Establishment of a CRISPR/dCas9 Activation Library for Screening Transcription Factors Co-Regulating OCT4 with GATA4 in Pig Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Culture
2.2. Plasmid Construction
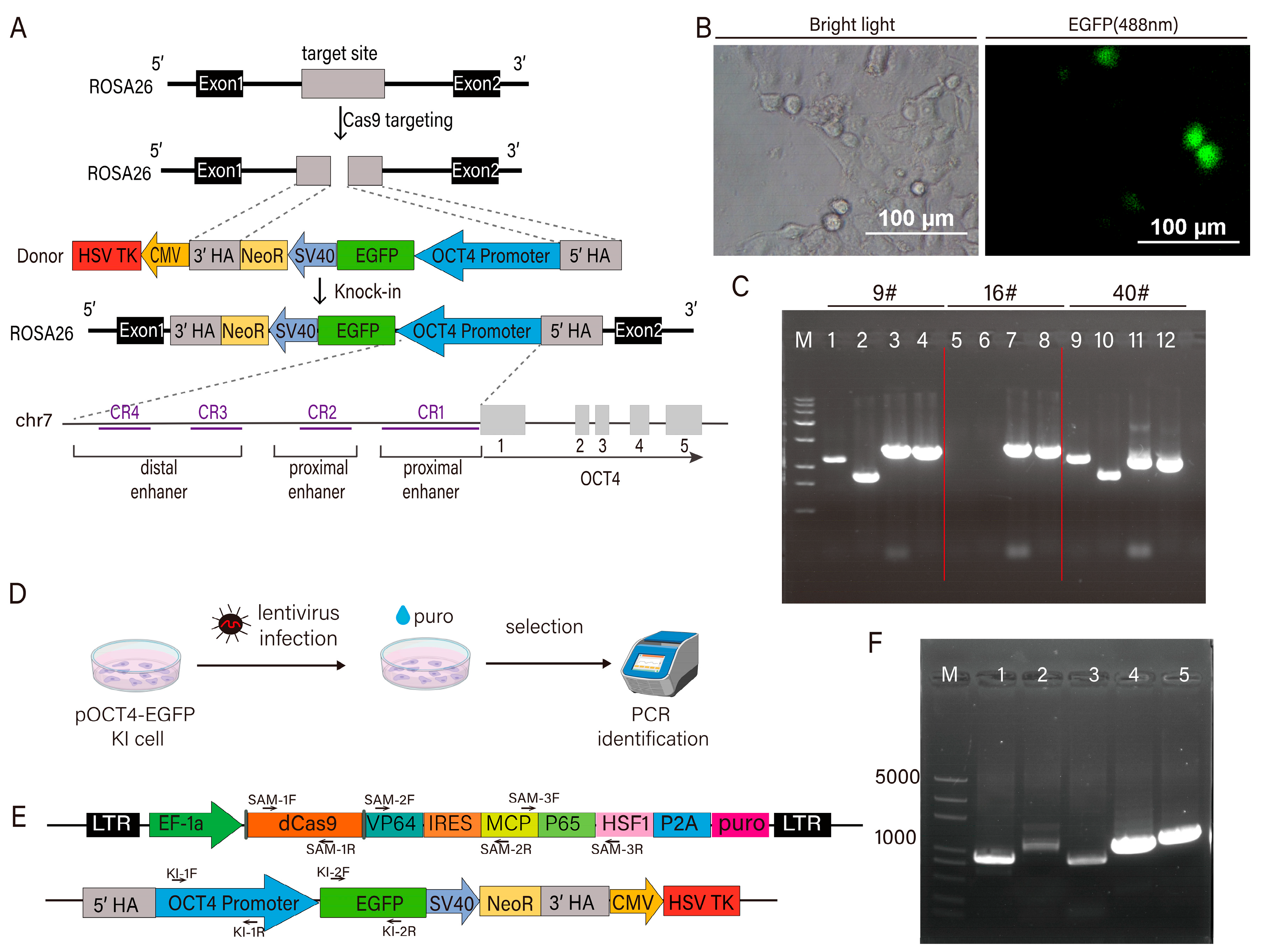
2.2.1. Construction of pOCT4-EGFP-TK Knock-In Plasmid
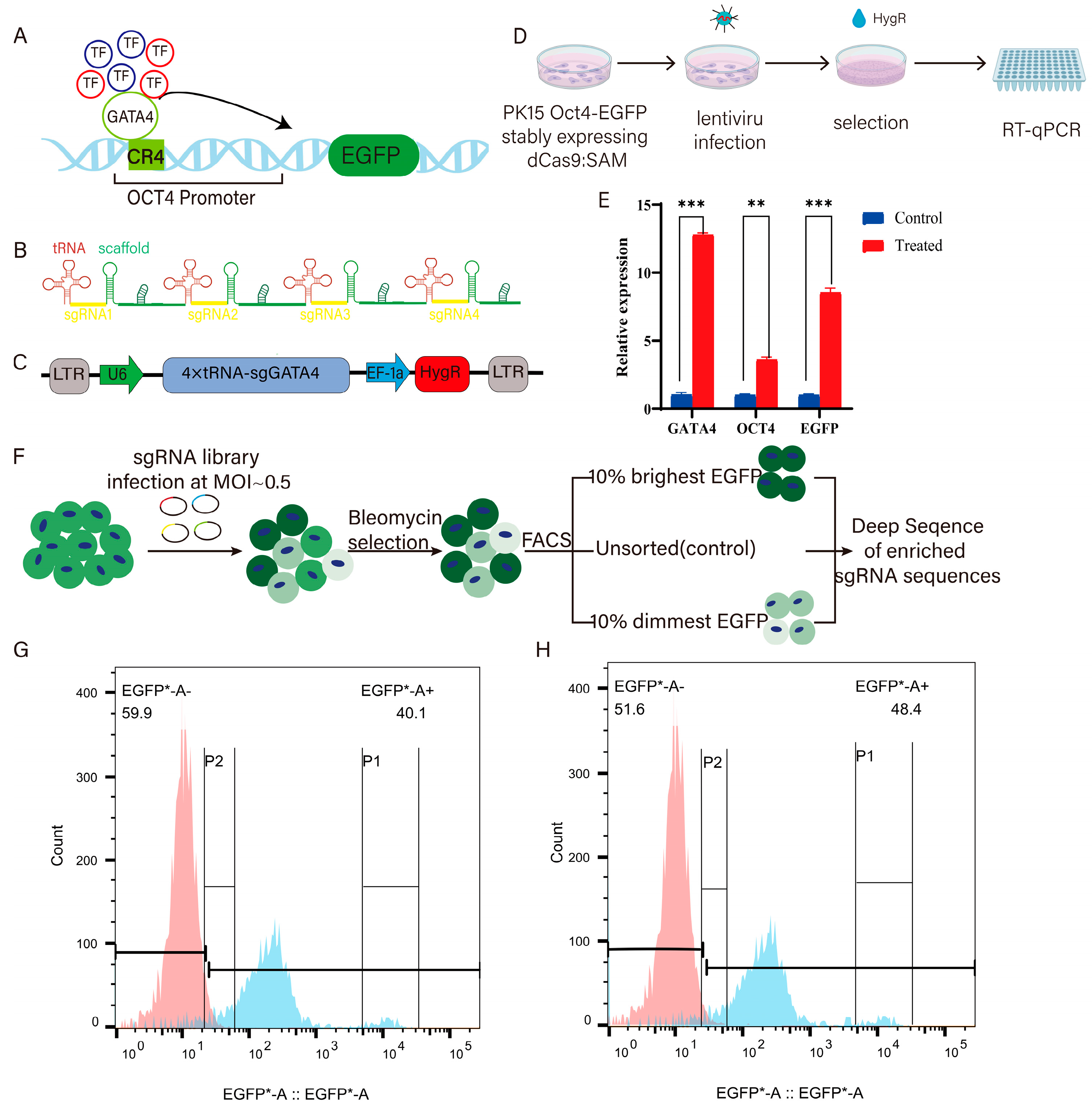
2.2.2. Construction of pLenti 4×tRNA-sgGATA4 Plasmid
