Identification and Characterization of a Thermotolerant TILLING Allele of Heat Shock Binding Protein 1 in Tomato
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material and Stress Treatment
2.2. TILLING Screening
2.3. Genotyping of Mutant Plants
2.4. Seedling Thermotolerance
2.5. Image Based Phenotyping: Data Acquisition and Processing
2.6. Expression Constructs
2.7. Protoplast Preparation and GUS Reporter Assays
2.8. RNA Extraction and Transcript Analysis
2.9. Microscopy
2.10. Orthology Search and in Silico Structure Prediction
2.11. Transcriptome Data
3. Results
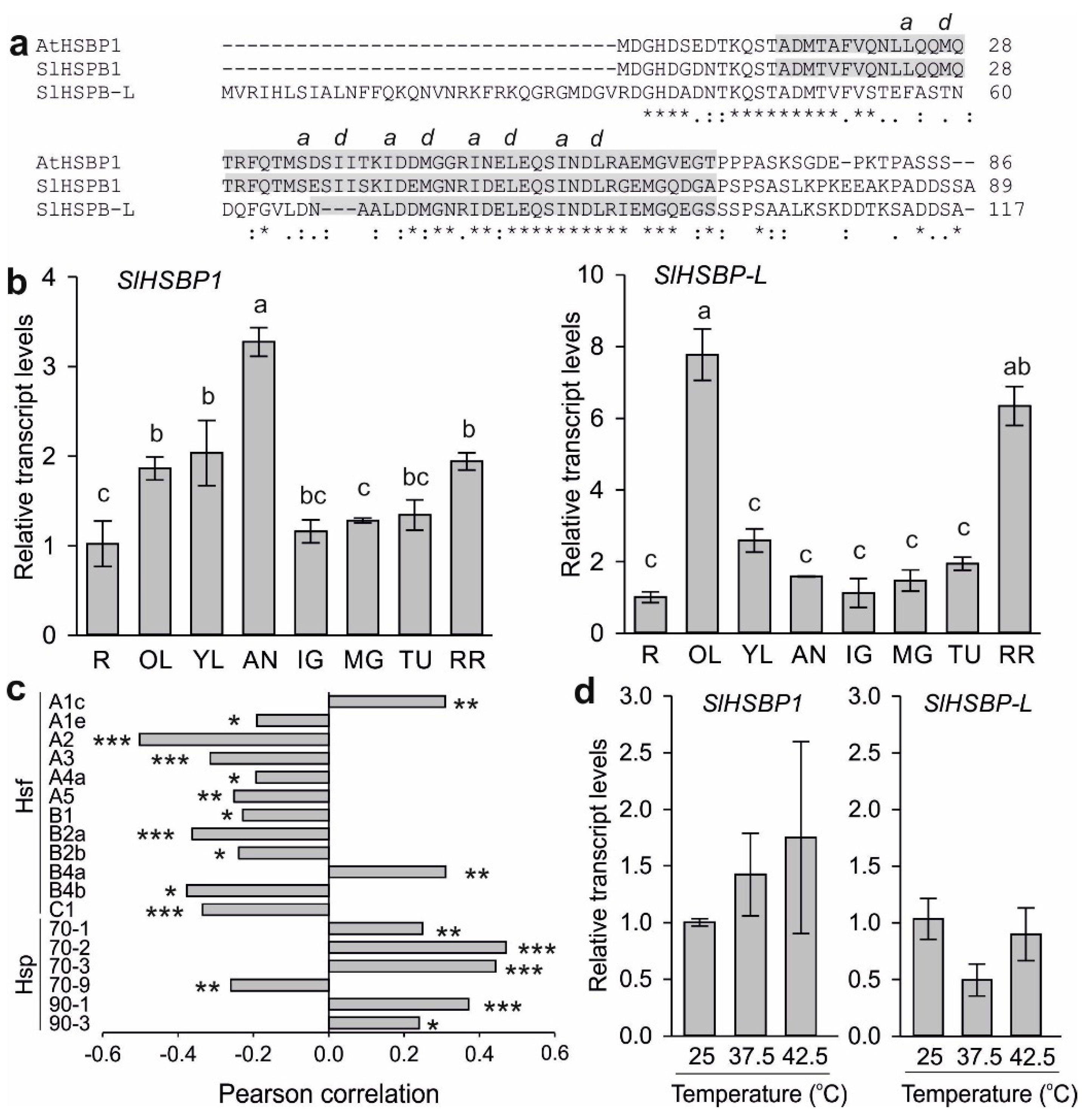
3.1. Expression of Putative HSBP Genes in Tomato
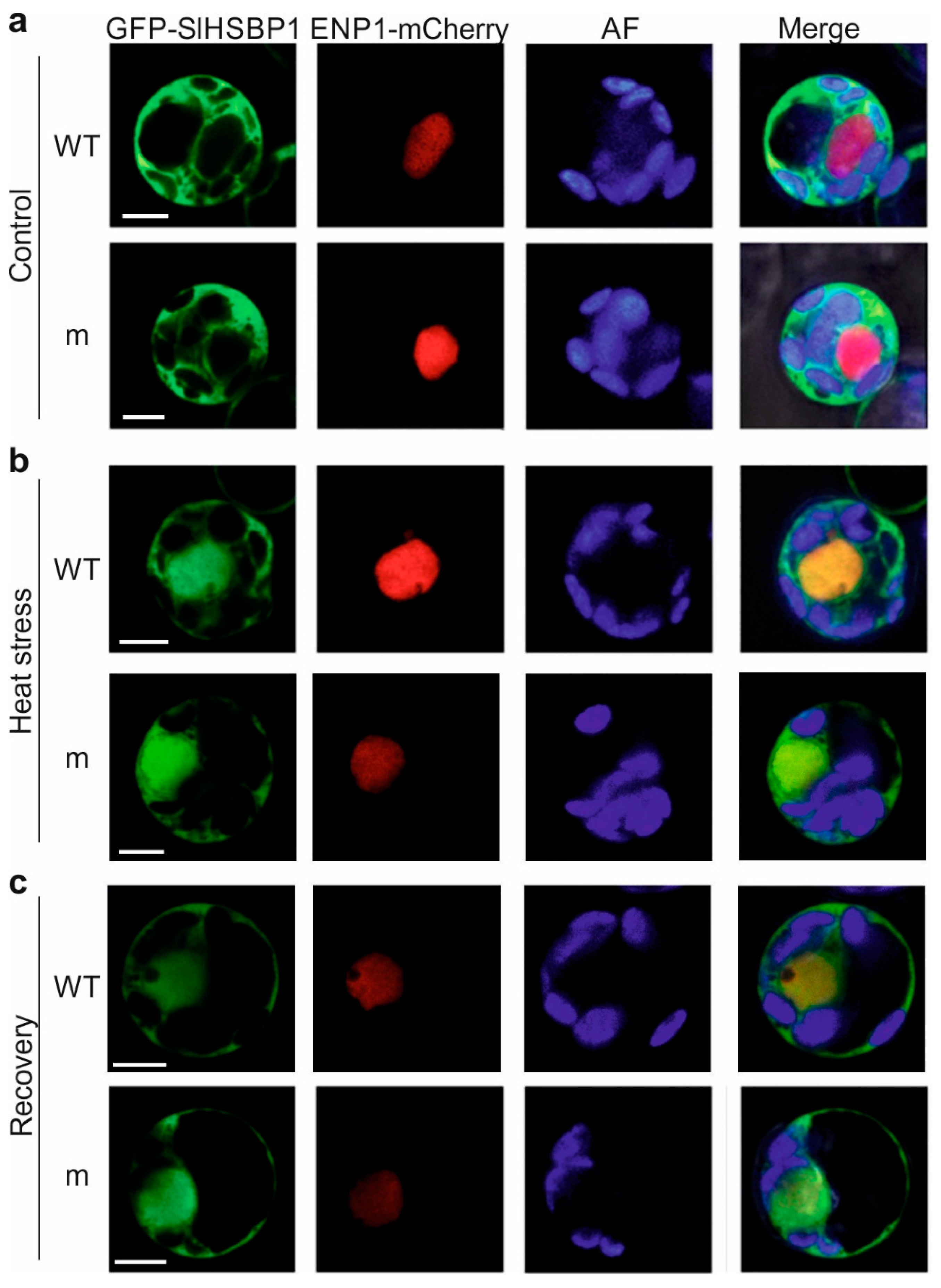
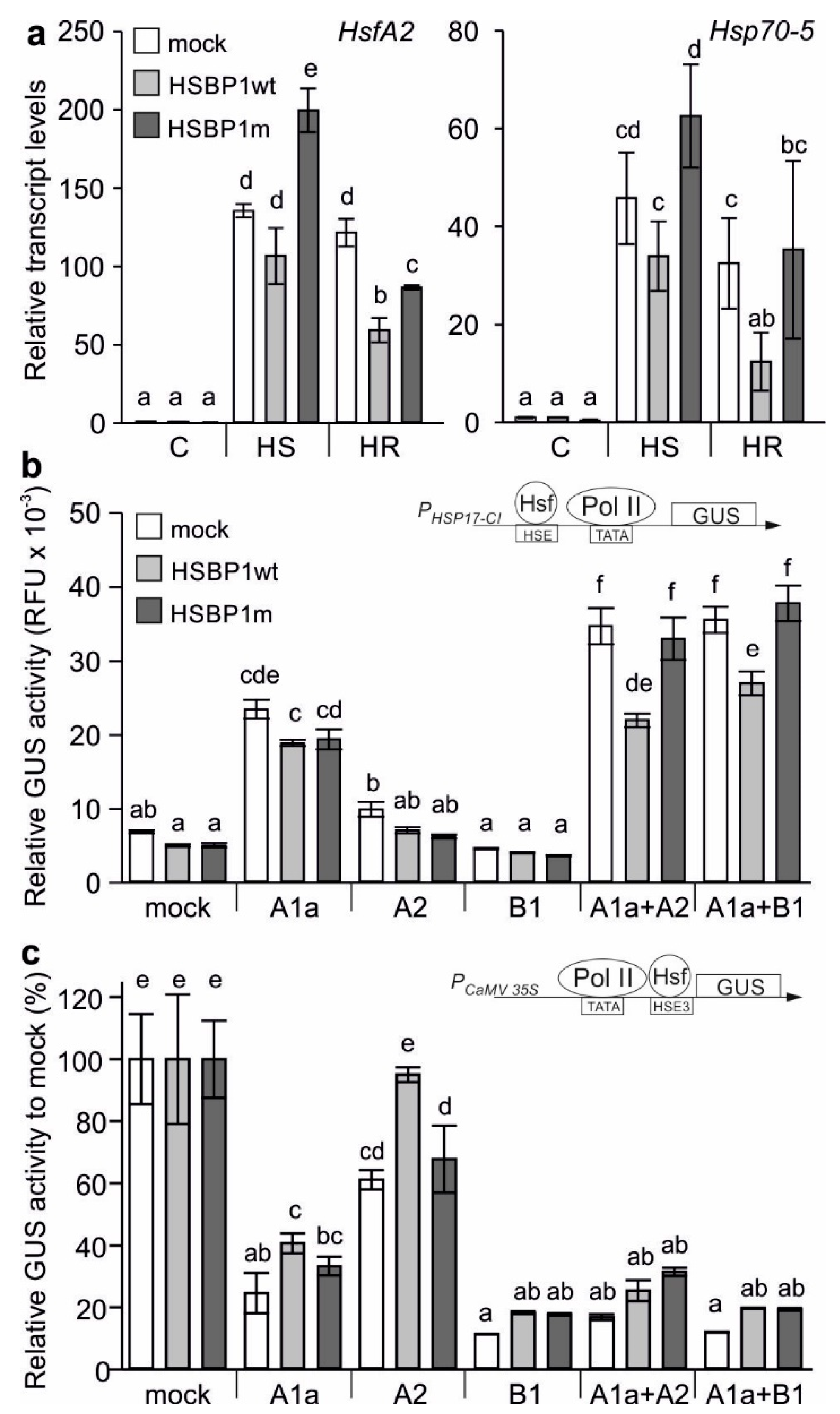
3.2. A Mutation of the Zipper Affects the HSBP Functionality
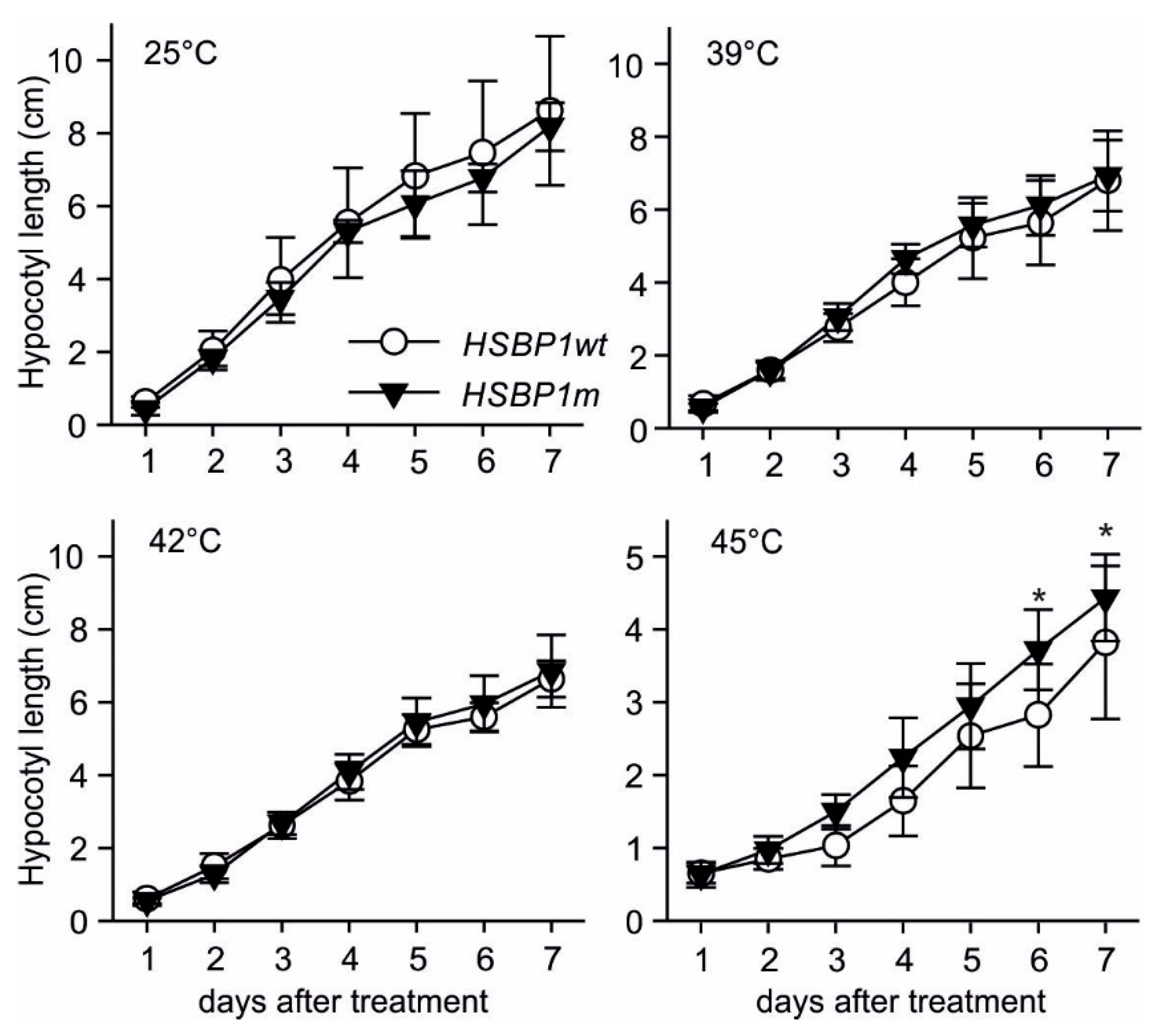
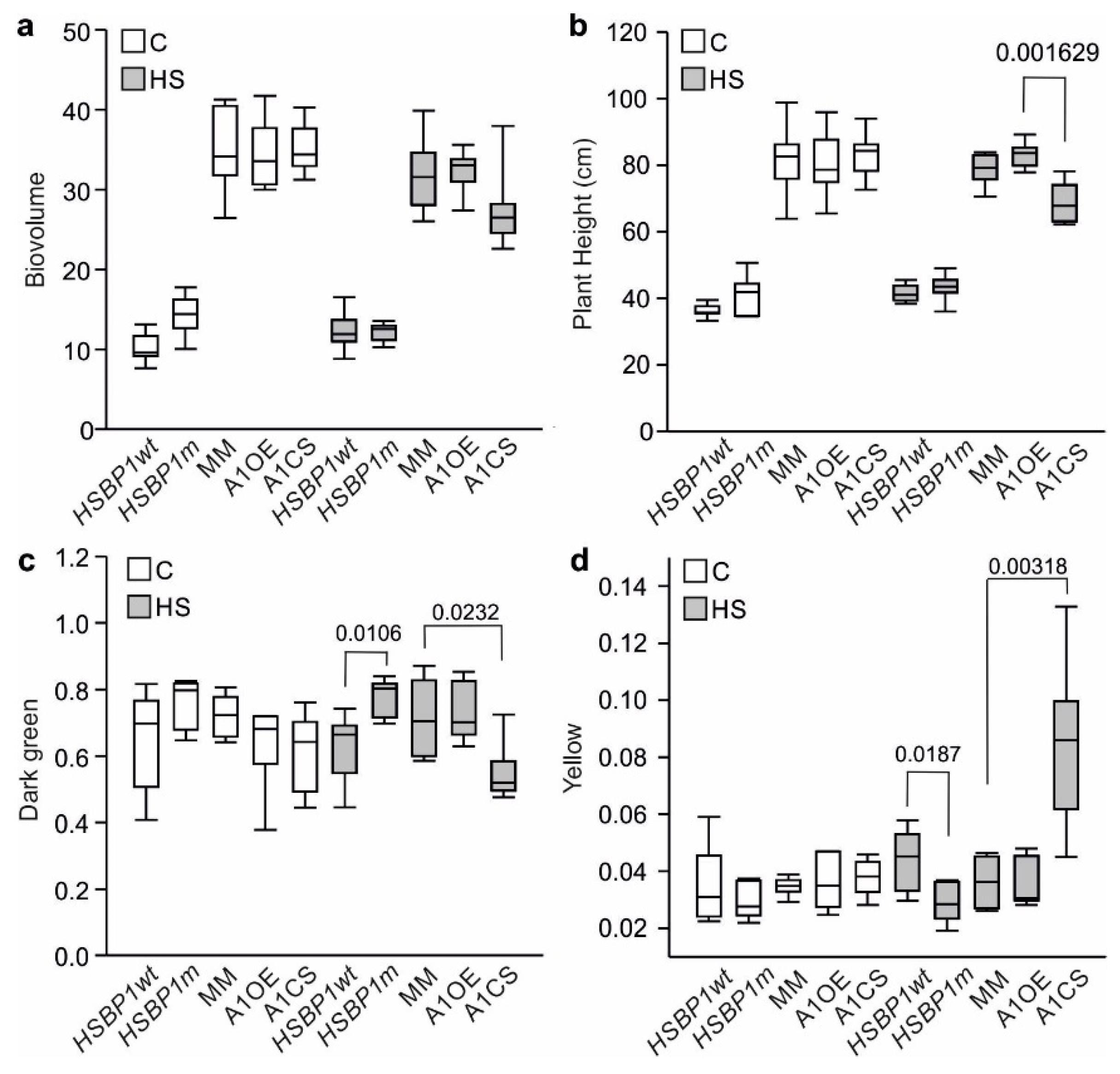
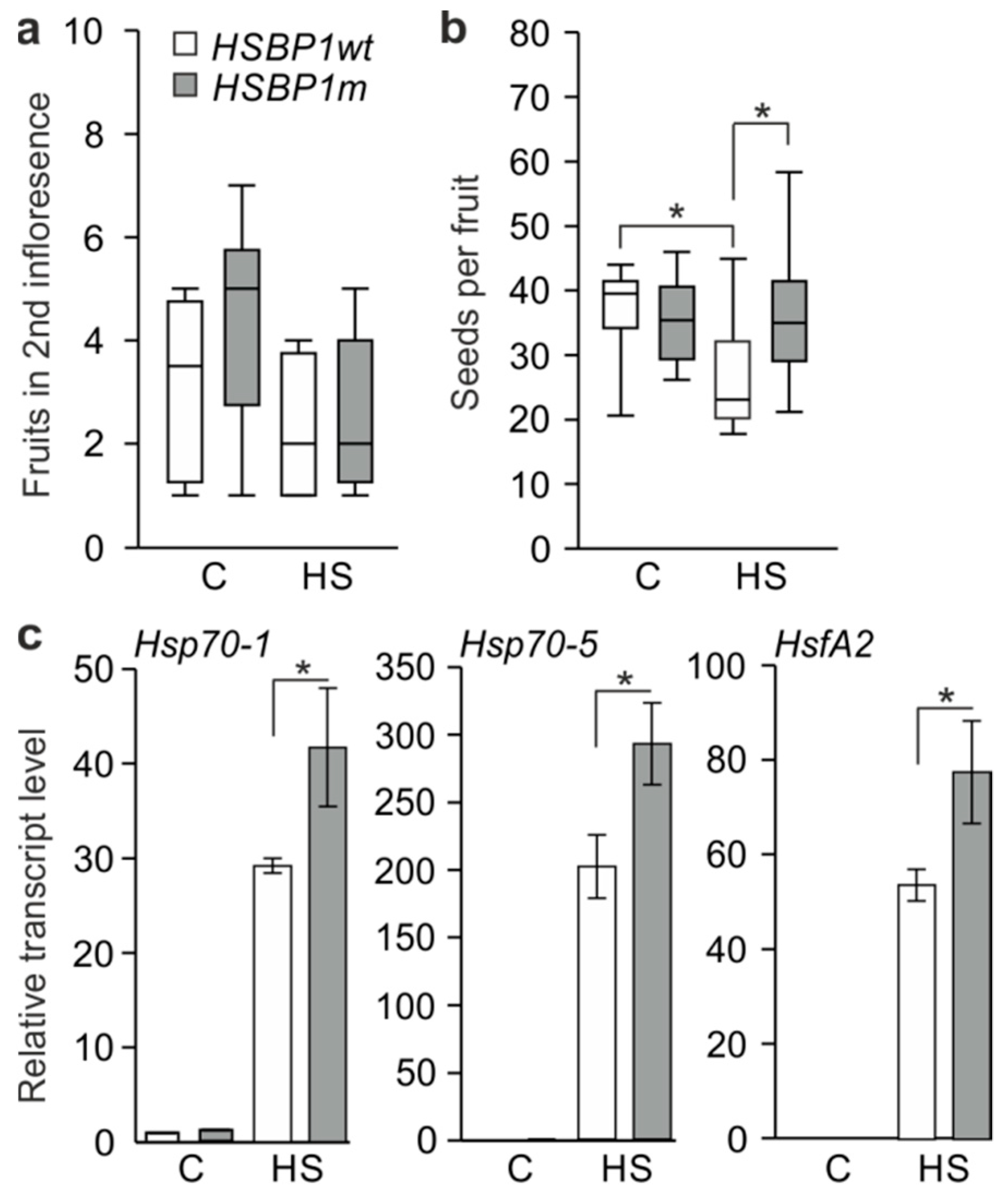
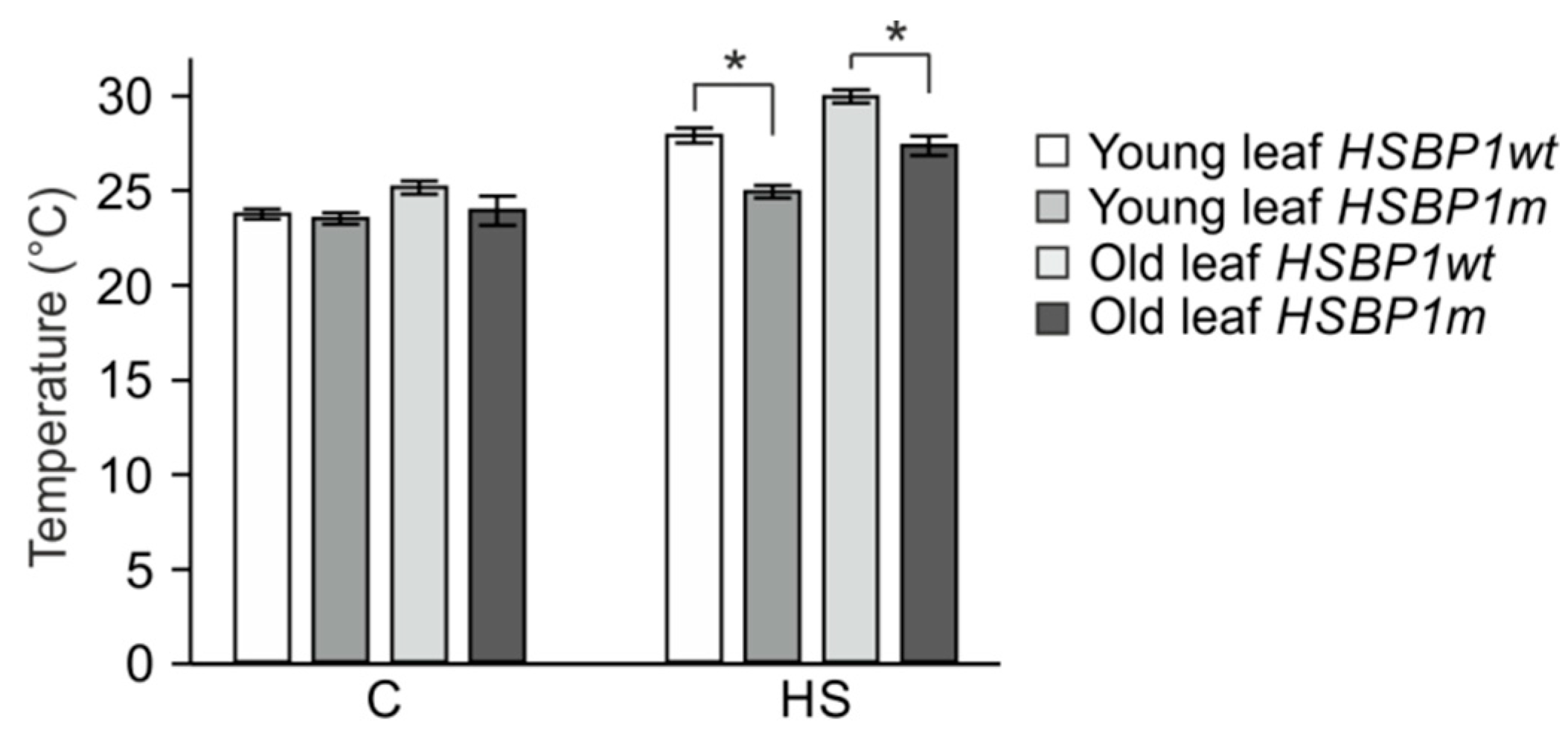
3.3. Thermotolerance of SlHSBP1 Plants
4. Discussion
4.1. Characterization of Tomato HSBP1 Protein
4.2. SlHSBP1m Plants Exhibit Higher Thermotolerance
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bokszczanin, K.L.; Fragkostefanakis, S. Perspectives on deciphering mechanisms underlying plant heat stress response and thermotolerance. Front. Plant Sci. 2013, 4, 315. [Google Scholar] [CrossRef] [PubMed]
- Wahid, A.; Gelani, S.; Ashraf, M.; Foolad, M. Heat tolerance in plants: An overview. Environ. Exp. Bot. 2007, 61, 199–223. [Google Scholar] [CrossRef]
- Mittler, R.; Finka, A.; Goloubinoff, P. How do plants feel the heat? Trends Biochem. Sci. 2012, 37, 118–125. [Google Scholar] [CrossRef] [PubMed]
- Rieu, I.; Twell, D.; Firon, N. Pollen Development at High Temperature: From Acclimation to Collapse. Plant Physiol. 2017, 173, 1967–1976. [Google Scholar] [CrossRef] [PubMed]
- Müller, F.; Xu, J.; Kristensen, L.; Wolters-Arts, M.; De Groot, P.F.M.; Jansma, S.Y.; Mariani, C.; Park, S.; Rieu, I. High-temperature-induced defects in tomato (Solanum lycopersicum) anther and pollen development are associated with reduced expression of B-class floral patterning genes. PLoS ONE 2016, 11, e0167614. [Google Scholar] [CrossRef] [PubMed]
