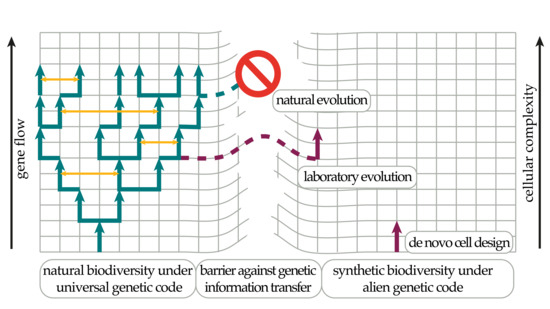
Alternative Biochemistries for Alien Life: Basic Concepts and Requirements for the Design of a Robust Biocontainment System in Genetic Isolation
Abstract
1. Introduction
2. Basic Considerations for an Alien Central Dogma
3. Altering the Meaning of the Genetic Code
3.1. Stop Codon Reassignment
3.2. Sense Codon Reassignment
3.3. Minimal Cell Design
4. Altering the Identity of the Genetic Code
4.1. Frameshift Codons
4.2. Synthetic Base Pairs
4.3. Alternative DNA Backbone Motifs
5. The Farther the Safer
6. Alienation Far beyond the Canonical Chemistries of Life
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Keasling, J.D. Manufacturing Molecules Through Metabolic Engineering. Science 2010, 330, 1355–1358. [Google Scholar] [CrossRef] [PubMed]
- Agostini, F.; Völler, J.S.; Koksch, B.; Acevedo-Rocha, C.G.; Kubyshkin, V.; Budisa, N. Biocatalysis with Unnatural Amino Acids: Enzymology Meets Xenobiology. Angew. Chem. Int. Ed. 2017, 56, 9680–9703. [Google Scholar] [CrossRef] [PubMed]
- Gao, G.-P.; Alvira, M.R.; Wang, L.; Calcedo, R.; Johnston, J.; Wilson, J.M. Novel adeno-associated viruses from rhesus monkeys as vectors for human gene therapy. Proc. Natl. Acad. Sci. USA 2002, 99, 11854–11859. [Google Scholar] [CrossRef]
- Noble, C.; Adlam, B.; Church, G.M.; Esvelt, K.M.; Nowak, M.A. Current CRISPR gene drive systems are likely to be highly invasive in wild populations. eLife 2018, 7, 1–30. [Google Scholar] [CrossRef] [PubMed]
- Hlihor, R.M.; Gavrilescu, M.; Tavares, T.; Favier, L.; Olivieri, G. Bioremediation: An Overview on Current Practices, Advances, and New Perspectives in Environmental Pollution Treatment. Biomed Res. Int. 2017, 2017, 3–5. [Google Scholar] [CrossRef] [PubMed]
- Dunbar, W.S. Biotechnology and the Mine of Tomorrow. Trends Biotechnol. 2017, 35, 79–89. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Rasheed, A.; Hickey, L.T.; He, Z. Fast-Forwarding Genetic Gain. Trends Plant Sci. 2018, 23, 184–186. [Google Scholar] [CrossRef]
- Gust, D.; Moore, T.A.; Moore, A.L. Mimicking photosynthetic solar energy transduction. Acc. Chem. Res. 2001, 34, 40–48. [Google Scholar] [CrossRef]
- Lynd, L.R.; Van Zyl, W.H.; McBride, J.E.; Laser, M. Consolidated bioprocessing of cellulosic biomass: An update. Curr. Opin. Biotechnol. 2005, 16, 577–583. [Google Scholar] [CrossRef]
- He, L.; Du, P.; Chen, Y.; Lu, H.; Cheng, X.; Chang, B.; Wang, Z. Advances in microbial fuel cells for wastewater treatment. Renew. Sustain. Energy Rev. 2017, 71, 388–403. [Google Scholar] [CrossRef]
- Rivnay, J.; Owens, R.M.; Malliaras, G.G. The rise of organic bioelectronics. Chem. Mater. 2014, 26, 679–685. [Google Scholar] [CrossRef]
- Shipman, S.L.; Nivala, J.; Macklis, J.D.; Church, G.M. CRISPR-Cas encoding of a digital movie into the genomes of a population of living bacteria. Nature 2017, 547, 345–349. [Google Scholar] [CrossRef]
