Multilevel and Multiscale Deep Neural Network for Retinal Blood Vessel Segmentation
Abstract
1. Introduction
- A better pre-processing approach that highlights the blood vessels;
- Incorporation of multilevel and multiscale deep supervision (DS) networks that can dive deep into the final layers of the four convolutional layers with two different scale initializations i.e., 0.001 and 0.0002;
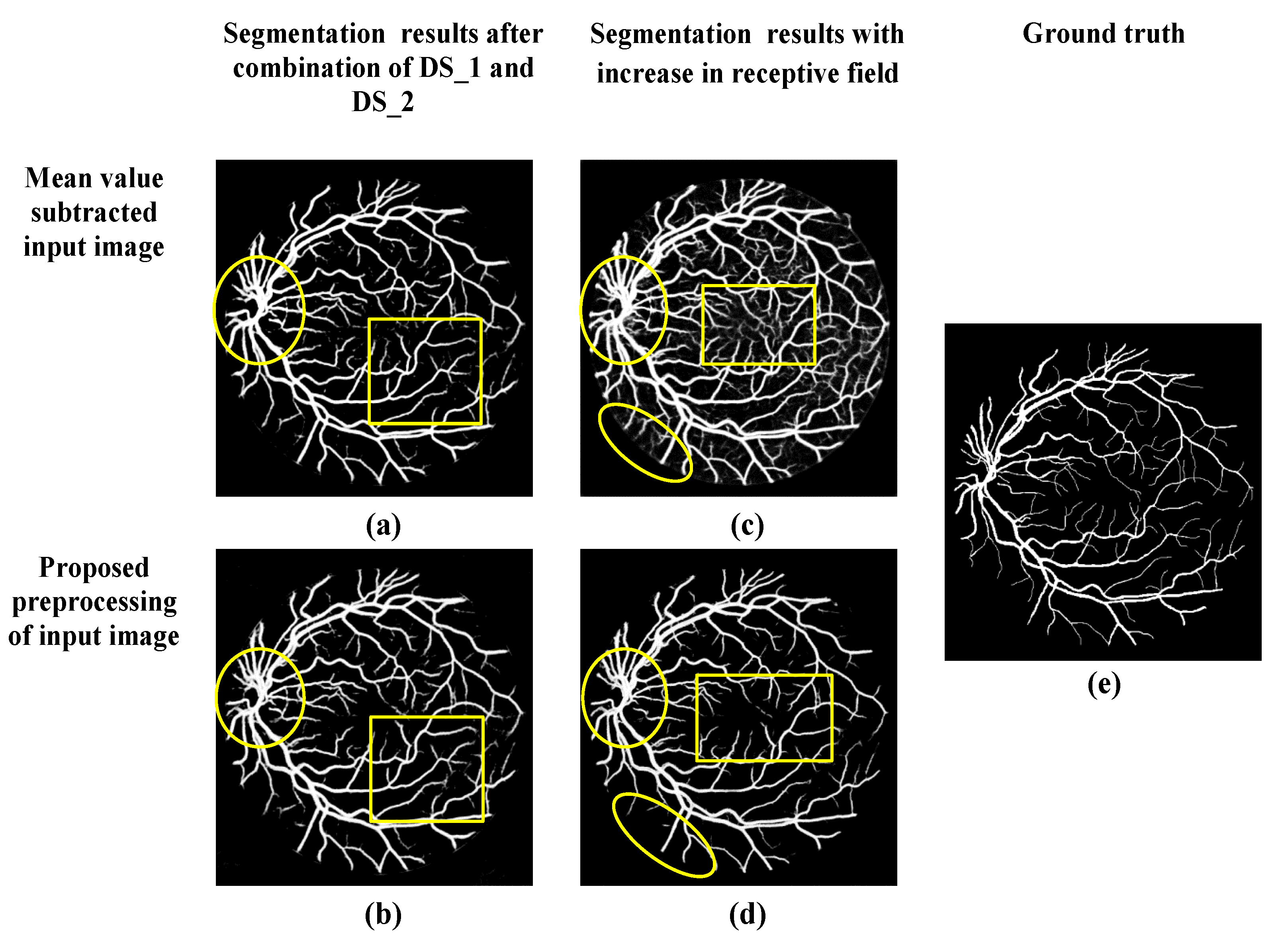
- Furthermore, the receptive field of this multilevel and multiscale deep supervision (DS) network is increased to refine and localize the blood vessels. Therefore, the probability map obtained consists clearly of blood vessels with fewer false predictions.
2. Materials and Methods
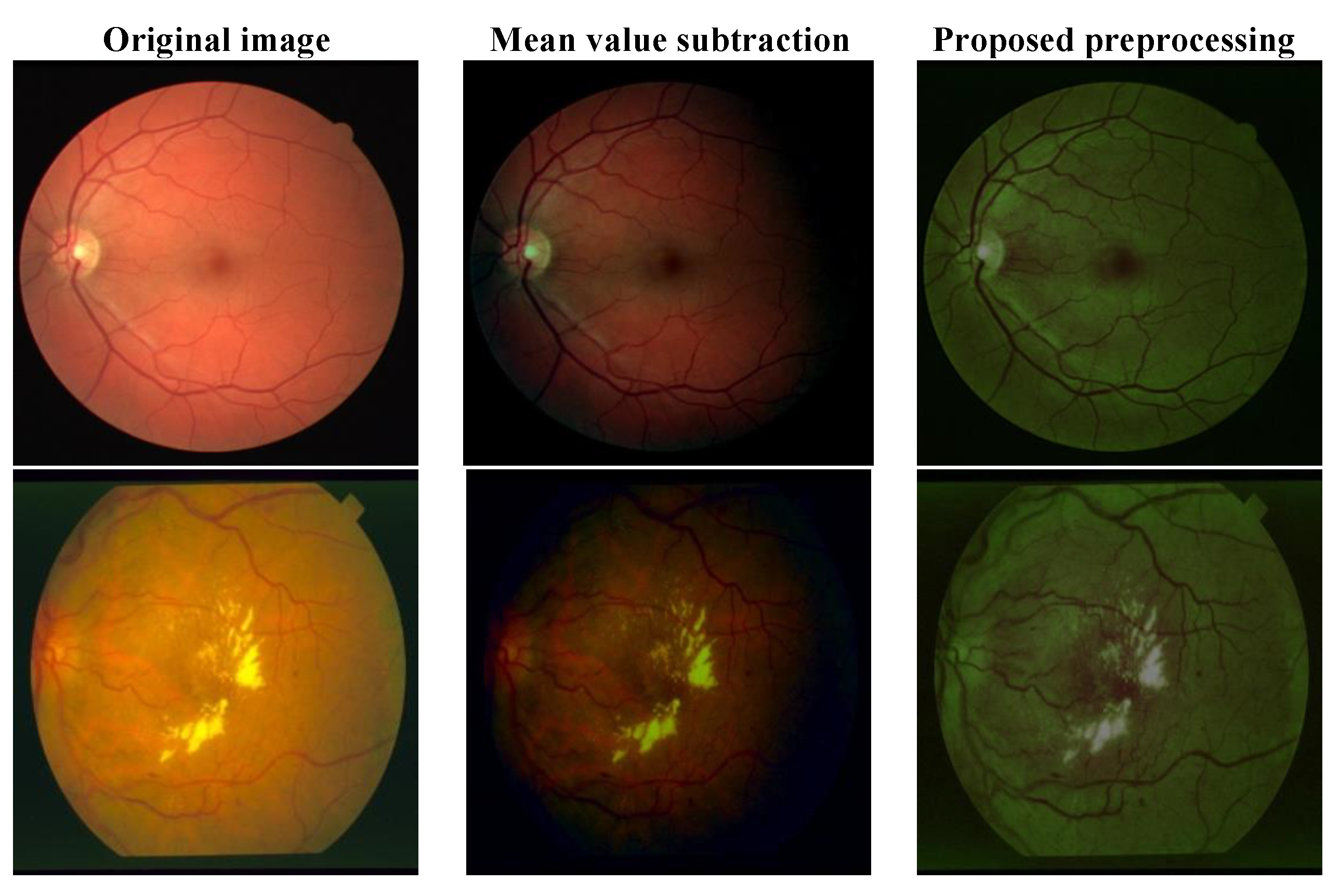
2.1. Proposed Preprocessing
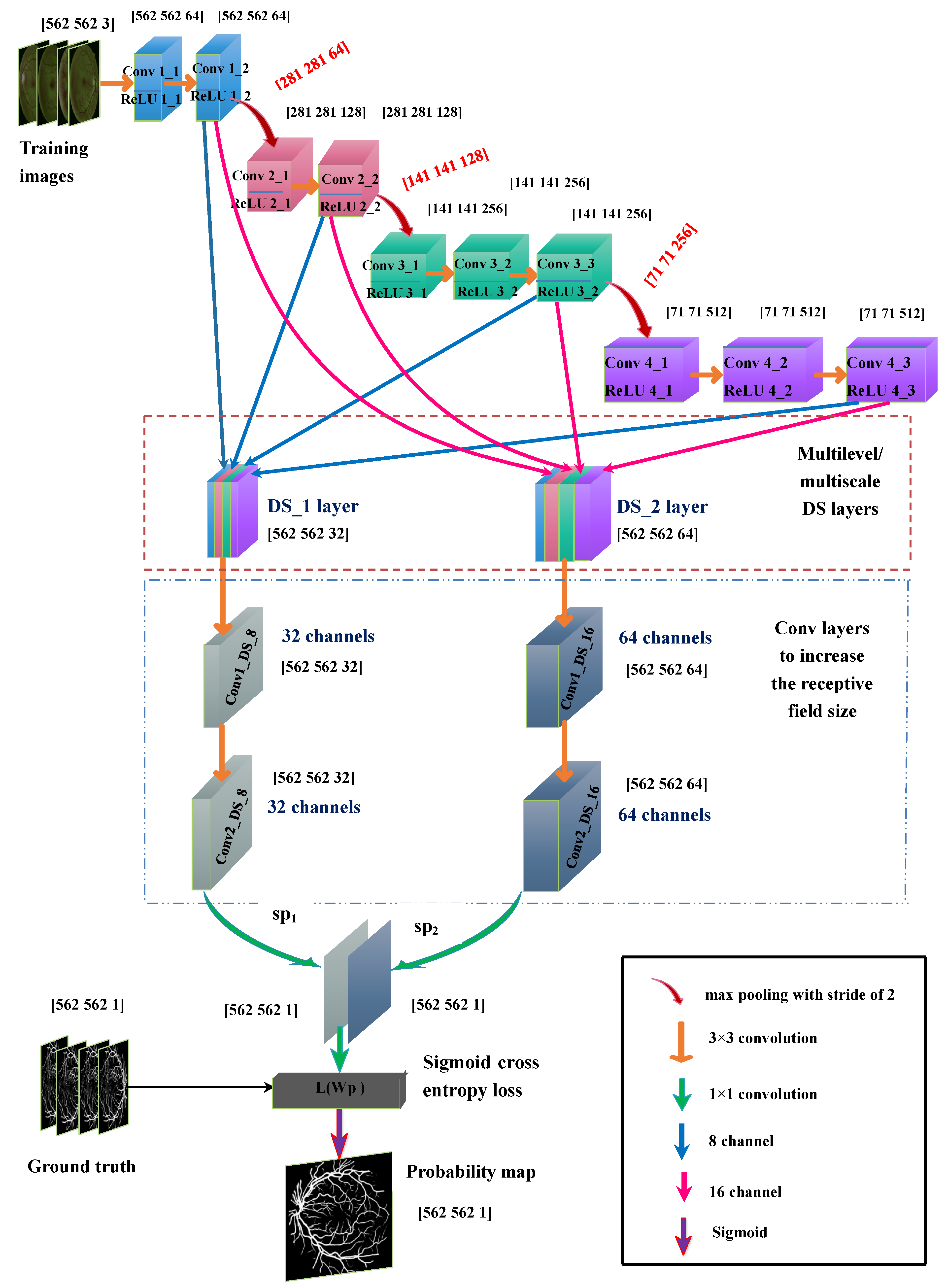
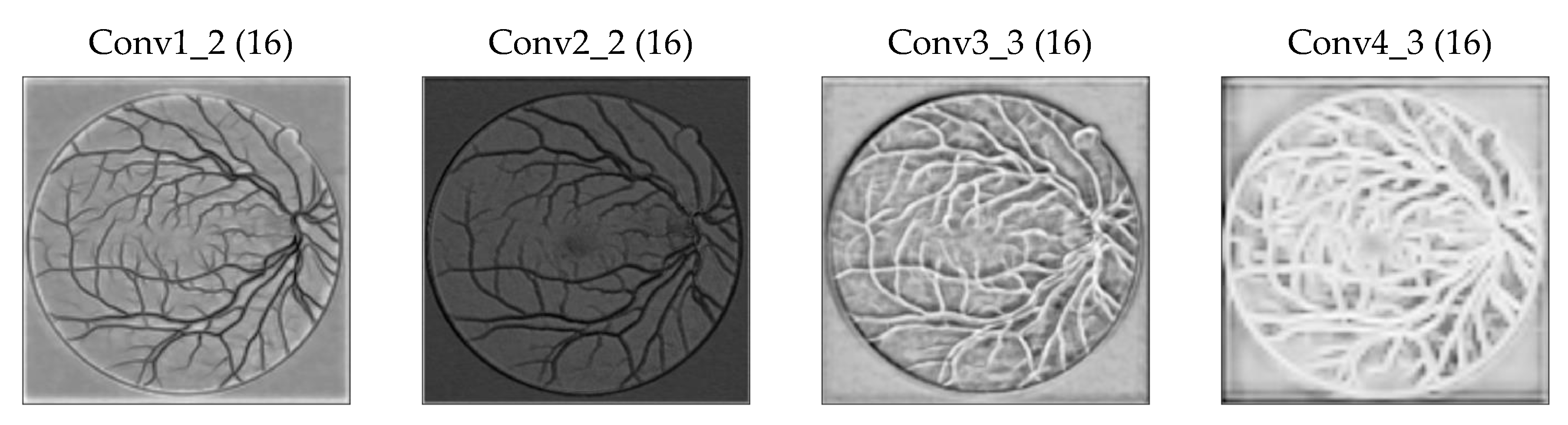
2.2. Proposed Multilevel/Multiscale Deep Neural Network (DNN)
2.2.1. Base Network
2.2.2. Deep Supervision (DS_1) Layer
2.2.3. Deep Supervision DS_2 Layer
- DS_1 layer parameters = (3 × 3 × 32 + 1) × 32 = 9248
- DS_2 layer parameters = (3 × 3 × 64 + 1) × 64 = 36,928
2.2.4. Increase in the Receptive Field of View of DS Layers
2.3. Input Image Augmentation
- Preprocessing the image using the method described in Section 2.1;
- Rotation of the image to 15 different angles;
- Flipping every rotated image;
- Cropping the region of interest in the rotated and flipped images;
- Scaling the rotated and flipped input image to 0.5 and 1.5, respectively.
2.4. Loss Function and Optimization
3. Results
3.1. Training
3.2. Testing
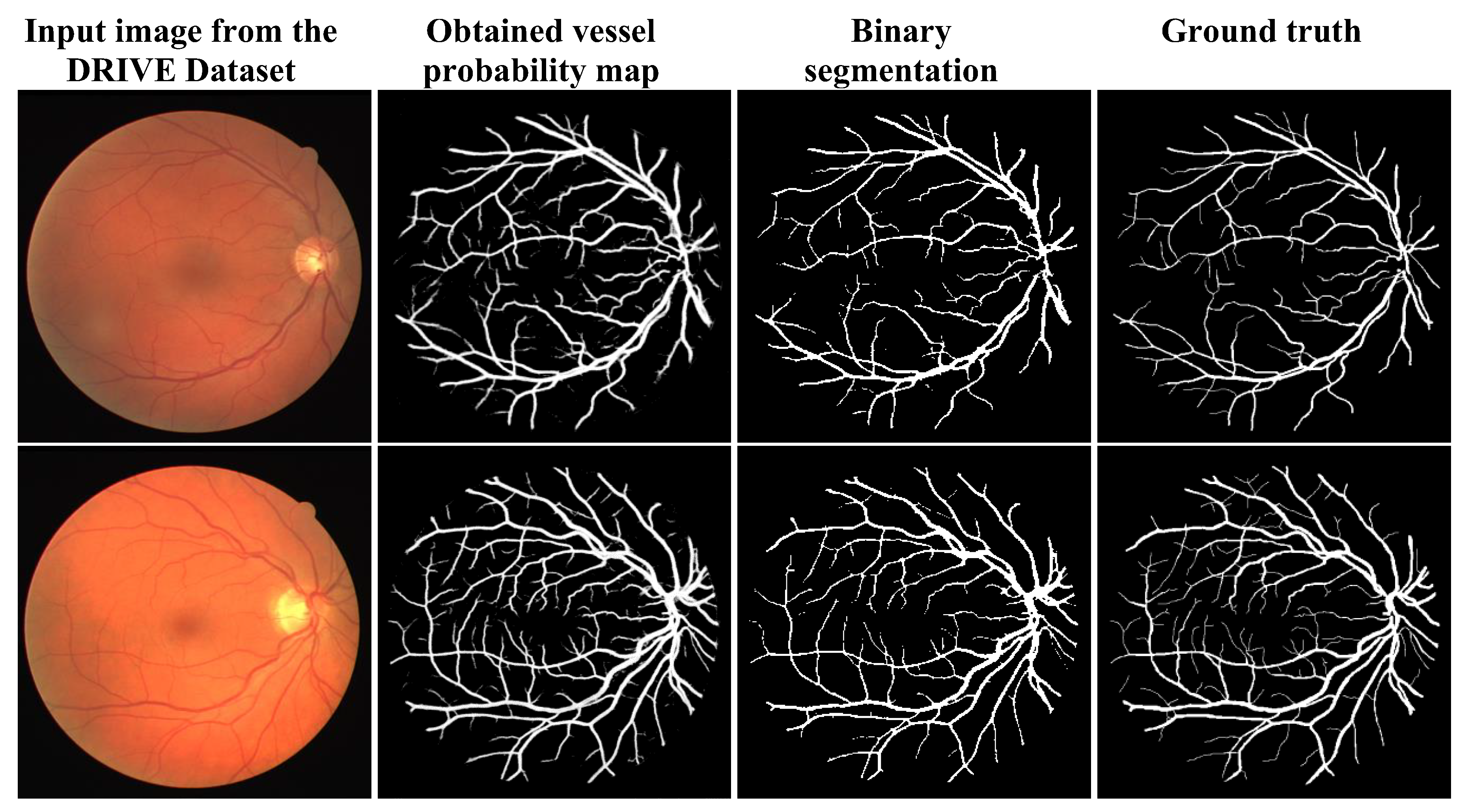
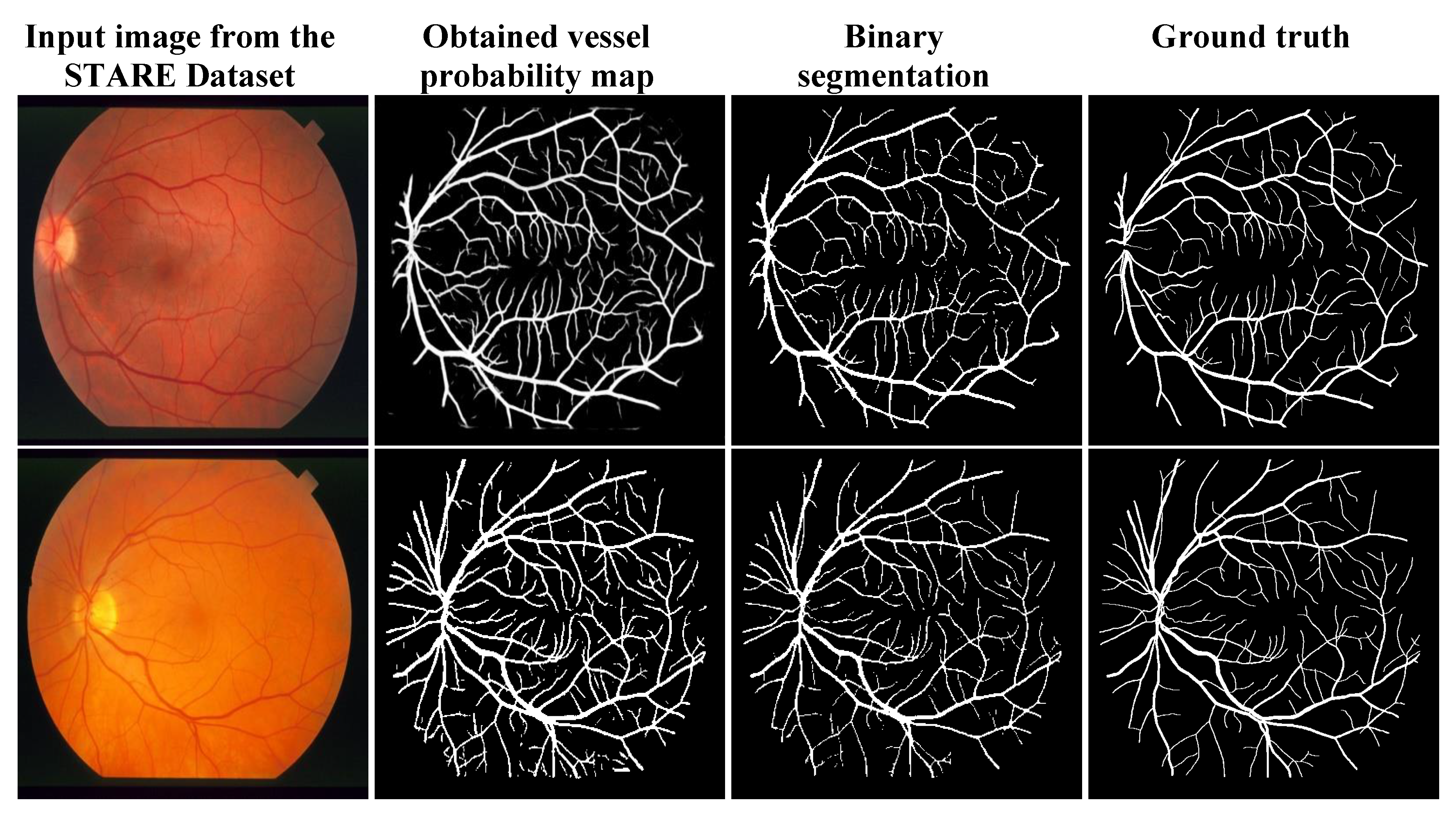
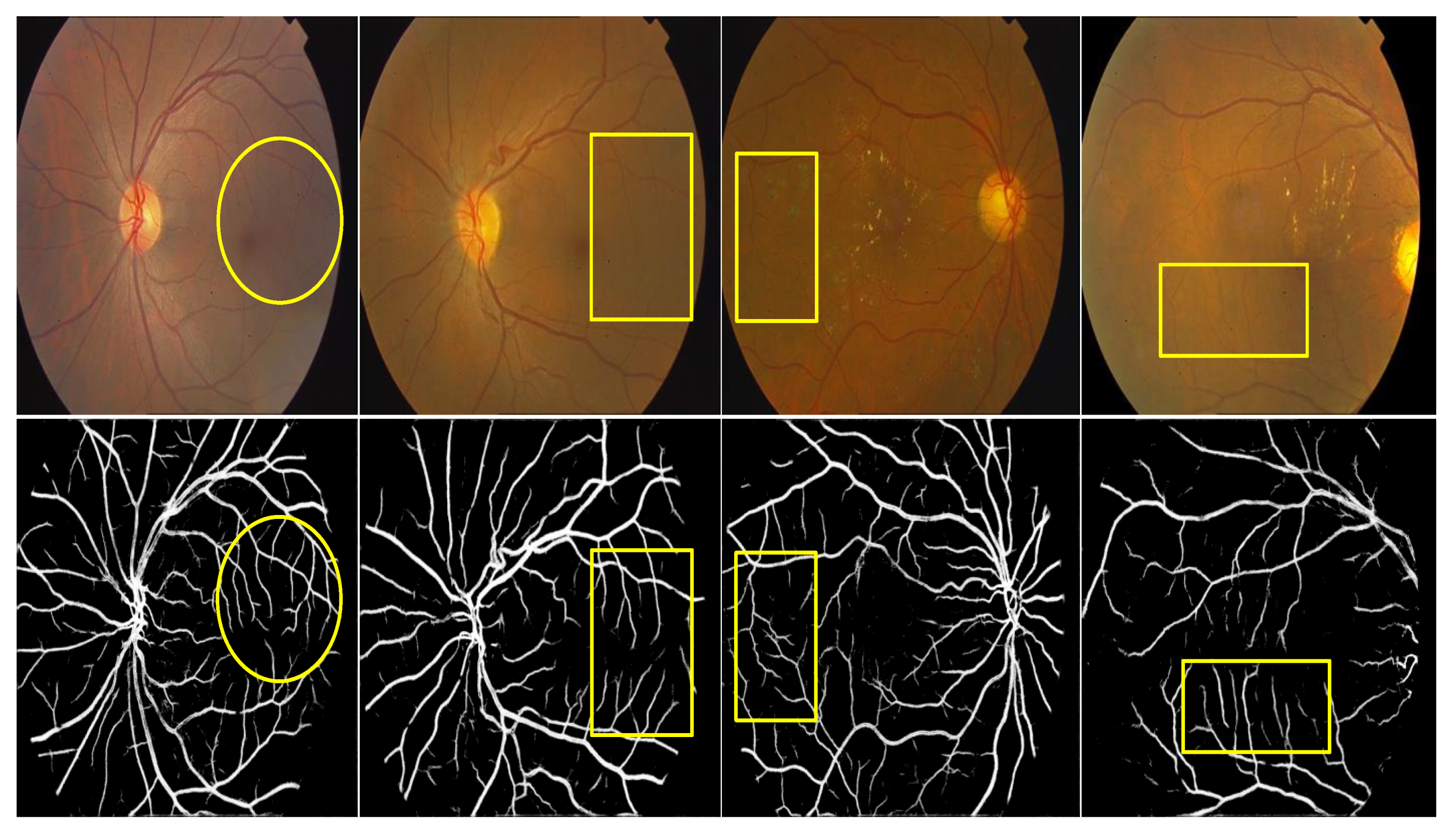
3.3. Qualitative Analysis
3.4. Quantitative Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Taylor, H.R.; Keeffe, J.E. World blindness: A 21st century perspective. Br. J. Ophthalmol. 2001, 85, 261–266. [Google Scholar] [CrossRef] [PubMed]
- Mateen, M.; Wen, J.; Nasrullah; Song, S.; Huang, Z. Fundus image classification using VGG-19 architecture with PCA and SVD. Symmetry 2019, 11, 1. [Google Scholar] [CrossRef]
- Popescu, D.; Ichim, L. Intelligent image processing system for detection and segmentation of regions of interest in retinal images. Symmetry 2018, 10, 73. [Google Scholar] [CrossRef]
- Han, H.C. Twisted blood vessels: Symptoms, etiology and biomechanical mechanisms. J. Vasc. Res. 2012, 49, 185–197. [Google Scholar] [CrossRef] [PubMed]
