stim2b Knockout Induces Hyperactivity and Susceptibility to Seizures in Zebrafish Larvae
Abstract
1. Introduction
2. Materials and Methods
2.1. Animal Maintenance
2.2. Generation and Genotyping of stim2b−/− Mutant Zebrafish
2.3. Drug Treatments
2.4. Behavioral Experiments
2.4.1. Open Field Test Adopted for Zebrafish Larvae
2.4.2. Light Preference Test
2.4.3. Visual-Motor Response Test
2.5. In Vivo Imaging of Neuronal Activity
2.6. RNA Sequencing and Real-Time Polymerase Chain Reaction Expression Analysis
3. Results
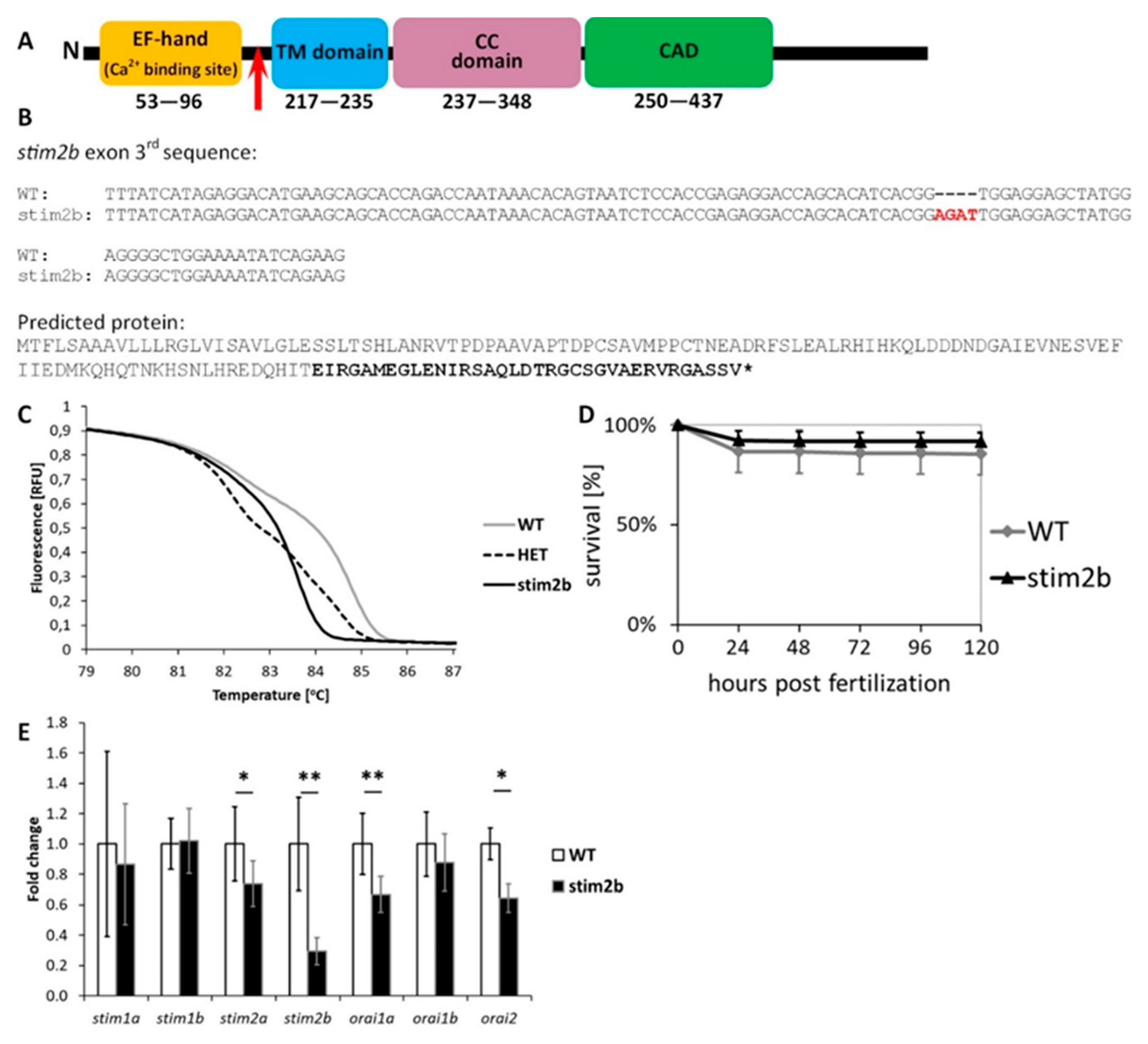
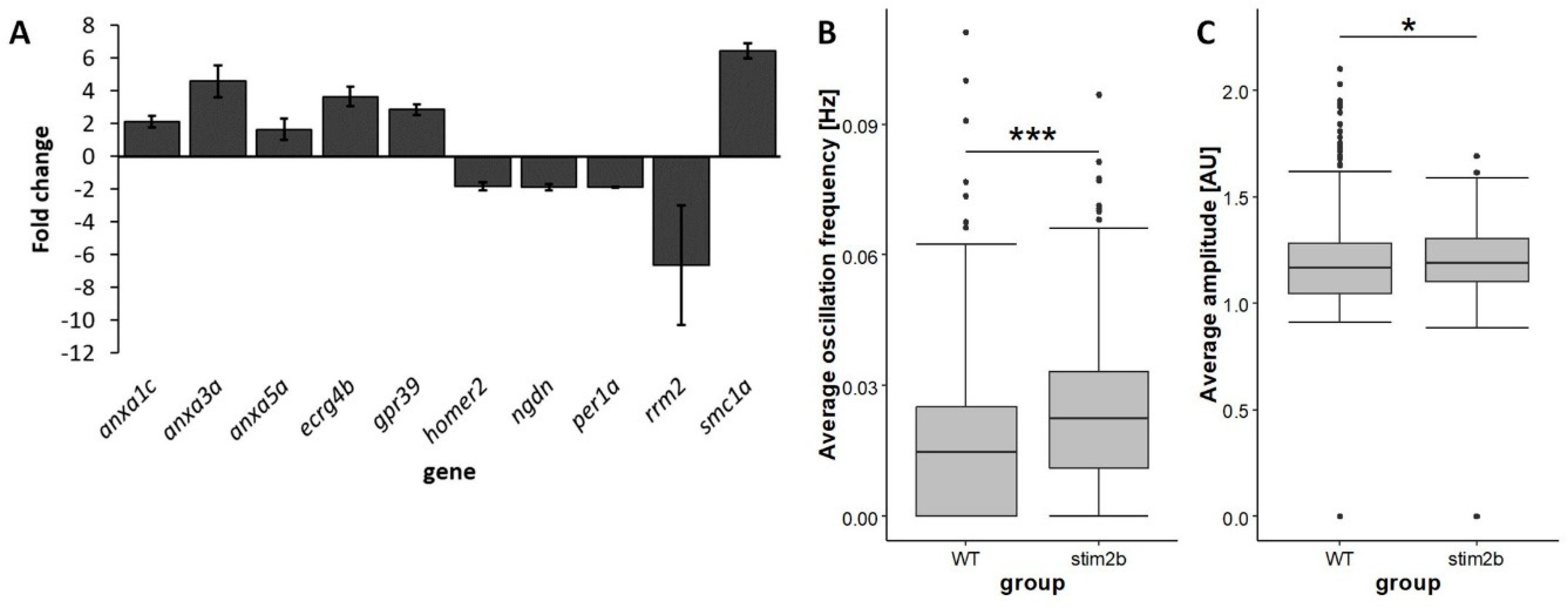
3.1. stim2b Knockout Does Not Induce a Severe Phenotype in Zebrafish Larvae but Affects Gene Expression and Neuronal Activity
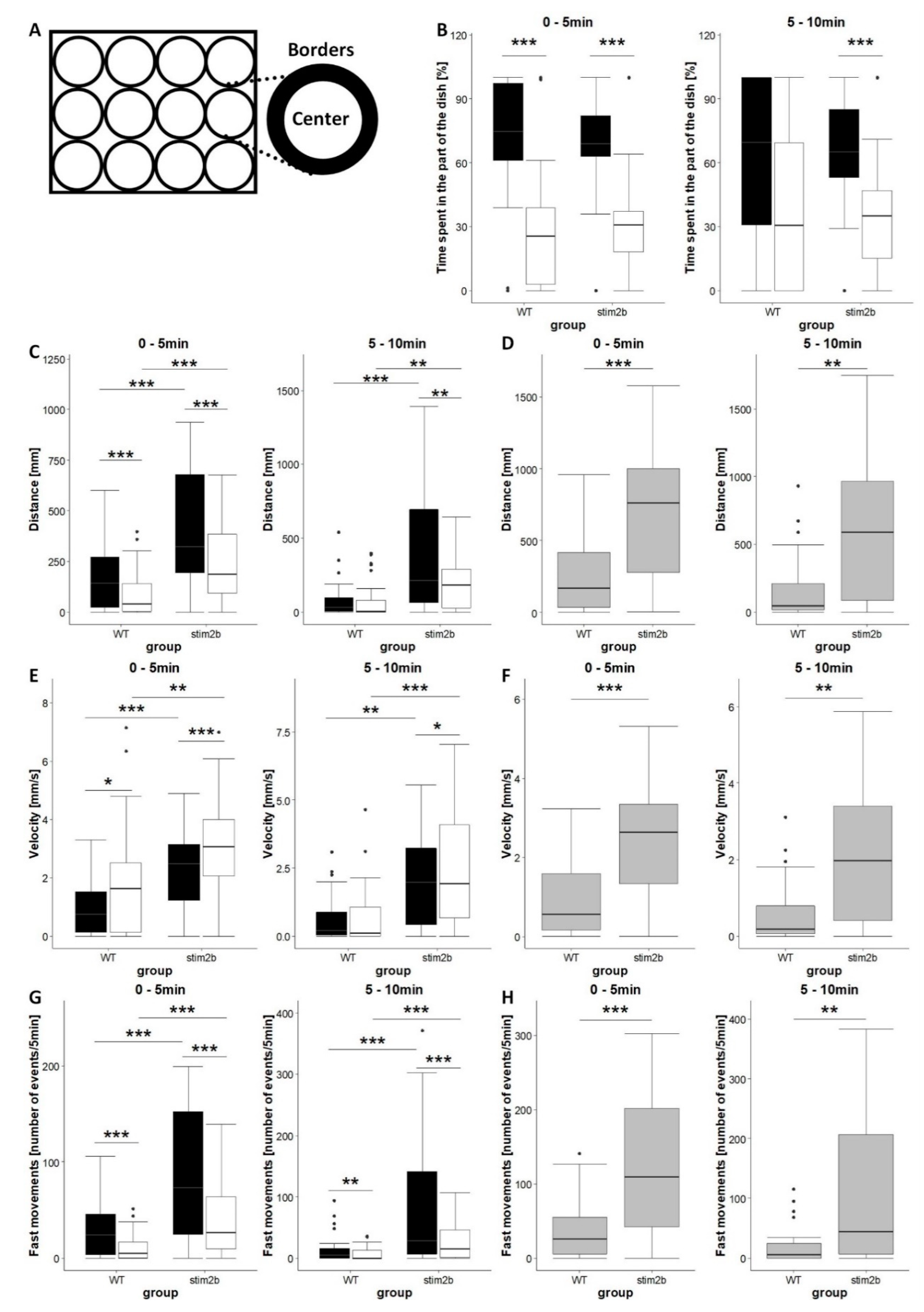
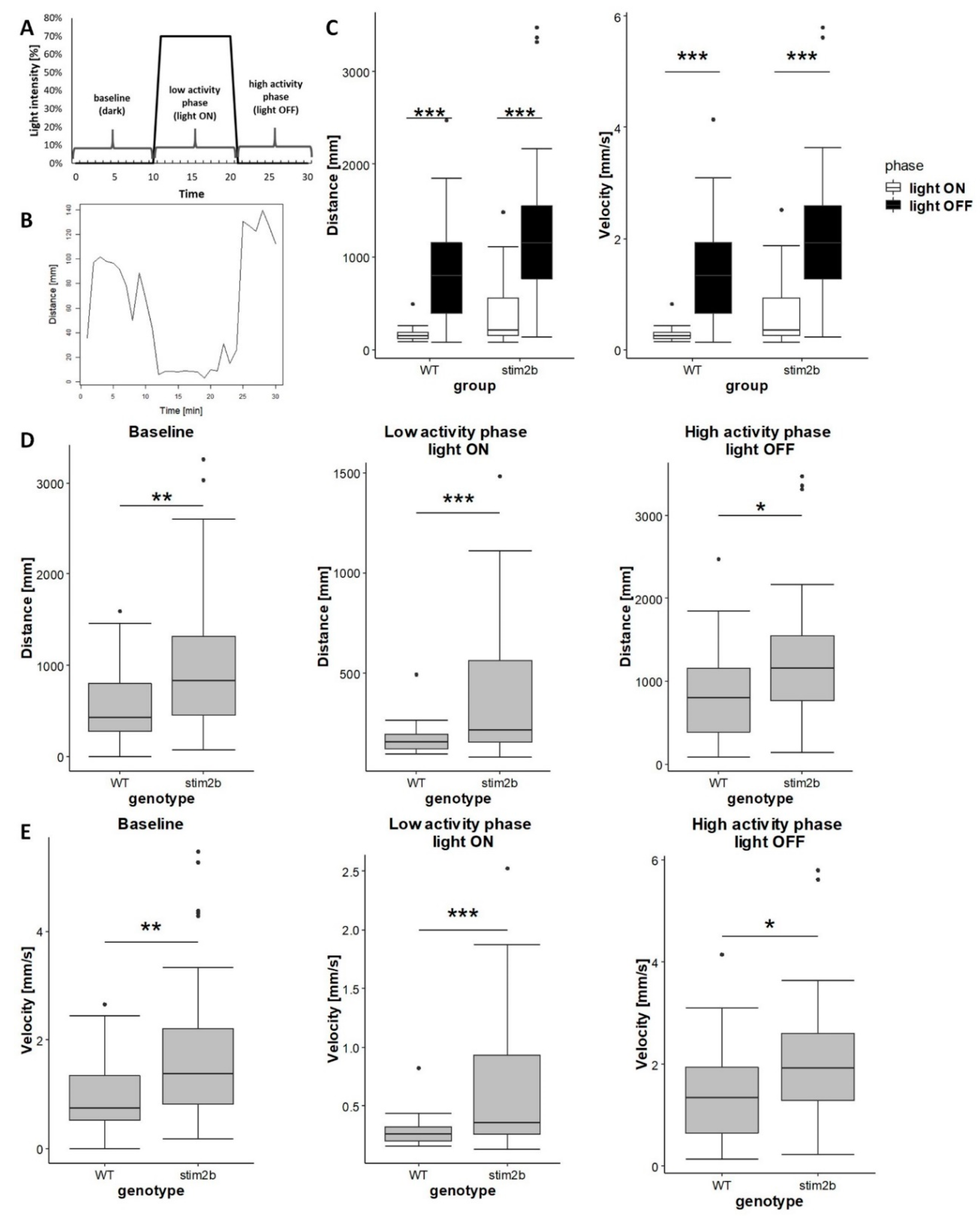
3.2. stim2b Knockout Increases Mobility and Thigmotaxis in Zebrafish Larvae
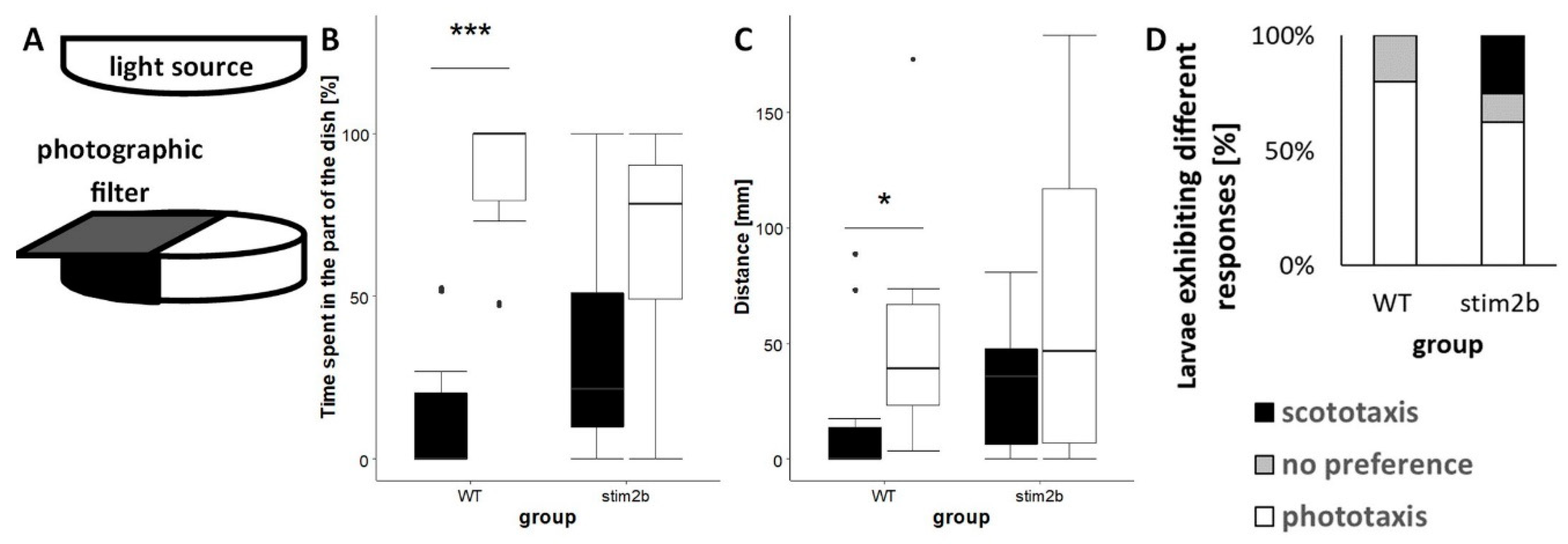
3.3. stim2b−/− Mutants Exhibited Disruptions of Phototaxis but a Normal Visual-Motor Response
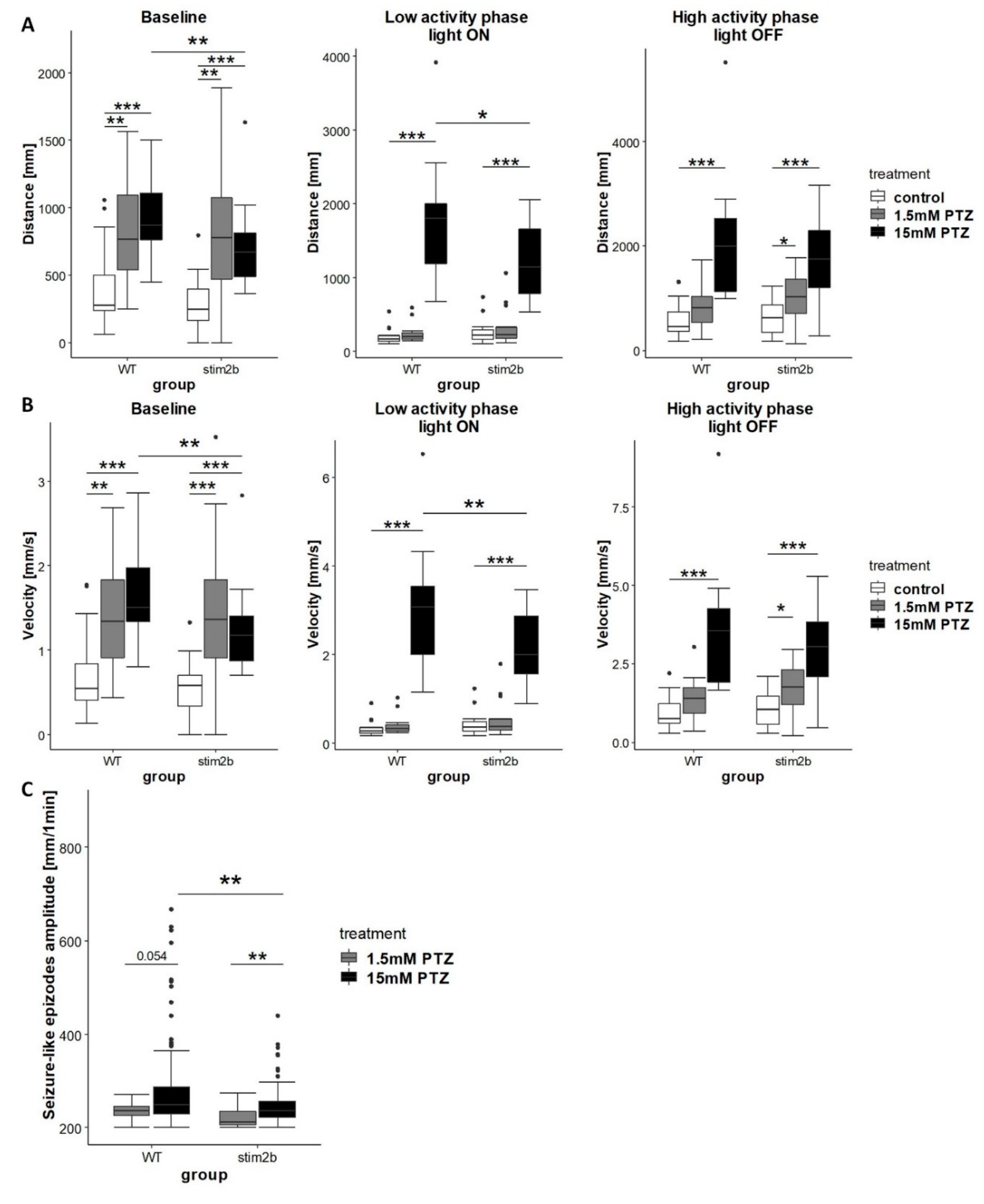
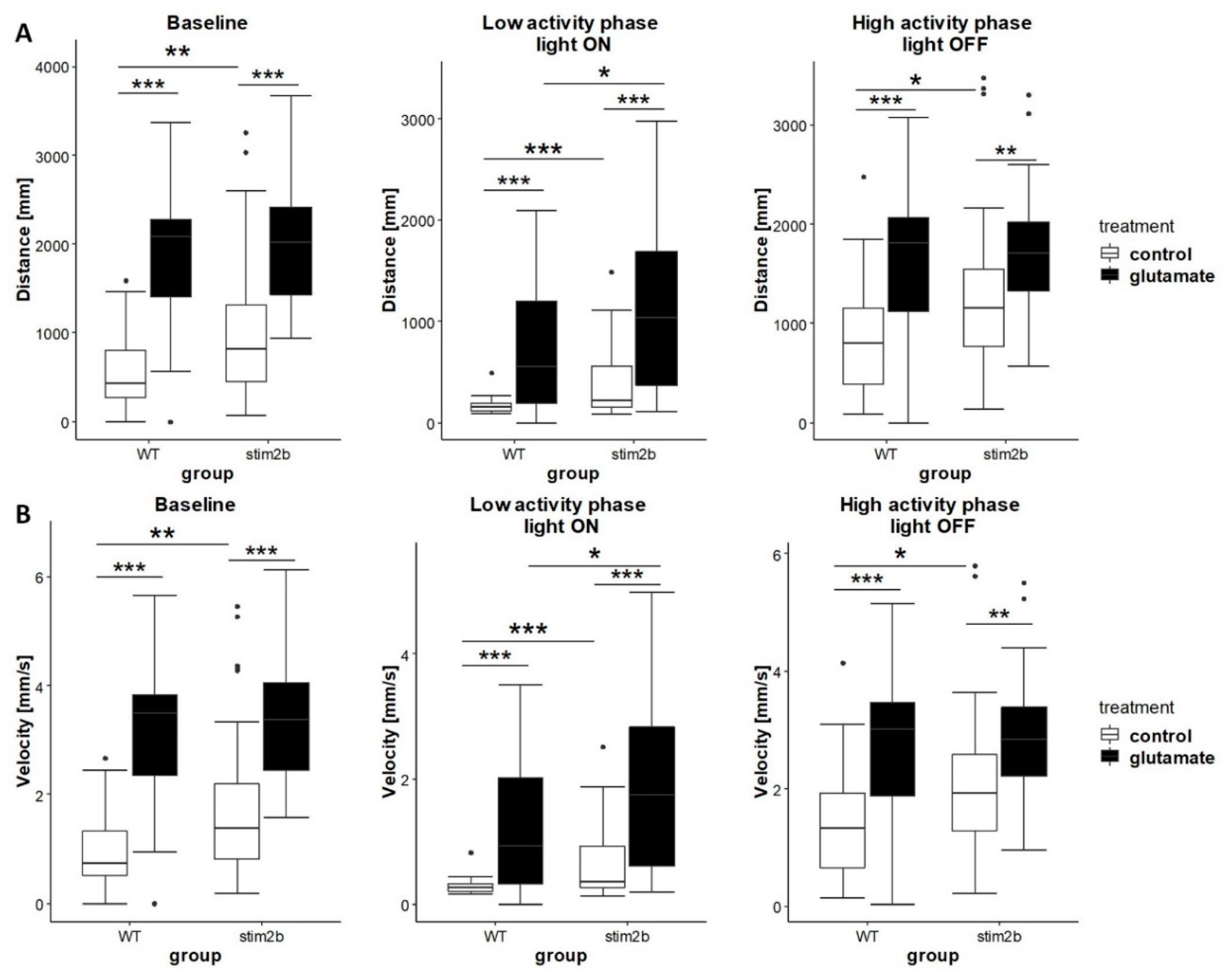
3.4. Response to GABAergic and Glutamatergic Signaling Modulators is Affected by Stim2b Knockout
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Williams, R.T.; Manji, S.S.M.; Parker, N.J.; Hancock, M.S.; Van Stekelenburg, L.; Eid, J.-P.; Senior, P.V.; Kazenwadel, J.; Shandala, T.; Saint, R.; et al. Identification and characterization of the STIM (stromal interaction molecule) gene family: Coding for a novel class of transmembrane proteins. Biochem. J. 2001, 357, 673. [Google Scholar] [CrossRef] [PubMed]
- Erro, A.B.; Jardin, I.; Salido, G.M.; Rosado, J.A. Role of STIM2 in cell function and physiopathology. J. Physiol. 2017, 595, 3111–3128. [Google Scholar] [CrossRef] [PubMed]
- Cai, X. Molecular Evolution and Functional Divergence of the Ca2+ Sensor Protein in Store-operated Ca2+ Entry: Stromal Interaction Molecule. PLoS ONE 2007, 2, e609. [Google Scholar] [CrossRef] [PubMed]
- Wasilewska, I.; Gupta, R.K.; Palchevska, O.; Kuznicki, J. Identification of Zebrafish Calcium Toolkit Genes and their Expression in the Brain. Genes 2019, 10, 230. [Google Scholar] [CrossRef]
- Skibinska-Kijek, A.; Wisniewska, M.B.; Gruszczynska-Biegala, J.; Methner, A.; Kuznicki, J. Immunolocalization of STIM1 in the mouse brain. Acta Neurobiol. Exp. 2009, 69, 413–428. [Google Scholar]
- Steinbeck, J.A.; Henke, N.; Opatz, J.; Gruszczynska-Biegala, J.; Schneider, L.; Theiss, S.; Hamacher, N.; Steinfarz, B.; Golz, S.; Brüstle, O.; et al. Store-operated calcium entry modulates neuronal network activity in a model of chronic epilepsy. Exp. Neurol. 2011, 232, 185–194. [Google Scholar] [CrossRef]
