Insights into Inflammatory Priming of Adipose-Derived Mesenchymal Stem Cells: Validation of Extracellular Vesicles-Embedded miRNA Reference Genes as A Crucial Step for Donor Selection
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. ASCs Isolation and Expansion
2.3. ASC Characterization
2.4. iASC-EV Isolation and Characterization
2.5. Candidate RGs Selection
2.6. Selection of OA-Related miRNAs
2.7. Total RNA Isolation and miRNA Profiling
2.8. Data Analysis
2.9. Statistical Analysis
3. Results
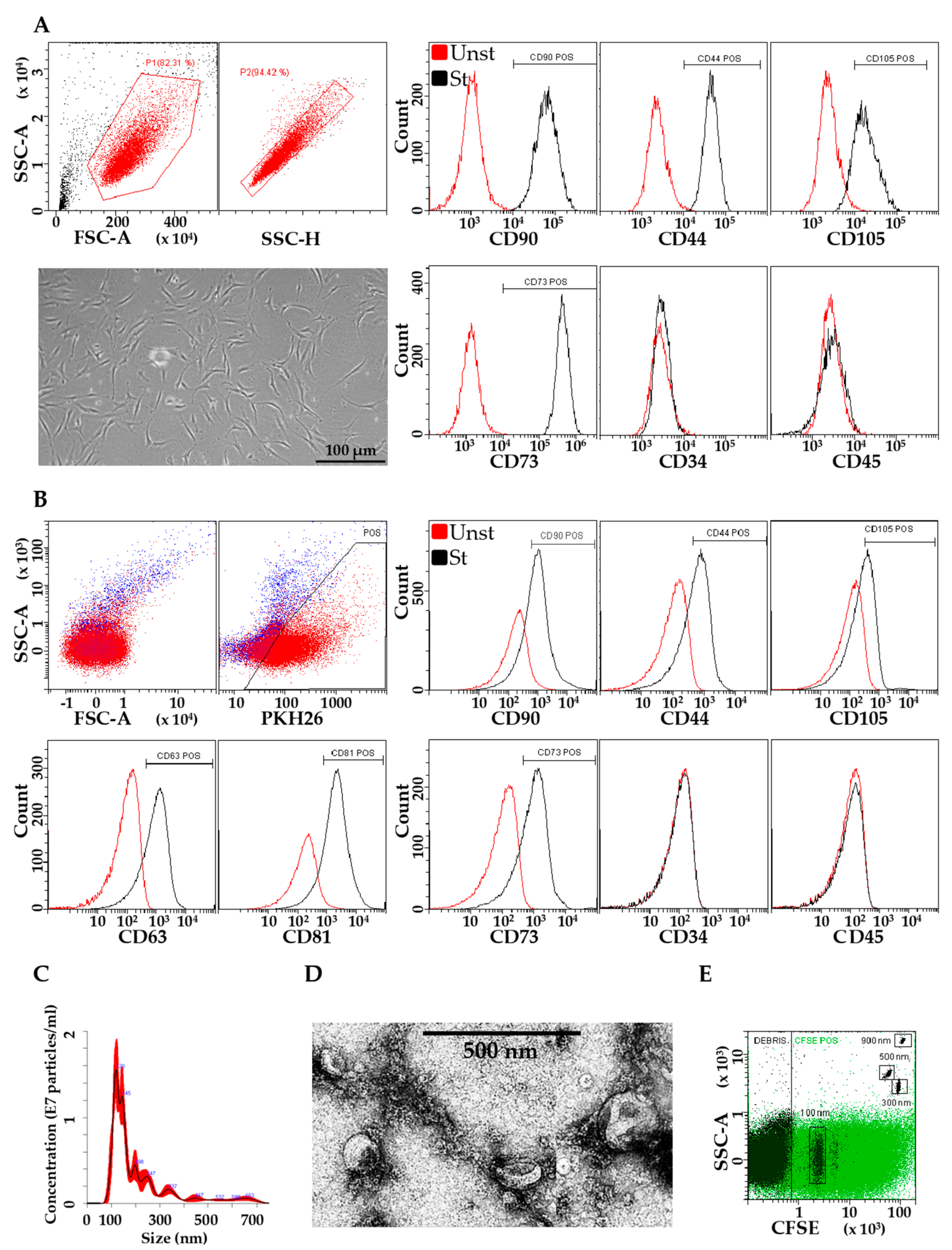
3.1. Characterization of ASCs and EVs
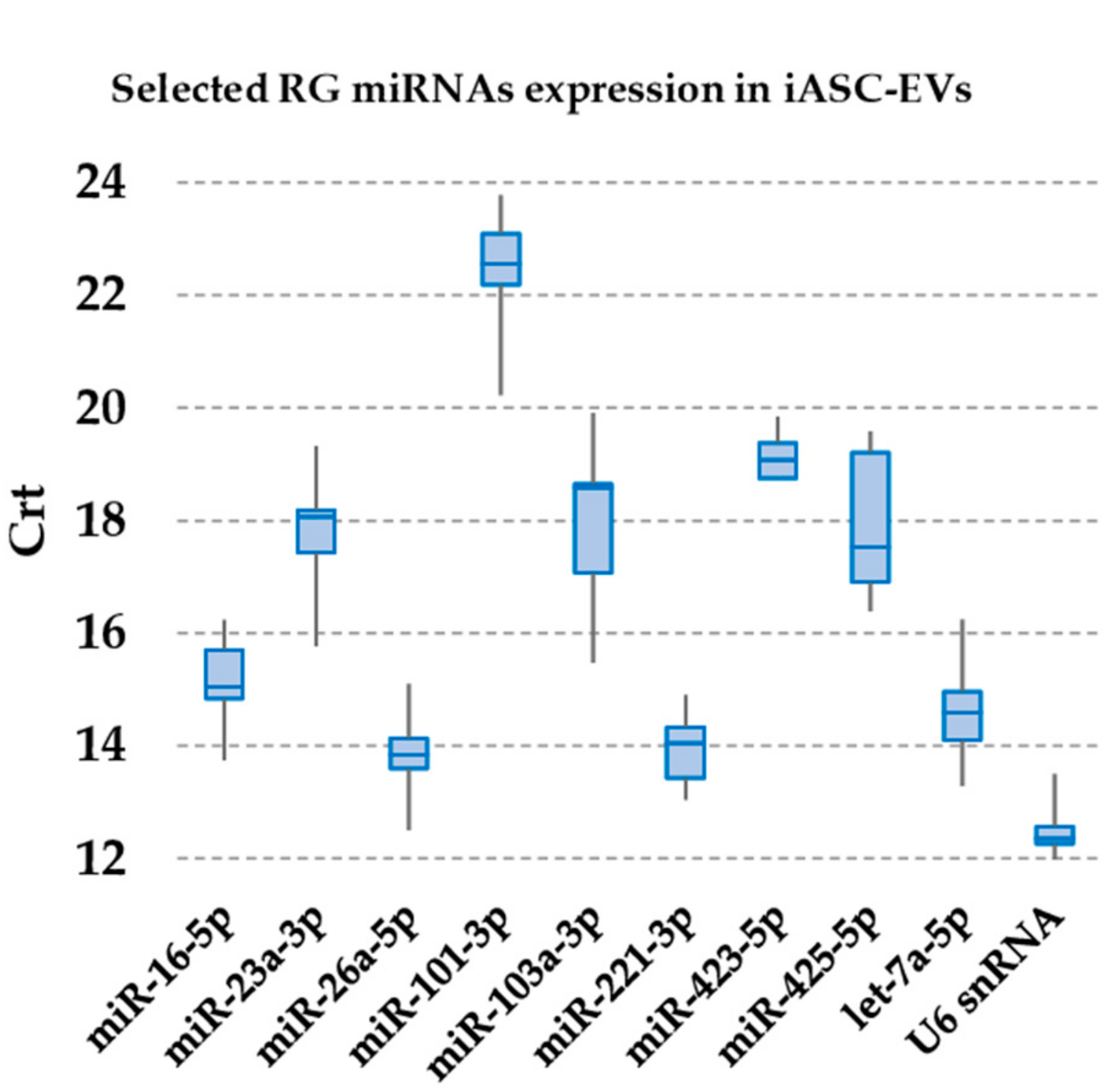
3.2. Expression of Candidate Reference miRNAs
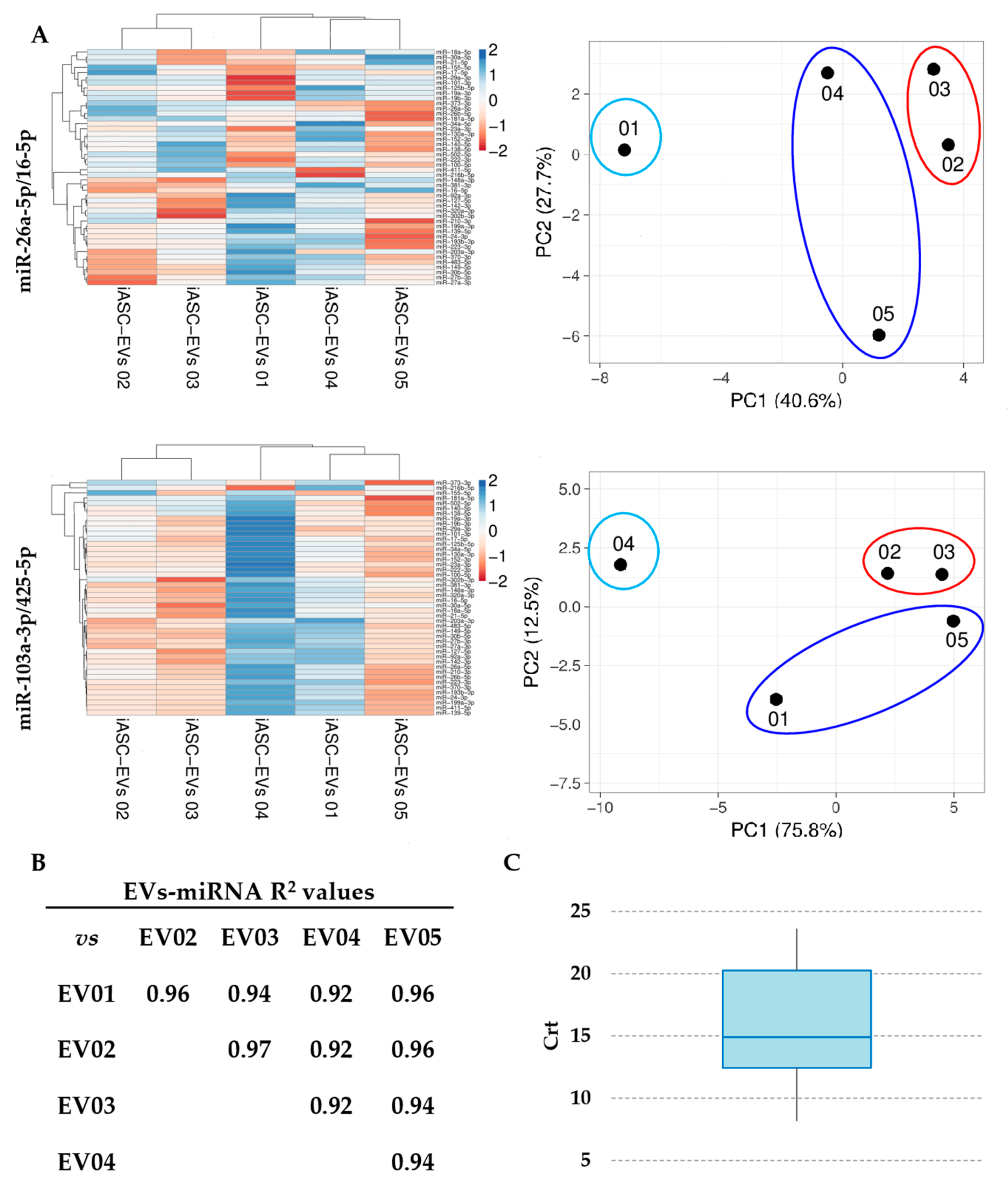
3.3. Expression Stability Analysis of RG miRNAs
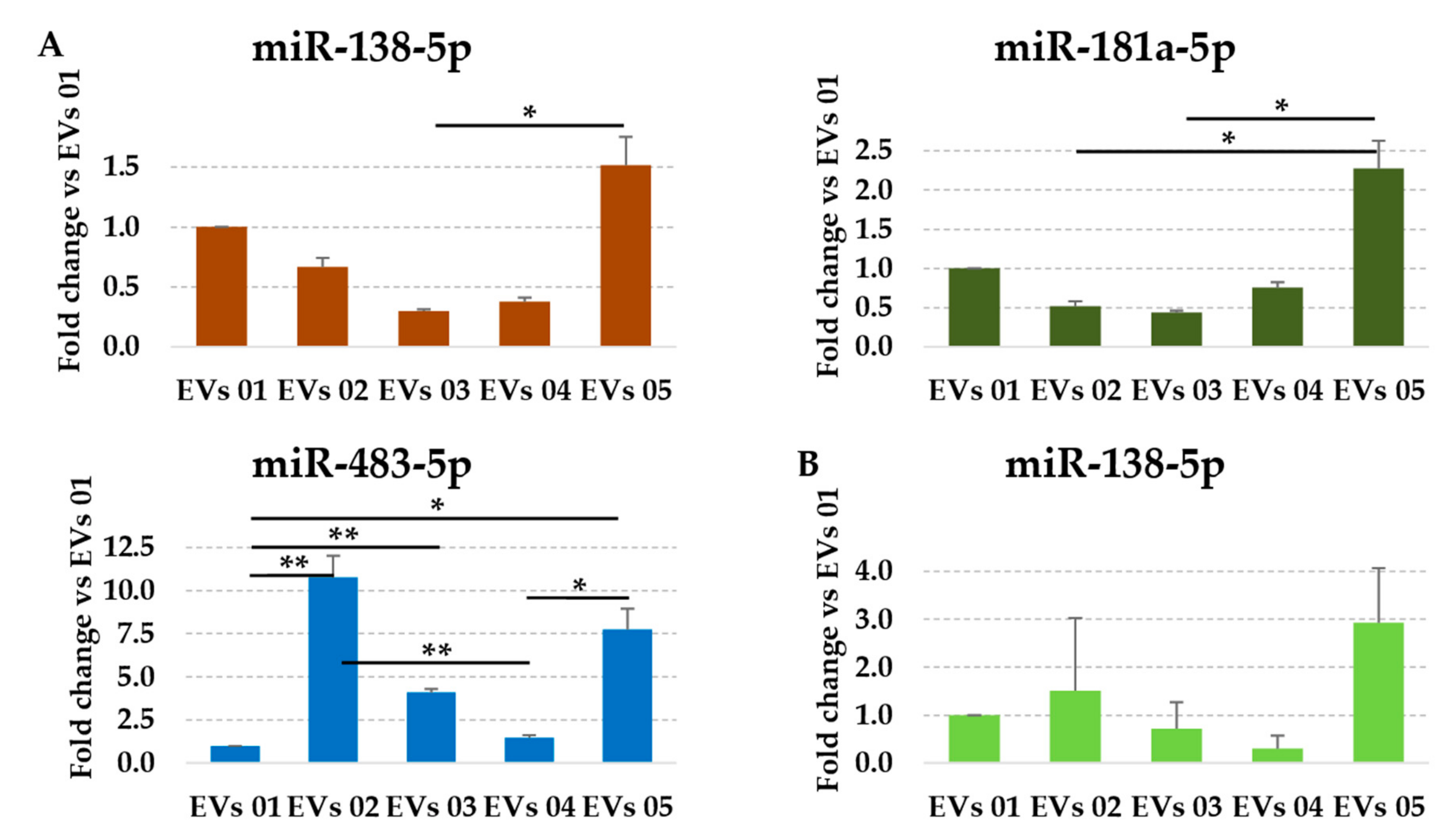
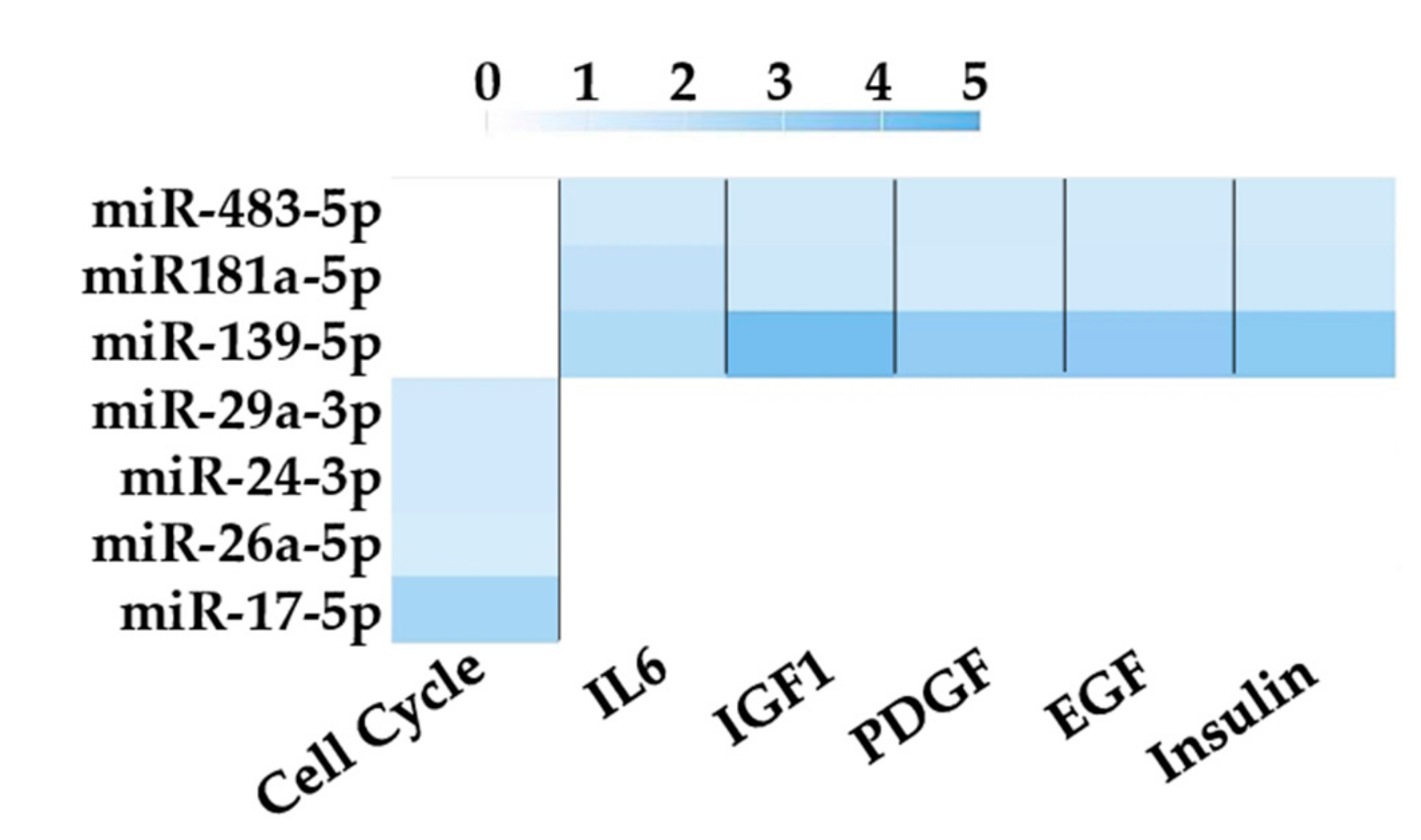
3.4. Assessment of Identified RGs on OA-Related miRNAs
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Heathman, T.R.; Nienow, A.W.; McCall, M.J.; Coopman, K.; Kara, B.; Hewitt, C.J. The translation of cell-based therapies: Clinical landscape and manufacturing challenges. Regen. Med. 2015, 10, 49–64. [Google Scholar] [CrossRef] [PubMed]
- Fitzsimmons, R.E.B.; Mazurek, M.S.; Soos, A.; Simmons, C.A. Mesenchymal Stromal/Stem Cells in Regenerative Medicine and Tissue Engineering. Stem Cells Int. 2018, 18, 8031718. [Google Scholar] [CrossRef] [PubMed]
- Vizoso, F.J.; Eiro, N.; Cid, S.; Schneider, J.; Perez-Fernandez, R. Mesenchymal Stem Cell Secretome: Toward Cell-Free Therapeutic Strategies in Regenerative Medicine. Int. J. Mol. Sci. 2017, 18, 1852. [Google Scholar] [CrossRef] [PubMed]
- Caplan, A.I. Mesenchymal Stem Cells: Time to Change the Name! Stem Cells Transl. Med. 2017, 6, 1445–1451. [Google Scholar] [CrossRef]
- Dumont, J.; Euwart, D.; Mei, B.; Estes, S.; Kshirsagar, R. Human cell lines for biopharmaceutical manufacturing: History, status, and future perspectives. Crit. Rev. Biotechnol. 2016, 36, 1110–1122. [Google Scholar] [CrossRef]
- Williams, D.J.; Thomas, R.J.; Hourd, P.C.; Chandra, A.; Ratcliffe, E.; Liu, Y.; Rayment, E.A.; Archer, J.R. Precision manufacturing for clinical-quality regenerative medicines. Philos. Trans. A Math. Phys. Eng. Sci. 2012, 370, 3924–3949. [Google Scholar] [CrossRef] [PubMed]
- Carmen, J.; Burger, S.R.; McCaman, M.; Rowley, J.A. Developing assays to address identity, potency, purity and safety: Cell characterization in cell therapy process development. Regen. Med. 2012, 7, 85–100. [Google Scholar] [CrossRef] [PubMed]
- Heathman, T.R.J.; Rafiq, Q.A.; Chan, A.K.C.; Coopman, K.; Nienow, A.W.; Kara, B.; Hewitt, C.J. Characterization of human mesenchymal stem cells from multiple donors and the implications for large scale bioprocess development. Biochem. Engin. J. 2016, 108, 14–23. [Google Scholar] [CrossRef]
- Redondo-Castro, E.; Cunningham, C.; Miller, J.; Martuscelli, L.; Aoulad-Ali, S.; Rothwell, N.J.; Kielty, C.M.; Allan, S.M.; Pinteaux, E. Interleukin-1 primes human mesenchymal stem cells towards an anti-inflammatory and pro-trophic phenotype in vitro. Stem Cell Res. Ther. 2017, 8, 79. [Google Scholar] [CrossRef]