- Schmidt, M.; de Lorenzo, V. Synthetic bugs on the loose: Containment options for deeply engineered (micro)organisms. Curr. Opin. Biotechnol. 2016, 38, 90–96. [Google Scholar] [CrossRef] [PubMed]
- Cohen, S.N.; Chang, A.C.Y.; Boyer, H.W.; Helling, R.B. Construction of Biologically Functional Bacterial Plasmids In Vitro. Proc. Natl. Acad. Sci. USA 1973, 70, 3240–3244. [Google Scholar] [CrossRef] [PubMed]
- Agapakis, C.M.; Silver, P.A. Synthetic biology: Exploring and exploiting genetic modularity through the design of novel biological networks. Mol. Biosyst. 2009, 5, 704–713. [Google Scholar] [CrossRef] [PubMed]
- Endy, D. Foundations for engineering biology. Nature 2005, 438, 449–453. [Google Scholar] [CrossRef] [PubMed]
- Kubyshkin, V.; Budisa, N. Synthetic alienation of microbial organisms by using genetic code engineering: Why and how? Biotechnol. J. 2017, 12, 1–8. [Google Scholar] [CrossRef]
- Kitano, H. Towards a theory of biological robustness. Mol. Syst. Biol. 2007, 3. [Google Scholar] [CrossRef]
- Ruiz-Mirazo, K.; Briones, C.; De La Escosura, A. Chemical roots of biological evolution: The origins of life as a process of development of autonomous functional systems. Open Biol. 2017, 7. [Google Scholar] [CrossRef]
- Syvanen, M. Cross-species gene transfer; implications for a new theory of evolution. J. Theor. Biol. 1985, 112, 333–343. [Google Scholar] [CrossRef]
- Vetsigian, K.; Woese, C.; Goldenfeld, N. Collective evolution and the genetic code. Proc. Natl. Acad. Sci. USA 2006, 103, 10696–10701. [Google Scholar] [CrossRef] [PubMed]
- Marliere, P. The farther, the safer: A manifesto for securely navigating synthetic species away from the old living world. Syst. Synth. Biol. 2009, 3, 77–84. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, M. Diffusion of synthetic biology: A challenge to biosafety. Syst. Synth. Biol. 2008, 2, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Mandell, D.J.; Lajoie, M.J.; Mee, M.T.; Takeuchi, R.; Kuznetsov, G.; Norville, J.E.; Gregg, C.J.; Stoddard, B.L.; Church, G.M. Biocontainment of genetically modified organisms by synthetic protein design. Nature 2015, 518, 55–60. [Google Scholar] [CrossRef] [PubMed]
- Agmon, N.; Tang, Z.; Yang, K.; Sutter, B.; Ikushima, S.; Cai, Y.; Caravelli, K.; Martin, J.A.; Sun, X.; Choi, W.J.; et al. Low escape-rate genome safeguards with minimal molecular perturbation of Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 2017, 114, E1470–E1479. [Google Scholar] [CrossRef] [PubMed]
- Wilson, D.J. Nih guidelines for research involving recombinant dna molecules. Account. Res. 1993, 3, 177–185. [Google Scholar] [CrossRef] [PubMed]
- Crick, F. Central dogma of molecular biology. Nature 1970, 227, 561. [Google Scholar] [CrossRef]
- Lee, J.W.; Chan, C.T.Y.; Slomovic, S.; Collins, J.J. Next-generation biocontainment systems for engineered organisms. Nat. Chem. Biol. 2018, 14, 530–537. [Google Scholar] [CrossRef]