- Tu, C.-C.; Wan, B.-Y.; Zeng, Y. STIM2 knockdown protects against ischemia/reperfusion injury through reducing mitochondrial calcium overload and preserving mitochondrial function. Life Sci. 2020, 247, 116560. [Google Scholar] [CrossRef]
- Brandman, O.; Liou, J.; Park, W.S.; Meyer, T. STIM2 Is a Feedback Regulator that Stabilizes Basal Cytosolic and Endoplasmic Reticulum Ca2+ Levels. Cell 2007, 131, 1327–1339. [Google Scholar] [CrossRef]
- Majewski, L.; Kuznicki, J. SOCE in neurons: Signaling or just refilling? Biochim. Biophys. Acta 2015, 1853, 1940–1952. [Google Scholar] [CrossRef]
- Wegierski, T.; Kuznicki, J. Neuronal calcium signaling via store-operated channels in health and disease. Cell Calcium 2018, 74, 102–111. [Google Scholar] [CrossRef]
- Berridge, M.J. Neuronal Calcium Signaling. Neuron 1998, 21, 13–26. [Google Scholar] [CrossRef]
- Moccia, F.; Zuccolo, E.; Soda, T.; Tanzi, F.; Guerra, G.; Mapelli, L.; Lodola, F.; D’Angelo, E. Stim and Orai proteins in neuronal Ca2+ signaling and excitability. Front. Cell. Neurosci. 2015, 9, 153. [Google Scholar] [CrossRef] [PubMed]
- Gruszczynska-Biegala, J.; Pomorski, P.; Wisniewska, M.B.; Kuznicki, J. Differential Roles for STIM1 and STIM2 in Store-Operated Calcium Entry in Rat Neurons. PLoS ONE 2011, 6, e19285. [Google Scholar] [CrossRef] [PubMed]
- Gruszczynska-Biegala, J.; Kuznicki, J. Native STIM2 and ORAI1 proteins form a calcium-sensitive and thapsigargin-insensitive complex in cortical neurons. J. Neurochem. 2013, 126, 727–738. [Google Scholar] [CrossRef] [PubMed]
- Baba, A.; Yasui, T.; Fujisawa, S.; Yamada, R.X.; Yamada, M.; Nishiyama, N.; Matsuki, N.; Ikegaya, Y. Activity-Evoked Capacitative Ca2+ Entry: Implications in Synaptic Plasticity. J. Neurosci. 2003, 23, 7737–7741. [Google Scholar] [CrossRef] [PubMed]
- LaLonde, J.; Saia, G.; Gill, G. Store-Operated Calcium Entry Promotes the Degradation of the Transcription Factor Sp4 in Resting Neurons. Sci. Signal. 2014, 7, ra51. [Google Scholar] [CrossRef]
- Sun, S.; Zhang, H.; Liu, J.; Popugaeva, E.; Xu, N.-J.; Feske, S.; White, C.L.; Bezprozvanny, I.B. Reduced synaptic STIM2 expression and impaired store-operated calcium entry cause destabilization of mature spines in mutant presenilin mice. Neuron 2014, 82, 79–93. [Google Scholar] [CrossRef]
- Park, C.Y.; Shcheglovitov, A.; Dolmetsch, R. The CRAC Channel Activator STIM1 Binds and Inhibits L-Type Voltage-Gated Calcium Channels. Science 2010, 330, 101–105. [Google Scholar] [CrossRef]
- Garcia-Alvarez, G.; Lu, B.; Yap, K.A.F.; Wong, L.C.; Thevathasan, J.V.; Lim, L.; Ji, F.; Tan, K.W.; Mancuso, J.J.; Tang, W.; et al. STIM2 regulates PKA-dependent phosphorylation and trafficking of AMPARs. Mol. Biol. Cell 2015, 26, 1141–1159. [Google Scholar] [CrossRef]
