Enhancing the Nucleoside Analog Response with Translational Therapeutic Approaches to Overcome Resistance
Highlights
- The anti-tumor efficacy of nucleoside analogs is dependent on transport, enzymatic activation, nucleotide pools and metabolic responses, which are often aberrant in cancer cells, rendering these chemotherapeutic drugs ineffective.
- Small molecule inhibitors are being tested/developed to target or downregulate specific mechanisms responsible for resistance to nucleoside analogs. The combination of small molecule inhibitors with nucleoside analogs has enhanced the efficacy of nucleoside analogs and improved patient outcomes.
Abstract
1. Introduction
2. Reduced Cellular Uptake in Nucleoside Analog Resistant Cancer Cells
2.1. Cellular Uptake Mediated by ENT1
2.2. Strategies to Overcome Transport-Mediated Resistance
3. Enzymatic Determinants of Nucleoside Analog Efficacy
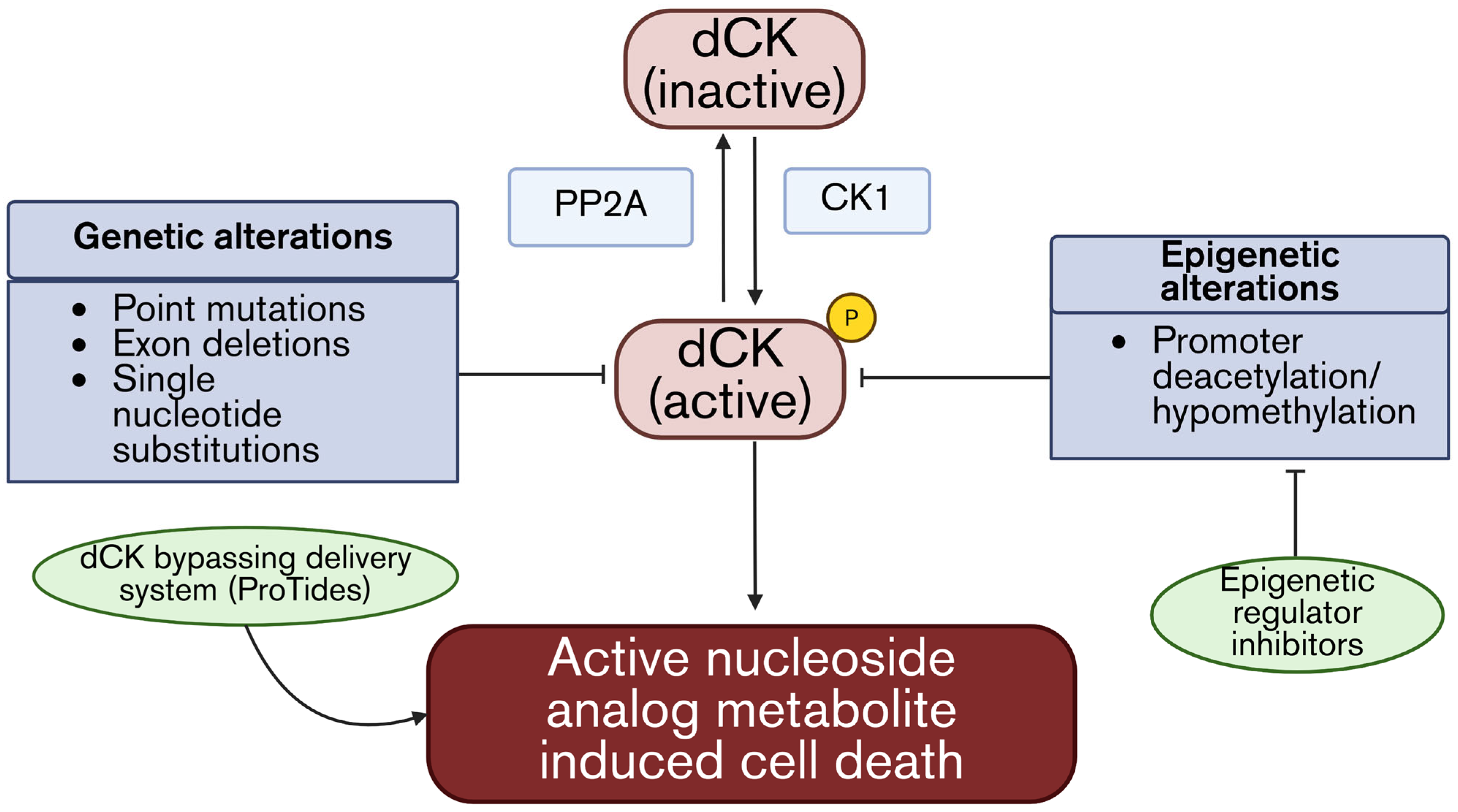
3.1. Enzymatic Activation—Defects in dCK Cause Resistance
3.2. Enzymatic Inactivation—CDA Activity Causes Resistance
3.3. Advancement in CDA Small Molecule Inhibitors
4. Altered Nucleotide Metabolism and dNTP Pool Rewiring
4.1. RNR
4.2. SAMHD1
4.3. Therapeutic Targeting of the RNR–SAMHD1 Axis to Overcome Nucleoside Analog Resistance
5. DNA Damage Response
5.1. Key Components of the DDR
5.2. Targeting Key DDR Components to Overcome Resistance to Nucleoside Analogs
| SMI | Target | Trial Details: ID; Phase; Cancer Type; Drug Combination (If Any) | Study Duration | Status/Outcomes |
|---|---|---|---|---|
| Berzosertib | ATR | NCT04216316 Phase 1b; NSCLC; combination with carboplatin, gemcitabine, and pembrolizumab | 2021–2024 | Completed; combination was tolerable with clinical activity [82] |
| ATG-018 | ATR | NCT05338346 Phase I open label; solid and hematological malignancies; monotherapy | 2022–2024 | Terminated due to lack of activity |
| Camonsertib (RP-3500) | ATR | NCT04497116 Phase 1/2a multi-center, open-label, dose-escalation, and expansion study; advanced solid tumor; combination with PARP inhibitor or GEM | 2020–2025 | Completed; clinical benefit observed with increased benefit in ovarian cancer [94]. |
| Elimusertib | ATR | NCT04514497 Phase I; NSCLC; combination with chemotherapy | 2021–ongoing | Ongoing |
| Prexasertib | CHK1 | NCT02873975 Phase II open label; advanced solid tumors with replicative stress, homologous recombination deficiency, or CCNE1 amplification; monotherapy | 2016–2021 | Completed; results are expected soon |
| GDC-0575 | CHK1 | NCT01564251 Phase I open label dose escalation study; refractory solid tumors or lymphoma; combination with GEM | 2012–2018 | Completed; observed antitumor activity with manageable hematological toxicities [95]. |
| APR-1051 | WEE1 | NCT06260514 Phase 1, open-label, multicenter, first-in-human study; advanced solid tumors; monotherapy | 2024–ongoing | Recruitment phase |
| Adavosertib (AZD-1775) | WEE1 | NCT02037230 Phase I dose escalation study; pancreatic cancer; combination with gemcitabine (+radiation) | 2014–2018 | Completed; overall survival is substantially higher than gemcitabine with radiation therapy [96]. |
6. Mitochondrial and Metabolic Reprogramming
6.1. Mitochondrial Metabolism in Resistance
6.2. Mitochondrial Targeting Strategies
6.3. Anti-Apoptotic BCL-2 Family Proteins-Mediated Resistance
6.4. Targeting BCL-2 with Venetoclax + Nucleoside Analogs
7. Conclusions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sandler, A.B.; Nemunaitis, J.; Denham, C.; Pawel, J.; Cormier, Y.; Gatzemeier, U.; Mattson, K.; Manegold, C.; Palmer, M.C.; Gregor, A.; et al. Phase III Trial of Gemcitabine Plus Cisplatin Versus Cisplatin Alone in Patients With Locally Advanced or Metastatic Non–Small-Cell Lung Cancer. J. Clin. Oncol. 2000, 18, 122–130. [Google Scholar] [CrossRef]
- de Gregorio, A.; Haberle, L.; Fasching, P.A.; Muller, V.; Schrader, I.; Lorenz, R.; Forstbauer, H.; Friedl, T.W.P.; Bauer, E.; de Gregorio, N.; et al. Gemcitabine as adjuvant chemotherapy in patients with high-risk early breast cancer-results from the randomized phase III SUCCESS-A trial. Breast Cancer Res. 2020, 22, 111. [Google Scholar] [CrossRef] [PubMed]
- Iwamoto, K.; Nakashiro, K.-I.; Tanaka, H.; Tokuzen, N.; Hamakawa, H. Ribonucleotide reductase M2 is a promising molecular target for the treatment of oral squamous cell carcinoma. Int. J. Oncol. 2015, 46, 1971–1977. [Google Scholar] [CrossRef] [PubMed]
- Samanta, K.; Setua, S.; Kumari, S.; Jaggi, M.; Yallapu, M.M.; Chauhan, S.C. Gemcitabine Combination Nano Therapies for Pancreatic Cancer. Pharmaceutics 2019, 11, 574. [Google Scholar] [CrossRef]
- Zeng, J.; Funk, J.; Lee, B.R.; Hsu, C.H.; Messing, E.M.; Chipollini, J. Gemcitabine as first-line therapy for high-grade non-muscle invasive bladder cancer: Results from a tertiary center in the contemporary BCG-shortage era. Transl. Androl. Urol. 2023, 12, 960–966. [Google Scholar] [CrossRef] [PubMed]
- Di Francia, R.; Crisci, S.; De Monaco, A.; Cafiero, C.; Re, A.; Iaccarino, G.; De Filippi, R.; Frigeri, F.; Corazzelli, G.; Micera, A.; et al. Response and Toxicity to Cytarabine Therapy in Leukemia and Lymphoma: From Dose Puzzle to Pharmacogenomic Biomarkers. Cancers 2021, 13, 966. [Google Scholar] [CrossRef]
- Faderl, S.; Gandhi, V.; Keating, M.J.; Jeha, S.; Plunkett, W.; Kantarjian, H.M. The role of clofarabine in hematologic and solid malignancies--development of a next-generation nucleoside analog. Cancer 2005, 103, 1985–1995. [Google Scholar] [CrossRef]
