Patterns, Mechanisms and Genetics of Speciation in Reptiles and Amphibians
Abstract
1. Synopsis
2. Aspects of Amphibian and Reptile Speciation
2.1. Integrative Taxonomy Builds the Foundation of Modern Speciation Research
2.2. The Importance of Biogeography and the Climatic Niche
2.3. Ecological Speciation
2.4. Speciation Rates and Variable Traits
2.5. Genome Properties and Processes
3. Case Studies
3.1. Liolaemid Lizards—From Poorly Known Taxonomic Groups to Evolutionary Radiations
3.2. Lizard Speciation across the South American Dry Biomes
3.3. An Early Stage of Adaptive Ecological Speciation in European Fire Salamanders.
3.4. Body Size and Speciation Rates in Mantellid and Other Frogs
3.5. Convergent Phenotypic and Genomic Adaptations to Elevational Clines in a Caribbean Anolis Species
3.6. Galápagos Giant Tortoises: Dispersal, Allopatry and the Fusion-Fission Dynamics of Speciation
4. Conclusions and Outlook
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- University of California, Berkeley, CA, USA. Amphibia Web. Available online: https://amphibiaweb.org (accessed on 11 February 2019).
- Uetz, P. The Reptile Database. Available online: http://www.reptile-database.org/ (accessed on 11 May 2019).
- Mayr, E. Systematics and the Origin of Species, from the Viewpoint of a Zoologist; Harvard University Press: Cambridge, MA, USA, 1942; ISBN 9780674862500. [Google Scholar]
- Seehausen, O.; Butlin, R.K.; Keller, I.; Wagner, C.E.; Boughman, J.W.; Hohenlohe, P.A.; Peichel, C.L.; Saetre, G.-P.; Bank, C.; Brännström, A.; et al. Genomics and the origin of species. Nat. Rev. Genet. 2014, 15, 176–192. [Google Scholar] [CrossRef] [PubMed]
- Price, T. Speciation in Birds; Roberts & Company Publishers: Greenwood Village, CO, USA, 2008; ISBN 9780974707785. [Google Scholar]
- Vieites, D.R.; Wollenberg, K.C.; Andreone, F.; Köhler, J.; Glaw, F.; Vences, M. Vast underestimation of Madagascar’s biodiversity evidenced by an integrative amphibian inventory. Proc. Natl. Acad. Sci. USA 2009, 106, 8267–8272. [Google Scholar] [CrossRef] [PubMed]
- Barraclough, T.G.; Vogler, A.P.; Harvey, P.H. Revealing the factors that promote speciation. Philos. Trans. R. Soc. Lond. B Biol. Sci. 1998, 353, 241–249. [Google Scholar] [CrossRef]
- Buckley, L.B.; Jetz, W. Environmental and historical constraints on global patterns of amphibian richness. Proc. R. Soc. B Biol. Sci. 2007, 274, 1167–1173. [Google Scholar] [CrossRef] [PubMed]
- Roll, U.; Feldman, A.; Novosolov, M.; Allison, A.; Bauer, A.M.; Bernard, R.; Böhm, M.; Castro-Herrera, F.; Chirio, L.; Collen, B.; et al. The global distribution of tetrapods reveals a need for targeted reptile conservation. Nat. Ecol. Evol. 2017, 1, 1677–1682. [Google Scholar] [CrossRef] [PubMed]
- Qian, H. Environment–richness relationships for mammals, birds, reptiles, and amphibians at global and regional scales. Ecol. Res. 2010, 25, 629–637. [Google Scholar] [CrossRef]
- Orr, M.R.; Smith, T.B. Ecology and speciation. Trends Ecol. Evol. 1998, 13, 502–506. [Google Scholar] [CrossRef]
- Losos, J.B. Lizards in an Evolutionary Tree: Ecology and Adaptive Radiation of Anoles, 1st ed.; University of California Press: Berkeley, CA, USA, 2009; ISBN 9780520943735. [Google Scholar]
- Sobel, J.M.; Chen, G.F.; Watt, L.R.; Schemske, D.W. The biology of speciation. Evolution 2010, 64, 295–315. [Google Scholar] [CrossRef] [PubMed]
- Feder, M.E.; Burggren, W.W. Environmental Physiology of the Amphibians; University of Chicago Press: Chicago, IL, USA, 1992; ISBN 9780226239446. [Google Scholar]
- Tamarin, R.H.; Krebs, C.J. Microtus Population Biology. II. Genetic changes at the Transferrin locus in fluctuating populations of two vole species. Evolution 1969, 23, 183–211. [Google Scholar] [CrossRef] [PubMed]
- Gaines, M.S.; Krebs, C.J. Genetic changes in fluctuating vole populations. Evolution 1971, 25, 702–723. [Google Scholar] [CrossRef] [PubMed]
- Sinervo, B.; Svensson, E.; Comendant, T. Density cycles and an offspring quantity and quality game driven by natural selection. Nature 2000, 406, 985–988. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, A.A.; Rieseberg, L.H. Revisiting the impact of inversions in evolution: From population genetic markers to drivers of adaptive shifts and speciation? Annu. Rev. Ecol. Evol. Syst. 2008, 39, 21–42. [Google Scholar] [CrossRef] [PubMed]
- Sites, J.W.; Davis, S.K. Phylogenetic relationships and molecular variability within and among six chromosome races of Sceloporus grammicus (Sauria, Iguanidae), based on nuclear and mitochondrial markers. Evolution 1989, 43, 296–317. [Google Scholar] [CrossRef] [PubMed]
- Olmo, E. Rate of chromosome changes and speciation in reptiles. Genetica 2005, 125, 185–203. [Google Scholar] [CrossRef] [PubMed]
- Leaché, A.D.; Sites, J.W., Jr. Chromosome evolution and diversification in North American spiny lizards (genus Sceloporus). Cytogenetic Genome Res. 2009, 127, 166–181. [Google Scholar] [CrossRef] [PubMed]
- Kitano, J.; Peichel, C.L. Turnover of sex chromosomes and speciation in fishes. Environ. Biol. Fishes 2012, 94, 549–558. [Google Scholar] [CrossRef] [PubMed]
- Adler, P.H.; Yadamsuren, O.; Procunier, W.S. Chromosomal translocations in black flies (Diptera: Simuliidae)—Facilitators of adaptive radiation? PLoS ONE 2016, 11, e0158272. [Google Scholar] [CrossRef] [PubMed]
- Blair, W.F. Mating call in the speciation of anuran amphibians. Am. Nat. 1958, 92, 27–51. [Google Scholar] [CrossRef]
- Zeh, D.W.; Zeh, J.A. Reproductive mode and speciation: The viviparity-driven conflict hypothesis. Bioessays 2000, 22, 938–946. [Google Scholar] [CrossRef]
- Hoskin, C.J.; Higgie, M.; McDonald, K.R.; Moritz, C. Reinforcement drives rapid allopatric speciation. Nature 2005, 437, 1353–1356. [Google Scholar] [CrossRef]
- Tilley, S.G.; Verrell, P.A.; Arnold, S.J. Correspondence between sexual isolation and allozyme differentiation: A test in the salamander Desmognathus ochrophaeus. Proc. Natl. Acad. Sci. USA 1990, 87, 2715–2719. [Google Scholar] [CrossRef]
- Garland, T., Jr.; Adolph, S.C. Physiological differentiation of vertebrate populations. Annu. Rev. Ecol. Syst. 1991, 22, 193–228. [Google Scholar] [CrossRef]
- Rodríguez-Robles, J.A.; de Jesús-Escobar, J.M. Molecular systematics of New World lampropeltinine snakes (Colubridae): Implications for biogeography and evolution of food habits. Biol. J. Linn. Soc. Lond. 1999, 68, 355–385. [Google Scholar] [CrossRef]
- Shaffer, H.B.; McKnight, M.L. The polytypic species revisited: Genetic differentiation and molecular phylogenetics of the tiger salamander Ambystoma tigrinum (Amphibia: Caudata) complex. Evolution 1996, 50, 417–433. [Google Scholar] [CrossRef]
- Wiens, J.J.; Reeder, T.W.; Montes De Oca, A.N. Molecular phylogenetics and evolution of sexual dichromatism among populations of the Yarrow’s spiny lizard (Sceloporus jarrovii). Evolution 1999, 53, 1884–1897. [Google Scholar]
- Malone, J.H.; Fontenot, B.E. Patterns of reproductive isolation in toads. PLoS ONE 2008, 3, e3900. [Google Scholar] [CrossRef]
- Wollenberg, K.C.; Vieites, D.R.; van der Meijden, A.; Glaw, F.; Cannatella, D.C.; Vences, M. Patterns of endemism and species richness in Malagasy cophyline frogs support a key role of mountainous areas for speciation. Evolution 2008, 62, 1890–1907. [Google Scholar] [CrossRef]
- Wollenberg, K.C.; Vieites, D.R.; Glaw, F.; Vences, M. Speciation in little: The role of range and body size in the diversification of Malagasy mantellid frogs. BMC Evol. Biol. 2011, 11, 217. [Google Scholar] [CrossRef]
- Kaffenberger, N.; Wollenberg, K.C.; Köhler, J.; Glaw, F.; Vieites, D.R.; Vences, M. Molecular phylogeny and biogeography of Malagasy frogs of the genus Gephyromantis. Mol. Phylogenet. Evol. 2012, 62, 555–560. [Google Scholar] [CrossRef]
- Wollenberg, K.C.; Wang, I.J.; Glor, R.E.; Losos, J.B. Determinism in the diversification of Hispaniolan trunk-ground anoles (Anolis cybotes species complex). Evolution 2013, 67, 3175–3190. [Google Scholar] [CrossRef]
- Moritz, C.; Schneider, C.J.; Wake, D.B. Evolutionary relationships within the Ensatina eschscholtzii complex confirm the ring species interpretation. Syst. Biol. 1992, 41, 273–291. [Google Scholar] [CrossRef]
- Dettman, J.R.; Sirjusingh, C.; Kohn, L.M.; Anderson, J.B. Incipient speciation by divergent adaptation and antagonistic epistasis in yeast. Nature 2007, 447, 585–588. [Google Scholar] [CrossRef]
- Shafer, A.B.A.; Wolf, J.B.W. Widespread evidence for incipient ecological speciation: A meta-analysis of isolation-by-ecology. Ecol. Lett. 2013, 16, 940–950. [Google Scholar] [CrossRef]
- Rice, W.R.; Hostert, E.E. Laboratory experiments on speciation: What have we learned in 40 years? Evolution 1993, 47, 1637–1653. [Google Scholar] [CrossRef]
- Byrne, K.; Nichols, R.A. Culex pipiens in London Underground tunnels: Differentiation between surface and subterranean populations. Heredity 1999, 82, 7–15. [Google Scholar] [CrossRef]
- Nosil, P.; Funk, D.J.; Ortiz-Barrientos, D. Divergent selection and heterogeneous genomic divergence. Mol. Ecol. 2009, 18, 375–402. [Google Scholar] [CrossRef]
- Camargo, A.; Sinervo, B.; Sites, J.W., Jr. Lizards as model organisms for linking phylogeographic and speciation studies. Mol. Ecol. 2010, 19, 3250–3270. [Google Scholar] [CrossRef]
- Florio, A.M.; Ingram, C.M.; Rakotondravony, H.A.; Louis, E.E.; Raxworthy, C.J. Detecting cryptic speciation in the widespread and morphologically conservative carpet chameleon (Furcifer lateralis) of Madagascar. J. Evol. Biol. 2012, 25, 1399–1414. [Google Scholar] [CrossRef]
- Vasconcelos, R.; Perera, A.; Geniez, P.; Harris, D.J.; Carranza, S. An integrative taxonomic revision of the Tarentola geckos (Squamata, Phyllodactylidae) of the Cape Verde Islands. Zool. J. Linn. Soc. 2012, 164, 328–360. [Google Scholar] [CrossRef]
- Fujita, M.K.; Leaché, A.D.; Burbrink, F.T.; McGuire, J.A.; Moritz, C. Coalescent-based species delimitation in an integrative taxonomy. Trends Ecol. Evol. 2012, 27, 480–488. [Google Scholar] [CrossRef]
- Chan, L.M.; Brown, J.L.; Yoder, A.D. Integrating statistical genetic and geospatial methods brings new power to phylogeography. Mol. Phylogenet. Evol. 2011, 59, 523–537. [Google Scholar] [CrossRef]
- Padial, J.M.; de La Riva, I. A response to recent proposals for integrative taxonomy. Biol. J. Linn. Soc. Lond. 2010, 101, 747–756. [Google Scholar] [CrossRef]
- Padial, J.M.; Miralles, A.; De la Riva, I.; Vences, M. The integrative future of taxonomy. Front. Zool. 2010, 7, 16. [Google Scholar] [CrossRef]
- McKay, B.D.; Mays, H.L., Jr.; Yao, C.-T.; Wan, D.; Higuchi, H.; Nishiumi, I. Incorporating color into integrative taxonomy: Analysis of the varied tit (Sittiparus varius) complex in East Asia. Syst. Biol. 2014, 63, 505–517. [Google Scholar] [CrossRef]
- Pante, E.; Schoelinck, C.; Puillandre, N. From integrative taxonomy to species description: One step beyond. Syst. Biol. 2015, 64, 152–160. [Google Scholar] [CrossRef]
- Schlick-Steiner, B.C.; Steiner, F.M.; Seifert, B.; Stauffer, C.; Christian, E.; Crozier, R.H. Integrative taxonomy: A multisource approach to exploring biodiversity. Annu. Rev. Entomol. 2010, 55, 421–438. [Google Scholar] [CrossRef]
- Yeates, D.K.; Seago, A.; Nelson, L.; Cameron, S.L.; Joseph, L.; Trueman, J.W.H. Integrative taxonomy, or iterative taxonomy? Syst. Entomol. 2011, 36, 209–217. [Google Scholar] [CrossRef]
- Edwards, D.L.; Knowles, L.L. Species detection and individual assignment in species delimitation: Can integrative data increase efficacy? Proc. Biol. Sci. 2014, 281, 20132765. [Google Scholar] [CrossRef]
- Solís-Lemus, C.; Knowles, L.L.; Ané, C. Bayesian species delimitation combining multiple genes and traits in a unified framework. Evolution 2015, 69, 492–507. [Google Scholar] [CrossRef]
- Aguilar, C.; Wood, P.L., Jr.; Belk, M.C.; Duff, M.H.; Sites, J.W., Jr. Different roads lead to Rome: Integrative taxonomic approaches lead to the discovery of two new lizard lineages in the Liolaemus montanus group (Squamata: Liolaemidae). Biol. J. Linn. Soc. Lond. 2016, 120, 448–467. [Google Scholar] [CrossRef][Green Version]
