Evidence of the Physical Interaction between Rpl22 and the Transposable Element Doc5, a Heterochromatic Transposon of Drosophila melanogaster
Abstract
1. Introduction
2. Materials and Methods
2.1. Plasmids Construction
2.2. PCR Amplification
2.3. One Hybrid Screening
2.4. Protein Expression and Purification
2.5. Electrophoretic Mobility Shift Assay (EMSA)
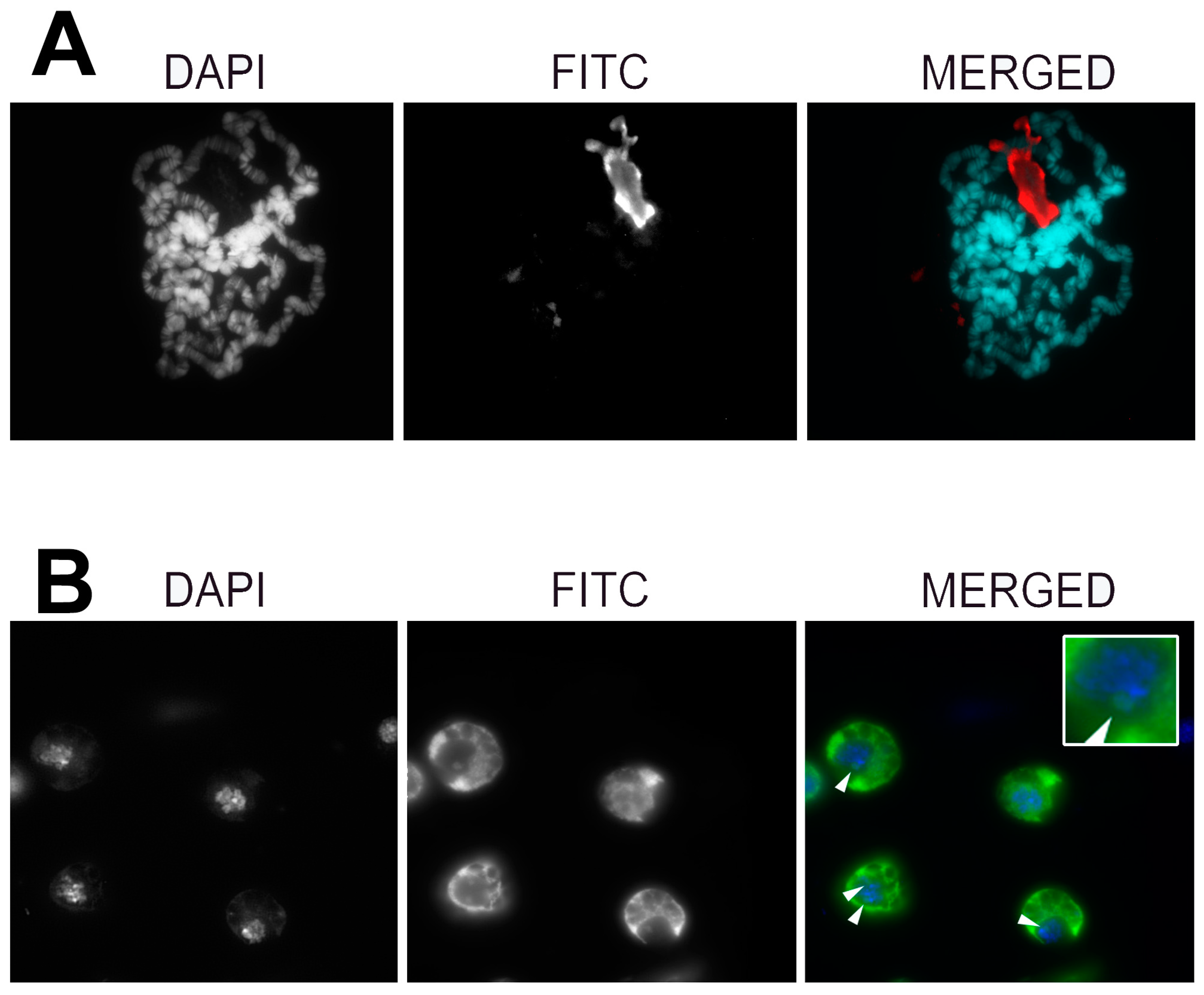
2.6. Fluorescence In Situ Hybridization and Immunofluorescence on Polytene Chromosomes
2.7. Other Methods
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kornberg, R.D. Chromatin Structure: A Repeating Unit of Histones and DNA. Science 1974, 184, 868. [Google Scholar] [CrossRef] [PubMed]
- Lindsay, S. Chromatin control of gene expression: The simplest model. Biophys. J. 2007, 92, 1113. [Google Scholar] [CrossRef][Green Version]
- Schmitt, A.D.; Hu, M.; Ren, B. Genome-wide mapping and analysis of chromosome architecture. Nat. Rev. Mol. Cell Biol. 2016, 17, 743–755. [Google Scholar] [CrossRef] [PubMed]
- Simpson, R.T. Structure of the chromatosome, a chromatin particle containing 160 base pairs of DNA and all the histones. Biochemistry 1978, 17, 5524–5531. [Google Scholar] [CrossRef]
- Consortium, E.P. An integrated encyclopedia of DNA elements in the human genome. Nature 2012, 489, 57–74. [Google Scholar] [CrossRef]
- Chen, Z.X.; Sturgill, D.; Qu, J.; Jiang, H.; Park, S.; Boley, N.; Suzuki, A.M.; Fletcher, A.R.; Plachetzki, D.C.; FitzGerald, P.C.; et al. Comparative validation of the D. melanogaster modENCODE transcriptome annotation. Genome Res. 2014, 24, 1209–1223. [Google Scholar] [CrossRef] [PubMed]
- Arunkumar, G.; Melters, D.P. Centromeric Transcription: A Conserved Swiss-Army Knife. Genes 2020, 11, 911. [Google Scholar] [CrossRef]
- Tachiwana, H.; Yamamoto, T.; Saitoh, N. Gene regulation by non-coding RNAs in the 3D genome architecture. Curr. Opin. Genet. Dev. 2020, 61, 69–74. [Google Scholar] [CrossRef] [PubMed]
- Barral, A.; Déjardin, J. Telomeric Chromatin and TERRA. J. Mol. Biol. 2020, 432, 4244–4256. [Google Scholar] [CrossRef]
- Zeng, C.; Fukunaga, T.; Hamada, M. Identification and analysis of ribosome-associated lncRNAs using ribosome profiling data. BMC Genom. 2018, 19, 414. [Google Scholar] [CrossRef]
- Mélèse, T.; Xue, Z. The nucleolus: An organelle formed by the act of building a ribosome. Curr. Opin. Cell Biol. 1995, 7, 319–324. [Google Scholar] [CrossRef]
- Marsano, R.M.; Giordano, E.; Messina, G.; Dimitri, P. A New Portrait of Constitutive Heterochromatin: Lessons from Drosophila melanogaster. Trends Genet. 2019, 35, 615–631. [Google Scholar] [CrossRef] [PubMed]
- Larracuente, A.M.; Presgraves, D.C. The selfish Segregation Distorter gene complex of Drosophila melanogaster. Genetics 2012, 192, 33–53. [Google Scholar] [CrossRef]