- Szabó, E.; Fajka-Boja, R.; Kriston-Pál, É.; Hornung, Á.; Makra, I.; Kudlik, G.; Uher, F.; Katona, R.L.; Monostori, É.; Czibula, Á. Licensing by Inflammatory Cytokines Abolishes Heterogeneity of Immunosuppressive Function of Mesenchymal Stem Cell Population. Stem Cells Dev. 2015, 24, 2171–2180. [Google Scholar] [CrossRef]
- Krampera, M. Mesenchymal stromal cell ‘licensing’: A multistep process. Leukemia 2011, 25, 1408–1414. [Google Scholar] [CrossRef]
- Polchert, D.; Sobinsky, J.; Douglas, G.; Kidd, M.; Moadsiri, A.; Reina, E.; Genrich, K.; Mehrotra, S.; Setty, S.; Smith, B.; et al. IFN-gamma activation of mesenchymal stem cells for treatment and prevention of graft versus host disease. Eur. J. Immunol. 2008, 38, 1745–1755. [Google Scholar] [CrossRef]
- Ruiz, M.; Cosenza, S.; Maumus, M.; Jorgensen, C.; Noël, D. Therapeutic application of mesenchymal stem cells in osteoarthritis. Expert Opin. Biol. Ther. 2016, 16, 33–42. [Google Scholar] [CrossRef]
- Manferdini, C.l.; Maumus, M.; Gabusi, E.; Paolella, F.; Grassi, F.; Jorgensen, C.; Fleury-Cappellesso, S.; Noël, D.; Lisignoli, G. Lack of anti-inflammatory and anti-catabolic effects on basal inflamed osteoarthritic chondrocytes or synoviocytes by adipose stem cell-conditioned medium. Osteoarthritis Cartilage 2015, 23, 2045–2057. [Google Scholar] [CrossRef]
- De Luca, P.; Kouroupis, D.; Viganò, M.; Perucca-Orfei, C.; Kaplan, L.; Zagra, L.; de Girolamo, L.; Correa, D.; Colombini, A. Human Diseased Articular Cartilage Contains a Mesenchymal Stem Cell-Like Population of Chondroprogenitors with Strong Immunomodulatory Responses. J. Clin. Med. 2019, 8, 423. [Google Scholar] [CrossRef] [PubMed]
- Crop, M.J.; Baan, C.C.; Korevaar, S.; Ijzermans, J.N.; Pescatori, M.; Stubbs, A.P.; van Ijcken, W.F.; Dahlke, M.H.; Eggenhofer, E.; Weimar, W.; et al. Inflammatory conditions affect gene expression and function of human adipose tissue-derived mesenchymal stem cells. Clin. Exp. Immunol. 2010, 162, 474–486. [Google Scholar] [CrossRef] [PubMed]
- Maumus, M.; Roussignol, G.; Toupet, K.; Penarier, G.; Bentz, I.; Teixeira, S.; Oustric, D.; Jung, M.; Lepage, O.; Steinberg, R.; et al. Utility of a Mouse Model of Osteoarthritis to Demonstrate Cartilage Protection by IFNγ-Primed Equine Mesenchymal Stem Cells. Front. Immunol. 2016, 7, 392. [Google Scholar] [CrossRef] [PubMed]
- Serejo, T.R.T.; Silva-Carvalho, A.É.; Braga, L.D.C.F.; Neves, F.A.R.; Pereira, R.W.; Carvalho, J.L.; Saldanha-Araujo, F. Assessment of the Immunosuppressive Potential of INF-γ Licensed Adipose Mesenchymal Stem Cells, Their Secretome and Extracellular Vesicles. Cells 2019, 8, 22. [Google Scholar] [CrossRef] [PubMed]
- Lopa, S.; Colombini, A.; Stanco, D.; de Girolamo, L.; Sansone, V.; Moretti, M. Donor-matched mesenchymal stem cells from knee infrapatellar and subcutaneous adipose tissue of osteoarthritic donors display differential chondrogenic and osteogenic commitment. Eur. Cell Mater. 2014, 27, 298–311. [Google Scholar]
