EcoTILLING Reveals Natural Allelic Variations in Starch Synthesis Key Gene TaSSIV and Its Haplotypes Associated with Higher Thousand Grain Weight
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material and DNA Extraction
2.2. DNA Isolation and Molecular Analyses
2.3. Primers Development
2.4. EcoTILLING
2.5. Sequencing of Variants
2.6. Development of Functional Markers (FMs) Using KASP
2.7. Association between SNPs and Agronomic Traits
3. Results
3.1. Natural Variation of TaSSIV in Accessions
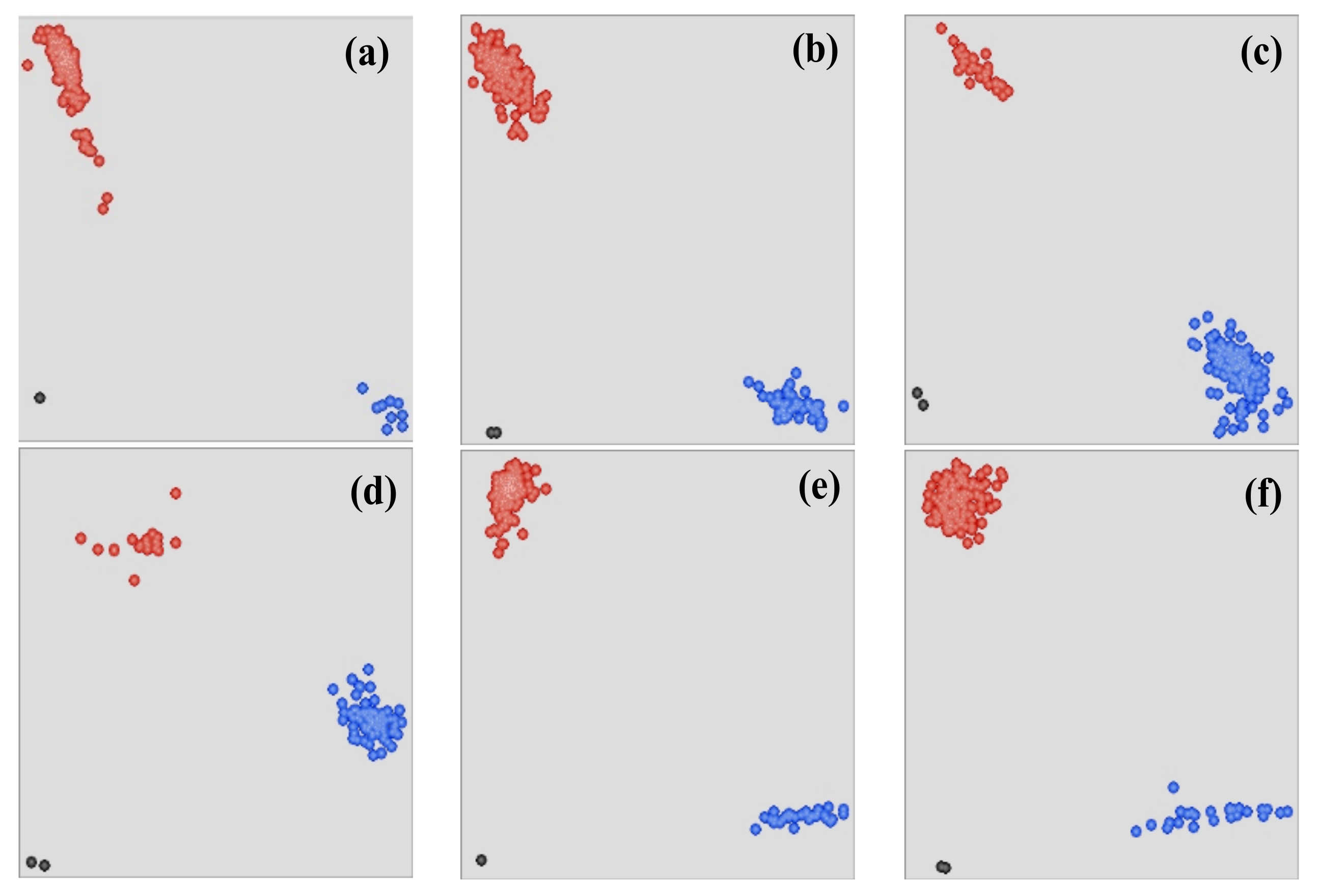
3.2. KASP Marker Development
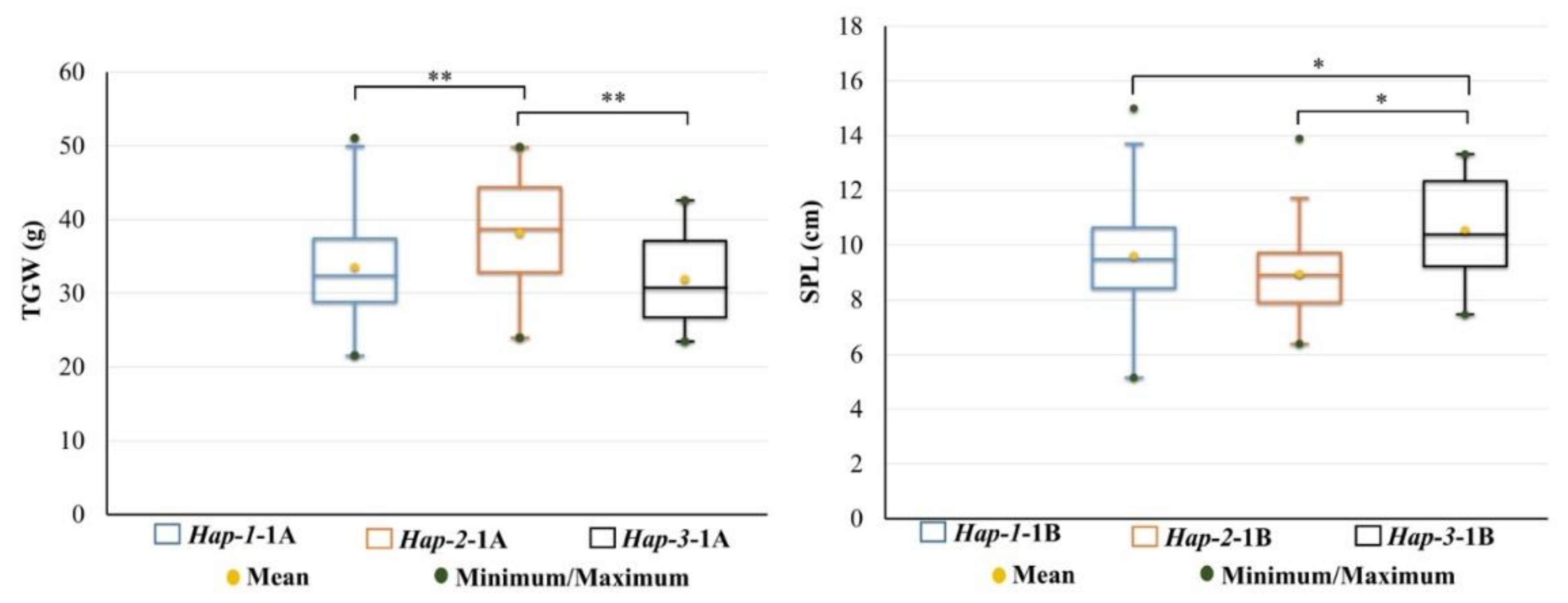
3.3. Association Between Haplotypes and Yield-Related Traits
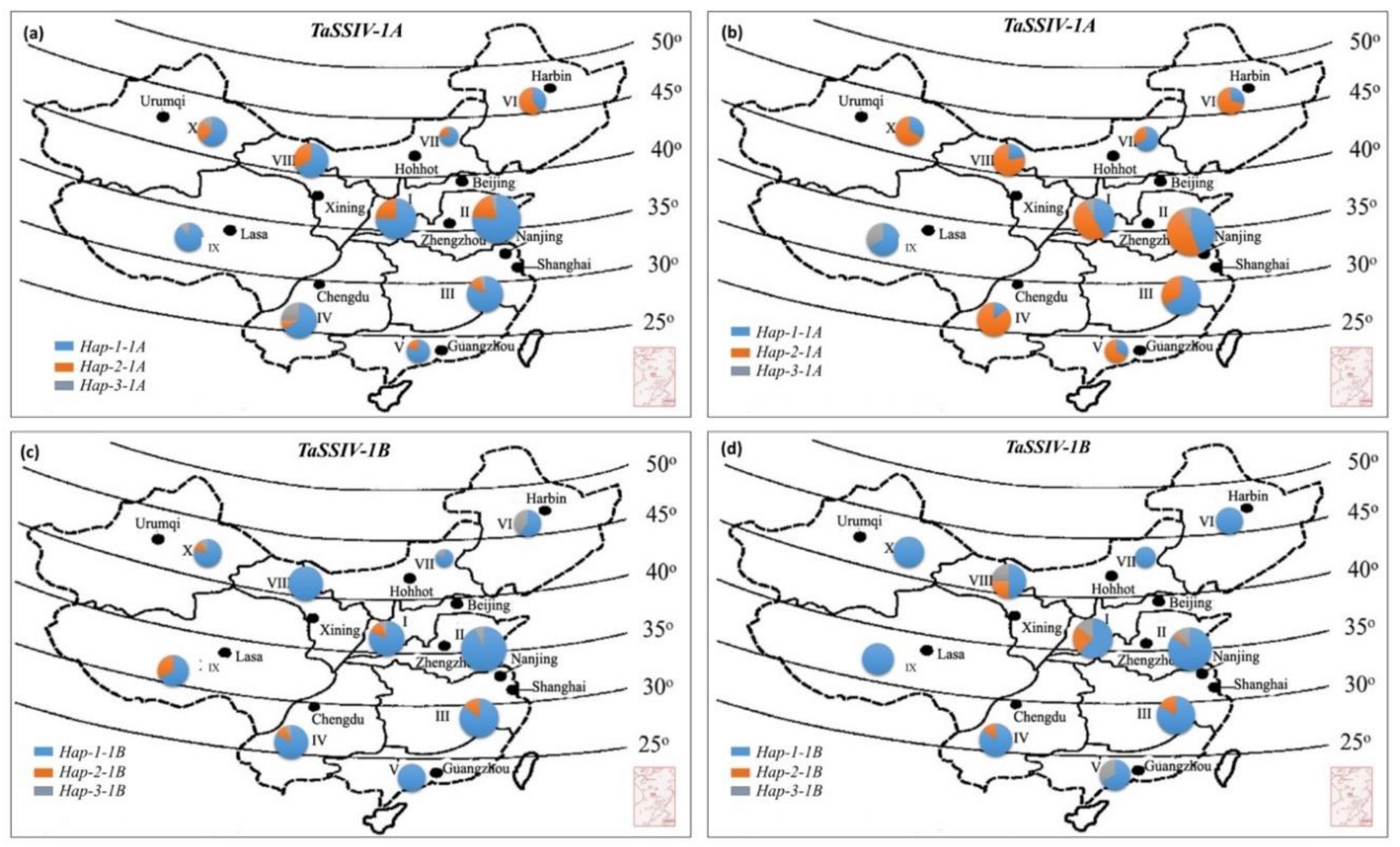
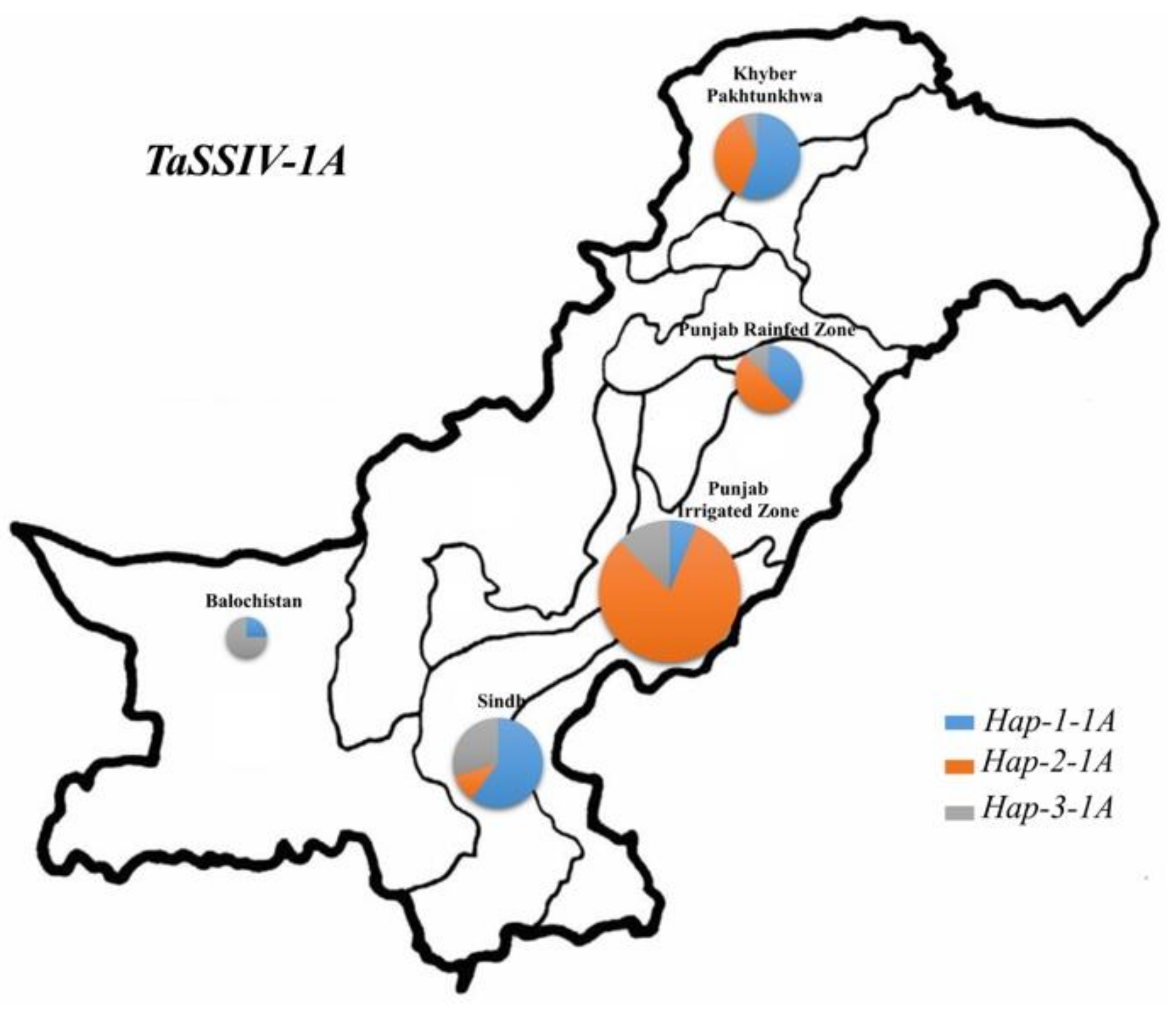
3.4. Geographic Distribution of TaSSIV-A and TaSSIV-B Haplotypes
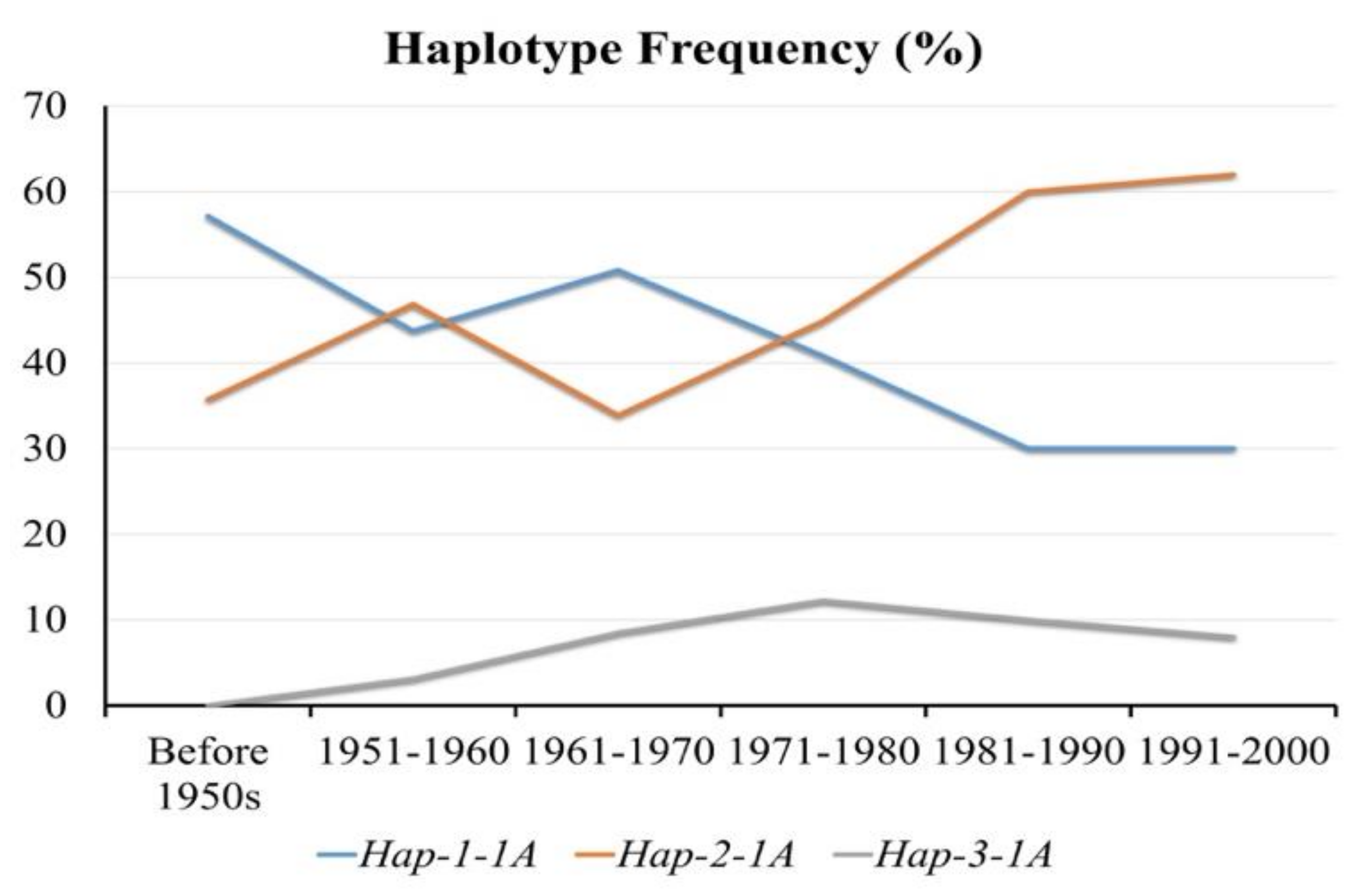
3.5. Positive Selection of Hap-2-1A in China’s Wheat Breeding Process
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Martin, C.; Smith, A.M. Starch biosynthesis. Plant Cell 1995, 7, 971. [Google Scholar] [CrossRef] [PubMed]
- Rossi, A.; Kontarakis, Z.; Gerri, C.; Nolte, H.; Hölper, S.; Krüger, M.; Stainier, D.Y. Genetic compensation induced by deleterious mutations but not gene knockdowns. Nature 2015, 524, 230. [Google Scholar] [CrossRef] [PubMed]
- Serrago, R.A.; Alzueta, I.; Savin, R.; Slafer, G.A. Understanding grain yield responses to source–sink ratios during grain filling in wheat and barley under contrasting environments. Field Crops Res. 2013, 150, 42–51. [Google Scholar] [CrossRef]
- Hou, J.; Jiang, Q.; Hao, C.; Wang, Y.; Zhang, H.; Zhang, X. Global selection on sucrose synthase haplotypes during a century of wheat breeding. Plant Physiol. 2014, 164, 1918–1929. [Google Scholar] [CrossRef] [PubMed]
- Ball, S.G.; Morell, M.K. From bacterial glycogen to starch: Understanding the biogenesis of the plant starch granule. Annu. Rev. Plant Biol. 2003, 54, 207–233. [Google Scholar] [CrossRef]
- Smith, A.M.; Zeeman, S.C.; Smith, S.M. Starch degradation. Annu. Rev. Plant Biol. 2005, 56, 73–98. [Google Scholar] [CrossRef] [PubMed]
