1. Introduction
The flu virus has the characteristics of high infectivity and high transmission speed. In addition, the flu is a disease that seriously threatens the health of human beings. Vaccination is universally regarded as the most important method to prevent influenza and eventually eradicate the disease. Vaccines are made using the influenza virus, which is cultured in living hatching eggs before being inactivated. Immunization can be applied to people after vaccination. A key step in the production of vaccines is to inject the virus into special egg embryos. Some egg embryos may die because of their individual differences. The dead egg embryos must be removed in time, otherwise they may contaminate other egg embryos in the same batch and even cause a serious medical safety accident. Therefore, the efficient detection and separation of necrotic hatching eggs is important for the production of vaccines.
Currently, most manufacturers still use the manual method that detects the integrity of blood vessels in hatching eggs under strong light. This method requires large-scale personnel costs, and the result is easily affected by subjective factors. In addition, because workers perform their duties under high-intensity pressure, there are many shortcomings, such as visual fatigue and low detection efficiency, which make it difficult to meet the high standard requirements of the modern hatching eggs detection and classification industry. Therefore, companies need a new way to replace work to reduce costs and improve the quality of products.
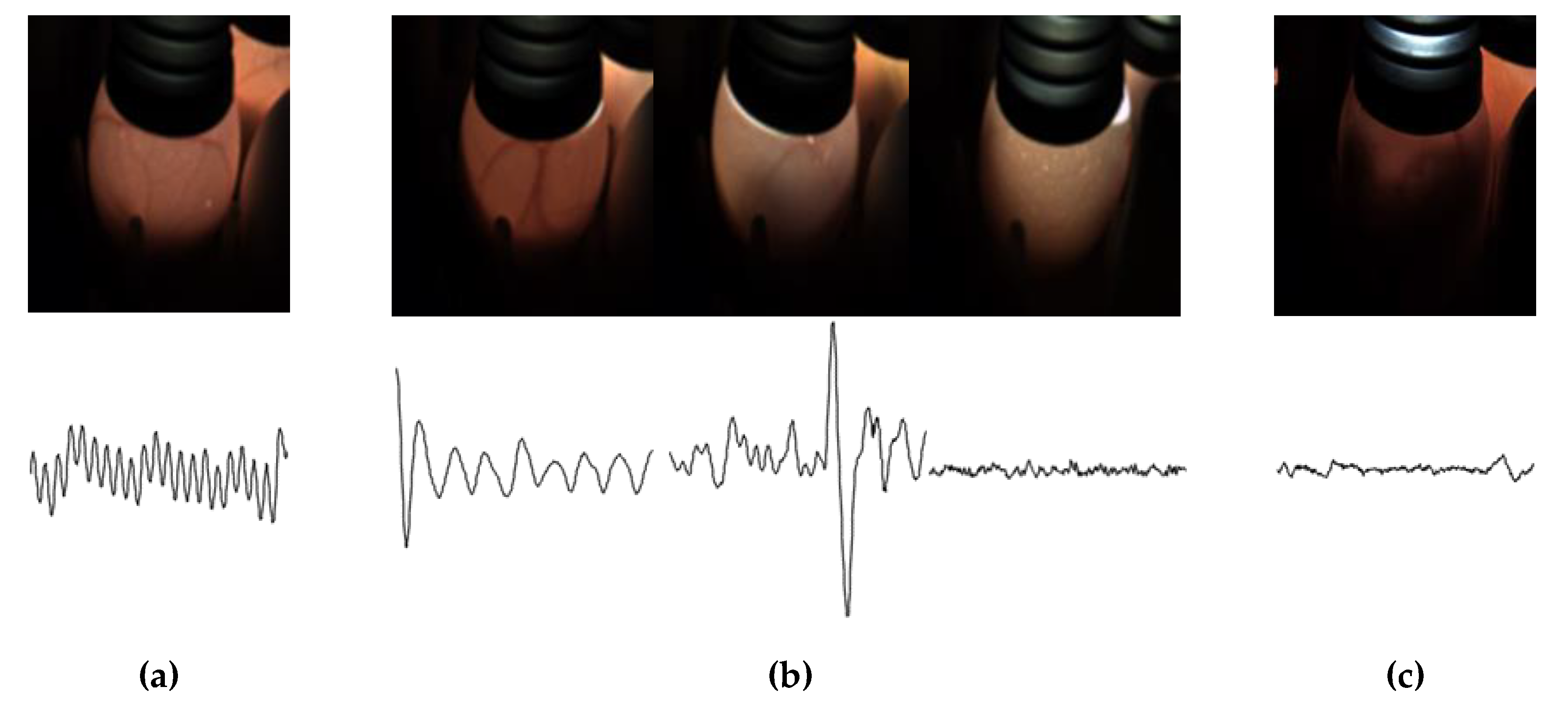
The detection of hatching eggs is usually divided into four periods: 5 days, 9 days, 14 days, and 16 days. Hatching eggs have different blood-vessel features and heartbeat features during the different periods. As such, there are different classification standards in different periods. In particular, because 16 days is the final period of hatching eggs, detection is more rigorous. The 16-day embryos are divided into three categories: Fertile embryos, recovered embryos, and waste embryos. Fertile embryos were used to extract the vaccine. The recovered embryos were recycled for further processing, and the qualified embryos were selected for vaccine extraction. Waste embryos were treated harmlessly. As illustrated in
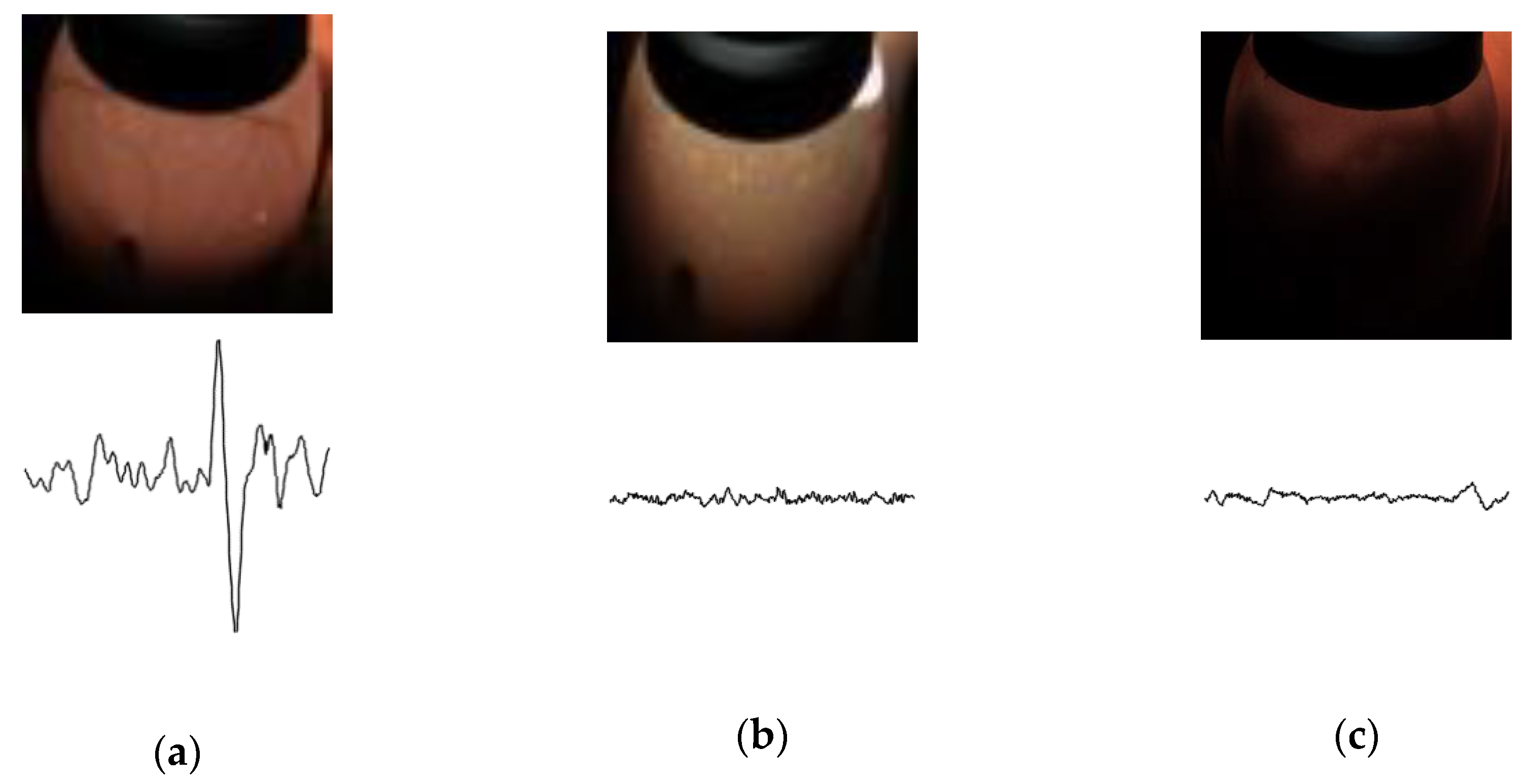
Figure 1, the fertile eggs have regular heartbeats and strong blood vessels. Recovered eggs have three sets of characteristics. The first set is that hatching eggs have slow heartbeats and strong blood vessels. The second kind of hatching eggs have irregular heartbeats and the blood vessels that begin to constrict. In the third set, egg embryos have no heartbeats and the blood vessels begin to constrict and may even disappear completely. The waste eggs have no heartbeat. All blood vessels disappear completely, and the insides of the eggs begin to rot. Approximately 10% of the recovered eggs have the same blood vessel characteristics as fertile eggs. In addition, 50% of the recovered eggs have the same heartbeat signal characteristics as waste eggs. Because different categories may have the same heartbeat or image features, they are difficult to classify with a single heartbeat or image signal. As such, it is of great significance to improve the technical level of classifying 16-day hatching eggs.