- Moss, H.E. Retinal vascular changes are a marker for cerebral vascular diseases. Curr. Neurol. Neurosci. Rep. 2015, 15, 40. [Google Scholar] [CrossRef] [PubMed]
- Hassan, S.S.A.; Bong, D.B.L.; Premsenthil, M. Detect on of neovascularization in diabetic retinopathy. J. Digit. Imaging 2012, 25, 437–444. [Google Scholar] [CrossRef] [PubMed]
- Wong, T.Y.; Klein, R.; Klein, B.E.K.; Tielsch, J.M.; Hubbard, L.; Nieto, F.J. Retinal microvascular abnormalities and their relationship with hypertension, cardiovascular disease, and mortality. Surv. Ophthalmol. 2001, 46, 59–80. [Google Scholar] [CrossRef]
- Nowilaty, S.; Al-Shamsi, H.; Al-Khars, W. Idiopathic juxtafoveolar retinal telangiectasis: A current review. Middle East Afr. J. Ophthalmol. 2010, 17, 224. [Google Scholar] [CrossRef]
- Niemeijer, M.; Xu, X.; Dumitrescu, A.V.; Gupta, P.; Van Ginneken, B.; Folk, J.C.; Abramoff, M.D. Automated measurement of the arteriolar-to-venular width ratio in digital color fundus photographs. IEEE Trans. Med. Imaging 2011, 30, 1941–1950. [Google Scholar] [CrossRef]
- Ünver, H.; Kökver, Y.; Duman, E.; Erdem, O. Statistical edge detection and circular hough transform for optic disk localization. Appl. Sci. 2019, 9, 350. [Google Scholar] [CrossRef]
- Al-Bander, B.; Williams, B.M.; Al-Nuaimy, W.; Al-Taee, M.A.; Pratt, H.; Zheng, Y. Dense fully convolutional segmentation of the optic disc and cup in colour fundus for glaucoma diagnosis. Symmetry 2018, 10, 87. [Google Scholar] [CrossRef]
- Sarathi, M.P.; Dutta, M.K.; Singh, A.; Travieso, C.M. Blood vessel inpainting based technique for efficient localization and segmentation of optic disc in digital fundus images. Biomed. Signal Process. Control 2016, 25, 108–117. [Google Scholar] [CrossRef]
- Almotiri, J.; Elleithy, K.; Elleithy, A. Retinal vessels Segmentation techniques and algorithms: A survey. Appl. Sci. 2018, 8, 155. [Google Scholar] [CrossRef]
- Yang, Y.; Huang, S.; Rao, N. An automatic hybrid method for retinal blood vessel extraction. Int. J. Appl. Math. Comput. Sci. 2008, 18, 399–407. [Google Scholar] [CrossRef]
- Chaudhuri, S.; Chatterjee, S.; Katz, N.; Nelson, M.; Goldbaum, M. Detection of blood vessels in retinal images using two-dimensional matched filters. IEEE Trans. Med. Imaging 1989, 8, 263–269. [Google Scholar] [CrossRef] [PubMed]
- Al-Rawi, M.; Qutaishat, M.; Arrar, M. An improved matched filter for blood vessel detection of digital retinal images. Comput. Biol. Med. 2007, 37, 262–267. [Google Scholar] [CrossRef] [PubMed]
