Microbial Biofilm Diversity and Prevalence of Antibiotic Resistance Genes in Drinking Water Distribution System of Peshawar, Pakistan
Abstract
:1. Introduction
2. Materials and Methods
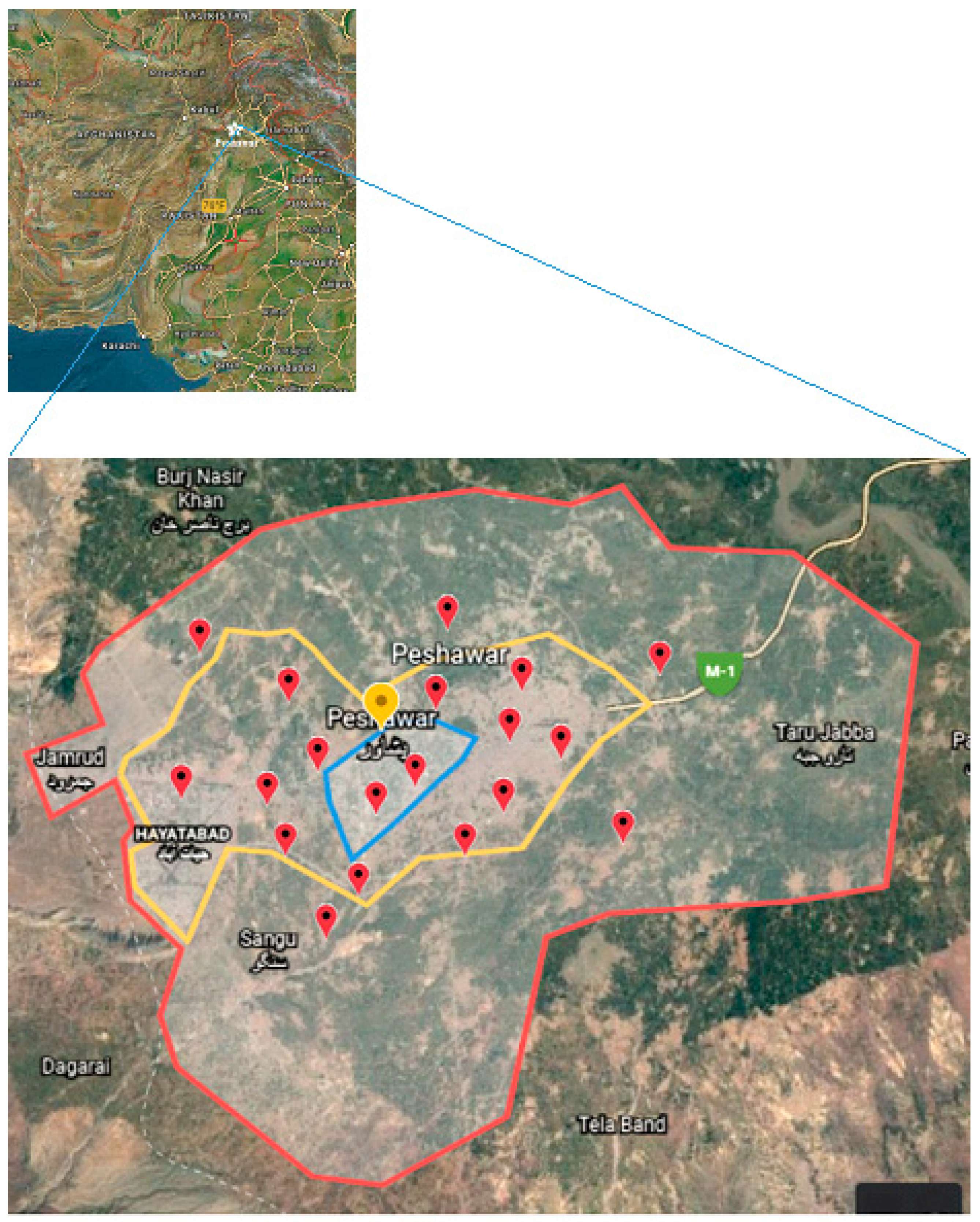
2.1. Study Area and Sampling
