Ameloblastomas Exhibit Stem Cell Potential, Possess Neurotrophic Properties, and Establish Connections with Trigeminal Neurons
Abstract
1. Introduction
2. Materials and Methods
2.1. Collection of Human Material
2.2. Immunostaining on Tissue Sections
2.3. Gene Expression Analysis—Real Time PCR
2.4. Preparation of Microfluidic Devices
2.5. Mouse Handling and Trigeminal Ganglia Dissection
2.6. Co-Culture of Trigeminal Ganglia and AB10 Cells in Microfluidic Devices
2.7. Immunofluorescent Staining in Microfluidic Devices
3. Results
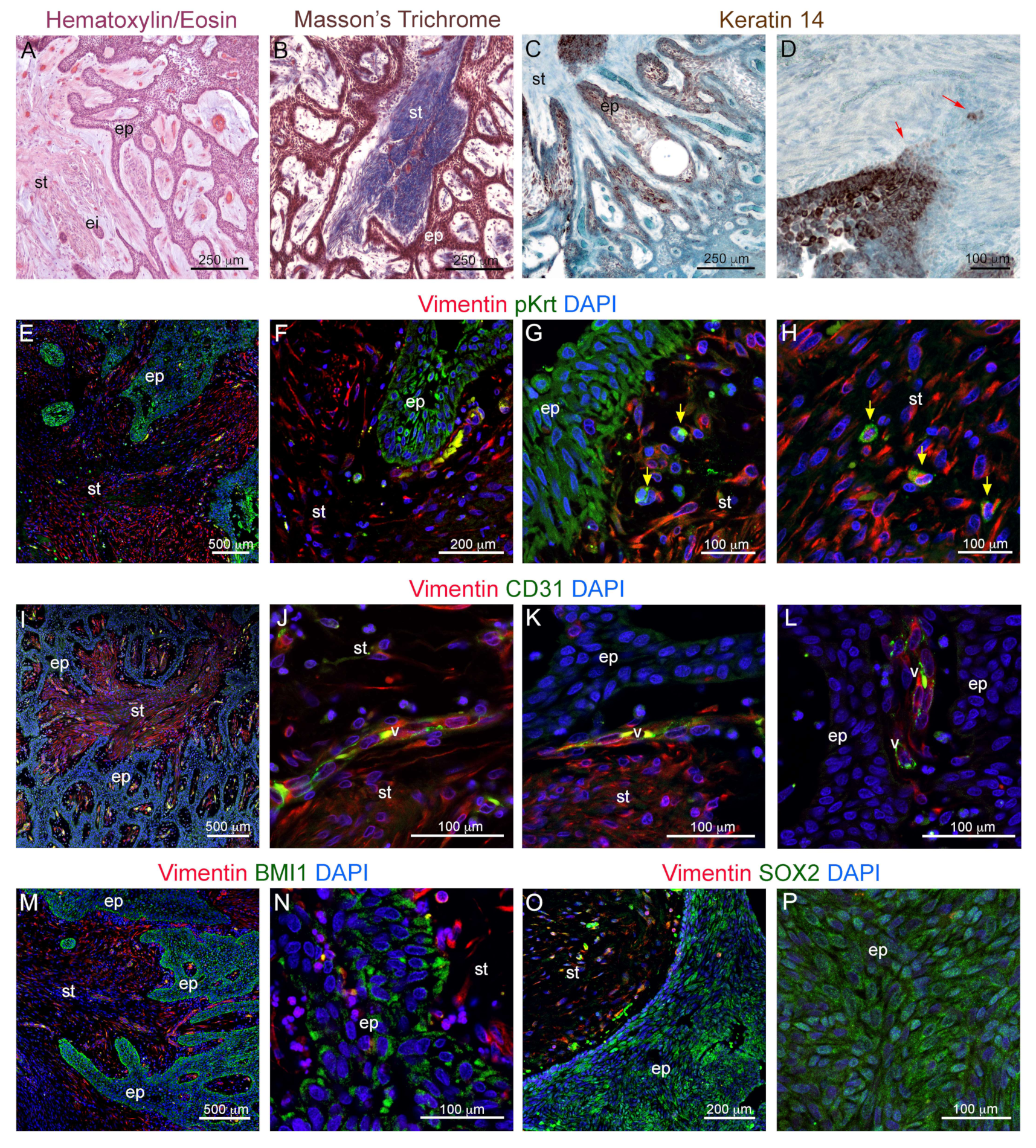
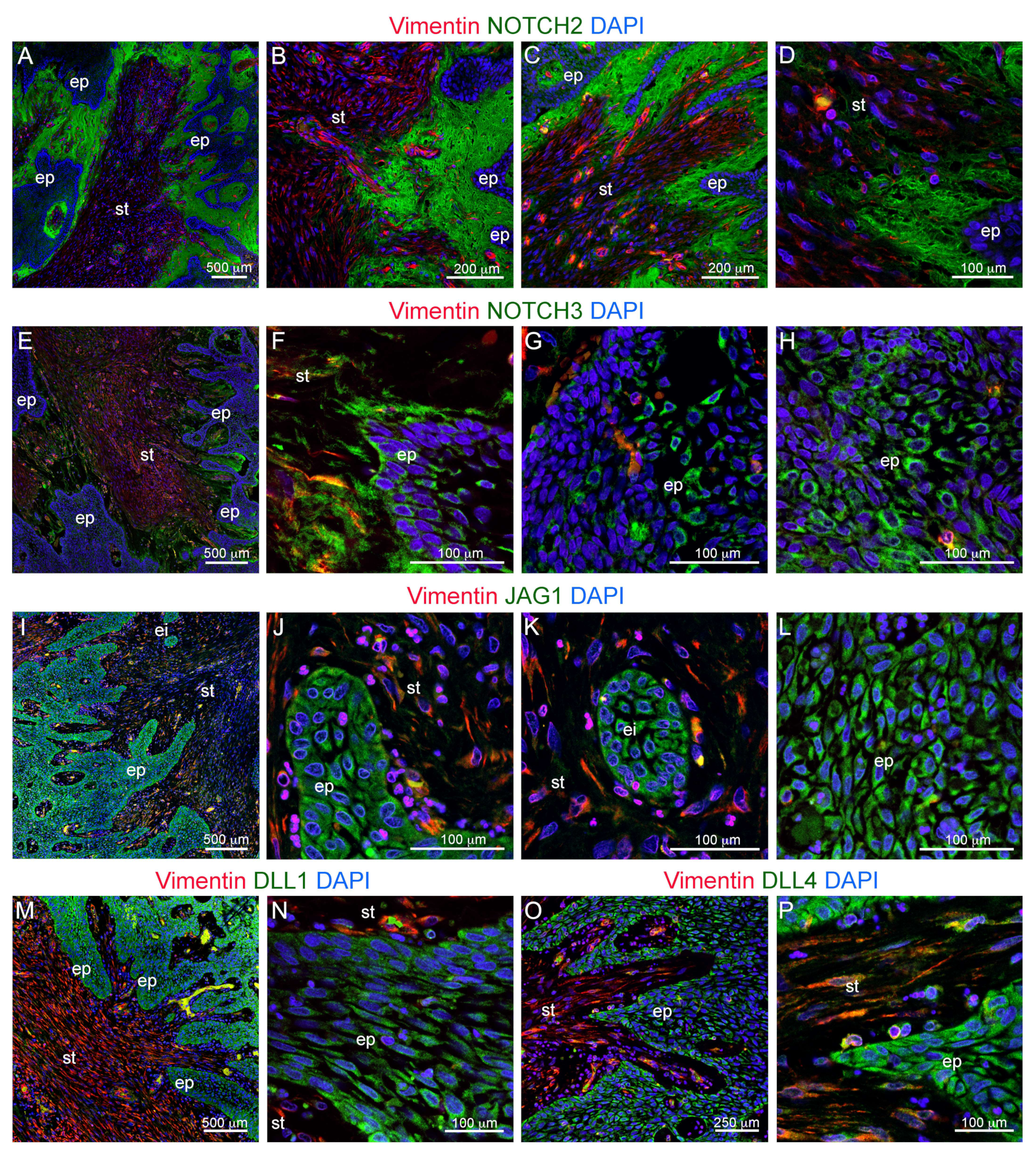
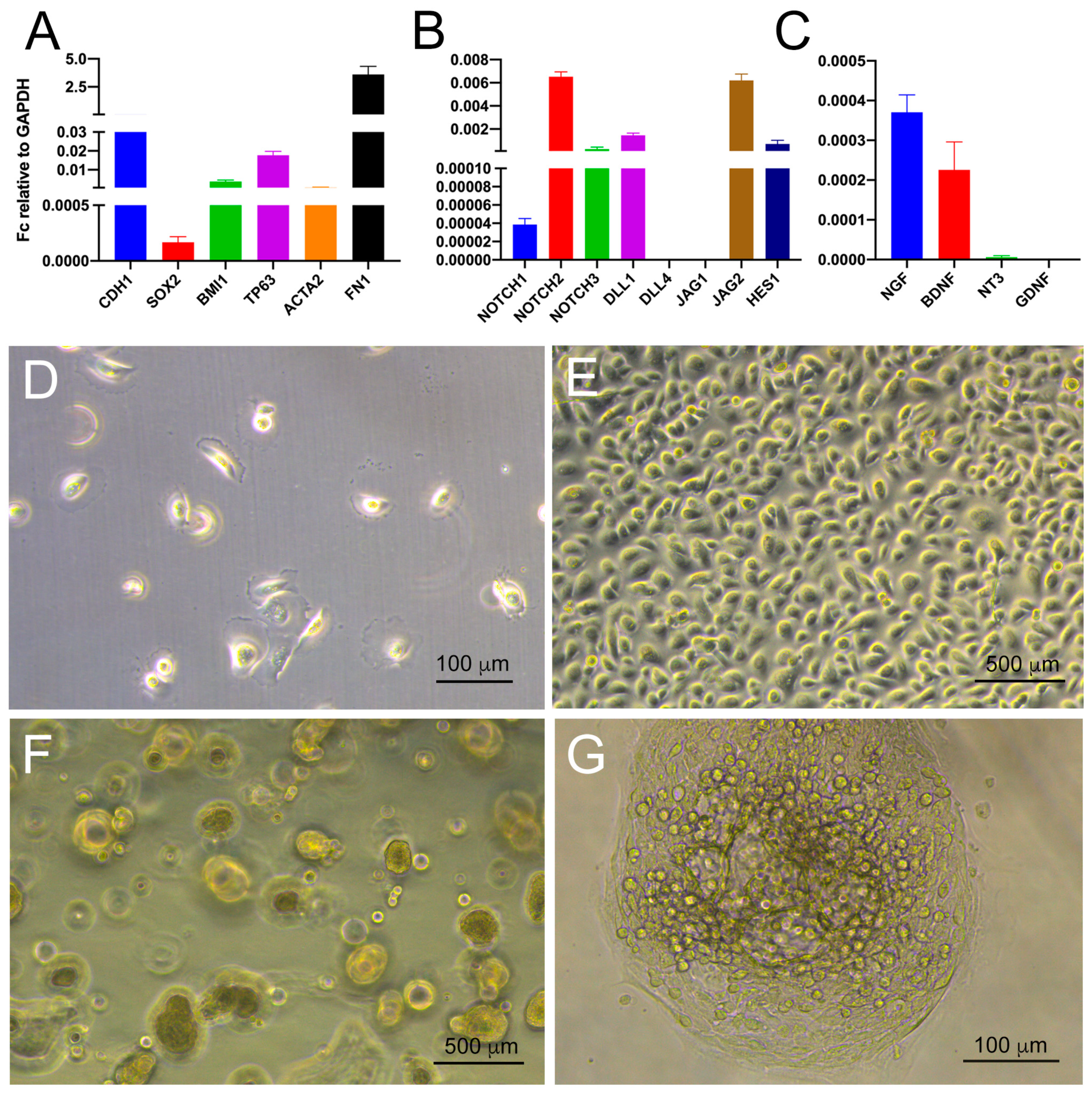
3.1. Histological and Molecular Characterization of Human Ameloblastomas
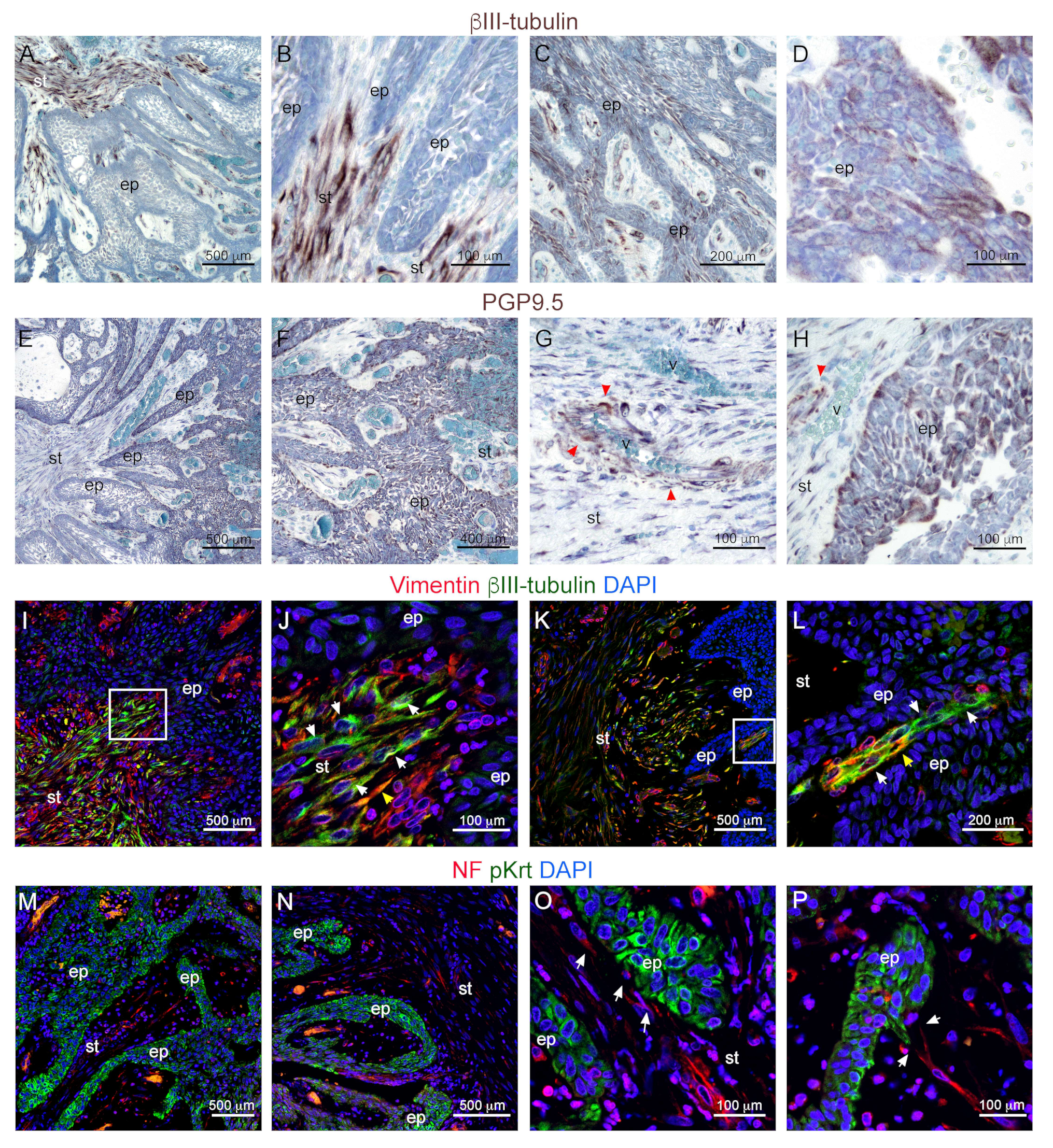
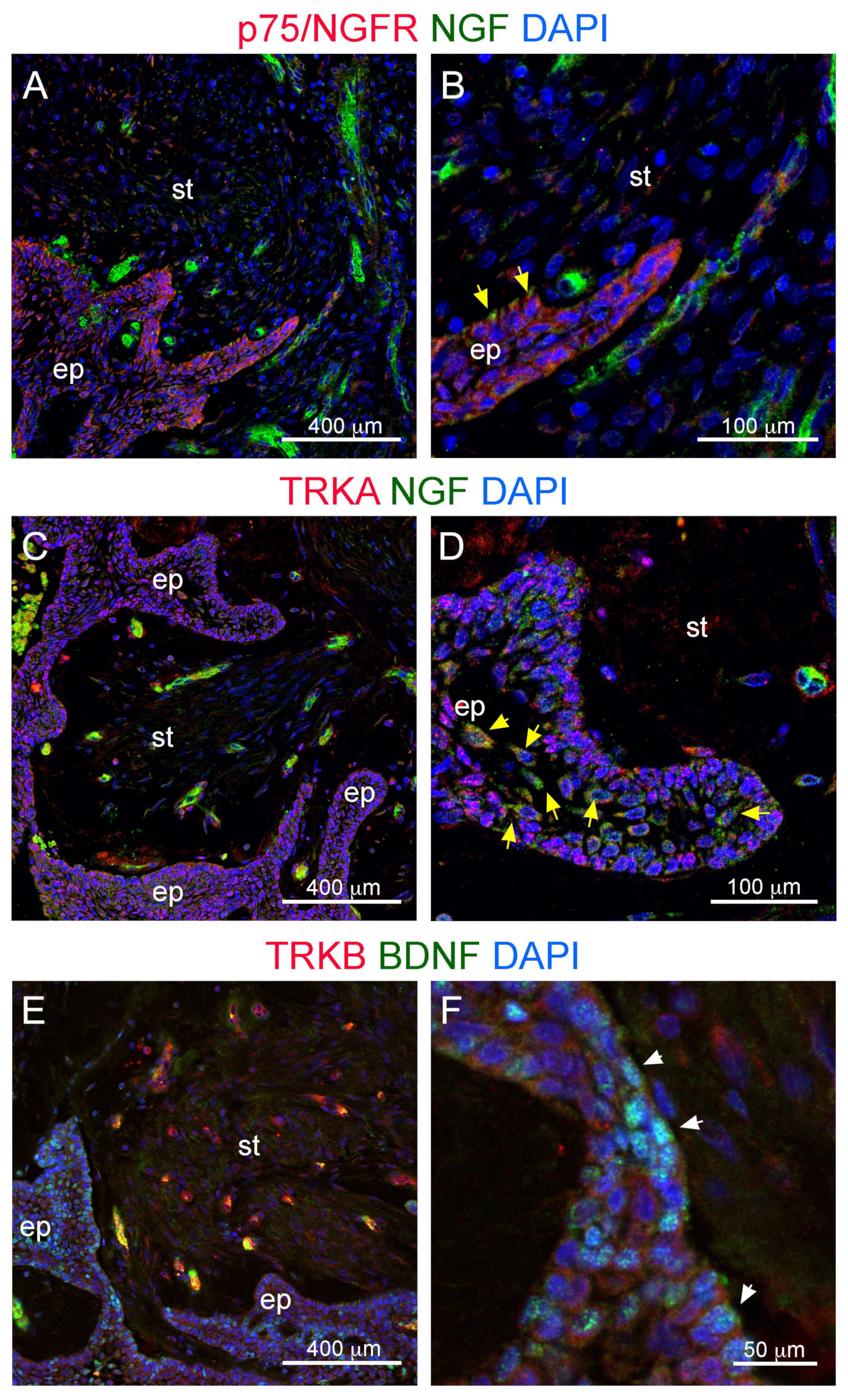
3.2. Innervation and Vascularization of Human Ameloblastomas
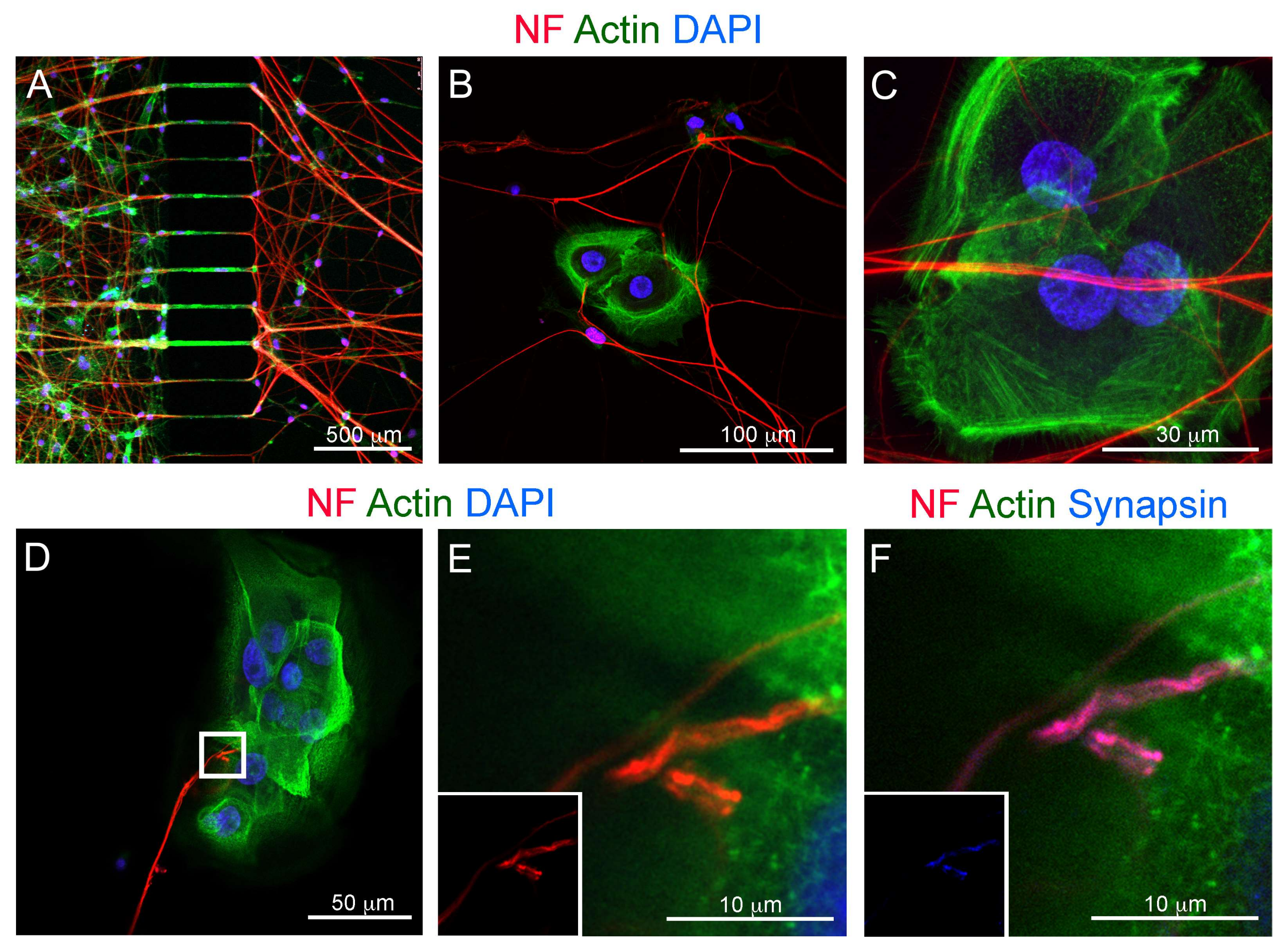
3.3. Co-Culture of Trigeminal Ganglia and Human Ameloblastoma Cells in a Microfluidics Co-Culture System
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Mendenhall, W.M.; Werning, J.W.; Fernandes, R.; Malyapa, R.S.; Mendenhall, N.P. Ameloblastoma. Am. J. Clin. Oncol 2007, 30, 645–648. [Google Scholar] [CrossRef] [PubMed]
- Wright, J.M.; Vered, M. Update from the 4th Edition of the World Health Organization Classification of Head and Neck Tumours: Odontogenic and Maxillofacial Bone Tumors. Head Neck Pathol. 2017, 11, 68–77. [Google Scholar] [CrossRef] [PubMed]
- Heikinheimo, K.; Kurppa, K.J.; Elenius, K. Novel targets for the treatment of ameloblastoma. J. Dent. Res. 2015, 94, 237–240. [Google Scholar] [CrossRef] [PubMed]
- Heikinheimo, K.; Huhtala, J.M.; Thiel, A.; Kurppa, K.J.; Heikinheimo, H.; Kovac, M.; Kragelund, C.; Warfvinge, G.; Dawson, H.; Elenius, K.; et al. The Mutational Profile of Unicystic Ameloblastoma. J. Dent. Res. 2019, 98, 54–60. [Google Scholar] [CrossRef]
- Heikinheimo, K.; Kurppa, K.J.; Laiho, A.; Peltonen, S.; Berdal, A.; Bouattour, A.; Ruhin, B.; Caton, J.; Thesleff, I.; Leivo, I.; et al. Early dental epithelial transcription factors distinguish ameloblastoma from keratocystic odontogenic tumor. J. Dent. Res. 2015, 94, 101–111. [Google Scholar] [CrossRef]
- Kurppa, K.J.; Caton, J.; Morgan, P.R.; Ristimaki, A.; Ruhin, B.; Kellokoski, J.; Elenius, K.; Heikinheimo, K. High frequency of BRAF V600E mutations in ameloblastoma. J. Pathol. 2014, 232, 492–498. [Google Scholar] [CrossRef]
- Sweeney, R.T.; McClary, A.C.; Myers, B.R.; Biscocho, J.; Neahring, L.; Kwei, K.A.; Qu, K.; Gong, X.; Ng, T.; Jones, C.D.; et al. Identification of recurrent SMO and BRAF mutations in ameloblastomas. Nat. Genet. 2014, 46, 722–725. [Google Scholar] [CrossRef]