In recent years, people have been exploring new methods to classify hatching eggs, such as machine vision technology, hyperspectral imaging technology, and multi-information fusion technology. In 2010, Shan et al. [
1] introduced a method to detect the fertility of middle-stage hatching eggs. They used image processing to enhance the picture and obtain the major embryo blood vessels of the hatching egg. Then, they used the weighted fuzzy c-means clustering algorithm to obtain a threshold to detect the fertility. In 2005, Lawrence et al. [
2] first used hyperspectral images to detect the development of egg embryos. They designed a hyperspectral imaging system to detect the development of brown- and white-shelled eggs. The detection accuracy was 91% for white-shelled eggs and 83% for brown-shelled eggs. In 2014, Liu et al. [
3] proposed a method for detecting infertile eggs using near-infrared hyperspectral imaging. They segmented the region of interest (ROI) of each hyperspectral image and extracted information in the hyperspectral images using the Gabor filter. They used principal component analysis (PCA) to reduce the dimensionality of the spectral transmission characteristics. The final classification accuracy rates were 78.8% on the first day, 74.1% on the second day, 81.8% on the third day, and 84.1% on the fourth day. In 2014, Xu et al. [
4] designed a non-destructive method for detecting the fertility of eggs prior to virus cultivation. Due to the high transmission through the holes in the eggshell, they used a method based on the smallest univalue segment assimilating nucleus to distinguish high-brightness speckle noise pixels in egg images. Additionally, they used the smallest univalve segment assimilating nucleus (SUSAN) principle to detect speckle noise. Then, the blood vessels were restored, and binarized images of the main blood vessels were obtained. By calculating the percentage of the image that the blood vessel area occupies in the ROI image, fertility was evaluated. The final classification accuracy rate was 97.78%.
With the development of deep learning, convolutional neural networks (CNNs) show good performance in solving classification problems. CNNs such as Alexnet [
5], GoogLeNet [
6], and ResNet [
7] are widely used in image classification. In 2018, Geng et al. [
8] designed a method for detecting 5-day infertile eggs using a CNN and images of hatching eggs. In 2019, Geng et al. [
9] designed a method for detecting 9-day infertile eggs using a CNN and heartbeat signal. Huang [
10] designed a CNN architecture in a small images dataset to classify 5- to 7-day embryos, but 5-day to 9-day embryos have no recovered eggs, and there is no overlap between the characteristics of different categories, so using a single heartbeat signal or images of hatching eggs can achieve good results. Therefore, these three CNN methods have achieved good results.