- Leterrier, M.; Holappa, L.D.; Broglie, K.E.; Beckles, D.M. Cloning, characterisation and comparative analysis of a starch synthase IV gene in wheat: Functional and evolutionary implications. BMC Plant Biol. 2008, 8, 98. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Wang, J.; Zhang, W.; Li, C.; Yu, J.; Wang, S. Molecular order and functional properties of starches from three waxy wheat varieties grown in China. Food Chem. 2015, 181, 43–50. [Google Scholar] [CrossRef]
- Roldán, I.; Wattebled, F.; Mercedes Lucas, M.; Delvallé, D.; Planchot, V.; Jiménez, S.; Pérez, R.; Ball, S.; D’hulst, C.; Mérida, Á. The phenotype of soluble starch synthase IV defective mutants of Arabidopsis thaliana suggests a novel function of elongation enzymes in the control of starch granule formation. Plant J. 2007, 49, 492–504. [Google Scholar] [CrossRef]
- Szydlowski, N.; Ragel, P.; Raynaud, S.; Lucas, M.M.; Roldán, I.; Montero, M.; Muñoz, F.J.; Ovecka, M.; Bahaji, A.; Planchot, V. Starch granule initiation in Arabidopsis requires the presence of either class IV or class III starch synthases. Plant Cell 2009, 21, 2443–2457. [Google Scholar] [CrossRef]
- Dian, W.; Jiang, H.; Chen, Q.; Liu, F.; Wu, P. Cloning and characterization of the granule-bound starch synthase II gene in rice: Gene expression is regulated by the nitrogen level, sugar and circadian rhythm. Planta 2003, 218, 261–268. [Google Scholar] [CrossRef]
- Jiang, H.; Dian, W.; Liu, F.; Wu, P. Molecular cloning and expression analysis of three genes encoding starch synthase II in rice. Planta 2004, 218, 1062–1070. [Google Scholar] [CrossRef]
- Smith, A.M.; Stitt, M. Coordination of carbon supply and plant growth. Plant Cell Environ. 2007, 30, 1126–1149. [Google Scholar] [CrossRef]
- Gámez-Arjona, F.M.; Li, J.; Raynaud, S.; Baroja-Fernández, E.; Muñoz, F.J.; Ovecka, M.; Ragel, P.; Bahaji, A.; Pozueta-Romero, J.; Mérida, Á. Enhancing the expression of starch synthase class IV results in increased levels of both transitory and long-term storage starch. Plant Biotechnol. J. 2011, 9, 1049–1060. [Google Scholar] [CrossRef] [PubMed]
- Schwarte, S.; Brust, H.; Steup, M.; Tiedemann, R. Intraspecific sequence variation and differential expression in starch synthase genes of Arabidopsis thaliana. BMC Res. Notes 2013, 6, 84. [Google Scholar] [CrossRef] [PubMed]