- Firon, N.; Shaked, R.; Peet, M.M.; Pharr, D.M.; Zamski, E.; Rosenfeld, K.; Althan, L.; Pressman, E. Pollen grains of heat tolerant tomato cultivars retain higher carbohydrate concentration under heat stress conditions. Sci. Hortic. 2006, 109, 212–217. [Google Scholar] [CrossRef]
- Sato, S.; Peet, M.M.; Thomas, J.F. Physiological factors limit fruit set of tomato (Lycopersicon esculentum Mill.) under chronic, mild heat stress. Plant Cell Environ. 2000, 23, 719–726. [Google Scholar] [CrossRef]
- Hartl, F.U.; Bracher, A.; Hayer-Hartl, M. Molecular chaperones in protein folding and proteostasis. Nature 2011, 475, 324–332. [Google Scholar] [CrossRef]
- Fragkostefanakis, S.; Simm, S.; Paul, P.; Bublak, D.; Scharf, K.D.; Schleiff, E. Chaperone network composition in Solanum lycopersicum explored by transcriptome profiling and microarray meta-analysis. Plant Cell Environ. 2015, 38, 693–709. [Google Scholar] [CrossRef]
- Keller, M.; Consortium, S.; Simm, S. The coupling of transcriptome and proteome adaptation during development and heat stress response of tomato pollen. BMC Genom. 2018, 19, 447. [Google Scholar] [CrossRef]
- Liu, H.-C.; Charng, Y.-Y. Common and Distinct Functions of Arabidopsis Class A1 and A2 Heat Shock Factors in Diverse Abiotic Stress Responses and Development. Plant Physiol. 2013, 163, 276–290. [Google Scholar] [CrossRef] [PubMed]
- Iwata, Y.; Sakiyama, M.; Lee, M.H.; Koizumi, N. Transcriptomic response of Arabidopsis thaliana to tunicamycin-induced endoplasmic reticulum stress. Plant Biotechnol. 2010, 27, 161–171. [Google Scholar] [CrossRef]
- Jiang, J.; Liu, X.; Liu, C.; Liu, G.; Li, S.; Wang, L. Integrating Omics and Alternative Splicing Reveals Insights into Grape Response to High Temperature. Plant Physiol. 2017, 173, 1502–1518. [Google Scholar] [CrossRef] [PubMed]
- Frank, G.; Pressman, E.; Ophir, R.; Althan, L.; Shaked, R.; Freedman, M.; Shen, S.; Firon, N. Transcriptional profiling of maturing tomato (Solanum lycopersicum L.) microspores reveals the involvement of heat shock proteins, ROS scavengers, hormones, and sugars in the heat stress response. J. Exp. Bot. 2009, 60, 3891–3908. [Google Scholar] [CrossRef] [PubMed]
