Dermatopontin in Skeletal Muscle Extracellular Matrix Regulates Myogenesis
Abstract
:1. Introduction
2. Materials and Methods
2.1. Cell Culture
2.2. Gene Knockdown
2.3. MTT Assay
2.4. RNA Isolation and qPCR
2.5. Scratch Experiment
2.6. Western Blot Analysis
2.7. Immunocytochemistry
2.8. Immunohistochemistry
2.9. Fusion Index
2.10. Plate Coating with ECM Proteins
2.11. Animal Experiment
2.12. Statistical Analysis
2.13. 3D Model Generation of DPT
2.14. Protein-Protein Interaction
3. Results
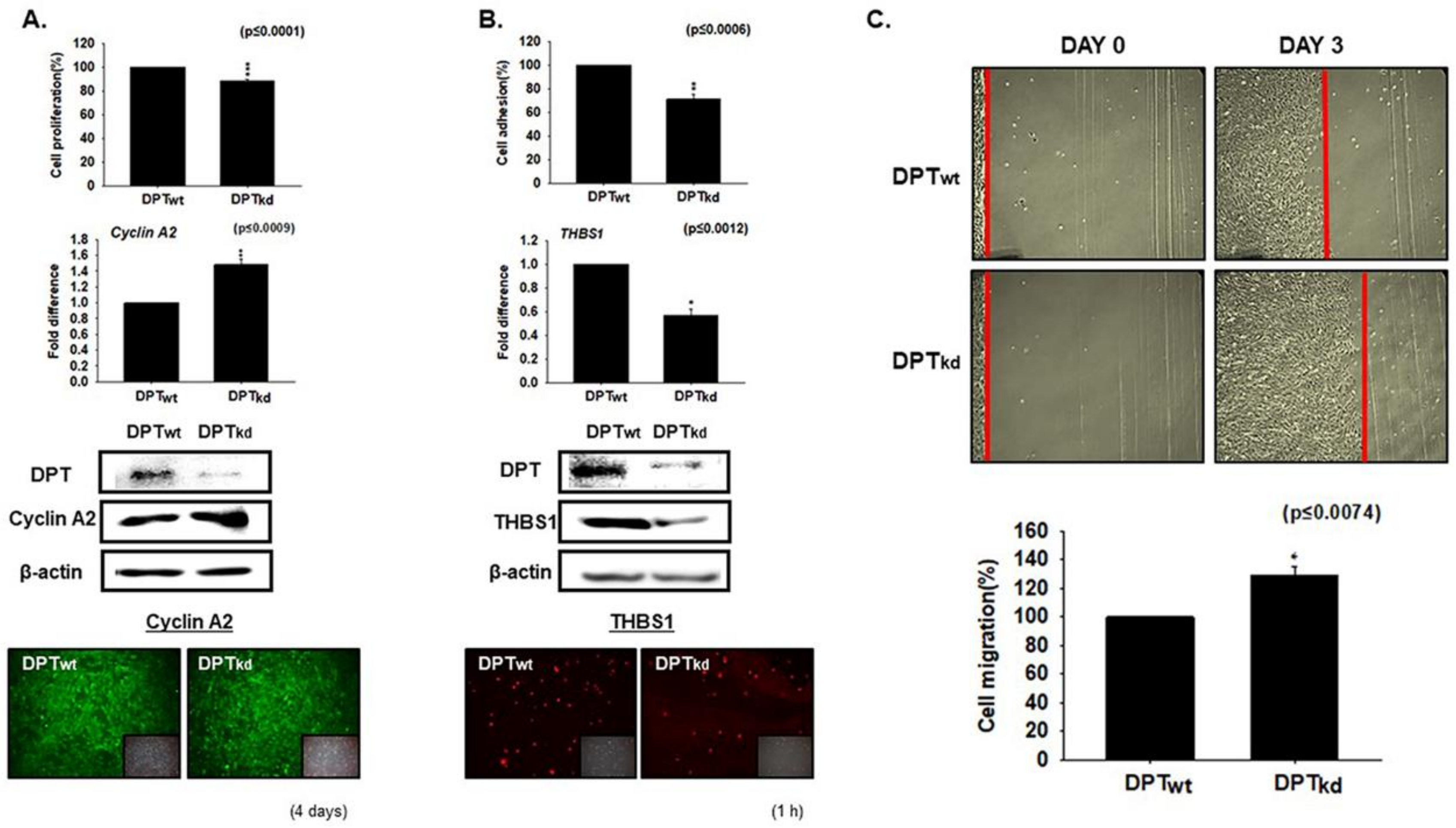
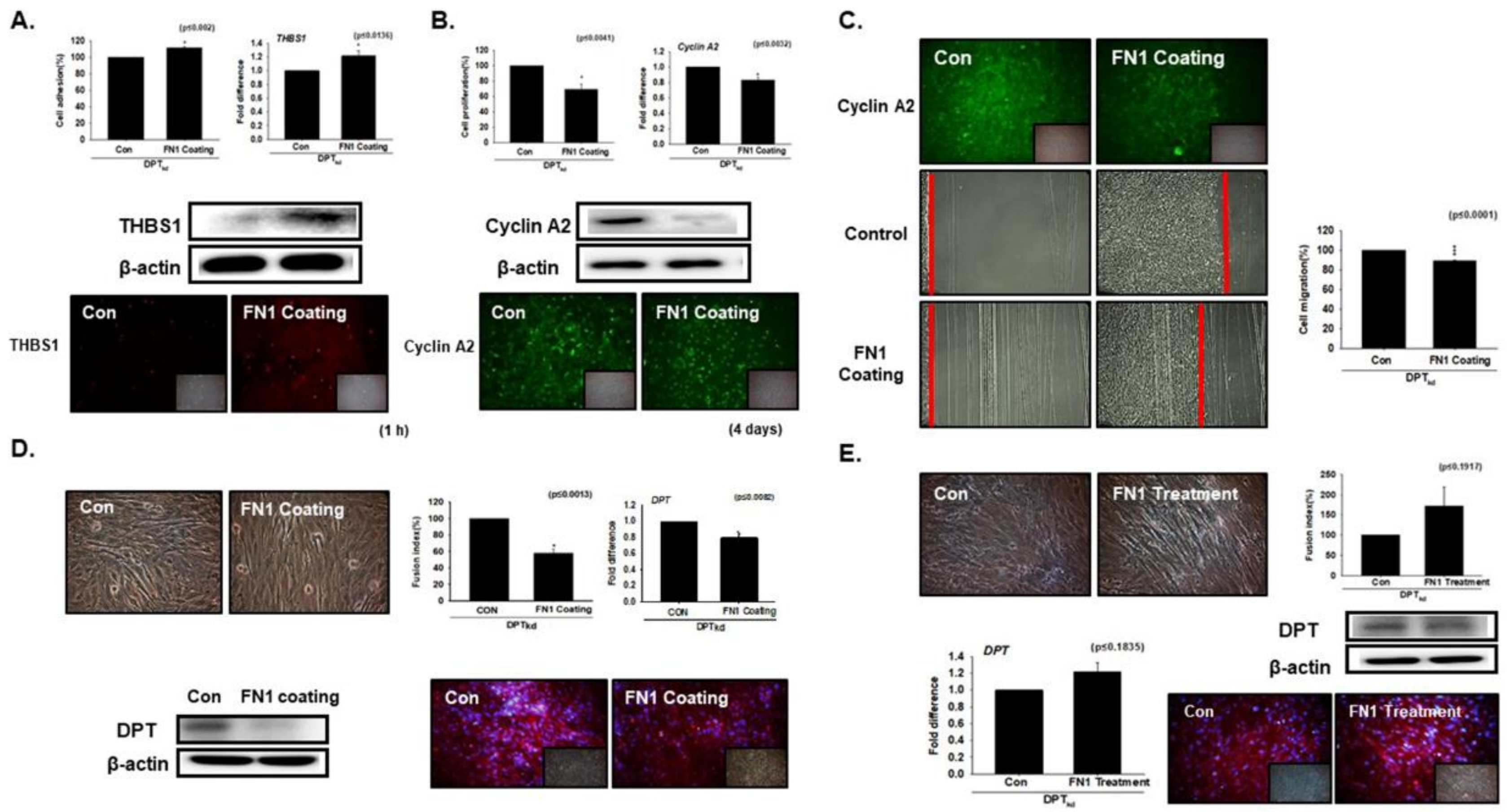
3.1. DPT Enhances the Cell Adhesion and Reduces Cell Proliferation
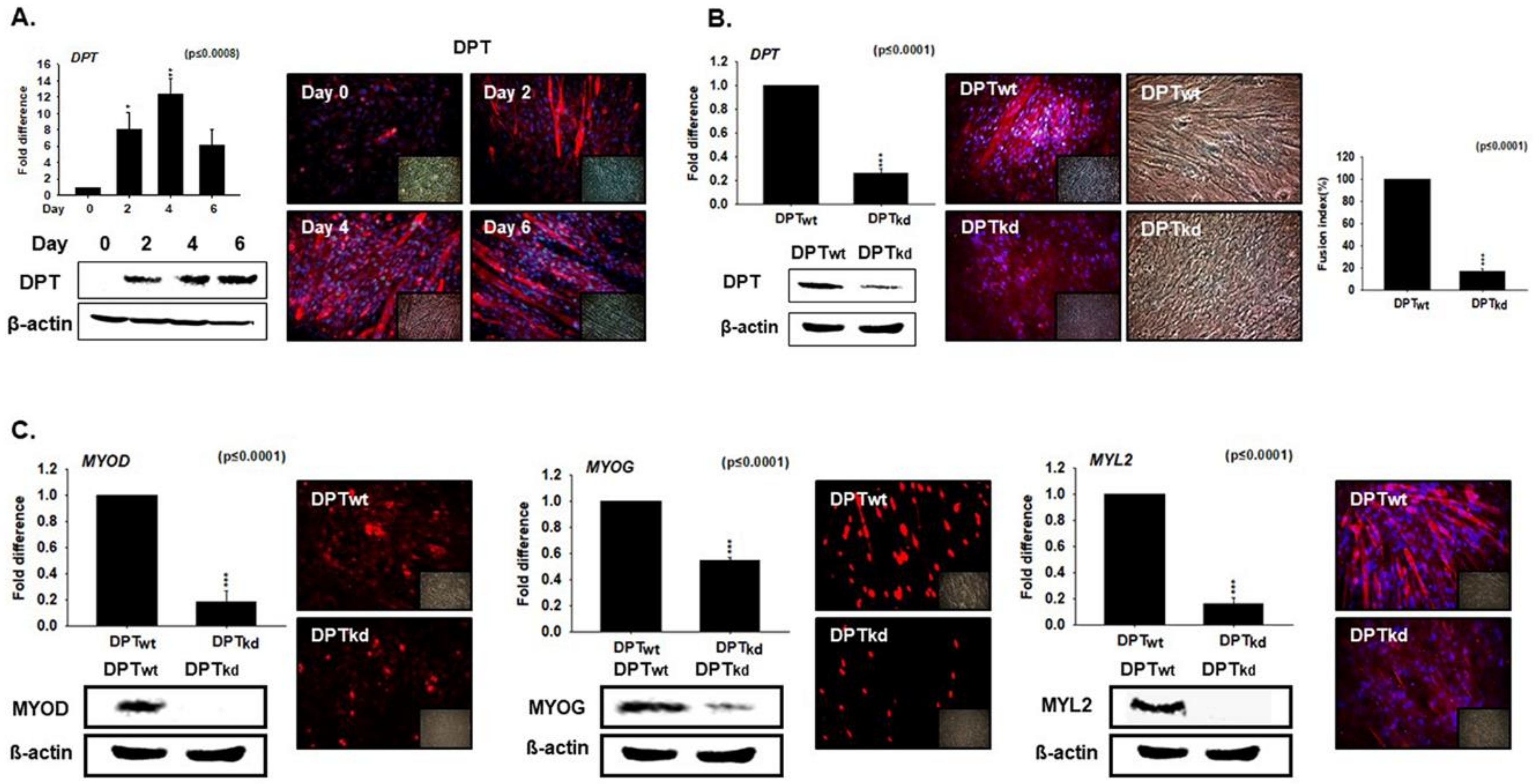
3.2. DPT Expression during Myoblast Differentiation
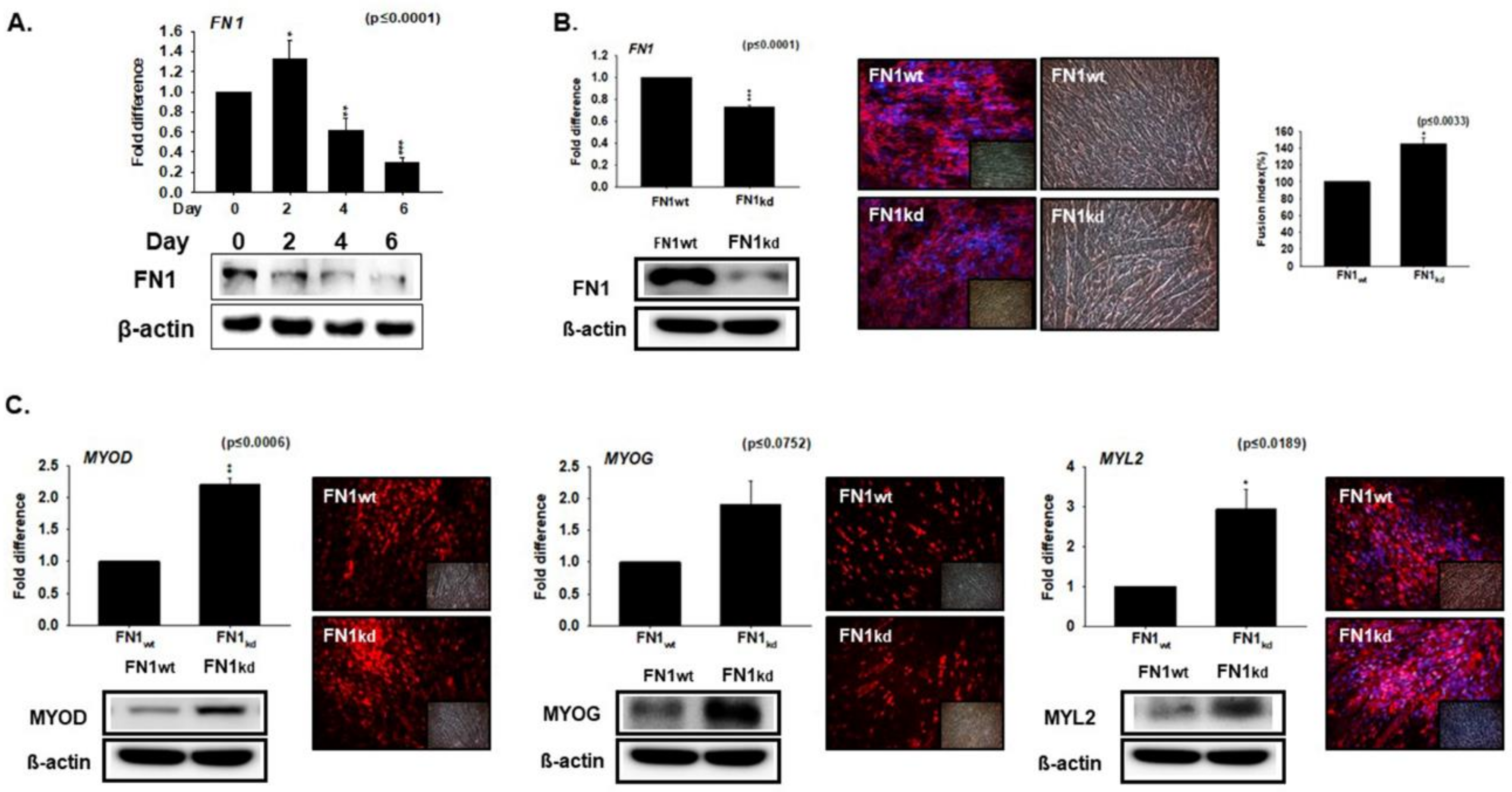
3.3. Knockdown Effect of FN during Myoblast Differentiation
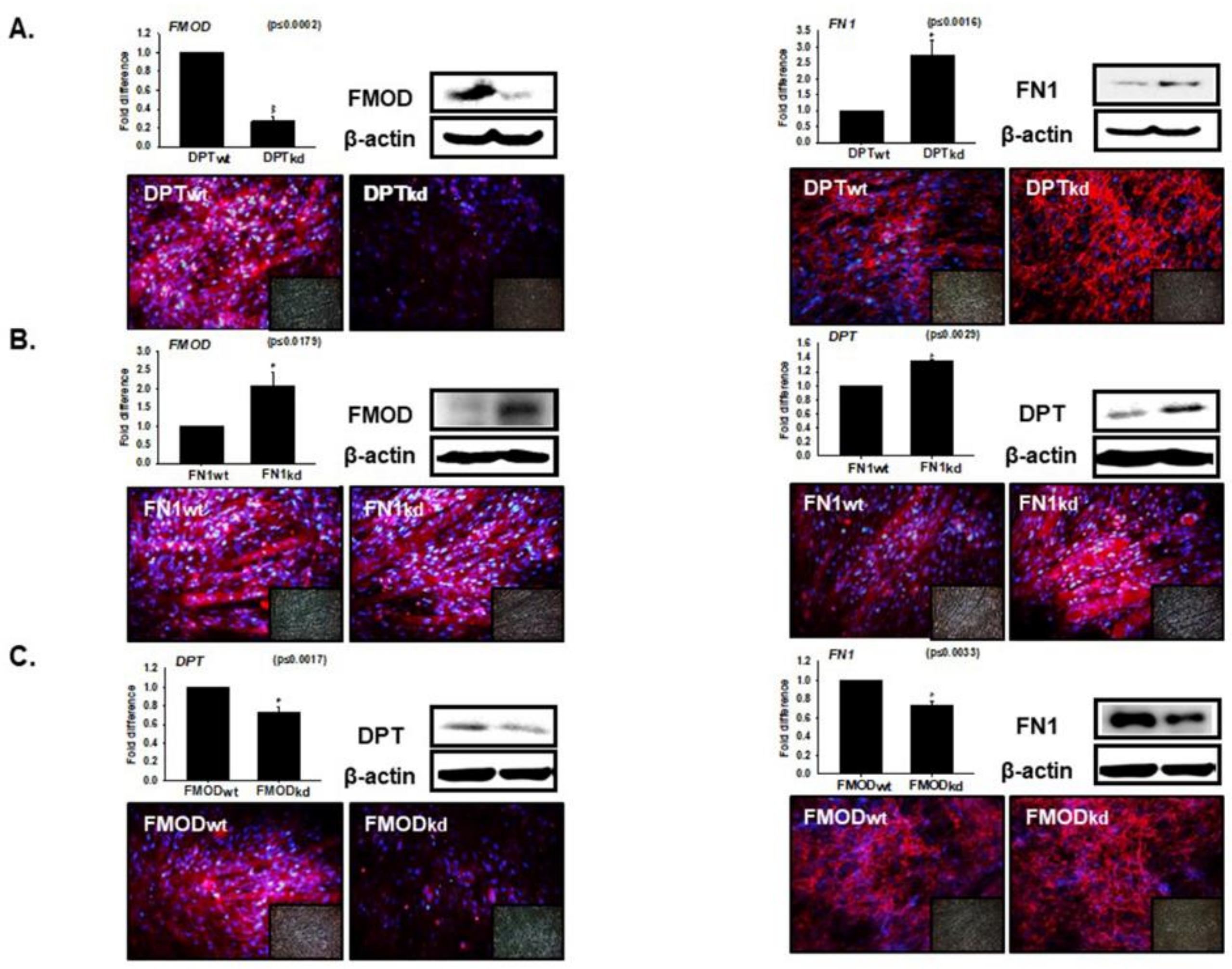
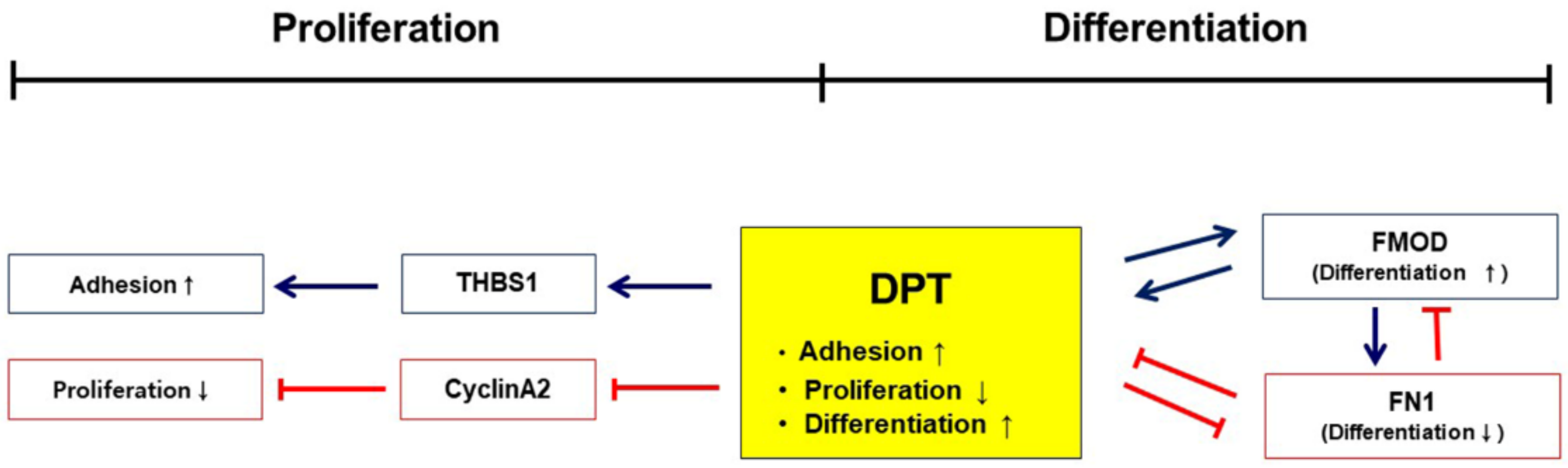
3.4. Interaction of DPT with FN and FMOD during Differentiation
3.5. Compensation Effect of Fibronectin with DPT
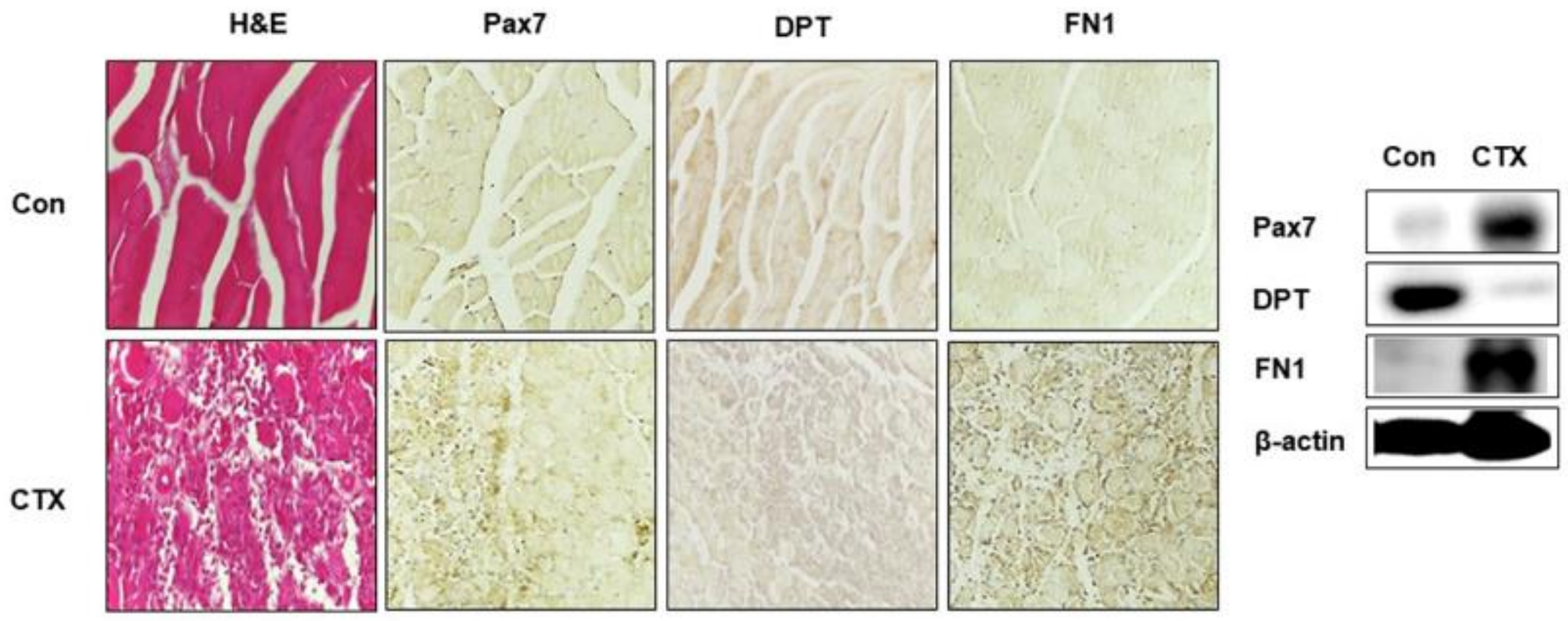
3.6. DPT and FN1 in Muscle Regeneration
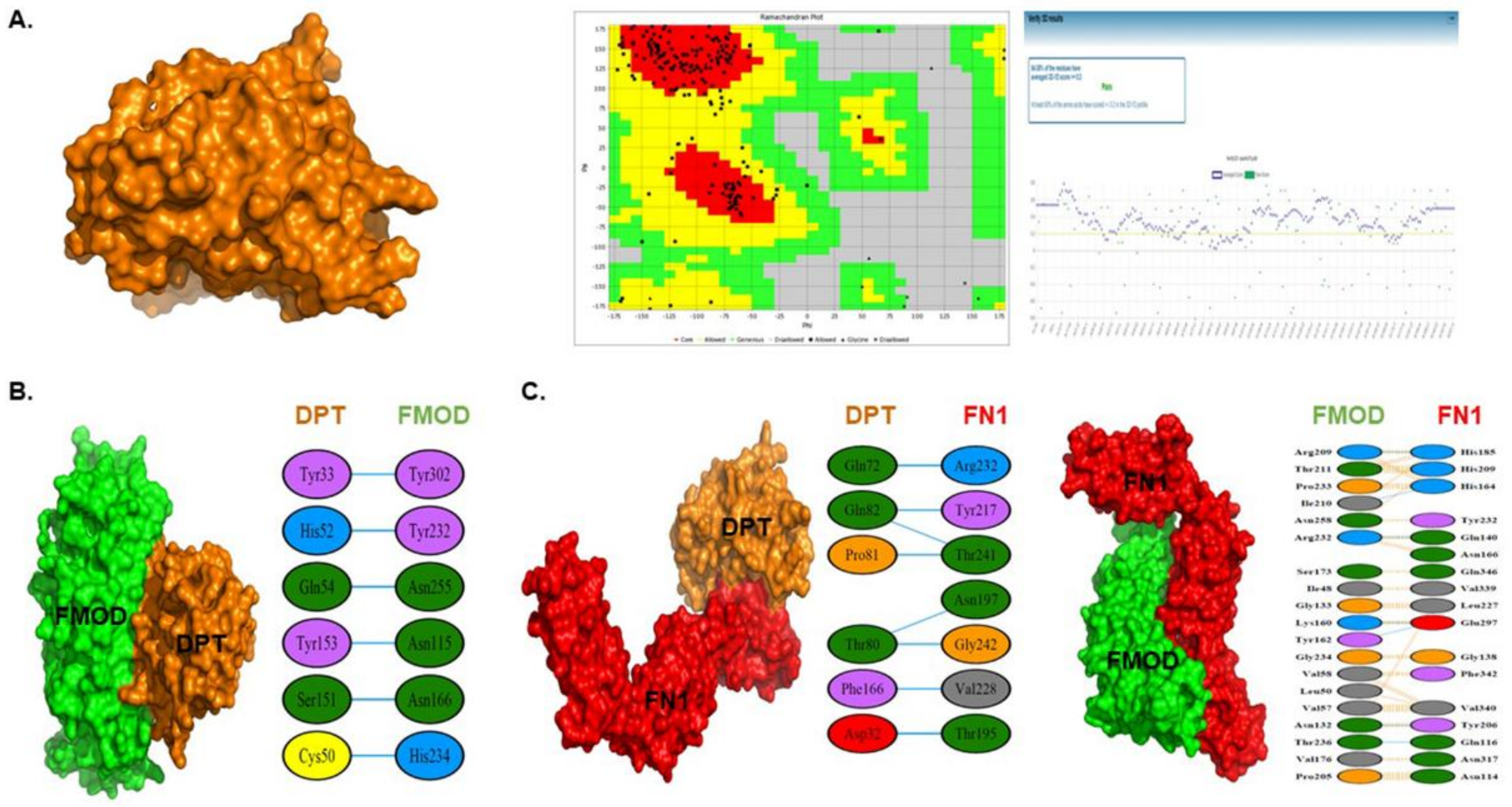
3.7. 3D Protein Modeling of DPT
3.8. Protein-Protein Interaction
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Frontera, W.R.; Ochala, J. Skeletal muscle: A brief review of structure and function. Calcif. Tissue Int. 2015, 96, 183–195. [Google Scholar] [CrossRef] [PubMed]