- Khan, M.; Phulara, N.R.; Seneviratne, H.K. Fludarabine metabolism by cytosolic 5′-nucleotidase III and its tissue-specific localization mapping using mass spectrometry imaging. Drug Metab. Dispos. 2025, 53, 100178. [Google Scholar] [CrossRef]
- Carter, J.L.; Su, Y.; Qiao, X.; Zhao, J.; Wang, G.; Howard, M.; Edwards, H.; Bao, X.; Li, J.; Huttemann, M.; et al. Acquired resistance to venetoclax plus azacitidine in acute myeloid leukemia: In vitro models and mechanisms. Biochem. Pharmacol. 2023, 216, 115759. [Google Scholar] [CrossRef]
- Li, J.; Fu, S.; Ye, C.; Li, J. Combination therapy involving azacitidine for acute myeloid leukemia patients ineligible for intensive chemotherapy. Leuk. Res. 2025, 148, 107638. [Google Scholar] [CrossRef]
- Yabushita, T.; Goyama, S. Nucleic acid metabolism: The key therapeutic target for myeloid tumors. Exp. Hematol. 2025, 142, 104693. [Google Scholar] [CrossRef]
- Siegel, R.L.; Kratzer, T.B.; Giaquinto, A.N.; Sung, H.; Jemal, A. Cancer statistics, 2025. CA Cancer J. Clin. 2025, 75, 10–45. [Google Scholar] [CrossRef]
- Jin, H.; Wang, L.; Bernards, R. Rational combinations of targeted cancer therapies: Background, advances and challenges. Nat. Rev. Drug Discov. 2023, 22, 213–234. [Google Scholar] [CrossRef]
- Hruba, L.; Das, V.; Hajduch, M.; Dzubak, P. Nucleoside-based anticancer drugs: Mechanism of action and drug resistance. Biochem. Pharmacol. 2023, 215, 115741. [Google Scholar] [CrossRef]
- Espinoza, J.A.; García, P.; Bizama, C.; Leal, J.L.; Riquelme, I.; Weber, H.; Macanas, P.; Aguayo, G.; Viñuela, E.; Roa, J.C.; et al. Low expression of equilibrative nucleoside transporter 1 is associated with poor prognosis in chemotherapy-naïve pT2 gallbladder adenocarcinoma patients. Histopathology 2016, 68, 722–728. [Google Scholar] [CrossRef] [PubMed]
- Spratlin, J.; Sangha, R.; Glubrecht, D.; Dabbagh, L.; Young, J.D.; Dumontet, C.; Cass, C.; Lai, R.; Mackey, J.R. The absence of human equilibrative nucleoside transporter 1 is associated with reduced survival in patients with gemcitabine-treated pancreas adenocarcinoma. Clin. Cancer Res. 2004, 10, 6956–6961. [Google Scholar] [CrossRef]
- Nayak, D.; Weadick, B.; Persaud, A.K.; Raj, R.; Shakya, R.; Li, J.; Campbell, M.J.; Govindarajan, R. EMT alterations in the solute carrier landscape uncover SLC22A10/A15 imposed vulnerabilities in pancreatic cancer. iScience 2022, 25, 104193. [Google Scholar] [CrossRef]
- Eltzschig, H.K.; Abdulla, P.; Hoffman, E.; Hamilton, K.E.; Daniels, D.; Schonfeld, C.; Loffler, M.; Reyes, G.; Duszenko, M.; Karhausen, J.; et al. HIF-1-dependent repression of equilibrative nucleoside transporter (ENT) in hypoxia. J. Exp. Med. 2005, 202, 1493–1505. [Google Scholar] [CrossRef] [PubMed]
- McNamara, M.G.; Bridgewater, J.; Palmer, D.H.; Faluyi, O.; Wasan, H.; Patel, A.; Ryder, W.D.; Barber, S.; Gnanaranjan, C.; Ghazaly, E.; et al. A Phase Ib Study of NUC-1031 in Combination with Cisplatin for the First-Line Treatment of Patients with Advanced Biliary Tract Cancer (ABC-08). Oncologist 2021, 26, e669–e678. [Google Scholar] [CrossRef] [PubMed]
- Kazmi, F.; Nicum, S.; Roux, R.L.; Spiers, L.; Gnanaranjan, C.; Sukumaran, A.; Gabra, H.; Ghazaly, E.; McCracken, N.W.; Harrison, D.J.; et al. A Phase Ib Open-Label, Dose-Escalation Study of NUC-1031 in Combination with Carboplatin for Recurrent Ovarian Cancer. Clin. Cancer Res. 2021, 27, 3028–3038. [Google Scholar] [CrossRef]
- Knox, J.J.; McNamara, M.G.; Bazin, I.S.; Oh, D.-Y.; Zubkov, O.; Breder, V.; Bai, L.-Y.; Christie, A.; Goyal, L.; Cosgrove, D.P.; et al. A phase III randomized study of first-line NUC-1031/cisplatin vs. gemcitabine/cisplatin in advanced biliary tract cancer. J. Hepatol. 2025, 83, 358–366. [Google Scholar] [CrossRef]
- Spiliopoulou, P.; Kazmi, F.; Aroldi, F.; Holmes, T.; Thompson, D.; Griffiths, L.; Qi, C.; Parkes, M.; Lord, S.; Veal, G.J.; et al. A phase I open-label, dose-escalation study of NUC-3373, a targeted thymidylate synthase inhibitor, in patients with advanced cancer (NuTide:301). J. Exp. Clin. Cancer Res. 2024, 43, 100. [Google Scholar] [CrossRef]
- Zhang, L.; Jia, G.; Li, Z.; Sun, S.; Chen, Y.; Zhao, J.; Wang, X.; Xu, W.; Jing, F.; Jiang, Y.; et al. Design, synthesis, and anti-cancer evaluation of the novel conjugate of gemcitabine’s ProTide prodrug based on CD13. Bioorg Chem. 2025, 157, 108293. [Google Scholar] [CrossRef]
- Siciliano, S.; Bernardi, C.; Finetti, F.; Guerrini, A.; Monti, M.C.; Morretta, E.; Petricci, E.; Poggialini, F.; Romagnoli, G.; Taddei, M.; et al. ProTide-enabled antibody-drug conjugates: A novel platform for the targeted delivery of phosphorylated drugs. Bioorg Chem. 2025, 167, 109260. [Google Scholar] [CrossRef]
- Alanazi, A.S.; James, E.; Mehellou, Y. The ProTide Prodrug Technology: Where Next? ACS Med. Chem. Lett. 2019, 10, 2–5. [Google Scholar] [CrossRef] [PubMed]
- Shigdar, S.; Schrand, B.; Giangrande, P.H.; de Franciscis, V. Aptamers: Cutting edge of cancer therapies. Mol. Ther. 2021, 29, 2396–2411. [Google Scholar] [CrossRef]
- Lee, Y.J.; Han, S.R.; Kim, N.Y.; Lee, S.-H.; Jeong, J.-S.; Lee, S.-W. An RNA aptamer that binds carcinoembryonic antigen inhibits hepatic metastasis of colon cancer cells in mice. Gastroenterology 2012, 143, 155–165.e8. [Google Scholar] [CrossRef] [PubMed]