2.2.3. Validation of sgRNA Expression Vectors
2.3. Establishment and Verification of EGFP Reporter Cell Line
2.4. Establishment of GATA4 Activating Expression Cell Lines
2.5. sgRNA Library Infection
2.6. Validation of Candidate Transcription Factors at the Transcriptional Level
2.7. Data Analysis
3. Results
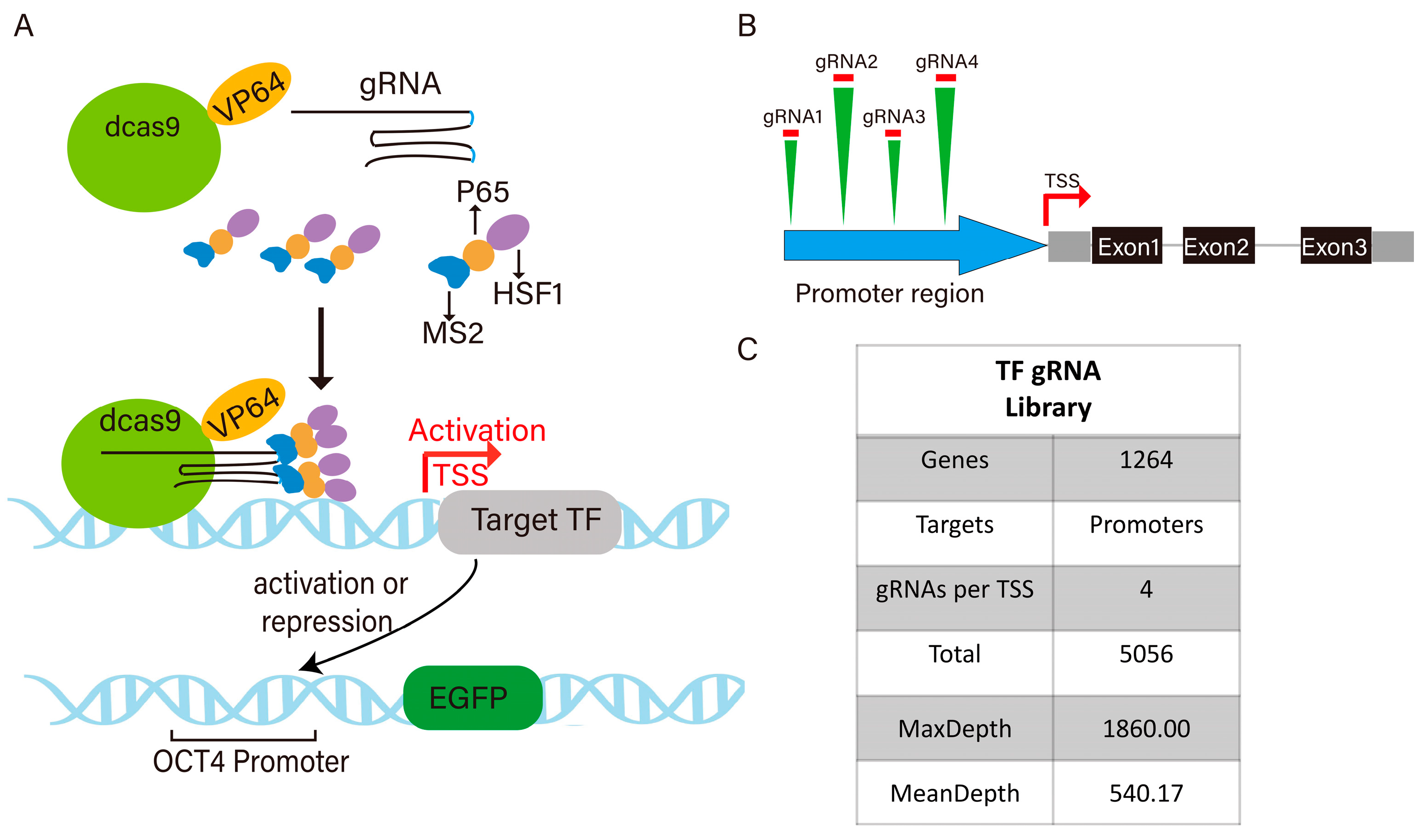
3.1. Construction of Pigs Transcription Factor sgRNA Library and Pigs Cell Line Expressing pOCT4-EGFP and dCas9 SAM System