- Dominici, M.; Le Blanc, K.; Mueller, I.; Slaper-Cortenbach, I.; Marini, F.; Krause, D.; Deans, R.; Keating, A.; Prockop, Dj.; Horwitz, E. Minimal criteria for defining multipotent mesenchymal stromal cells. The International Society for Cellular Therapy position statement. Cytotherapy 2006, 8, 315–317. [Google Scholar] [CrossRef] [PubMed]
- Barilani, M.; Banfi, F.; Sironi, S.; Ragni, E.; Guillaumin, S.; Polveraccio, F.; Rosso, L.; Moro, M.; Astori, G.; Pozzobon, M.; et al. Low-affinity Nerve Growth Factor Receptor (CD271) Heterogeneous Expression in Adult and Fetal Mesenchymal Stromal Cells. Sci. Rep. 2018, 8, 9321. [Google Scholar] [CrossRef]
- Ragni, E.; Banfi, F.; Barilani, M.; Cherubini, A.; Parazzi, V.; Larghi, P.; Dolo, V.; Bollati, V.; Lazzari, L. Extracellular Vesicle-Shuttled mRNA in Mesenchymal Stem Cell Communication. Stem Cells 2017, 35, 1093–1105. [Google Scholar] [CrossRef] [PubMed]
- Gouin, K.; Peck, K.; Antes, T.; Johnson, J.L.; Li, C.; Vaturi, S.D.; Middleton, R.; de Couto, G.; Walravens, A.S.; Rodriguez-Borlado, L.; et al. A comprehensive method for identification of suitable reference genes in extracellular vesicles. J. Extracell. Vesicles 2017, 6, 1347019. [Google Scholar] [CrossRef]
- Lv, C.; Yang, T. Effective enrichment of urinary exosomes by polyethylene glycol for RNA detection. Biomed. Res. 2018, 29. [Google Scholar] [CrossRef]
- Li, Y.; Zhang, L.; Liu, F.; Xiang, G.; Jiang, D.; Pu, X. Identification of endogenous controls for analyzing serum exosomal miRNA in patients with hepatitis B or hepatocellular carcinoma. Dis. Markers 2015, 2015, 893594. [Google Scholar] [CrossRef] [PubMed]
- Kennel, P.J.; Saha, A.; Maldonado, D.A.; Givens, R.; Brunjes, D.L.; Castillero, E.; Zhang, X.; Ji, R.; Yahi, A.; George, I.; et al. Serum exosomal protein profiling for the non-invasive detection of cardiac allograft rejection. J. Heart Lung Transplant. 2018, 37, 409–417. [Google Scholar] [CrossRef] [PubMed]
- Ge, Q.; Zhou, Y.; Lu, J.; Bai, Y.; Xie, X.; Lu, Z. miRNA in plasma exosome is stable under different storage conditions. Molecules 2014, 19, 1568–1575. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Xiang, G.M.; Liu, L.L.; Liu, C.; Liu, F.; Jiang, D.N.; Pu, X.Y. Assessment of endogenous reference gene suitability for serum exosomal microRNA expression analysis in liver carcinoma resection studies. Mol. Med. Rep. 2015, 12, 4683–4691. [Google Scholar] [CrossRef] [PubMed]
- Santovito, D.; De Nardis, V.; Marcantonio, P.; Mandolini, C.; Paganelli, C.; Vitale, E.; Buttitta, F.; Bucci, M.; Mezzetti, A.; Consoli, A.; et al. Plasma exosome microRNA profiling unravels a new potential modulator of adiponectin pathway in diabetes: Effect of glycemic control. J. Clin. Endocrinol. Metab. 2014, 99, E1681–E1685. [Google Scholar] [CrossRef]