- Berloco, M.; Fanti, L.; Fau-Breiling, A.; Breiling, A.; Fau-Orlando, V.; Orlando, V.; Fau-Pimpinelli, S.; Pimpinelli, S. The maternal effect gene, abnormal oocyte (abo), of Drosophila melanogaster encodes a specific negative regulator of histones. Proc. Natl. Acad. Sci. USA 2001, 98, 12126–12131. [Google Scholar] [CrossRef]
- Tritto, P.; Specchia, V.; Fanti, L.; Berloco, M.; D’Alessandro, R.; Pimpinelli, S.; Palumbo, G.; Pia Bozzetti, M. Structure, Regulation and Evolution of the Crystal–Stellate System of Drosophila. Genetica 2003, 117, 247–257. [Google Scholar] [CrossRef]
- Caizzi, R.; Caggese, C.; Pimpinelli, S. Bari-1, a new transposon-like family in Drosophila melanogaster with a unique heterochromatic organization. Genetics 1993, 133, 335–345. [Google Scholar] [CrossRef]
- Palazzo, A.; Lovero, D.; D’Addabbo, P.; Caizzi, R.; Marsano, R.M. Identification of Bari Transposons in 23 Sequenced Drosophila Genomes Reveals Novel Structural Variants, MITEs and Horizontal Transfer. PLoS ONE 2016, 11, e0156014. [Google Scholar] [CrossRef]
- McGurk, M.P.; Barbash, D.A. Double insertion of transposable elements provides a substrate for the evolution of satellite DNA. Genome Res. 2018, 28, 714–725. [Google Scholar] [CrossRef] [PubMed]
- Caggese, C.; Pimpinelli, S.; Barsanti, P.; Caizzi, R. The distribution of the transposable element Bari-1 in the Drosophila melanogaster and Drosophila simulans genomes. Genetica 1995, 96, 269–283. [Google Scholar] [CrossRef]
- Palazzo, A.; Marconi, S.; Specchia, V.; Bozzetti, M.P.; Ivics, Z.; Caizzi, R.; Marsano, R.M. Functional Characterization of the Bari1 Transposition System. PLoS ONE 2013, 8, e79385. [Google Scholar] [CrossRef]
- Palazzo, A.; Moschetti, R.; Caizzi, R.; Marsano, R.M. The Drosophila mojavensis Bari3 transposon: Distribution and functional characterization. Mob. DNA 2014, 5, 21. [Google Scholar] [CrossRef]
- Sandler, L.; Hiraizumi, Y.; Fau-Sandler, I.; Sandler, I. Meiotic Drive in Natural Populations of Drosophila melanogaster. I. the Cytogenetic Basis of Segregation-Distortion. Genetics 1959, 44, 233. [Google Scholar] [CrossRef] [PubMed]
- Pimpinelli, S.; Dimitri, P. Cytogenetic analysis of segregation distortion in Drosophila melanogaster: The cytological organization of the Responder (Rsp) locus. Genetics 1989, 121, 765–772. [Google Scholar] [CrossRef] [PubMed]
- Moschetti, R.; Palazzo, A.; Lorusso, P.; Viggiano, L.; Marsano, R.M. “What You Need, Baby, I Got It”: Transposable Elements as Suppliers of Cis-Operating Sequences in Drosophila. Biology 2020, 9, 25. [Google Scholar] [CrossRef] [PubMed]
- Marsano, R.M.; Milano, R.; Minervini, C.; Moschetti, R.; Caggese, C.; Barsanti, P.; Caizzi, R. Organization and possible origin of the Bari-1 cluster in the heterochromatic h39 region of Drosophila melanogaster. Genetica 2003, 117, 281–289. [Google Scholar] [CrossRef]
- Moschetti, R.; Caizzi, R.; Pimpinelli, S. Segregation distortion in Drosophila melanogaster: Genomic organization of Responder sequences. Genetics 1996, 144, 1365–1371. [Google Scholar] [CrossRef] [PubMed]