- Leavitt, S.D.; Moreau, C.S.; Thorsten Lumbsch, H. The dynamic discipline of species delimitation: Progress toward effectively recognizing species boundaries in natural populations. In Recent Advances in Lichenology: Modern Methods and Approaches in Lichen Systematics and Culture Techniques; Upreti, D.K., Divakar, P.K., Shukla, V., Bajpai, R., Eds.; Springer India: New Delhi, India, 2015; Volume 2, pp. 11–44. ISBN 9788132222354. [Google Scholar]
- Olave, M.; Martinez, L.E.; Avila, L.J.; Sites, J.W., Jr.; Morando, M. Evidence of hybridization in the Argentinean lizards Liolaemus gracilis and Liolaemus bibronii (Iguania: Liolaemini): An integrative approach based on genes and morphology. Mol. Phylogenet. Evol. 2011, 61, 381–391. [Google Scholar] [CrossRef]
- Faria, R.; Johannesson, K.; Butlin, R.K.; Westram, A.M. Evolving inversions. Trends Ecol. Evol. 2019, 34, 239–248. [Google Scholar] [CrossRef]
- Campbell, C.R.; Poelstra, J.W.; Yoder, A.D. What is Speciation Genomics? The roles of ecology, gene flow, and genomic architecture in the formation of species. Biol. J. Linn. Soc. Lond. 2018, 124, 561–583. [Google Scholar] [CrossRef]
- Frantz, L.A.F.; Schraiber, J.G.; Madsen, O.; Megens, H.-J.; Bosse, M.; Paudel, Y.; Semiadi, G.; Meijaard, E.; Li, N.; Crooijmans, R.P.M.A.; et al. Genome sequencing reveals fine scale diversification and reticulation history during speciation in Sus. Genome Biol. 2013, 14, R107. [Google Scholar] [CrossRef]
- Árnason, E.; Halldórsdóttir, K. Codweb: Whole-genome sequencing uncovers extensive reticulations fueling adaptation among Atlantic, Arctic, and Pacific gadids. Sci. Adv. 2019, 5, eaat8788. [Google Scholar] [CrossRef]
- Solís-Lemus, C.; Bastide, P.; Ané, C. PhyloNetworks: A package for Phylogenetic Networks. Mol. Biol. Evol. 2017, 34, 3292–3298. [Google Scholar] [CrossRef]
- Wen, D.; Yu, Y.; Zhu, J.; Nakhleh, L. Inferring Phylogenetic Networks using PhyloNet. Syst. Biol. 2018, 67, 735–740. [Google Scholar] [CrossRef]
- Zhang, C.; Ogilvie, H.A.; Drummond, A.J.; Stadler, T. Bayesian inference of species networks from multilocus sequence data. Mol. Biol. Evol. 2018, 35, 504–517. [Google Scholar] [CrossRef]
- MacLeod, N.; Benfield, M.; Culverhouse, P. Time to automate identification. Nature 2010, 467, 154–155. [Google Scholar] [CrossRef]
- Wäldchen, J.; Mäder, P. Machine learning for image based species identification. Methods Ecol. Evol. 2018, 9, 2216–2225. [Google Scholar] [CrossRef]
- Botto Nuñez, G.; Lemus, G.; Muñoz Wolf, M.; Rodales, A.L.; González, E.M.; Crisci, C. The first artificial intelligence algorithm for identification of bat species in Uruguay. Ecol. Inform. 2018, 46, 97–102. [Google Scholar] [CrossRef]
- Mitra, R.; Marchitto, T.M.; Ge, Q.; Zhong, B.; Kanakiya, B.; Cook, M.S.; Fehrenbacher, J.S.; Ortiz, J.D.; Tripati, A.; Lobaton, E. Automated species-level identification of planktic foraminifera using convolutional neural networks, with comparison to human performance. Mar. Micropaleontol. 2019, 147, 16–24. [Google Scholar] [CrossRef]
- Xie, J.; Towsey, M.; Zhang, J.; Roe, P. Acoustic classification of Australian frogs based on enhanced features and machine learning algorithms. Appl. Acoust. 2016, 113, 193–201. [Google Scholar] [CrossRef]
- Colonna, J.G.; Gama, J.; Nakamura, E.F. A comparison of hierarchical multi-output recognition approaches for anuran classification. Mach. Learn. 2018, 107, 1651–1671. [Google Scholar] [CrossRef]
- Ceríaco, L.M.P.; Gutiérrez, E.E.; Dubois, A. Photography-based taxonomy is inadequate, unnecessary, and potentially harmful for biological sciences. Zootaxa 2016, 4196, 435–445. [Google Scholar] [CrossRef]
- Wearn, O.R.; Freeman, R.; Jacoby, D.M.P. Responsible AI for conservation. Nat. Mach. Intell. 2019, 1, 72–73. [Google Scholar] [CrossRef]
- Hillis, D.M. Species delimitation in herpetology. J. Herpetol. 2019, 53, 3–13. [Google Scholar] [CrossRef]
- Nicholson, K.E.; Crother, B.I.; Guyer, C.; Savage, J.M. It is time for a new classification of anoles (Squamata: Dactyloidae). Zootaxa 2012, 3477, 1–108. [Google Scholar] [CrossRef]
- Hutchinson, G.E. Concluding remarks. Cold Spring Harb. Symp. Quant. Biol. 1957, 22, 415–427. [Google Scholar] [CrossRef]
- Holt, R.D. Bringing the Hutchinsonian niche into the 21st century: Ecological and evolutionary perspectives. Proc. Natl. Acad. Sci. USA 2009, 106 (Suppl. 2), 19659–19665. [Google Scholar] [CrossRef]
- Moritz, C.; Patton, J.L.; Schneider, C.J.; Smith, T.B. Diversification of rainforest faunas: An integrated molecular approach. Annu. Rev. Ecol. Syst. 2000, 31, 533–563. [Google Scholar] [CrossRef]
- Hua, X.; Wiens, J.J. How does climate influence speciation? Am. Nat. 2013, 182, 1–12. [Google Scholar] [CrossRef]
- Wiens, J.J. Speciation and ecology revisited: Phylogenetic niche conservatism and the origin of species. Evolution 2004, 58, 193–197. [Google Scholar] [CrossRef]
- Wiens, J.J.; Ackerly, D.D.; Allen, A.P.; Anacker, B.L.; Buckley, L.B.; Cornell, H.V.; Damschen, E.I.; Jonathan Davies, T.; Grytnes, J.-A.; Harrison, S.P.; et al. Niche conservatism as an emerging principle in ecology and conservation biology. Ecol. Lett. 2010, 13, 1310–1324. [Google Scholar] [CrossRef]
- Kozak, K.H.; Wiens, J.J. Climatic zonation drives latitudinal variation in speciation mechanisms. Proc. Biol. Sci. 2007, 274, 2995–3003. [Google Scholar] [CrossRef]
- Kozak, K.H.; Wiens, J.J. Accelerated rates of climatic-niche evolution underlie rapid species diversification. Ecol. Lett. 2010, 13, 1378–1389. [Google Scholar] [CrossRef]
- Janzen, D.H. Why mountain passes are higher in the tropics. Am. Nat. 1967, 101, 233–249. [Google Scholar] [CrossRef]
- Ghalambor, C.K.; Huey, R.B.; Martin, P.R.; Tewksbury, J.J.; Wang, G. Are mountain passes higher in the tropics? Janzen’s hypothesis revisited. Integr. Comp. Biol. 2006, 46, 5–17. [Google Scholar] [CrossRef]
- Quintero, I.; Wiens, J.J. What determines the climatic niche width of species? The role of spatial and temporal climatic variation in three vertebrate clades. Glob. Ecol. Biogeogr. 2013, 22, 422–432. [Google Scholar] [CrossRef]
- Cadena, C.D.; Kozak, K.H.; Gómez, J.P.; Parra, J.L.; McCain, C.M.; Bowie, R.C.K.; Carnaval, A.C.; Moritz, C.; Rahbek, C.; Roberts, T.E.; et al. Latitude, elevational climatic zonation and speciation in New World vertebrates. Proc. Biol. Sci. 2012, 279, 194–201. [Google Scholar] [CrossRef]
- Gómez-Rodríguez, C.; Baselga, A.; Wiens, J.J. Is diversification rate related to climatic niche width? Glob. Ecol. Biogeogr. 2015, 24, 383–395. [Google Scholar] [CrossRef]
- Moen, D.S.; Wiens, J.J. Microhabitat and climatic niche change explain patterns of diversification among frog families. Am. Nat. 2017, 190, 29–44. [Google Scholar] [CrossRef]
- Hua, X.; Wiens, J.J. Latitudinal variation in speciation mechanisms in frogs. Evolution 2010, 64, 429–443. [Google Scholar] [CrossRef]
- Knouft, J.H.; Losos, J.B.; Glor, R.E.; Kolbe, J.J. Phylogenetic analysis of the evolution of the niche in lizards of the Anolis sagrei group. Ecology 2006, 87, S29–S38. [Google Scholar] [CrossRef]
- Pyron, A.R.; Burbrink, F.T. Lineage diversification in a widespread species: Roles for niche divergence and conservatism in the common kingsnake, Lampropeltis getula. Mol. Ecol. 2009, 18, 3443–3457. [Google Scholar] [CrossRef]
- Schneider, C.J.; Smith, T.B.; Larison, B.; Moritz, C. A test of alternative models of diversification in tropical rainforests: Ecological gradients vs. rainforest refugia. Proc. Natl. Acad. Sci. USA 1999, 96, 13869–13873. [Google Scholar] [CrossRef]
- Ogden, R.; Thorpe, R.S. Molecular evidence for ecological speciation in tropical habitats. Proc. Natl. Acad. Sci. USA 2002, 99, 13612–13615. [Google Scholar] [CrossRef]
- Muñoz, M.M.; Crawford, N.G.; McGreevy, T.J., Jr.; Messana, N.J.; Tarvin, R.D.; Revell, L.J.; Zandvliet, R.M.; Hopwood, J.M.; Mock, E.; Schneider, A.L.; et al. Divergence in coloration and ecological speciation in the Anolis marmoratus species complex. Mol. Ecol. 2013, 22, 2668–2682. [Google Scholar] [CrossRef]
- Kozak, K.H.; Wiens, J.J. Does niche conservatism promote speciation? A case study in North American salamanders. Evolution 2006, 60, 2604–2621. [Google Scholar] [CrossRef]
- Kozak, K.H.; Wiens, J.J. Niche conservatism drives elevational diversity patterns in Appalachian salamanders. Am. Nat. 2010, 176, 40–54. [Google Scholar] [CrossRef]
- Hoskin, C.J.; Tonione, M.; Higgie, M.; Mackenzie, J.B.; Williams, S.E.; Vanderwal, J.; Moritz, C. Persistence in peripheral refugia promotes phenotypic divergence and speciation in a rainforest frog. Am. Nat. 2011, 178, 561–578. [Google Scholar] [CrossRef]
- Jezkova, T.; Wiens, J.J. Testing the role of climate in speciation: New methods and applications to squamate reptiles (lizards and snakes). Mol. Ecol. 2018, 27, 2754–2769. [Google Scholar] [CrossRef]
- Todd Streelman, J.; Danley, P.D. The stages of vertebrate evolutionary radiation. Trends Ecol. Evol. 2003, 18, 126–131. [Google Scholar] [CrossRef]
- Adams, D.C.; Berns, C.M.; Kozak, K.H.; Wiens, J.J. Are rates of species diversification correlated with rates of morphological evolution? Proc. Biol. Sci. 2009, 276, 2729–2738. [Google Scholar] [CrossRef]
- Wiens, J.J. Explaining large-scale patterns of vertebrate diversity. Biol. Lett. 2015, 11, 20150506. [Google Scholar] [CrossRef]
- Bars-Closel, M.; Kohlsdorf, T.; Moen, D.S.; Wiens, J.J. Diversification rates are more strongly related to microhabitat than climate in squamate reptiles (lizards and snakes). Evolution 2017, 71, 2243–2261. [Google Scholar] [CrossRef]
- Cyriac, V.P.; Kodandaramaiah, U. Digging their own macroevolutionary grave: Fossoriality as an evolutionary dead end in snakes. J. Evol. Biol. 2018, 31, 587–598. [Google Scholar] [CrossRef]
- Coyne, J.A.; Allen Orr, H. Speciation; Sinauer: Sunderland, MA, USA, 2004; ISBN 9780878930890. [Google Scholar]
- Stephens, P.R.; Wiens, J.J. Explaining species richness from continents to communities: The time-for-speciation effect in emydid turtles. Am. Nat. 2003, 161, 112–128. [Google Scholar] [CrossRef]
- Schluter, D. Ecology and the origin of species. Trends Ecol. Evol. 2001, 16, 372–380. [Google Scholar] [CrossRef]
- Dieckmann, U.; Doebeli, M.; Metz, J.A.J.; Tautz, D. Adaptive Speciation; Cambridge University Press: Cambridge, UK, 2004; ISBN 9780521828420. [Google Scholar]
- Hendry, A.P. Ecological speciation! Or the lack thereof? Can. J. Fish. Aquat. Sci. 2009, 66, 1383–1398. [Google Scholar] [CrossRef]
- Nosil, P. Ecological Speciation; Oxford University Press: Oxford, UK, 2012; ISBN 9780199587100. [Google Scholar]
- Thorpe, R.S.; Surget-Groba, Y.; Johansson, H. Genetic tests for ecological and allopatric speciation in anoles on an island archipelago. PLoS Genet. 2010, 6, e1000929. [Google Scholar] [CrossRef]
- Thorpe, R.S.; Surget-Groba, Y.; Johansson, H. The relative importance of ecology and geographic isolation for speciation in anoles. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2008, 363, 3071–3081. [Google Scholar] [CrossRef]
- Schluter, D. Ecological causes of adaptive radiation. Am. Nat. 1996, 148, S40–S64. [Google Scholar] [CrossRef]
- Grant, P.R.; Rosemary Grant, B. 40 Years of Evolution: Darwin’s Finches on Daphne Major Island; Princeton University Press: Princeton, NJ, USA, 2014; ISBN 9781400851300. [Google Scholar]
- McKinnon, J.S.; Mori, S.; Blackman, B.K.; David, L.; Kingsley, D.M.; Jamieson, L.; Chou, J.; Schluter, D. Evidence for ecology’s role in speciation. Nature 2004, 429, 294–298. [Google Scholar] [CrossRef]
- Vines, T.H.; Schluter, D. Strong assortative mating between allopatric sticklebacks as a by-product of adaptation to different environments. Proc. Biol. Sci. 2006, 273, 911–916. [Google Scholar] [CrossRef]
- Tautz, D. Phylogeography and patterns of incipient speciation. In Adaptive Speciation; Dieckmann, U., Doebeli, M., Metz, J.A.J., Tautz, D., Eds.; Cambridge University Press: Cambridge, UK, 2004; pp. 305–321. ISBN 1139379534. [Google Scholar]
- Lamichhaney, S.; Berglund, J.; Almén, M.S.; Maqbool, K.; Grabherr, M.; Martinez-Barrio, A.; Promerová, M.; Rubin, C.-J.; Wang, C.; Zamani, N.; et al. Evolution of Darwin’s finches and their beaks revealed by genome sequencing. Nature 2015, 518, 371–375. [Google Scholar] [CrossRef]
- Schliewen, U.K.; Klee, B. Reticulate sympatric speciation in Cameroonian crater lake cichlids. Front. Zool. 2004, 1, 5. [Google Scholar] [CrossRef]