- Chakraborti, T.; Jha, D.K.; Chowdhury, A.S.; Jiang, X. A self-adaptive matched filter for retinal blood vessel detection. Mach. Vis. Appl. 2014, 26, 55–68. [Google Scholar] [CrossRef]
- Singh, N.P.; Srivastava, R. Retinal blood vessels segmentation by using Gumbel probability distribution function based matched filter. Comput. Methods Programs Biomed. 2016, 129, 40–50. [Google Scholar] [CrossRef]
- Dharmawan, D.A.; Ng, B.P.; Rahardja, S. A modified Dolph-Chebyshev type II function matched filter for retinal vessels segmentation. Symmetry 2018, 10, 257. [Google Scholar] [CrossRef]
- Frangi, A.; Niessen, W.; Vincken, K.; Viergever, M. Multiscale vessel enhancement filtering medical image computing and computer-assisted interventation—MICCAI. In Medical Image Computing and Computer-Assisted Interventation—MICCAI’98; Springer: Berlin/Heidelberg, Germany, 1998; Volume 1496, pp. 130–137. ISBN 978-3-540-65136-9. [Google Scholar]
- Sofka, M.; Stewart, C.V. Retinal vessel centerline extraction using multiscale matched filters, confidence and edge measures. IEEE Trans. Med. Imaging 2006, 25, 1531–1546. [Google Scholar] [CrossRef]
- Saffarzadeh, V.M.; Osareh, A.; Shadgar, B. Vessel segmentation in retinal images using multi-scale line operator and K-means clustering. J. Med. Signals Sens. 2014, 4, 122–129. [Google Scholar] [PubMed]
- Zhang, L.; Fisher, M.; Wang, W. Retinal vessel segmentation using multi-scale textons derived from keypoints. Comput. Med. Imaging Graph. 2015, 45, 47–56. [Google Scholar] [CrossRef] [PubMed]
- Joshi, V.S.; Reinhardt, J.M.; Garvin, M.K.; Abramoff, M.D. Automated method for identification and artery-venous classification of vessel trees in retinal vessel networks. PLoS ONE 2014, 9, e88061. [Google Scholar] [CrossRef] [PubMed]
- Lázár, I.; Hajdu, A. Segmentation of retinal vessels by means of directional response vector similarity and region growing. Comput. Biol. Med. 2015, 66, 209–221. [Google Scholar] [CrossRef] [PubMed]
- Roychowdhury, S.; Koozekanani, D.D.; Parhi, K.K. Iterative Vessel segmentation of fundus images. IEEE Trans. Biomed. Eng. 2015, 62, 1738–1749. [Google Scholar] [CrossRef] [PubMed]
- Al-Diri, B.; Hunter, A.; Steel, D. An active contour model for segmenting and measuring retinal vessels. IEEE Trans. Med. Imaging 2009, 28, 1488–1497. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Rada, L.; Chen, K.; Harding, S.P.; Zheng, Y. Automated Vessel segmentation using infinite perimeter active contour model with hybrid region Information with application to retinal images. IEEE Trans. Med. Imaging 2015, 34, 1797–1807. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Zhao, J.; Yang, J.; Liu, Y.; Zhao, Y.; Zheng, Y.; Xia, L.; Wang, Y. Saliency driven vasculature segmentation with infinite perimeter active contour model. Neurocomputing 2017, 259, 201–209. [Google Scholar] [CrossRef]