- Gemes, G.; Bangaru, M.L.Y.; Wu, H.-E.; Tang, Q.; Weihrauch, R.; Koopmeiners, A.S.; Cruikshank, J.M.; Kwok, W.-M.; Hogan, Q.H. Store-operated Ca2+ entry in sensory neurons: Functional role and the effect of painful nerve injury. J. Neurosci. 2011, 31, 3536–3549. [Google Scholar] [CrossRef]
- Bojarski, L.; Pomorski, P.; Szybinska, A.; Drab, M.; Skibinska-Kijek, A.; Gruszczynska-Biegala, J.; Kuznicki, J. Presenilin-dependent expression of STIM proteins and dysregulation of capacitative Ca2+ entry in familial Alzheimer’s disease. Biochim. Biophys. Acta. 2009, 1793, 1050–1057. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Shih, H.-P.; Vigont, V.; Hrdlicka, L.; Diggins, L.; Singh, C.; Mahoney, M.; Chesworth, R.; Shapiro, G.; Zimina, O.; et al. Neuronal Store-Operated Calcium Entry Pathway as a Novel Therapeutic Target for Huntington’s Disease Treatment. Chem. Biol. 2011, 18, 777–793. [Google Scholar] [CrossRef] [PubMed]
- Czeredys, M.; Vigont, V.A.; Boeva, V.A.; Mikoshiba, K.; Kaznacheyeva, E.V.; Kuznicki, J. Huntingtin-Associated Protein 1A Regulates Store-Operated Calcium Entry in Medium Spiny Neurons From Transgenic YAC128 Mice, a Model of Huntington’s Disease. Front. Cell. Neurosci. 2018, 12. [Google Scholar] [CrossRef]
- Secondo, A.; Bagetta, G.; Amantea, D. On the Role of Store-Operated Calcium Entry in Acute and Chronic Neurodegenerative Diseases. Front. Mol. Neurosci. 2018, 11, 87. [Google Scholar] [CrossRef] [PubMed]
- Erro, A.B.; Braun, A.; Kraft, R.; Kleinschnitz, C.; Schuhmann, M.K.; Stegner, D.; Wultsch, T.; Eilers, J.; Meuth, S.G.; Stoll, G.; et al. STIM2 Regulates Capacitive Ca2+ Entry in Neurons and Plays a Key Role in Hypoxic Neuronal Cell Death. Sci. Signal. 2009, 2, ra67. [Google Scholar] [CrossRef]
- Oh-Hora, M.; Yamashita, M.; Hogan, P.G.; Sharma, S.; Lamperti, E.; Chung, W.; Prakriya, M.; Feske, S.; Rao, A. Dual functions for the endoplasmic reticulum calcium sensors STIM1 and STIM2 in T cell activation and tolerance. Nat. Immunol. 2008, 9, 432–443. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Alvarez, G.; Shetty, M.S.; Lu, B.; Yap, K.A.F.; Oh-Hora, M.; Sajikumar, S.; Bichler, Z.; Fivaz, M. Impaired spatial memory and enhanced long-term potentiation in mice with forebrain-specific ablation of the Stim genes. Front. Behav. Neurosci. 2015, 9, 180. [Google Scholar] [CrossRef] [PubMed]
- Kalueff, A.V.; Gebhardt, M.; Stewart, A.M.; Cachat, J.M.; Brimmer, M.; Chawla, J.S.; Craddock, C.; Kyzar, E.J.; Roth, A.; Landsman, S.; et al. Towards a Comprehensive Catalog of Zebrafish Behavior 1.0 and Beyond. Zebrafish 2013, 10, 70–86. [Google Scholar] [CrossRef] [PubMed]
- Orger, M.B.; De Polavieja, G.G. Zebrafish Behavior: Opportunities and Challenges. Annu. Rev. Neurosci. 2017, 40, 125–147. [Google Scholar] [CrossRef] [PubMed]
- Schnörr, S.; Steenbergen, P.; Richardson, M.; Champagne, D. Measuring thigmotaxis in larval zebrafish. Behav. Brain Res. 2012, 228, 367–374. [Google Scholar] [CrossRef]
