Driving Fullerene Toxicity. Toxicol. Sci. 2010, 114, 162–182. [Google Scholar] [CrossRef]
- Evstigneev, M.P.; Buchelnikov, A.S.; Voronin, D.P.; Rubin, Y.V.; Belous, L.F.; Prylutskyy, Y.I.; Ritter, U. Complexation of C60 Fullerene with Aromatic Drugs. Chem Phys Chem 2013, 14, 568–578. [Google Scholar] [CrossRef] [PubMed]
- Czeleń, P. Molecular dynamics study on inhibition mechanism of CDK-2 and GSK-3β by CHEMBL272026 molecule. Struct. Chem. 2016, 27, 1807–1818. [Google Scholar] [CrossRef]
- Czeleń, P. Inhibition mechanism of CDK-2 and GSK-3β by a sulfamoylphenyl derivative of indoline—A molecular dynamics study. J. Mol. Model. 2017, 23, 230. [Google Scholar] [CrossRef]
- Besson, A.; Dowdy, S.F.; Roberts, J.M. CDK Inhibitors: Cell Cycle Regulators and Beyond. Dev. Cell 2008, 14, 159–169. [Google Scholar] [CrossRef] [PubMed]
- Canavese, M.; Santo, L.; Raje, N. Cyclin dependent kinases in cancer. Cancer Biol. Ther. 2012, 13, 451–457. [Google Scholar] [CrossRef] [PubMed]