- Ricklefs, R.E.; Losos, J.B.; Townsend, T.M. Evolutionary diversification of clades of squamate reptiles. J. Evol. Biol. 2007, 20, 1751–1762. [Google Scholar] [CrossRef]
- Hedges, S.B.; Marin, J.; Suleski, M.; Paymer, M.; Kumar, S. Tree of life reveals clock-like speciation and diversification. Mol. Biol. Evol. 2015, 32, 835–845. [Google Scholar] [CrossRef]
- Greenberg, D.A.; Mooers, A.Ø. Linking speciation to extinction: Diversification raises contemporary extinction risk in amphibians. Evol. Lett. 2017, 1, 40–48. [Google Scholar] [CrossRef]
- Scholl, J.P.; Wiens, J.J. Diversification rates and species richness across the Tree of Life. Proc. Biol. Sci. 2016, 283, 20161334. [Google Scholar] [CrossRef]
- Butlin, R.; Debelle, A.; Kerth, C.; Snook, R.R.; Beukeboom, L.W.; Castillo Cajas, R.F.; Diao, W.; Maan, M.E.; Paolucci, S.; Weissing, F.J.; et al. What do we need to know about speciation? Trends Ecol. Evol. 2012, 27, 27–39. [Google Scholar]
- Mooers, A.O.; Heard, S.B. Inferring evolutionary process from phylogenetic tree shape. Q. Rev. Biol. 1997, 72, 31–54. [Google Scholar] [CrossRef]
- FitzJohn, R.G. Diversitree: Comparative phylogenetic analyses of diversification in R. Methods Ecol. Evol. 2012, 3, 1084–1092. [Google Scholar] [CrossRef]
- Jezkova, T.; Wiens, J.J. What explains patterns of diversification and richness among animal phyla? Am. Nat. 2017, 189, 201–212. [Google Scholar] [CrossRef]
- Takezaki, N. Global rate variation in Bony Vertebrates. Genome Biol. Evol. 2018, 10, 1803–1815. [Google Scholar] [CrossRef]
- Meiri, S. Evolution and ecology of lizard body sizes. Glob. Ecol. Biogeogr. 2008, 17, 724–734. [Google Scholar] [CrossRef]
- Bambach, R.K. Ecospace utilization and guilds in marine communities through the Phanerozoic. In Biotic Interactions in Recent and Fossil Benthic Communities; Tevesz, M.J.S., McCall, P.L., Eds.; Springer US: Boston, MA, USA, 1983; pp. 719–746. ISBN 9781475707403. [Google Scholar]
- Gomez-Mestre, I.; Pyron, R.A.; Wiens, J.J. Phylogenetic analyses reveal unexpected patterns in the evolution of reproductive modes in frogs. Evolution 2012, 66, 3687–3700. [Google Scholar] [CrossRef]
- Zimkus, B.M.; Lawson, L.; Loader, S.P.; Hanken, J. Terrestrialization, miniaturization and rates of diversification in African puddle frogs (Anura: Phrynobatrachidae). PLoS ONE 2012, 7, e35118. [Google Scholar] [CrossRef]
- Gonzalez-Voyer, A.; Padial, J.M.; Castroviejo-Fisher, S.; De La Riva, I.; Vilà, C. Correlates of species richness in the largest Neotropical amphibian radiation. J. Evol. Biol. 2011, 24, 931–942. [Google Scholar] [CrossRef]
- Van Bocxlaer, I.; Loader, S.P.; Roelants, K.; Biju, S.D.; Menegon, M.; Bossuyt, F. Gradual adaptation toward a range-expansion phenotype initiated the global radiation of toads. Science 2010, 327, 679–682. [Google Scholar] [CrossRef]
- Pizzatto, L.; Dubey, S. Colour-polymorphic snake species are older. Biol. J. Linn. Soc. Lond. 2012, 107, 210–218. [Google Scholar] [CrossRef]
- West-Eberhard, M.J. Alternative adaptations, speciation, and phylogeny (a review). Proc. Natl. Acad. Sci. USA 1986, 83, 1388–1392. [Google Scholar] [CrossRef]
- Forsman, A.; Ahnesjö, J.; Caesar, S.; Karlsson, M. A model of ecological and evolutionary consequences of color polymorphism. Ecology 2008, 89, 34–40. [Google Scholar] [CrossRef]
- Hugall, A.F.; Stuart-Fox, D. Accelerated speciation in colour-polymorphic birds. Nature 2012, 485, 631–634. [Google Scholar] [CrossRef]
- Bolton, P.E.; Rollins, L.A.; Griffith, S.C. The danger within: The role of genetic, behavioural and ecological factors in population persistence of colour polymorphic species. Mol. Ecol. 2015, 24, 2907–2915. [Google Scholar] [CrossRef]
- West-Eberhard, M.J. Sexual selection, social competition, and speciation. Q. Rev. Biol. 1983, 58, 155–183. [Google Scholar] [CrossRef]
- Ritchie, M.G. Sexual selection and speciation. Annu. Rev. Ecol. Evol. Syst. 2007, 38, 79–102. [Google Scholar] [CrossRef]
- Gray, S.M.; McKinnon, J.S. Linking color polymorphism maintenance and speciation. Trends Ecol. Evol. 2007, 22, 71–79. [Google Scholar] [CrossRef]
- Corl, A.; Lancaster, L.T.; Sinervo, B. Rapid formation of reproductive isolation between two populations of side-blotched lizards, Uta stansburiana. Copeia 2012, 2012, 593–602. [Google Scholar] [CrossRef]
- Kolora, S.R.R.; Weigert, A.; Saffari, A.; Kehr, S.; Walter Costa, M.B.; Spröer, C.; Indrischek, H.; Chintalapati, M.; Lohse, K.; Doose, G.; et al. Divergent evolution in the genomes of closely related lacertids, Lacerta viridis and L. bilineata, and implications for speciation. Gigascience 2019, 8, giy160. [Google Scholar] [CrossRef]
- Rykena, S. Kreuzungsexperimente zur Prüfung der Artgrenzen im Genus Lacerta sensu stricto. Mitt. Mus. Naturk. Berlin 1991, 67, 55–68. [Google Scholar] [CrossRef]
- Rykena, S. Experimental hybridization in green lizards (Lacerta s. str.), a tool to study species boundaries. Mertensiella 2001, 13, 78–88. [Google Scholar]
- Andersson, M. Sexual Selection; Princeton University Press: Princeton, NJ, USA, 1994; ISBN 9780691000572. [Google Scholar]
- Sinervo, B.; Zamudio, K.R. The evolution of alternative reproductive strategies: Fitness differential, heritability, and genetic correlation between the sexes. J. Hered. 2001, 92, 198–205. [Google Scholar] [CrossRef]
- Olsson, M.; Healey, M.; Wapstra, E.; Schwartz, T.; Lebas, N.; Uller, T. Mating system variation and morph fluctuations in a polymorphic lizard. Mol. Ecol. 2007, 16, 5307–5315. [Google Scholar] [CrossRef]
- Huyghe, K.; Small, M.; Vanhooydonck, B.; Herrel, A.; Tadić, Z.; Van Damme, R.; Backeljau, T. Genetic divergence among sympatric colour morphs of the Dalmatian wall lizard (Podarcis melisellensis). Genetica 2010, 138, 387–393. [Google Scholar] [CrossRef]
- Andrade, P.; Pinho, C.; Pérez I de Lanuza, G.; Afonso, S.; Brejcha, J.; Rubin, C.-J.; Wallerman, O.; Pereira, P.; Sabatino, S.J.; Bellati, A.; et al. Regulatory changes in pterin and carotenoid genes underlie balanced color polymorphisms in the wall lizard. Proc. Natl. Acad. Sci. USA 2019, 116, 5633–5642. [Google Scholar] [CrossRef]
- Haisten, D.C.; Paranjpe, D.; Loveridge, S.; Sinervo, B. The cellular basis of polymorphic coloration in common side-blotched lizards, Uta stansburiana. Herpetologica 2015, 71, 125–135. [Google Scholar] [CrossRef]
- San-Jose, L.M.; Granado-Lorencio, F.; Sinervo, B.; Fitze, P.S. Iridophores and not carotenoids account for chromatic variation of carotenoid-based coloration in common lizards (Lacerta vivipara). Am. Nat. 2013, 181, 396–409. [Google Scholar] [CrossRef]
- Rand, M.S. Courtship and aggressive behaviour in male lizards exhibiting 2 different sexual colorations. Proc. Am. Zool. 1988, 28, A153. [Google Scholar]
- Thompson, C.W.; Moore, M.C. Throat colour reliably signals status in male tree lizards, Urosaurus ornatus. Anim. Behav. 1991, 42, 745–753. [Google Scholar] [CrossRef]
- Sinervo, B.; Lively, C.M. The rock–paper–scissors game and the evolution of alternative male strategies. Nature 1996, 380, 240–243. [Google Scholar] [CrossRef]
- Bastiaans, E.; Morinaga, G.; Castañeda Gaytán, J.G.; Marshall, J.C.; Sinervo, B. Male aggression varies with throat color in 2 distinct populations of the mesquite lizard. Behav. Ecol. 2013, 24, 968–981. [Google Scholar] [CrossRef]
- Vercken, E.; Massot, M.; Sinervo, B.; Clobert, J. Colour variation and alternative reproductive strategies in females of the common lizard Lacerta vivipara. J. Evol. Biol. 2007, 20, 221–232. [Google Scholar] [CrossRef]
- Huyghe, K.; Vanhooydonck, B.; Herrel, A.; Tadic, Z.; Van Damme, R. Morphology, performance, behavior and ecology of three color morphs in males of the lizard Podarcis melisellensis. Integr. Comp. Biol. 2007, 47, 211–220. [Google Scholar] [CrossRef]
- Vignoli, L.; Vuerich, V.; Bologna, M.A. Experimental study of dispersal behaviour in a wall lizard species (Podarcis sicula) (Sauria Lacertidae). Ethol. Ecol. Evol. 2012, 24, 244–256. [Google Scholar] [CrossRef]
- Healey, M.; Uller, T.; Olsson, M. Seeing red: Morph-specific contest success and survival rates in a colour-polymorphic agamid lizard. Anim. Behav. 2007, 74, 337–341. [Google Scholar] [CrossRef]
- Yewers, M.S.C.; Pryke, S.; Stuart-Fox, D. Behavioural differences across contexts may indicate morph-specific strategies in the lizard Ctenophorus decresii. Anim. Behav. 2016, 111, 329–339. [Google Scholar] [CrossRef]
- Sinervo, B.; Chaine, A.; Clobert, J.; Calsbeek, R.; Hazard, L.; Lancaster, L.; McAdam, A.G.; Alonzo, S.; Corrigan, G.; Hochberg, M.E. Self-recognition, color signals, and cycles of greenbeard mutualism and altruism. Proc. Natl. Acad. Sci. USA 2006, 103, 7372–7377. [Google Scholar] [CrossRef]
- Vercken, E.; Clobert, J. Ventral colour polymorphism correlates with alternative behavioural patterns in female common lizards (Lacerta vivipara). Écoscience 2008, 15, 320–326. [Google Scholar] [CrossRef]
- Vercken, E.; Clobert, J. The role of colour polymorphism in social encounters among female common lizards. Herpetol. J. 2008, 18, 223–230. [Google Scholar]
- Zajitschek, S.R.K.; Zajitschek, F.; Miles, D.B.; Clobert, J. The effect of coloration and temperature on sprint performance in male and female wall lizards. Biol. J. Linn. Soc. Lond. 2012, 107, 573–582. [Google Scholar] [CrossRef]
- Zamudio, K.R.; Sinervo, B. Polygyny, mate-guarding, and posthumous fertilization as alternative male mating strategies. Proc. Natl. Acad. Sci. USA 2000, 97, 14427–14432. [Google Scholar] [CrossRef]
- Sinervo, B. Runaway social games, genetic cycles driven by alternative male and female strategies, and the origin of morphs. In Microevolution Rate, Pattern, Process; Hendry, A.P., Kinnison, M.T., Eds.; Springer Netherlands: Dordrecht, The Netherlands, 2001; pp. 417–434. ISBN 9789401005852. [Google Scholar]
- Bleay, C.; Sinervo, B. Discrete genetic variation in mate choice and a condition-dependent preference function in the side-blotched lizard: Implications for the formation and maintenance of coadapted gene complexes. Behav. Ecol. 2007, 18, 304–310. [Google Scholar] [CrossRef][Green Version]
- Lancaster, L.T.; McAdam, A.G.; Wingfield, J.C.; Sinervo, B.R. Adaptive social and maternal induction of antipredator dorsal patterns in a lizard with alternative social strategies. Ecol. Lett. 2007, 10, 798–808. [Google Scholar] [CrossRef]
- Lancaster, L.T.; Hipsley, C.A.; Sinervo, B. Female choice for optimal combinations of multiple male display traits increases offspring survival. Behav. Ecol. 2009, 20, 993–999. [Google Scholar] [CrossRef][Green Version]
- Lancaster, L.T.; McAdam, A.G.; Sinervo, B. Maternal adjustment of egg size organizes alternative escape behaviors, promoting adaptive phenotypic integration. Evolution 2010, 64, 1607–1621. [Google Scholar] [CrossRef]
- Pérez i de Lanuza, G.; Font, E.; Carazo, P. Color-assortative mating in a color-polymorphic lacertid lizard. Behav. Ecol. 2013, 24, 273–279. [Google Scholar] [CrossRef]
- Lattanzio, M.S.; Metro, K.J.; Miles, D.B. Preference for male traits differ in two female morphs of the tree lizard, Urosaurus ornatus. PLoS ONE 2014, 9, e101515. [Google Scholar] [CrossRef]
- Wellenreuther, M.; Svensson, E.I.; Hansson, B. Sexual selection and genetic colour polymorphisms in animals. Mol. Ecol. 2014, 23, 5398–5414. [Google Scholar] [CrossRef]
- Cox, C.L.; Chippindale, P.T. Patterns of genetic diversity in the polymorphic ground snake (Sonora semiannulata). Genetica 2014, 142, 361–370. [Google Scholar] [CrossRef]
- Lattanzio, M.S.; Miles, D.B. Ecological divergence among colour morphs mediated by changes in spatial network structure associated with disturbance. J. Anim. Ecol. 2014, 83, 1490–1500. [Google Scholar] [CrossRef]
- McLean, C.A.; Stuart-Fox, D.; Moussalli, A. Environment, but not genetic divergence, influences geographic variation in colour morph frequencies in a lizard. BMC Evol. Biol. 2015, 15, 156. [Google Scholar] [CrossRef]
- Runemark, A.; Hansson, B.; Pafilis, P.; Valakos, E.D.; Svensson, E.I. Island biology and morphological divergence of the Skyros wall lizard Podarcis gaigeae: A combined role for local selection and genetic drift on color morph frequency divergence? BMC Evol. Biol. 2010, 10, 269. [Google Scholar] [CrossRef]