- Park, J.Y.; Cho, Y.L.; Chae, J.R.; Moon, S.H.; Cho, W.G.; Choi, Y.J.; Lee, S.J.; Kang, W.J. Gemcitabine-Incorporated G-Quadruplex Aptamer for Targeted Drug Delivery into Pancreas Cancer. Mol. Ther. Nucleic Acids 2018, 12, 543–553. [Google Scholar] [CrossRef] [PubMed]
- Mahmoudian, F.; Ahmari, A.; Shabani, S.; Sadeghi, B.; Fahimirad, S.; Fattahi, F. Aptamers as an approach to targeted cancer therapy. Cancer Cell Int. 2024, 24, 108. [Google Scholar] [CrossRef]
- Gavas, S.; Quazi, S.; Karpinski, T.M. Nanoparticles for Cancer Therapy: Current Progress and Challenges. Nanoscale Res. Lett. 2021, 16, 173. [Google Scholar] [CrossRef]
- Altammar, K.A. A review on nanoparticles: Characteristics, synthesis, applications, and challenges. Front. Microbiol. 2023, 14, 1155622. [Google Scholar] [CrossRef] [PubMed]
- Arvejeh, P.M.; Chermahini, F.A.; Marincola, F.; Taheri, F.; Mirzaei, S.A.; Alizadeh, A.; Deris, F.; Jafari, R.; Amiri, N.; Soltani, A.; et al. A novel approach for the co-delivery of 5-fluorouracil and everolimus for breast cancer combination therapy: Stimuli-responsive chitosan hydrogel embedded with mesoporous silica nanoparticles. J. Transl. Med. 2025, 23, 382. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Chen, Q.; Zheng, R.; Shao, Z.; Wei, X.; Gu, X.; Zang, Y.; Yang, X.; Wu, J.; Dai, M.; et al. Nano-enabled repurposing of Benproperine phosphate enhances pancreatic Cancer chemotherapy through lethal autophagy arrest and immune activation. J. Control. Release 2025, 387, 114187. [Google Scholar] [CrossRef]
- Wang, Y.; Fan, H.; Chen, Q.; Song, H.; Li, X.; Su, B.; Sun, T.; Cheng, L.; Jiang, C. Mitocytosis-inducing nanoparticles alleviate gemcitabine resistance via dual disruption of pyrimidine synthesis and redox homeostasis in pancreatic ductal adenocarcinoma. Biomaterials 2026, 325, 123630. [Google Scholar] [CrossRef]
- Alyami, H.; Alharthi, S.; Alqahtani, A.J.; Ebrahimi Shahmabadi, H.; Alavi, S.E. Enhanced antitumor efficacy of nanostructured lipid carrier co-loaded with docetaxel and 5-fluorouracil for targeted gastric cancer therapy. Med. Oncol. 2025, 42, 53. [Google Scholar] [CrossRef]
- Mezher, A.H.; Salehpour, M.; Saadati, Z. Folic acid-functionalized and acetyl-terminated dendrimers as nanovectors for co-delivery of sorafenib and 5-fluorouracil. Arch. Biochem. Biophys. 2024, 762, 110176. [Google Scholar] [CrossRef]
- Mehrotra, N.; Anees, M.; Tiwari, S.; Kharbanda, S.; Singh, H. Polylactic acid based polymeric nanoparticle mediated co-delivery of navitoclax and decitabine for cancer therapy. Nanomed. Nanotechnol. Biol. Med. 2023, 47, 102627. [Google Scholar] [CrossRef]
- Hughes, S.J.; Cravetchi, X.; Vilas, G.; Hammond, J.R. Adenosine A1 receptor activation modulates human equilibrative nucleoside transporter 1 (hENT1) activity via PKC-mediated phosphorylation of serine-281. Cell. Signal. 2015, 27, 1008–1018. [Google Scholar] [CrossRef]
- Leisewitz, A.V.; Zimmerman, E.I.; Huang, M.; Jones, S.Z.; Yang, J.; Graves, L.M. Regulation of ENT1 expression and ENT1-dependent nucleoside transport by c-Jun N-terminal kinase. Biochem. Biophys. Res. Commun. 2011, 404, 370–375. [Google Scholar] [CrossRef]
- Knoll, L.; Hamm, J.; Stroebel, P.; Jovan, T.; Goetze, R.; Singh, S.; Hessmann, E.; Ellenrieder, V.; Ammer-Herrmenau, C.; Neesse, A. Expression of gemcitabine metabolizing enzymes and stromal components reveal complexities of preclinical pancreatic cancer models for therapeutic testing. Neoplasia 2024, 53, 101002. [Google Scholar] [CrossRef] [PubMed]
- Abbaspour, A.; Dehghani, M.; Setayesh, M.; Tavakkoli, M.; Rostamipour, H.A.; Ghorbani, M.; Ramzi, M.; Omidvari, S.; Moosavi, F.; Firuzi, O. Cytidine deaminase enzyme activity is a predictive biomarker in gemcitabine-treated cancer patients. Cancer Chemother. Pharmacol. 2023, 92, 475–483. [Google Scholar] [CrossRef]
- Wu, B.; Mao, Z.J.; Wang, Z.; Wu, P.; Huang, H.; Zhao, W.; Zhang, L.; Zhang, Z.; Yin, H.; Gale, R.P.; et al. Deoxycytidine Kinase (DCK) Mutations in Human Acute Myeloid Leukemia Resistant to Cytarabine. Acta Haematol. 2021, 144, 534–541. [Google Scholar] [CrossRef]
- Kurata, M.; Rathe, S.K.; Bailey, N.J.; Aumann, N.K.; Jones, J.M.; Veldhuijzen, G.W.; Moriarity, B.S.; Largaespada, D.A. Using genome-wide CRISPR library screening with library resistant DCK to find new sources of Ara-C drug resistance in AML. Sci. Rep. 2016, 6, 36199. [Google Scholar] [CrossRef]
- Rathe, S.K.; Moriarity, B.S.; Stoltenberg, C.B.; Kurata, M.; Aumann, N.K.; Rahrmann, E.P.; Bailey, N.J.; Melrose, E.G.; Beckmann, D.A.; Liska, C.R.; et al. Using RNA-seq and targeted nucleases to identify mechanisms of drug resistance in acute myeloid leukemia. Sci. Rep. 2014, 4, 6048. [Google Scholar] [CrossRef]
- Remmerie, M.; Dok, R.; Wang, Z.; Omella, J.D.; Alen, S.; Cokelaere, C.; Lenaerts, L.; Dreesen, E.; Nuyts, S.; Derua, R.; et al. The PPP2R1A cancer hotspot mutant p.R183W increases clofarabine resistance in uterine serous carcinoma cells by a gain-of-function mechanism. Cell. Oncol. 2024, 47, 1811–1829. [Google Scholar] [CrossRef]
- Hosokawa, M.; Saito, M.; Nakano, A.; Iwashita, S.; Ishizaka, A.; Ueda, K.; Iwakawa, S. Acquired resistance to decitabine and cross-resistance to gemcitabine during the long-term treatment of human HCT116 colorectal cancer cells with decitabine. Oncol. Lett. 2015, 10, 761–767. [Google Scholar] [CrossRef]
- Kaszycki, J.; Kim, M. Epigenetic Drivers of Chemoresistance in Nucleobase and Nucleoside Analog Therapies. Biology 2025, 14, 838. [Google Scholar] [CrossRef]