- Queitsch, C.; Hong, S.W.; Vierling, E.; Lindquist, S. Heat shock protein 101 plays a crucial role in thermotolerance in Arabidopsis. Plant Cell 2000, 12, 479–492. [Google Scholar] [CrossRef] [PubMed]
- Mishra, R.C.; Grover, A. ClpB/Hsp100 proteins and heat stress tolerance in plants. Crit. Rev. Biotechnol. 2015, 36, 862–874. [Google Scholar] [CrossRef] [PubMed]
- McLoughlin, F.; Basha, E.; Fowler, M.E.; Kim, M.; Bordowitz, J.; Katiyar-Agarwal, S.; Vierling, E. Class I and II small heat-shock proteins protect protein translation factors during heat stress. Plant Physiol. 2016, 172, 1221–1236. [Google Scholar] [CrossRef]
- Scharf, K.D.; Berberich, T.; Ebersberger, I.; Nover, L. The plant heat stress transcription factor (Hsf) family: Structure, function and evolution. Biochim. Biophys. Acta 2012, 1819, 104–119. [Google Scholar] [CrossRef]
- Berz, J.; Simm, S.; Schuster, S.; Scharf, K.; Schleiff, E.; Ebersberger, I. HEATSTER: A Database and Web Server for Identification and Classification of Heat Stress Transcription Factors in Plants. Bioinform. Biol. Insights 2019, 13. [Google Scholar] [CrossRef]
- Nover, L.; Bharti, K.; Döring, P.; Mishra, S.K.; Ganguli, A.; Scharf, K.D. Arabidopsis and the heat stress transcription factor world: How many heat stress transcription factors do we need? Cell Stress Chaperones 2001, 6, 177–189. [Google Scholar] [CrossRef]
- von Koskull-Döring, P.; Scharf, K.-D.; Nover, L. The diversity of plant heat stress transcription factors. Trends Plant Sci. 2007, 12, 452–457. [Google Scholar] [CrossRef] [PubMed]
- Mishra, S.K.; Tripp, J.; Winkelhaus, S.; Tschiersch, B.; Theres, K.; Nover, L.; Scharf, K.-D. In the complex family of heat stress transcription factors, HsfA1 has a unique role as master regulator of thermotolerance in tomato. Genes Dev. 2002, 16, 1555–1567. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.C.; Liao, H.T.; Charng, Y.Y. The role of class A1 heat shock factors (HSFA1s) in response to heat and other stresses in Arabidopsis. Plant Cell Environ. 2011, 34, 738–751. [Google Scholar] [CrossRef] [PubMed]
- Schramm, F.; Ganguli, A.; Kiehlmann, E.; Englich, G.; Walch, D.; Von Koskull-Döring, P. The heat stress transcription factor HsfA2 serves as a regulatory amplifier of a subset of genes in the heat stress response in Arabidopsis. Plant Mol. Biol. 2006, 60, 759–772. [Google Scholar] [CrossRef] [PubMed]