- Toyosawa, Y.; Kawagoe, Y.; Matsushima, R.; Crofts, N.; Ogawa, M.; Fukuda, M.; Kumamaru, T.; Okazaki, Y.; Kusano, M.; Saito, K. Deficiency of starch synthase IIIa and IVb alters starch granule morphology from polyhedral to spherical in rice endosperm. Plant Physiol. 2016, 170, 1255–1270. [Google Scholar] [CrossRef]
- Li, Z.; Mouille, G.; Kosar-Hashemi, B.; Rahman, S.; Clarke, B.; Gale, K.R.; Appels, R.; Morell, M.K. The structure and expression of the wheat starch synthase III gene. Motifs in the expressed gene define the lineage of the starch synthase III gene family. Plant Physiol. 2000, 123, 613–624. [Google Scholar] [CrossRef] [PubMed]
- Guo, H.; Liu, Y.; Li, X.; Yan, Z.; Xie, Y.; Xiong, H.; Zhao, L.; Gu, J.; Zhao, S.; Liu, L. Novel mutant alleles of the starch synthesis gene TaSSIVb-D result in the reduction of starch granule number per chloroplast in wheat. BMC Genom. 2017, 18, 358. [Google Scholar] [CrossRef]
- Mccallum, C.M.; Comai, L.; Greene, E.A.; Henikoff, S. Targeting induced local lesions IN genomes (TILLING) for plant functional genomics. Plant Physiol. 2000, 123, 439. [Google Scholar] [CrossRef] [PubMed]
- Comai, L.; Young, K.; Till, B.J.; Reynolds, S.H.; Greene, E.A.; Codomo, C.A.; Enns, L.C.; Johnson, J.E.; Burtner, C.; Odden, A.R. Efficient discovery of DNA polymorphisms in natural populations by EcoTILLING. Plant J. 2004, 37, 778–786. [Google Scholar] [CrossRef] [PubMed]
- Till, B.J.; Comai, L.; Henikoff, S.; Zerr, T. A protocol for TILLING and EcoTILLING in plants and animals. Nat. Protoc. 2006, 1, 2465. [Google Scholar] [CrossRef]
- Till, B.J.; Cooper, J.; Tai, T.H.; Colowit, P.; Greene, E.A.; Henikoff, S.; Comai, L. Discovery of chemically induced mutations in rice by TILLING. BMC Plant Biol. 2007, 7, 19. [Google Scholar] [CrossRef] [PubMed]
- Raghavan, C.; Naredo, M.E.B.; Wang, H.; Atienza, G.; Liu, B.; Qiu, F.; McNally, K.L.; Leung, H. Rapid method for detecting SNPs on agarose gels and its application in candidate gene mapping. Mol. Breed. 2006, 19, 87–101. [Google Scholar] [CrossRef]
- Wang, T.L.; Uauy, C.; Robson, F.; Till, B. TILLING in extremis. Plant Biotech. J. 2012, 10, 761–772. [Google Scholar] [CrossRef] [PubMed]
- Xia, Y.; Ning, Z.; Bai, G.; Li, R.; Yan, G.; Siddique, K.H.; Baum, M.; Guo, P. Allelic variations of a light harvesting chlorophyll a/b-binding protein gene (Lhcb1) associated with agronomic traits in barley. PLoS ONE 2012, 7, e37573. [Google Scholar] [CrossRef] [PubMed]