- Yoshida, K.; Fujita, A.; Narazaki, H.; Asano, T.; Itoh, Y. Drug resistance to nelarabine in leukemia cell lines might be caused by reduced expression of deoxycytidine kinase through epigenetic mechanisms. Cancer Chemother. Pharmacol. 2022, 89, 83–91. [Google Scholar] [CrossRef]
- Fanciullino, R.; Farnault, L.; Donnette, M.; Imbs, D.C.; Roche, C.; Venton, G.; Berda-Haddad, Y.; Ivanov, V.; Ciccolini, J.; Ouafik, L.; et al. CDA as a predictive marker for life-threatening toxicities in patients with AML treated with cytarabine. Blood Adv. 2018, 2, 462–469. [Google Scholar] [CrossRef]
- Mahfouz, R.Z.; Jankowska, A.; Ebrahem, Q.; Gu, X.; Visconte, V.; Tabarroki, A.; Terse, P.; Covey, J.; Chan, K.; Ling, Y.; et al. Increased CDA expression/activity in males contributes to decreased cytidine analog half-life and likely contributes to worse outcomes with 5-azacytidine or decitabine therapy. Clin. Cancer Res. 2013, 19, 938–948. [Google Scholar] [CrossRef]
- Lumeau, A.; Bery, N.; Frances, A.; Gayral, M.; Labrousse, G.; Ribeyre, C.; Lopez, C.; Nevot, A.; El Kaoutari, A.; Hanoun, N.; et al. Cytidine Deaminase Resolves Replicative Stress and Protects Pancreatic Cancer from DNA-Targeting Drugs. Cancer Res. 2024, 84, 1013–1028. [Google Scholar] [CrossRef]
- Frances, A.; Cordelier, P. The Emerging Role of Cytidine Deaminase in Human Diseases: A New Opportunity for Therapy? Mol. Ther. 2020, 28, 357–366. [Google Scholar] [CrossRef]
- Lemaire, M.; Momparler, L.F.; Raynal, N.J.M.; Bernstein, M.L.; Momparler, R.L. Inhibition of cytidine deaminase by zebularine enhances the antineoplastic action of 5-aza-2′-deoxycytidine. Cancer Chemother. Pharmacol. 2009, 63, 411–416. [Google Scholar] [CrossRef]
- Patki, M.; Saraswat, A.; Bhutkar, S.; Dukhande, V.; Patel, K. In vitro assessment of a synergistic combination of gemcitabine and zebularine in pancreatic cancer cells. Exp. Cell Res. 2021, 405, 112660. [Google Scholar] [CrossRef]
- Kim, N.; Norsworthy, K.J.; Subramaniam, S.; Chen, H.; Manning, M.L.; Kitabi, E.; Earp, J.; Ehrlich, L.A.; Okusanya, O.O.; Vallejo, J.; et al. FDA Approval Summary: Decitabine and Cedazuridine Tablets for Myelodysplastic Syndromes. Clin. Cancer Res. 2022, 28, 3411–3416. [Google Scholar] [CrossRef]
- Helleberg, H.; Bjornsdottir, I.; Offenberg, H.K. Effects of Tetrahydrouridine Pre-dosing on Absorption, Metabolism, and Excretion of Decitabine in the Mouse. Eur. J. Pharm. Sci. 2025, 216, 107353. [Google Scholar] [CrossRef]
- Ureshino, H.; Kurahashi, Y.; Watanabe, T.; Yamashita, S.; Kamachi, K.; Yamamoto, Y.; Fukuda-Kurahashi, Y.; Yoshida-Sakai, N.; Hattori, N.; Hayashi, Y.; et al. Silylation of Deoxynucleotide Analog Yields an Orally Available Drug with Antileukemia Effects. Mol. Cancer Ther. 2021, 20, 1412–1421. [Google Scholar] [CrossRef]
- Moorthy, R.; Grillo, M.J.; Baur, J.W.; Schmidt, S.A.; Passow, K.T.; Demir, O.; Tang, J.; Olson, M.E.; Amaro, R.E.; Harki, D.A. Development of a Real-Time Fluorescence-Based Deamination Assay and Identification of Inhibitors of Human Cytidine Deaminase. ACS Chem. Biol. 2025, 20, 2593–2600. [Google Scholar] [CrossRef]
- Chabes, A.; Thelander, L. Controlled protein degradation regulates ribonucleotide reductase activity in proliferating mammalian cells during the normal cell cycle and in response to DNA damage and replication blocks. J. Biol. Chem. 2000, 275, 17747–17753. [Google Scholar] [CrossRef]
- Franzolin, E.; Pontarin, G.; Rampazzo, C.; Miazzi, C.; Ferraro, P.; Palumbo, E.; Reichard Reichard, P.; Bianchi, V. The deoxynucleotide triphosphohydrolase SAMHD1 is a major regulator of DNA precursor pools in mammalian cells. Proc. Natl. Acad. Sci. USA 2013, 110, 14272–14277. [Google Scholar] [CrossRef]
- Tian, L.; Chen, C.; Guo, Y.; Zhang, F.; Mi, J.; Feng, Q.; Lin, S.; Xi, N.; Tian, J.; Yu, L.; et al. mTORC2 regulates ribonucleotide reductase to promote DNA replication and gemcitabine resistance in non-small cell lung cancer. Neoplasia 2021, 23, 643–652. [Google Scholar] [CrossRef]
- Liu, X.; Zhang, H.; Lai, L.; Wang, X.; Loera, S.; Xue, L.; He, H.; Zhang, K.; Hu, S.; Huang, Y.; et al. Ribonucleotide reductase small subunit M2 serves as a prognostic biomarker and predicts poor survival of colorectal cancers. Clin. Sci. 2013, 124, 567–578. [Google Scholar] [CrossRef]
- Kato, T.; Ono, H.; Fujii, M.; Akahoshi, K.; Ogura, T.; Ogawa, K.; Ban, D.; Kudo, A.; Tanaka, S.; Tanabe, M. Cytoplasmic RRM1 activation as an acute response to gemcitabine treatment is involved in drug resistance of pancreatic cancer cells. PLoS ONE 2021, 16, e0252917. [Google Scholar] [CrossRef]
- Herold, N.; Rudd, S.G.; Ljungblad, L.; Sanjiv, K.; Myrberg, I.H.; Paulin, C.B.; Heshmati, Y.; Hagenkort, A.; Kutzner, J.; Page, B.D.; et al. Targeting SAMHD1 with the Vpx protein to improve cytarabine therapy for hematological malignancies. Nat. Med. 2017, 23, 256–263. [Google Scholar] [CrossRef]
- Schneider, C.; Oellerich, T.; Baldauf, H.A.-O.X.; Schwarz, S.M.; Thomas, D.; Flick, R.; Bohnenberger, H.; Kaderali, L.; Stegmann, L.; Cremer, A.; et al. SAMHD1 is a biomarker for cytarabine response and a therapeutic target in acute myeloid leukemia. Nat. Med. 2017, 23, 250–255. [Google Scholar] [CrossRef]
- Knecht, K.M.; Buzovetsky, O.; Schneider, C.; Thomas, D.; Srikanth, V.; Kaderali, L.; Tofoleanu, F.; Reiss, K.; Ferreirós, N.; Geisslinger, G.; et al. The structural basis for cancer drug interactions with the catalytic and allosteric sites of SAMHD1. Proc. Natl. Acad. Sci. USA 2018, 115, E10022–E10031. [Google Scholar] [CrossRef]