- Kande, G.B.; Subbaiah, P.V.; Savithri, T.S. Unsupervised fuzzy based vessel segmentation in pathological digital fundus images. J. Med. Syst. 2010, 34, 849–858. [Google Scholar] [CrossRef]
- Allen, K.; Joshi, N.; Noble, J.A. Tramline and NP windows estimation for enhanced unsupervised retinal vessel segmentation. In Proceedings of the International Symposium on Biomedical Imaging, Chicago, IL, USA, 30 March–2 April 2011; pp. 1387–1390. [Google Scholar]
- Soares, J.V.B.; Leandro, J.J.G.; Cesar, R.M.; Jelinek, H.F.; Cree, M.J. Retinal vessel segmentation using the 2-D Gabor wavelet and supervised classification. IEEE Trans. Med. Imaging 2006, 25, 1214–1222. [Google Scholar] [CrossRef]
- Rahebi, J.; Hardalaç, F. Retinal blood vessel segmentation with neural network by using gray-level co-occurrence matrix-based features patient facing systems. J. Med. Syst. 2014, 38, 85. [Google Scholar] [CrossRef] [PubMed]
- Orlando, J.I.; Blaschko, M. Learning fully-connected CRFs for blood vessel segmentation in retinal images. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Boston, MA, USA, 14–18 September 2014. Lecture Notes in Computer Science (including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics). [Google Scholar]
- Aslani, S.; Sarnel, H. A new supervised retinal vessel segmentation method based on robust hybrid features. Biomed. Signal Process. Control 2016, 30, 1–12. [Google Scholar] [CrossRef]
- Zhang, J.; Chen, Y.; Bekkers, E.; Wang, M.; Dashtbozorg, B.; ter Haar Romeny, B.M. Retinal vessel delineation using a brain-inspired wavelet transform and random forest. Pattern Recognit. 2017, 69, 107–123. [Google Scholar] [CrossRef]
- Guo, Y.; Budak, Ü.; Şengür, A.; Smarandache, F. A retinal Vessel detection approach based on Shearlet transform and indeterminacy filtering on fundus images. Symmetry 2017, 9, 235. [Google Scholar] [CrossRef]
- Li, Q.; Feng, B.; Xie, L.; Liang, P.; Zhang, H.; Wang, T. A cross-modality learning approach for vessel segmentation in retinal images. IEEE Trans. Med. Imaging 2016, 35, 109–118. [Google Scholar] [CrossRef] [PubMed]
- Liskowski, P.; Krawiec, K. Segmenting retinal blood vessels with deep neural networks. IEEE Trans. Med. Imaging 2016, 35, 2369–2380. [Google Scholar] [CrossRef] [PubMed]