- Lange, T.; Stracke, S.; Rettig, R.; Lendeckel, U.; Kuhn, J.; Schlüter, R.; Rippe, V.; Endlich, K.; Endlich, N. Identification of miR-16 as an endogenous reference gene for the normalization of urinary exosomal miRNA expression data from CKD patients. PLoS ONE 2017, 12, e0183435. [Google Scholar] [CrossRef]
- Cazzoli, R.; Buttitta, F.; di Nicola, M.; Malatesta, S.; Marchetti, A.; Rom, W.N.; Pass, H.I. microRNAs derived from circulating exosomes as noninvasive biomarkers for screening and diagnosing lung cancer. J. Thorac. Oncol. 2013, 8, 1156–1162. [Google Scholar] [CrossRef] [PubMed]
- Ragni, E.; Perucca Orfei, C.; De Luca, P.; Colombini, A.; Viganò, M.; Lugano, G.; Bollati, V.; de Girolamo, L. Identification of miRNA Reference Genes in Extracellular Vesicles from Adipose Derived Mesenchymal Stem Cells for Studying Osteoarthritis. Int. J. Mol. Sci. 2019, 20, 1108. [Google Scholar] [CrossRef]
- Endisha, H.; Rockel, J.; Jurisica, I.; Kapoor, M. The complex landscape of microRNAs in articular cartilage: Biology, pathology, and therapeutic targets. JCI Insight 2018, 3, 121630. [Google Scholar] [CrossRef]
- Cavalleri, T.; Angelici, L.; Favero, C.; Dioni, L.; Mensi, C.; Bareggi, C.; Palleschi, A.; Rimessi, A.; Consonni, D.; Bordini, L.; et al. Author information Plasmatic extracellular vesicle microRNAs in malignant pleural mesothelioma and asbestos-exposed subjects suggest a 2-miRNA signature as potential biomarker of disease. PLoS ONE 2017, 12, e0176680. [Google Scholar] [CrossRef]
- Vandesompele, J.; de Preter, K.; Pattyn, F.; Poppe, B.; van Roy, N.; De Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3, RESEARCH0034. [Google Scholar] [CrossRef] [PubMed]
- Andersen, C.L.; Jensen, J.L.; Ørntoft, T.F. Normalization of real-time quantitative reverse transcription-PCR data: A model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 2004, 64, 5245–5250. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W.; Tichopad, A.; Prgomet, C.; Neuvians, T.P. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper--Excel-based tool using pair-wise correlations. Biotechnol. Lett. 2004, 26, 509–515. [Google Scholar] [CrossRef]
- Silver, N.; Best, S.; Jiang, J.; Thein, S.L. Selection of housekeeping genes for gene expression studies in human reticulocytes using real-time PCR. BMC Mol. Biol. 2006, 7, 33. [Google Scholar] [CrossRef]
- Metsalu, T.; Vilo, J. ClustVis: A web tool for visualizing clustering of multivariate data using Principal Component Analysis and heatmap. Nucleic Acids Res. 2015, 43, W566–W570. [Google Scholar] [CrossRef]
- Backes, C.; Kehl, T.; Stöckel, D.; Fehlmann, T.; Schneider, L.; Meese, E.; Lenhof, H.P.; Keller, A. miRPathDB: A new dictionary on microRNAs and target pathways. Nucleic Acids Res. 2017, 45, D90–D96. [Google Scholar] [CrossRef] [PubMed]
- Nishimura, D. BioCarta. Biotech Soft. Int. Rep. 2001. [Google Scholar] [CrossRef]
- Mieczkowska, A.; Schumacher, A.; Filipowicz, N.; Wardowska, A.; Zieliński, M.; Madanecki, P.; Nowicka, E.; Langa, P.; Deptuła, M.; Zieliński, J.; et al. Immunophenotyping and transcriptional profiling of in vitro cultured human adipose tissue derived stem cells. Sci. Rep. 2018, 8, 11339. [Google Scholar] [CrossRef] [PubMed]