- Whitford, C.M.; Dymek, S.; Kerkhoff, D.; März, C.; Schmidt, O.; Edich, M.; Droste, J.; Pucker, B.; Rückert, C.; Kalinowski, J. Auxotrophy to Xeno-DNA: An exploration of combinatorial mechanisms for a high-fidelity biosafety system for synthetic biology applications. J. Biol. Eng. 2018, 12. [Google Scholar] [CrossRef]
- Schwille, P. Bottom-Up Synthetic Biology: Engineering in a Tinkerer’s World. Science 2011, 333, 1252–1254. [Google Scholar] [CrossRef]
- Blain, J.C.; Szostak, J.W. Progress Toward Synthetic Cells. Annu. Rev. Biochem. 2014, 83, 615–640. [Google Scholar] [CrossRef] [PubMed]
- Hoesl, M.G.; Oehm, S.; Durkin, P.; Darmon, E.; Peil, L.; Aerni, H.R.; Rappsilber, J.; Rinehart, J.; Leach, D.; Söll, D.; et al. Chemical Evolution of a Bacterial Proteome. Angew. Chem. Int. Ed. 2015, 54, 10030–10034. [Google Scholar] [CrossRef] [PubMed]
- Hutchison, C.A.; Chuang, R.Y.; Noskov, V.N.; Assad-Garcia, N.; Deerinck, T.J.; Ellisman, M.H.; Gill, J.; Kannan, K.; Karas, B.J.; Ma, L.; et al. Design and synthesis of a minimal bacterial genome. Science 2016, 351. [Google Scholar] [CrossRef] [PubMed]
- Kubyshkin, V.; Acevedo-Rocha, C.G.; Budisa, N. On universal coding events in protein biogenesis. BioSystems 2018, 164, 16–25. [Google Scholar] [CrossRef] [PubMed]
- Ciliberti, S.; Martin, O.C.; Wagner, A. Robustness can evolve gradually in complex regulatory gene networks with varying topology. PLoS Comput. Biol. 2007, 3, e15. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.C.; Jewett, M.C.; Chin, J.W.; Voigt, C.A. Toward an orthogonal central dogma. Nat. Chem. Biol. 2018, 14, 103–106. [Google Scholar] [CrossRef] [PubMed]
- Thomas, C.M.; Nielsen, K.M. Mechanisms of, and barriers to, horizontal gene transfer between bacteria. Nat. Rev. Microbiol. 2005, 3, 711–721. [Google Scholar] [CrossRef]
- Herdewijn, P.; Marlière, P. Toward safe genetically modified organisms through the chemical diversification of nucleic acids. Chem. Biodivers. 2009, 6, 791–808. [Google Scholar] [CrossRef]
- Vargas-Rodriguez, O.; Sevostyanova, A.; Söll, D.; Crnković, A. Upgrading aminoacyl-tRNA synthetases for genetic code expansion. Curr. Opin. Chem. Biol. 2018, 46, 115–122. [Google Scholar] [CrossRef]
- Lajoie, M.J.; Kosuri, S.; Mosberg, J.A.; Gregg, C.J.; Zhang, D.; Church, G.M. Probing the limits of genetic recoding in essential genes. Science 2013, 342, 361–363. [Google Scholar] [CrossRef]
- Rovner, A.J.; Haimovich, A.D.; Katz, S.R.; Li, Z.; Grome, M.W.; Gassaway, B.M.; Amiram, M.; Patel, J.R.; Gallagher, R.R.; Rinehart, J.; et al. Recoded organisms engineered to depend on synthetic amino acids. Nature 2015, 518, 89–93. [Google Scholar] [CrossRef] [PubMed]
- Ostrov, N.; Landon, M.; Guell, M.; Kuznetsov, G.; Teramoto, J.; Cervantes, N.; Zhou, M.; Singh, K.; Napolitano, M.G.; Moosburner, M.; et al. Design, synthesis, and testing toward a 57-codon genome. Science 2016, 353, 819–822. [Google Scholar] [CrossRef] [PubMed]