- Juuri, E.; Isaksson, S.; Jussila, M.; Heikinheimo, K.; Thesleff, I. Expression of the stem cell marker, SOX2, in ameloblastoma and dental epithelium. Eur. J. Oral Sci. 2013, 121, 509–516. [Google Scholar] [CrossRef]
- Kumamoto, H.; Ohki, K. Detection of Notch signaling molecules in ameloblastomas. J. Oral Pathol. Med. 2008, 37, 228–234. [Google Scholar] [CrossRef]
- Nakano, K.; Takabatake, K.; Kawai, H.; Yoshida, S.; Maeda, H.; Kawakami, T.; Nagatsuka, H. Notch Signaling Affects Oral Neoplasm Cell Differentiation and Acquisition of Tumor-Specific Characteristics. Int. J. Mol. Sci 2019, 20, 1973. [Google Scholar] [CrossRef]
- Siar, C.H.; Nakano, K.; Han, P.P.; Nagatsuka, H.; Ng, K.H.; Kawakami, T. Differential expression of Notch receptors and their ligands in desmoplastic ameloblastoma. J. Oral Pathol. Med. 2010, 39, 552–558. [Google Scholar] [CrossRef] [PubMed]
- Artavanis-Tsakonas, S.; Matsuno, K.; Fortini, M.E. Notch signaling. Science 1995, 268, 225–232. [Google Scholar] [CrossRef] [PubMed]
- Artavanis-Tsakonas, S.; Rand, M.D.; Lake, R.J. Notch signaling: Cell fate control and signal integration in development. Science 1999, 284, 770–776. [Google Scholar] [CrossRef] [PubMed]
- Ho, D.M.; Artavanis-Tsakonas, S. The Notch-Mediated Proliferation Circuitry. Curr. Top. Dev. Biol 2016, 116, 17–33. [Google Scholar] [CrossRef]
- Cancer Genome Atlas, N. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 2015, 517, 576–582. [Google Scholar] [CrossRef] [PubMed]
- Fukusumi, T.; Califano, J.A. The NOTCH Pathway in Head and Neck Squamous Cell Carcinoma. J. Dent. Res. 2018, 97, 645–653. [Google Scholar] [CrossRef]
- Nyman, P.E.; Buehler, D.; Lambert, P.F. Loss of Function of Canonical Notch Signaling Drives Head and Neck Carcinogenesis. Clin. Cancer Res. 2018, 24, 6308–6318. [Google Scholar] [CrossRef]
- Porcheri, C.; Meisel, C.T.; Mitsiadis, T. Multifactorial Contribution of Notch Signaling in Head and Neck Squamous Cell Carcinoma. Int. J. Mol. Sci 2019, 20, 1520. [Google Scholar] [CrossRef]
- Sun, W.; Gaykalova, D.A.; Ochs, M.F.; Mambo, E.; Arnaoutakis, D.; Liu, Y.; Loyo, M.; Agrawal, N.; Howard, J.; Li, R.; et al. Activation of the NOTCH pathway in head and neck cancer. Cancer Res. 2014, 74, 1091–1104. [Google Scholar] [CrossRef]
- Boilly, B.; Faulkner, S.; Jobling, P.; Hondermarck, H. Nerve Dependence: From Regeneration to Cancer. Cancer Cell 2017, 31, 342–354. [Google Scholar] [CrossRef]
- Magnon, C.; Hall, S.J.; Lin, J.; Xue, X.; Gerber, L.; Freedland, S.J.; Frenette, P.S. Autonomic nerve development contributes to prostate cancer progression. Sci. 2013, 341, 1236361. [Google Scholar] [CrossRef] [PubMed]