- Charng, Y.-Y.; Liu, H.-C.; Liu, N.-Y.; Chi, W.-T.; Wang, C.-N.; Chang, S.-H.; Wang, T.-T. A Heat-Inducible Transcription Factor, HsfA2, Is Required for Extension of Acquired Thermotolerance in Arabidopsis. Plant Physiol. 2006, 143, 251–262. [Google Scholar] [CrossRef] [PubMed]
- Fragkostefanakis, S.; Mesihovic, A.; Simm, S.; Paupière, M.J.; Hu, Y.; Paul, P.; Mishra, S.K.; Tschiersch, B.; Theres, K.; Bovy, A.; et al. HsfA2 Controls the Activity of Developmentally and Stress-Regulated Heat Stress Protection Mechanisms in Tomato Male Reproductive Tissues. Plant Physiol. 2016, 170, 2461–2477. [Google Scholar] [CrossRef]
- Bharti, K.; Von Koskull-Döring, P.; Bharti, S.; Kumar, P.; Tintschl-Körbitzer, A.; Treuter, E.; Nover, L. Tomato Heat Stress Transcription Factor HsfB1 Represents a Novel Type of General Transcription Coactivator with a Histone-Like Motif Interacting with the Plant CREB Binding Protein Ortholog HAC1. Plant Cell 2004, 16, 1521–1535. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, M.; Mitsuda, N.; Ohme-Takagi, M. Arabidopsis HsfB1 and HsfB2b act as repressors of the expression of heat-inducible Hsfs but positively regulate the acquired thermotolerance. Plant Physiol. 2011, 157, 1243–1254. [Google Scholar] [CrossRef]
- Kumar, M.; Busch, W.; Birke, H.; Kemmerling, B.; Nürnberger, T.; Schöffl, F. Heat shock factors HsfB1 and HsfB2b are involved in the regulation of Pdf1.2 expression and pathogen resistance in Arabidopsis. Mol. Plant 2009, 2, 152–165. [Google Scholar] [CrossRef]
- Fragkostefanakis, S.; Simm, S.; El-Shershaby, A.; Hu, Y.; Bublak, D.; Mesihovic, A.; Darm, K.; Mishra, S.K.; Tschiersch, B.; Theres, K.; et al. The repressor and co-activator HsfB1 regulates the major heat stress transcription factors in tomato. Plant Cell Environ. 2018, 42, 874–890. [Google Scholar] [CrossRef] [PubMed]
- Wunderlich, M.; Groß-Hardt, R.; Schöffl, F. Heat shock factor HSFB2a involved in gametophyte development of Arabidopsis thaliana and its expression is controlled by a heat-inducible long non-coding antisense RNA. Plant Mol. Biol. 2014, 85, 541–550. [Google Scholar] [CrossRef] [PubMed]