- Child, E.S.; Hendrychov, T.; McCague, K.; Futreal, A.; Otyepka, M.; Mann, D.J. A cancer-derived mutation in the PSTAIRE helix of cyclin-dependent kinase 2 alters the stability of cyclin binding. Biochim. Biophys. Acta Mol. Cell Res. 2010, 1803, 858–864. [Google Scholar] [CrossRef] [PubMed]
- Malumbres, M.; Barbacid, M. Cell cycle, CDKs and cancer: A changing paradigm. Nat. Rev. Cancer 2009, 9, 153–166. [Google Scholar] [CrossRef] [PubMed]
- Malumbres, M.; Barbacid, M. To cycle or not to cycle: A critical decision in cancer. Nat. Rev. Cancer 2001, 1, 222–231. [Google Scholar] [CrossRef] [PubMed]
- Diudea, M.V.; Lungu, C.N.; Nagy, C.L.; Diudea, M.V.; Lungu, C.N.; Nagy, C.L. Cube-Rhombellane Related Structures: A Drug Perspective. Molecules 2018, 23, 2533. [Google Scholar] [CrossRef] [PubMed]
- Szefler, B.; Czeleń, P.; Diudea, M. V Docking of indolizine derivatives on cube rhombellane functionalized homeomorphs. Stud. Univ. Babes-Bolyai Chem. 2018, 63, 7–18. [Google Scholar] [CrossRef]
- ChEMBL. Available online: https://www.ebi.ac.uk/chembl/ (accessed on 1 March 2016).
- Kim, K.H.; Ko, D.K.; Kim, Y.T.; Kim, N.H.; Paul, J.; Zhang, S.Q.; Murray, C.B.; Acharya, R.; Kim, Y.H.; DeGrado, W.F.; et al. Protein-directed self-assembly of a fullerene crystal. Nat. Commun. 2016, 7, 11429. [Google Scholar] [CrossRef] [PubMed]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Bartashevich, E.V.; Potemkin, V.A.; Grishina, M.A.; Belik, A.V. A Method for Multiconformational Modeling of the Three-Dimensional Shape of a Molecule. J. Struct. Chem. 2002, 43, 1033–1039. [Google Scholar] [CrossRef]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
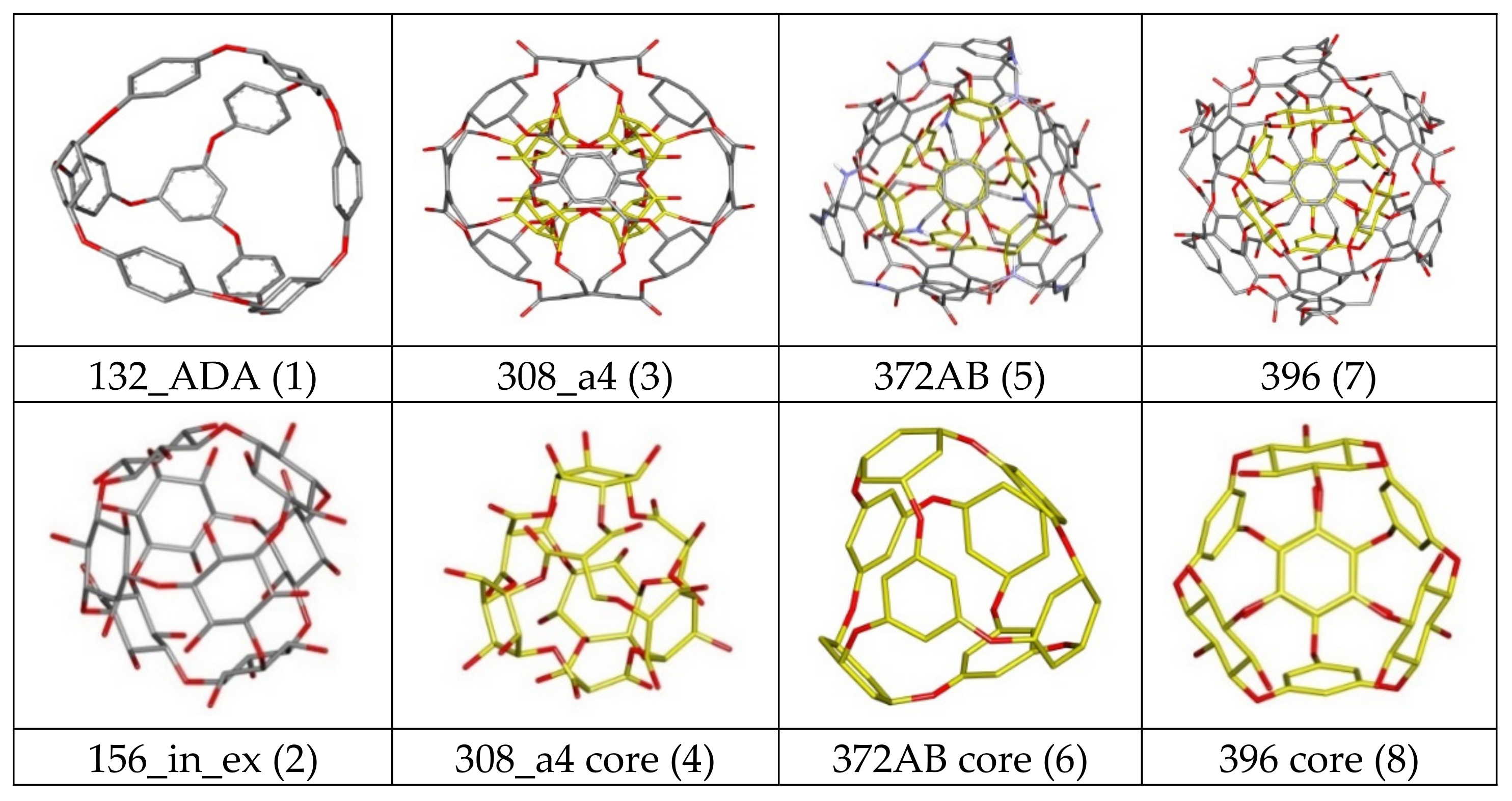
| Nanostructure Name | Structure Type | Chem.Type Core/ex. Shell | Atoms Quantity | ||||