- Campbell, K.P.; Stull, J.T. Skeletal muscle basement membrane-sarcolemma-cytoskeleton interaction minireview series. J. Biol. Chem. 2003, 278, 12599–12600. [Google Scholar] [CrossRef]
- Chal, J.; Pourquie, O. Making muscle: Skeletal myogenesis in vivo and in vitro. Development 2017, 144, 2104–2122. [Google Scholar] [CrossRef] [PubMed]
- Pallafacchina, G.; Francois, S.; Regnault, B.; Czarny, B.; Dive, V.; Cumano, A.; Montarras, D.; Buckingham, M. An adult tissue-specific stem cell in its niche: A gene profiling analysis of in vivo quiescent and activated muscle satellite cells. Stem Cell Res. 2010, 4, 77–91. [Google Scholar] [CrossRef]
- Karalaki, M.; Fili, S.; Philippou, A.; Koutsilieris, M. Muscle regeneration: Cellular and molecular events. In Vivo 2009, 23, 779–796. [Google Scholar] [PubMed]
- Collins, C.A.; Olsen, I.; Zammit, P.S.; Heslop, L.; Petrie, A.; Partridge, T.A.; Morgan, J.E. Stem cell function, self-renewal, and behavioral heterogeneity of cells from the adult muscle satellite cell niche. Cell 2005, 122, 289–301. [Google Scholar] [CrossRef]
- Zammit, P.; Beauchamp, J. The skeletal muscle satellite cell: Stem cell or son of stem cell? Differentiation 2001, 68, 193–204. [Google Scholar] [CrossRef]