- Margaritopoulou, T.; Kryovrysanaki, N.; Megkoula, P.; Prassinos, C.; Samakovli, D.; Milioni, D.; Hatzopoulos, P. HSP90 canonical content organizes a molecular scaffold mechanism to progress flowering. Plant J. 2016, 87, 174–187. [Google Scholar] [CrossRef] [PubMed]
- Meiri, D.; Breiman, A. Arabidopsis ROF1 (FKBP62) modulates thermotolerance by interacting with HSP90.1 and affecting the accumulation of HsfA2-regulated sHSPs. Plant J. 2009, 59, 387–399. [Google Scholar] [CrossRef] [PubMed]
- Meiri, D.; Tazat, K.; Cohen-Peer, R.; Farchi-Pisanty, O.; Aviezer-Hagai, K.; Avni, A.; Breiman, A. Involvement of arabidopsis ROF2 (FKBP65) in thermotolerance. Plant Mol. Biol. 2010, 72, 191–203. [Google Scholar] [CrossRef]
- Ohama, N.; Kusakabe, K.; Mizoi, J.; Zhao, H.; Kidokoro, S.; Koizumi, S.; Takahashi, F.; Ishida, T.; Yanagisawa, S.; Shinozaki, K.; et al. The transcriptional cascade in the heat stress response of Arabidopsis is strictly regulated at the expression levels of transcription factors. Plant Cell 2016, 28, 181–201. [Google Scholar] [CrossRef]
- Hahn, A.; Bublak, D.; Schleiff, E.; Scharf, K.-D. Crosstalk between Hsp90 and Hsp70 chaperones and heat stress transcription factors in tomato. Plant Cell 2011, 23, 741–755. [Google Scholar] [CrossRef] [PubMed]
- Baniwal, S.K.; Bharti, K.; Chan, K.Y.; Fauth, M.; Ganguli, A.; Kotak, S.; Mishra, S.K.; Nover, L.; Port, M.; Scharf, K.D.; et al. Heat stress response in plants: A complex game with chaperones and more than twenty heat stress transcription factors. J. Biosci. 2004, 29, 471–487. [Google Scholar] [CrossRef]
- Röth, S.; Mirus, O.; Bublak, D.; Klaus-Dieter, S.; Schleiff, E. DNA-binding and repressor function are prerequisite for the turnover of the tomato heat stress transcription factor HsfB1. Plant J. 2016, 89, 31–44. [Google Scholar] [CrossRef]
- Fox, S.G.; Morimoto, R.I.; Satyal, S.H.; Chen, D.; Kramer, J.M. Negative regulation of the heat shock transcriptional response by HSBP1. Genes Dev. 2008, 12, 1962–1974. [Google Scholar]
- Liu, X.; Xu, L.; Liu, Y.; Tong, X.; Zhu, G.; Zhang, X.C.; Li, X.; Rao, Z. Crystal structure of the hexamer of human heat shock factor binding protein 1. Proteins Struct. Funct. Bioinforma. 2009, 75, 1–11. [Google Scholar] [CrossRef]