- Ye, Y.; Dang, D.; Zhang, J.; Viet, C.T.; Lam, D.K.; Dolan, J.C.; Gibbs, J.L.; Schmidt, B.L. Nerve growth factor links oral cancer progression, pain, and cachexia. Mol. Cancer Ther. 2011, 10, 1667–1676. [Google Scholar] [CrossRef] [PubMed]
- Vermeer, P.D. Exosomal Induction of Tumor Innervation. Cancer Res. 2019, 79, 3529–3535. [Google Scholar] [CrossRef] [PubMed]
- Renz, B.W.; Takahashi, R.; Tanaka, T.; Macchini, M.; Hayakawa, Y.; Dantes, Z.; Maurer, H.C.; Chen, X.; Jiang, Z.; Westphalen, C.B.; et al. beta2 Adrenergic-Neurotrophin Feedforward Loop Promotes Pancreatic Cancer. Cancer Cell 2018, 34, 863–867. [Google Scholar] [CrossRef] [PubMed]
- Hayakawa, Y.; Sakitani, K.; Konishi, M.; Asfaha, S.; Niikura, R.; Tomita, H.; Renz, B.W.; Tailor, Y.; Macchini, M.; Middelhoff, M.; et al. Nerve Growth Factor Promotes Gastric Tumorigenesis through Aberrant Cholinergic Signaling. Cancer Cell 2017, 31, 21–34. [Google Scholar] [CrossRef]
- Sinha, S.; Fu, Y.Y.; Grimont, A.; Ketcham, M.; Lafaro, K.; Saglimbeni, J.A.; Askan, G.; Bailey, J.M.; Melchor, J.P.; Zhong, Y.; et al. PanIN Neuroendocrine Cells Promote Tumorigenesis via Neuronal Cross-talk. Cancer Res. 2017, 77, 1868–1879. [Google Scholar] [CrossRef]
- Zhao, C.M.; Hayakawa, Y.; Kodama, Y.; Muthupalani, S.; Westphalen, C.B.; Andersen, G.T.; Flatberg, A.; Johannessen, H.; Friedman, R.A.; Renz, B.W.; et al. Denervation suppresses gastric tumorigenesis. Sci. Transl. Med. 2014, 6, 250ra115. [Google Scholar] [CrossRef]
- Gerbershagen, H.J.; Ozgur, E.; Straub, K.; Dagtekin, O.; Gerbershagen, K.; Petzke, F.; Heidenreich, A.; Lehmann, K.A.; Sabatowski, R. Prevalence, severity, and chronicity of pain and general health-related quality of life in patients with localized prostate cancer. Eur. J. Pain 2008, 12, 339–350. [Google Scholar] [CrossRef]
- Jimenez-Andrade, J.M.; Ghilardi, J.R.; Castaneda-Corral, G.; Kuskowski, M.A.; Mantyh, P.W. Preventive or late administration of anti-NGF therapy attenuates tumor-induced nerve sprouting, neuroma formation, and cancer pain. Pain 2011, 152, 2564–2574. [Google Scholar] [CrossRef]
- Effiom, O.A.; Ogundana, O.M.; Akinshipo, A.O.; Akintoye, S.O. Ameloblastoma: Current etiopathological concepts and management. Oral Dis. 2018, 24, 307–316. [Google Scholar] [CrossRef]
- Ingber, D.E. Developmentally inspired human ‘organs on chips’. Development 2018, 145. [Google Scholar] [CrossRef]
- Pagella, P.; Neto, E.; Lamghari, M.; Mitsiadis, T.A. Investigation of orofacial stem cell niches and their innervation through microfluidic devices. Eur. Cell Mater. 2015, 29, 213–223. [Google Scholar] [CrossRef] [PubMed]