- Feldman, C.R.; Flores-Villela, O.; Papenfuss, T.J. Phylogeny, biogeography, and display evolution in the tree and brush lizard genus Urosaurus (Squamata: Phrynosomatidae). Mol. Phylogenet. Evol. 2011, 61, 714–725. [Google Scholar] [CrossRef]
- Hews, D.K.; Thompson, C.W.; Moore, I.T.; Moore, M.C. Population frequencies of alternative male phenotypes in tree lizards: Geographic variation and common-garden rearing studies. Behav. Ecol. Sociobiol. 1997, 41, 371–380. [Google Scholar] [CrossRef]
- Sacchi, R.; Scali, S.; Pupin, F.; Gentilli, A.; Galeotti, P.; Fasola, M. Microgeographic variation of colour morph frequency and biometry of common wall lizards. J. Zool. 2007, 273, 389–396. [Google Scholar] [CrossRef]
- Corl, A.; Davis, A.R.; Kuchta, S.R.; Sinervo, B. Selective loss of polymorphic mating types is associated with rapid phenotypic evolution during morphic speciation. Proc. Natl. Acad. Sci. USA 2010, 107, 4254–4259. [Google Scholar] [CrossRef]
- McLean, C.A.; Stuart-Fox, D. Geographic variation in animal colour polymorphisms and its role in speciation. Biol. Rev. Camb. Philos. Soc. 2014, 89, 860–873. [Google Scholar] [CrossRef]
- Friedman, D.; Magnani, J.; Paranjpe, D.; Sinervo, B. Evolutionary games, climate and the generation of diversity. PLoS ONE 2017, 12, e0184052. [Google Scholar] [CrossRef]
- Corl, A.; Davis, A.R.; Kuchta, S.R.; Comendant, T.; Sinervo, B. Alternative mating strategies and the evolution of sexual size dimorphism in the side-blotched lizard, Uta stansburiana: A population-level comparative analysis. Evolution 2010, 64, 79–96. [Google Scholar] [CrossRef]
- Alonzo, S.H.; Sinervo, B. Mate choice games, context-dependent good genes, and genetic cycles in the side-blotched lizard, Uta stansburiana. Behav. Ecol. Sociobiol. 2001, 49, 176–186. [Google Scholar] [CrossRef]
- McLean, C.A.; Stuart-Fox, D.; Moussalli, A. Phylogeographic structure, demographic history and morph composition in a colour polymorphic lizard. J. Evol. Biol. 2014, 27, 2123–2137. [Google Scholar] [CrossRef]
- Bastiaans, E.; Bastiaans, M.J.; Morinaga, G.; Castañeda Gaytán, J.G.; Marshall, J.C.; Bane, B.; de la Cruz, F.M.; Sinervo, B. Female preference for sympatric vs. allopatric male throat color morphs in the mesquite lizard (Sceloporus grammicus) species complex. PLoS ONE 2014, 9, e93197. [Google Scholar] [CrossRef]
- Ryan, M.J.; Rand, A.S. Species recognition and sexual selection as a unitary problem in animal communication. Evolution 1993, 47, 647–657. [Google Scholar] [CrossRef]
- Boul, K.E.; Funk, W.C.; Darst, C.R.; Cannatella, D.C.; Ryan, M.J. Sexual selection drives speciation in an Amazonian frog. Proc. Biol. Sci. 2007, 274, 399–406. [Google Scholar] [CrossRef]
- Rice, A.M.; Pfennig, D.W. Does character displacement initiate speciation? Evidence of reduced gene flow between populations experiencing divergent selection. J. Evol. Biol. 2010, 23, 854–865. [Google Scholar] [CrossRef]
- Pfennig, K.S.; Rice, A.M. Reinforcement generates reproductive isolation between neighbouring conspecific populations of spadefoot toads. Proc. R. Soc. B Biol. Sci. 2014, 281, 20140949. [Google Scholar] [CrossRef]
- Lemmon, E.M. Diversification of conspecific signals in sympatry: Geographic overlap drives multidimensional reproductive character displacement in frogs. Evolution 2009, 63, 1155–1170. [Google Scholar] [CrossRef]
- Lemmon, E.M.; Juenger, T.E. Geographic variation in hybridization across a reinforcement contact zone of chorus frogs (Pseudacris). Ecol. Evol. 2017, 7, 9485–9502. [Google Scholar] [CrossRef]
- Mayr, E. Populations, Species, and Evolution: An Abridgment of Animal Species and Evolution, 1st ed.; Harvard University Press: Cambridge, MA, USA, 1970; ISBN 9780674690134. [Google Scholar]
- Pyron, R.A.; Burbrink, F.T. Hard and soft allopatry: Physically and ecologically mediated modes of geographic speciation. J. Biogeogr. 2010, 37, 2005–2015. [Google Scholar]
- Wollenberg, K.C.; Harvey, J. First assessment of the male territorial vocal behaviour of a Malagasy leaf litter frog (Gephyromantis thelenae). Herpetol. Notes 2010, 3, 141–150. [Google Scholar]
- Wollenberg Valero, K.C. Evidence for an intrinsic factor promoting landscape genetic divergence in Madagascan leaf-litter frogs. Front. Genet. 2015, 6, 155. [Google Scholar] [CrossRef]
- Schaefer, H.-C.; Vences, M.; Veith, M. Molecular phylogeny of Malagasy poison frogs, genus Mantella (Anura: Mantellidae): Homoplastic evolution of colour pattern in aposematic amphibians. Org. Divers. Evol. 2002, 2, 97–105. [Google Scholar] [CrossRef]
- Brown, J.L.; Cameron, A.; Yoder, A.D.; Vences, M. A necessarily complex model to explain the biogeography of the amphibians and reptiles of Madagascar. Nat. Commun. 2014, 5, 5046. [Google Scholar] [CrossRef]
- Hohenlohe, P.A.; Bassham, S.; Etter, P.D.; Stiffler, N.; Johnson, E.A.; Cresko, W.A. Population genomics of parallel adaptation in threespine stickleback using sequenced RAD tags. PLoS Genet. 2010, 6, e1000862. [Google Scholar] [CrossRef]
- Daub, J.T.; Hofer, T.; Cutivet, E.; Dupanloup, I.; Quintana-Murci, L.; Robinson-Rechavi, M.; Excoffier, L. Evidence for polygenic adaptation to pathogens in the human genome. Mol. Biol. Evol. 2013, 30, 1544–1558. [Google Scholar] [CrossRef]
- Via, S. Sympatric speciation in animals: The ugly duckling grows up. Trends Ecol. Evol. 2001, 16, 381–390. [Google Scholar] [CrossRef]
- Butlin, R.K.; Galindo, J.; Grahame, J.W. Review. Sympatric, parapatric or allopatric: The most important way to classify speciation? Philos. Trans. R. Soc. Lond. B Biol. Sci. 2008, 363, 2997–3007. [Google Scholar] [CrossRef]
- Fitzpatrick, B.M.; Fordyce, J.A.; Gavrilets, S. What, if anything, is sympatric speciation? J. Evol. Biol. 2008, 21, 1452–1459. [Google Scholar] [CrossRef]
- Fitzpatrick, B.M.; Fordyce, J.A.; Gavrilets, S. Pattern, process and geographic modes of speciation. J. Evol. Biol. 2009, 22, 2342–2347. [Google Scholar] [CrossRef]
- Mallet, J.; Meyer, A.; Nosil, P.; Feder, J.L. Space, sympatry and speciation. J. Evol. Biol. 2009, 22, 2332–2341. [Google Scholar] [CrossRef]
- Feder, J.L.; Flaxman, S.M.; Egan, S.P.; Comeault, A.A.; Nosil, P. Geographic mode of speciation and genomic divergence. Annu. Rev. Ecol. Evol. Syst. 2013, 44, 73–97. [Google Scholar] [CrossRef]
- Gavrilets, S. Fitness Landscapes and the Origin of Species (MPB-41); Princeton University Press: Princeton, NJ, USA, 2004; ISBN 9780691119830. [Google Scholar]
- Bolnick, D.I.; Fitzpatrick, B.M. Sympatric speciation: Models and empirical evidence. Annu. Rev. Ecol. Evol. Syst. 2007, 38, 459–487. [Google Scholar] [CrossRef]
- Van Doorn, G.S.; Edelaar, P.; Weissing, F.J. On the origin of species by natural and sexual selection. Science 2009, 326, 1704–1707. [Google Scholar] [CrossRef]
- Flaxman, S.M.; Wacholder, A.C.; Feder, J.L.; Nosil, P. Theoretical models of the influence of genomic architecture on the dynamics of speciation. Mol. Ecol. 2014, 23, 4074–4088. [Google Scholar] [CrossRef]
- Nosil, P. Speciation with gene flow could be common. Mol. Ecol. 2008, 17, 2103–2106. [Google Scholar] [CrossRef]
- Feder, J.L.; Egan, S.P.; Nosil, P. The genomics of speciation-with-gene-flow. Trends Genet. 2012, 28, 342–350. [Google Scholar] [CrossRef]
- Wu, C.-I.; Ting, C.-T. Genes and speciation. Nat. Rev. Genet. 2004, 5, 114–122. [Google Scholar] [CrossRef]
- Via, S.; West, J. The genetic mosaic suggests a new role for hitchhiking in ecological speciation. Mol. Ecol. 2008, 17, 4334–4345. [Google Scholar] [CrossRef]
- Via, S. Divergence hitchhiking and the spread of genomic isolation during ecological speciation-with-gene-flow. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2012, 367, 451–460. [Google Scholar] [CrossRef]
- Ellegren, H.; Smeds, L.; Burri, R.; Olason, P.I.; Backström, N.; Kawakami, T.; Künstner, A.; Mäkinen, H.; Nadachowska-Brzyska, K.; Qvarnström, A.; et al. The genomic landscape of species divergence in Ficedula flycatchers. Nature 2012, 491, 756–760. [Google Scholar] [CrossRef]
- Nosil, P.; Gompert, Z.; Farkas, T.E.; Comeault, A.A.; Feder, J.L.; Buerkle, C.A.; Parchman, T.L. Genomic consequences of multiple speciation processes in a stick insect. Proc. Biol. Sci. 2012, 279, 5058–5065. [Google Scholar] [CrossRef]
- Guarnizo, C.E.; Cannatella, D.C. Geographic determinants of gene flow in two sister species of tropical Andean frogs. J. Hered. 2014, 105, 216–225. [Google Scholar] [CrossRef]
- Richter-Boix, A.; Teplitsky, C.; Rogell, B.; Laurila, A. Local selection modifies phenotypic divergence among Rana temporaria populations in the presence of gene flow. Mol. Ecol. 2010, 19, 716–731. [Google Scholar] [CrossRef]
- Richter-Boix, A.; Quintela, M.; Kierczak, M.; Franch, M.; Laurila, A. Fine-grained adaptive divergence in an amphibian: Genetic basis of phenotypic divergence and the role of nonrandom gene flow in restricting effective migration among wetlands. Mol. Ecol. 2013, 22, 1322–1340. [Google Scholar] [CrossRef]
- Lind, M.I.; Ingvarsson, P.K.; Johansson, H.; Hall, D.; Johansson, F. Gene flow and selection on phenotypic plasticity in an island system of Rana temporaria. Evolution 2011, 65, 684–697. [Google Scholar] [CrossRef]
- Van Buskirk, J. Incipient habitat race formation in an amphibian. J. Evol. Biol. 2014, 27, 585–592. [Google Scholar] [CrossRef]
- Streicher, J.W.; Devitt, T.J.; Goldberg, C.S.; Malone, J.H.; Blackmon, H.; Fujita, M.K. Diversification and asymmetrical gene flow across time and space: Lineage sorting and hybridization in polytypic barking frogs. Mol. Ecol. 2014, 23, 3273–3291. [Google Scholar] [CrossRef]
- Funk, W.C.; Murphy, M.A.; Hoke, K.L.; Muths, E.; Amburgey, S.M.; Lemmon, E.M.; Lemmon, A.R. Elevational speciation in action? Restricted gene flow associated with adaptive divergence across an altitudinal gradient. J. Evol. Biol. 2016, 29, 241–252. [Google Scholar] [CrossRef]
- Nadachowska, K.; Babik, W. Divergence in the face of gene flow: The case of two newts (Amphibia: Salamandridae). Mol. Biol. Evol. 2009, 26, 829–841. [Google Scholar] [CrossRef]
- Pereira, R.J.; Martínez-Solano, I.; Buckley, D. Hybridization during altitudinal range shifts: Nuclear introgression leads to extensive cyto-nuclear discordance in the fire salamander. Mol. Ecol. 2016, 25, 1551–1565. [Google Scholar] [CrossRef]
- Niemiller, M.L.; Fitzpatrick, B.M.; Miller, B.T. Recent divergence with gene flow in Tennessee cave salamanders (Plethodontidae: Gyrinophilus) inferred from gene genealogies. Mol. Ecol. 2008, 17, 2258–2275. [Google Scholar] [CrossRef]
- Stenson, A.G.; Malhotra, A.; Thorpe, R.S. Population differentiation and nuclear gene flow in the Dominican anole (Anolis oculatus). Mol. Ecol. 2002, 11, 1679–1688. [Google Scholar] [CrossRef]
- Calsbeek, R.; Smith, T.B.; Bardeleben, C. Intraspecific variation in Anolis sagrei mirrors the adaptive radiation of Greater Antillean anoles. Biol. J. Linn. Soc. 2007, 90, 189–199. [Google Scholar] [CrossRef]
- Pinho, C.; Harris, D.J.; Ferrand, N. Non-equilibrium estimates of gene flow inferred from nuclear genealogies suggest that Iberian and North African wall lizards (Podarcis spp.) are an assemblage of incipient species. BMC Evol. Biol. 2008, 8, 63. [Google Scholar] [CrossRef]
- Rosenblum, E.B.; Hickerson, M.J.; Moritz, C. A multilocus perspective on colonization accompanied by selection and gene flow. Evolution 2007, 61, 2971–2985. [Google Scholar] [CrossRef]
- Leaché, A.D. Multi-locus estimates of population structure and migration in a fence lizard hybrid zone. PLoS ONE 2011, 6, e25827. [Google Scholar] [CrossRef]
- Leache, A.D.; Harris, R.B.; Maliska, M.E.; Linkem, C.W. Comparative species divergence across eight triplets of spiny lizards (Sceloporus) using genomic sequence data. Genome Biol. Evol. 2013, 5, 2410–2419. [Google Scholar] [CrossRef][Green Version]
- Leaché, A.D.; Palacios, J.A.; Minin, V.N.; Bryson, R.W. Phylogeography of the Trans-Volcanic bunchgrass lizard (Sceloporus bicanthalis) across the highlands of south-eastern Mexico. Biol. J. Linn. Soc. 2013, 110, 852–865. [Google Scholar] [CrossRef]
- Grummer, J.A.; Calderón-Espinosa, M.L.; Nieto-Montes de Oca, A.; Smith, E.N.; Méndez-de la Cruz, F.R.; Leaché, A.D. Estimating the temporal and spatial extent of gene flow among sympatric lizard populations (genus Sceloporus) in the southern Mexican highlands. Mol. Ecol. 2015, 24, 1523–1542. [Google Scholar] [CrossRef]