- Negrao, S.; Almadanim, M.C.; Pires, I.S.; Abreu, I.A.; Maroco, J.; Courtois, B.; Gregorio, G.B.; McNally, K.L.; Oliveira, M.M. New allelic variants found in key rice salt-tolerance genes: An association study. Plant Biotechnol. J. 2013, 11, 87–100. [Google Scholar] [CrossRef]
- Zeng, Y.D.; Sun, J.L.; Bu, S.H.; Deng, K.S.; Tao, T.; Zhang, Y.M.; Zhang, T.Z.; Du, X.M.; Zhou, B.L. EcoTILLING revealed SNPs in GhSus genes that are associated with fiber- and seed-related traits in upland cotton. Sci. Rep. 2016, 6, 29250. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ma, X.; Sajjad, M.; Wang, J.; Yang, W.; Sun, J.; Li, X.; Zhang, A.; Liu, D. Diversity, distribution of Puroindoline genes and their effect on kernel hardness in a diverse panel of Chinese wheat germplasm. BMC Plant Biol. 2017, 17, 158. [Google Scholar] [CrossRef] [PubMed]
- Russell, J.; Mascher, M.; Dawson, I.K.; Kyriakidis, S.; Calixto, C.; Freund, F.; Bayer, M.; Milne, I.; Marshall-Griffiths, T.; Heinen, S. Exome sequencing of geographically diverse barley landraces and wild relatives gives insights into environmental adaptation. Nat. Genet. 2016, 48, 1024. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Li, P.; Zou, C.; Lu, Y.; Xie, C.; Zhang, X.; Prasanna, B.M.; Olsen, M.S. Enhancing genetic gain in the era of molecular breeding. J. Exp. Bot. 2017, 68, 2641–2666. [Google Scholar] [CrossRef]
- Wang, L.; Ge, H.; Hao, C.; Dong, Y.; Zhang, X. Identifying loci influencing 1,000-kernel weight in wheat by microsatellite screening for evidence of selection during breeding. PLoS ONE 2012, 7, e29432. [Google Scholar] [CrossRef]
- Murray, M.G.; Thompson, W.F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 1980, 8, 4321–4325. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Sun, J.; Liu, D.; Yang, W.; Wang, D.; Tong, Y.; Zhang, A. Analysis of Pina and Pinb alleles in the micro-core collections of Chinese wheat germplasm by EcoTILLING and identification of a novel Pinb allele. J. Cereal Sci. 2008, 48, 836–842. [Google Scholar] [CrossRef]
- Ng, P.C.; Henikoff, S. SIFT: Predicting amino acid changes that affect protein function. Nucleic Acids Res. 2003, 31, 3812–3814. [Google Scholar] [CrossRef]