- Lajoie, M.J.; Rovner, A.J.; Goodman, D.B.; Aerni, H.-R.; Haimovich, A.D.; Kuznetsov, G.; Mercer, J.A.; Wang, H.H.; Carr, P.A.; Mosberg, J.A.; et al. Genomically Recoded Organisms Expand Biological Functions. Science 2013, 342, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Acevedo-Rocha, C.G.; Budisa, N. Xenomicrobiology: a roadmap for genetic code engineering. Microb. Biotechnol. 2016, 9, 666–676. [Google Scholar] [CrossRef] [PubMed]
- Kunjapur, A.M.; Stork, D.A.; Kuru, E.; Vargas-Rodriguez, O.; Landon, M.; Söll, D.; Church, G.M. Engineering posttranslational proofreading to discriminate nonstandard amino acids. Proc. Natl. Acad. Sci. USA 2018, 115, 619–624. [Google Scholar] [CrossRef] [PubMed]
- Tack, D.S.; Ellefson, J.W.; Thyer, R.; Wang, B.; Gollihar, J.; Forster, M.T.; Ellington, A.D. Addicting diverse bacteria to a noncanonical amino acid. Nat. Chem. Biol. 2016, 12, 138–140. [Google Scholar] [CrossRef] [PubMed]
- Lajoie, M.J.; Söll, D.; Church, G.M. Overcoming Challenges in Engineering the Genetic Code. J. Mol. Biol. 2016, 428, 1004–1021. [Google Scholar] [CrossRef]
- Biddle, W.; Schmitt, M.A.; Fisk, J.D. Modification of orthogonal tRNAs: Unexpected consequences for sense codon reassignment. Nucleic Acids Res. 2016, 44, 10042–10050. [Google Scholar] [CrossRef]
- Ma, N.J.; Isaacs, F.J. Genomic Recoding Broadly Obstructs the Propagation of Horizontally Transferred Genetic Elements. Cell Syst. 2016, 3, 199–207. [Google Scholar] [CrossRef]
- Hammerling, M.J.; Ellefson, J.W.; Boutz, D.R.; Marcotte, E.M.; Ellington, A.D.; Barrick, J.E. Bacteriophages use an expanded genetic code on evolutionary paths to higher fitness. Nat. Chem. Biol. 2014, 10, 178–180. [Google Scholar] [CrossRef]
- Ma, N.J.; Hemez, C.F.; Barber, K.W.; Rinehart, J.; Isaacs, F.J. Organisms with alternative genetic codes resolve unassigned codons via mistranslation and ribosomal rescue. eLife 2018, 7, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Wan, W.; Huang, Y.; Wang, Z.; Russell, W.K.; Pai, P.J.; Russell, D.H.; Liu, W.R. A facile system for genetic incorporation of two different noncanonical amino acids into one protein in Escherichia coli. Angew. Chem. Int. Ed. 2010, 49, 3211–3214. [Google Scholar] [CrossRef] [PubMed]
- Santos, M.A.S.; Cheesman, C.; Costa, V.; Moradas-Ferreira, P.; Tuite, M.F. Selective advantages created by codon ambiguity allowed for the evolution of an alternative genetic code in Candida spp. Mol. Microbiol. 1999, 31, 937–947. [Google Scholar] [CrossRef] [PubMed]
- Kollmar, M.; Mühlhausen, S. How tRNAs dictate nuclear codon reassignments: Only a few can capture non-cognate codons. RNA Biol. 2017, 14, 293–299. [Google Scholar] [CrossRef] [PubMed]
- Schmitt, M.A.; Biddle, W.; Fisk, J.D. Mapping the Plasticity of the Escherichia coli Genetic Code with Orthogonal Pair-Directed Sense Codon Reassignment. Biochemistry 2018, 57, 2762–2774. [Google Scholar] [CrossRef] [PubMed]
- Hoesl, M.G.; Budisa, N. Expanding and Engineering the Genetic Code in a Single Expression Experiment. ChemBioChem 2011, 12, 552–555. [Google Scholar] [CrossRef] [PubMed]
- Wong, J.T. Membership mutation of the genetic code: loss of fitness by tryptophan. Proc. Natl. Acad. Sci. USA 1983, 80, 6303–6306. [Google Scholar] [CrossRef] [PubMed]