- Oliveira, E.F.; Gehara, M.; São-Pedro, V.A.; Chen, X.; Myers, E.A.; Burbrink, F.T.; Mesquita, D.O.; Garda, A.A.; Colli, G.R.; Rodrigues, M.T.; et al. Speciation with gene flow in whiptail lizards from a Neotropical xeric biome. Mol. Ecol. 2015, 24, 5957–5975. [Google Scholar] [CrossRef]
- Schield, D.R.; Card, D.C.; Adams, R.H.; Jezkova, T.; Reyes-Velasco, J.; Proctor, F.N.; Spencer, C.L.; Herrmann, H.-W.; Mackessy, S.P.; Castoe, T.A. Incipient speciation with biased gene flow between two lineages of the Western Diamondback Rattlesnake (Crotalus atrox). Mol. Phylogenet. Evol. 2015, 83, 213–223. [Google Scholar] [CrossRef]
- Gibbs, H.L.; Corey, S.J.; Blouin-Demers, G.; Prior, K.A.; Weatherhead, P.J. Hybridization between mtDNA-defined phylogeographic lineages of black ratsnakes (Pantherophis sp.). Mol. Ecol. 2006, 15, 3755–3767. [Google Scholar] [CrossRef]
- Fitzpatrick, B.M.; Placyk, J.S., Jr.; Niemiller, M.L.; Casper, G.S.; Burghardt, G.M. Distinctiveness in the face of gene flow: Hybridization between specialist and generalist gartersnakes. Mol. Ecol. 2008, 17, 4107–4117. [Google Scholar] [CrossRef]
- Placyk, J.S.; Fitzpatrick, B.M.; Casper, G.S.; Small, R.L.; Graham Reynolds, R.; Noble, D.W.A.; Brooks, R.J.; Burghardt, G.M. Hybridization between two gartersnake species (Thamnophis) of conservation concern: A threat or an important natural interaction? Conserv. Genet. 2012, 13, 649–663. [Google Scholar] [CrossRef]
- Nadachowska, K. Divergence with gene flow—The amphibian perspective. Herpetol. J. 2010, 20, 7–15. [Google Scholar]
- Riesch, R.; Muschick, M.; Lindtke, D.; Villoutreix, R.; Comeault, A.A.; Farkas, T.E.; Lucek, K.; Hellen, E.; Soria-Carrasco, V.; Dennis, S.R.; et al. Transitions between phases of genomic differentiation during stick-insect speciation. Nat. Ecol. Evol. 2017, 1, 82. [Google Scholar] [CrossRef]
- Rhymer, J.M.; Simberloff, D. Extinction by hybridization and introgression. Annu. Rev. Ecol. Syst. 1996, 27, 83–109. [Google Scholar] [CrossRef]
- Austin, J.D.; Gorman, T.A.; Bishop, D.; Moler, P. Genetic evidence of contemporary hybridization in one of North America’s rarest anurans, the Florida bog frog. Anim. Conserv. 2011, 14, 553–561. [Google Scholar] [CrossRef]
- Macculloch, R.D.; Murphy, R.W.; Kupriyanova, L.A.; Darevsky, I.S. The Caucasian rock lizard Lacerta rostombekovi: A monoclonal parthenogenetic vertebrate. Biochem. Syst. Ecol. 1997, 25, 33–37. [Google Scholar] [CrossRef]
- Ryskov, A.P.; Osipov, F.A.; Omelchenko, A.V.; Semyenova, S.K.; Girnyk, A.E.; Korchagin, V.I.; Vergun, A.A.; Murphy, R.W. The origin of multiple clones in the parthenogenetic lizard species Darevskia rostombekowi. PLoS ONE 2017, 12, e0185161. [Google Scholar] [CrossRef]
- Sinclair, E.A.; Pramuk, J.B.; Bezy, R.L.; Crandall, K.A.; Sites, J.W., Jr. DNA evidence for nonhybrid origins of parthenogenesis in natural populations of vertebrates. Evolution 2010, 64, 1346–1357. [Google Scholar] [CrossRef]
- Rogers, R.L.; Zhou, L.; Chu, C.; Márquez, R.; Corl, A.; Linderoth, T.; Freeborn, L.; MacManes, M.D.; Xiong, Z.; Zheng, J.; et al. Genomic takeover by Transposable Elements in the Strawberry Poison Frog. Mol. Biol. Evol. 2018, 35, 2913–2927. [Google Scholar] [CrossRef]
- Bourgeois, Y.; Boissinot, S. On the population dynamics of junk: A review on the Population Genomics of Transposable Elements. Genes 2019, 10, 419. [Google Scholar] [CrossRef]
- Ricci, M.; Peona, V.; Guichard, E.; Taccioli, C.; Boattini, A. Transposable Elements activity is positively related to rate of speciation in mammals. J. Mol. Evol. 2018, 86, 303–310. [Google Scholar] [CrossRef]
- Feiner, N. Accumulation of transposable elements in Hox gene clusters during adaptive radiation of Anolis lizards. Proc. Biol. Sci. 2016, 283, 20161555. [Google Scholar] [CrossRef]
- Grant, B.R.; Grant, P.R. Fission and fusion of Darwin’s finches populations. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2008, 363, 2821–2829. [Google Scholar] [CrossRef]
- Geist, D.J.; Snell, H.; Snell, H.; Goddard, C.; Kurz, M.D. A paleogeographic model of the Galápagos Islands and biogeographical and evolutionary implications. In The Galápagos: A Natural Laboratory for the Earth Sciences; American Geophysical Union: Washington, DC, USA, 2014; pp. 145–166. [Google Scholar]
- Wagner, W.L.; Funk, V.A. Hawaiian Biogeography: Evolution on a Hot Spot Archipelago; Smithsonian Institution Press: Washington, DC, USA, 1995; ISBN 9781560984627. [Google Scholar]
- Pritchard, P.C.H. The Galápagos Tortoises: Nomenclatural and Survival Status; Chelonian Research Foundation: Lunenburg, MA, USA, 1996; ISBN 9780965354004. [Google Scholar]
- Abdala, C.S. Phylogeny of the boulengeri group (Iguania: Liolaemidae, Liolaemus) based on morphological and molecular characters. Zootaxa 2007, 1538, 1–84. [Google Scholar] [CrossRef]
- Lobo, F.; Espinoza, R.E.; Quinteros, S. A critical review and systematic discussion of recent classification proposals for liolaemid lizards. Zootaxa 2010, 2549, 1–30. [Google Scholar] [CrossRef]
- Olave, M.A.; González-Marín, L.J.; Avila, J.W.; Sites, M., Jr. Morando Disparate patterns of diversification within Liolaemini lizards. In Neotropical Diversification; Rull, V., Carnaval, A.C., Eds.; Springer Nature: Berlin, Germany, in press.
- Gómez, J.M.D.; Lobo, F. Historical Biogeography of a clade of Liolaemus (Iguania: Liolaemidae) based on ancestral areas and dispersal-vicariance analysis (DIVA). Pap. Avulsos Zool. 2006, 46. [Google Scholar] [CrossRef]
- Schulte, J.A.; Macey, J.R.; Espinoza, R.E.; Larson, A. Phylogenetic relationships in the iguanid lizard genus Liolaemus: Multiple origins of viviparous reproduction and evidence for recurring Andean vicariance and dispersal. Biol. J. Linn. Soc. 2000, 69, 75–102. [Google Scholar] [CrossRef]
- Morando, M.; Avila, L.J.; Sites, J.W. Sampling strategies for delimiting species: Genes, individuals, and populations in the Liolaemus elongatus-kriegi complex (Squamata: Liolaemidae) in Andean–Patagonian South America. Syst. Biol. 2003, 52, 159–185. [Google Scholar] [CrossRef]
- Minoli, I.; Morando, M.; Avila, L.J. Reptiles of Chubut province, Argentina: Richness, diversity, conservation status and geographic distribution maps. Zookeys 2015, 498, 103–126. [Google Scholar] [CrossRef]
- Breitman, M.F.; Minoli, I.; Avila, L.J.; Medina, C.D.; Sites, J.W., Jr.; Morando, M. Lagartijas de la provincia de Santa Cruz, Argentina: Distribución geográfica, diversidad genética y estado de conservación. Cuad. Herpetol. 2014, 28, 83–110. [Google Scholar]
- Medina, C.D.; Avila, L.J.; Sites, J.W.; Morando, M. Multilocus phylogeography of the Patagonian lizard complex Liolaemus kriegi (Iguania: Liolaemini). Biol. J. Linn. Soc. 2014, 113, 256–269. [Google Scholar] [CrossRef]
- Medina, C.D.; Avila, L.J.; Sites, J.W., Jr.; Morando, M. Phylogeographic history of Patagonian lizards of the Liolaemus elongatus complex (Iguania: Liolaemini) based on mitochondrial and nuclear DNA sequences. J. Zool. Syst. Evol. Res. 2017, 55, 238–249. [Google Scholar] [CrossRef]
- Aguilar, C.; Wood, P.; Cusi, J.C.; Guzman, A.; Huari, F.; Lundberg, M.; Mortensen, E.; Ramirez, C.; Robles, D.; Suarez, J.; et al. Integrative taxonomy and preliminary assessment of species limits in the Liolaemus walkeri complex (Squamata, Liolaemidae) with descriptions of three new species from Peru. ZooKeys 2013, 364, 47–91. [Google Scholar] [CrossRef]
- Minoli, I.; Morando, M.; Avila, L.J. Integrative taxonomy in the Liolaemus fitzingerii complex (Squamata: Liolaemini) based on morphological analyses and niche modeling. Zootaxa 2014, 3856, 501–528. [Google Scholar] [CrossRef]
- Breitman, M.F.; Martinez, R.J.N.; Avila, L.J.; Sites, J.W.; Morando, M. Phylogeography and morphological variation of the northernmost distributed species of the Liolaemus lineomaculatus section (Liolaemini) from Patagonia. Amphib-Reptilia 2015, 36, 373–387. [Google Scholar] [CrossRef]
- Breitman, M.F.; Bonino, M.F.; Sites, J.W.; Avila, L.J.; Morando, M. Morphological variation, niche divergence, and phylogeography of lizards of the Liolaemus lineomaculatus section (Liolaemini) from Southern Patagonia. Herp. Monogr. 2015, 29, 65–88. [Google Scholar] [CrossRef]
- Olave, M.; Avila, L.J.; Sites, J.W.; Morando, M. Hidden diversity within the lizard genus Liolaemus: Genetic vs. morphological divergence in the L. rothi complex (Squamata: Liolaeminae). Mol. Phylogenet. Evol. 2017, 107, 56–63. [Google Scholar] [CrossRef]
- Merilä, J.; Sheldon, B.C.; Kruuk, L.E. Explaining stasis: Microevolutionary studies in natural populations. Genetica 2001, 112–113, 199–222. [Google Scholar] [CrossRef]
- Estes, S.; Arnold, S.J. Resolving the paradox of stasis: Models with stabilizing selection explain evolutionary divergence on all timescales. Am. Nat. 2007, 169, 227–244. [Google Scholar] [CrossRef]
- Futuyma, D.J. Evolutionary constraint and ecological consequences. Evolution 2010, 64, 1865–1884. [Google Scholar] [CrossRef]
- Morando, M.; Avila, L.J.; Baker, J.; Sites, J.W., Jr. Phylogeny and phylogeography of the Liolaemus darwinii complex (Squamata: Liolaemidae): Evidence for introgression and incomplete lineage sorting. Evolution 2004, 58, 842–861. [Google Scholar] [CrossRef]
- Breitman, M.F.; Avila, L.J.; Sites, J.W., Jr.; Morando, M. Lizards from the end of the world: Phylogenetic relationships of the Liolaemus lineomaculatus section (Squamata: Iguania: Liolaemini). Mol. Phylogenet. Evol. 2011, 59, 364–376. [Google Scholar] [CrossRef]
- Morando, M.; Avila, L.J.; Perez, C.H.F.; Hawkins, M.A.; Sites, J.W. A molecular phylogeny of the lizard genus Phymaturus (Squamata, Liolaemini): Implications for species diversity and historical biogeography of southern South America. Mol. Phylogenet. Evol. 2013, 66, 694–714. [Google Scholar] [CrossRef]
- Rabosky, D.L.; Donnellan, S.C.; Talaba, A.L.; Lovette, I.J. Exceptional among-lineage variation in diversification rates during the radiation of Australia’s most diverse vertebrate clade. Proc. Biol. Sci. 2007, 274, 2915–2923. [Google Scholar] [CrossRef]
- Rabosky, D.L.; Slater, G.J.; Alfaro, M.E. Clade age and species richness are decoupled across the eukaryotic tree of life. PLoS Biol. 2012, 10, e1001381. [Google Scholar] [CrossRef]
- Kozak, K.H.; Wiens, J.J. Testing the relationships between diversification, species richness, and trait evolution. Syst. Biol. 2016, 65, 975–988. [Google Scholar] [CrossRef]
- Olave, M.; Avila, L.J.; Sites, J.W.; Morando, M. Model-based approach to test hard polytomies in the Eulaemus clade of the most diverse South American lizard genus Liolaemus (Liolaemini, Squamata). Zool. J. Linn. Soc. 2015, 174, 169–184. [Google Scholar] [CrossRef]
- Olave, M.; Avila, L.J.; Sites, J.W.; Morando, M. How important is it to consider lineage diversification heterogeneity in macroevolutionary studies: Lessons from the lizard family Liolaemidae. bioRxiv 2019. [Google Scholar] [CrossRef]
- Pincheira-Donoso, D.; Harvey, L.P.; Ruta, M. What defines an adaptive radiation? Macroevolutionary diversification dynamics of an exceptionally species-rich continental lizard radiation. BMC Evol. Biol. 2015, 15, 153. [Google Scholar] [CrossRef]
- Esquerré, D.; Brennan, I.G.; Catullo, R.A.; Torres-Pérez, F.; Scott Keogh, J. How mountains shape biodiversity: The role of the Andes in biogeography, diversification, and reproductive biology in South America’s most species-rich lizard radiation (Squamata: Liolaemidae). Evolution 2019, 73, 214–230. [Google Scholar] [CrossRef]
- Hennig, C.; Hausdorf, B. Prabclus: Functions for clustering of presence-absence, abundance and multilocus genetic data. R Package Vers. 2010, 2, 2. [Google Scholar]
- Avila, L.J.; Morando, M.; Sites, J.W. Congeneric phylogeography: Hypothesizing species limits and evolutionary processes in Patagonian lizards of the Liolaemus boulengeri group (Squamata: Liolaemini). Biol. J. Linn. Soc. 2006, 89, 241–275. [Google Scholar] [CrossRef]
- Breitman, M.F.; Florencia Breitman, M.; Avila, L.J.; Sites, J.W.; Morando, M. How lizards survived blizzards: Phylogeography of the Liolaemus lineomaculatus group (Liolaemidae) reveals multiple breaks and refugia in southern Patagonia and their concordance with other codistributed taxa. Mol. Ecol. 2012, 21, 6068–6085. [Google Scholar] [CrossRef]