- Wu, A.; Xu, Z.; Gao, M.; Buty, M.; Mollura, D.J. Deep vessel tracking: A generalized probabilistic approach via deep learning. In Proceedings of the International Symposium on Biomedical Imaging, Prague, Czech Republic, 13–16 April 2016; pp. 1363–1367. [Google Scholar]
- Xie, S.; Tu, Z. Holistically-Nested Edge Detection. Int. J. Comput. Vis. 2017, 1–16. [Google Scholar] [CrossRef]
- Fu, H.; Xu, Y.; Wong, D.W.K.; Liu, J. Retinal vessel segmentation via deep learning network and fully-connected conditional random fields. In Proceedings of the International Symposium on Biomedical Imaging, Prague, Czech Republic, 13–16 April 2016; pp. 698–701. [Google Scholar]
- Maninis, K.K.; Pont-Tuset, J.; Arbeláez, P.; Van Gool, L. Deep Retinal Image Understanding; Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Cham, Switzerland, 2016; Volume 9901 LNCS, pp. 140–148. [Google Scholar]
- Mo, J.; Zhang, L. Multi-level deep supervised networks for retinal vessel segmentation. Int. J. Comput. Assist. Radiol. Surg. 2017, 12, 2181–2193. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Yu, Q.; Xu, X.; Gu, Y.; Yang, J. Improving dense conditional random field for retinal vessel segmentation by discriminative feature learning and thin-vessel enhancement. Comput. Methods Programs Biomed. 2017, 148, 13–25. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y. A Labeling-free approach to supervising deep neural networks for retinal blood Vessel segmentation. arXiv 2017, arXiv:1704.07502. [Google Scholar]
- Yan, Z.; Yang, X.; Cheng, K.T.T. A Three-stage deep learning model for accurate retinal Vessel segmentation. IEEE J. Biomed. Health Inform. 2019, 23, 1427–1436. [Google Scholar] [CrossRef] [PubMed]
- Niemeijer, M.; Staal, J.; van Ginneken, B.; Loog, M.; Abràmoff, M.D. Comparative study of retinal vessel segmentation methods on a new publicly available database. In Proceedings of the Medical Imaging 2004, Medical Imaging 2004: Image Processing, San Diego, CA, USA, 12 May 2004; Volume 5370, pp. 648–656. [Google Scholar]
- Hoover, A. Locating blood vessels in retinal images by piecewise threshold probing of a matched filter response. IEEE Trans. Med. Imaging 2000, 19, 203–210. [Google Scholar] [CrossRef] [PubMed]
- Odstrcilik, J.; Kolar, R.; Kubena, T.; Cernosek, P.; Budai, A.; Hornegger, J.; Gazarek, J.; Svoboda, O.; Jan, J.; Angelopoulou, E. Retinal vessel segmentation by improved matched filtering: Evaluation on a new high-resolution fundus image database. IET Image Process. 2013, 7, 373–383. [Google Scholar] [CrossRef]