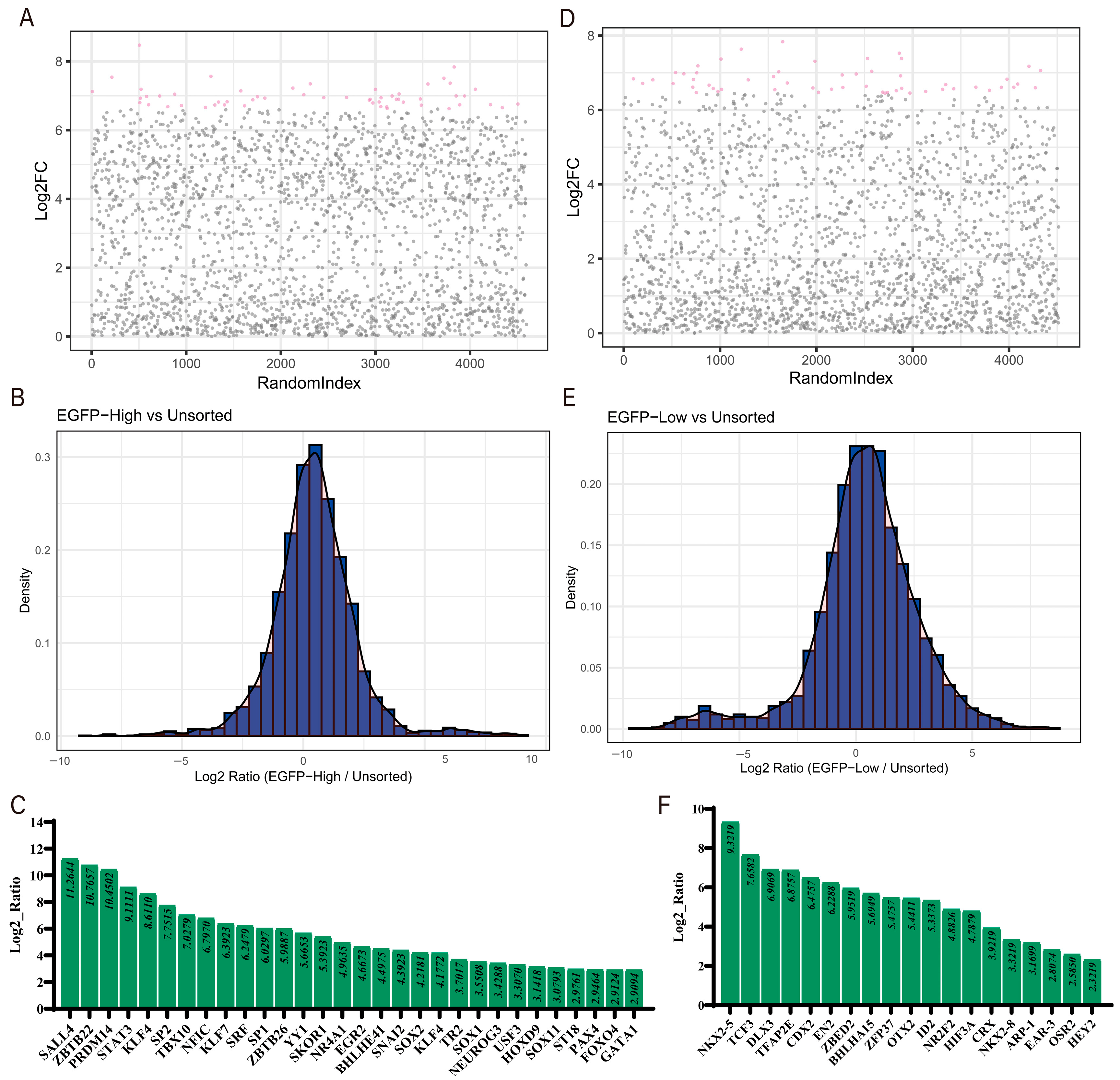
3.2. Acquisition of GATA4-Activated Cells and Screening with sgRNA Library
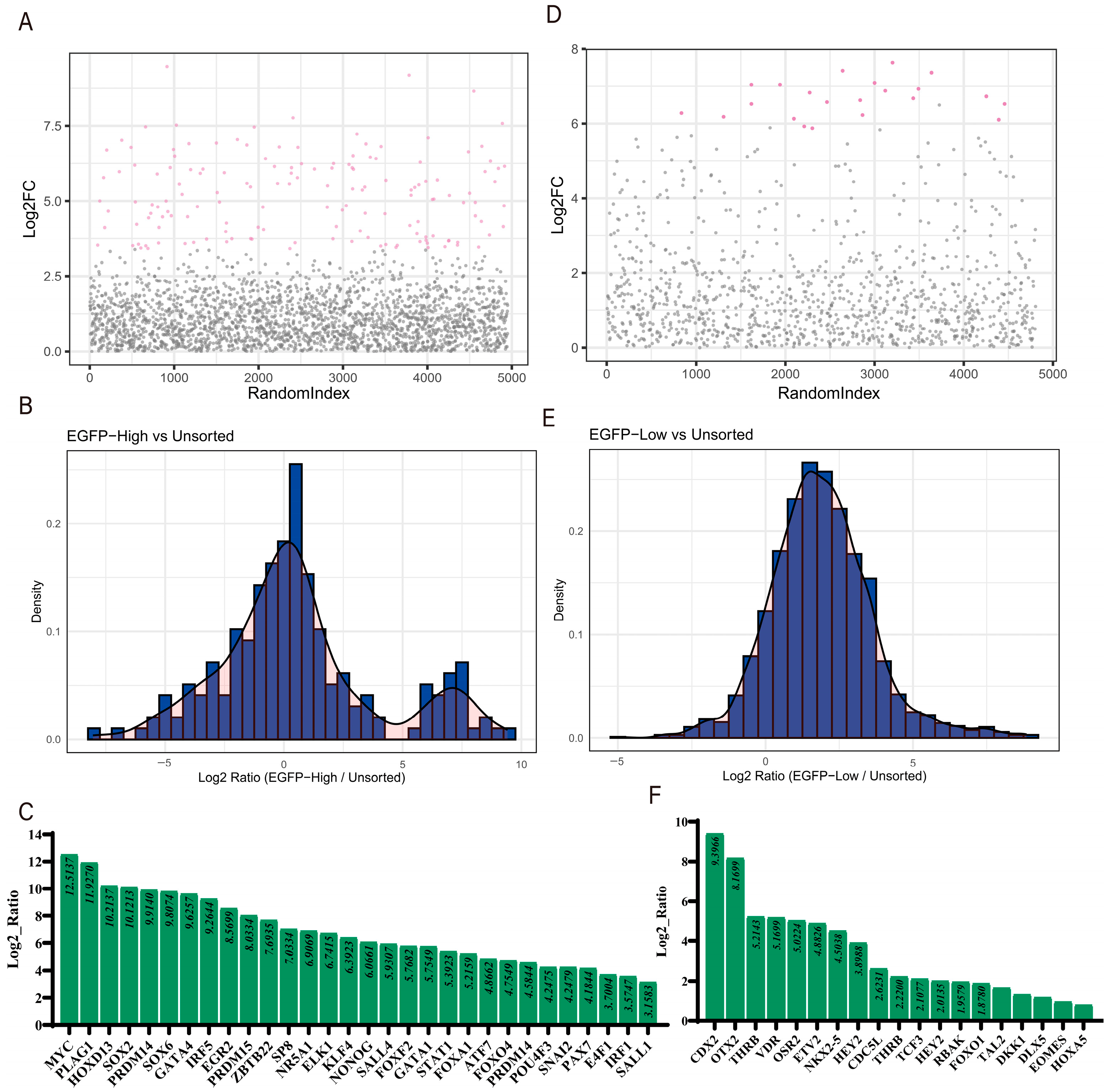
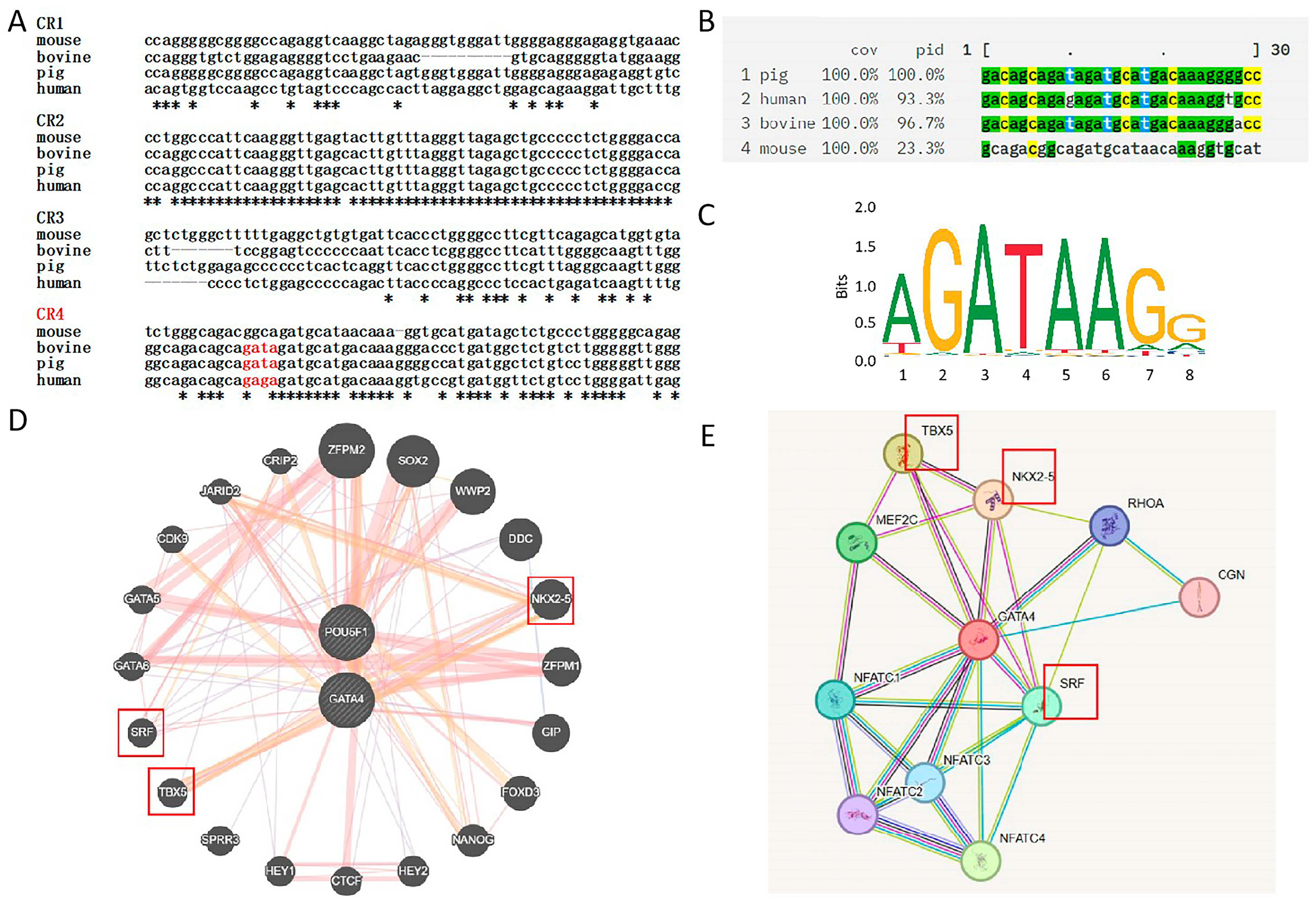