- Day, K.; Paterson, B.; Yablonka-Reuveni, Z. A distinct profile of myogenic regulatory factor detection within Pax7+ cells at S phase supports a unique role of Myf5 during posthatch chicken myogenesis. Dev. Dyn. 2009, 238, 1001–1009. [Google Scholar] [CrossRef] [PubMed]
- Baig, M.H.; Rashid, I.; Srivastava, P.; Ahmad, K.; Jan, A.T.; Rabbani, G.; Choi, D.; Barreto, G.E.; Ashraf, G.M.; Lee, E.J.; et al. NeuroMuscleDB: A Database of Genes Associated with Muscle Development, Neuromuscular Diseases, Ageing, and Neurodegeneration. Mol. Neurobiol. 2019. [Google Scholar] [CrossRef]
- Ahmad, K.; Lee, E.J.; Moon, J.S.; Park, S.Y.; Choi, I. Multifaceted Interweaving Between Extracellular Matrix, Insulin Resistance, and Skeletal Muscle. Cells 2018, 7, 148. [Google Scholar] [CrossRef] [PubMed]
- Thorsteinsdottir, S.; Deries, M.; Cachaco, A.S.; Bajanca, F. The extracellular matrix dimension of skeletal muscle development. Dev. Biol. 2011, 354, 191–207. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, S.; Jan, A.T.; Baig, M.H.; Lee, E.J.; Choi, I. Matrix gla protein: An extracellular matrix protein regulates myostatin expression in the muscle developmental program. Life Sci. 2017, 172, 55–63. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.J.; Nam, J.H.; Choi, I. Fibromodulin modulates myoblast differentiation by controlling calcium channel. Biochem. Biophys. Res. Commun. 2018, 503, 580–585. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.J.; Jan, A.T.; Baig, M.H.; Ashraf, J.M.; Nahm, S.S.; Kim, Y.W.; Park, S.Y.; Choi, I. Fibromodulin: A master regulator of myostatin controlling progression of satellite cells through a myogenic program. FASEB J. 2016, 30, 2708–2719. [Google Scholar] [CrossRef]
- Lee, E.J.; Jan, A.T.; Baig, M.H.; Ahmad, K.; Malik, A.; Rabbani, G.; Kim, T.; Lee, I.K.; Lee, Y.H.; Park, S.Y.; et al. Fibromodulin and regulation of the intricate balance between myoblast differentiation to myocytes or adipocyte-like cells. FASEB J. 2018, 32, 768–781. [Google Scholar] [CrossRef]
- Okamoto, O.; Fujiwara, S. Dermatopontin, a novel player in the biology of the extracellular matrix. Connect. Tissue Res. 2006, 47, 177–189. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Meng, L.; Shi, Q.; Liu, S.; Cui, C.; Hu, S.; Wei, Y. Dermatopontin promotes adhesion, spreading and migration of cardiac fibroblasts in vitro. Matrix Biol. 2013, 32, 23–31. [Google Scholar] [CrossRef]
- Okamoto, O.; Hozumi, K.; Katagiri, F.; Takahashi, N.; Sumiyoshi, H.; Matsuo, N.; Yoshioka, H.; Nomizu, M.; Fujiwara, S. Dermatopontin promotes epidermal keratinocyte adhesion via alpha3beta1 integrin and a proteoglycan receptor. Biochemistry 2010, 49, 147–155. [Google Scholar] [CrossRef]