- Mat, W.K.; Xue, H.; Wong, J.T.F. Genetic code mutations: The breaking of a three billion year invariance. PLoS ONE 2010, 5, e12206. [Google Scholar] [CrossRef]
- Yu, A.C.S.; Yim, A.K.Y.; Mat, W.K.; Tong, A.H.Y.; Lok, S.; Xue, H.; Tsui, S.K.W.; Wong, J.T.F.; Chan, T.F. Mutations enabling displacement of tryptophan by 4-fluorotryptophan as a canonical amino acid of the genetic code. Genome Biol. Evol. 2014, 6, 629–641. [Google Scholar] [CrossRef]
- Crick, F.H.C. Codon—anticodon pairing: The wobble hypothesis. J. Mol. Biol. 1966, 19, 548–555. [Google Scholar] [CrossRef]
- Bohlke, N.; Budisa, N. Sense codon emancipation for proteome-wide incorporation of noncanonical amino acids: Rare isoleucine codon AUA as a target for genetic code expansion. FEMS Microbiol. Lett. 2014, 351, 133–144. [Google Scholar] [CrossRef] [PubMed]
- Mukai, T.; Yamaguchi, A.; Ohtake, K.; Takahashi, M.; Hayashi, A.; Iraha, F.; Kira, S.; Yanagisawa, T.; Yokoyama, S.; Hoshi, H.; et al. Reassignment of a rare sense codon to a non-canonical amino acid in Escherichia coli. Nucleic Acids Res. 2015, 43, 8111–8122. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.S.; Shin, S.; Jeon, J.Y.; Jang, K.S.; Lee, B.Y.; Choi, S.; Yoo, T.H. Incorporation of Unnatural Amino Acids in Response to the AGG Codon. ACS Chem. Biol. 2015, 10, 1648–1653. [Google Scholar] [CrossRef] [PubMed]
- Fu, X.; Söll, D.; Sevostyanova, A. Challenges of site-specific selenocysteine incorporation into proteins by Escherichia coli. RNA Biol. 2018, 15, 461–470. [Google Scholar] [CrossRef] [PubMed]
- Bröcker, M.J.; Ho, J.M.L.; Church, G.M.; Söll, D.; O’Donoghue, P. Recoding the genetic code with selenocysteine. Angew. Chem. Int. Ed. 2014, 53, 319–323. [Google Scholar] [CrossRef] [PubMed]
- Torres, L.; Krüger, A.; Csibra, E.; Gianni, E.; Pinheiro, V.B. Synthetic biology approaches to biological containment: pre-emptively tackling potential risks. Essays Biochem. 2016, 60, 393–410. [Google Scholar] [CrossRef] [PubMed]
- Grosjean, H.; Westhof, E. An integrated, structure- and energy-based view of the genetic code. Nucleic Acids Res. 2016, 44, 8020–8040. [Google Scholar] [CrossRef]
- Posfai, G. Emergent Properties of Reduced-Genome Escherichia coli. Science 2006, 312, 1044–1046. [Google Scholar] [CrossRef]
- Mukai, T.; Lajoie, M.J.; Englert, M.; Söll, D. Rewriting the Genetic Code. Annu. Rev. Microbiol. 2017, 71, 557–577. [Google Scholar] [CrossRef]
- Chari, R.; Church, G.M. Beyond editing to writing large genomes. Nat. Rev. Genet. 2017, 18, 749–760. [Google Scholar] [CrossRef]
- Kuo, J.; Stirling, F.; Lau, Y.H.; Shulgina, Y.; Way, J.C.; Silver, P.A. Synthetic genome recoding: new genetic codes for new features. Curr. Genet. 2018, 64, 327–333. [Google Scholar] [CrossRef] [PubMed]
- Rackham, O.; Chin, J.W. A network of orthogonal ribosome·mrna pairs. Nat. Chem. Biol. 2005, 1, 159–166. [Google Scholar] [CrossRef] [PubMed]