2.2. DNA Extraction
2.3. Shotgun Metagenomic Sequencing and Quality Filtering
2.4. Bioinformatics Analysis
2.5. Statistical Analysis
3. Results
3.1. Taxonomic Diversity
3.1.1. Alpha Diversity
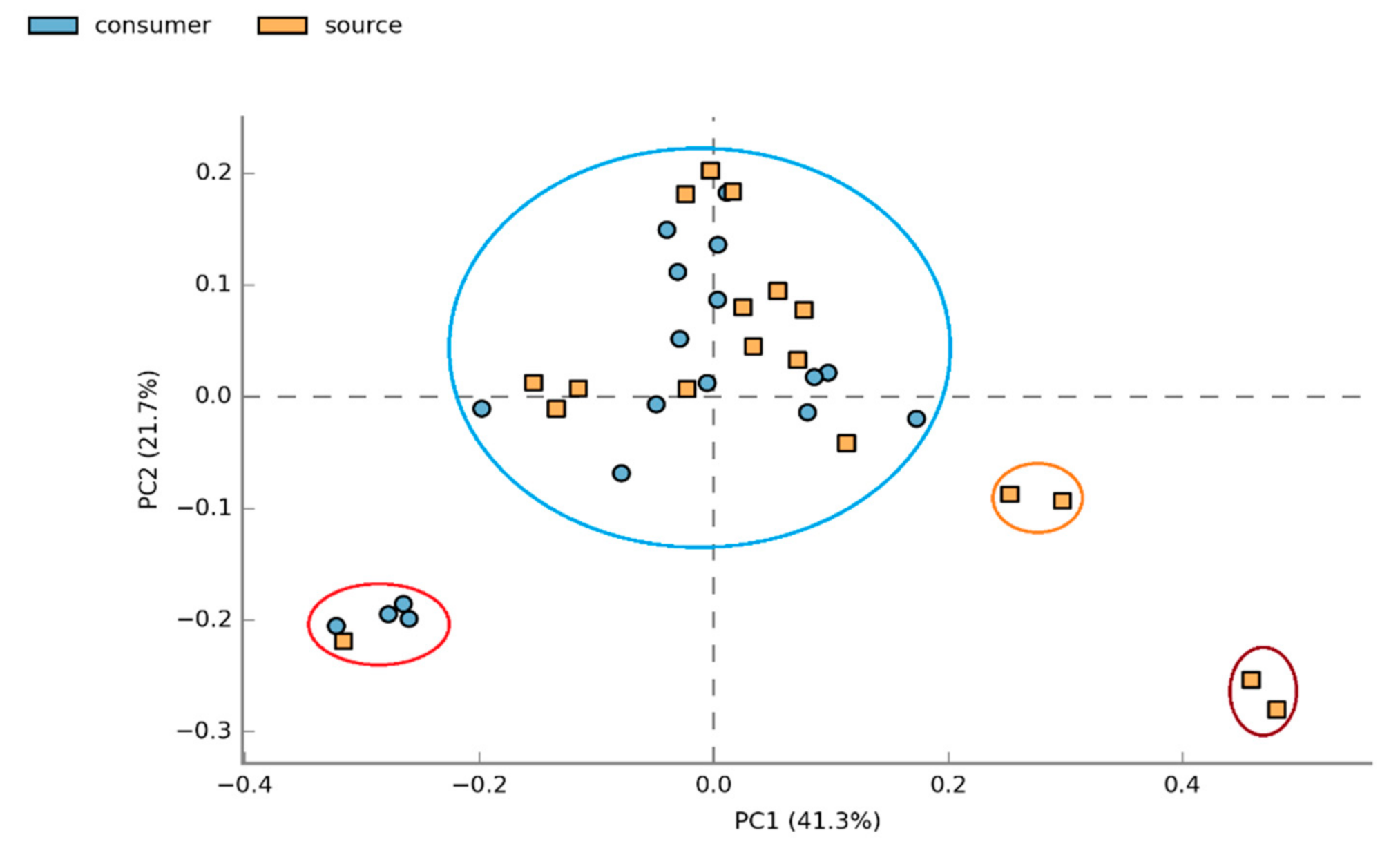
3.1.2. Beta Diversity