- Zuiderveld, K. Contrast Limited Adaptive Histogram Equalization. In Graphics Gems; Academic Press Professional, Inc.: San Diego, CA, USA, 1994; pp. 474–485. ISBN 0-12-336155-9. [Google Scholar]
- Simonyan, K.; Zisserman, A. Very Deep Convolutional Networks for Large-Scale Image Recognition. In Proceedings of the 3rd International Conference on Learning Representations, San Diego, CA, USA, 7–9 May 2015. [Google Scholar]
- Zhang, Y.D.; Dong, Z.; Chen, X.; Jia, W.; Du, S.; Muhammad, K.; Wang, S.H. Image based fruit category classification by 13-layer deep convolutional neural network and data augmentation. Multimed. Tools Appl. 2019, 78, 3613–3632. [Google Scholar] [CrossRef]
- Wang, S.; Sun, J.; Mehmood, I.; Pan, C.; Chen, Y.; Zhang, Y.D. Cerebral micro-bleeding identification based on a nine-layer convolutional neural network with stochastic pooling. Concurr. Comput. 2019, e5130. [Google Scholar] [CrossRef]
- Jia, Y.; Shelhamer, E.; Donahue, J.; Karayev, S.; Long, J.; Girshick, R.; Guadarrama, S.; Darrell, T. Caffe: Convolutional architecture for fast feature embedding. In Proceedings of the 22nd ACM International Conference on Multimedia (ACM 2014), Orlando, FL, USA, 3–7 November 2014; pp. 675–678. [Google Scholar]





| Formation of DS_1 Layer | Size [Width, Height, Depth] | Formation of DS_2 Layer | Size [Width, Height, Depth] |
|---|---|---|---|
| Crop | Crop | ||
| Concatenate (DS_1) | Concatenate (DS_2) |
| Layers in the Proposed DNN | Output Size [Width, Height, Depth] | Activation Maps | Parameters (Weights) |
|---|---|---|---|
| Input image | [562 562 3] | 3 planes | |
| Conv 1_1 | [562 562 64] | 64 | (3 × 3 × 3 + 1) × 64 = 1792 |
| Conv 1_2 | [562 562 64] | 64 | (3 × 3 × 64 + 1) × 64 = 36,928 |
| Max pooling | [281 281 64] | 64 | 0 |
| Conv 2_1 | [281 281 128] | 128 | (3 × 3 × 64 + 1) × 128 = 73,856 |
| Conv 2_2 | [281 281 128] | 128 | (3 × 3 × 128 + 1) × 128 = 147,584 |
| Max pooling | [141 141 128] | 128 | 0 |
| Conv 3_1 | [141 141 256] | 256 | (3 × 3 × 128 + 1) × 256 = 295,168 |
| Conv 3_2 | [141 141 256] | 256 | (3 × 3 × 256 + 1) × 256 = 590,080 |
| Conv 3_3 | [141 141 256] | 256 | (3 × 3 × 256 + 1) × 256 = 590,080 |
| Max pooling | [71 71 256] | 256 | 0 |
| Conv 4_1 | [71 71 512] | 512 | (3 × 3 × 256 + 1) × 512 = 1,180,160 |
| Conv 4_2 | [71 71 512] | 512 | (3 × 3 × 512 + 1) × 512 = 2,359,808 |
| Conv 4_3 | [71 71 512] | 512 | (3 × 3 × 512 + 1) × 512 = 2,359,808 |