- Ramos, L.T.; Sánchez-Abarca, L.I.; Muntión, S.; Preciado, S.; Puig, N.; López-Ruano, G.; Hernández-Hernández, Á.; Redondo, A.; Ortega, R.; Rodríguez, C.; et al. MSC surface markers (CD44, CD73, and CD90) can identify human MSC-derived extracellular vesicles by conventional flow cytometry. Cell Commun. Signal. 2016, 14, 2. [Google Scholar] [CrossRef]
- Mestdagh, P.; van Vlierberghe, P.; de Weer, A.; Muth, D.; Westermann, F.; Speleman, F.; Vandesompele, J. A novel and universal method for microRNA RT-qPCR data normalization. Genome Biol. 2009, 10, R64. [Google Scholar] [CrossRef]
- Toh, W.S.; Lai, R.C.; Zhang, B.; Lim, S.K. MSC exosome works through a protein-based mechanism of action. Biochem. Soc. Trans. 2018, 46, 843–853. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Takagawa, J.; Lam, V.C.; Haddad, D.J.; Tobler, D.L.; Mok, P.Y.; Zhang, Y.; Clifford, B.T.; Pinnamaneni, K.; Saini, S.A.; et al. Donor myocardial infarction impairs the therapeutic potential of bone marrow cells by an interleukin-1-mediated inflammatory response. Sci. Transl. Med. 2011, 3, 100ra90. [Google Scholar] [CrossRef] [PubMed]
- Sanz-Nogués, C.; O’Brien, T. MSCs isolated from patients with ischemic vascular disease have normal angiogenic potential. Mol. Ther. 2014, 22, 1888–1889. [Google Scholar] [CrossRef] [PubMed]
- Bocelli-Tyndall, C.; Bracci, L.; Spagnoli, G.; Braccini, A.; Bouchenaki, M.; Ceredig, R.; Pistoia, V.; Martin, I.; Tyndall, A. Bone marrow mesenchymal stromal cells (BM-MSCs) from healthy donors and auto-immune disease patients reduce the proliferation of autologous- and allogeneic-stimulated lymphocytes in vitro. Rheumatology 2007, 46, 403–408. [Google Scholar] [CrossRef]
- Bacigalupo, A.; Valle, M.; Podestà, M.; Pitto, A.; Zocchi, E.; De Flora, A.; Pozzi, S.; Luchetti, S.; Frassoni, F.; van Lint, M.T.; et al. T-cell suppression mediated by mesenchymal stem cells is deficient in patients with severe aplastic anemia. Exp. Hematol. 2005, 33, 819–827. [Google Scholar] [CrossRef] [PubMed]
- Murphy, J.M.; Dixon, K.; Beck, S.; Fabian, D.; Feldman, A.; Barry, F. Reduced chondrogenic and adipogenic activity of mesenchymal stem cells from patients with advanced osteoarthritis. Arthritis Rheum. 2002, 46, 704–713. [Google Scholar] [CrossRef]
- Del Papa, N.; Quirici, N.; Soligo, D.; Scavullo, C.; Cortiana, M.; Borsotti, C.; Maglione, W.; Comina, D.P.; Vitali, C.; Fraticelli, P.; et al. Bone marrow endothelial progenitors are defective in systemic sclerosis. Arthritis Rheum. 2006, 54, 2605–2615. [Google Scholar] [CrossRef]
- Kastrinaki, M.C.; Sidiropoulos, P.; Roche, S.; Ringe, J.; Lehmann, S.; Kritikos, H.; Vlahava, V.M.; Delorme, B.; Eliopoulos, G.D.; Jorgensen, C.; et al. Functional, molecular and proteomic characterisation of bone marrow mesenchymal stem cells in rheumatoid arthritis. Ann. Rheum. Dis. 2008, 67, 741–749. [Google Scholar] [CrossRef]