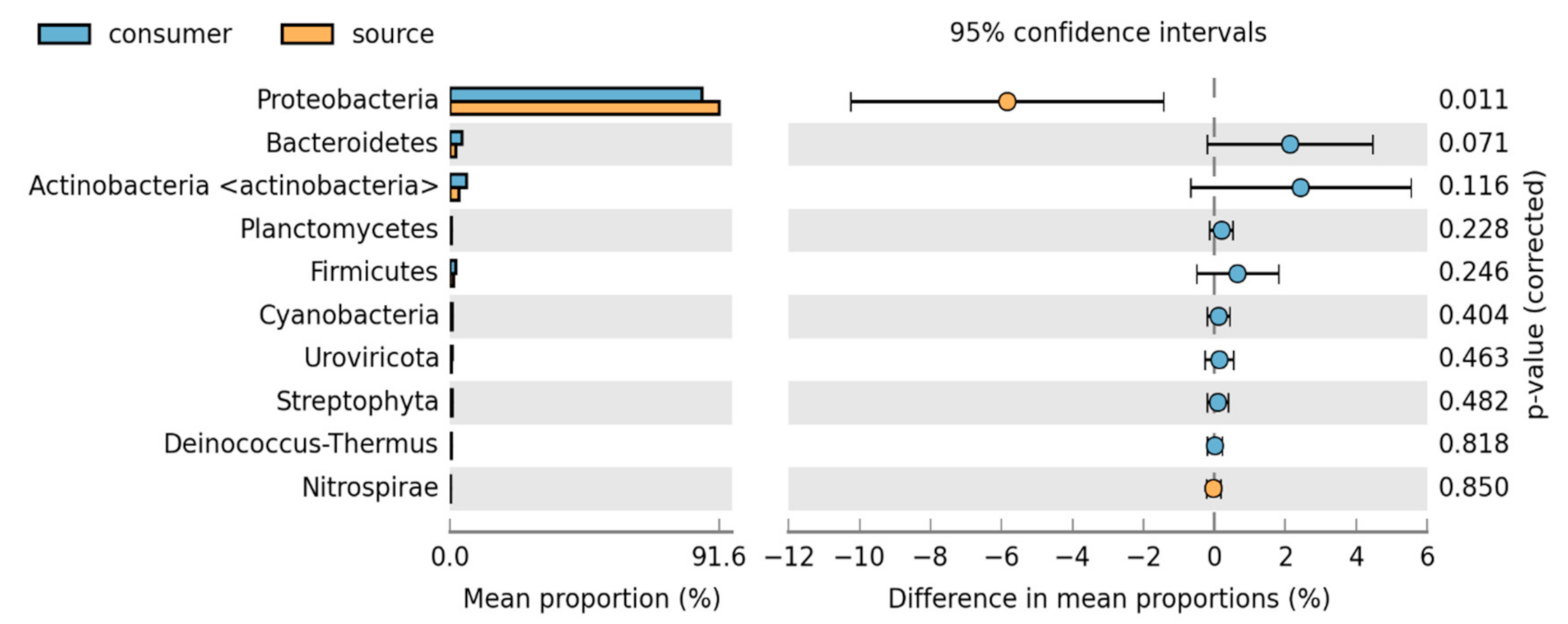
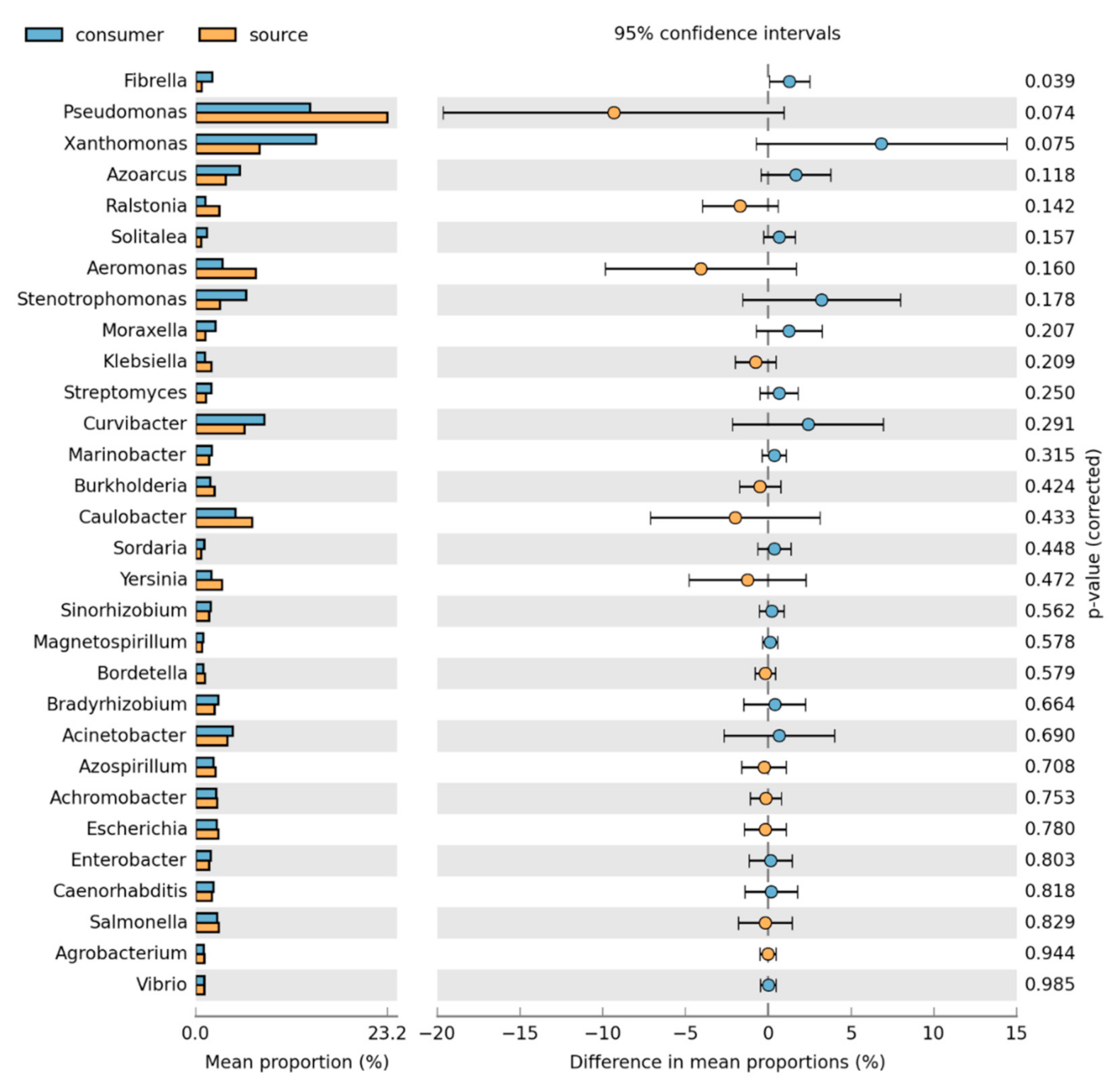
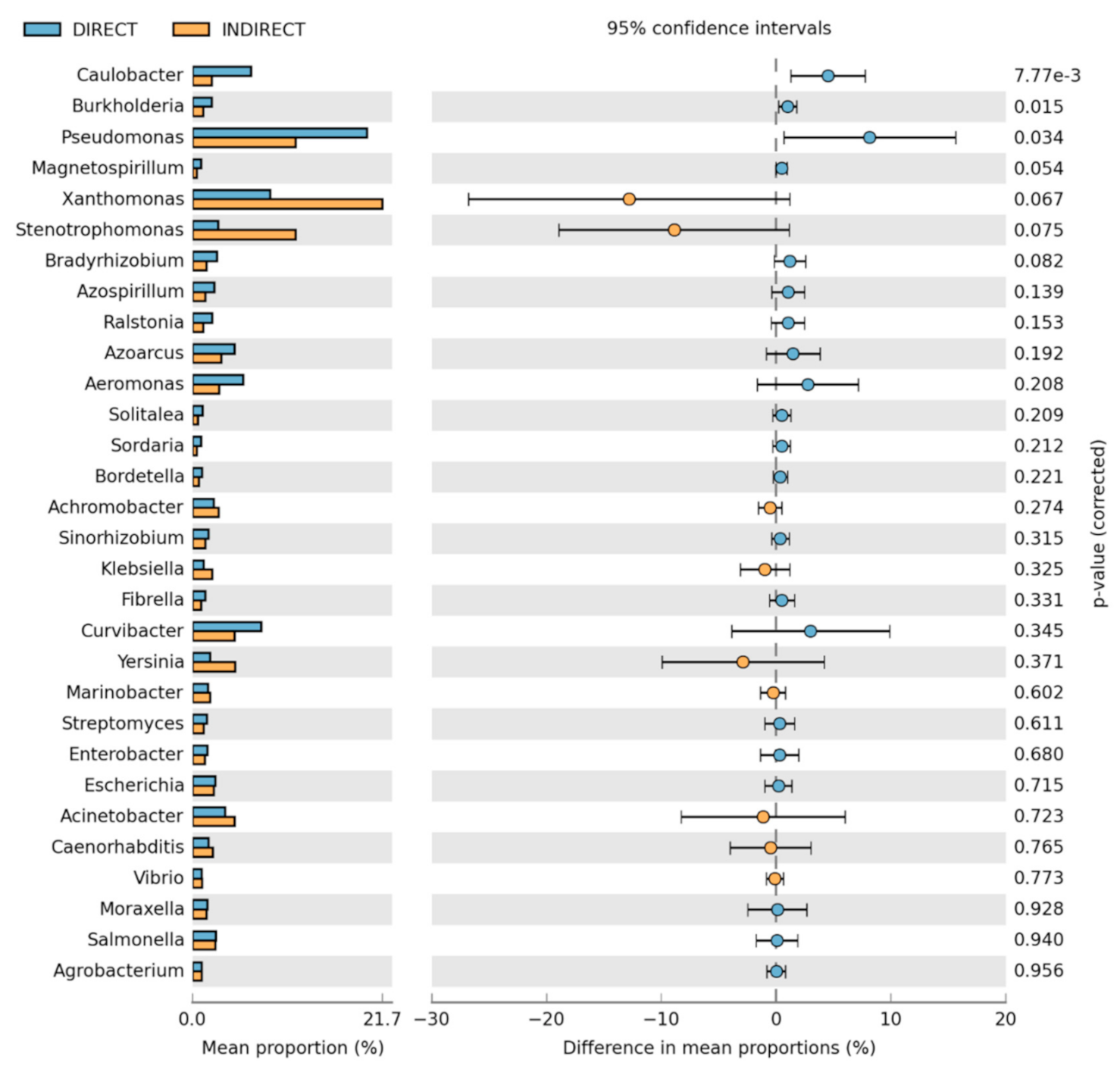
3.2. Bacteria Community Composition
Taxonomic and Differential Abundance Profiles
3.3. DWDS Features and Their Relationship with Bacterial Biofilm Community