- Morando, M.; Avila, L.J.; Turner, C.R.; Sites, J.W. Molecular evidence for a species complex in the patagonian lizard Liolaemus bibronii and phylogeography of the closely related Liolaemus gracilis (Squamata: Liolaemini). Mol. Phylogenet. Evol. 2007, 43, 952–973. [Google Scholar] [CrossRef]
- Camargo, A.; Morando, M.; Avila, L.J.; Sites, J.W., Jr. Species delimitation using ABC: Accounting for speciation with gene flow in lizards of the Liolaemus darwinii complex (Squamata: Liolaemidae). Evolution 2012, 66, 2834–2849. [Google Scholar] [CrossRef]
- Olave, M.; Avila, L.J.; Sites, J.W., Jr.; Morando, M. Hybridization could be a common phenomenon within the highly diverse lizard genus Liolaemus. J. Evol. Biol. 2018, 31, 893–903. [Google Scholar] [CrossRef]
- Grummer, J.A.; Morando, M.M.; Avila, L.J.; Sites, J.W., Jr.; Leaché, A.D. Phylogenomic evidence for a recent and rapid radiation of lizards in the Patagonian Liolaemus fitzingerii species group. Mol. Phylogenet. Evol. 2018, 125, 243–254. [Google Scholar] [CrossRef]
- Sukumaran, J.; Knowles, L.L. Multispecies coalescent delimits structure, not species. Proc. Natl. Acad. Sci. USA 2017, 114, 1607–1612. [Google Scholar] [CrossRef]
- Leaché, A.D.; Zhu, T.; Rannala, B.; Yang, Z. The spectre of too many species. Syst. Biol. 2019, 68, 168–181. [Google Scholar] [CrossRef]
- Heath, T.A.; Zwickl, D.J.; Kim, J.; Hillis, D.M. Taxon sampling affects inferences of macroevolutionary processes from phylogenetic trees. Syst. Biol. 2008, 57, 160–166. [Google Scholar] [CrossRef]
- Venditti, C.; Meade, A.; Pagel, M. Detecting the node-density artifact in phylogeny reconstruction. Syst. Biol. 2006, 55, 637–643. [Google Scholar] [CrossRef]
- Webster, A.J.; Payne, R.J.H.; Pagel, M. Molecular phylogenies link rates of evolution and speciation. Science 2003, 301, 478. [Google Scholar] [CrossRef]
- Venditti, C.; Meade, A.; Pagel, M. Phylogenetic mixture models can reduce node-density artifacts. Syst. Biol. 2008, 57, 286–293. [Google Scholar] [CrossRef]
- Fontanella, F.M.; Feltrin, N.; Avila, L.J.; Sites, J.W.; Morando, M. Early stages of divergence: Phylogeography, climate modeling, and morphological differentiation in the South American lizard Liolaemus petrophilus (Squamata: Liolaemidae). Ecol. Evol. 2012, 2, 792–808. [Google Scholar] [CrossRef]
- Edwards, D.L.; Melville, J.; Joseph, L.; Keogh, J.S. Ecological divergence, adaptive diversification, and the evolution of social signaling traits: An empirical study in arid Australian lizards. Am. Nat. 2015, 186, E144–E161. [Google Scholar] [CrossRef]
- Grummer, J.A. Evolutionary History of the Patagonian Liolaemus fitzingerii Species Group of Lizards. Ph.D. Thesis, University of Washington, Seattle, WA, USA, 2017. [Google Scholar]
- Hey, J.; Chung, Y.; Sethuraman, A.; Lachance, J.; Tishkoff, S.; Sousa, V.C.; Wang, Y. Phylogeny estimation by integration over isolation with migration models. Mol. Biol. Evol. 2018, 35, 2805–2818. [Google Scholar] [CrossRef]
- Abdala, V.; Tulli, M.J.; Russell, A.P.; Powell, G.L.; Cruz, F.B. Anatomy of the crus and pes of neotropical iguanian lizards in relation to habitat use and digitally based grasping capabilities. Anat. Rec. 2014, 297, 397–409. [Google Scholar] [CrossRef]
- Medina, C.D.; Avila, L.J.; Sites, J.W.; Morando, M. Molecular phylogeny of the Liolaemus kriegi complex (Iguania, Liolaemini). Herpetologica 2015, 71, 143–151. [Google Scholar] [CrossRef]
- Martínez, L.E. Métodos Empíricos Para Delimitar Especies: El Complejo Liolaemus bibronii (Squamata: Liolaemini) Como Ejemplo. Unpublished Ph.D. Thesis, Universidad Nacional de Córdoba, Córdoba, Argentina, 2012. [Google Scholar]
- Bell, R.C.; MacKenzie, J.B.; Hickerson, M.J.; Chavarría, K.L.; Cunningham, M.; Williams, S.; Moritz, C. Comparative multi-locus phylogeography confirms multiple vicariance events in co-distributed rainforest frogs. Proc. Biol. Sci. 2012, 279, 991–999. [Google Scholar] [CrossRef]
- Fujita, M.K.; McGuire, J.A.; Donnellan, S.C.; Moritz, C. Diversification and persistence at the arid-monsoonal interface: Australia-wide biogeography of the Bynoe’s gecko (Heteronotia binoei; Gekkonidae). Evolution 2010, 64, 2293–2314. [Google Scholar] [CrossRef]
- Pepper, M.; Fujita, M.K.; Moritz, C.; Keogh, J.S. Palaeoclimate change drove diversification among isolated mountain refugia in the Australian arid zone. Mol. Ecol. 2011, 20, 1529–1545. [Google Scholar] [CrossRef]
- Werneck, F.P.; Gamble, T.; Colli, G.R.; Rodrigues, M.T.; Sites, J.W., Jr. Deep diversification and long-term persistence in the South American “Dry diagonal”: Integrating continent-wide phylogeography and distribution modeling of geckos. Evolution 2012, 66, 3014–3034. [Google Scholar] [CrossRef]
- Nogueira, C.; Ribeiro, S.; Costa, G.C.; Colli, G.R. Vicariance and endemism in a Neotropical savanna hotspot: Distribution patterns of Cerrado squamate reptiles. J. Biogeogr. 2011, 38, 1907–1922. [Google Scholar] [CrossRef]
- Werneck, F.P. The diversification of eastern South American open vegetation biomes: Historical biogeography and perspectives. Quat. Sci. Rev. 2011, 30, 1630–1648. [Google Scholar] [CrossRef]
- Valdujo, P.H.; Silvano, D.L.; Colli, G.; Martins, M. Anuran species composition and distribution patterns in Brazilian Cerrado, a neotropical hotspot. S. Am. J. Herpetol. 2012, 7, 63–78. [Google Scholar] [CrossRef]
- Recoder, R.S.; De Pinho Werneck, F.; Teixeira, M.; Colli, G.R.; Sites, J.W.; Rodrigues, M.T. Geographic variation and systematic review of the lizard genus Vanzosaura (Squamata, Gymnophthalmidae), with the description of a new species. Zool. J. Linn. Soc. 2014, 171, 206–225. [Google Scholar] [CrossRef]
- Werneck, F.P.; Leite, R.N.; Geurgas, S.R.; Rodrigues, M.T. Biogeographic history and cryptic diversity of saxicolous Tropiduridae lizards endemic to the semiarid Caatinga. BMC Evol. Biol. 2015, 15, 94. [Google Scholar] [CrossRef]
- Prado, C.P.A.; Haddad, C.F.B.; Zamudio, K.R. Cryptic lineages and Pleistocene population expansion in a Brazilian Cerrado frog. Mol. Ecol. 2012, 21, 921–941. [Google Scholar] [CrossRef]
- Machado, T.; Silva, V.X.; Silva, M.J. Phylogenetic relationships within Bothrops neuwiedi group (Serpentes, Squamata): Geographically highly-structured lineages, evidence of introgressive hybridization and Neogene/Quaternary diversification. Mol. Phylogenet. Evol. 2014, 71, 1–14. [Google Scholar] [CrossRef]
- Santos, M.G.; Nogueira, C.; Giugliano, L.G.; Colli, G.R. Landscape evolution and phylogeography of Micrablepharus atticolus (Squamata, Gymnophthalmidae), an endemic lizard of the Brazilian Cerrado. J. Biogeogr. 2014, 41, 1506–1519. [Google Scholar] [CrossRef]
- Guarnizo, C.E.; Werneck, F.P.; Giugliano, L.G.; Santos, M.G.; Fenker, J.; Sousa, L.; D’Angiolella, A.B.; dos Santos, A.R.; Strüssmann, C.; Rodrigues, M.T.; et al. Cryptic lineages and diversification of an endemic anole lizard (Squamata, Dactyloidae) of the Cerrado hotspot. Mol. Phylogenet. Evol. 2016, 94, 279–289. [Google Scholar] [CrossRef]
- Thomé, M.T.C.; Sequeira, F.; Brusquetti, F.; Carstens, B.; Haddad, C.F.B.; Rodrigues, M.T.; Alexandrino, J. Recurrent connections between Amazon and Atlantic forests shaped diversity in Caatinga four-eyed frogs. J. Biogeogr. 2016, 43, 1045–1056. [Google Scholar] [CrossRef]
- Davis, M.B.; Shaw, R.G.; Etterson, J.R. Evolutionary responses to changing climate. Ecology 2005, 86, 1704–1714. [Google Scholar] [CrossRef]
- Steinfartz, S.; Veith, M.; Tautz, D. Mitochondrial sequence analysis of Salamandra taxa suggests old splits of major lineages and postglacial recolonizations of central Europe from distinct source populations of Salamandra salamandra. Mol. Ecol. 2000, 9, 397–410. [Google Scholar] [CrossRef]
- Vences, M.; Sanchez, E.; Susanne Hauswaldt, J.; Eikelmann, D.; Rodríguez, A.; Carranza, S.; Donaire, D.; Gehara, M.; Helfer, V.; Lötters, S.; et al. Nuclear and mitochondrial multilocus phylogeny and survey of alkaloid content in true salamanders of the genus Salamandra (Salamandridae). Mol. Phylogenet. Evol. 2014, 73, 208–216. [Google Scholar] [CrossRef]
- Eiselt, J. Der Feuersalamander Salamandra salamandra (L.): Beiträge zu Einer Taxonomischen Synthese; Abhandlungen und Berichte für Naturkunde und Vorgeschichte; Museum für Kulturgeschichte: Magdeburg, Germany, 1958; pp. 1–78. [Google Scholar]
- Thiesmeier, B. Salamandra salamandra (Linnaeus, 1758)—Feuersalamander. In Handbuch der Reptilien und Amphibien Europas Schwanzlurche IIB; Thiesmeier, B., Grossenbacher, K., Eds.; Aula: Wiebelsheim, Germany, 2004; pp. 1059–1132. ISBN 389104674X. [Google Scholar]
- Weitere, M.; Tautz, D.; Neumann, D.; Steinfartz, S. Adaptive divergence vs. environmental plasticity: Tracing local genetic adaptation of metamorphosis traits in salamanders. Mol. Ecol. 2004, 13, 1665–1677. [Google Scholar] [CrossRef]
- Reinhardt, T.; Steinfartz, S.; Paetzold, A.; Weitere, M. Linking the evolution of habitat choice to ecosystem functioning: Direct and indirect effects of pond-reproducing fire salamanders on aquatic-terrestrial subsidies. Oecologia 2013, 173, 281–291. [Google Scholar] [CrossRef]
- Reinhardt, T. New Home, New Life: The Influence of Shifts in Fire-Salamander Larval Habitat Choice on Population Perfomance and Their Effect on Structure and Functioning of Pond Invertebrate Communities. Ph.D. Thesis, Technische Universität Dresden, Dresden, Germany, 2014. [Google Scholar]
- Steinfartz, S.; Weitere, M.; Tautz, D. Tracing the first step to speciation: Ecological and genetic differentiation of a salamander population in a small forest. Mol. Ecol. 2007, 16, 4550–4561. [Google Scholar] [CrossRef]
- Schmidt, B.R.; Schaub, M.; Steinfartz, S. Apparent survival of the salamander Salamandra salamandra is low because of high migratory activity. Front. Zool. 2007, 4, 19. [Google Scholar] [CrossRef]
- Schulte, U.; Küsters, D.; Steinfartz, S. A PIT tag based analysis of annual movement patterns of adult fire salamanders (Salamandra salamandra) in a Middle European habitat. Amphibia Reptilia 2007, 28, 531–536. [Google Scholar] [CrossRef]
- Steinfartz, S.; Caspers, B.A. Preference for the other sex: Olfactory sex recognition in terrestrial fire salamanders (Salamandra salamandra). Amphib. Reptil. 2011, 32, 503–508. [Google Scholar] [CrossRef]
- Caspers, B.A.; Steinfartz, S.; Tobias Krause, E. Larval deposition behaviour and maternal investment of females reflect differential habitat adaptation in a genetically diverging salamander population. Behav. Ecol. Sociobiol. 2015, 69, 407–413. [Google Scholar] [CrossRef]
- Hendrix, R.; Schmidt, B.R.; Schaub, M.; Krause, E.T.; Steinfartz, S. Differentiation of movement behaviour in an adaptively diverging salamander population. Mol. Ecol. 2017, 26, 6400–6413. [Google Scholar] [CrossRef]
- Czypionka, T.; Krugman, T.; Altmüller, J.; Blaustein, L.; Steinfartz, S.; Templeton, A.R.; Nolte, A.W. Ecological transcriptomics—A non-lethal sampling approach for endangered fire salamanders. Methods Ecol. Evol. 2015, 6, 1417–1425. [Google Scholar] [CrossRef]
- Goedbloed, D.J.; Czypionka, T.; Altmüller, J.; Rodriguez, A.; Küpfer, E.; Segev, O.; Blaustein, L.; Templeton, A.R.; Nolte, A.W.; Steinfartz, S. Parallel habitat acclimatization is realized by the expression of different genes in two closely related salamander species (genus Salamandra). Heredity 2017, 119, 429–437. [Google Scholar] [CrossRef]
- Czypionka, T.; Goedbloed, D.J.; Steinfartz, S.; Nolte, A.W. Plasticity and evolutionary divergence in gene expression associated with alternative habitat use in larvae of the European Fire Salamander. Mol. Ecol. 2018, 27, 2698–2713. [Google Scholar] [CrossRef]
- Nowoshilow, S.; Schloissnig, S.; Fei, J.-F.; Dahl, A.; Pang, A.W.C.; Pippel, M.; Winkler, S.; Hastie, A.R.; Young, G.; Roscito, J.G.; et al. The axolotl genome and the evolution of key tissue formation regulators. Nature 2018, 554, 50–55. [Google Scholar] [CrossRef]
- Wollenberg, K.C.; Glaw, F.; Meyer, A.; Vences, M. Molecular phylogeny of Malagasy reed frogs, Heterixalus, and the relative performance of bioacoustics and color-patterns for resolving their systematics. Mol. Phylogenet. Evol. 2007, 45, 14–22. [Google Scholar] [CrossRef]