Now, recurrent neural networks (RNNs) are also widely used in the field of processing sequences, such as speech recognition [
11]. More and more researchers are combining CNN with RNN to solve new problems. In reference [
12], they use CNN-long short-term memory (LSTM) for non-invasive behavior analysis.
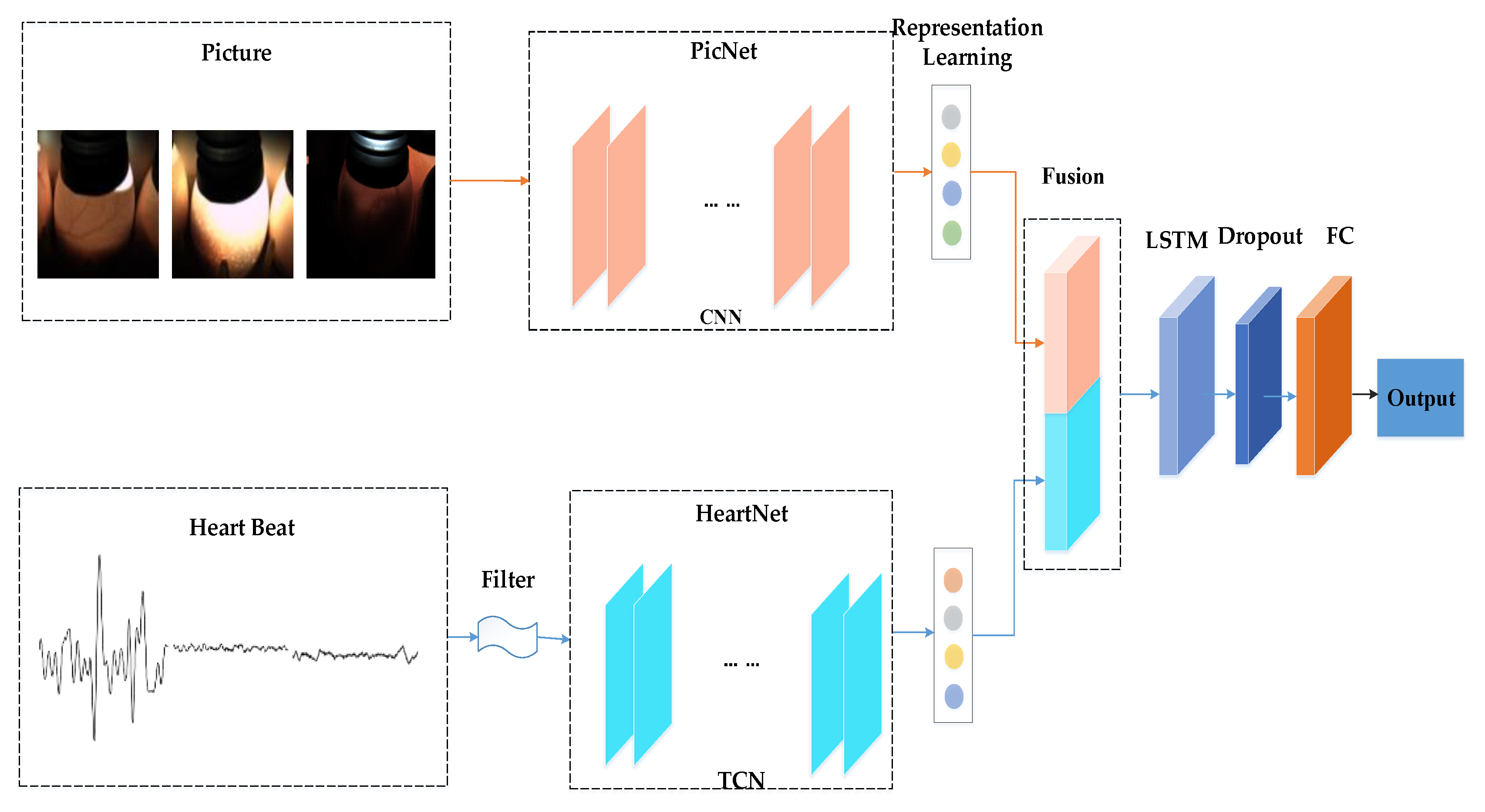
The 16-day hatching eggs are divided into three categories. Since different kinds of eggs may have the same heartbeat signal or blood-vessel features, it is not possible to judge the waste eggs and the recovered eggs by the heartbeat signal only, and the embryo image signals cannot be used in isolation to determine the fertile eggs and recovered eggs. As such, we propose an end-to-end, multimodal hatching eggs classification method. Our main contributions are listed as follows:
In order to solve the problem of different categories possibly having the same image or heartbeat characteristics, this paper designed a network structure that can simultaneously use the time series heartbeat signals and the egg embryo images.