|---|---|---|---|---|---|---|---|
| All | C | H | O | N | |||
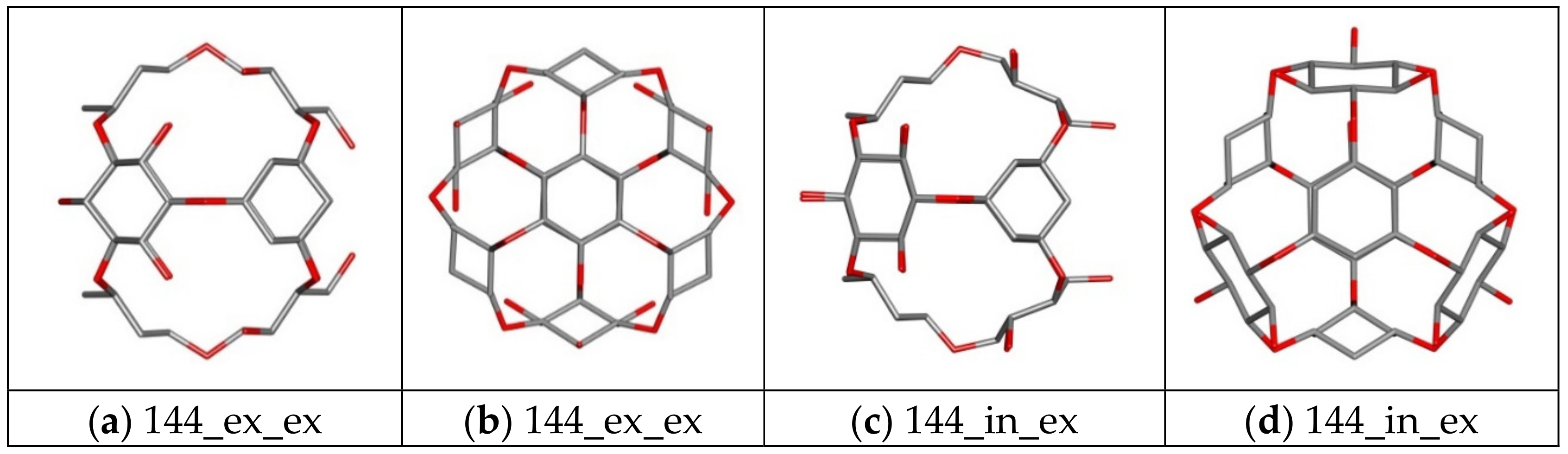
| 144 ex_ex/in_ex | Core | Ether | 144 | 48 | 84 | 24 | 0 |
| 156 ex_ex/in_ex | Core | Ether | 156 | 48 | 84 | 36 | 0 |
| 308 a4/b4 | C-rbl | Ether/Ester | 308 | 100 | 124 | 84 | 0 |
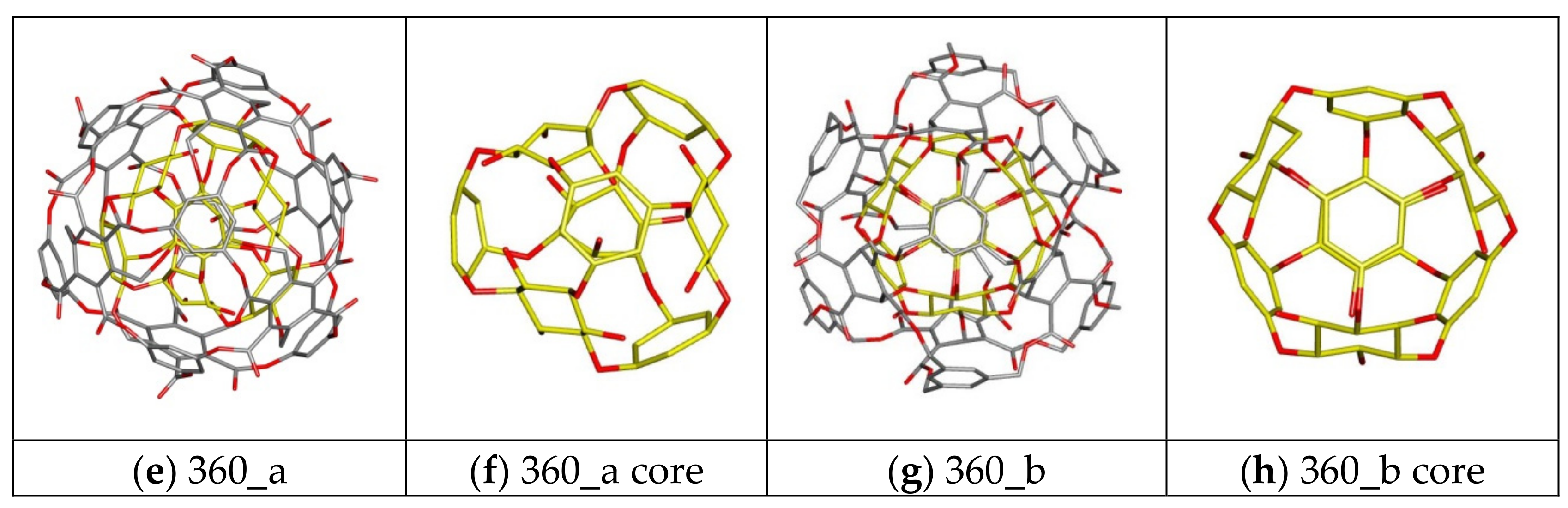
| 360 a/b | C-rbl | Ether/Ester | 360 | 168 | 108 | 84 | 0 |
| 372AB | C-rbl | Ether/Amide; Ester | 372 | 180 | 120 | 60 | 12 |
| 396 | C-rbl | Ether/Ester | 396 | 192 | 132 | 72 | 0 |
| 420 | C-rbl | Ether/Amide | 420 | 192 | 156 | 48 | 24 |
| 444 | C-rbl | Ether/Amide | 444 | 192 | 180 | 48 | 24 |
| 456 | C-rbl | Ether/Amide | 456 | 192 | 180 | 60 | 24 |
| ADA_132 | ADA/rbl | Ether | 132 | 60 | 60 | 12 | 0 |
| Nanostructure Name | ΔG [kcal/mol] | Binding Constant [Kmax] | Difference of Kmax Relative to C60 [%] | |||
|---|---|---|---|---|---|---|
| MAX | MIN | AVERAGE | SD | |||