- Glaw, F.; Vences, M. A Field Guide to the Amphibians and Reptiles of Madagascar; Vences & Glaw: Cologne, Germany, 2007; ISBN 392944903X. [Google Scholar]
- Allnutt, T.F.; Ferrier, S.; Manion, G.; Powell, G.V.N.; Ricketts, T.H.; Fisher, B.L.; Harper, G.J.; Irwin, M.E.; Kremen, C.; Labat, J.-N.; et al. A method for quantifying biodiversity loss and its application to a 50-year record of deforestation across Madagascar. Cons. Lett. 2008, 1, 173–181. [Google Scholar] [CrossRef]
- Brown, J.L.; Sillero, N.; Glaw, F.; Bora, P.; Vieites, D.R.; Vences, M. Spatial Biodiversity patterns of Madagascar’s Amphibians and Reptiles. PLoS ONE 2016, 11, e0144076. [Google Scholar] [CrossRef]
- Vences, M.; Wollenberg, K.C.; Vieites, D.R.; Lees, D.C. Madagascar as a model region of species diversification. Trends Ecol. Evol. 2009, 24, 456–465. [Google Scholar] [CrossRef]
- Turelli, M.; Barton, N.H.; Coyne, J.A. Theory and speciation. Trends Ecol. Evol. 2001, 16, 330–343. [Google Scholar] [CrossRef]
- De Queiroz, K. Species concepts and species delimitation. Syst. Biol. 2007, 56, 879–886. [Google Scholar] [CrossRef]
- Vellend, M. Species diversity and genetic diversity: Parallel processes and correlated patterns. Am. Nat. 2005, 166, 199–215. [Google Scholar] [CrossRef]
- Vellend, M.; Geber, M.A. Connections between species diversity and genetic diversity. Ecol. Lett. 2005, 8, 767–781. [Google Scholar] [CrossRef]
- Pabijan, M.; Wollenberg, K.C.; Vences, M. Small body size increases the regional differentiation of populations of tropical mantellid frogs (Anura: Mantellidae). J. Evol. Biol. 2012, 25, 2310–2324. [Google Scholar] [CrossRef]
- Vences, M.; Köhler, J.; Crottini, A.; Glaw, F. High mitochondrial sequence divergence meets morphological and bioacoustic conservatism: Boophis quasiboehmei sp. n., a new cryptic treefrog species from south-eastern Madagascar. Bonn Zool. Bull. 2010, 57, 241–255. [Google Scholar]
- Martin, R.A. Body size in (mostly) mammals: Mass, speciation rates and the translation of gamma to alpha diversity on evolutionary timescales. Hist. Biol. 2017, 29, 576–593. [Google Scholar] [CrossRef]
- Feldman, A.; Sabath, N.; Pyron, R.A.; Mayrose, I.; Meiri, S. Body sizes and diversification rates of lizards, snakes, amphisbaenians and the tuatara. Glob. Ecol. Biogeogr. 2016, 25, 187–197. [Google Scholar] [CrossRef]
- Owens, I.P.F.; Bennett, P.M.; Harvey, P.H. Species richness among birds: Body size, life history, sexual selection or ecology? Proc. R. Soc. Lond. B Biol. Sci. 1999, 266, 933–939. [Google Scholar] [CrossRef]
- Sodhi, N.S.; Bickford, D.; Diesmos, A.C.; Lee, T.M.; Koh, L.P.; Brook, B.W.; Sekercioglu, C.H.; Bradshaw, C.J.A. Measuring the meltdown: Drivers of global amphibian extinction and decline. PLoS ONE 2008, 3, e1636. [Google Scholar] [CrossRef]
- Kisel, Y.; Barraclough, T.G. Speciation has a spatial scale that depends on levels of gene flow. Am. Nat. 2010, 175, 316–334. [Google Scholar] [CrossRef]
- Claramunt, S.; Derryberry, E.P.; Remsen, J.V.; Brumfield, R.T. High dispersal ability inhibits speciation in a continental radiation of passerine birds. Proc. R. Soc. B Biol. Sci. 2012, 279, 1567–1574. [Google Scholar] [CrossRef]
- Etienne, R.S.; Olff, H. How dispersal limitation shapes species–body size distributions in local communities. Am. Nat. 2004, 163, 69–83. [Google Scholar] [CrossRef]
- Price, J.P.; Wagner, W.L. Speciation in Hawaiian angiosperm lineages: Cause, consequence, and mode. Evolution 2004, 58, 2185–2200. [Google Scholar] [CrossRef]
- Agnarsson, I.; Kuntner, M. The generation of a biodiversity hotspot: Biogeography and phylogeography of the western Indian Ocean islands. In Current Topics in Phylogenetics and Phylogeography of Terrestrial and Aquatic Systems; IntechOpen: London, UK, 2012; pp. 32–82. ISBN 9789535102175. [Google Scholar]
- Wake, D.B. What salamanders have taught us about evolution. Annu. Rev. Ecol. Evol. Syst. 2009, 40, 333–352. [Google Scholar] [CrossRef]
- Rovito, S.M.; Parra-Olea, G.; Hanken, J.; Bonett, R.M.; Wake, D.B. Adaptive radiation in miniature: The minute salamanders of the Mexican highlands (Amphibia: Plethodontidae: Thorius). Biol. J. Linn. Soc. 2013, 109, 622–643. [Google Scholar] [CrossRef]
- Wells, K.D. The Ecology and Behavior of Amphibians; The University of Chicago Press: Chicago, IL, USA, 2007; p. 1400. ISBN 978-0226893341. [Google Scholar]
- Rodríguez, A.; Börner, M.; Pabijan, M.; Gehara, M.; Haddad, C.F.B.; Vences, M. Genetic divergence in tropical anurans: Deeper phylogeographic structure in forest specialists and in topographically complex regions. Evol. Ecol. 2015, 29, 765–785. [Google Scholar] [CrossRef]
- Guarnizo, C.E.; Cannatella, D.C. Genetic divergence within frog species is greater in topographically more complex regions. J. Zool. Syst. Evol. Res. 2013, 51, 333–340. [Google Scholar] [CrossRef]
- Hutter, C.R.; Lambert, S.M.; Wiens, J.J. Rapid diversification and time explain amphibian richness at different scales in the tropical Andes, earth’s most biodiverse hotspot. Am. Nat. 2017, 190, 828–843. [Google Scholar] [CrossRef]
- Paz, A.; Ibáñez, R.; Lips, K.R.; Crawford, A.J. Testing the role of ecology and life history in structuring genetic variation across a landscape: A trait-based phylogeographic approach. Mol. Ecol. 2015, 24, 3723–3737. [Google Scholar] [CrossRef]
- Fouquet, A.; Ledoux, J.-B.; Dubut, V.; Noonan, B.P.; Scotti, I. The interplay of dispersal limitation, rivers, and historical events shapes the genetic structure of an Amazonian frog. Biol. J. Linn. Soc. 2012, 106, 356–373. [Google Scholar] [CrossRef]
- Santos, J.C. Fast molecular evolution associated with high active metabolic rates in poison frogs. Mol. Biol. Evol. 2012, 29, 2001–2018. [Google Scholar] [CrossRef]
- Chong, R.A.; Mueller, R.L. Low metabolic rates in salamanders are correlated with weak selective constraints on mitochondrial genes. Evolution 2013, 67, 894–899. [Google Scholar] [CrossRef]
- Eo, S.H.; DeWoody, J.A. Evolutionary rates of mitochondrial genomes correspond to diversification rates and to contemporary species richness in birds and reptiles. Proc. Biol. Sci. 2010, 277, 3587–3592. [Google Scholar] [CrossRef]
- Chong, R.A.; Mueller, R.L. Evolution along the mutation gradient in the dynamic mitochondrial genome of salamanders. Genome Biol. Evol. 2013, 5, 1652–1660. [Google Scholar] [CrossRef]
- Bogart, J.P. The influence of life history on karyotypic evolution in frogs. In Amphibian Cytogenetics and Evolution; Academic Press: Cambridge, MA, USA, 1991; pp. 233–258. ISBN 978-0-12-297880-7. [Google Scholar]
- Schmid, M.; Steinlein, C.; Bogart, J.P.; Feichtinger, W.; León, P.; La Marca, E.; Diaz, L.M.; Sanz, A.; -H- Chen, S.; Hedges, S.B. The chromosomes of Terraranan frogs. Insights into vertebrate cytogenetics. Cytogenet. Genome Res. 2010, 130–131, 1–14. [Google Scholar] [CrossRef]
- Wiens, J.J.; Brandley, M.C.; Reeder, T.W. Why does a trait evolve multiple times within a clade? Repeated evolution of snakelike body form in squamate reptiles. Evolution 2006, 60, 123. [Google Scholar] [CrossRef]
- Moen, D.S.; Morlon, H.; Wiens, J.J. Testing convergence versus history: Convergence dominates phenotypic evolution for over 150 million years in frogs. Syst. Biol. 2016, 65, 146–160. [Google Scholar] [CrossRef]
- Blom, M.P.K.; Horner, P.; Moritz, C. Convergence across a continent: Adaptive diversification in a recent radiation of Australian lizards. Proc. Biol. Sci. 2016, 283, 20160181. [Google Scholar] [CrossRef]
- Esquerré, D.; Scott Keogh, J. Parallel selective pressures drive convergent diversification of phenotypes in pythons and boas. Ecol. Lett. 2016, 19, 800–809. [Google Scholar] [CrossRef]
- Losos, J.B.; Jackman, T.R.; Larson, A.; Queiroz, K.; Rodriguez-Schettino, L. Contingency and determinism in replicated adaptive radiations of island lizards. Science 1998, 279, 2115–2118. [Google Scholar] [CrossRef]
- Mahler, D.L.; Revell, L.J.; Glor, R.E.; Losos, J.B. Ecological opportunity and the rate of morphological evolution in the diversification of Greater Antillean anoles. Evolution 2010, 64, 2731–2745. [Google Scholar] [CrossRef]
- Williams, E.E. The origin of faunas. Evolution of lizard congeners in a complex island fauna: A trial analysis. In Evolutionary Biology; Dobzhansky, T., Hecht, M.K., Steere, W.C., Eds.; Springer: New York, NY, USA, 1972; Volume 6, pp. 47–89. ISBN 9781468490633. [Google Scholar]
- Glor, R.E.; Kolbe, J.J.; Powell, R.; Larson, A.; Losos, J.B. Phylogenetic analysis of ecological and morphological diversification in Hispaniolan trunk-ground anoles (Anolis cybotes group). Evolution 2003, 57, 2383–2397. [Google Scholar] [CrossRef]
- Ng, J.; Landeen, E.L.; Logsdon, R.M.; Glor, R.E. Correlation between Anolis lizard dewlap phenotype and environmental variation indicates adaptive divergence of a signal important to sexual selection and species recognition. Evolution 2013, 67, 573–582. [Google Scholar] [CrossRef]
- Darwin, C. The Variation of Animals and Plants under Domestication; D. Appleton: New York, NY, USA, 1894; Volume 2, ISBN 9781406842500. [Google Scholar]
- Gould, S.J. Wonderful Life: The Burgess Shale and the Nature Of History; WW Norton & Company: New York, NY, USA, 1989; ISBN 039330700X. [Google Scholar]
- Osborn, H.F. The Titanotheres of Ancient Wyoming, Dakota, and Nebraska; U.S. Geological Survey Monograph: Sioux Falls, SD, USA, 1929; p. 55. [Google Scholar]
- Schluter, D.; McPhail, J.D. Character displacement and replicate adaptive radiation. Trends Ecol. Evol. 1993, 8, 197–200. [Google Scholar] [CrossRef]
- Danley, P.D.; Kocher, T.D. Speciation in rapidly diverging systems: Lessons from Lake Malawi. Mol. Ecol. 2001, 10, 1075–1086. [Google Scholar] [CrossRef]
- Conway Morris, S. Life’s Solution: Inevitable Humans in a Lonely Universe; Cambridge University Press: Cambridge, UK, 2003; ISBN 0521603250. [Google Scholar]
- Losos, J.B. Adaptive radiation, ecological opportunity, and evolutionary determinism. American Society of Naturalists E. O. Wilson award address. Am. Nat. 2010, 175, 623–639. [Google Scholar] [CrossRef]
- Schwartz, A.; Henderson, R.W. Amphibians and Reptiles of the West Indies: Descriptions, Distributions, and Natural History; University Press of Florida: Gainesville, FL, USA, 1991; ISBN 9780813010496. [Google Scholar]
- Cochran, D.M. Herpetological collections made in Hispaniola by the Utowana Expedition, 1934. Occ. Pap. Boston Soc. Nat. Hist. 1934, 8, 163–188. [Google Scholar]
- Rodríguez, A.; Rusciano, T.; Hamilton, R.; Holmes, L.; Jordan, D.; Wollenberg Valero, K.C. Genomic and phenotypic signatures of climate adaptation in an Anolis lizard. Ecol. Evol. 2017, 7, 6390–6403. [Google Scholar] [CrossRef]
- Endler, J.A. Geographic Variation, Speciation, and Clines; Princeton University Press: Princeton, NJ, USA, 1977; ISBN 0691081875. [Google Scholar]
- Wake, D.B.; Papenfuss, T.J.; Lynch, J.F. Distribution of salamanders along elevational transects in Mexico and Guatemala. Tulane Stud. Zool. Bot. 1992, 303–319. [Google Scholar]
- Valero, K.C.W.; Pathak, R.; Prajapati, I.; Bankston, S. A candidate multimodal functional genetic network for thermal adaptation. PeerJ 2014, 2, e578. [Google Scholar] [CrossRef]
- Munoz, M.M.; Stimola, M.A.; Algar, A.C.; Conover, A.; Rodriguez, A.J.; Landestoy, M.A.; Bakken, G.S.; Losos, J.B. Evolutionary stasis and lability in thermal physiology in a group of tropical lizards. Proc. R. Soc. Lond. B Biol. Sci. 2014, 281, 20132433. [Google Scholar] [CrossRef]
- Araújo, M.B.; Ferri-Yáñez, F.; Bozinovic, F.; Marquet, P.A.; Valladares, F.; Chown, S.L. Heat freezes niche evolution. Ecol. Lett. 2013, 16, 1206–1219. [Google Scholar] [CrossRef]
- Dacquin, R.; Davey, R.A.; Laplace, C.; Levasseur, R.; Morris, H.A.; Goldring, S.R.; Gebre-Medhin, S.; Galson, D.L.; Zajac, J.D.; Karsenty, G. Amylin inhibits bone resorption while the calcitonin receptor controls bone formation in vivo. J. Cell Biol. 2004, 164, 509–514. [Google Scholar] [CrossRef]
- Lee, H.-J.; Kim, S.-Y.; Kim, G.S.; Hwang, J.-Y.; Kim, Y.-J.; Jeong, B.; Kim, T.-H.; Park, E.K.; Lee, S.H.; Kim, H.-L.; et al. Fracture, bone mineral density, and the effects of calcitonin receptor gene in postmenopausal Koreans. Osteoporos. Int. 2010, 21, 1351–1360. [Google Scholar] [CrossRef]