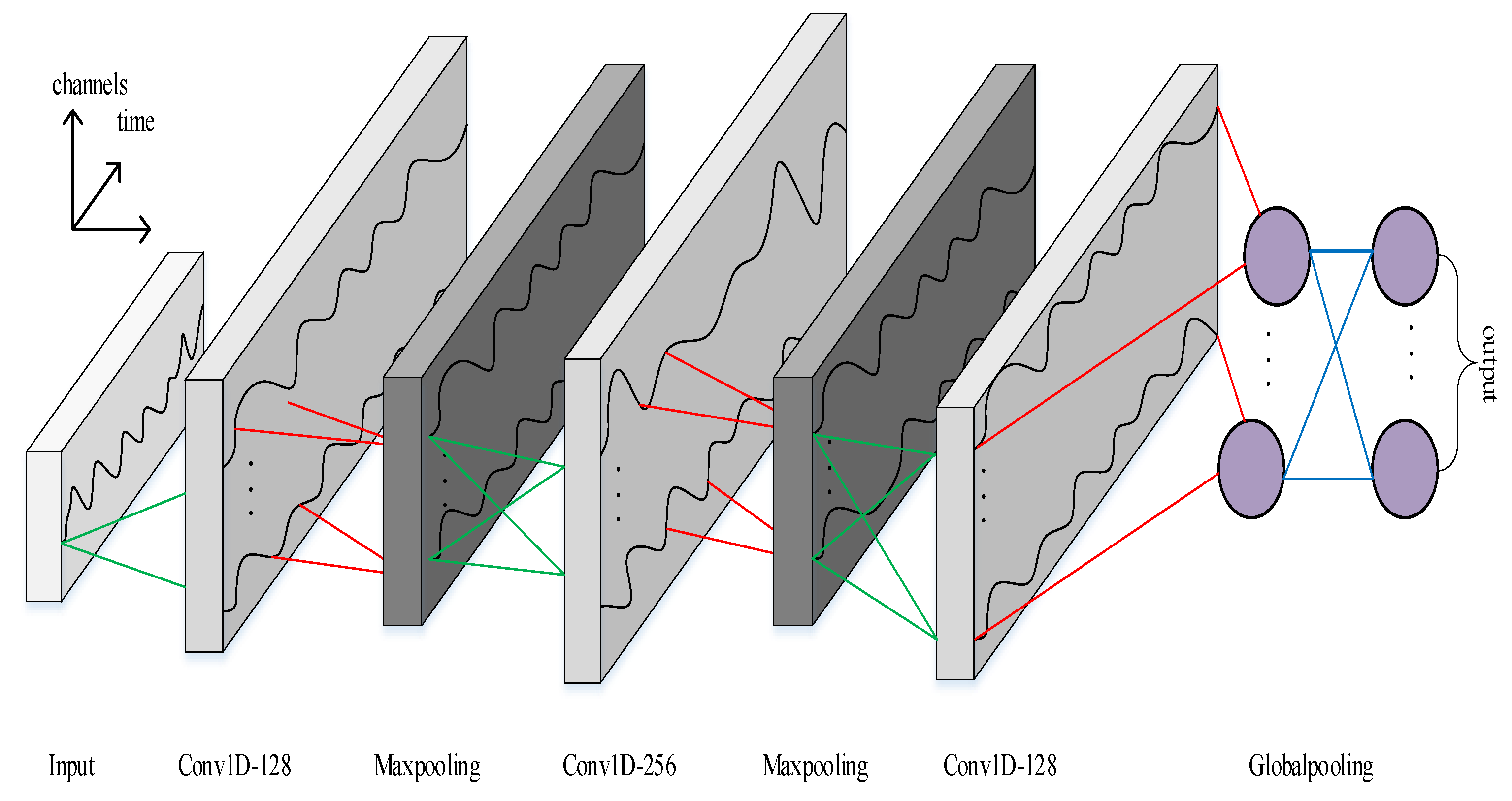
In order to solve time-series classification problems, this paper designed a six layer-deep temporal convolutional network (TCN) architecture that can model the heartbeat signal.
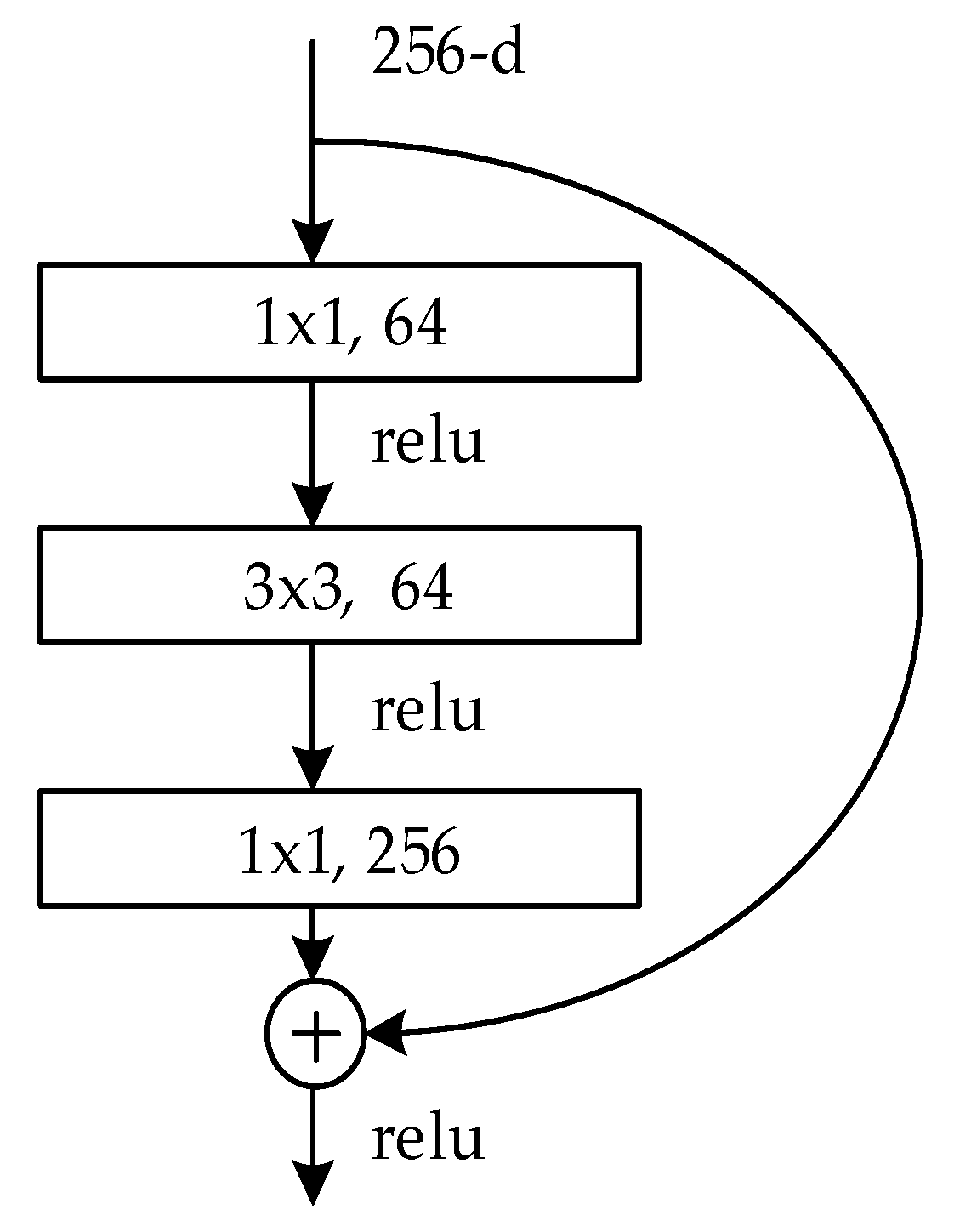
We used a pre-training ResNet model to shorten the training time and create a more accurate image classification model.