| DS_1 layer | [562 562 32] | (8 × 4 = 32) | [(3 × 3 × 64 + 1) × 8 + (3 × 3 × 128 + 1) × 8 + (3 × 3 × 256 + 1) × 8 + (3 × 3 × 512 + 1) × 8] = 69,152 |
| DS_2 layer | [562 562 64] | (16 × 4 = 64) | [(3 × 3 × 64 + 1) × 16 + (3 × 3 × 128 + 1) × 16 + (3 × 3 × 256 + 1) × 16 + (3 × 3 × 512 + 1) × 16] = 138,304 |
| Conv1_DS_8/16 layer | [562 562 32/64] | 32/64 | (3 × 3 × 32 + 1) × 32 = 9248/3 × 3× 32 + 1) × 64 = 18,496 |
| Conv2_DS_8/16 layer | [562 562 32/64] | 32/64 | (3 × 3 × 32 + 1) × 32 = 9248/3 × 3 × 32 + 1) × 64 = 18,496 |
| sp1/sp2 | [562 562 1/1] | 1/1 | (1 × 1 × 32 + 1) × 1 = 33/(1 × 1 × 64 + 1) × 1 = 65 |
| Final 1 × 1 conv output | [562 562 1] | 1 | (1 × 1 × 2 + 1) × 1 = 3 |
| Method | Author/Year/Ref. | Metrics Obtained from DRIVE Dataset | Metrics Obtained from STARE Dataset | ||||||
|---|---|---|---|---|---|---|---|---|---|
| SN | SP | Acc | AUC | SN | SP | Acc | AUC | ||
| Ophthalmologist | 0.7763 | 0.9723 | 0.947 | - | 0.8951 | 0.9384 | 0.9348 | - | |
| Matched filter | Chakraborti et al. (2014) [17] | 0.7205 | 0.9579 | 0.9370 | 0.9419 | 0.6786 | 0.9586 | 0.9379 | - |
| Singh, Srivatsava (2016) [18] | 0.7594 | 0.9708 | 0.9522 | 0.9287 | 0.7939 | 0.9376 | 0.9270 | 0.9140 | |
| Multi-scale approach | Saffarzadeh et al. (2014) [22] | - | - | 0.9387 | 0.9303 | - | - | 0.9483 | 0.9431 |
| Zhang, Fisher, et al. (2015) [23] | 0.7812 | 0.9668 | 0.9504 | - | - | - | - | - | |
| Region growing method | Lazar and Hajdu (2015) [25] | 0.7646 | 0.9723 | 0.9458 | - | 0.7248 | 0.9751 | 0.9492 | - |
| Roychowdhury et al. (2015) [26] | 0.739 | 0.978 | 0.949 | 0.967 | 0.732 | 0.984 | 0.956 | 0.967 | |
| Active contour model | Zhao, Rada, et al. (2015) [28] | 0.742 | 0.982 | 0.954 | 0.862 | 0.780 | 0.978 | 0.956 | 0.874 |
| Zhao, Zhao, et al. (2017) [29] | 0.782 | 0.979 | 0.957 | 0.886 | 0.789 | 0.978 | 0.956 | 0.885 | |
| Unsupervised method | Kande et al. (2010) [30] | - | - | 0.8911 | 0.9518 | - | - | 0.8976 | 0.9298 |
| Allen et al. (2011) [31] | - | - | 0.9342 | - | - | - | - | ||
| Supervised method | Aslani and Sarnel (2016) [35] | 0.7545 | 0.9801 | 0.9513 | 0.9682 | 0.7556 | 0.9837 | 0.9605 | 0.9789 |
| Zhang, Chen, et al. (2017) [36] | 0.7861 | 0.9712 | 0.9466 | 0.9703 | 0.7882 | 0.9729 | 0.9547 | 0.9740 | |
| Deep learning method | Li et al. (2016) [38] | 0.7569 | 0.9816 | 0.9527 | 0.9738 | 0.7726 | 0.9844 | 0.9628 | 0.9879 |
| Liskowski and Krawiec (2016) [39] | 0.7520 | 0.9806 | 0.9515 | 0.9710 | 0.8145 | 0.9866 | 0.9696 | 0.9880 | |