- Neumann, H.; Wang, K.; Davis, L.; Garcia-Alai, M.; Chin, J.W. Encoding multiple unnatural amino acids via evolution of a quadruplet-decoding ribosome. Nature 2010, 464, 441–444. [Google Scholar] [CrossRef] [PubMed]
- Anderson, J.C.; Wu, N.; Santoro, S.W.; Lakshman, V.; King, D.S.; Schultz, P.G. An expanded genetic code with a functional quadruplet codon. Proc. Natl. Acad. Sci. USA 2004, 101, 7566–7571. [Google Scholar] [CrossRef] [PubMed]
- Hohsaka, T.; Ashizuka, Y.; Murakami, H.; Sisido, M. Five-base codons for incorporation of nonnatural amino acids into proteins. Nucleic Acids Res. 2001, 29, 3646–3651. [Google Scholar] [CrossRef] [PubMed]
- Watson, J.D.; Crick, F.H.C. Molecular Structure of Nucleic Acids: A Structure for Deoxyribose Nucleic Acid. Nature 1953, 171, 737–738. [Google Scholar] [CrossRef] [PubMed]
- Pinheiro, V.B.; Holliger, P. The XNA world: Progress towards replication and evolution of synthetic genetic polymers. Curr. Opin. Chem. Biol. 2012, 16, 245–252. [Google Scholar] [CrossRef]
- Warren, R.A.J. Modified Bases in Bacteriophage DNAs. Annu. Rev. Microbiol. 1980, 34, 137–158. [Google Scholar] [CrossRef]
- Yang, Z.; Sismour, A.M.; Sheng, P.; Puskar, N.L.; Benner, S.A. Enzymatic incorporation of a third nucleobase pair. Nucleic Acids Res. 2007, 35, 4238–4249. [Google Scholar] [CrossRef]
- Sismour, A.M. PCR amplification of DNA containing non-standard base pairs by variants of reverse transcriptase from Human Immunodeficiency Virus-1. Nucleic Acids Res. 2004, 32, 728–735. [Google Scholar] [CrossRef]
- Leal, N.A.; Kim, H.J.; Hoshika, S.; Kim, M.J.; Carrigan, M.A.; Benner, S.A. Transcription, reverse transcription, and analysis of RNA containing artificial genetic components. ACS Synth. Biol. 2015, 4, 407–413. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Zhang, Y.; Daugherty, A.B.; Yang, Z.; Shaw, R.; Dong, M.; Lutz, S.; Benner, S.A. Biological phosphorylation of an Unnatural Base Pair (UBP) using a Drosophila melanogaster deoxynucleoside kinase (DmdNK) mutant. PLoS ONE 2017, 12, e0174163. [Google Scholar] [CrossRef] [PubMed]
- Marlière, P.; Patrouix, J.; Döring, V.; Herdewijn, P.; Tricot, S.; Cruveiller, S.; Bouzon, M.; Mutzel, R. Chemical evolution of a bacterium’s genome. Angew. Chem. Int. Ed. 2011, 50, 7109–7114. [Google Scholar] [CrossRef] [PubMed]
- Malyshev, D.A.; Dhami, K.; Lavergne, T.; Chen, T.; Dai, N.; Foster, J.M.; Corrêa, I.R.; Romesberg, F.E. A semi-synthetic organism with an expanded genetic alphabet. Nature 2014, 509, 385–388. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Romesberg, F.E. Semisynthetic Organisms with Expanded Genetic Codes. Biochemistry 2018, 57, 2177–2178. [Google Scholar] [CrossRef]
- Zhang, Y.; Ptacin, J.L.; Fischer, E.C.; Aerni, H.R.; Caffaro, C.E.; San Jose, K.; Feldman, A.W.; Turner, C.R.; Romesberg, F.E. A semi-synthetic organism that stores and retrieves increased genetic information. Nature 2017, 551, 644–647. [Google Scholar] [CrossRef]