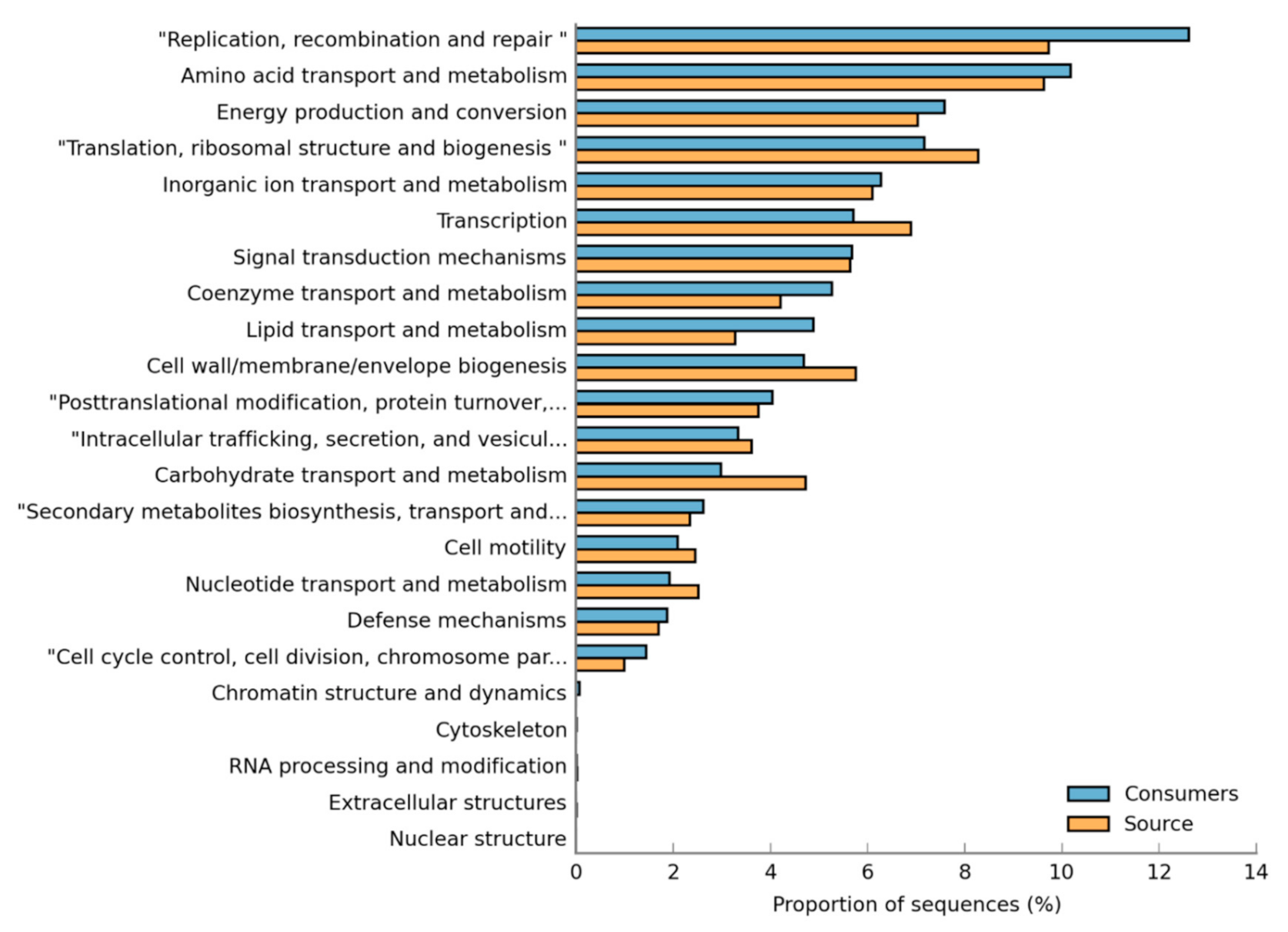
3.4. Functional Annotation of DWDS Biofilm Metagenome
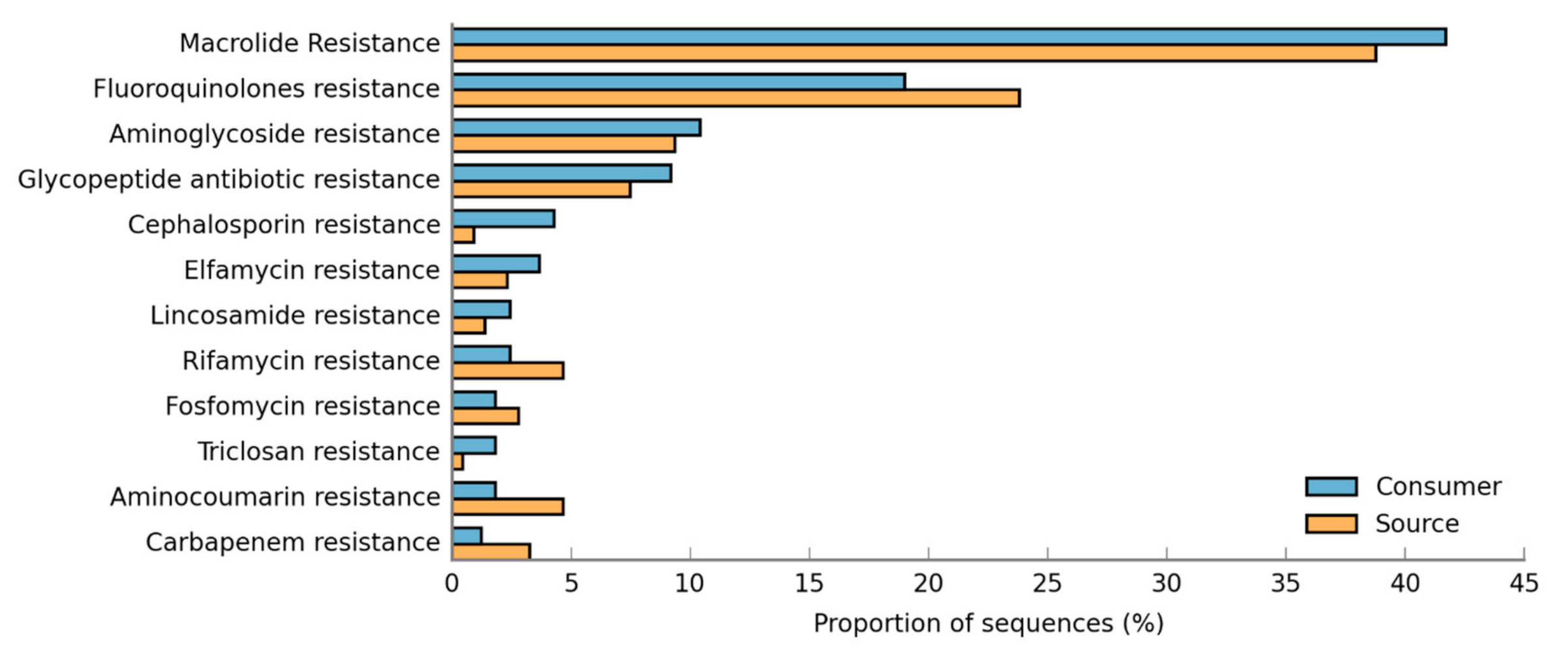
3.5. Resistance Genes Prevalence in DWDS Biofilms
4. Discussion
4.1. Genral Discussion
4.2. Limitations of the Study
4.3. Practical Implications and Future Directions
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Declaration of Alma-Ata. Available online: https://www.who.int/teams/social-determinants-of-health/declaration-of-alma-ata (accessed on 22 May 2021).