- Goda, T.; Doi, M.; Umezaki, Y.; Murai, I.; Shimatani, H.; Chu, M.L.; Nguyen, V.H.; Okamura, H.; Hamada, F.N. Calcitonin receptors are ancient modulators for rhythms of preferential temperature in insects and body temperature in mammals. Genes Dev. 2018, 32, 140–155. [Google Scholar] [CrossRef]
- Cowie, R.H.; Holland, B.S. Molecular biogeography and diversification of the endemic terrestrial fauna of the Hawaiian Islands. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2008, 363, 3363–3376. [Google Scholar] [CrossRef]
- Emerson, B.C. Speciation on islands: What are we learning? Biol. J. Linn. Soc. 2008, 95, 47–52. [Google Scholar] [CrossRef][Green Version]
- Gillespie, R.G.; Claridge, E.M.; Goodacre, S.L. Biogeography of the fauna of French Polynesia: Diversification within and between a series of hot spot archipelagos. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2008, 363, 3335–3346. [Google Scholar] [CrossRef]
- Parent, C.E.; Caccone, A.; Petren, K. Colonization and diversification of Galápagos terrestrial fauna: A phylogenetic and biogeographical synthesis. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2008, 363, 3347–3361. [Google Scholar] [CrossRef]
- Grant, P.R.; Rosemary Grant, B. How and Why Species Multiply: The Radiation of Darwin’s Finches; Princeton University Press: Princeton, NJ, USA, 2011; ISBN 9780691149998. [Google Scholar]
- Larson, E.J. Evolution’s Workshop: God and Science on the Galápagos Islands; Basic Books: New York, NY, USA, 2002; p. 93. ISBN 0465038115. [Google Scholar]
- Van Valen, L. A new evolutionary law. Evol. Theory 1973, 1, 1–30. [Google Scholar]
- Caccone, A.; Gibbs, J.P.; Ketmaier, V.; Suatoni, E.; Powell, J.R. Origin and evolutionary relationships of giant Galápagos tortoises. Proc. Natl. Acad. Sci. USA 1999, 96, 13223–13228. [Google Scholar] [CrossRef]
- Román-Palacios, C.; Wiens, J.J. The Tortoise and the Finch: Testing for island effects on diversification using two iconic Galápagos radiations. J. Biogeogr. 2018, 45, 1701–1712. [Google Scholar] [CrossRef]
- Poulakakis, N.; Edwards, D.L.; Chiari, Y.; Garrick, R.C.; Russello, M.A.; Benavides, E.; Watkins-Colwell, G.J.; Glaberman, S.; Tapia, W.; Gibbs, J.P.; et al. Description of a new Galapagos Giant Tortoise species (Chelonoidis; Testudines: Testudinidae) from Cerro Fatal on Santa Cruz island. PLoS ONE 2015, 10, e0138779. [Google Scholar] [CrossRef]
- Beheregaray, L.B.; Gibbs, J.P.; Havill, N.; Fritts, T.H.; Powell, J.R.; Caccone, A. Giant tortoises are not so slow: Rapid diversification and biogeographic consensus in the Galápagos. Proc. Natl. Acad. Sci. USA 2004, 101, 6514–6519. [Google Scholar] [CrossRef]
- Poulakakis, N.; Russello, M.; Geist, D.; Caccone, A. Unravelling the peculiarities of island life: Vicariance, dispersal and the diversification of the extinct and extant giant Galápagos tortoises. Mol. Ecol. 2012, 21, 160–173. [Google Scholar] [CrossRef]
- Jensen, E.L.; Mooers, A.Ø.; Caccone, A.; Russello, M.A. I-HEDGE: Determining the optimum complementary sets of taxa for conservation using evolutionary isolation. PeerJ 2016, 4, e2350. [Google Scholar] [CrossRef]
- Abbott, R.; Albach, D.; Ansell, S.; Arntzen, J.W.; Baird, S.J.E.; Bierne, N.; Boughman, J.; Brelsford, A.; Buerkle, C.A.; Buggs, R.; et al. Hybridization and speciation. J. Evol. Biol. 2013, 26, 229–246. [Google Scholar] [CrossRef]
- Schwenk, K.; Brede, N.; Streit, B. Introduction. Extent, processes and evolutionary impact of interspecific hybridization in animals. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2008, 363, 2805–2811. [Google Scholar] [CrossRef]
- Garrick, R.C.; Benavides, E.; Russello, M.A.; Hyseni, C.; Edwards, D.L.; Gibbs, J.P.; Tapia, W.; Ciofi, C.; Caccone, A. Lineage fusion in Galápagos giant tortoises. Mol. Ecol. 2014, 23, 5276–5290. [Google Scholar] [CrossRef]
- Caccone, A.; Gentile, G.; Gibbs, J.P.; Frirts, T.H.; Snell, H.L.; Betts, J.; Powell, J.R. Phylogeography and history of giant Galápagos tortoises. Evolution 2002, 56, 2052–2066. [Google Scholar]
- Russello, M.A.; Beheregaray, L.B.; Gibbs, J.P.; Fritts, T.; Havill, N.; Powell, J.R.; Caccone, A. Lonesome George is not alone among Galápagos tortoises. Curr. Biol. 2007, 17, R317–R318. [Google Scholar] [CrossRef]
- Poulakakis, N.; Glaberman, S.; Russello, M.; Beheregaray, L.B.; Ciofi, C.; Powell, J.R.; Caccone, A. Historical DNA analysis reveals living descendants of an extinct species of Galápagos tortoise. Proc. Natl. Acad. Sci. USA 2008, 105, 15464–15469. [Google Scholar] [CrossRef]
- Russello, M.A.; Poulakakis, N.; Gibbs, J.P.; Tapia, W.; Benavides, E.; Powell, J.R.; Caccone, A. DNA from the past informs ex situ conservation for the future: An “extinct” species of Galápagos Tortoise identified in captivity. PLoS ONE 2010, 5, e8683. [Google Scholar] [CrossRef]
- Garrick, R.C.; Benavides, E.; Russello, M.A.; Gibbs, J.P.; Poulakakis, N.; Dion, K.B.; Hyseni, C.; Kajdacsi, B.; Márquez, L.; Bahan, S.; et al. Genetic rediscovery of an “extinct” Galápagos giant tortoise species. Curr. Biol. 2012, 22, R10–R11. [Google Scholar] [CrossRef]
- Edwards, D.L.; Benavides, E.; Garrick, R.C.; Gibbs, J.P.; Russello, M.A.; Dion, K.B.; Hyseni, C.; Flanagan, J.P.; Tapia, W.; Caccone, A. The genetic legacy of Lonesome George survives: Giant tortoises with Pinta Island ancestry identified in Galápagos. Biol. Conserv. 2013, 157, 225–228. [Google Scholar] [CrossRef]
- Raia, P.; Guarino, F.M.; Turano, M.; Polese, G.; Rippa, D.; Carotenuto, F.; Monti, D.M.; Cardi, M.; Fulgione, D. The blue lizard spandrel and the island syndrome. BMC Evol. Biol. 2010, 10, 289. [Google Scholar] [CrossRef]
- Wang, S.; Liu, C.; Wu, J.; Xu, C.; Zhang, J.; Bai, C.; Gao, X.; Liu, X.; Li, X.; Zhu, W.; et al. Propagule pressure and hunting pressure jointly determine genetic evolution in insular populations of a global frog invader. Sci. Rep. 2019, 9, 448. [Google Scholar] [CrossRef]
- Miller, J.M.; Quinzin, M.C.; Poulakakis, N.; Gibbs, J.P.; Beheregaray, L.B.; Garrick, R.C.; Russello, M.A.; Ciofi, C.; Edwards, D.L.; Hunter, E.A.; et al. Identification of genetically important individuals of the rediscovered Floreana Galápagos Giant Tortoise (Chelonoidis elephantopus) provides founders for species restoration program. Sci. Rep. 2017, 7, 11471. [Google Scholar] [CrossRef]
- Quinzin, M.C.; Sandoval-Castillo, J.; Miller, J.M.; Beheregaray, L.B.; Russello, M.A.; Hunter, E.A.; Gibbs, J.P.; Tapia, W.; Villalva, F.; Caccone, A. Genetically informed captive breeding of hybrids of an extinct species of Galapagos tortoise. Conserv. Biol. 2019. [Google Scholar] [CrossRef]
- Quesada, V.; Freitas-Rodríguez, S.; Miller, J.; Pérez-Silva, J.G.; Jiang, Z.-F.; Tapia, W.; Santiago-Fernández, O.; Campos-Iglesias, D.; Kuderna, L.F.K.; Quinzin, M.; et al. Giant tortoise genomes provide insights into longevity and age-related disease. Nat. Ecol. Evol. 2019, 3, 87–95. [Google Scholar] [CrossRef]
- Gaughran, S.J.; Quinzin, M.C.; Miller, J.M.; Garrick, R.C.; Edwards, D.L.; Russello, M.A.; Poulakakis, N.; Ciofi, C.; Beheregaray, L.B.; Caccone, A. Theory, practice, and the conservation of Galápagos giant tortoises in the age of genomics. Evol. Appl. 2017, 11, 1084–1093. [Google Scholar] [CrossRef]
- Jensen, E.L.; Miller, J.M.; Edwards, D.L.; Garrick, R.C.; Tapia, W.; Caccone, A.; Russello, M.A. Temporal mitogenomics of the Galapagos Giant Tortoise from Pinzón reveals potential biases in population genetic inference. J. Hered. 2018, 109, 631–640. [Google Scholar] [CrossRef]
- Jensen, E.L.; Edwards, D.L.; Garrick, R.C.; Miller, J.M.; Gibbs, J.P.; Cayot, L.J.; Tapia, W.; Caccone, A.; Russello, M.A. Population genomics through time provides insights into the consequences of decline and rapid demographic recovery through head-starting in a Galapagos giant tortoise. Evol. Appl. 2018, 11, 1811–1821. [Google Scholar] [CrossRef]
- Miller, J.M.; Quinzin, M.C.; Edwards, D.L.; Eaton, D.A.R.; Jensen, E.L.; Russello, M.A.; Gibbs, J.P.; Tapia, W.; Rueda, D.; Caccone, A. Genome-wide assessment of diversity and divergence among extant Galapagos Giant Tortoise species. J. Hered. 2018, 109, 611–619. [Google Scholar] [CrossRef]
- Vences, M.; Wake, D.B. Speciation, species boundaries and phylogeography of amphibians. In Amphibian Biology; Heatwole, H., Ed.; Surrey Beatty & Sons: Clayton South, Australia, 2007; pp. 2613–2675. ISBN 9780980311389. [Google Scholar]
- Shaffer, H.B.; Gidiş, M.; McCartney-Melstad, E.; Neal, K.M.; Oyamaguchi, H.M.; Tellez, M.; Toffelmier, E.M. Conservation genetics and genomics of amphibians and reptiles. Annu. Rev. Anim. Biosci. 2015, 3, 113–138. [Google Scholar] [CrossRef]
- AmphibiaWeb Species Numbers. Available online: http://amphibiaweb.org:8000/amphibian/speciesnums.html (accessed on 8 April 2019).
- Scheele, B.C.; Pasmans, F.; Skerratt, L.F.; Berger, L.; Martel, A.; Beukema, W.; Acevedo, A.A.; Burrowes, P.A.; Carvalho, T.; Catenazzi, A.; et al. Amphibian fungal panzootic causes catastrophic and ongoing loss of biodiversity. Science 2019, 363, 1459–1463. [Google Scholar] [CrossRef]
- Gibbons, J.W.; Scott, D.E.; Ryan, T.J.; Buhlmann, K.A.; Tuberville, T.D.; Metts, B.S.; Greene, J.L.; Mills, T.; Leiden, Y.; Poppy, S.; et al. The Global Decline of Reptiles, Déjà Vu Amphibians: Reptile species are declining on a global scale. Six significant threats to reptile populations are habitat loss and degradation, introduced invasive species, environmental pollution, disease, unsustainable use, and global climate change. Bioscience 2000, 50, 653–666. [Google Scholar]
- Donihue, C.M.; Lambert, M.R. Adaptive evolution in urban ecosystems. Ambio 2015, 44, 194–203. [Google Scholar] [CrossRef]
- Winchell, K.M.; Reynolds, R.G.; Prado-Irwin, S.R.; Puente-Rolón, A.R.; Revell, L.J. Phenotypic shifts in urban areas in the tropical lizard Anolis cristatellus. Evolution 2016, 70, 1009–1022. [Google Scholar] [CrossRef]
- Thompson, K.A.; Rieseberg, L.H.; Schluter, D. Speciation and the City. Trends Ecol. Evol. 2018, 33, 815–826. [Google Scholar] [CrossRef]
- Campbell-Staton, S.C.; Cheviron, Z.A.; Rochette, N.; Catchen, J.; Losos, J.B.; Edwards, S.V. Winter storms drive rapid phenotypic, regulatory, and genomic shifts in the green anole lizard. Science 2017, 357, 495–498. [Google Scholar] [CrossRef]
- Cyriac, V.P.; Kodandaramaiah, U. Paleoclimate determines diversification patterns in the fossorial snake family Uropeltidae Cuvier, 1829. Mol. Phylogenet. Evol. 2017, 116, 97–107. [Google Scholar] [CrossRef]
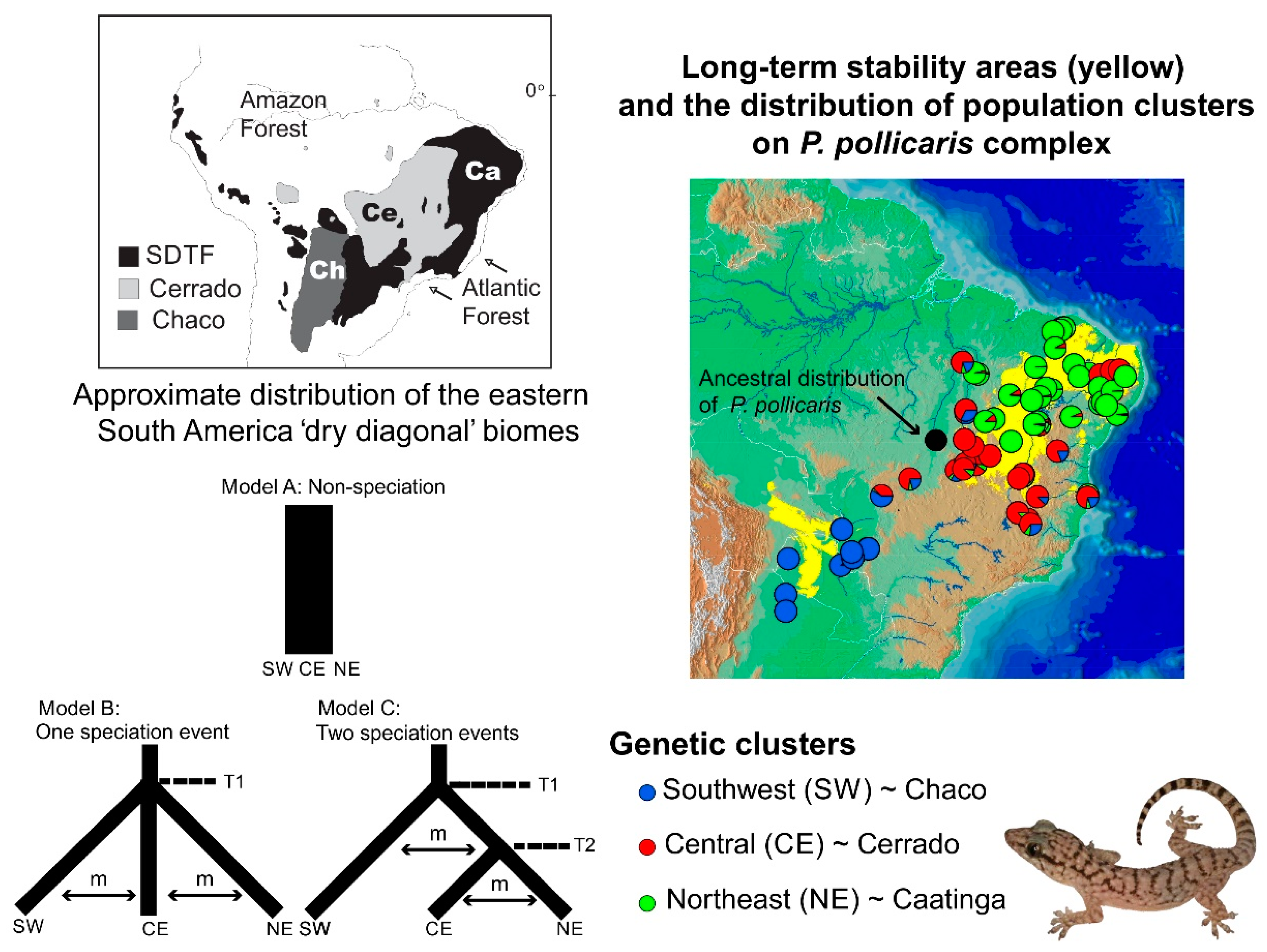
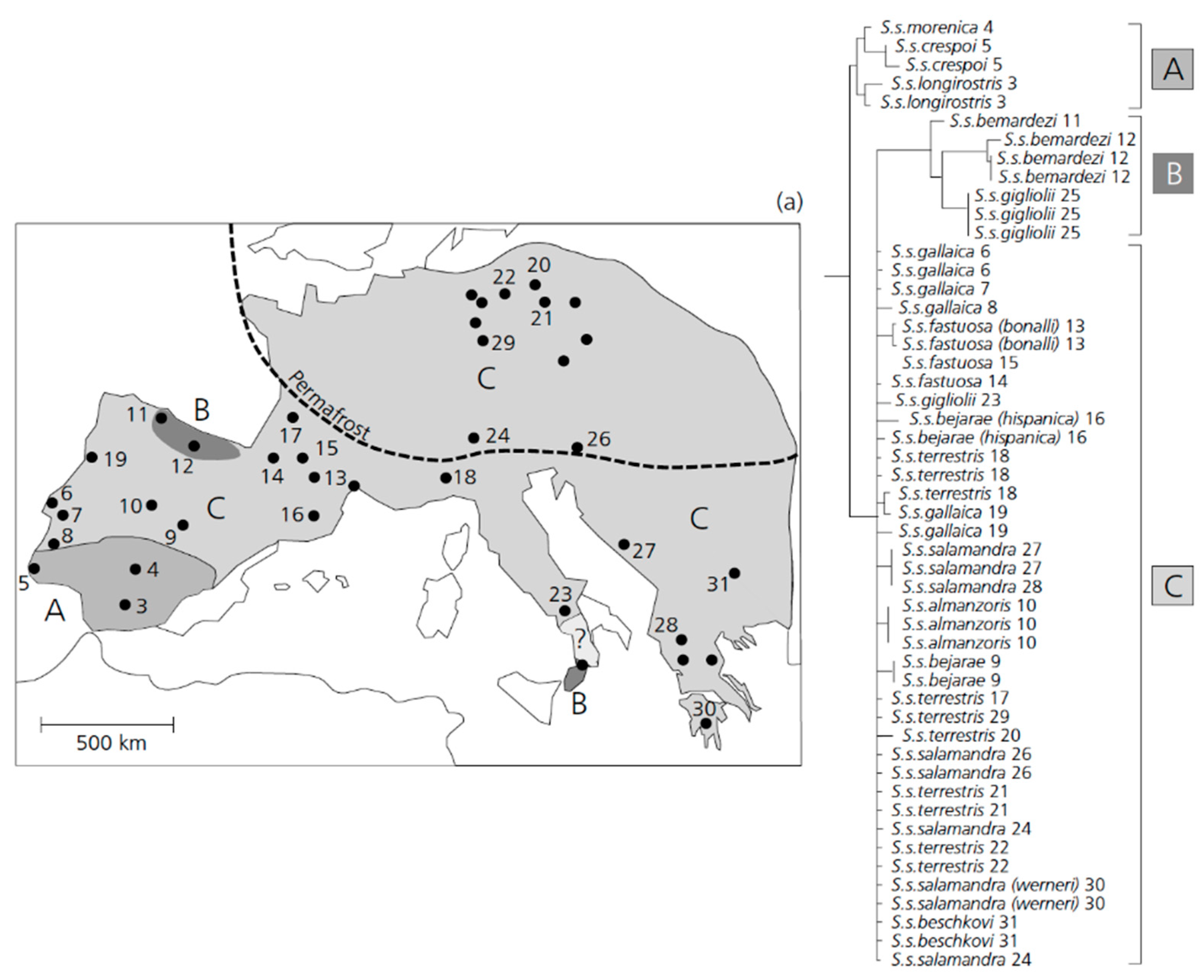
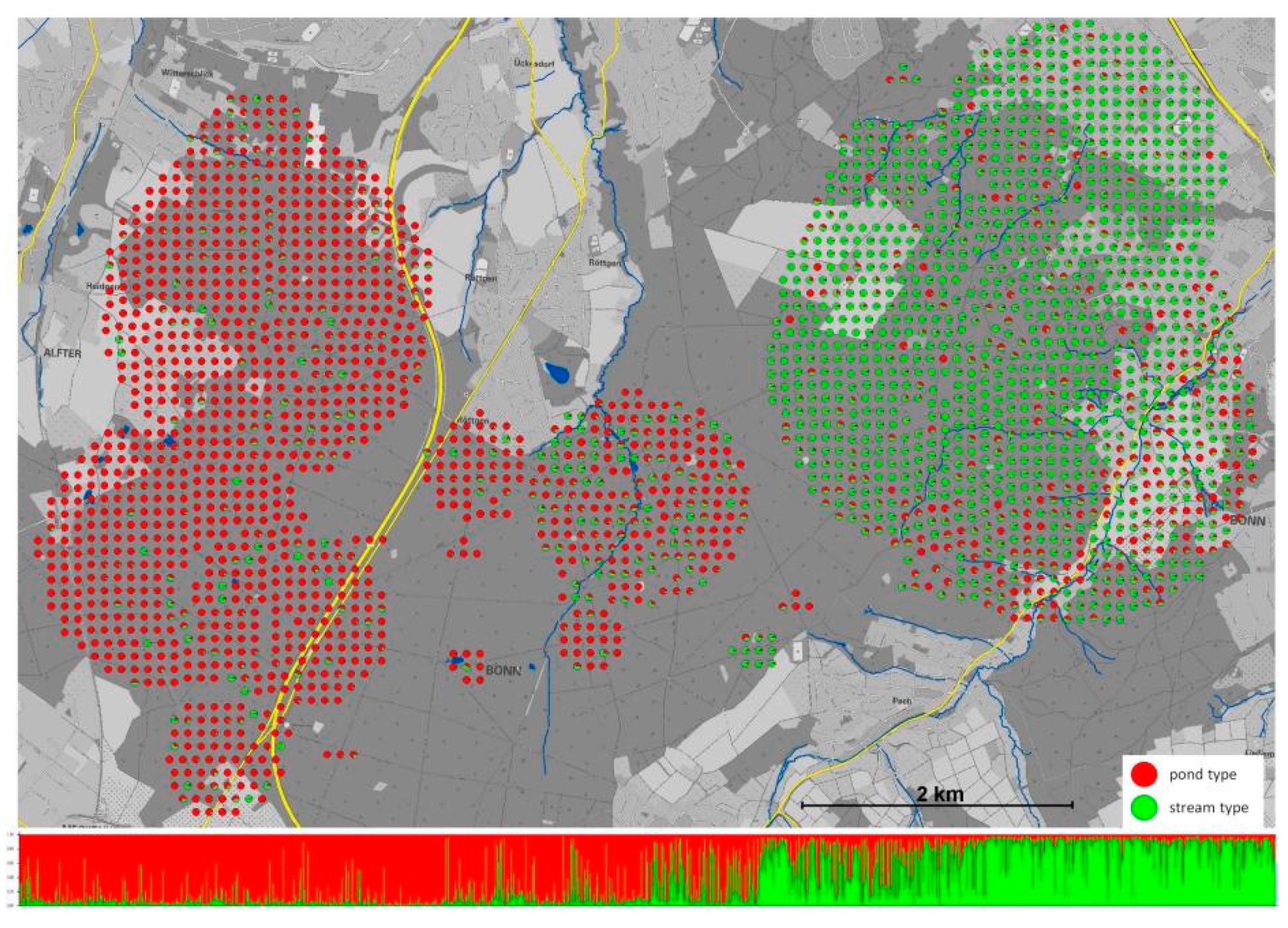
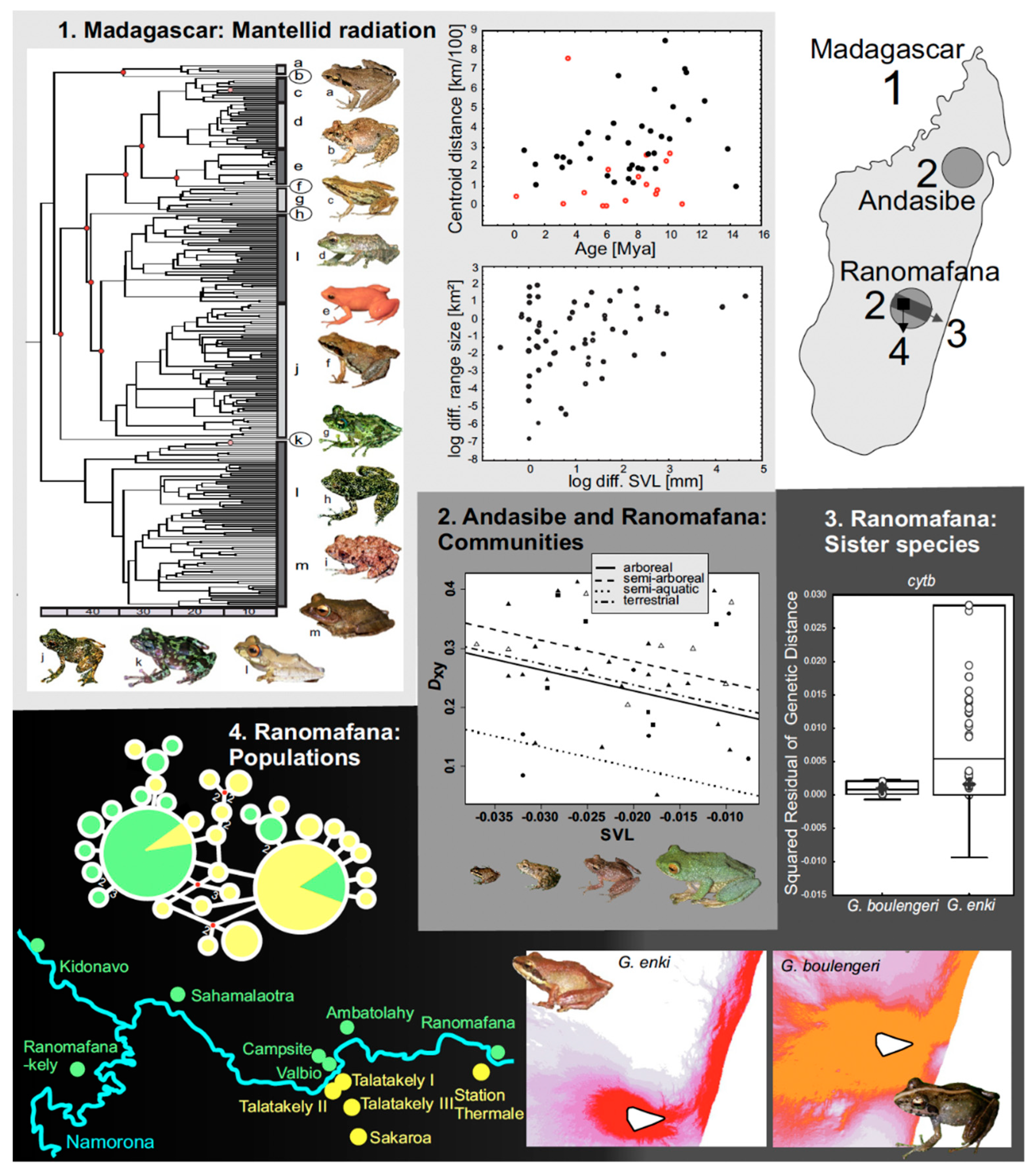
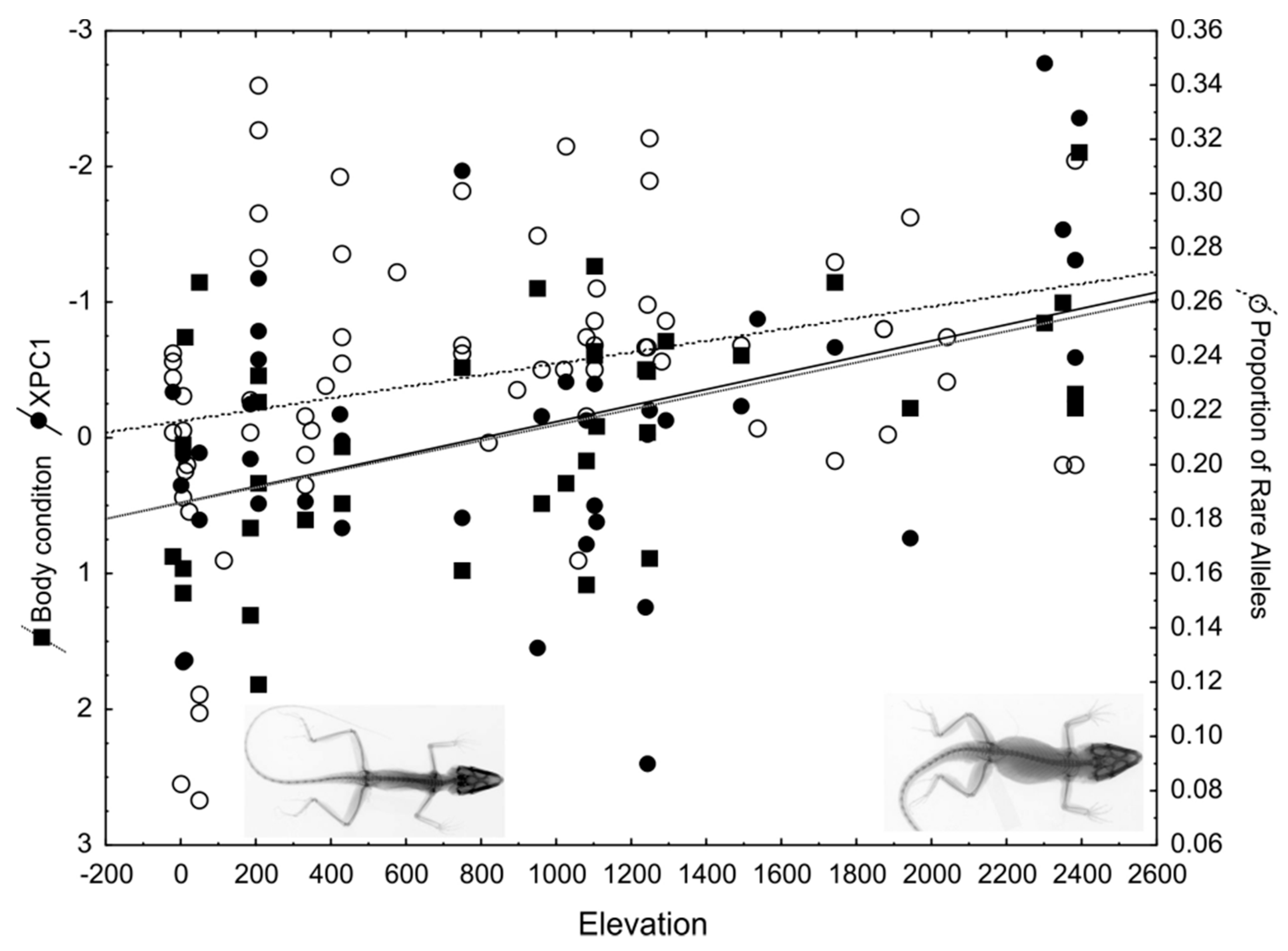
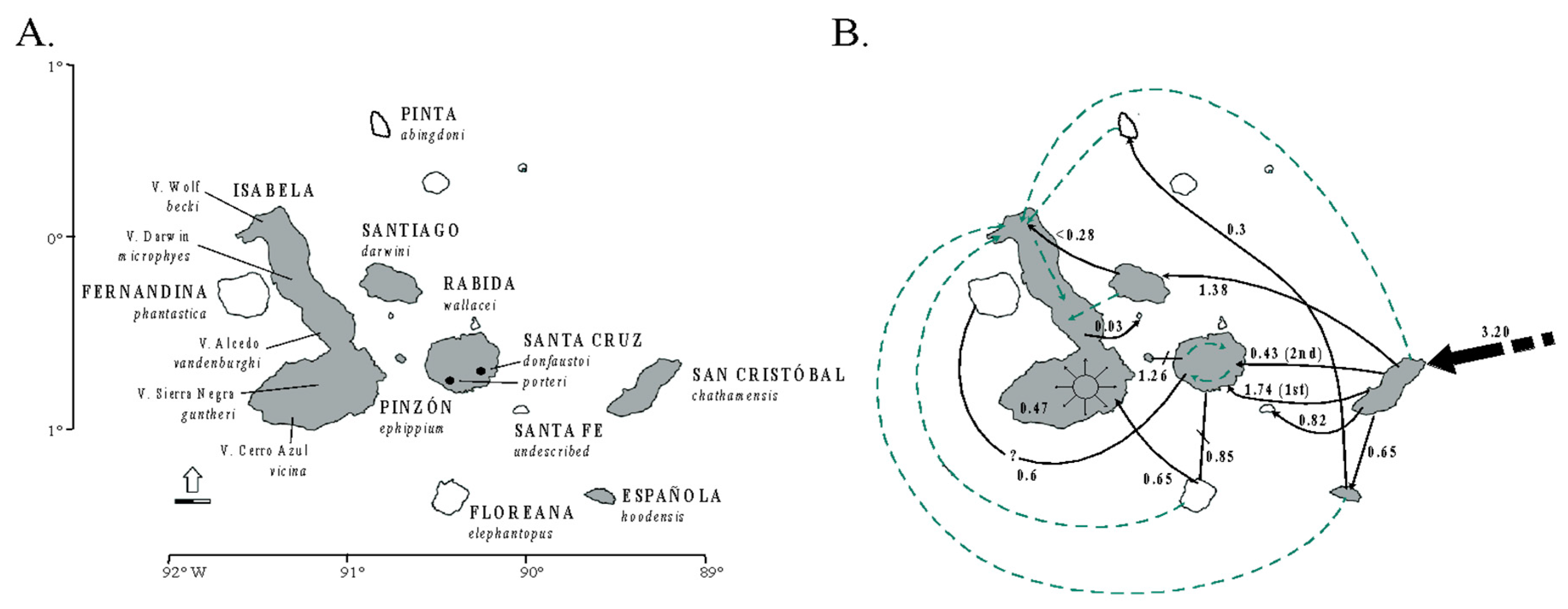
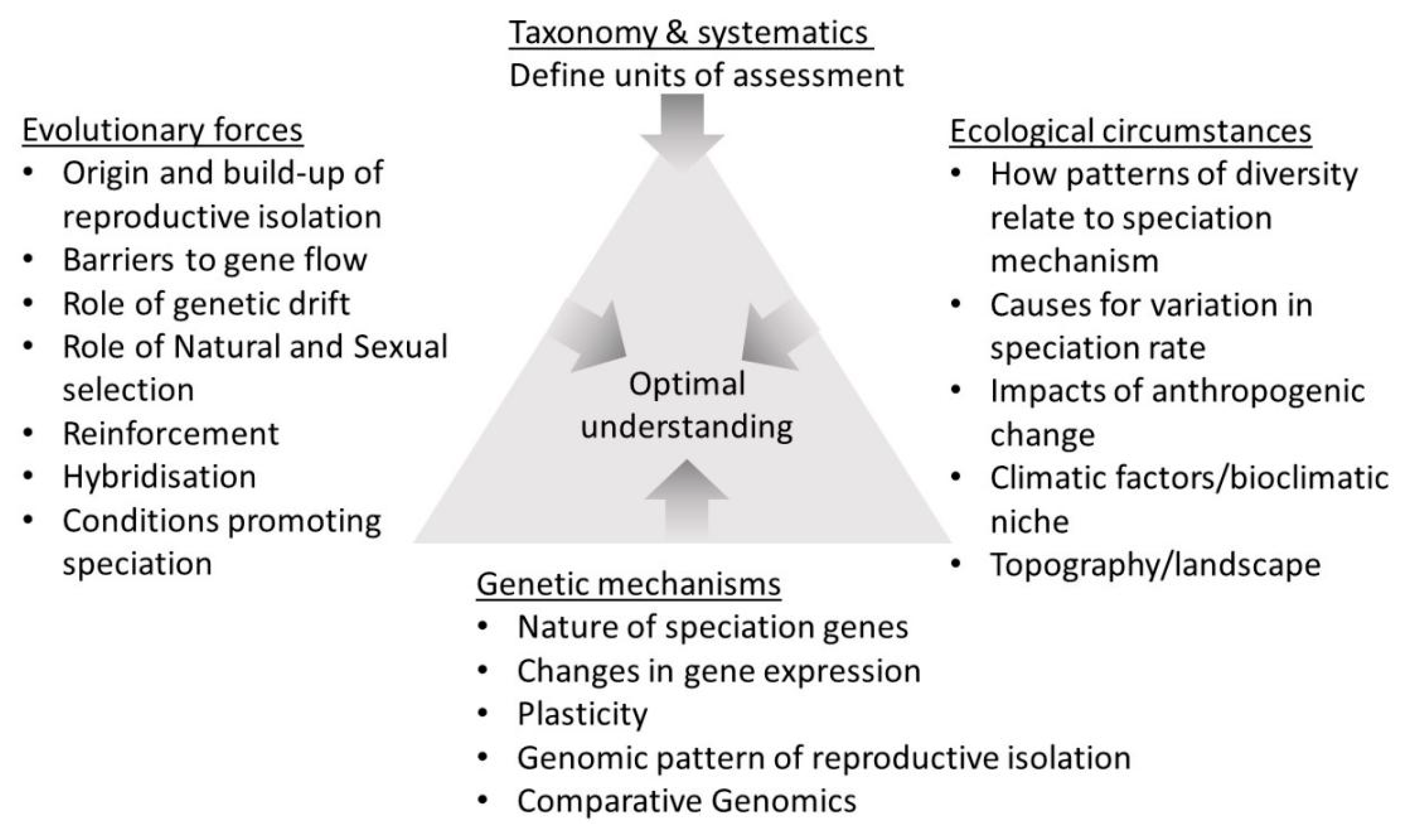
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wollenberg Valero, K.C.; Marshall, J.C.; Bastiaans, E.; Caccone, A.; Camargo, A.; Morando, M.; Niemiller, M.L.; Pabijan, M.; Russello, M.A.; Sinervo, B.; et al. Patterns, Mechanisms and Genetics of Speciation in Reptiles and Amphibians. Genes 2019, 10, 646. https://doi.org/10.3390/genes10090646
Wollenberg Valero KC, Marshall JC, Bastiaans E, Caccone A, Camargo A, Morando M, Niemiller ML, Pabijan M, Russello MA, Sinervo B, et al. Patterns, Mechanisms and Genetics of Speciation in Reptiles and Amphibians. Genes. 2019; 10(9):646. https://doi.org/10.3390/genes10090646
Chicago/Turabian StyleWollenberg Valero, Katharina C., Jonathon C. Marshall, Elizabeth Bastiaans, Adalgisa Caccone, Arley Camargo, Mariana Morando, Matthew L. Niemiller, Maciej Pabijan, Michael A. Russello, Barry Sinervo, and et al. 2019. "Patterns, Mechanisms and Genetics of Speciation in Reptiles and Amphibians" Genes 10, no. 9: 646. https://doi.org/10.3390/genes10090646
APA StyleWollenberg Valero, K. C., Marshall, J. C., Bastiaans, E., Caccone, A., Camargo, A., Morando, M., Niemiller, M. L., Pabijan, M., Russello, M. A., Sinervo, B., Werneck, F. P., Sites, J. W., Jr., Wiens, J. J., & Steinfartz, S. (2019). Patterns, Mechanisms and Genetics of Speciation in Reptiles and Amphibians. Genes, 10(9), 646. https://doi.org/10.3390/genes10090646