- Caton, J.; Mitsiadis, T.A.; Morgan, P.R. In vitro studies on odontogenic tumors. Methods Mol. Biol. 2012, 887, 167–177. [Google Scholar] [CrossRef] [PubMed]
- Natsiou, D.; Granchi, Z.; Mitsiadis, T.A.; Jimenez-Rojo, L. Generation of Spheres from Dental Epithelial Stem Cells. Front. Physiol. 2017, 8, 7. [Google Scholar] [CrossRef]
- Mitsiadis, T.A.; Lardelli, M.; Lendahl, U.; Thesleff, I. Expression of Notch 1, 2 and 3 is regulated by epithelial-mesenchymal interactions and retinoic acid in the developing mouse tooth and associated with determination of ameloblast cell fate. J. Cell Biol. 1995, 130, 407–418. [Google Scholar] [CrossRef] [PubMed]
- Pagella, P.; Miran, S.; Mitsiadis, T. Analysis of Developing Tooth Germ Innervation Using Microfluidic Co-culture Devices. J. Vis. Exp. 2015, e53114. [Google Scholar] [CrossRef] [PubMed]
- Pagella, P.; Neto, E.; Jimenez-Rojo, L.; Lamghari, M.; Mitsiadis, T.A. Microfluidics co-culture systems for studying tooth innervation. Front. Physiol. 2014, 5, 326. [Google Scholar] [CrossRef] [PubMed]
- Mahler Convenor, M.; Berard, M.; Feinstein, R.; Gallagher, A.; Illgen-Wilcke, B.; Pritchett-Corning, K.; Raspa, M. FELASA recommendations for the health monitoring of mouse, rat, hamster, guinea pig and rabbit colonies in breeding and experimental units. Lab. Anim. 2014, 48, 178–192. [Google Scholar] [CrossRef]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. The hallmarks of cancer. Cell 2000, 100, 57–70. [Google Scholar] [CrossRef]
- Carmeliet, P.; Tessier-Lavigne, M. Common mechanisms of nerve and blood vessel wiring. Nature 2005, 436, 193–200. [Google Scholar] [CrossRef] [PubMed]
- Thompson, R.J.; Doran, J.F.; Jackson, P.; Dhillon, A.P.; Rode, J. PGP 9.5—A new marker for vertebrate neurons and neuroendocrine cells. Brain Res. 1983, 278, 224–228. [Google Scholar] [CrossRef]
- Campbell, L.K.; Thomas, J.R.; Lamps, L.W.; Smoller, B.R.; Folpe, A.L. Protein gene product 9.5 (PGP 9.5) is not a specific marker of neural and nerve sheath tumors: An immunohistochemical study of 95 mesenchymal neoplasms. Mod. Pathol. 2003, 16, 963–969. [Google Scholar] [CrossRef]
- You, Z.; Liu, S.P.; Du, J.; Wu, Y.H.; Zhang, S.Z. Advancements in MAPK signaling pathways and MAPK-targeted therapies for ameloblastoma: A review. J. Oral Pathol. Med. 2019, 48, 201–205. [Google Scholar] [CrossRef]
- Nassar, D.; Blanpain, C. Cancer Stem Cells: Basic Concepts and Therapeutic Implications. Annu. Rev. Pathol. 2016, 11, 47–76. [Google Scholar] [CrossRef]