3. Experiments and Results
In this section, we compare our multimodal classification method with a single-mode classification method based on our dataset. Additionally, we evaluate previous methods and the method proposed herein. To evaluate the performance of different methods, we use micro-averaged recall score, micro-averaged precision score and micro-averaged F1 score, which are defined as follows,
where
(true positives) is the number of eggs correctly classified into category
;
is the total number of instances;
is the number of categories,
where
(false positives) is the number of eggs that do not belong to class
but are misclassified to class
;
(true negatives) is the number of eggs that do not belong to class
and not classified to class
;
(false negatives) is the number of eggs that belong to class
but were misclassified.
3.1. Dataset
To capture image data, we used a color industrial camera with an 8 mm lens to take pictures of hatching eggs. We used lamps with adjustable brightness to provide a light source and covered the tops of the eggs with a rubber sleeve to prevent light leakage. The size of the original image was 1280 × 960 pixels. We used the photoplethysmography (PPG) technique to acquire the corresponding heartbeat signal. PPG can be used to detect blood volume changes in a microvascular bed of tissue [
19]. Because the volume of blood in the blood vessels of egg embryos changes with the heart activity cycle, the light intensity absorbed by the vessels changes synchronously with the beating of the heart. As such, the A/D module can convert light that passes through the tissue into an electrical signal. The signal acquisition equipment is shown in
Figure 5. The hatching egg is placed between a laser and a receiving terminal module, which receives light that passes through the egg and converts light into an electrical signal. Finally, the PPG signal is transferred to the microcontroller. The PPG signal is a sequence of 500 data points and the sampling rate is 62.5 Hz.
Because the background area of the original image was too large, we extracted the region of interest (ROI) to make the embryonic characteristics more obvious. We binarized the image to highlight the outline of the of the egg embryo. For different types of embryos, we used different gray values as thresholds. Then, the maximum contour of the binary image was extracted as the boundary of the ROI region. Finally, all the processed images were scaled to 224 × 224 pixels to fit the required input size of ResNet-50. We designed a second order Butterworth high-pass filter to denoise the heartbeat data and take the last 350 filtered points as the sampling points. The processed egg embryo pictures and corresponding heartbeat signals are shown in
Figure 6.
The dataset in this study has a total of 7128 egg embryo images, named the egg picture dataset. Each picture corresponds to a heartbeat signal, and these heartbeat signals are called the heartbeat dataset. In this dataset, there are 2088 samples of fertile eggs, 2160 samples of waste eggs, and 2880 recovered egg samples. The number of embryos in each category is roughly the same, ensuring the balance of the data. All datasets are divided into training sets, validation sets, and testing sets.
Table 2 contains more details for each portion of our dataset.
3.2. Unimodal Training
We trained PicNet and HeartNet separately on our dataset and compared them to other network structures. The results are as follows.
3.2.1. PicNet Training
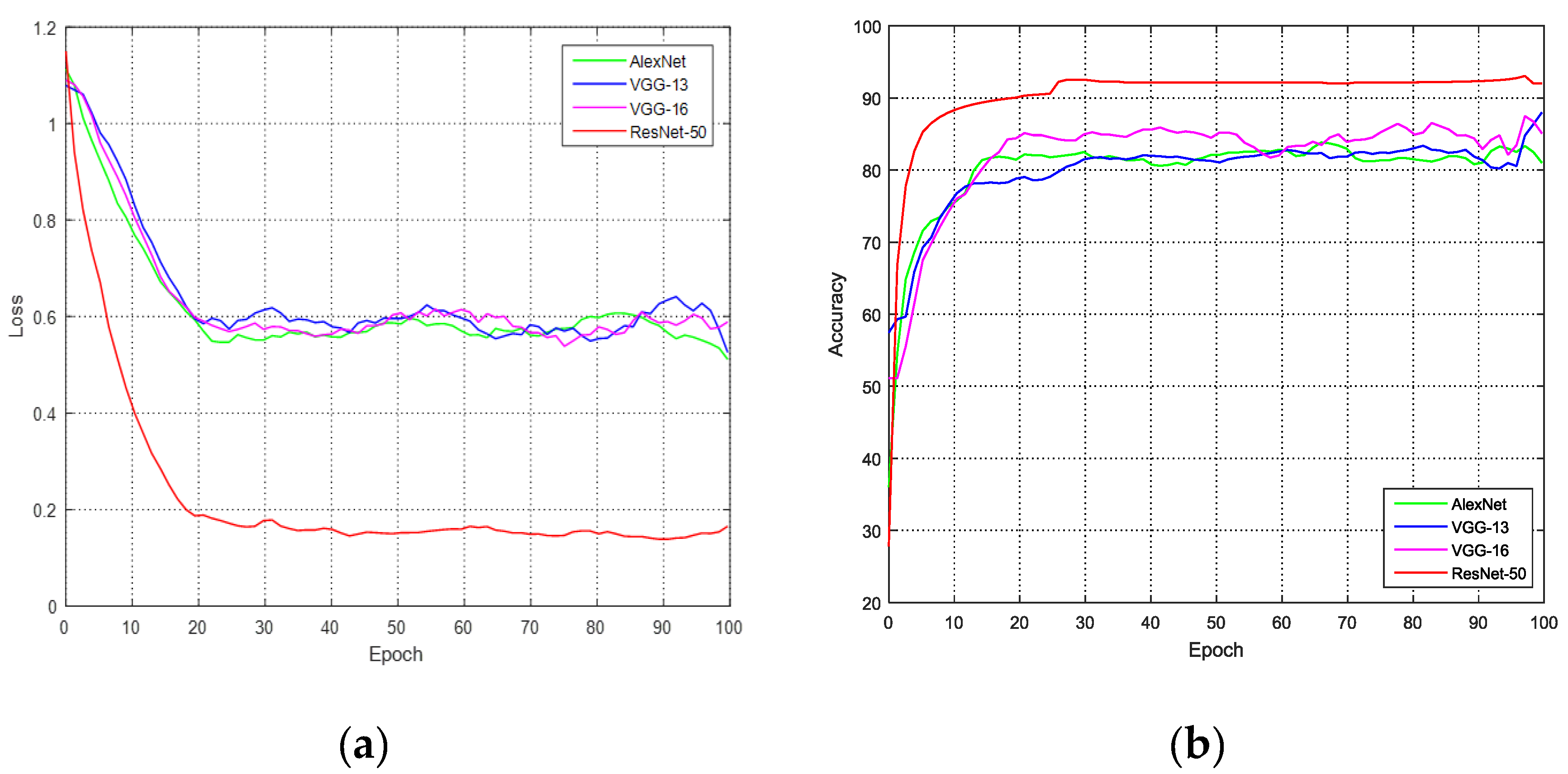
We compared existing CNNs on the hatching egg picture dataset. The model was trained for, at most, 100 epochs. The batch size was 32. With eight NVIDIA GTX 1080 Ti GPUs, it took approximately 2 minutes for one epoch. We used the cross-entropy loss function to compute the loss of the PicNet. The varying curves of loss and accuracy are shown in
Figure 7.