- Lundegaard, P.R.; Anastasaki, C.; Grant, N.J.; Sillito, R.R.; Zich, J.; Zeng, Z.; Paranthaman, K.; Larsen, A.P.; Armstrong, D.; Porteous, D.J.; et al. MEK Inhibitors Reverse cAMP-Mediated Anxiety in Zebrafish. Chem. Biol. 2015, 22, 1335–1346. [Google Scholar] [CrossRef] [PubMed]
- Bai, Y.; Liu, H.; Huang, B.; Wagle, M.; Guo, S. Identification of environmental stressors and validation of light preference as a measure of anxiety in larval zebrafish. BMC Neurosci. 2016, 17, 63. [Google Scholar] [CrossRef] [PubMed]
- Kedra, M.; Banasiak, K.; Kisielewska, K.; Wolinska-Niziol, L.; Jaworski, J.; Zmorzynska, J. TrkB hyperactivity contributes to brain dysconnectivity, epileptogenesis, and anxiety in zebrafish model of Tuberous Sclerosis Complex. Proc. Natl. Acad. Sci. USA 2020, 117, 2170–2179. [Google Scholar] [CrossRef]
- Wiley, D.; Redfield, S.; Zon, L. Chemical screening in zebrafish for novel biological and therapeutic discovery. Methods Cell Biol. 2016, 138, 651–679. [Google Scholar] [CrossRef]
- Baraban, S.C.; Dinday, M.T.; Castro, P.A.; Chege, S.; Guyenet, S.; Taylor, M.R. A large-scale mutagenesis screen to identify seizure-resistant zebrafish. Epilepsia 2007, 48, 1151–1157. [Google Scholar] [CrossRef] [PubMed]
- Ensembl Genome Browser. Available online: www.ensembl.org (accessed on 1 April 2020).
- Chan, C.M.; Chen, Y.; Hung, T.S.; Miller, A.; Shipley, A.M.; Webb, S. Inhibition of SOCE disrupts cytokinesis in zebrafish embryos via inhibition of cleavage furrow deepening. Int. J. Dev. Biol. 2015, 59, 289–301. [Google Scholar] [CrossRef] [PubMed]
- Pavez, M.; Thompson, A.C.; Arnott, H.J.; Mitchell, C.B.; D’Atri, I.; Don, E.K.; Chilton, J.K.; Scott, E.K.; Lin, J.Y.; Young, K.M.; et al. STIM1 Is Required for Remodeling of the Endoplasmic Reticulum and Microtubule Cytoskeleton in Steering Growth Cones. J. Neurosci. 2019, 39, 5095–5114. [Google Scholar] [CrossRef]
- Motiani, R.K.; Tanwar, J.; Raja, D.A.; Vashisht, A.; Khanna, S.; Sharma, S.; Srivastava, S.; Sivasubbu, S.; Natarajan, V.T.; Gokhale, A.R.S. STIM 1 activation of adenylyl cyclase 6 connects Ca2+ and cAMP signaling during melanogenesis. EMBO J. 2018, 37, e97597. [Google Scholar] [CrossRef]
- Ahrens, M.B.; Orger, M.B.; Robson, D.; Li, J.; Keller, P.J. Whole-brain functional imaging at cellular resolution using light-sheet microscopy. Nat. Methods 2013, 10, 413–420. [Google Scholar] [CrossRef]
- Matthews, M.; Trevarrow, B.; Matthews, J. A virtual tour of the Guide for zebrafish users. Lab Anim. 2002, 31, 34–40. [Google Scholar]
- Liu, Y.; Carmer, R.; Zhang, G.; Venkatraman, P.; Brown, S.A.; Pang, C.-P.; Zhang, M.; Ma, P.; Leung, Y.F. Statistical Analysis of Zebrafish Locomotor Response. PLoS ONE 2015, 10, e0139521. [Google Scholar] [CrossRef] [PubMed]