- Marsano, R.M.; Moschetti, R.; Barsanti, P.; Caggese, C.; Caizzi, R. A survey of the DNA sequences surrounding the Bari1 repeats in the pericentromeric h39 region of Drosophila melanogaster. Gene 2003, 307, 167–174. [Google Scholar] [CrossRef]
- Zhao, W.; Bidwai, A.P.; Glover, C.V.C. Interaction of casein kinase II with ribosomal protein L22 of Drosophila melanogaster. Biochem. Biophys. Res. Commun. 2002, 298, 60–66. [Google Scholar] [CrossRef]
- Bayona-Feliu, A.; Casas-Lamesa, A.; Reina, O.; Bernués, J.; Azorín, F. Linker histone H1 prevents R-loop accumulation and genome instability in heterochromatin. Nat. Commun. 2017, 8, 283. [Google Scholar] [CrossRef]
- Koyama, Y.; Katagiri, S.; Hanai, S.; Uchida, K.; Miwa, M. Poly(ADP-ribose) polymerase interacts with novel Drosophila ribosomal proteins, L22 and L23a, with unique histone-like amino-terminal extensions. Gene 1999, 226, 339–345. [Google Scholar] [CrossRef]
- Kearse, M.G.; Ireland, J.A.; Prem, S.M.; Chen, A.S.; Ware, V.C. RpL22e, but not RpL22e-like-PA, is SUMOylated and localizes to the nucleoplasm of Drosophila meiotic spermatocytes. Nucleus 2013, 4, 241–258. [Google Scholar] [CrossRef] [PubMed]
- Mageeney, C.M.; Ware, V.C. Specialized eRpL22 paralogue-specific ribosomes regulate specific mRNA translation in spermatogenesis in Drosophila melanogaster. Mol. Biol. Cell 2019, 30, 2240–2253. [Google Scholar] [CrossRef]
- Gietz, R.D. Yeast transformation by the LiAc/SS carrier DNA/PEG method. Methods Mol. Biol. 2014, 1205, 1–12. [Google Scholar]
- Colloms, S.D.; van Luenen, H.G.; Plasterk, R.H. DNA binding activities of the Caenorhabditis elegans Tc3 transposase. Nucleic Acids Res. 1994, 22, 5548–5554. [Google Scholar] [CrossRef]
- Marsano, R.M.; Moschetti, R.; Caggese, C.; Lanave, C.; Barsanti, P.; Caizzi, R. The complete Tirant transposable element in Drosophila melanogaster shows a structural relationship with retrovirus-like retrotransposons. Gene 2000, 247, 87–95. [Google Scholar] [CrossRef]
- James, T.C.; Eissenberg, J.C.; Craig, C.; Dietrich, V.; Hobson, A.; Elgin, S.C. Distribution patterns of HP1, a heterochromatin-associated nonhistone chromosomal protein of Drosophila. Eur. J. Cell Biol. 1989, 50, 170–180. [Google Scholar]
- Marck, C. ‘DNA Strider’: A ‘C’ program for the fast analysis of DNA and protein sequences on the Apple Macintosh family of computers. Nucleic Acids Res. 1988, 16, 1829–1836. [Google Scholar] [CrossRef]
- Kosugi, S.; Hasebe, M.; Fau-Tomita, M.; Tomita, M.; Fau-Yanagawa, H.; Yanagawa, H. Systematic identification of cell cycle-dependent yeast nucleocytoplasmic shuttling proteins by prediction of composite motifs. Proc. Natl. Acad. Sci. USA 2009, 106, 10171–10176. [Google Scholar] [CrossRef]
- Brameier, M.; Krings A Fau-MacCallum, R.M.; MacCallum, R.M. NucPred—Predicting nuclear localization of proteins. Bioinformatics 2007, 23, 1159–1160. [Google Scholar] [CrossRef] [PubMed]