- Hsu, S.-F.; Lai, H.-C.; Jinn, T.-L. Cytosol-Localized Heat Shock Factor-Binding Protein, AtHSBP, Functions as a Negative Regulator of Heat Shock Response by Translocation to the Nucleus and Is Required for Seed Development in Arabidopsis. Plant Physiol. 2010, 153, 773–784. [Google Scholar] [CrossRef] [PubMed]
- Fu, S. empty pericarp2 Encodes a Negative Regulator of the Heat Shock Response and Is Required for Maize Embryogenesis. Plant Cell 2002, 14, 3119–3132. [Google Scholar] [CrossRef] [PubMed]
- Fu, S.; Rogowsky, Æ.P.; Nover, Æ.L.; Scanlon, M.J. The maize heat shock factor-binding protein paralogs EMP2 and HSBP2 interact non-redundantly with specific heat shock factors. Planta 2006, 224, 42–52. [Google Scholar] [CrossRef] [PubMed]
- Rana, R.M.; Dong, S.; Tang, H.; Ahmad, F.; Zhang, H. Functional analysis of OsHSBP1 and OsHSBP2 revealed their involvement in the heat shock response in rice (Oryza sativa L.). J. Exp. Bot. 2012, 63, 6003–6016. [Google Scholar] [CrossRef] [PubMed]
- Tomato, T.; Consortium, G. The tomato genome sequence provides insights into fleshy fruit evolution. Nature 2012, 485, 635–641. [Google Scholar]
- Minoia, S.; Cellini, F.; Bendahmane, A.; D’Onofrio, O.; Petrozza, A.; Carriero, F.; Piron, F.; Mosca, G.; Sozio, G. A new mutant genetic resource for tomato crop improvement by TILLING technology. BMC Res. Notes 2010, 3, 69. [Google Scholar] [CrossRef] [PubMed]
- Dalmais, M.; Schmidt, J.; Le Signor, C.; Moussy, F.; Burstin, J.; Savois, V.; Aubert, G.; Brunaud, V.; de Oliveira, Y.; Guichard, C.; et al. UTILLdb, a Pisum sativum in silico forward and reverse genetics tool. Genome Biol. 2008, 9, R43. [Google Scholar] [CrossRef]
- Triques, K.; Sturbois, B.; Gallais, S.; Dalmais, M.; Chauvin, S.; Clepet, C.; Aubourg, S.; Rameau, C.; Caboche, M.; Bendahmane, A. Characterization of Arabidopsis thaliana mismatch specific endonucleases: Application to mutation discovery by TILLING in pea. Plant J. 2007, 51, 1116–1125. [Google Scholar] [CrossRef]
- Ng, P.C.; Henikoff, S. SIFT: Predicting amino acid changes that affect protein function. Nucleic Acids Res. 2003, 31, 3812–3814. [Google Scholar] [CrossRef]