Table 3 contains the accuracies of different CNNs.
Because our egg picture dataset has three types, and approximately 10 percent of the recovered eggs have the same blood vessel characteristics as fertile eggs, the accuracy of using only the picture signal is not high. The best CNN is ResNet-50, which has an accuracy of 90.92%. Based on the results, we used ResNet-50 as the picture network.
3.2.2. HeartNet Training
We studied the effects of different filter sizes
used by each layer of our TCN architecture. We used the cross-entropy loss function to compute the loss of the HeartNet. We performed a series of controlled experiments on the egg heart dataset, the results of which are shown in
Table 4. The experimental results show that the TCN model performs best when filter size
, so our model’s 1D convolution kernel size is 5.
We also compared canonical recurrent neural network architectures, such as LSTM and gated recurrent unit (GRU) [
20], with the TCN architecture based on our egg heart dataset. To compare all three architectures fairly, the LSTM and GRU architectures have up to six layers so that each model has approximately the same number of parameters, and the optimizers are chosen from adaptive moment estimation (Adam) [
21], stochastic gradient descent (SGD) [
22], and adaptive gradient algorithm (Adagrad) [
23]. The details of the LSTM and GRU architectures are given in
Table 5 and
Table 6.
All models were trained for, at most, 100 epochs. The batch size was 32. With eight NVIDIA GTX 1080Ti GPUs, it took approximately 1 minute for one epoch.
Table 7 contains the accuracies of different networks.
The experimental results show that our TCN architecture performs better than other RNN architectures such as LSTM and GRU. As such, we use our TCN architecture as the HeartNet architecture.
Because our egg heart dataset has three types, and approximately 50% of the recovered eggs have the same heartbeat signal characteristics as fertile eggs, the accuracy of using only the heartbeat signal is low.
3.3. Multimodal Training
We trained the multimodal network and compared it to HeartNet and PicNet. The optimization method we used to train our model is the Adam optimizer. The fixed learning rate is 10
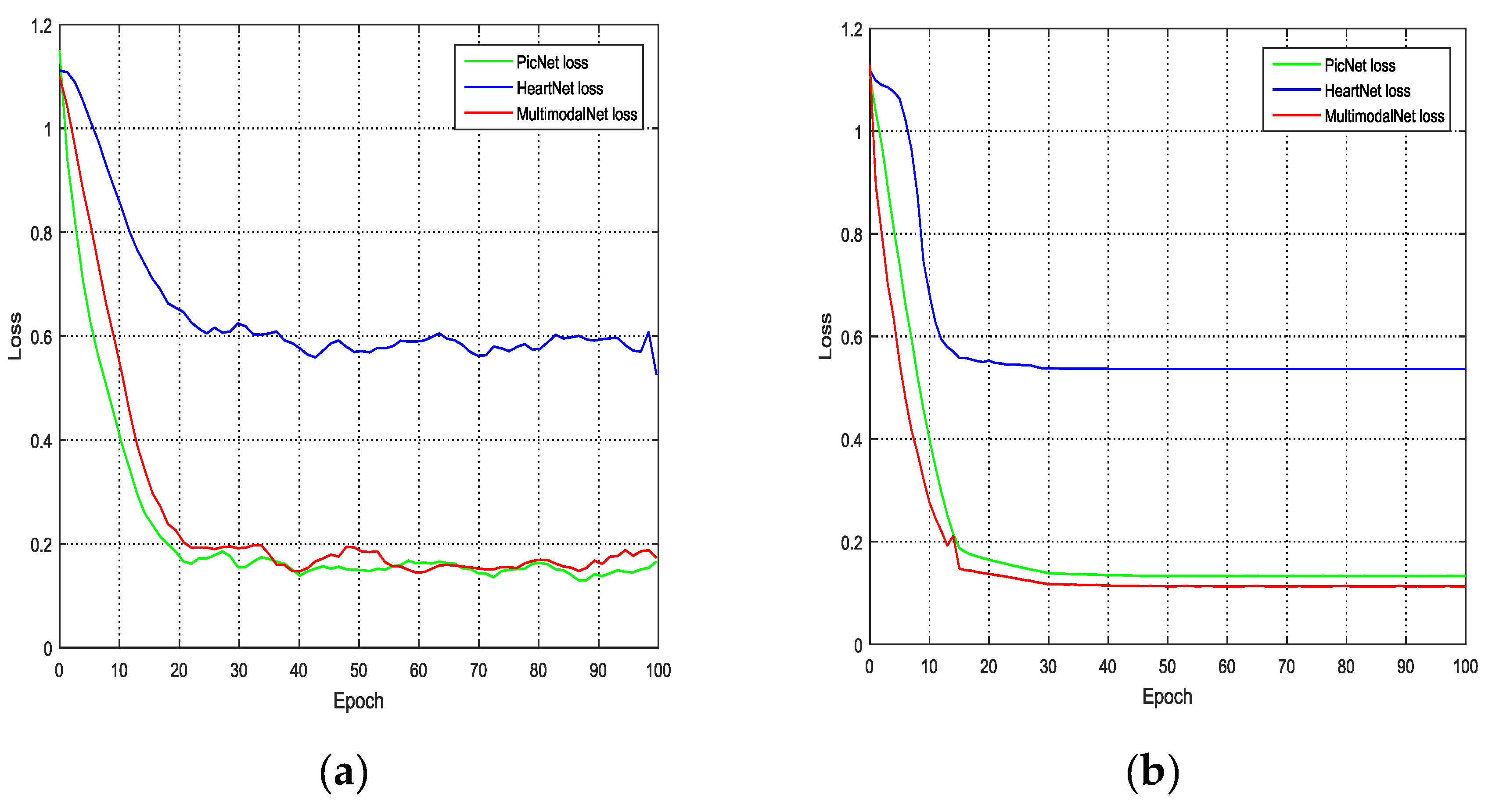
−4, the decay rate is 0.9, and the momentum is 0.1. The batch size is 32. With eight NVIDIA GTX 1080 Ti GPUs, it took approximately 3 minutes for one epoch. The loss curve of the training process is shown in
Figure 8.