- Coelho, P.A.; Nurminsky, D.; Hartl, D.; Sunkel, C.E. Identification of Porto-1, a new repeated sequence that localises close to the centromere of chromosome 2 of Drosophila melanogaster. Chromosoma 1996, 105, 211–222. [Google Scholar] [CrossRef] [PubMed]
- Csink, A.K.; Henikoff, S. Something from nothing: The evolution and utility of satellite repeats. Trends Genet. 1998, 14, 200–204. [Google Scholar] [CrossRef]
- Zhou, R.; Hotta, I.; Denli, A.M.; Hong, P.; Perrimon, N.; Hannon, G.J. Comparative analysis of argonaute-dependent small RNA pathways in Drosophila. Mol. Cell 2008, 32, 592–599. [Google Scholar] [CrossRef]
- Anger, A.M.; Armache, J.-P.; Berninghausen, O.; Habeck, M.; Subklewe, M.; Wilson, D.N.; Beckmann, R. Structures of the human and Drosophila 80S ribosome. Nature 2013, 497, 80–85. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Xu, S.; Xia, L.; Wang, J.; Wen, S.; Jin, P.; Chen, D. The Bantam microRNA Is Associated with Drosophila Fragile X Mental Retardation Protein and Regulates the Fate of Germline Stem Cells. PLoS Genet. 2009, 5, e1000444. [Google Scholar] [CrossRef]
- Celniker, S.E.; Dillon, L.A.L.; Gerstein, M.B.; Gunsalus, K.C.; Henikoff, S.; Karpen, G.H.; Kellis, M.; Lai, E.C.; Lieb, J.D.; MacAlpine, D.M.; et al. Unlocking the secrets of the genome. Nature 2009, 459, 927–930. [Google Scholar] [CrossRef]
- Caizzi, R.; Moschetti, R.; Piacentini, L.; Fanti, L.; Marsano, R.M.; Dimitri, P. Comparative Genomic Analyses Provide New Insights into the Evolutionary Dynamics of Heterochromatin in Drosophila. PLoS Genet. 2016, 12, e1006212. [Google Scholar] [CrossRef]
- Tzika, E.; Dreker, T.; Imhof, A. Epigenetics and Metabolism in Health and Disease. Front. Genet. 2018, 9, 361. [Google Scholar] [CrossRef]
- Signor, S. Transposable elements in individual genotypes of Drosophila simulans. Ecol. Evol. 2020, 10, 3402–3412. [Google Scholar] [CrossRef]
- Spana, C.; Harrison, D.A.; Corces, V.G. The Drosophila melanogaster suppressor of Hairy-wing protein binds to specific sequences of the gypsy retrotransposon. Genes Dev. 1988, 2, 1414–1423. [Google Scholar] [CrossRef] [PubMed]
- Georgiev, P.G.; Corces, V.G. The su(Hw) protein bound to gypsy sequences in one chromosome can repress enhancer-promoter interactions in the paired gene located in the other homolog. Proc. Natl. Acad. Sci. USA 1995, 92, 5184–5188. [Google Scholar] [CrossRef]
- Minervini, C.F.; Marsano, R.M.; Casieri, P.; Fanti, L.; Caizzi, R.; Pimpinelli, S.; Rocchi, M.; Viggiano, L. Heterochromatin protein 1 interacts with 5’UTR of transposable element ZAM in a sequence-specific fashion. Gene 2007, 393, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Bhavsar, R.B.; Makley, L.N.; Tsonis, P.A. The other lives of ribosomal proteins. Hum. Genom. 2010, 4, 327. [Google Scholar] [CrossRef] [PubMed]
- Brogna, S.; Sato, T.-A.; Rosbash, M. Ribosome Components Are Associated with Sites of Transcription. Mol. Cell 2002, 10, 93–104. [Google Scholar] [CrossRef]