- Okamoto, O.; Fujiwara, S.; Abe, M.; Sato, Y. Dermatopontin interacts with transforming growth factor beta and enhances its biological activity. Biochem. J. 1999, 337, 537–541. [Google Scholar] [CrossRef]
- Kato, A.; Okamoto, O.; Ishikawa, K.; Sumiyoshi, H.; Matsuo, N.; Yoshioka, H.; Nomizu, M.; Shimada, T.; Fujiwara, S. Dermatopontin interacts with fibronectin, promotes fibronectin fibril formation, and enhances cell adhesion. J. Biol. Chem. 2011, 286, 14861–14869. [Google Scholar] [CrossRef]
- Lee, E.J.; Pokharel, S.; Jan, A.T.; Huh, S.; Galope, R.; Lim, J.H.; Lee, D.M.; Choi, S.W.; Nahm, S.S.; Kim, Y.W.; et al. Transthyretin: A Transporter Protein Essential for Proliferation of Myoblast in the Myogenic Program. Int. J. Mol. Sci. 2017, 18, 115. [Google Scholar] [CrossRef]
- Kim, D.S.; Cha, H.N.; Jo, H.J.; Song, I.H.; Baek, S.H.; Dan, J.M.; Kim, Y.W.; Kim, J.Y.; Lee, I.K.; Seo, J.S.; et al. TLR2 deficiency attenuates skeletal muscle atrophy in mice. Biochem. Biophys. Res. Commun. 2015, 459, 534–540. [Google Scholar] [CrossRef] [PubMed]
- Tan, K.; Lawler, J. The interaction of Thrombospondins with extracellular matrix proteins. J. Cell Commun. Signal. 2009, 3, 177–187. [Google Scholar] [CrossRef] [PubMed]