- Da’ana, D.A.; Zouari, N.; Ashfaq, M.Y.; Abu-Dieyeh, M.; Khraisheh, M.; Hijji, Y.M.; Al-Ghouti, M.A. Removal of Toxic Elements and Microbial Contaminants from Groundwater Using Low-Cost Treatment Options. Curr. Pollut. Rep. 2021, 1–25. [Google Scholar] [CrossRef]
- World Health Organization (WHO). WHO|Progress on Drinking Water, Sanitation and Hygiene. Available online: http://www.who.int/water_sanitation_health/publications/jmp-2017/en/ (accessed on 26 May 2021).
- Vestby, L.K.; Grønseth, T.; Simm, R.; Nesse, L.L. Bacterial Biofilm and its Role in the Pathogenesis of Disease. Antibiotics 2020, 9, 59. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stewart, P.S.; Costerton, J.W. Antibiotic resistance of bacteria in biofilms. Lancet Lond. Engl. 2001, 358, 135–138. [Google Scholar] [CrossRef]
- Lof, M.; Janus, M.M.; Krom, B.P. Metabolic Interactions between Bacteria and Fungi in Commensal Oral Biofilms. J. Fungi 2017, 3, 40. [Google Scholar]
- Chan, S.; Pullerits, K.; Keucken, A.; Persson, K.M.; Paul, C.J.; Rådström, P. Bacterial release from pipe biofilm in a full-scale drinking water distribution system. Npj Biofilms Microbiomes 2019, 5, 9. [Google Scholar] [CrossRef] [PubMed]
- Bai, X.; Ma, X.; Xu, F.; Li, J.; Zhang, H.; Xiao, X. The drinking water treatment process as a potential source of affecting the bacterial antibiotic resistance. Sci. Total Environ. 2015, 533, 24–31. [Google Scholar] [CrossRef] [PubMed]
- Anh, H.Q.; Le, T.P.Q.; Da Le, N.; Lu, X.X.; Duong, T.T.; Garnier, J.; Rochelle-Newall, E.; Zhang, S.; Oh, N.-H.; Oeurng, C.; et al. Antibiotics in surface water of East and Southeast Asian countries: A focused review on contamination status, pollution sources, potential risks, and future perspectives. Sci. Total Environ. 2021, 764, 142865. [Google Scholar] [CrossRef] [PubMed]
- Wu, D.-L.; Zhang, M.; He, L.-X.; Zou, H.-Y.; Liu, Y.-S.; Li, B.-B.; Yang, Y.-Y.; Liu, C.; He, L.-Y.; Ying, G.-G. Contamination profile of antibiotic resistance genes in ground water in comparison with surface water. Sci. Total Environ. 2020, 715, 136975. [Google Scholar] [CrossRef]
- Wang, H.; Wang, N.; Wang, B.; Zhao, Q.; Fang, H.; Fu, C.; Tang, C.; Jiang, F.; Zhou, Y.; Chen, Y.; et al. Antibiotics in Drinking Water in Shanghai and Their Contribution to Antibiotic Exposure of School Children. Environ. Sci. Technol. 2016, 50, 2692–2699. [Google Scholar] [CrossRef]
- Ribeiro, A.F.; Bodilis, J.; Alonso, L.; Buquet, S.; Feuilloley, M.; Dupont, J.P.; Pawlak, B. Occurrence of multi-antibiotic resistant Pseudomonas spp. in drinking water produced from karstic hydrosystems. Sci. Total Environ. 2014, 490, 370–378. [Google Scholar] [CrossRef]
- Cosgrove, S.E. The relationship between antimicrobial resistance and patient outcomes: Mortality, length of hospital stay, and health care costs. Clin. Infect Dis. Off. Publ. Infect Dis. Soc. Am. 2006, 42 (Suppl. 2), S82–S89. [Google Scholar] [CrossRef] [Green Version]
- O’Neill, J. Tackling Drug-Resistant Infections Globally: Final Report and Recommendations. Government of the United Kingdom. May 2016. Available online: https://apo.org.au/node/63983 (accessed on 16 June 2021).