- Schroder, P.A.; Moore, M.J. Association of ribosomal proteins with nascent transcripts in S. cerevisiae. RNA 2005, 11, 1521–1529. [Google Scholar] [CrossRef]
- Wool, I.G. Extraribosomal functions of ribosomal proteins. Trends Biochem. Sci. 1996, 21, 164–165. [Google Scholar] [CrossRef]
- Warner, J.R.; McIntosh, K.B. How Common Are Extraribosomal Functions of Ribosomal Proteins? Mol. Cell 2009, 34, 3–11. [Google Scholar] [CrossRef]
- De, S.; Varsally, W.; Falciani, F.; Brogna, S. Ribosomal proteins’ association with transcription sites peaks at tRNA genes in Schizosaccharomyces pombe. RNA 2011, 17, 1713–1726. [Google Scholar] [CrossRef]
- Dai, M.-S.; Arnold, H.; Sun, X.-X.; Sears, R.; Lu, H. Inhibition of c-Myc activity by ribosomal protein L11. Embo J. 2007, 26, 3332–3345. [Google Scholar] [CrossRef]
- Dai, M.-S.; Sun, X.-X.; Lu, H. Ribosomal protein L11 associates with c-Myc at 5 S rRNA and tRNA genes and regulates their expression. J. Biol. Chem. 2010, 285, 12587–12594. [Google Scholar] [CrossRef]
- Wan, F.; Anderson, D.E.; Barnitz, R.A.; Snow, A.; Bidere, N.; Zheng, L.; Hegde, V.; Lam, L.T.; Staudt, L.M.; Levens, D.; et al. Ribosomal Protein S3: A KH Domain Subunit in NF-κB Complexes that Mediates Selective Gene Regulation. Cell 2007, 131, 927–939. [Google Scholar] [CrossRef] [PubMed]
- Wan, F.; Weaver, A.; Gao, X.; Bern, M.; Hardwidge, P.R.; Lenardo, M.J. IKKβ phosphorylation regulates RPS3 nuclear translocation and NF-κB function during infection with Escherichia coli strain O157:H7. Nat. Immunol. 2011, 12, 335–343. [Google Scholar] [CrossRef] [PubMed]
- Ni, J.-Q.; Liu, L.-P.; Hess, D.; Rietdorf, J.; Sun, F.-L. Drosophila ribosomal proteins are associated with linker histone H1 and suppress gene transcription. Genes Dev. 2006, 20, 1959–1973. [Google Scholar] [CrossRef]
- Kharchenko, P.V.; Alekseyenko, A.A.; Schwartz, Y.B.; Minoda, A.; Riddle, N.C.; Ernst, J.; Sabo, P.J.; Larschan, E.; Gorchakov, A.A.; Gu, T.; et al. Comprehensive analysis of the chromatin landscape in Drosophila melanogaster. Nature 2011, 471, 480–485. [Google Scholar] [CrossRef]
- Bonnet, J.; Lindeboom, R.G.H.; Pokrovsky, D.; Stricker, G.; Çelik, M.H.; Rupp, R.A.W.; Gagneur, J.; Vermeulen, M.; Imhof, A.; Müller, J. Quantification of Proteins and Histone Marks in Drosophila Embryos Reveals Stoichiometric Relationships Impacting Chromatin Regulation. Dev. Cell 2019, 51, 632–644.e6. [Google Scholar] [CrossRef] [PubMed]
- Sinzelle, L.; Izsvak, Z.; Ivics, Z. Molecular domestication of transposable elements: From detrimental parasites to useful host genes. Cell Mol. Life Sci. 2009, 66, 1073–1093. [Google Scholar] [CrossRef]
- Cosby, R.L.; Chang, N.C.; Feschotte, C. Host-transposon interactions: Conflict, cooperation, and cooption. Genes Dev. 2019, 33, 1098–1116. [Google Scholar] [CrossRef] [PubMed]
- Pane, A.; Jiang, P.; Zhao, D.Y.; Singh, M.; Schupbach, T. The Cutoff protein regulates piRNA cluster expression and piRNA production in the Drosophila germline. EMBO J. 2011, 30, 4601–4615. [Google Scholar] [CrossRef]