- Makena, M.R.; Ranjan, A.; Thirumala, V.; Reddy, A.P. Cancer stem cells: Road to therapeutic resistance and strategies to overcome resistance. Biochim. Biophys. Acta Mol. Basis Dis. 2018, 165339. [Google Scholar] [CrossRef]
- Batlle, E.; Clevers, H. Cancer stem cells revisited. Nat. Med. 2017, 23, 1124–1134. [Google Scholar] [CrossRef]
- Guruharsha, K.G.; Kankel, M.W.; Artavanis-Tsakonas, S. The Notch signalling system: Recent insights into the complexity of a conserved pathway. Nat. Rev. Genet. 2012, 13, 654–666. [Google Scholar] [CrossRef]
- Clara, J.A.; Monge, C.; Yang, Y.; Takebe, N. Targeting signalling pathways and the immune microenvironment of cancer stem cells—A clinical update. Nat. Rev. Clin. Oncol. 2019. [Google Scholar] [CrossRef]
- Rajakulendran, N.; Rowland, K.J.; Selvadurai, H.J.; Ahmadi, M.; Park, N.I.; Naumenko, S.; Dolma, S.; Ward, R.J.; So, M.; Lee, L.; et al. Wnt and Notch signaling govern self-renewal and differentiation in a subset of human glioblastoma stem cells. Genes Dev. 2019, 33, 498–510. [Google Scholar] [CrossRef] [PubMed]
- Ouadah, Y.; Rojas, E.R.; Riordan, D.P.; Capostagno, S.; Kuo, C.S.; Krasnow, M.A. Rare Pulmonary Neuroendocrine Cells Are Stem Cells Regulated by Rb, p53, and Notch. Cell 2019, 179, 403–416 e423. [Google Scholar] [CrossRef] [PubMed]
- Meurette, O.; Mehlen, P. Notch Signaling in the Tumor Microenvironment. Cancer Cell 2018, 34, 536–548. [Google Scholar] [CrossRef] [PubMed]
- Mitsiadis, T.A.; Graf, D.; Luder, H.; Gridley, T.; Bluteau, G. BMPs and FGFs target Notch signalling via jagged 2 to regulate tooth morphogenesis and cytodifferentiation. Development 2010, 137, 3025–3035. [Google Scholar] [CrossRef]
- Mucchielli, M.L.; Mitsiadis, T.A. Correlation of asymmetric Notch2 expression and mouse incisor rotation. Mech. Dev. 2000, 91, 379–382. [Google Scholar] [CrossRef]
- Mitsiadis, T.A.; Hirsinger, E.; Lendahl, U.; Goridis, C. Delta-notch signaling in odontogenesis: Correlation with cytodifferentiation and evidence for feedback regulation. Dev. Biol. 1998, 204, 420–431. [Google Scholar] [CrossRef][Green Version]
- Mitsiadis, T.A.; Henrique, D.; Thesleff, I.; Lendahl, U. Mouse Serrate-1 (Jagged-1): Expression in the developing tooth is regulated by epithelial-mesenchymal interactions and fibroblast growth factor-4. Development 1997, 124, 1473–1483. [Google Scholar]
- Mitsiadis, T.A.; Thesleff, I.; Lendahl, U. Mouse Serrate-1 and Notch genes differ in their response to signaling molecules during odontogenesis. Int. J. Dev. Biol. 1996, (Suppl. 1), 113S–114S. [Google Scholar]
- Mitsiadis, T.A.; Graf, D. Cell fate determination during tooth development and regeneration. Birth Defects Res. C. Embryo Today 2009, 87, 199–211. [Google Scholar] [CrossRef]