- Papadaki, H.A.; Tsagournisakis, M.; Mastorodemos, V.; Pontikoglou, C.; Damianaki, A.; Pyrovolaki, K.; Stamatopoulos, K.; Fassas, A.; Plaitakis, A.; Eliopoulos, G.D. Normal bone marrow hematopoietic stem cell reserves and normal stromal cell function support the use of autologous stem cell transplantation in patients with multiple sclerosis. Bone Marrow Transplant. 2005, 36, 1053–1063. [Google Scholar] [CrossRef]
- Mallam, E.; Kemp, K.; Wilkins, A.; Rice, C.; Scolding, N. Characterization of in vitro expanded bone marrow-derived mesenchymal stem cells from patients with multiple sclerosis. Mult. Scler. 2010, 16, 909–918. [Google Scholar] [CrossRef]
- Mazzanti, B.; Aldinucci, A.; Biagioli, T.; Barilaro, A.; Urbani, S.; Dal Pozzo, S.; Amato, M.P.; Siracusa, G.; Crescioli, C.; Manuelli, C.; et al. Differences in mesenchymal stem cell cytokine profiles between MS patients and healthy donors: Implication for assessment of disease activity and treatment. J. Neuroimmunol. 2008, 199, 142–150. [Google Scholar] [CrossRef] [PubMed]
- Srijaya, T.C.; Ramasamy, T.S.; Kasim, N.H. Advancing stem cell therapy from bench to bedside: Lessons from drug therapies. J. Transl. Med. 2014, 12, 243. [Google Scholar] [CrossRef] [PubMed]
- Trounson, A.; McDonald, C. Stem cell therapies in clinical trials: Progress and challenges. Cell Stem Cell 2015, 17, 11–22. [Google Scholar] [CrossRef]
- Galderisi, U.; Calarco, A.; Melone, M.; Peluso, G. Is it possible to improve the success rate of cellular therapies based on mesenchymal stem cells? J. Stem Cells Res. Rev. Rep. 2014, 1, 1–3. [Google Scholar]
- Zhukareva, V.; Obrocka, M.; Houle, J.D.; Fischer, I.; Neuhuber, B. Secretion profile of human bone marrow stromal cells: Donor variability and response to inflammatory stimuli. Cytokine 2010, 50, 317–321. [Google Scholar] [CrossRef]
- Gray, A.; Schloss, R.S.; Yarmush, M. Donor variability among anti-inflammatory pre-activated mesenchymal stromal cells. Technology 2016, 4, 201–215. [Google Scholar] [CrossRef]
- Phinney, D.G.; Pittenger, M.F. Concise Review: MSC-Derived Exosomes for Cell-Free Therapy. Stem Cells 2017, 35, 851–858. [Google Scholar] [CrossRef]
- Ferguson, S.W.; Wang, J.; Lee, C.J.; Liu, M.; Neelamegham, S.; Canty, J.M.; Nguyen, J. The microRNA regulatory landscape of MSC-derived exosomes: A systems view. Sci Rep. 2018, 8, 1419. [Google Scholar] [CrossRef] [PubMed]
- Cheng, L.; Zhang, K.; Wu, S.; Cui, M.; Xu, T. Focus on Mesenchymal Stem Cell-Derived Exosomes: Opportunities and Challenges in Cell-Free Therapy. Stem Cells Int. 2017, 2017, 6305295. [Google Scholar] [CrossRef] [PubMed]
- Toh, W.S.; Lai, R.C.; Hui, J.H.; Lim, S.K. MSC exosome as a cell-free MSC therapy for cartilage regeneration: Implications for osteoarthritis treatment. Semin. Cell Dev. Biol. 2017, 67, 56–64. [Google Scholar] [CrossRef] [PubMed]
- Schwarzenbach, H.; da Silva, A.M.; Calin, G.; Pantel, K. Data Normalization Strategies for MicroRNA Quantification. Clin. Chem. 2015, 61, 1333–1342. [Google Scholar] [CrossRef] [PubMed]