- Li, J.; Cheng, W.; Xu, L.; Strong, P.J.; Chen, H. Antibiotic-resistant genes and antibiotic-resistant bacteria in the effluent of urban residential areas, hospitals, and a municipal wastewater treatment plant system. Environ. Sci. Pollut. Res. Int. 2015, 22, 4587–4596. [Google Scholar] [CrossRef]
- Azizullah, A.; Khattak, M.N.K.; Richter, P.; Häder, D.-P. Water pollution in Pakistan and its impact on public health—A review. Environ. Int. 2011, 37, 479–497. [Google Scholar] [CrossRef]
- WHO. Guidelines for Drinking-Water Quality, 4th ed., Incorporating the 1st Addendum. 2017. Available online: https://www.who.int/publications-detail-redirect/9789241549950 (accessed on 27 May 2021).
- Lührig, K.; Canbäck, B.; Paul, C.J.; Johansson, T.; Persson, K.M.; Rådström, P. Bacterial Community Analysis of Drinking Water Biofilms in Southern Sweden. Microbes Environ. 2015, 30, 99–107. [Google Scholar] [CrossRef] [Green Version]
- PBS. District and Tehsil Level Population Summary with Region Breakup. Pakistan Bureau of Statistics, Islamabad, Pakistan. 2017. Available online: https://www.pbs.gov.pk/content/final-results-census-2017 (accessed on 27 May 2021).
- Hamida, A.; Javed, A.; Mohammad, N.A. Bacteriological analysis of drinking water of hand pumps in different schools of District Peshawar (Pakistan). J. Food Sci. 2006, 16, 34–38. [Google Scholar]
- Zahoorullah, T.A. Quality of drinking water in rural Peshawar. Pak. J. Med. Res. 2003, 42, 85–89. [Google Scholar]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data–Science Open. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 26 May 2021).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinform. Oxf. Engl. 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pruitt, K.D.; Tatusova, T.; Maglott, D.R. NCBI Reference Sequence (RefSeq): A curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res. 2005, 33, D501–D504. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods. 2015, 12, 59–60. [Google Scholar] [CrossRef]
- Huson, D.H.; Auch, A.F.; Qi, J.; Schuster, S.C. MEGAN analysis of metagenomic data. Genome Res. 2007, 17, 377–386. [Google Scholar] [CrossRef] [Green Version]
- Hunt, M.; Mather, A.E.; Sánchez-Busó, L.; Page, A.J.; Parkhill, J.; Keane, J.A.; Harris, S.R. ARIBA: Rapid antimicrobial resistance genotyping directly from sequencing reads. Microb. Genom. 2017, 3, e000131. [Google Scholar] [CrossRef]
- Parks, D.H.; Tyson, G.W.; Hugenholtz, P.; Beiko, R.G. STAMP: Statistical analysis of taxonomic and functional profiles. Bioinf. Oxf. Engl. 2014, 30, 3123–3124. [Google Scholar] [CrossRef] [Green Version]
- Bray, J.R.; Curtis, J.T. An Ordination of the Upland Forest Communities of Southern Wisconsin. Ecol. Monogr. 1957, 27, 325–349. [Google Scholar] [CrossRef]
- Rice, E.W.; Baird, R.B.; Eaton, A.D. Standard Methods for the Examination of Water and Wastewater, 23rd ed.; American Public Health Association, American Water Works Association, Water Environment Federation: Alexandria, VA, USA, 2017. [Google Scholar]
- Field, K.G.; Samadpour, M. Fecal source tracking, the indicator paradigm, and managing water quality. Water Res. 2007, 41, 3517–3538. [Google Scholar] [CrossRef] [PubMed]
- Acharya, K.; Khanal, S.; Pantha, K.; Amatya, N.; Davenport, R.J.; Werner, D. A comparative assessment of conventional and molecular methods, including MinION nanopore sequencing, for surveying water quality. Sci. Rep. 2019, 9, 15726. [Google Scholar] [CrossRef]
- Besemer, K.; Singer, G.; Quince, C.; Bertuzzo, E.; Sloan, W.; Battin, T.J. Headwaters are critical reservoirs of microbial diversity for fluvial networks. Proc. R. Soc. B Biol. Sci. 2013, 280, 20131760. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Dong, D.; Wan, K.; Chen, C.; Yu, X.; Lin, H. Potential shift of bacterial community structure and corrosion-related bacteria in drinking water distribution pipeline driven by water source switching. Front. Environ. Sci. Eng. 2020, 15, 28. [Google Scholar] [CrossRef]
- Zhou, Z.; Xu, L.; Zhu, L.; Liu, Y.; Shuai, X.; Lin, Z.; Chen, H. Metagenomic analysis of microbiota and antibiotic resistome in household activated carbon drinking water purifiers. Environ. Int. 2021, 148, 106394. [Google Scholar] [CrossRef]
- Liu, R.; Yu, Z.; Zhang, H.; Yang, M.; Shi, B.; Liu, X. Diversity of bacteria and mycobacteria in biofilms of two urban drinking water distribution systems. Can. J. Microbiol. 2012, 58, 261–270. [Google Scholar] [CrossRef] [Green Version]