- Mazzucato, A.; Cellini, F.; Bouzayen, M.; Zouine, M.; Mila, I.; Minoia, S.; Petrozza, A.; Picarella, M.E.; Ruiu, F.; Carriero, F. A TILLING allele of the tomato Aux/IAA9 gene offers new insights into fruit set mechanisms and perspectives for breeding seedless tomatoes. Mol. Breed. 2015, 35, 22. [Google Scholar] [CrossRef]
- Petrozza, A.; Santaniello, A.; Summerer, S.; Di Tommaso, G.; Di Tommaso, D.; Paparelli, E.; Piaggesi, A.; Perata, P.; Cellini, F. Physiological responses to Megafol® treatments in tomato plants under drought stress: A phenomic and molecular approach. Sci. Hortic. 2014, 174, 185–192. [Google Scholar] [CrossRef]
- Eberius, M.; Lima-Guerra, J. High-throughput plant phenotyping-Data acquisition, transformation, and analysis. In Bioinformatics: Tools and Applications; Springer: New York, NY, USA, 2007; ISBN 9783540241669. [Google Scholar]
- Treuter, E.; Nover, L.; Ohme, K.; Scharf, K.D. Promoter specificity and deletion analysis of three heat stress transcription factors of tomato. Mol. Gen. Genet. 1993, 240, 113–125. [Google Scholar] [CrossRef] [PubMed]
- Chan-Schaminet, K.Y.; Baniwal, S.K.; Bublak, D.; Nover, L.; Scharf, K.-D. Specific interaction between tomato HsfA1 and HsfA2 creates hetero-oligomeric superactivator complexes for synergistic activation of heat stress gene expression. J. Biol. Chem. 2009, 284, 20848–20857. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Missbach, S.; Weis, B.L.; Martin, R.; Simm, S.; Bohnsack, M.T.; Schleiff, E. 40S Ribosome Biogenesis Co-Factors Are Essential for Gametophyte and Embryo Development. PLoS ONE 2013, 8, e54084. [Google Scholar] [CrossRef] [PubMed]
- Zdobnov, E.M.; Tegenfeldt, F.; Kuznetsov, D.; Waterhouse, R.M.; Simao, F.A.; Ioannidis, P.; Seppey, M.; Loetscher, A.; Kriventseva, E.V. OrthoDB v9.1: Cataloging evolutionary and functional annotations for animal, fungal, plant, archaeal, bacterial and viral orthologs. Nucleic Acids Res. 2017, 45, D744–D749. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Yan, R.; Roy, A.; Xu, D.; Poisson, J.; Zhang, Y. The I-TASSER suite: Protein structure and function prediction. Nat. Methods 2014, 12, 7–8. [Google Scholar] [CrossRef]