| Fu et al. (2016) [42] | 0.7294 | - | 0.947 | - | 0.714 | - | 0.9545 | - | |
| Maninis et al. (2016) [43] | 0.9497 | 0.9377 | 0.9386 | 0.9862 | 0.9403 | 0.9552 | 0.9543 | 0.9748 | |
| Mo and Zhang (2017) [44] | 0.7779 | 0.9780 | 0.9521 | 0.9782 | 0.8147 | 0.9844 | 0.9674 | 0.9885 | |
| Zhou et al. (2017) [45] | 0.8078 | 0.9674 | 0.9469 | - | 0.8065 | 0.9761 | 0.9585 | - | |
| Chen (2017) [46] | 0.7426 | 0.9735 | 0.9453 | 0.9516 | 0.7295 | 0.9696 | 0.9449 | 0.9557 | |
| Yan et al. (2018) [47] | 0.7631 | 0.9820 | 0.9538 | 0.9750 | 0.7735 | 0.9857 | 0.9638 | 0.9833 | |
| Proposed method | 0.8282 | 0.9738 | 0.9609 | 0.9786 | 0.8979 | 0.9701 | 0.9646 | 0.9892 | |
| DNN Framework | Preprocessed with Mean Value Subtraction | Preprocessed with the Proposed Preprocessing | ||||
|---|---|---|---|---|---|---|
| SN | SP | Acc | SN | SP | Acc | |
| Front end: 4 stages of VGG-16 Fine-tuning phase: DS_1 and DS_2 layers with a conv1_DS_8/16 layer | 0.8474 | 0.9652 | 0.9547 | 0.8428 | 0.9677 | 0.9560 |
| Front end: 4 stages of VGG-16 Fine-tuning phase: DS_1 and DS_2 layers with conv1_DS_8/16 & conv2_DS_8/16 layers (our model) | 0.9058 | 0.9514 | 0.9472 | 0.8282 | 0.9738 | 0.9609 |
| DNN Framework | Preprocessed with Mean Value Subtraction | Preprocessed with the Proposed Preprocessing | ||||
|---|---|---|---|---|---|---|
| SN | SP | Acc | SN | SP | Acc | |
| Front end: 4 stages of VGG-16 Fine-tuning phase: DS_1 and DS_2 layers with a conv1_DS_ 8/16 layer | 0.6581 | 0.9581 | 0.9379 | 0.9199 | 0.9630 | 0.9599 |
| Front end: 4 stages of VGG-16 Fine-tuning phase: DS_1 and DS_2 layers with conv1_DS_8/16 and conv2_DS_8/16 layers (our model) | 0.4184 | 0.9875 | 0.9461 | 0.8979 | 0.9701 | 0.9645 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Samuel, P.M.; Veeramalai, T. Multilevel and Multiscale Deep Neural Network for Retinal Blood Vessel Segmentation. Symmetry 2019, 11, 946. https://doi.org/10.3390/sym11070946
Samuel PM, Veeramalai T. Multilevel and Multiscale Deep Neural Network for Retinal Blood Vessel Segmentation. Symmetry. 2019; 11(7):946. https://doi.org/10.3390/sym11070946
Chicago/Turabian StyleSamuel, Pearl Mary, and Thanikaiselvan Veeramalai. 2019. "Multilevel and Multiscale Deep Neural Network for Retinal Blood Vessel Segmentation" Symmetry 11, no. 7: 946. https://doi.org/10.3390/sym11070946
APA StyleSamuel, P. M., & Veeramalai, T. (2019). Multilevel and Multiscale Deep Neural Network for Retinal Blood Vessel Segmentation. Symmetry, 11(7), 946. https://doi.org/10.3390/sym11070946