- Mitsiadis, T.A.; Caton, J.; Pagella, P.; Orsini, G.; Jimenez-Rojo, L. Monitoring Notch Signaling-Associated Activation of Stem Cell Niches within Injured Dental Pulp. Front. Physiol. 2017, 8, 372. [Google Scholar] [CrossRef]
- Mitsiadis, T.A.; Feki, A.; Papaccio, G.; Caton, J. Dental pulp stem cells, niches, and notch signaling in tooth injury. Adv. Dent. Res. 2011, 23, 275–279. [Google Scholar] [CrossRef] [PubMed]
- Lovschall, H.; Mitsiadis, T.A.; Poulsen, K.; Jensen, K.H.; Kjeldsen, A.L. Coexpression of Notch3 and Rgs5 in the pericyte-vascular smooth muscle cell axis in response to pulp injury. Int. J. Dev. Biol. 2007, 51, 715–721. [Google Scholar] [CrossRef] [PubMed]
- Mitsiadis, T.A.; Romeas, A.; Lendahl, U.; Sharpe, P.T.; Farges, J.C. Notch2 protein distribution in human teeth under normal and pathological conditions. Exp. Cell Res. 2003, 282, 101–109. [Google Scholar] [CrossRef]
- Mitsiadis, T.A.; Fried, K.; Goridis, C. Reactivation of Delta-Notch signaling after injury: Complementary expression patterns of ligand and receptor in dental pulp. Exp. Cell Res. 1999, 246, 312–318. [Google Scholar] [CrossRef] [PubMed]
- Shen, K.; Luk, S.; Hicks, D.F.; Elman, J.S.; Bohr, S.; Iwamoto, Y.; Murray, R.; Pena, K.; Wang, F.; Seker, E.; et al. Resolving cancer-stroma interfacial signalling and interventions with micropatterned tumour-stromal assays. Nat. Commun. 2014, 5, 5662. [Google Scholar] [CrossRef]
- Jackstadt, R.; van Hooff, S.R.; Leach, J.D.; Cortes-Lavaud, X.; Lohuis, J.O.; Ridgway, R.A.; Wouters, V.M.; Roper, J.; Kendall, T.J.; Roxburgh, C.S.; et al. Epithelial NOTCH Signaling Rewires the Tumor Microenvironment of Colorectal Cancer to Drive Poor-Prognosis Subtypes and Metastasis. Cancer Cell 2019, 36, 319–336. [Google Scholar] [CrossRef]
- Siar, C.H.; Nagatsuka, H.; Chuah, K.S.; Rivera, R.S.; Nakano, K.; Ng, K.H.; Kawakami, T. Notch4 overexpression in ameloblastoma correlates with the solid/multicystic phenotype. Oral Surg. Oral Med. Oral Pathol. Oral Radiol. Endod. 2010, 110, 224–233. [Google Scholar] [CrossRef]
- Pagella, P.; Jimenez-Rojo, L.; Mitsiadis, T.A. Roles of innervation in developing and regenerating orofacial tissues. Cell Mol. Life Sci. 2014, 71, 2241–2251. [Google Scholar] [CrossRef]
- Portenoy, R.K. Treatment of cancer pain. Lancet 2011, 377, 2236–2247. [Google Scholar] [CrossRef]
- Adriaenssens, E.; Vanhecke, E.; Saule, P.; Mougel, A.; Page, A.; Romon, R.; Nurcombe, V.; Le Bourhis, X.; Hondermarck, H. Nerve growth factor is a potential therapeutic target in breast cancer. Cancer Res. 2008, 68, 346–351. [Google Scholar] [CrossRef]