- Wood, C.W.; Woolfson, D.N. CCBuilder 2.0: Powerful and accessible coiled-coil modeling. Protein Sci. 2018, 27, 103–111. [Google Scholar] [CrossRef] [PubMed]
- Zouine, M.; Maza, E.; Djari, A.; Lauvernier, M.; Frasse, P.; Smouni, A.; Pirrello, J.; Bouzayen, M. TomExpress, a unified tomato RNA-Seq platform for visualization of expression data, clustering and correlation networks. Plant J. 2017, 92, 727–735. [Google Scholar] [CrossRef] [PubMed]
- Marko, D.; Briglia, N.; Summerer, S.; Petrozza, A.; Cellini, F.; Iannacone, R. High-Throughput Phenotyping in Plant Stress Response: Methods and Potential Applications to Polyamine Field. In Polyamines; Humana Press: New York, NY, USA, 2018; Volume 1694, pp. 373–388. ISBN 978-1-4939-7397-2. [Google Scholar]
- Giorno, F.; Wolters-Arts, M.; Grillo, S.; Scharf, K.D.; Vriezen, W.H.; Mariani, C. Developmental and heat stress-regulated expression of HsfA2 and small heat shock proteins in tomato anthers. J. Exp. Bot. 2010, 61, 453–462. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Marko, D.; El-shershaby, A.; Carriero, F.; Summerer, S.; Petrozza, A.; Iannacone, R.; Schleiff, E.; Fragkostefanakis, S. Identification and Characterization of a Thermotolerant TILLING Allele of Heat Shock Binding Protein 1 in Tomato. Genes 2019, 10, 516. https://doi.org/10.3390/genes10070516
Marko D, El-shershaby A, Carriero F, Summerer S, Petrozza A, Iannacone R, Schleiff E, Fragkostefanakis S. Identification and Characterization of a Thermotolerant TILLING Allele of Heat Shock Binding Protein 1 in Tomato. Genes. 2019; 10(7):516. https://doi.org/10.3390/genes10070516
Chicago/Turabian StyleMarko, Dominik, Asmaa El-shershaby, Filomena Carriero, Stephan Summerer, Angelo Petrozza, Rina Iannacone, Enrico Schleiff, and Sotirios Fragkostefanakis. 2019. "Identification and Characterization of a Thermotolerant TILLING Allele of Heat Shock Binding Protein 1 in Tomato" Genes 10, no. 7: 516. https://doi.org/10.3390/genes10070516
APA StyleMarko, D., El-shershaby, A., Carriero, F., Summerer, S., Petrozza, A., Iannacone, R., Schleiff, E., & Fragkostefanakis, S. (2019). Identification and Characterization of a Thermotolerant TILLING Allele of Heat Shock Binding Protein 1 in Tomato. Genes, 10(7), 516. https://doi.org/10.3390/genes10070516