| 144_ex_ex | −3.50 | −3.20 | −3.30 | 0.12 | 367.73 | −97.11 |
| 144_in_ex | −3.90 | −3.70 | −3.81 | 0.09 | 722.33 | −94.33 |
| 156_ex_ex | −4.10 | −3.70 | −3.88 | 0.11 | 1012.37 | −92.05 |
| 156_in_ex | −4.20 | −3.80 | −3.92 | 0.14 | 1198.51 | −90.59 |
| 308a4 | −6.00 | −5.60 | −5.76 | 0.13 | 25,006.81 | 96.43 |
| 308b4 | −5.90 | −5.50 | −5.74 | 0.14 | 21,123.08 | 65.92 |
| 360a | −6.20 | −5.90 | −6.02 | 0.11 | 35,047.76 | 175.30 |
| 360b | −5.50 | −5.10 | −5.26 | 0.11 | 10,753.59 | −15.53 |
| 372AB | −6.10 | −5.50 | −5.67 | 0.21 | 29,604.61 | 132.54 |
| 396 | −5.60 | −5.20 | −5.37 | 0.13 | 12,730.76 | 0.00 |
| 420 | −5.70 | −5.20 | −5.42 | 0.15 | 15,071.46 | 18.39 |
| 444 | −5.30 | −5.00 | −5.12 | 0.10 | 7672.76 | −39.73 |
| 456 | −5.50 | −5.00 | −5.26 | 0.17 | 10,753.59 | −15.53 |
| ADA_132 | −5.80 | −5.50 | −5.58 | 0.10 | 17,842.53 | 40.15 |
| C60 | −5.60 | −5.40 | −5.50 | 0.07 | 12,730.76 | — |