- Aloe, L.; Rocco, M.L.; Balzamino, B.O.; Micera, A. Nerve growth factor: Role in growth, differentiation and controlling cancer cell development. J. Exp. Clin. Cancer Res. 2016, 35, 116. [Google Scholar] [CrossRef] [PubMed]
- Chopin, V.; Lagadec, C.; Toillon, R.A.; Le Bourhis, X. Neurotrophin signaling in cancer stem cells. Cell Mol. Life Sci. 2016, 73, 1859–1870. [Google Scholar] [CrossRef] [PubMed]
- Griffin, N.; Faulkner, S.; Jobling, P.; Hondermarck, H. Targeting neurotrophin signaling in cancer: The renaissance. Pharmacol. Res. 2018, 135, 12–17. [Google Scholar] [CrossRef] [PubMed]
- Vanhecke, E.; Adriaenssens, E.; Verbeke, S.; Meignan, S.; Germain, E.; Berteaux, N.; Nurcombe, V.; Le Bourhis, X.; Hondermarck, H. Brain-derived neurotrophic factor and neurotrophin-4/5 are expressed in breast cancer and can be targeted to inhibit tumor cell survival. Clin. Cancer Res. 2011, 17, 1741–1752. [Google Scholar] [CrossRef]
- Amit, M.; Takahashi, H.; Dragomir, M.P.; Lindemann, A.; Gleber-Netto, F.O.; Pickering, C.R.; Anfossi, S.; Osman, A.A.; Cai, Y.; Wang, R.; et al. Loss of p53 drives neuron reprogramming in head and neck cancer. Nature 2020, 578, 449–454. [Google Scholar] [CrossRef]
- Saloman, J.L.; Albers, K.M.; Li, D.; Hartman, D.J.; Crawford, H.C.; Muha, E.A.; Rhim, A.D.; Davis, B.M. Ablation of sensory neurons in a genetic model of pancreatic ductal adenocarcinoma slows initiation and progression of cancer. Proc. Natl. Acad. Sci. USA 2016, 113, 3078–3083. [Google Scholar] [CrossRef]
- Kamiya, A.; Hayama, Y.; Kato, S.; Shimomura, A.; Shimomura, T.; Irie, K.; Kaneko, R.; Yanagawa, Y.; Kobayashi, K.; Ochiya, T. Genetic manipulation of autonomic nerve fiber innervation and activity and its effect on breast cancer progression. Nat. Neurosci. 2019, 22, 1289–1305. [Google Scholar] [CrossRef]
- Madeo, M.; Colbert, P.L.; Vermeer, D.W.; Lucido, C.T.; Cain, J.T.; Vichaya, E.G.; Grossberg, A.J.; Muirhead, D.; Rickel, A.P.; Hong, Z.; et al. Cancer exosomes induce tumor innervation. Nat. Commun. 2018, 9, 4284. [Google Scholar] [CrossRef]
- Allen, J.K.; Armaiz-Pena, G.N.; Nagaraja, A.S.; Sadaoui, N.C.; Ortiz, T.; Dood, R.; Ozcan, M.; Herder, D.M.; Haemmerle, M.; Gharpure, K.M.; et al. Sustained Adrenergic Signaling Promotes Intratumoral Innervation through BDNF Induction. Cancer Res. 2018, 78, 3233–3242. [Google Scholar] [CrossRef]
- Lagadec, C.; Meignan, S.; Adriaenssens, E.; Foveau, B.; Vanhecke, E.; Romon, R.; Toillon, R.A.; Oxombre, B.; Hondermarck, H.; Le Bourhis, X. TrkA overexpression enhances growth and metastasis of breast cancer cells. Oncogene 2009, 28, 1960–1970. [Google Scholar] [CrossRef]