- Meyer, S.U.; Pfaffl, M.W.; Ulbrich, S.E. Normalization strategies for microRNA profiling experiments: A ‘normal’ way to a hidden layer of complexity? Biotechnol. Lett. 2010, 32, 1777–1788. [Google Scholar] [CrossRef]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef]
- Gray, W.D.; French, K.M.; Ghosh-Choudhary, S.; Maxwell, J.T.; Brown, M.E.; Platt, M.O.; Searles, C.D.; Davis, M.E. Identification of therapeutic covariant microRNA clusters in hypoxia-treated cardiac progenitor cell exosomes using systems biology. Circ. Res. 2015, 116, 255–263. [Google Scholar] [CrossRef]
- Hayashi, T.; Lombaert, I.M.; Hauser, B.R.; Patel, V.N.; Hoffman, M.P. Exosomal MicroRNA Transport from Salivary Mesenchyme Regulates Epithelial Progenitor Expansion during Organogenesis. Dev. Cell 2017, 40, 95–103. [Google Scholar] [CrossRef]
- Lee, Y.; Ahn, C.; Han, J.; Choi, H.; Kim, J.; Yim, J.; Lee, J.; Provost, P.; Rådmark, O.; Kim, S.; et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 2003, 425, 415–419. [Google Scholar] [CrossRef]
- Yuan, Y.; Zhang, G.Q.; Chai, W.; Ni, M.; Xu, C.; Chen, J.Y. Silencing of microRNA-138-5p promotes IL-1β-induced cartilage degradation in human chondrocytes by targeting FOXC1: miR-138 promotes cartilage degradation. Bone Joint Res. 2016, 5, 523–530. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, M.; Hata, K.; Takashima, R.; Ono, K.; Nakamura, E.; Takahata, Y.; Murakami, T.; Iseki, S.; Takano-Yamamoto, T.; Nishimura, R.; et al. The transcription factor Foxc1 is necessary for Ihh-Gli2-regulated endochondral ossification. Nat. Commun. 2015, 6, 6653. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, A.; Rampersaud, Y.R.; Sharma, A.; Lewis, S.J.; Wu, B.; Datta, P.; Sundararajan, K.; Endisha, H.; Rossomacha, E.; Rockel, J.S.; et al. Identification of microRNA-181a-5p and microRNA-4454 as mediators of facet cartilage degeneration. JCI Insight 2016, 1, e86820. [Google Scholar] [CrossRef]
- Wang, H.; Zhang, H.; Sun, Q.; Wang, Y.; Yang, J.; Yang, J.; Zhang, T.; Luo, S.; Wang, L.; Jiang, Y.; et al. Intra-articular Delivery of Antago-miR-483-5p Inhibits Osteoarthritis by Modulating Matrilin 3 and Tissue Inhibitor of Metalloproteinase 2. Mol. Ther. 2017, 25, 715–727. [Google Scholar] [CrossRef] [PubMed]
- Philipot, D.; Guérit, D.; Platano, D.; Chuchana, P.; Olivotto, E.; Espinoza, F.; Dorandeu, A.; Pers, Y.M.; Piette, J.; Borzi, R.M.; et al. p16INK4a and its regulator miR-24 link senescence and chondrocyte terminal differentiation-associated matrix remodeling in osteoarthritis. Arthritis Res. Ther. 2014, 16, R58. [Google Scholar] [CrossRef]
- Ge, F.X.; Li, H.; Yin, X. Upregulation of microRNA-125b-5p is involved in the pathogenesis of osteoarthritis by downregulating SYVN1. Oncol. Rep. 2017, 37, 2490–2496. [Google Scholar] [CrossRef][Green Version]
- Song, J.; Jin, E.H.; Kim, D.; Kim, K.Y.; Chun, C.H.; Jin, E.J. MicroRNA-222 regulates MMP-13 via targeting HDAC-4 during osteoarthritis pathogenesis. BBA Clin. 2014, 3, 79–89. [Google Scholar] [CrossRef]