- Kalamajski, S.; Bihan, D.; Bonna, A.; Rubin, K.; Farndale, R.W. Fibromodulin Interacts with Collagen Cross-linking Sites and Activates Lysyl Oxidase. J. Biol. Chem. 2016, 291, 7951–7960. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Z.; Zhang, X.; Dang, C.; Beanes, S.; Chang, G.X.; Chen, Y.; Li, C.S.; Lee, K.S.; Ting, K.; Soo, C. Fibromodulin Is Essential for Fetal-Type Scarless Cutaneous Wound Healing. Am. J. Pathol. 2016, 186, 2824–2832. [Google Scholar] [CrossRef]
- Kalamajski, S.; Oldberg, A. The role of small leucine-rich proteoglycans in collagen fibrillogenesis. Matrix Biol. 2010, 29, 248–253. [Google Scholar] [CrossRef] [PubMed]
- Lu, B.; Mahmud, H.; Maass, A.H.; Yu, B.; van Gilst, W.H.; de Boer, R.A.; Sillje, H.H. The Plk1 inhibitor BI 2536 temporarily arrests primary cardiac fibroblasts in mitosis and generates aneuploidy in vitro. PLoS ONE 2010, 5, e12963. [Google Scholar] [CrossRef] [PubMed]
- Arslan, A.A.; Gold, L.I.; Mittal, K.; Suen, T.C.; Belitskaya-Levy, I.; Tang, M.S.; Toniolo, P. Gene expression studies provide clues to the pathogenesis of uterine leiomyoma: New evidence and a systematic review. Hum. Reprod. 2005, 20, 852–863. [Google Scholar] [CrossRef]
- MacBeath, J.R.; Shackleton, D.R.; Hulmes, D.J. Tyrosine-rich acidic matrix protein (TRAMP) accelerates collagen fibril formation in vitro. J. Biol. Chem. 1993, 268, 19826–19832. [Google Scholar]
- Catherino, W.H.; Leppert, P.C.; Stenmark, M.H.; Payson, M.; Potlog-Nahari, C.; Nieman, L.K.; Segars, J.H. Reduced dermatopontin expression is a molecular link between uterine leiomyomas and keloids. Genes Chromosomes Cancer 2004, 40, 204–217. [Google Scholar] [CrossRef]