For the training dataset, the loss values of PicNet are slightly lower than those of MultimodalNet. As such, PicNet showed a slightly better performance than MultimodalNet on the training dataset, but for the validation dataset, MultimodalNet had the best performance. Therefore, our proposed method provided the lowest loss among all methods on the validation dataset.
3.4. Results Evaluation
To verify the feasibility of the network proposed in this paper, we compared the accuracy of single modal networks and the multimodal network. The results are shown in
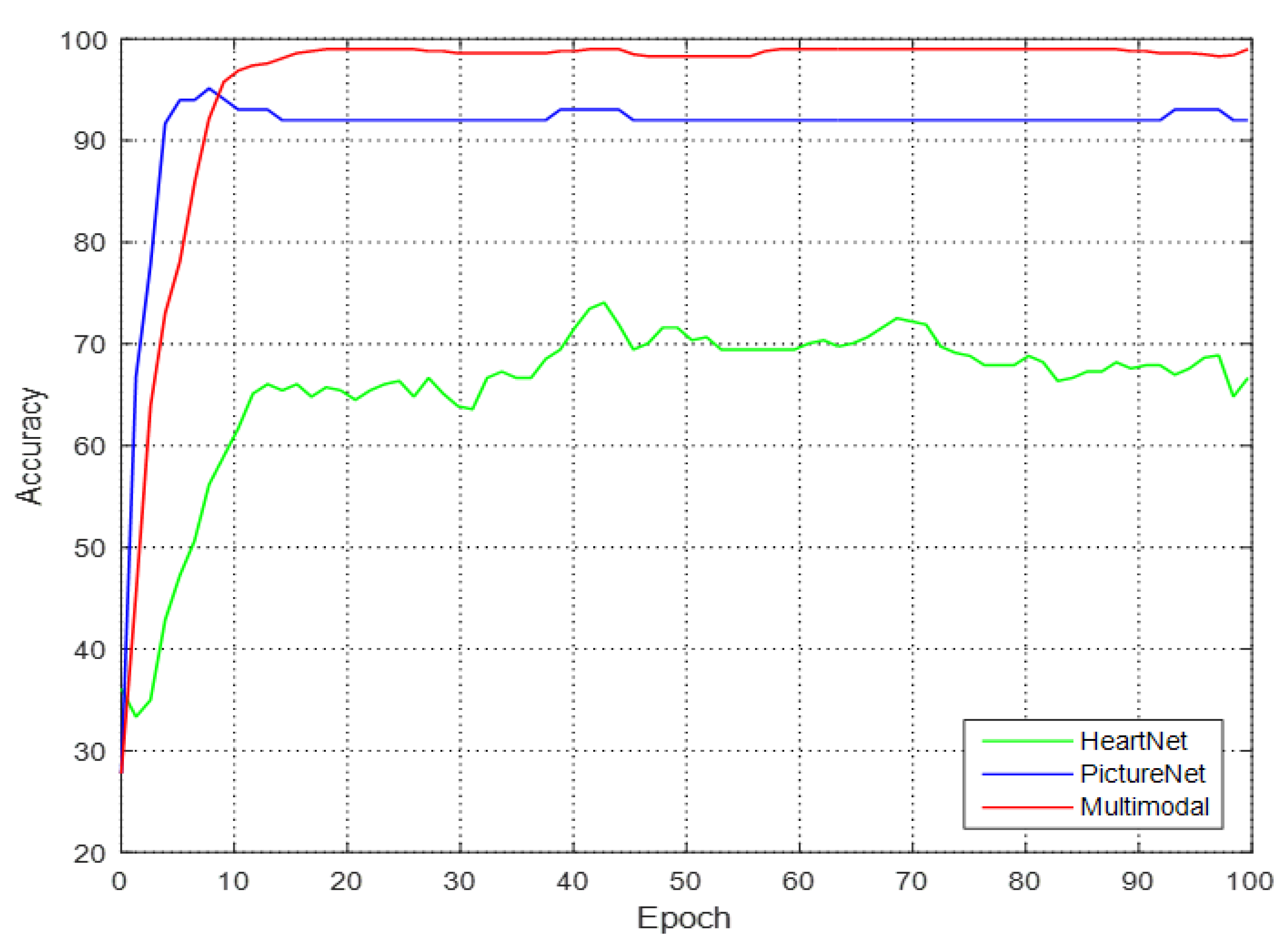
Table 8 and the accuracy curve is shown in
Figure 9.
From
Table 8, it is apparent that HeartNet has the lowest accuracy because most of the waste embryos and recovered embryos have no heartbeats, and a small portion of the recovered embryos have heartbeats. Therefore, it is difficult to distinguish the waste embryos and recovered embryos by relying only on the heartbeat signal. Therefore, the accuracy of using HeartNet is very low. There is a small number of embryos with blood vessels but abnormal heartbeats in the recovered embryos, so the use of PicNet led to the inaccurate classification of recovered embryos and fertile embryos. Only by using both signals at the same time can the three types of embryo be correctly classified.