- Hettinger, T.P. Helix instability and self-pairing prevent unnatural base pairs from expanding the genetic alphabet. Proc. Natl. Acad. Sci. USA 2017, 114, E6476–E6477. [Google Scholar] [CrossRef]
- Pinheiro, V.B.; Taylor, A.I.; Cozens, C.; Abramov, M.; Renders, M.; Zhang, S.; Chaput, J.C.; Wengel, J.; Peak-Chew, S.Y.; McLaughlin, S.H.; et al. Synthetic genetic polymers capable of heredity and evolution. Science 2012, 336, 341–344. [Google Scholar] [CrossRef]
- Culbertson, M.C.; Temburnikar, K.W.; Sau, S.P.; Liao, J.; Bala, S.; Chaput, J.C. Bioorganic & Medicinal Chemistry Letters Evaluating TNA stability under simulated physiological conditions. Bioorg. Med. Chem. Lett. 2016, 26, 2418–2421. [Google Scholar] [CrossRef]
- Ichida, J.K.; Horhota, A.; Zou, K.; Mclaughlin, L.W.; Szostak, J.W. High fidelity TNA synthesis by Therminator polymerase. Nucleic Acids Res. 2005, 33, 5219–5225. [Google Scholar] [CrossRef]
- Dunn, M.R.; Chaput, J.C. Reverse Transcription of Threose Nucleic Acid by a Naturally Occurring DNA Polymerase. Chembiochem 2016, 1804–1808. [Google Scholar] [CrossRef] [PubMed]
- Dunn, M.R.; Otto, C.; Fenton, K.E.; Chaput, J.C. Improving Polymerase Activity with Unnatural Substrates by Sampling Mutations in Homologous Protein Architectures. ACS Chem. Biol. 2016. [Google Scholar] [CrossRef] [PubMed]
- Mei, H.; Shi, C.; Jimenez, R.M.; Wang, Y.; Kardouh, M.; Chaput, C. Synthesis and polymerase activity of a fluorescent cytidine TNA triphosphate analogue. Nucleic Acids Res. 2017, 45, 5629–5638. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, M. Xenobiology: A new form of life as the ultimate biosafety tool. BioEssays 2010, 32, 322–331. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Cozens, C.; Jaziri, F.; Rozenski, J.; Maréchal, A.; Dumbre, S.; Pezo, V.; Marlière, P.; Pinheiro, V.B.; Groaz, E.; et al. Phosphonomethyl Oligonucleotides as Backbone-Modified Artificial Genetic Polymers. J. Am. Chem. Soc. 2018, 140, 6690–6699. [Google Scholar] [CrossRef] [PubMed]
- Hartman, H.; Smith, T. The Evolution of the Ribosome and the Genetic Code. Life 2014, 4, 227–249. [Google Scholar] [CrossRef] [PubMed]
- Smith, T.F.; Hartman, H. The evolution of Class II Aminoacyl-tRNA synthetases and the first code. FEBS Lett. 2015, 589, 3499–3507. [Google Scholar] [CrossRef]
- Granold, M.; Hajieva, P.; Toşa, M.I.; Irimie, F.-D.; Moosmann, B. Modern diversification of the amino acid repertoire driven by oxygen. Proc. Natl. Acad. Sci. USA 2017, 115, 201717100. [Google Scholar] [CrossRef]
- Forster, A.C.; Church, G.M. Synthetic biology projects in vitro. Genome Res. 2007, 1–6. [Google Scholar] [CrossRef]
- Adamala, K.P.; Martin-Alarcon, D.A.; Guthrie-Honea, K.R.; Boyden, E.S. Engineering genetic circuit interactions within and between synthetic minimal cells. Nat. Chem. 2017, 9, 431–439. [Google Scholar] [CrossRef]
- Van den Bergh, B.; Swings, T.; Fauvart, M.; Michiels, J. Experimental Design, Population Dynamics, and Diversity in Microbial Experimental Evolution. Microbiol. Mol. Biol. Rev. 2018, 82, e00008-18. [Google Scholar] [CrossRef] [PubMed]