- Okamoto, O.; Suzuki, Y.; Kimura, S.; Shinkai, H. Extracellular matrix 22-kDa protein interacts with decorin core protein and is expressed in cutaneous fibrosis. J. Biochem. 1996, 119, 106–114. [Google Scholar] [CrossRef] [PubMed]
- Sidgwick, G.; Bayat, A. Extracellular matrix molecules implicated in hypertrophic and keloid scarring. J. Eur. Acad. Dermatol. Venereol. 2012, 26, 141–152. [Google Scholar] [CrossRef]
- Guo, Y.; Li, H.; Guan, H.; Ke, W.; Liang, W.; Xiao, H.; Li, Y. Dermatopontin inhibits papillary thyroid cancer cell proliferation through MYC repression. Mol. Cell Endocrinol. 2018. [Google Scholar] [CrossRef]
- Resovi, A.; Pinessi, D.; Chiorino, G.; Taraboletti, G. Current understanding of the thrombospondin-1 interactome. Matrix Biol. 2014, 37, 83–91. [Google Scholar] [CrossRef] [PubMed]
- Gardner, J.M.; Fambrough, D.M. Fibronectin expression during myogenesis. J. Cell Biol. 1983, 96, 474–485. [Google Scholar] [CrossRef] [PubMed]
- Grounds, M.D. Complexity of extracellular matrix and skeletal muscle regeneration. In Skeletal Muscle Repair and Regeneration; Springer: Dordrecht, The Netherlands, 2008; pp. 269–302. [Google Scholar]