- Rudd, S.G.; Tsesmetzis, N.; Sanjiv, K.; Paulin, C.B.; Sandhow, L.; Kutzner, J.; Hed Myrberg, I.; Bunten, S.S.; Axelsson, H.; Zhang, S.M.; et al. Ribonucleotide reductase inhibitors suppress SAMHD1 ara-CTPase activity enhancing cytarabine efficacy. EMBO Mol. Med. 2020, 12, e10419. [Google Scholar] [CrossRef]
- McCown, C.; Yu, C.H.; Ivanov, D.N. SAMHD1 shapes deoxynucleotide triphosphate homeostasis by interconnecting the depletion and biosynthesis of different dNTPs. Nat. Commun. 2025, 16, 793. [Google Scholar] [CrossRef]
- Huff, S.E.; Winter, J.M.; Dealwis, C.G. Inhibitors of the Cancer Target Ribonucleotide Reductase, Past and Present. Biomolecules 2022, 12, 815. [Google Scholar] [CrossRef]
- Jadersten, M.; Lilienthal, I.; Tsesmetzis, N.; Lourda, M.; Bengtzen, S.; Bohlin, A.; Arnroth, C.; Erkers, T.; Seashore-Ludlow, B.; Giraud, G.; et al. Targeting SAMHD1 with hydroxyurea in first-line cytarabine-based therapy of newly diagnosed acute myeloid leukaemia: Results from the HEAT-AML trial. J. Intern. Med. 2022, 292, 925–940. [Google Scholar] [CrossRef]
- Nair, R.; Salinas-Illarena, A.; Sponheimer, M.; Wullkopf, I.; Schreiber, Y.; Corte-Real, J.V.; Del Pozo Ben, A.; Marterer, H.; Thomas, D.; Geisslinger, G.; et al. Novel Vpx virus-like particles to improve cytarabine treatment response against acute myeloid leukemia. Clin. Exp. Med. 2024, 24, 155. [Google Scholar] [CrossRef]
- De Mel, S.; Lee, A.R.; Tan, J.H.I.; Tan, R.Z.Y.; Poon, L.M.; Chan, E.; Lee, J.; Chee, Y.L.; Lakshminarasappa, S.R.; Jaynes, P.W.; et al. Targeting the DNA damage response in hematological malignancies. Front. Oncol. 2024, 14, 1307839. [Google Scholar] [CrossRef]
- Saldivar, J.C.; Cortez, D.; Cimprich, K.A. The essential kinase ATR: Ensuring faithful duplication of a challenging genome. Nat. Rev. Mol. Cell Biol. 2017, 18, 622–636. [Google Scholar] [CrossRef]
- Saxena, S.; Zou, L. Hallmarks of DNA replication stress. Mol. Cell 2022, 82, 2298–2314. [Google Scholar] [CrossRef]
- Qi, W.; Xie, C.; Li, C.; Caldwell, J.T.; Edwards, H.; Taub, J.W.; Wang, Y.; Lin, H.; Ge, Y. CHK1 plays a critical role in the anti-leukemic activity of the wee1 inhibitor MK-1775 in acute myeloid leukemia cells. J. Hematol. Oncol. 2014, 7, 53. [Google Scholar] [CrossRef]
- Esposito, F.A.-O.; Giuffrida, R.A.-O.; Raciti, G.; Puglisi, C.A.-O.; Forte, S.A.-O. Wee1 Kinase: A Potential Target to Overcome Tumor Resistance to Therapy. Int. J. Mol. Sci. 2021, 22, 10689. [Google Scholar] [CrossRef]
- Patil, M.; Pabla, N.; Dong, Z. Checkpoint kinase 1 in DNA damage response and cell cycle regulation. Cell Mol. Life Sci. 2013, 70, 4009–4021. [Google Scholar] [CrossRef]
- Saldanha, J.; Rageul, J.; Patel, J.A.; Kim, H. The Adaptive Mechanisms and Checkpoint Responses to a Stressed DNA Replication Fork. Int. J. Mol. Sci. 2023, 24, 10488. [Google Scholar] [CrossRef]
- Wang, M.; Chen, S.; Ao, D. Targeting DNA repair pathway in cancer: Mechanisms and clinical application. MedComm 2021, 2, 654–691. [Google Scholar] [CrossRef]
- Morano, L.; Vie, N.; Aissanou, A.; Hodroj, D.; Garambois, V.; Fauvre, A.; Promonet, A.; Egger, T.; Bordignon, B.; Hassen-Khodja, C.; et al. Shining light on drug discovery: Optogenetic screening for TopBP1 biomolecular condensate inhibitors. NAR Cancer 2025, 7, zcaf041. [Google Scholar] [CrossRef]
- Wallez, Y.; Dunlop, C.R.; Johnson, T.I.; Koh, S.B.; Fornari, C.; Yates, J.W.T.; Bernaldo de Quiros Fernandez, S.; Lau, A.; Richards, F.M.; Jodrell, D.I. The ATR Inhibitor AZD6738 Synergizes with Gemcitabine In Vitro and In Vivo to Induce Pancreatic Ductal Adenocarcinoma Regression. Mol. Cancer Ther. 2018, 17, 1670–1682. [Google Scholar] [CrossRef]
- Villaruz, L.C.; Schluger, B.; Wang, H.; Ohr, J.; Petro, D.; Fuld, A.D.; Herzberg, B.; Dawar, R.; Nieva, J.; Ruiz, J.; et al. Phase Ib study of berzosertib, carboplatin, gemcitabine, and pembrolizumab in patients with squamous nonsmall lung cancer (ETCTN 10313). Oncologist 2025, 30, oyaf373. [Google Scholar] [CrossRef]
- Konstantinopoulos, P.A.; Lee, J.M.; Gao, B.; Miller, R.; Lee, J.Y.; Colombo, N.; Vergote, I.; Credille, K.M.; Young, S.R.; McNeely, S.; et al. A Phase 2 study of prexasertib (LY2606368) in platinum resistant or refractory recurrent ovarian cancer. Gynecol. Oncol. 2022, 167, 213–225. [Google Scholar] [CrossRef]
- Rodriguez-Vazquez, G.O.; Diaz-Quinones, A.O.; Chorna, N.; Salgado-Villanueva, I.K.; Tang, J.; Ortiz, W.I.S.; Maldonado, H.M. Synergistic interactions of cytarabine-adavosertib in leukemic cell lines proliferation and metabolomic endpoints. Biomed. Pharmacother. 2023, 166, 115352. [Google Scholar] [CrossRef] [PubMed]
- Bell, H.L.; Blair, H.J.; Singh, M.; Moorman, A.V.; Heidenreich, O.; van Delft, F.W.; Lunec, J.; Irving, J.A.E. Targeting WEE1 kinase as a p53-independent therapeutic strategy in high-risk and relapsed acute lymphoblastic leukemia. Cancer Cell Int. 2023, 23, 202. [Google Scholar] [CrossRef]
- Chen, C.P.; Hung, T.H.; Hsu, P.C.; Yeh, C.N.; Huang, W.K.; Pan, Y.R.; Hsiao, Y.T.; Lo, C.H.; Wu, C.E. Synergistic effects of MK-1775 and gemcitabine on cytotoxicity in non-small cell lung cancer. Heliyon 2024, 10, e40299. [Google Scholar] [CrossRef]