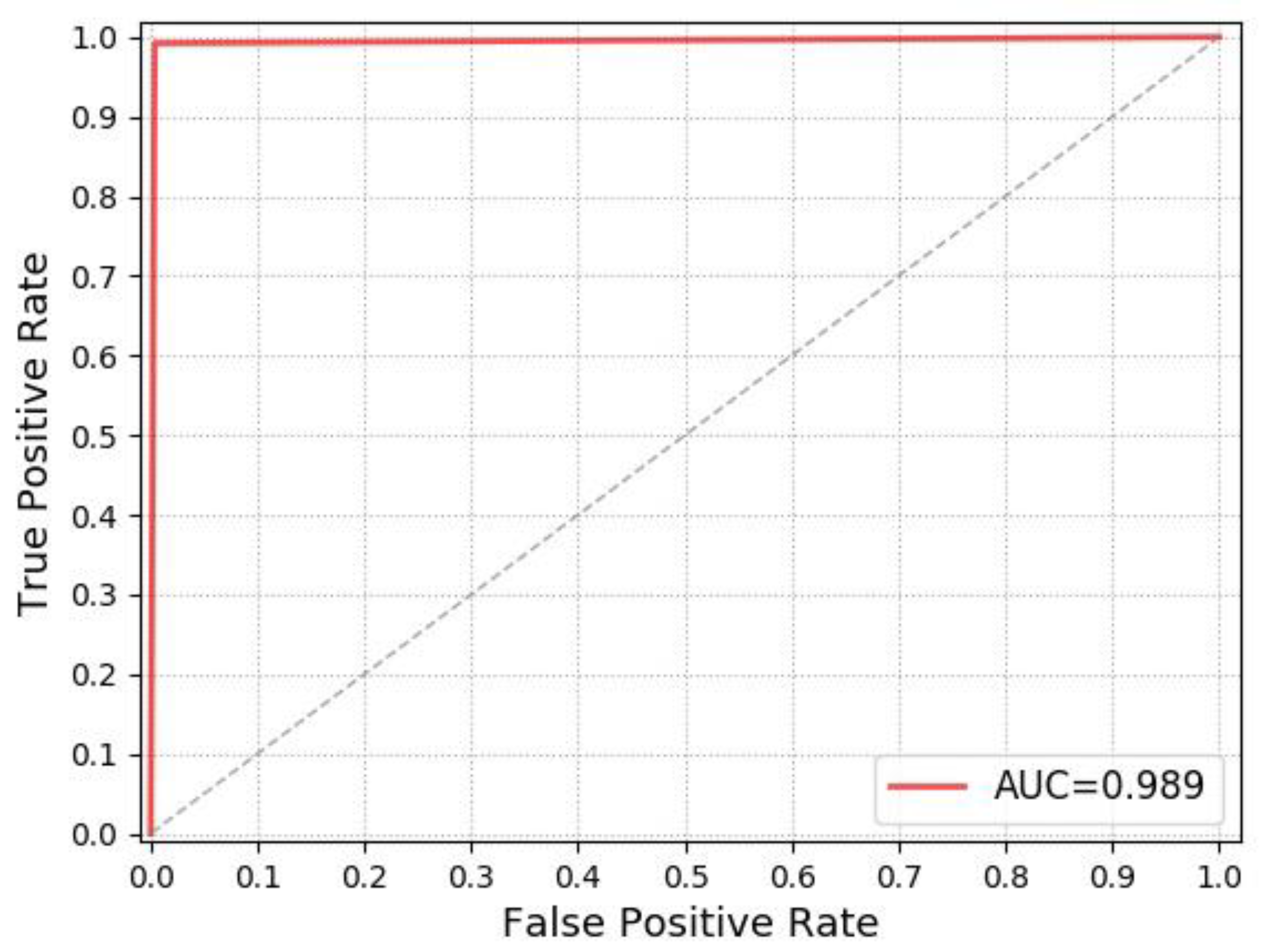
Receiver operating characteristic (ROC) curve can illustrate the diagnostic ability of a classifier system. The larger the area under the ROC curve (AUC), the better the classifier performance, so we also use the ROC chart to illustrate the performance of our model. The ROC chart is shown in
Figure 10.
As can be seen from
Figure 10, the AUC indicator of our model is 0.989, which indicates that the performance of our model is outstanding for the 16-day hatching eggs Classification.
We tested our model on the testing sets. The confusion matrix is shown in
Figure 11. As can be seen from
Figure 11, two of 417 fertile hatching eggs were classified as recovered embryos, five of 432 waste embryos were classified as recovered embryos, and five recovered embryos were also misclassified. A total of 12 embryos were misclassified. The accuracy on the testing sets reached 99.15%.
The proposed multimodal network structure inputs two modalities of data at the same time, respectively processes the heartbeat signal and embryo image, and finally fuses them together. The method we proposed can achieve a higher accuracy rate than using a single type of signal.
4. Conclusions
In this paper, we propose an end-to-end multimodal hatching eggs classification method. We designed a deep learning network that includes a picture processing network and a heartbeat signal processing network. We fed both the heartbeat signals and the egg embryo images into our deep learning network, which overcame the problems that only using heartbeat signals cannot correctly distinguish recovered embryos from waste embryos and that using single-mode embryo images cannot correctly distinguish recovery embryos from fertile embryos. Based on the results of our experiments, the accuracy reached 98.98%. Our method has obvious advantages over other methods that use single modal signals. Additionally, the results show that the proposed method is more suitable for multi-classification of egg embryos.
Our method can replace workers in production and maintain stable operation. This method is not only suitable for hatching eggs classification but also suitable for other aspects. For example, in the fields of face recognition and emotion recognition, video, audio, and other forms of signals can be used for recognition at the same time. In the medical field, we can also combine electrocardiogram and CT images and other signals to improve the accuracy of recognition. Therefore, the method we proposed is very meaningful.
In future work, we will expand our dataset in terms of both the categories of embryos and the amount of experimental data. In addition, we will add more modalities and continue to optimize the network structure to improve its accuracy.