- Bentzinger, C.F.; Wang, Y.X.; von Maltzahn, J.; Soleimani, V.D.; Yin, H.; Rudnicki, M.A. Fibronectin regulates Wnt7a signaling and satellite cell expansion. Cell Stem Cell 2013, 12, 75–87. [Google Scholar] [CrossRef]
- Chaturvedi, V.; Dye, D.E.; Kinnear, B.F.; van Kuppevelt, T.H.; Grounds, M.D.; Coombe, D.R. Interactions between skeletal muscle myoblasts and their extracellular matrix revealed by a serum free culture system. PLoS ONE 2015, 10, e0127675. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, T.; Ahmad, K.; Shaikh, S.; Jan, A.T.; Seo, M.-G.; Lee, E.J.; Choi, I. Dermatopontin in Skeletal Muscle Extracellular Matrix Regulates Myogenesis. Cells 2019, 8, 332. https://doi.org/10.3390/cells8040332
Kim T, Ahmad K, Shaikh S, Jan AT, Seo M-G, Lee EJ, Choi I. Dermatopontin in Skeletal Muscle Extracellular Matrix Regulates Myogenesis. Cells. 2019; 8(4):332. https://doi.org/10.3390/cells8040332
Chicago/Turabian StyleKim, Taeyeon, Khurshid Ahmad, Sibhghatulla Shaikh, Arif Tasleem Jan, Myung-Gi Seo, Eun Ju Lee, and Inho Choi. 2019. "Dermatopontin in Skeletal Muscle Extracellular Matrix Regulates Myogenesis" Cells 8, no. 4: 332. https://doi.org/10.3390/cells8040332
APA StyleKim, T., Ahmad, K., Shaikh, S., Jan, A. T., Seo, M.-G., Lee, E. J., & Choi, I. (2019). Dermatopontin in Skeletal Muscle Extracellular Matrix Regulates Myogenesis. Cells, 8(4), 332. https://doi.org/10.3390/cells8040332