- Knafo, S.; Fidelin, K.; Prendergast, A.; Tseng, P.-E.B.; Parrin, A.; Dickey, C.W.; Böhm, U.L.; Figueiredo, S.N.; Thouvenin, O.; Pascal-Moussellard, H.; et al. Mechanosensory neurons control the timing of spinal microcircuit selection during locomotion. eLife 2017, 6, 6. [Google Scholar] [CrossRef]
- Panier, T.; Romano, S.; Olive, R.; Pietri, T.; Sumbre, G.; Candelier, R.; Debrégeas, G. Fast functional imaging of multiple brain regions in intact zebrafish larvae using selective plane illumination microscopy. Front. Neural Circuit. 2013, 7, 65. [Google Scholar] [CrossRef] [PubMed]
- Peterson, S.M.; Freeman, J.L. RNA Isolation from Embryonic Zebrafish and cDNA Synthesis for Gene Expression Analysis. J. Vis. Exp. 2009. [Google Scholar] [CrossRef] [PubMed]
- Burgess, H.A.; Granato, M. Modulation of locomotor activity in larval zebrafish during light adaptation. J. Exp. Biol. 2007, 210, 2526–2539. [Google Scholar] [CrossRef]
- Meldrum, B.S. The role of glutamate in epilepsy and other CNS disorders. Neurology 1994, 44, 14–23. [Google Scholar]
- Gruszczynska-Biegala, J.; Sladowska, M.; Kuznicki, J. AMPA Receptors Are Involved in Store-Operated Calcium Entry and Interact with STIM Proteins in Rat Primary Cortical Neurons. Front. Cell. Neurosci. 2016, 10, 1289. [Google Scholar] [CrossRef]
- Gruszczynska-Biegala, J.; Strucinska, K.; Maciąg, F.; Majewski, L.; Sladowska, M.; Kuznicki, J. STIM Protein-NMDA2 Receptor Interaction Decreases NMDA-Dependent Calcium Levels in Cortical Neurons. Cells 2020, 9, 160. [Google Scholar] [CrossRef]
- Zhou, S.; Li, C.-S.; Liu, L.-Q.; Li, Y.; Wang, X.-F.; Shen, L. Increased expression of annexin A7 in temporal lobe tissue of patients with refractory epilepsy. Histol. Histopathol. 2011, 26, 571–579. [Google Scholar]
- Zub, E.; Canet, G.; Garbelli, R.; Blaquiere, M.; Rossini, L.; Pastori, C.; Sheikh, M.; Reutelingsperger, C.; Klement, W.; De Bock, F.; et al. The GR-ANXA1 pathway is a pathological player and a candidate target in epilepsy. FASEB J. 2019, 33, 13998–14009. [Google Scholar] [CrossRef]
- Chong, K.W.Y.; Chen, M.J.; Koay, E.S.; Wong, B.-S.; Lee, A.Y.-W.; Russo-Marie, F.; Cheung, N.S. Annexin A3 is associated with cell death in lactacystin-mediated neuronal injury. Neurosci. Lett. 2010, 485, 129–133. [Google Scholar] [CrossRef] [PubMed]
- Simats, A.; García-Berrocoso, T.; Montaner, J. Neuroinflammatory biomarkers: From stroke diagnosis and prognosis to therapy. Biochim. Biophys. Acta 2016, 1862, 411–424. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, A.M.; Podvin, S.; Lin, S.; Miller, M.C.; Botfield, H.; E Leadbeater, W.; Roberton, A.; Dang, X.; E Knowling, S.; Cardenas-Galindo, E.; et al. Ecrg4 expression and its product augurin in the choroid plexus: Impact on fetal brain development, cerebrospinal fluid homeostasis and neuroprogenitor cell response to CNS injury. Fluids Barriers CNS 2011, 8, 6. [Google Scholar] [CrossRef] [PubMed]