- Brumfield, K.D.; Hasan, N.A.; Leddy, M.B.; Cotruvo, J.A.; Rashed, S.M.; Colwell, R.R.; Huq, A. A comparative analysis of drinking water employing metagenomics. PLoS ONE 2020, 15, e0231210. [Google Scholar] [CrossRef] [Green Version]
- Mi, Z.; Dai, Y.; Xie, S.; Chen, C.; Zhang, X. Impact of disinfection on drinking water biofilm bacterial community. J. Environ. Sci. China 2015, 37, 200–205. [Google Scholar] [CrossRef]
- Rothballer, M.; Picot, M.; Sieper, T.; Arends, J.B.; Schmid, M.; Hartmann, A.; Boon, N.; Buisman, C.J.; Barrière, F.; Strik, D.P. Monophyletic group of unclassified γ-Proteobacteria dominates in mixed culture biofilm of high-performing oxygen reducing biocathode. Bioelectrochemistry 2015, 106 Pt A, 167–176. [Google Scholar] [CrossRef]
- Augimeri, R.V.; Varley, A.J.; Strap, J.L. Establishing a Role for Bacterial Cellulose in Environmental Interactions: Lessons Learned from Diverse Biofilm-Producing Proteobacteria. Front. Microbiol. 2015, 6, 1282. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hentchel, K.L.; Reyes Ruiz, L.M.; Curtis, P.D.; Fiebig, A.; Coleman, M.L.; Crosson, S. Genome-scale fitness profile of Caulobacter crescentus grown in natural freshwater. ISME J. 2019, 13, 523–536. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hwang, G.; Kang, S.; El-Din, M.G.; Liu, Y. Impact of an extracellular polymeric substance (EPS) precoating on the initial adhesion of Burkholderia cepacia and Pseudomonas aeruginosa. Biofouling 2012, 28, 525–538. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Kim, D.; Lee, T. Microbial diversity in biofilms on water distribution pipes of different materials. Water Sci. Technol. J. Int. Assoc. Water Pollut. Res. 2010, 61, 163–171. [Google Scholar] [CrossRef]
- Niquette, P.; Servais, P.; Savoir, R. Impacts of pipe materials on densities of fixed bacterial biomass in a drinking water distribution system. Water Res. 2000, 34, 1952–1956. [Google Scholar] [CrossRef]
- Papciak, D.; Tchórzewska-Cieślak, B.; Domoń, A.; Wojtuś, A.; Żywiec, J.; Konkol, J. The Impact of the Quality of Tap Water and the Properties of Installation Materials on the Formation of Biofilms. Water 2019, 11, 1903. [Google Scholar] [CrossRef] [Green Version]
- USAID. Planning and Engineering Services for Master Plan in Peshawar Khyber Pakhtunkhwa: Drinking Water, Sanitation/Storm Water and Solid Waste Services-Executive Summary—Consolidation of Key Findings of Volumes 1–3—Deliverable No. 24. 2014. Available online: http://urbanpolicyunit.gkp.pk/wp-content/uploads/2018/02/Water-Sanitation-Peshawar-Report.pdf (accessed on 27 May 2021).
- Xu, L.; Ouyang, W.; Qian, Y.; Su, C.; Su, J.; Chen, H. High-throughput profiling of antibiotic resistance genes in drinking water treatment plants and distribution systems. Environ. Pollut. 2016, 213, 119–126. [Google Scholar] [CrossRef]
- Lv, L.; Jiang, T.; Zhang, S.; Yu, X. Exposure to Mutagenic Disinfection Byproducts Leads to Increase of Antibiotic Resistance in Pseudomonas aeruginosa. Environ. Sci. Technol. 2014, 48, 8188–8195. [Google Scholar] [CrossRef]

| Taxonomical Levels | Shannon−Weaver Index | Simpsons’s Reciprocal Index | ||
|---|---|---|---|---|
| Source End | Consumer End | Source End | Consumer End | |
| Phylum level | 0.728 ± 0.326 | 1.186 ± 0.197 | 0.933 ± 0.376 | 1.353 ± 0.296 |
| Genus level | 4.716 ± 0.753 | 5.147 ± 0.574 | 14.168 ± 7.131 | 18.09 ± 7.773 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ullah, H.; Shahzad, M.; Saleem, F.; Ali, T.; Azim, M.K.; Khan, H.; Ali, J.; Ahmed, J. Microbial Biofilm Diversity and Prevalence of Antibiotic Resistance Genes in Drinking Water Distribution System of Peshawar, Pakistan. Water 2021, 13, 1788. https://doi.org/10.3390/w13131788
Ullah H, Shahzad M, Saleem F, Ali T, Azim MK, Khan H, Ali J, Ahmed J. Microbial Biofilm Diversity and Prevalence of Antibiotic Resistance Genes in Drinking Water Distribution System of Peshawar, Pakistan. Water. 2021; 13(13):1788. https://doi.org/10.3390/w13131788
Chicago/Turabian StyleUllah, Habib, Muhammad Shahzad, Faizan Saleem, Taj Ali, Muhammad Kamran Azim, Haris Khan, Johar Ali, and Jawad Ahmed. 2021. "Microbial Biofilm Diversity and Prevalence of Antibiotic Resistance Genes in Drinking Water Distribution System of Peshawar, Pakistan" Water 13, no. 13: 1788. https://doi.org/10.3390/w13131788
APA StyleUllah, H., Shahzad, M., Saleem, F., Ali, T., Azim, M. K., Khan, H., Ali, J., & Ahmed, J. (2021). Microbial Biofilm Diversity and Prevalence of Antibiotic Resistance Genes in Drinking Water Distribution System of Peshawar, Pakistan. Water, 13(13), 1788. https://doi.org/10.3390/w13131788