- Zhang, Y.; Jia, J.; Yang, S.; Liu, X.; Ye, S.; Tian, H. MicroRNA-21 controls the development of osteoarthritis by targeting GDF-5 in chondrocytes. Exp. Mol. Med. 2014, 46, e79. [Google Scholar] [CrossRef] [PubMed]
- Chang, Z.K.; Meng, F.G.; Zhang, Z.Q.; Mao, G.P.; Huang, Z.Y.; Liao, W.M.; He, A.S. MicroRNA-193b-3p regulates matrix metalloproteinase 19 expression in interleukin-1β-induced human chondrocytes. J. Cell Biochem. 2018, 119, 4775–4782. [Google Scholar] [CrossRef]
| Gene name | GeNorm M-Value | NormFinder Stability | BestKeeper SD ± CP | ΔCt Mean | Mean Crt | GeoMean |
|---|---|---|---|---|---|---|
| miR-26a-5p | 0.228 (1) | 0.079 (2) | 0.626 (4) | 0.536 (1) | 13.81 (2) | 1.7 |
| miR-16-5p | 0.228 (1) | 0.060 (1) | 0.683 (5) | 0.562 (2) | 15.09 (5) | 2.2 |
| 221-3p | 0.313 (3) | 0.282 (4) | 0.571 (3) | 0.648 (3) | 13.94 (3) | 3.2 |
| U6 snRNA | 0.414 (4) | 0.423 (7) | 0.401 (2) | 0.757 (7) | 12.53 (1) | 3.3 |
| let-7a-5p | 0.497 (6) | 0.240 (3) | 0.773 (6) | 0.663 (4) | 14.61 (4) | 4.4 |
| miR-423-5p | 0.439 (5) | 0.511 (8) | 0.368 (1) | 0.834 (8) | 19.15 (9) | 4.9 |
| miR-23a-3p | 0.548 (7) | 0.336 (6) | 0.926 (8) | 0.716 (6) | 17.72 (6) | 6.6 |
| miR-101-3p | 0.583 (8) | 0.323 (5) | 0.926 (7) | 0.702 (5) | 22.33 (10) | 6.7 |
| miR-103a-3p | 0.674 (9) | 0.622 (10) | 1.332 (10) | 0.997 (9) | 17.87 (7) | 8.9 |
| miR-425-5p | 0.741 (10) | 0.620 (9) | 1.177 (9) | 1.026 (10) | 17.88 (8) | 9.2 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ragni, E.; De Luca, P.; Perucca Orfei, C.; Colombini, A.; Viganò, M.; Lugano, G.; Bollati, V.; de Girolamo, L. Insights into Inflammatory Priming of Adipose-Derived Mesenchymal Stem Cells: Validation of Extracellular Vesicles-Embedded miRNA Reference Genes as A Crucial Step for Donor Selection. Cells 2019, 8, 369. https://doi.org/10.3390/cells8040369
Ragni E, De Luca P, Perucca Orfei C, Colombini A, Viganò M, Lugano G, Bollati V, de Girolamo L. Insights into Inflammatory Priming of Adipose-Derived Mesenchymal Stem Cells: Validation of Extracellular Vesicles-Embedded miRNA Reference Genes as A Crucial Step for Donor Selection. Cells. 2019; 8(4):369. https://doi.org/10.3390/cells8040369
Chicago/Turabian StyleRagni, Enrico, Paola De Luca, Carlotta Perucca Orfei, Alessandra Colombini, Marco Viganò, Gaia Lugano, Valentina Bollati, and Laura de Girolamo. 2019. "Insights into Inflammatory Priming of Adipose-Derived Mesenchymal Stem Cells: Validation of Extracellular Vesicles-Embedded miRNA Reference Genes as A Crucial Step for Donor Selection" Cells 8, no. 4: 369. https://doi.org/10.3390/cells8040369
APA StyleRagni, E., De Luca, P., Perucca Orfei, C., Colombini, A., Viganò, M., Lugano, G., Bollati, V., & de Girolamo, L. (2019). Insights into Inflammatory Priming of Adipose-Derived Mesenchymal Stem Cells: Validation of Extracellular Vesicles-Embedded miRNA Reference Genes as A Crucial Step for Donor Selection. Cells, 8(4), 369. https://doi.org/10.3390/cells8040369