- Kujuro, Y.; Suzuki, N.; Kondo, T. Esophageal cancer-related gene 4 is a secreted inducer of cell senescence expressed by aged CNS precursor cells. Proc. Natl. Acad. Sci. USA 2010, 107, 8259–8264. [Google Scholar] [CrossRef] [PubMed]
- Hwang, D.Y.; Woo, J.-M.; Park, S.J.; Kang, H.I.; Kim, B.G.; Shim, S.B.; Jee, S.W.; Lee, S.H.; Sin, J.S.; Bae, C.J.; et al. Characterization of changes in global gene expression in the brain of neuron-specific enolase/human Tau23 transgenic mice in response to overexpression of Tau protein. Int. J. Mol. Med. 2010, 25, 667–675. [Google Scholar] [CrossRef][Green Version]
- Hershfinkel, M. The Zinc Sensing Receptor, ZnR/GPR39, in Health and Disease. Int. J. Mol. Sci. 2018, 19, 439. [Google Scholar] [CrossRef]
- Alavi, M.S.; Shamsizadeh, A.; Azhdari-Zarmehri, H.; Roohbakhsh, A. Orphan G protein-coupled receptors: The role in CNS disorders. Biomed. Pharmacother. 2018, 98, 222–232. [Google Scholar] [CrossRef]
- Fazio, G.; Gaston-Massuet, C.; Bettini, L.R.; Graziola, F.; Scagliotti, V.; Cereda, A.; Ferrari, L.; Mazzola, M.; Cazzaniga, G.; Giordano, A.; et al. CyclinD1 Down-Regulation and Increased Apoptosis Are Common Features of Cohesinopathies. J. Cell. Physiol. 2015, 231, 613–622. [Google Scholar] [CrossRef]
- Goldstein, J.H.; Tim-Aroon, T.; Shieh, J.; Merrill, M.; Deeb, K.K.; Zhang, S.; Bass, N.E.; Bedoyan, J.K. Novel SMC1A frameshift mutations in children with developmental delay and epilepsy. Eur. J. Med Genet. 2015, 58, 562–568. [Google Scholar] [CrossRef]
- Lebrun, N.; Lebon, S.; Jeannet, P.-Y.; Jacquemont, S.; Billuart, P.; Bienvenu, T. Early-onset encephalopathy with epilepsy associated with a novel splice site mutation inSMC1A. Am. J. Med Genet. Part A 2015, 167, 3076–3081. [Google Scholar] [CrossRef]
- Barresi, M.J.; Burton, S.; Dipietrantonio, K.; Amsterdam, A.; Hopkins, N.; Karlstrom, R.O. Essential genes for astroglial development and axon pathfinding during zebrafish embryogenesis. Dev. Dyn. 2010, 239, 2603–2618. [Google Scholar] [CrossRef] [PubMed]
- Udagawa, T.; Swanger, S.A.; Takeuchi, K.; Kim, J.H.; Nalavadi, V.; Shin, J.; Lorenz, L.J.; Zukin, R.S.; Bassell, G.J.; Richter, J.D. Bidirectional control of mRNA translation and synaptic plasticity by the cytoplasmic polyadenylation complex. Mol. Cell 2012, 47, 253–266. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Zhong, Z.; Wang, M.; Chen, X.; Tan, Y.; Zhang, S.; He, W.; He, X.; Huang, G.; Lu, H.; et al. Circadian Modulation of Dopamine Levels and Dopaminergic Neuron Development Contributes to Attention Deficiency and Hyperactive Behavior. J. Neurosci. 2015, 35, 2572–2587. [Google Scholar] [CrossRef]