- Tack, D.S.; Cole, A.C.; Shroff, R.; Morrow, B.R.; Ellington, A.D. Evolving Bacterial Fitness with an Expanded Genetic Code. Sci. Rep. 2018, 8, 3288. [Google Scholar] [CrossRef] [PubMed]
- Freeland, S.J.; Hurst, L.D. The genetic code is one in a million. J. Mol. Evol. 1998, 47, 238–248. [Google Scholar] [CrossRef] [PubMed]
- Knight, R.; Freeland, S.; Landweber, L. Rewiring the keyboard: evolvability ogf the genetic code. Nat. Rev. Genet. 2001, 2. [Google Scholar] [CrossRef] [PubMed]
- Palmer, J.D.; Adams, K.L.; Cho, Y.; Parkinson, C.L.; Qiu, Y.L.; Song, K. Dynamic evolution of plant mitochondrial genomes: mobile genes and introns and highly variable mutation rates. Proc. Natl. Acad. Sci. USA 2000, 97, 6960–6966. [Google Scholar] [CrossRef] [PubMed]
- Baranov, P.V.; Atkins, J.F.; Yordanova, M.M. Augmented genetic decoding: Global, local and temporal alterations of decoding processes and codon meaning. Nat. Rev. Genet. 2015, 16, 517–529. [Google Scholar] [CrossRef] [PubMed]
- Kowal, A.K.; Oliver, J.S. Exploiting unassigned codons in Micrococcus luteus for tRNA-based amino acid mutagenesis. Nucleic Acids Res. 1997, 25, 4685–4689. [Google Scholar] [CrossRef]
- Chandel, N.S. Evolution of Mitochondria as Signaling Organelles. Cell Metab. 2015, 22, 204–206. [Google Scholar] [CrossRef]
- Nagao, A.; Ohara, M.; Miyauchi, K.; Yokobori, S.I.; Yamagishi, A.; Watanabe, K.; Suzuki, T. Hydroxylation of a conserved tRNA modification establishes non-universal genetic code in echinoderm mitochondria. Nat. Struct. Mol. Biol. 2017, 24, 778–782. [Google Scholar] [CrossRef]
- Krieger, F.; Mo, A.; Kiefhaber, T. Effect of Proline and Glycine Residues on Dynamics and Barriers of Loop Formation in Polypeptide Chains. J. Am. Chem. Soc. 2005, 3346–3352. [Google Scholar] [CrossRef]
- Kubyshkin, V.; Grage, S.L.; Bürck, J.; Ulrich, A.S.; Budisa, N. Transmembrane Polyproline Helix. J. Phys. Chem. Lett. 2018, 9, 2170–2174. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Diwo, C.; Budisa, N. Alternative Biochemistries for Alien Life: Basic Concepts and Requirements for the Design of a Robust Biocontainment System in Genetic Isolation. Genes 2019, 10, 17. https://doi.org/10.3390/genes10010017
Diwo C, Budisa N. Alternative Biochemistries for Alien Life: Basic Concepts and Requirements for the Design of a Robust Biocontainment System in Genetic Isolation. Genes. 2019; 10(1):17. https://doi.org/10.3390/genes10010017
Chicago/Turabian StyleDiwo, Christian, and Nediljko Budisa. 2019. "Alternative Biochemistries for Alien Life: Basic Concepts and Requirements for the Design of a Robust Biocontainment System in Genetic Isolation" Genes 10, no. 1: 17. https://doi.org/10.3390/genes10010017
APA StyleDiwo, C., & Budisa, N. (2019). Alternative Biochemistries for Alien Life: Basic Concepts and Requirements for the Design of a Robust Biocontainment System in Genetic Isolation. Genes, 10(1), 17. https://doi.org/10.3390/genes10010017