| Nanostructure Name | ΔG [kcal/mol] | Binding Constant [Kmax] | Difference of Kmax Relative to C60 [%] | |||
|---|---|---|---|---|---|---|
| MAX | MIN | AVERAGE | SD | |||
| 144_ex_ex | −3.60 | −3.50 | −3.57 | 0.05 | 435.35 | −94.33 |
| 144_in_ex | −4.00 | −3.70 | −3.80 | 0.10 | 855.14 | −88.85 |
| 156_ex_ex | −3.90 | −3.70 | −3.79 | 0.06 | 722.33 | −90.59 |
| 156_in_ex | −4.20 | −3.90 | −4.03 | 0.09 | 1198.51 | −84.38 |
| 308a4 | −5.80 | −5.50 | −5.59 | 0.11 | 17,842.53 | 132.54 |
| 308b4 | −5.80 | −5.60 | −5.72 | 0.08 | 17,842.53 | 132.54 |
| 360a | −6.30 | −5.80 | −6.08 | 0.15 | 41,491.70 | 440.77 |
| 360b | −5.10 | −4.80 | −4.92 | 0.13 | 5474.56 | −28.65 |
| 372AB | −5.80 | −5.40 | −5.58 | 0.13 | 17,842.53 | 132.54 |
| 396 | −5.80 | −5.50 | −5.68 | 0.11 | 17,842.53 | 132.54 |
| 420 | −5.60 | −5.40 | −5.49 | 0.08 | 12,730.76 | 65.92 |
| 444 | −5.00 | −4.70 | −4.83 | 0.11 | 4624.33 | −39.73 |
| 456 | −5.00 | −4.80 | −4.91 | 0.08 | 4624.33 | −39.73 |