- Chen, C.P.; Yeh, C.N.; Pan, Y.R.; Huang, W.K.; Hsiao, Y.T.; Lo, C.H.; Wu, C.E. Wee1 inhibition by MK1775 potentiates gemcitabine through accumulated replication stress leading to apoptosis in biliary tract cancer. Biomed. Pharmacother. 2023, 166, 115389. [Google Scholar] [CrossRef]
- Lheureux, S.; Cristea, M.C.; Bruce, J.P.; Garg, S.; Cabanero, M.; Mantia-Smaldone, G.; Olawaiye, A.B.; Ellard, S.L.; Weberpals, J.I.; Wahner Hendrickson, A.E.; et al. Adavosertib plus gemcitabine for platinum-resistant or platinum-refractory recurrent ovarian cancer: A double-blind, randomised, placebo-controlled, phase 2 trial. Lancet 2021, 397, 281–292. [Google Scholar] [CrossRef]
- Zeng, Y.; Arisa, O.; Peer, C.J.; Fojo, A.; Figg, W.D. PARP inhibitors: A review of the pharmacology, pharmacokinetics, and pharmacogenetics. Semin. Oncol. 2024, 51, 19–24. [Google Scholar] [CrossRef]
- Jiang, Y.; Dai, H.; Li, Y.; Yin, J.; Guo, S.; Lin, S.Y.; McGrail, D.J. PARP inhibitors synergize with gemcitabine by potentiating DNA damage in non-small-cell lung cancer. Int. J. Cancer 2019, 144, 1092–1103. [Google Scholar] [CrossRef]
- Shabashvili, D.E.; Feng, Y.; Kaur, P.; Venugopal, K.; Guryanova, O.A. Combination strategies to promote sensitivity to cytarabine-induced replication stress in acute myeloid leukemia with and without DNMT3A mutations. Exp. Hematol. 2022, 110, 20–27. [Google Scholar] [CrossRef]
- Hilton, B.A.; Li, Z.; Musich, P.R.; Wang, H.; Cartwright, B.M.; Serrano, M.; Zhou, X.Z.; Lu, K.P.; Zou, Y. ATR Plays a Direct Antiapoptotic Role at Mitochondria, which Is Regulated by Prolyl Isomerase Pin1. Mol. Cell 2015, 60, 35–46. [Google Scholar] [CrossRef]
- Luo, Y.; Biswas, H.; Makinwa, Y.; Liu, S.-H.; Dong, Z.; Liu, J.-Y.; Zhang, J.-T.; Zou, Y. A PP2A-mtATR-tBid axis links DNA damage-induced CIP2A degradation to apoptotic dormancy and therapeutic resistance in PDAC. Cancer Lett. 2025, 627, 217790. [Google Scholar] [CrossRef]
- Yap, T.A.; Fontana, E.; Lee, E.K.; Spigel, D.R.; Hojgaard, M.; Lheureux, S.; Mettu, N.B.; Carneiro, B.A.; Carter, L.; Plummer, R.; et al. Camonsertib in DNA damage response-deficient advanced solid tumors: Phase 1 trial results. Nat. Med. 2023, 29, 1400–1411. [Google Scholar] [CrossRef]
- Italiano, A.; Infante, J.R.; Shapiro, G.I.; Moore, K.N.; LoRusso, P.M.; Hamilton, E.; Cousin, S.; Toulmonde, M.; Postel-Vinay, S.; Tolaney, S.; et al. Phase I study of the checkpoint kinase 1 inhibitor GDC-0575 in combination with gemcitabine in patients with refractory solid tumors. Ann. Oncol. 2018, 29, 1304–1311. [Google Scholar] [CrossRef]
- Cuneo, K.C.; Morgan, M.A.; Sahai, V.; Schipper, M.J.; Parsels, L.A.; Parsels, J.D.; Devasia, T.; Al-Hawaray, M.; Cho, C.S.; Nathan, H.; et al. Dose Escalation Trial of the Wee1 Inhibitor Adavosertib (AZD1775) in Combination With Gemcitabine and Radiation for Patients With Locally Advanced Pancreatic Cancer. J. Clin. Oncol. 2019, 37, 2643–2650. [Google Scholar] [CrossRef] [PubMed]
- Carter, J.L.; Hege, K.; Yang, J.; Kalpage, H.A.; Su, Y.; Edwards, H.; Huttemann, M.; Taub, J.W.; Ge, Y. Targeting multiple signaling pathways: The new approach to acute myeloid leukemia therapy. Signal Transduct. Target. Ther. 2020, 5, 288. [Google Scholar] [CrossRef] [PubMed]
- van Gastel, N.; Spinelli, J.B.; Sharda, A.; Schajnovitz, A.; Baryawno, N.; Rhee, C.; Oki, T.; Grace, E.; Soled, H.J.; Milosevic, J.; et al. Induction of a Timed Metabolic Collapse to Overcome Cancer Chemoresistance. Cell Metab. 2020, 32, 391–403 e396. [Google Scholar] [CrossRef]
- Ikeda, J.; Kunimoto, H.; Saito, Y.; Tsujimoto, S.I.; Takeuchi, M.; Miura, A.; Kurosawa, T.; Murakami, K.; Kato, I.; Funakoshi-Tago, M.; et al. Metabolic reprogramming by PRDM16 drives cytarabine resistance in acute myeloid leukemia. Haematologica 2025, 110, 2647–2660. [Google Scholar] [CrossRef] [PubMed]
- Dash, S.; Ueda, T.; Komuro, A.; Honda, M.; Sugisawa, R.; Okada, H. Deoxycytidine kinase inactivation enhances gemcitabine resistance and sensitizes mitochondrial metabolism interference in pancreatic cancer. Cell Death Dis. 2024, 15, 131. [Google Scholar] [CrossRef]
- Farge, T.; Saland, E.; de Toni, F.; Aroua, N.; Hosseini, M.; Perry, R.; Bosc, C.; Sugita, M.; Stuani, L.; Fraisse, M.; et al. Chemotherapy-Resistant Human Acute Myeloid Leukemia Cells Are Not Enriched for Leukemic Stem Cells but Require Oxidative Metabolism. Cancer Discov. 2017, 7, 716–735. [Google Scholar] [CrossRef] [PubMed]
- Baccelli, I.; Gareau, Y.; Lehnertz, B.; Gingras, S.; Spinella, J.F.; Corneau, S.; Mayotte, N.; Girard, S.; Frechette, M.; Blouin-Chagnon, V.; et al. Mubritinib Targets the Electron Transport Chain Complex I and Reveals the Landscape of OXPHOS Dependency in Acute Myeloid Leukemia. Cancer Cell 2019, 36, 84–99 e88. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Cubillos-Ruiz, J.R. Endoplasmic reticulum stress signals in the tumour and its microenvironment. Nat. Rev. Cancer 2021, 21, 71–88. [Google Scholar] [CrossRef] [PubMed]
- Fujiwara-Tani, R.; Sasaki, T.; Takagi, T.; Mori, S.; Kishi, S.; Nishiguchi, Y.; Ohmori, H.; Fujii, K.; Kuniyasu, H. Gemcitabine Resistance in Pancreatic Ductal Carcinoma Cell Lines Stems from Reprogramming of Energy Metabolism. Int. J. Mol. Sci. 2022, 23, 7824. [Google Scholar] [CrossRef]
- Yeon Chae, S.; Jang, S.Y.; Kim, J.; Hwang, S.; Malani, D.; Kallioniemi, O.; Yun, S.G.; Kim, J.S.; Kim, H.I. Mechanisms of chemotherapy failure in refractory/relapsed acute myeloid leukemia: The role of cytarabine resistance and mitochondrial metabolism. Cell Death Dis. 2025, 16, 331. [Google Scholar] [CrossRef]