- Shiraishi-Yamaguchi, Y.; Sato, Y.; Sakai, R.; Mizutani, A.; Knöpfel, T.; Mori, N.; Mikoshiba, K.; Furuichi, T. Interaction of Cupidin/Homer2 with two actin cytoskeletal regulators, Cdc42 small GTPase and Drebrin, in dendritic spines. BMC Neurosci. 2009, 10, 25. [Google Scholar] [CrossRef] [PubMed]
- Shin, D.M.; Dehoff, M.; Luo, X.; Kang, S.H.; Tu, J.; Nayak, S.K.; Ross, E.M.; Worley, P.; Muallem, S. Homer 2 tunes G protein–coupled receptors stimulus intensity by regulating RGS proteins and PLCβ GAP activities. J. Cell Biol. 2003, 162, 293–303. [Google Scholar] [CrossRef] [PubMed]
- Jardin, I.; Lopez, J.J.; Erro, A.B.; Salido, G.M.; Rosado, J.A. Homer Proteins in Ca2+ Entry. IUBMB Life 2013, 65, 497–504. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.-M.; Lee, J.; Jo, H.; Park, S.; Chang, I.; Muallem, S.; Shin, D.M. Homer2 Protein Regulates Plasma Membrane Ca2+-ATPase-mediated Ca2+Signaling in Mouse Parotid Gland Acinar Cells. J. Biol. Chem. 2014, 289, 24971–24979. [Google Scholar] [CrossRef] [PubMed]
- Majewski, L.; Maciąg, F.; Boguszewski, P.M.; Kuznicki, J. Transgenic Mice Overexpressing Human STIM2 and ORAI1 in Neurons Exhibit Changes in Behavior and Calcium Homeostasis but Show No Signs of Neurodegeneration. Int. J. Mol. Sci. 2020, 21, 842. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Chang, S.; Lu, Q.; Zhang, Y.; Wu, Z.; Sun, X.; Cao, Q.; Qian, Y.; Jia, T.; Xu, B.; et al. A new locus regulating MICALL2 expression was identified for association with executive inhibition in children with attention deficit hyperactivity disorder. Mol. Psychiatry 2017, 23, 1014–1020. [Google Scholar] [CrossRef]
- Lange, M.; Norton, W.H.; Coolen, M.; Chaminade, M.; Merker, S.; Proft, F.; Schmitt, A.; Vernier, P.; Lesch, K.-P.; Bally-Cuif, L. The ADHD-susceptibility gene lphn3.1 modulates dopaminergic neuron formation and locomotor activity during zebrafish development. Mol. Psychiatry 2012, 17, 946–954. [Google Scholar] [CrossRef]
- Taylor, A.; Steinberg, J.; Webber, C. Duplications in ADHD patients harbour neurobehavioural genes that are co-expressed with genes associated with hyperactivity in the mouse. Am. J. Med Genet. 2015, 168, 97–107. [Google Scholar] [CrossRef] [PubMed]
- De Calbiac, H.; Dabacan, A.; Marsan, E.; Tostivint, H.; Devienne, G.; Ishida, S.; LeGuern, E.; Baulac, S.; Muresan, R.C.; Kabashi, E.; et al. Depdc5 knockdown causes mTOR-dependent motor hyperactivity in zebrafish. Ann. Clin. Transl. Neurol. 2018, 5, 510–523. [Google Scholar] [CrossRef] [PubMed]
- Baraban, S.C.; Taylor, M.; Castro, P.; Baier, H. Pentylenetetrazole induced changes in zebrafish behavior, neural activity and c-fos expression. Neuroscience 2005, 131, 759–768. [Google Scholar] [CrossRef] [PubMed]