| ADA_132 | −5.40 | −5.10 | −5.19 | 0.11 | 9083.48 | 18.39 |
| C60 | −5.30 | −5.20 | −5.21 | 0.03 | 7672.76 | — |
| Lig1 | ||||||
| 308b4 | 308a4 | 360a | 372AB | ADA 132 | C60 | |
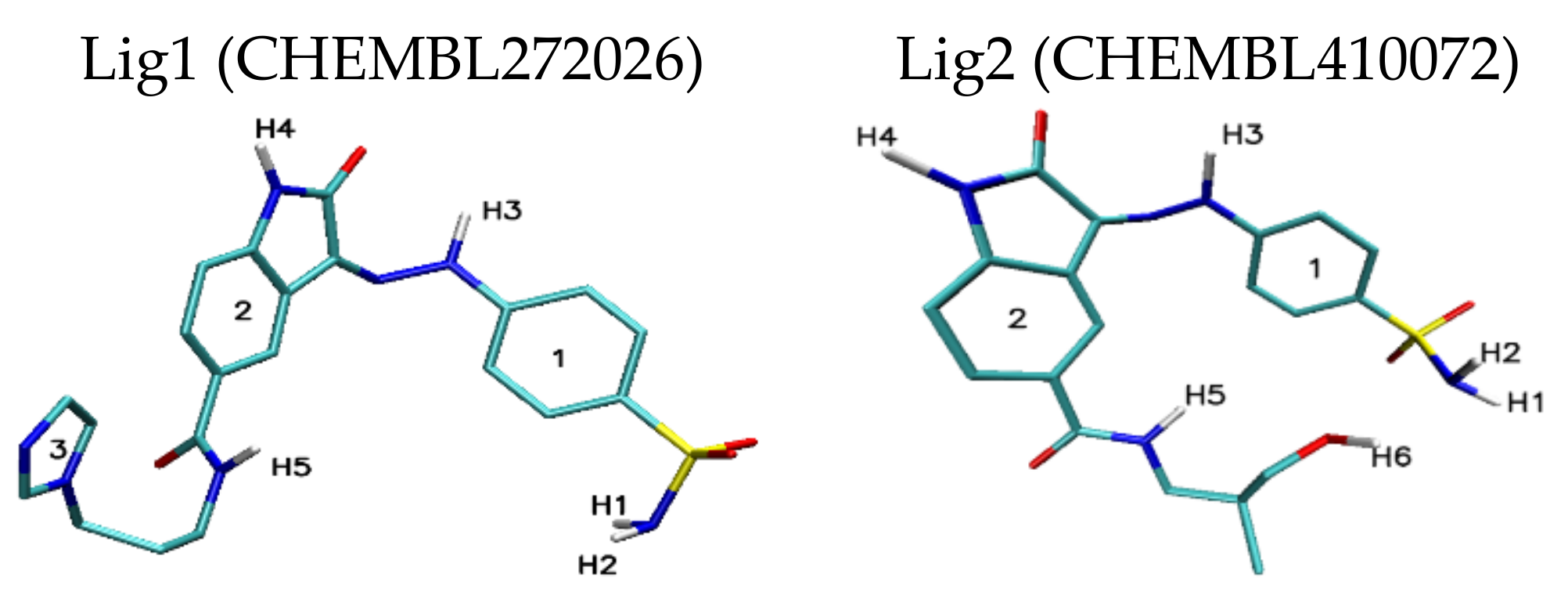
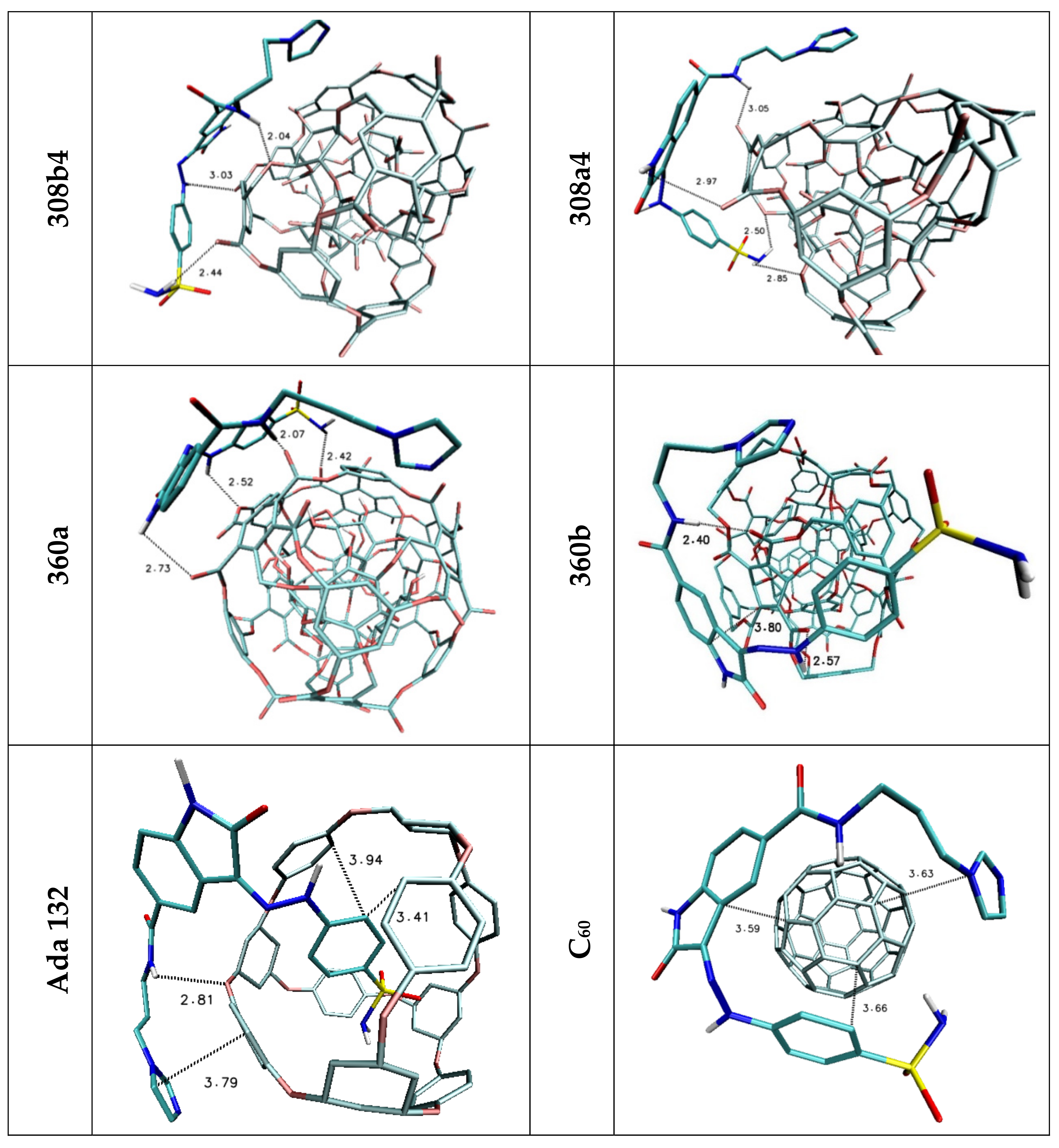
| H1/H2 | 2.44 | 2.50/2.85 | 2.42 | — | — | — |
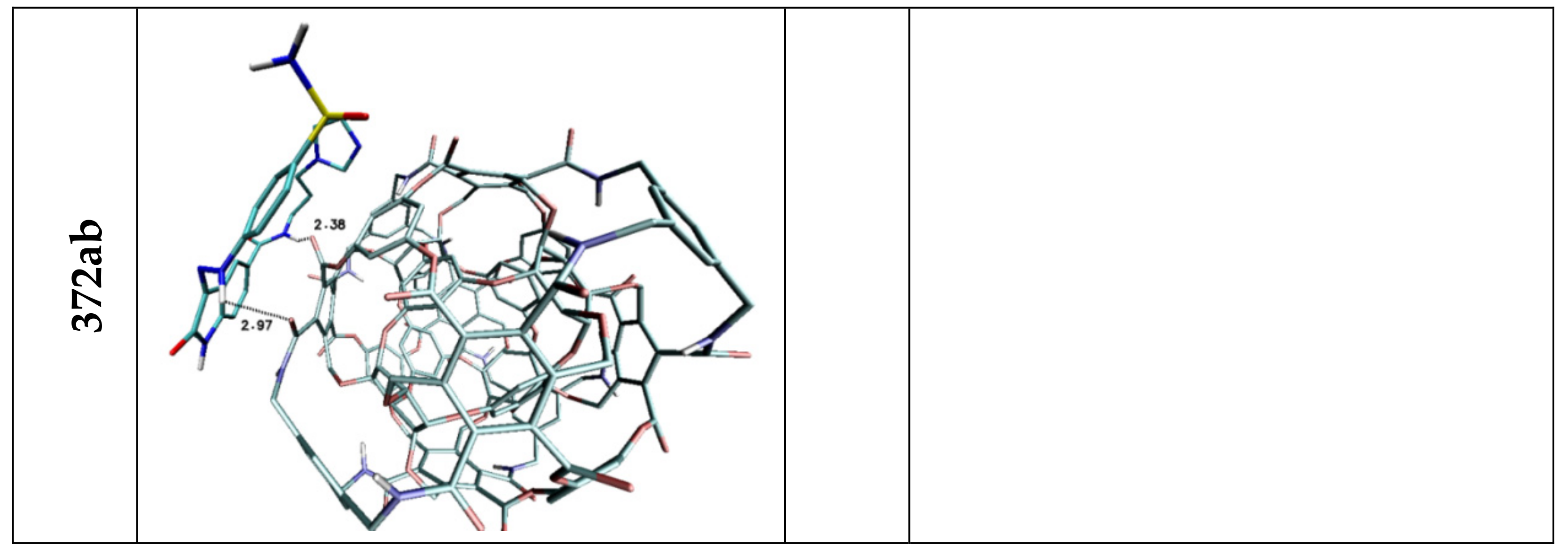
| H3 | 3.03 | — | 2.52 | 2.97 | — | — |
| H4 | — | 2.97 | 2.73 | — | — | — |
| H5 | 2.04 | 3.05 | 2.07 | 2.38 | 2.81 | — |
| Stacking 1/2/3 | 3.74/—/3.96 | —/3.74/4.34 | —/3.68/— | 3.55/3.95/— | 3.94; 3.41/—/3.79 | 3.66/3.59/3.63 |
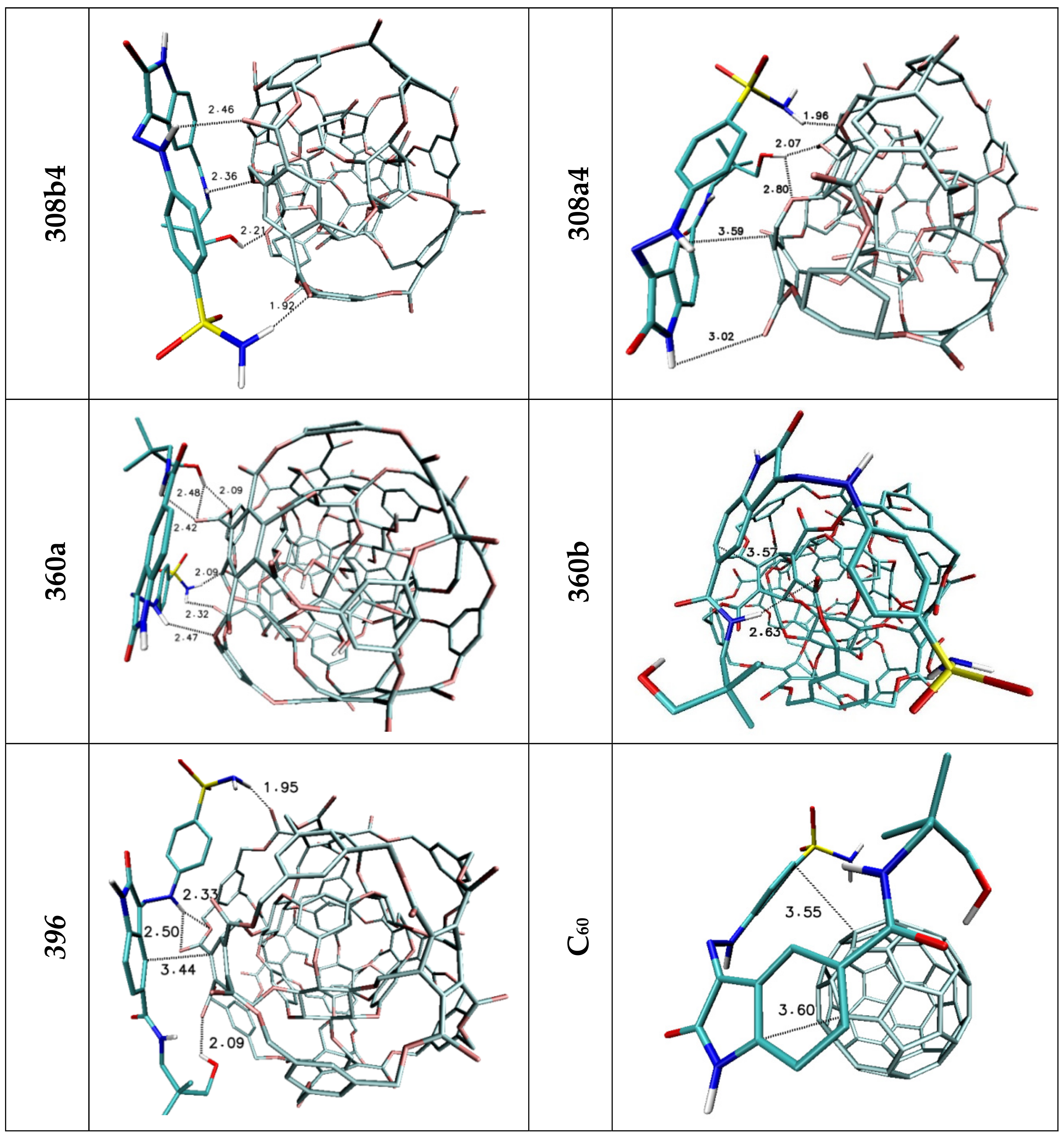
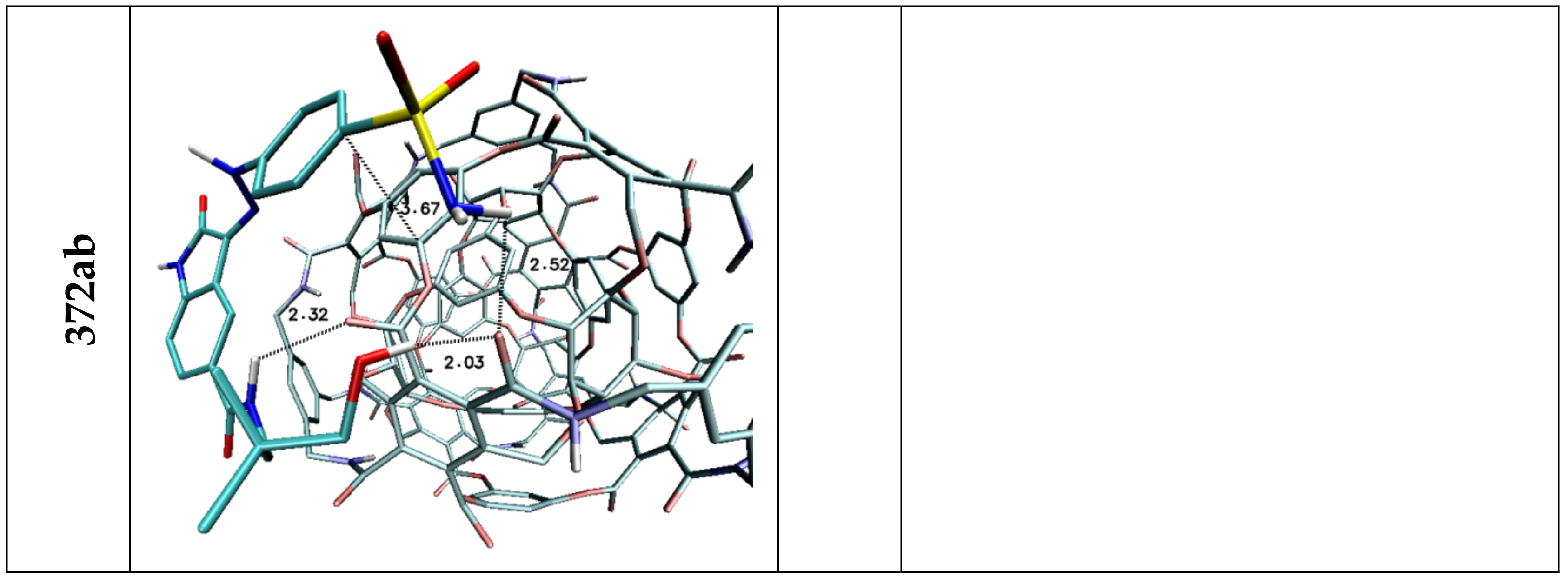
| Lig2 | ||||||
| 308b4 | 308a4 | 360a | 372AB | 396 | C60 | |
| H1/H2 | 1.92 | 1.96 | 2.09/2.32 | 2.52 | 1.95 | — |
| H3 | 2.46 | — | 2.47 | — | 2.33/2.50 | — |
| H4 | — | 3.02 | — | — | — | — |
| H5 | 2.36 | — | 2.42 | 2.32 | 3.15 | — |
| H6 | 2.21 | 2.07/2.80 | 2.09/2.48 | 2.03 | 2.09 | — |
| Stacking 1/2 | 3.59/— | —/3.59 | —/3.66 | 3.67/— | —/3.44 | 3.55/3.60 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Czeleń, P.; Szefler, B. The Immobilization of Oxindole Derivatives with Use of Cube Rhombellane Homeomorphs. Symmetry 2019, 11, 900. https://doi.org/10.3390/sym11070900
Czeleń P, Szefler B. The Immobilization of Oxindole Derivatives with Use of Cube Rhombellane Homeomorphs. Symmetry. 2019; 11(7):900. https://doi.org/10.3390/sym11070900
Chicago/Turabian StyleCzeleń, Przemysław, and Beata Szefler. 2019. "The Immobilization of Oxindole Derivatives with Use of Cube Rhombellane Homeomorphs" Symmetry 11, no. 7: 900. https://doi.org/10.3390/sym11070900
APA StyleCzeleń, P., & Szefler, B. (2019). The Immobilization of Oxindole Derivatives with Use of Cube Rhombellane Homeomorphs. Symmetry, 11(7), 900. https://doi.org/10.3390/sym11070900