- Contreras-Zarate, M.J.; Day, N.L.; Ormond, D.R.; Borges, V.F.; Tobet, S.; Gril, B.; Steeg, P.S.; Cittelly, D.M. Estradiol induces BDNF/TrkB signaling in triple-negative breast cancer to promote brain metastases. Oncogene 2019, 38, 4685–4699. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Hao, Q.; Liao, P.; Luo, S.; Zhang, M.; Hu, G.; Liu, H.; Zhang, Y.; Cao, B.; Baddoo, M.; et al. Nerve growth factor receptor negates the tumor suppressor p53 as a feedback regulator. Elife 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Di Donato, M.; Cernera, G.; Auricchio, F.; Migliaccio, A.; Castoria, G. Cross-talk between androgen receptor and nerve growth factor receptor in prostate cancer cells: Implications for a new therapeutic approach. Cell Death Discov. 2018, 4, 5. [Google Scholar] [CrossRef] [PubMed]
- Blondy, S.; Christou, N.; David, V.; Verdier, M.; Jauberteau, M.O.; Mathonnet, M.; Perraud, A. Neurotrophins and their involvement in digestive cancers. Cell Death Dis. 2019, 10, 123. [Google Scholar] [CrossRef]
- Saloman, J.L.; Albers, K.M.; Rhim, A.D.; Davis, B.M. Can Stopping Nerves, Stop Cancer? Trends Neurosci. 2016, 39, 880–889. [Google Scholar] [CrossRef]
- Melhem-Bertrandt, A.; Chavez-Macgregor, M.; Lei, X.; Brown, E.N.; Lee, R.T.; Meric-Bernstam, F.; Sood, A.K.; Conzen, S.D.; Hortobagyi, G.N.; Gonzalez-Angulo, A.M. Beta-blocker use is associated with improved relapse-free survival in patients with triple-negative breast cancer. J. Clin. Oncol. 2011, 29, 2645–2652. [Google Scholar] [CrossRef]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pagella, P.; Catón, J.; Meisel, C.T.; Mitsiadis, T.A. Ameloblastomas Exhibit Stem Cell Potential, Possess Neurotrophic Properties, and Establish Connections with Trigeminal Neurons. Cells 2020, 9, 644. https://doi.org/10.3390/cells9030644
Pagella P, Catón J, Meisel CT, Mitsiadis TA. Ameloblastomas Exhibit Stem Cell Potential, Possess Neurotrophic Properties, and Establish Connections with Trigeminal Neurons. Cells. 2020; 9(3):644. https://doi.org/10.3390/cells9030644
Chicago/Turabian StylePagella, Pierfrancesco, Javier Catón, Christian T. Meisel, and Thimios A. Mitsiadis. 2020. "Ameloblastomas Exhibit Stem Cell Potential, Possess Neurotrophic Properties, and Establish Connections with Trigeminal Neurons" Cells 9, no. 3: 644. https://doi.org/10.3390/cells9030644
APA StylePagella, P., Catón, J., Meisel, C. T., & Mitsiadis, T. A. (2020). Ameloblastomas Exhibit Stem Cell Potential, Possess Neurotrophic Properties, and Establish Connections with Trigeminal Neurons. Cells, 9(3), 644. https://doi.org/10.3390/cells9030644