- Boet, E.; Sarry, J.E. Targeting Metabolic Dependencies Fueling the TCA Cycle to Circumvent Therapy Resistance in Acute Myeloid Leukemia. Cancer Res. 2024, 84, 950–952. [Google Scholar] [CrossRef]
- Su, Y.; Carter, J.L.; Li, X.; Fukuda, Y.; Gray, A.; Lynch, J.; Edwards, H.; Ma, J.; Schreiner, P.; Polin, L.; et al. The Imipridone ONC213 Targets alpha-Ketoglutarate Dehydrogenase to Induce Mitochondrial Stress and Suppress Oxidative Phosphorylation in Acute Myeloid Leukemia. Cancer Res. 2024, 84, 1084–1100. [Google Scholar] [CrossRef] [PubMed]
- Zarei, M.; Hajihassani, O.; Hue, J.J.; Loftus, A.W.; Graor, H.J.; Nakazzi, F.; Naji, P.; Boutros, C.S.; Uppin, V.; Vaziri-Gohar, A.; et al. IDH1 Inhibition Potentiates Chemotherapy Efficacy in Pancreatic Cancer. Cancer Res. 2024, 84, 3072–3085. [Google Scholar] [CrossRef]
- Masoud, R.; Reyes-Castellanos, G.; Lac, S.; Garcia, J.; Dou, S.; Shintu, L.; Abdel Hadi, N.; Gicquel, T.; El Kaoutari, A.; Dieme, B.; et al. Targeting Mitochondrial Complex I Overcomes Chemoresistance in High OXPHOS Pancreatic Cancer. Cell Rep. Med. 2020, 1, 100143. [Google Scholar] [CrossRef]
- Mauro-Lizcano, M.; Di Pisa, F.; Larrea Murillo, L.; Sugden, C.J.; Sotgia, F.; Lisanti, M.P. High mitochondrial DNA content is a key determinant of stemness, proliferation, cell migration, and cancer metastasis in vivo. Cell Death Dis. 2024, 15, 745. [Google Scholar] [CrossRef]
- Wang, W.; Jiang, C.F.; Yin, H.S.; Gao, S.; Yu, B.P. Targeting mitochondrial transcription factor A sensitizes pancreatic cancer cell to gemcitabine. Hepatobiliary Pancreat. Dis. Int. 2023, 22, 519–527. [Google Scholar] [CrossRef]
- Tan, S.; Kim, S.; Kim, Y. Targeting mitochondrial RNAs enhances the efficacy of the DNA-demethylating agents. Sci. Rep. 2024, 14, 30767. [Google Scholar] [CrossRef]
- Carneiro, B.A.; El-Deiry, W.S. Targeting apoptosis in cancer therapy. Nat. Rev. Clin. Oncol. 2020, 17, 395–417. [Google Scholar] [CrossRef]
- Glaviano, A.; Weisberg, E.; Lam, H.Y.; Tan, D.J.J.; Innes, A.J.; Ge, Y.; Lai, C.E.; Stock, W.; Glytsou, C.; Smit, L.; et al. Apoptosis-targeting BH3 mimetics: Transforming treatment for patients with acute myeloid leukaemia. Nat. Rev. Clin. Oncol. 2025, 22, 847–868. [Google Scholar] [CrossRef] [PubMed]
- Qian, S.; Wei, Z.; Yang, W.; Huang, J.; Yang, Y.; Wang, J. The role of BCL-2 family proteins in regulating apoptosis and cancer therapy. Front. Oncol. 2022, 12, 985363. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Li, C.; Liu, Y. The role of metabolic reprogramming in pancreatic cancer chemoresistance. Front. Pharmacol. 2022, 13, 1108776. [Google Scholar] [CrossRef] [PubMed]
- Nishi, R.; Shigemi, H.; Negoro, E.; Okura, M.; Hosono, N.; Yamauchi, T. Venetoclax and alvocidib are both cytotoxic to acute myeloid leukemia cells resistant to cytarabine and clofarabine. BMC Cancer 2020, 20, 984. [Google Scholar] [CrossRef]
- Niu, X.; Zhao, J.; Ma, J.; Xie, C.; Edwards, H.; Wang, G.; Caldwell, J.T.; Xiang, S.; Zhang, X.; Chu, R.; et al. Binding of Released Bim to Mcl-1 is a Mechanism of Intrinsic Resistance to ABT-199 which can be Overcome by Combination with Daunorubicin or Cytarabine in AML Cells. Clin. Cancer Res. 2016, 22, 4440–4451. [Google Scholar] [CrossRef]
- DiNardo, C.D.; Jonas, B.A.; Pullarkat, V.; Thirman, M.J.; Garcia, J.S.; Wei, A.H.; Konopleva, M.; Döhner, H.; Letai, A.; Fenaux, P.; et al. Azacitidine and Venetoclax in Previously Untreated Acute Myeloid Leukemia. N. Engl. J. Med. 2020, 383, 617–629. [Google Scholar] [CrossRef]
- Asada, N.; Ando, J.; Takada, S.; Yoshida, C.; Usuki, K.; Shinagawa, A.; Ishizawa, K.; Miyamoto, T.; Iida, H.; Dobashi, N.; et al. Venetoclax plus low-dose cytarabine in patients with newly diagnosed acute myeloid leukemia ineligible for intensive chemotherapy: An expanded access study in Japan. Jpn. J. Clin. Oncol. 2023, 53, 595–603. [Google Scholar] [CrossRef]
- Wei, A.H.; Strickland, S.A.; Hou, J.-Z.; Fiedler, W.; Lin, T.L.; Walter, R.B.; Enjeti, A.; Tiong, I.S.; Savona, M.; Lee, S.; et al. Venetoclax Combined With Low-Dose Cytarabine for Previously Untreated Patients With Acute Myeloid Leukemia: Results From a Phase Ib/II Study. J. Clin. Oncol. 2019, 37, 1277–1284. [Google Scholar] [CrossRef] [PubMed]
- Kadia, T.M.; Reville, P.K.; Borthakur, G.; Yilmaz, M.; Kornblau, S.; Alvarado, Y.; Dinardo, C.D.; Daver, N.; Jain, N.; Pemmaraju, N.; et al. Venetoclax plus intensive chemotherapy with cladribine, idarubicin, and cytarabine in patients with newly diagnosed acute myeloid leukaemia or high-risk myelodysplastic syndrome: A cohort from a single-centre, single-arm, phase 2 trial. Lancet Haematol. 2021, 8, e552–e561. [Google Scholar] [CrossRef]
- Mantzaris, I.; Goldfinger, M.; Uriel, M.; Shastri, A.; Shah, N.; Gritsman, K.; Kornblum, N.S.; Shapiro, L.; Sica, R.A.; Munoz, A.; et al. Venetoclax plus daunorubicin and cytarabine for newly diagnosed acute myeloid leukemia: Results of a phase 1b study. Blood 2025, 145, 1870–1875. [Google Scholar] [CrossRef]
- Li, Y.Y.; Ge, S.S.; Huang, Y.H.; Xu, M.Z.; Wan, C.L.; Tan, K.W.; Tao, T.; Zhou, H.X.; Xue, S.L.; Dai, H.P. Efficacy and safety of cladribine, low-dose cytarabine and venetoclax in relapsed/refractory acute myeloid leukemia: Results of a pilot study. Blood Cancer J. 2024, 14, 12. [Google Scholar] [CrossRef] [PubMed]