- Rehman, S.U.; Wang, J.; Chang, X.; Zhang, X.; Mao, X.; Jing, R. A wheat protein kinase gene TaSnRK2. 9-5A associated with yield contributing traits. Theor. Appl. Genet. 2019, 132, 907–919. [Google Scholar] [CrossRef]
- He, Z. A History of Wheat Breeding in China; Cimmyt: Texcoco, Edo Mex, Mexico, 2001. [Google Scholar]
- Moose, S.P.; Mumm, R.H. Molecular plant breeding as the foundation for 21st century crop improvement. Plant Physiol. 2008, 147, 969–977. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Shi, L.; Tian, F.; Ning, H.; Wu, X.; Long, Y.; Meng, J. Assessment of FAE1 polymorphisms in three Brassica species using EcoTILLING and their association with differences in seed erucic acid contents. BMC Plant Biol. 2010, 10, 137. [Google Scholar] [CrossRef] [PubMed]
- Mejlhede, N.; Kyjovska, Z.; Backes, G.; Burhenne, K.; Rasmussen, S.K.; Jahoor, A. EcoTILLING for the identification of allelic variation in the powdery mildew resistance genes mlo and Mla of barley. Plant Breed. 2006, 125, 461–467. [Google Scholar] [CrossRef]
- Semagn, K.; Babu, R.; Hearne, S.; Olsen, M. Single nucleotide polymorphism genotyping using Kompetitive Allele Specific PCR (KASP): Overview of the technology and its application in crop improvement. Mol. Breed. 2014, 33, 1–14. [Google Scholar] [CrossRef]
- Rasheed, A.; Wen, W.; Gao, F.; Zhai, S.; Jin, H.; Liu, J.; Guo, Q.; Zhang, Y.; Dreisigacker, S.; Xia, X. Development and validation of KASP assays for genes underpinning key economic traits in bread wheat. Theor. Appl. Genet. 2016, 129, 1843–1860. [Google Scholar] [CrossRef]
- Nasu, S.; Suzuki, J.; Ohta, R.; Hasegawa, K.; Yui, R.; Kitazawa, N.; Monna, L.; Minobe, Y. Search for and analysis of single nucleotide polymorphisms (SNPs) in rice (Oryza sativa, Oryza rufipogon) and establishment of SNP markers. DNA Res. 2002, 9, 163–171. [Google Scholar] [CrossRef] [PubMed]
- Stephens, J.C.; Schneider, J.A.; Tanguay, D.A.; Choi, J.; Acharya, T.; Stanley, S.E.; Jiang, R.; Messer, C.J.; Chew, A.; Han, J.H. Haplotype variation and linkage disequilibrium in 313 human genes. Science 2001, 293, 489–493. [Google Scholar] [CrossRef] [PubMed]
- Lorenz, A.J.; Hamblin, M.T.; Jean-Luc, J. Performance of single nucleotide polymorphisms versus haplotypes for genome-wide association analysis in barley. PLoS ONE 2010, 5, e14079. [Google Scholar] [CrossRef]