- Zhao, K.; Cheng, S.; Miao, N.; Xu, P.; Lu, X.; Zhang, Y.; Wang, M.; Ouyang, X.; Yuan, X.; Liu, W.; et al. A Pandas complex adapted for piRNA-guided transcriptional silencing and heterochromatin formation. Nat. Cell Biol. 2019, 21, 1261–1272. [Google Scholar] [CrossRef]
- Cherbas, L.; Willingham, A.; Zhang, D.; Yang, L.; Zou, Y.; Eads, B.D.; Carlson, J.W.; Landolin, J.M.; Kapranov, P.; Dumais, J.; et al. The transcriptional diversity of 25 Drosophila cell lines. Genome Res. 2011, 21, 301–314. [Google Scholar] [CrossRef]
- Lee, H.; McManus, C.J.; Cho, D.-Y.; Eaton, M.; Renda, F.; Somma, M.P.; Cherbas, L.; May, G.; Powell, S.; Zhang, D.; et al. DNA copy number evolution in Drosophila cell lines. Genome Biol. 2014, 15, R70. [Google Scholar]
- Hammond Mp Fau-Laird, C.D.; Laird, C.D. Control of DNA replication and spatial distribution of defined DNA sequences in salivary gland cells of Drosophila melanogaster. Chromosoma 1985, 91, 279–286. [Google Scholar] [CrossRef] [PubMed]
- Spradling, A.; Orr-Weaver, T. Regulation of DNA replication during Drosophila development. Annu. Rev. Genet. 1987, 21, 373–403. [Google Scholar] [CrossRef]
- Goshima, G.; Wollman, R.; Goodwin, S.S.; Zhang, N.; Scholey, J.M.; Vale, R.D.; Stuurman, N. Genes required for mitotic spindle assembly in Drosophila S2 cells. Science 2007, 316, 417–421. [Google Scholar] [CrossRef] [PubMed]
- Lindström, M.S. Emerging functions of ribosomal proteins in gene-specific transcription and translation. Biochem. Biophys. Res. Commun. 2009, 379, 167–170. [Google Scholar] [CrossRef] [PubMed]
- Marsano, R.M.; Caizzi, R.; Moschetti, R.; Junakovic, N. Evidence for a functional interaction between the Bari1 transposable element and the cytochrome P450 cyp12a4 gene in Drosophila melanogaster. Gene 2005, 357, 122–128. [Google Scholar] [CrossRef]
- Palazzo, A.; Caizzi, R.; Viggiano, L.; Marsano, R.M. Does the Promoter Constitute a Barrier in the Horizontal Transposon Transfer Process? Insight from Bari Transposons. Genome Biol. Evol. 2017, 9, 1637–1645. [Google Scholar] [PubMed]
- Palazzo, A.; Lorusso, P.; Miskey, C.; Walisko, O.; Gerbino, A.; Marobbio, C.M.T.; Ivics, Z.; Marsano, R.M. Transcriptionally promiscuous “blurry” promoters in Tc1/mariner transposons allow transcription in distantly related genomes. Mob. DNA 2019, 10, 13. [Google Scholar] [CrossRef] [PubMed]
- Suresh, S.; Ahn, H.W.; Joshi, K.; Dakshinamurthy, A.; Kananganat, A.; Garfinkel, D.J.; Farabaugh, P.J. Ribosomal protein and biogenesis factors affect multiple steps during movement of the Saccharomyces cerevisiae Ty1 retrotransposon. Mob. DNA 2015, 6, 22. [Google Scholar] [CrossRef]
| Primer | Sequence | Usage |
|---|---|---|
| ADread | 5′-CTATTCGATGATGAAGAT-3′ | sequencing |
| pACT2seq | 5′-TACCACTACAATGGATG-3′ | sequencing |
| pACT2 up | 5′-CTATTCGATGATGAAGATACCCCACCAAACCC-3′ | Amplification/cloning |
| pACT2 low | 5′-GTGAACTTGCGGGGTTTTTCAGTATCTACGAT-3′ | Amplification/cloning |