- Vogler, M.; Braun, Y.; Smith, V.M.; Westhoff, M.-A.; Pereira, R.S.; Pieper, N.M.; Anders, M.; Callens, M.; Vervliet, T.; Abbas, M.; et al. The BCL2 family: From apoptosis mechanisms to new advances in targeted therapy. Signal Transduct. Target. Ther. 2025, 10, 91. [Google Scholar] [CrossRef] [PubMed]




| Drug | FDA Approval | Cancer Use | Mechanism of Action |
|---|---|---|---|
| Gemcitabine (GEM) | 1996 | Pancreatic cancer | Inhibition of DNA synthesis Inhibition of ribonucleotide reductase (RNR) |
| 1998 | Non-small cell lung cancer | ||
| 2004 | Breast cancer | ||
| Fluorouracil (5-FU) | 1962 | Adenocarcinoma of the colon, rectum, breast, stomach, and pancreas | Inhibition of RNA processing Inhibition of thymidylate synthase |
| Cytarabine (Ara-C) | 1969 | Acute myeloid leukemia Acute lymphocytic leukemia | Inhibition of DNA synthesis |
| Azacitidine (AZA) | 2004 | Myelodysplastic syndrome | Inhibition of RNA synthesis |
| 2020 | Acute myeloid leukemia | Inhibition of DNA methyltransferase (DNMT), causing DNA hypomethylation | |
| Decitabine (DAC) | 2006 | Myelodysplastic syndrome | Inhibition of DNMT causing DNA hypomethylation |
| Cladribine (CdA) | 1993 | Hairy cell leukemia | Inhibition of DNA synthesis |
| Fludarabine (FA) | 1991 | B-cell chronic lymphocytic leukemia | Inhibition of DNA synthesis Inhibition of DNA polymerase, RNR, and DNA primase |
| Nelarabine (Ara-G) | 2006 | T-cell acute lymphoblastic leukemia T-cell lymphoblastic lymphoma | Inhibition of DNA synthesis |
| Nanoparticle | Therapeutic Cargos | Purpose | Key Results |
|---|---|---|---|
| Mesoporous Silica Nanoparticle/5FU- Everolimus@ chitosan hydrogels (MSN/5FU-EVE@CSH) | 5-FU and EVE | MSN improves drug targeting and therapeutic efficacy by exhibiting high surface-to-volume ratio 5-FU inhibits DNA synthesis EVE targets mTOR pathway CSH sustains drug release | The combination treatment groups loaded on MSN and CSH nanoparticles showed a significant reduction in tumor size and lung metastasis compared to the monotherapy and control groups [32]. |
| Hyaluronic-acid-modified zeolitic imidazolate framework @benproperine phosphate/GEM (HA/ZIF-8@BPP/Gem) | GEM and BPP | BPP initiated autophagy but blocked autophagosome-lysosome fusion; thereby, GEM-induced protective autophagy was turned into a lethal autophagy. | Represents an efficient drug-repurposing nanoplatform for delivering a promising drug combination, which may sensitize pancreatic cancer to chemotherapy and improve patient survival outcomes [33]. |
| GE11 peptide-modified polyphenol-iron chelate nanoparticles/GEM- leflunomide (GE11-PEG-pPCA/Gem-Lef) | GEM and leflunomide | Leflunomide inhibits dihydroorotate dehydrogenase (DHODH) that is upregulated in GEM resistance | Demonstrated superior antitumor efficacy compared to the standard of care regimen. GEM concentration was lowered 6.3-fold in nanoparticles with increased benefit compared to free drug [34]. |
| Nanostructured lipid carrier- Docetaxel/5FU (NLC-DCT/5-FU) | 5-FU and DCT | DCT and 5-FU strongly cooperate to stop cell proliferation and induce cell death | In vitro cytotoxicity assays showed significantly lower IC50 values for nanoparticle combination compared to free DOX and FU. In vivo studies showed significantly reduced tumor size and low toxicity [35]. |
| Generation 4 (G4) acetyl-terminated poly(amidoamine) dendrimers conjugated with folic acid: 5FU: Sorafenib (G4ACE-FA:5-FU:SF) | 5-FU and SF | G4ACE-FA sustains strong drug binding and targeted release to tumor. SF is a multi-kinase inhibitor that suppresses angiogenesis and tumor proliferation. Combining with FU can synergize to kill cancer cells | Nanoparticles were successfully synthesized using a folic-acid-based platform. Use of this platform demonstrated synergy with the loaded cargo, offering a promising platform to deliver anti-cancer drugs [36]. |
| Navitoclax/decitabine nanoparticles (NAV/DCB NPs) | DAC and NAV | NAV is a pan BCL-2 inhibitor that synergizes with DAC in solid and liquid tumors | In vitro and in vivo evaluation of NAV/DCB demonstrated synergistic effects, increased drug accumulation, and decreased toxicity compared to free drug in both AML and breast cancer models [37]. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Thibodeau, J.; Hershberger, K.; Ramakrishna, S.S.M.; Su, Y.; Timmer, L.; Brophy, B.; Zhang, K.; Edwards, H.; Taub, J.W.; Ge, Y. Enhancing the Nucleoside Analog Response with Translational Therapeutic Approaches to Overcome Resistance. Cells 2026, 15, 130. https://doi.org/10.3390/cells15020130
Thibodeau J, Hershberger K, Ramakrishna SSM, Su Y, Timmer L, Brophy B, Zhang K, Edwards H, Taub JW, Ge Y. Enhancing the Nucleoside Analog Response with Translational Therapeutic Approaches to Overcome Resistance. Cells. 2026; 15(2):130. https://doi.org/10.3390/cells15020130
Chicago/Turabian StyleThibodeau, Jenna, Kian Hershberger, Sai Samanvitha M. Ramakrishna, Yongwei Su, Lauren Timmer, Bryce Brophy, Katherine Zhang, Holly Edwards, Jeffrey W. Taub, and Yubin Ge. 2026. "Enhancing the Nucleoside Analog Response with Translational Therapeutic Approaches to Overcome Resistance" Cells 15, no. 2: 130. https://doi.org/10.3390/cells15020130
APA StyleThibodeau, J., Hershberger, K., Ramakrishna, S. S. M., Su, Y., Timmer, L., Brophy, B., Zhang, K., Edwards, H., Taub, J. W., & Ge, Y. (2026). Enhancing the Nucleoside Analog Response with Translational Therapeutic Approaches to Overcome Resistance. Cells, 15(2), 130. https://doi.org/10.3390/cells15020130