- Li, B.; Li, Q.; Mao, X.; Li, A.; Wang, J.; Chang, X.; Hao, C.; Zhang, X.; Jing, R. Two Novel AP2/EREBP Transcription Factor GenesTaPARGHave Pleiotropic Functions on Plant Architecture and Yield-Related Traits in Common Wheat. Front. Plant Sci. 2016, 7, 1191. [Google Scholar]
- Miao, L.; Mao, X.; Wang, J.; Liu, Z.; Zhang, B.; Li, W.; Chang, X.; Reynolds, M.; Wang, Z.; Jing, R. Elite Haplotypes of a Protein Kinase GeneTaSnRK2.3 Associated with Important Agronomic Traits in Common Wheat. Front. Plant Sci. 2017, 8, 341. [Google Scholar] [CrossRef] [PubMed]
- Bevan, M.W.; Uauy, C.; Wulff, B.B.; Zhou, J.; Krasileva, K.; Clark, M.D. Genomic innovation for crop improvement. Nature 2017, 543, 346. [Google Scholar] [CrossRef] [PubMed]
- Parry, M.A.; Reynolds, M.; Salvucci, M.E.; Raines, C.; Andralojc, P.J.; Zhu, X.-G.; Price, G.D.; Condon, A.G.; Furbank, R.T. Raising yield potential of wheat. II. Increasing photosynthetic capacity and efficiency. J. Exp. Bot. 2010, 62, 453–467. [Google Scholar] [CrossRef] [PubMed]
- Zhu, C.; Gore, M.; Buckler, E.S.; Yu, J. Status and prospects of association mapping in plants. Plant Genome 2008, 1, 5–20. [Google Scholar] [CrossRef]
- Brancourt-Hulmel, M.; Doussinault, G.; Lecomte, C.; Berard, P.; Le Buanec, B.; Trottet, M. Genetic improvement of agronomic traits of winter wheat cultivars released in France from 1946 to 1992. Crop Sci. 2003, 43, 37–45. [Google Scholar] [CrossRef]
- Canevara, M.; Romani, M.; Corbellini, M.; Perenzin, M.; Borghi, B. Evolutionary trends in morphological, physiological, agronomical and qualitative traits of Triticum aestivum L. cultivars bred in Italy since 1900. Euro. J. Agron. 1994, 3, 175–185. [Google Scholar] [CrossRef]
- Zhou, Y.; Zhu, H.; Cai, S.; He, Z.; Zhang, X.; Xia, X.; Zhang, G. Genetic improvement of grain yield and associated traits in the southern China winter wheat region: 1949 to 2000. Euphytica 2007, 157, 465–473. [Google Scholar] [CrossRef]
- Gaju, O.; Reynolds, M.; Sparkes, D.; Foulkes, M. Relationships between large-spike phenotype, grain number, and yield potential in spring wheat. Crop Sci. 2009, 49, 961–973. [Google Scholar] [CrossRef]