| His1_up | 5′-GAGGCCCTTTCGTCTTCAA-3′ | Amplification/cloning |
| His1_low | 5′-CTAGGGCTTTCTGCTCTGTCATCT-3′ | Amplification/cloning |
| Doc5_up | 5′-ACGGCTATTATTGTTTCTTATTGCT-3′ | Amplification/cloning |
| Doc5_low | 5′-TTATCCTCATCCCTTATCCTATGT-3′ | Amplification/cloning |
| pETup | 5′-CACCATGGCTTACCCATA-3′ | Amplification/cloning |
| pETlow | 5′-ATAAAAGAAGGCAAAACGATG-3′ | Amplification/cloning |
| H5low | 5′-CTAACGCAGCACGTTCTTCTT-3′ | Amplification/cloning |
| L22up | 5′-CACCAAGGTGGTCAAGAAGAA-3′ | Amplification/cloning |
| Subject Accession | Start | End | Chromosome | % Identity | Alignment Length | Evalue | Bit Score | Chromosome Map Position |
|---|---|---|---|---|---|---|---|---|
| NT_033779.5 | 23,430,152 | 23,429,509 | 2L | 85.891 | 645 | 0 | 643 | h35–36 |
| NT_033779.5 | 23,037,532 | 23,037,717 | 2L | 91.237 | 194 | 2.03 × 10−67 | 257 | h35–36 |
| NW_001845128.1 | 3990 | 4435 | 2CEN | 100 | 446 | 0 | 824 | deep het |
| NW_001845128.1 | 3875 | 3990 | 2CEN | 100 | 116 | 1.24 × 10−54 | 215 | deep het |
| NW_001844967.1 | 11,017 | 10,572 | 2CEN | 100 | 446 | 0 | 824 | deep het |
| NW_001844967.1 | 11,132 | 11,017 | 2CEN | 100 | 116 | 1.24 × 10−54 | 215 | deep het |
| NW_007931075.1 | 7023 | 6491 | 2CEN | 82.655 | 565 | 2.43 × 10−121 | 436 | deep het |
| NW_007931075.1 | 9914 | 9382 | 2CEN | 82.655 | 565 | 2.43 × 10−121 | 436 | deep het |
| NT_033778.4 | 396,636 | 397,234 | 2R | 99.332 | 599 | 0 | 1083 | h41–h44 |
| NT_033778.4 | 165,422 | 164,833 | 2R | 95.326 | 599 | 0 | 942 | h41–h44 |
| NT_033778.4 | 74,075 | 74,637 | 2R | 92.833 | 586 | 0 | 824 | h41–h44 |
| NT_033778.4 | 298,134 | 297,689 | 2R | 100 | 446 | 0 | 824 | h41–h44 |
| NT_033778.4 | 872,342 | 871,810 | 2R | 82.655 | 565 | 2.43 × 10−121 | 436 | h41–h44 |
| NT_033778.4 | 875,233 | 874,701 | 2R | 82.655 | 565 | 2.43 × 10−121 | 436 | h41–h44 |
| NT_033778.4 | 1,413,093 | 1,413,384 | 2R | 91.333 | 300 | 2.47 × 10−111 | 403 | h45 |
| NT_033778.4 | 3,518,328 | 3,518,018 | 2R | 88.179 | 313 | 1.95 × 10−97 | 357 | h41–h44 |
| NT_033778.4 | 5,012,337 | 5,012,064 | 2R | 82.818 | 291 | 4.43 × 10−59 | 230 | h46 |
| NT_033778.4 | 298,249 | 298,134 | 2R | 100 | 116 | 1.24 × 10−54 | 215 | h41-h44 |
| NT_033778.4 | 5,012,596 | 5,012,448 | 2R | 84.302 | 172 | 4.59 × 10−34 | 147 | h46 |
| NT_037436.4 | 24,877,579 | 24,878,162 | 3L | 87.081 | 596 | 0.0 | 656 | h49 |
| NT_037436.4 | 24,914,416 | 24,913,834 | 3L | 86.745 | 596 | 0.0 | 645 | h49 |
| NT_037436.4 | 24,944,076 | 24,944,602 | 3L | 87.199 | 539 | 2.93 × 10−170 | 599 | h49 |
| NT_037436.4 | 24,461,859 | 24,462,331 | 3L | 85.443 | 474 | 6.71 × 10−127 | 455 | h47 |
| NT_037436.4 | 23,664,082 | 23,663,846 | 3L | 94.583 | 240 | 9.00 × 10−101 | 368 | 80F9 |
| NT_037436.4 | 23,663,844 | 23,663,639 | 3L | 92.754 | 207 | 1.20 × 10−79 | 298 | 80F9 |
| NT_037436.4 | 25,490,094 | 25,490,271 | 3L | 94.382 | 178 | 2.02 × 10−72 | 274 | h49–h50 |
| NT_037436.4 | 27,913,043 | 27,913,159 | 3L | 93.277 | 119 | 7.57 × 10−42 | 172 | h51 |