- McCutcheon, V.; Park, E.; Liu, E.; Wang, Y.; Wen, X.-Y.; Baker, A.J. A Model of Excitotoxic Brain Injury in Larval Zebrafish: Potential Application for High-Throughput Drug Evaluation to Treat Traumatic Brain Injury. Zebrafish 2016, 13, 161–169. [Google Scholar] [CrossRef] [PubMed]
| Type of Test | Genotype | Treatment | Number of Larvae | Number of Experiments |
|---|---|---|---|---|
| Open field test adopted for zebrafish larvae | WT | E3 | 32 | 3 |
| stim2b−/− | 33 | |||
| Light preference test | WT | E3 | 10 | 5 |
| stim2b−/− | 8 | |||
| Visual-motor response test–no treatment | WT | E3 | 36 | 3 |
| stim2b−/− | 35 | |||
| Visual-motor response test–glutamate treatment | WT | 600 µM glutamate | 36 | 3 |
| stim2b−/− | 35 | |||
| Visual-motor response test–PTZ treatment | WT | 1.5 mM PTZ | 14 | 3 |
| stim2b−/− | 18 | |||
| WT | 15 mM PTZ | 18 | ||
| stim2b−/− | 18 |
| Gene | Forward Primer | Reverse Primer |
|---|---|---|
| eef1a1l1 | AAAATCGGTGGTGCTGGCAA | GGAACGGTGTGATTGAGGGA |
| stim1a | TGAATTCGGATTGCCAGTCGT | TTCAAGTCCCTCTGCGAACC |
| stim1b | TGAGTTTTGAGGCCATCCGC | AACCCATCCGTCTCTGTCAC |
| stim2a | ATTACGGAGGCGGATCGATT | CCTCAATGCCTCCATCCTGA |
| stim2b | CTGGTGGAGTGGACGATCTT | CGTCAGAGGAGGTCGAATCA |
| orai1a | GTGCATTTTTACCGCTCGCT | TTGAAGAGGCATCTCCCCTC |
| orai1b | GCTGTAAGCAACGTGCACAA | TCCCGATGACGGTGGAAAAG |
| orai2 | CGAGCTAGCCTGGGGTTTTT | AGTCAACCGGCAGGAACTTG |
| Gene | Mean Fold Change | p | False Discovery Rate |
|---|---|---|---|
| anxa1c | 2.13 | 0.0027 | 0.0874 |
| anxa3a | 4.59 * | 0.0002 | 0.0332 |
| anxa5a | 1.64 | 0.0083 | 0.1387 |
| ecrg4b | 3.65 * | 0.0001 | 0.0271 |
| gpr39 | 2.87 * | 0.0003 | 0.0380 |
| homer2 | 0.54 * | 0.0004 | 0.0426 |
| ngdn | 0.54 * | 0.0004 | 0.0426 |
| per1a | 0.54 ** | <0.0001 | 0.0066 |
| rrm2 | 0.15 * | 0.0007 | 0.0500 |
| smc1a | 6.45 * | <0.0001 | 0.0142 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wasilewska, I.; Gupta, R.K.; Wojtaś, B.; Palchevska, O.; Kuźnicki, J. stim2b Knockout Induces Hyperactivity and Susceptibility to Seizures in Zebrafish Larvae. Cells 2020, 9, 1285. https://doi.org/10.3390/cells9051285
Wasilewska I, Gupta RK, Wojtaś B, Palchevska O, Kuźnicki J. stim2b Knockout Induces Hyperactivity and Susceptibility to Seizures in Zebrafish Larvae. Cells. 2020; 9(5):1285. https://doi.org/10.3390/cells9051285
Chicago/Turabian StyleWasilewska, Iga, Rishikesh Kumar Gupta, Bartosz Wojtaś, Oksana Palchevska, and Jacek Kuźnicki. 2020. "stim2b Knockout Induces Hyperactivity and Susceptibility to Seizures in Zebrafish Larvae" Cells 9, no. 5: 1285. https://doi.org/10.3390/cells9051285
APA StyleWasilewska, I., Gupta, R. K., Wojtaś, B., Palchevska, O., & Kuźnicki, J. (2020). stim2b Knockout Induces Hyperactivity and Susceptibility to Seizures in Zebrafish Larvae. Cells, 9(5), 1285. https://doi.org/10.3390/cells9051285