- Hanif, M.; Gao, F.; Liu, J.; Wen, W.; Zhang, Y.; Rasheed, A.; Cao, S. TaTGW6-A1 an ortholog of rice TGW6 is associated with the grain weight and yield in bread wheat. Mol. Breed. 2016, 36, 1–8. [Google Scholar] [CrossRef]
- Zhang, K.; Wang, J.; Zhang, L.; Rong, C.; Zhao, F.; Peng, T.; Li, H.; Cheng, D.; Liu, X.; Qin, H. Association analysis of genomic loci important for grain weight control in elite common wheat varieties cultivated with variable water and fertiliser supply. PLoS ONE 2013, 8, e57853. [Google Scholar] [CrossRef] [PubMed]
- Dong, L.; Wang, F.; Liu, T.; Dong, Z.; Li, A.; Jing, R.; Mao, L.; Li, Y.; Liu, X.; Zhang, K. Natural variation of TaGASR7-A1 affects grain length in common wheat under multiple cultivation conditions. Mol. Breed. 2014, 34, 937–947. [Google Scholar] [CrossRef]
- Jiang, Q.; Hou, J.; Hao, C.; Wang, L.; Ge, H.; Dong, Y.; Zhang, X. The wheat (T. aestivum) sucrose synthase 2 gene (TaSus2) active in endosperm development is associated with yield traits. Funct. Integr. Genom. 2011, 11, 49–61. [Google Scholar] [CrossRef]
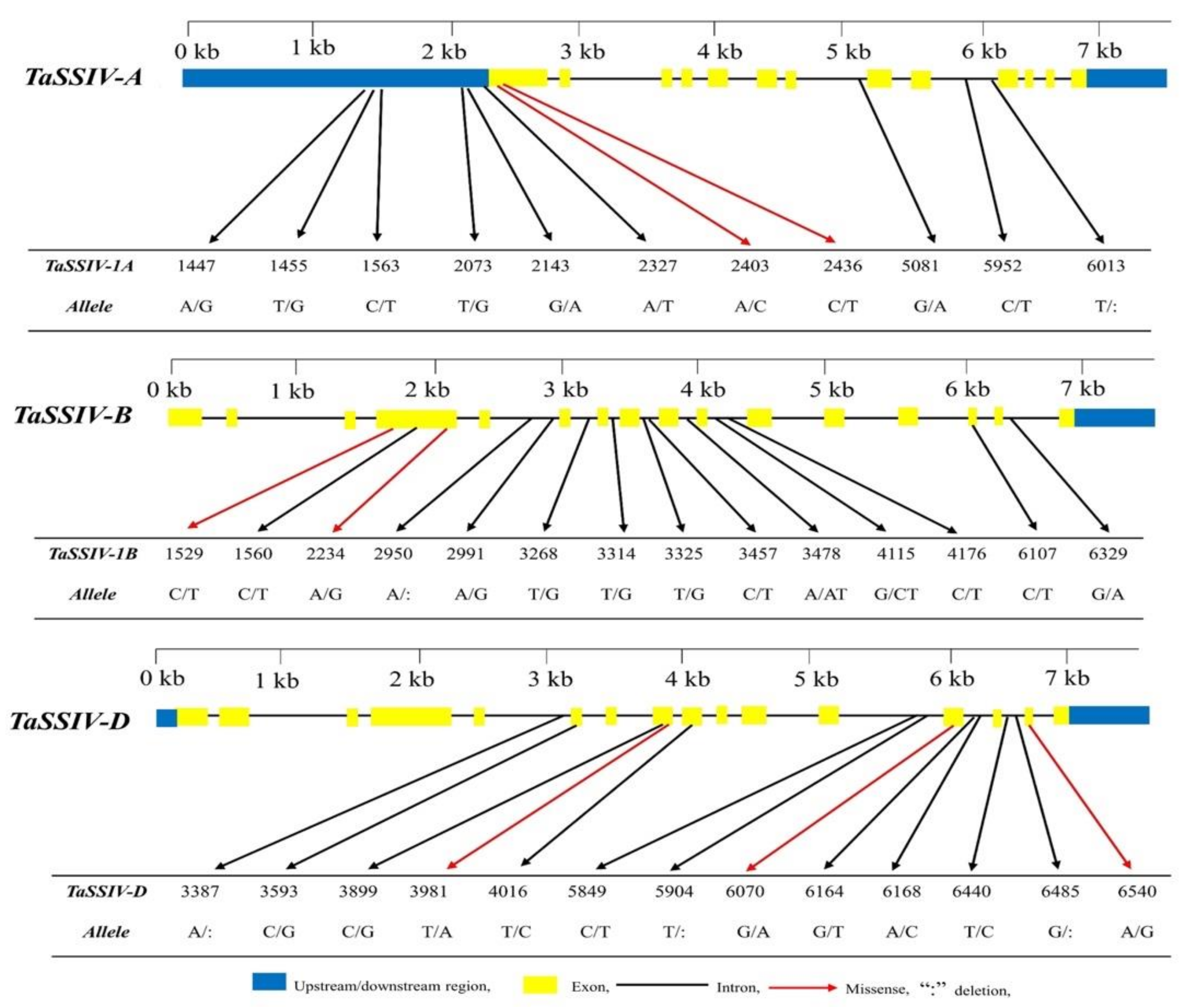
| No. | Gene | Genotype | Allele | Mutation Type | Variation in Amino Acid |
|---|---|---|---|---|---|
| 1 | TaSSIV-A | Laoqiaomai | A1447G | Non-coding | |
| 2 | Sankecun | T1455G | Non-coding | ||
| 3 | Wenmai-6 | C1563T | Non-coding | ||
| 4 | Yangmai-158 | A1673T | Non-coding | ||
| 5 | Yangmai-158 | C2436T | Missense | R77C | |
| 6 | Guinong-10 | T2073G | Non-coding | ||
| 6 | Zhongzhou-741 | A2403C | Missense | I66L | |
| 8 | Mahuaban | G5081A | Intron | ||
| 9 | Huoliyan | C5952T | Intron | ||
| 10 | Huangshuibai | T6013: | Intron | ||
| 11 | Guinong-11 | G2143A | Non-coding | ||
| 12 | TaSSIV-B | Dingxingzhai | C1529T | Missense | S149L |
| 13 | Pingyuan-50 | C1560T | Silent | F159= | |
| 14 | Dabaimai | A2234G | Missense | K384R | |
| 15 | Lumai-1 | T3314G | Intron | ||
| 16 | Lumai-1 | T3325G | Intron | ||
| 17 | Xinmai-18 | A3478ATT | Intron | ||
| 18 | Baipixiaomai | C6107T | Silent | I822= | |
| 19 | Xiaobaimai | G6329A | Intron | ||
| 20 | Shahkar-95 | A2950: | Intron | ||
| 21 | Drawar-97 | A2991G | Intron | ||
| 22 | Blue silver | C3457T | Intron | ||
| 23 | Lasani | G4115CTT | Intron | ||
| 24 | Kohistan | C4176T | Intron | ||
| 25 | Marvi | T3268G | Intron | ||
| 26 | TaSSIV-D | Zijuhong | T5904: | Intron | |