| NT_037436.4 | 24,502,780 | 24,502,891 | 3L | 92.035 | 113 | 2.12 × 10−37 | 158 | h48 |
| NT_037436.4 | 27,912,886 | 27,913,038 | 3L | 81.609 | 174 | 2.15 × 10−27 | 124 | h51 |
| NT_033777.3 | 646,928 | 646,337 | 3R | 86.612 | 605 | 0.0 | 649 | h54–h56 |
| NT_033777.3 | 4,042,323 | 4,042,066 | 3R | 94.961 | 258 | 6.86 × 10−112 | 405 | 81F |
| NT_033777.3 | 1,401,453 | 1,401,023 | 3R | 83.991 | 431 | 1.50 × 10−103 | 377 | h54–h56 |
| NT_033777.3 | 4,050,664 | 4,050,455 | 3R | 95.238 | 210 | 3.28 × 10−90 | 333 | 81F |
| NT_033777.3 | 3,992,575 | 3,992,365 | 3R | 94.787 | 211 | 1.53 × 10−88 | 327 | 81F |
| NT_033777.3 | 4,039,954 | 4,039,769 | 3R | 91.710 | 193 | 1.57 × 10−68 | 261 | 81F |
| NT_033777.3 | 2,453,123 | 2,453,400 | 3R | 80.357 | 280 | 4.53 × 10−44 | 180 | h56 |
| NT_033777.3 | 2,453,399 | 2,453,508 | 3R | 92.793 | 111 | 5.90 × 10−38 | 159 | h56 |
| NW_001845051.1 | 2554 | 2831 | UNK | 80.357 | 280 | 4.53 × 10−44 | 180 | deep het |
| NW_001845051.1 | 2830 | 2934 | UNK | 89.189 | 111 | 1.29 × 10−29 | 132 | deep het |
| Gene Name | FlyBase ID | Function | Inferred by | Reference |
|---|---|---|---|---|
| vig | FBgn0024183 | Heterochromatin organization | Co-IP | [42] |
| AGO1 | FBgn0262739 | transcriptional repression | Co-IP | [42] |
| AGO2 | FBgn0087035 | transcriptional repression | Co-IP | [42] |
| vig2 | FBgn0046214 | Heterochromatin organization | Mass-spec | [43] |
| Fmr1 | FBgn0028734 | piRNA biogenesis | Co-IP | [42] |
| ban | FBgn0262451 | piRNA biogenesis | Co-IP | [42] |
| esi2 | FBgn0285992 | Unknown | Co-IP | [42] |
| smt3 | FBgn0264922 | mitosis | Co-IP | [31] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Berloco, M.F.; Minervini, C.F.; Moschetti, R.; Palazzo, A.; Viggiano, L.; Marsano, R.M. Evidence of the Physical Interaction between Rpl22 and the Transposable Element Doc5, a Heterochromatic Transposon of Drosophila melanogaster. Genes 2021, 12, 1997. https://doi.org/10.3390/genes12121997
Berloco MF, Minervini CF, Moschetti R, Palazzo A, Viggiano L, Marsano RM. Evidence of the Physical Interaction between Rpl22 and the Transposable Element Doc5, a Heterochromatic Transposon of Drosophila melanogaster. Genes. 2021; 12(12):1997. https://doi.org/10.3390/genes12121997
Chicago/Turabian StyleBerloco, Maria Francesca, Crescenzio Francesco Minervini, Roberta Moschetti, Antonio Palazzo, Luigi Viggiano, and René Massimiliano Marsano. 2021. "Evidence of the Physical Interaction between Rpl22 and the Transposable Element Doc5, a Heterochromatic Transposon of Drosophila melanogaster" Genes 12, no. 12: 1997. https://doi.org/10.3390/genes12121997
APA StyleBerloco, M. F., Minervini, C. F., Moschetti, R., Palazzo, A., Viggiano, L., & Marsano, R. M. (2021). Evidence of the Physical Interaction between Rpl22 and the Transposable Element Doc5, a Heterochromatic Transposon of Drosophila melanogaster. Genes, 12(12), 1997. https://doi.org/10.3390/genes12121997