| 27 | Hongchumai | A6168C | Intron | ||
| 28 | Hongmai | G6485: | Intron | ||
| 29 | Hongmai | T6440C | Intron | ||
| 30 | Kabka-3 | A3387: | Intron | ||
| 31 | Chumai | C3593G | Silent | G454= | |
| 32 | Yangmai | C3899G | Silent | V505= | |
| 33 | Hongpidongmai | T3981A | Missense | S533T | |
| 34 | FSD-08 | C5849T | Intron | ||
| 35 | BWP-6309 | G6164T | Intron | ||
| 36 | Manthar | G6070A | Missense | V800I | |
| 37 | Kohistan-97 | A6540G | Missense | T841A | |
| 38 | Sehar | T4016C | Silent | Y533= |
| Gene | Haplotype | Position of Nucleotides | Gene | Haplotype | Position of Nucleotides | ||||
|---|---|---|---|---|---|---|---|---|---|
| TaSSIV-A | 1673 nt | 2403 nt | 2436 nt | 5952 nt | TaSSIV-B | 1560 nt | 6107 nt | ||
| Hap-1-1A | A | A | T | C | Hap-1-1B | C | C | ||
| Hap-2-1A | A | C | T | T | Hap-2-1B | T | C | ||
| Hap-3-1A | A | C | T | C | Hap-3-1B | C | T | ||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Irshad, A.; Guo, H.; Zhang, S.; Gu, J.; Zhao, L.; Xie, Y.; Xiong, H.; Zhao, S.; Ding, Y.; Ma, Y.; et al. EcoTILLING Reveals Natural Allelic Variations in Starch Synthesis Key Gene TaSSIV and Its Haplotypes Associated with Higher Thousand Grain Weight. Genes 2019, 10, 307. https://doi.org/10.3390/genes10040307
Irshad A, Guo H, Zhang S, Gu J, Zhao L, Xie Y, Xiong H, Zhao S, Ding Y, Ma Y, et al. EcoTILLING Reveals Natural Allelic Variations in Starch Synthesis Key Gene TaSSIV and Its Haplotypes Associated with Higher Thousand Grain Weight. Genes. 2019; 10(4):307. https://doi.org/10.3390/genes10040307
Chicago/Turabian StyleIrshad, Ahsan, Huijun Guo, Shunlin Zhang, Jiayu Gu, Linshu Zhao, Yongdun Xie, Hongchun Xiong, Shirong Zhao, Yuping Ding, Youzhi Ma, and et al. 2019. "EcoTILLING Reveals Natural Allelic Variations in Starch Synthesis Key Gene TaSSIV and Its Haplotypes Associated with Higher Thousand Grain Weight" Genes 10, no. 4: 307. https://doi.org/10.3390/genes10040307
APA StyleIrshad, A., Guo, H., Zhang, S., Gu, J., Zhao, L., Xie, Y., Xiong, H., Zhao, S., Ding, Y., Ma, Y., & Liu, L. (2019). EcoTILLING Reveals Natural Allelic Variations in Starch Synthesis Key Gene TaSSIV and Its Haplotypes Associated with Higher Thousand Grain Weight. Genes, 10(4), 307. https://doi.org/10.3390/genes10040307