3.3. Data Analysis and Identification of Co-Regulated Genes
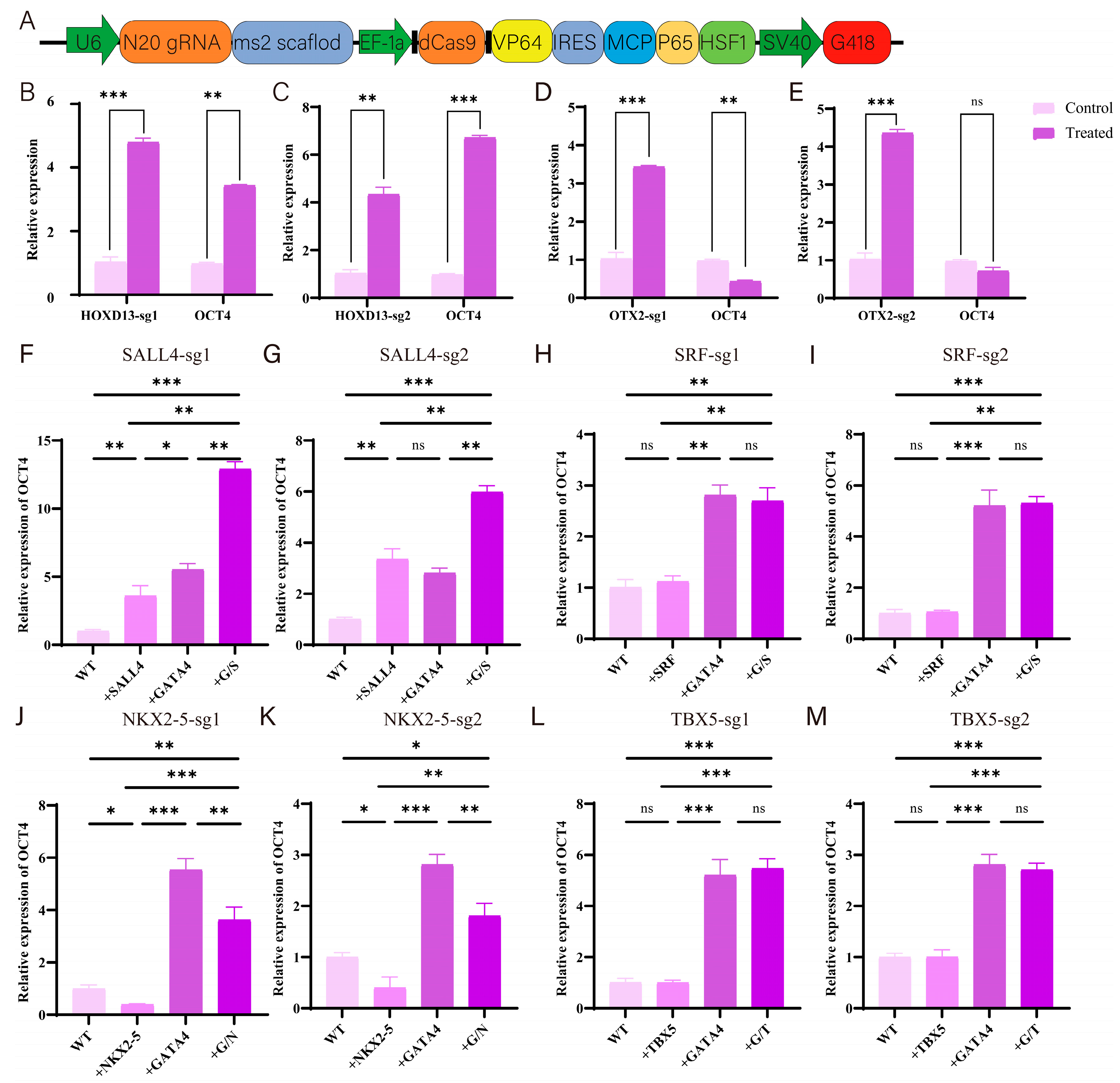
3.4. Experimental Validation of Gene Screening
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zang, X.; Gu, T.; Wang, W.; Zhou, C.; Ding, Y.; Gu, S.; Xu, Z.; Xie, Y.; Li, Z.; Cai, G. Integrated insight into the molecular mechanisms of spontaneous abortion during early pregnancy in pigs. Int. J. Mol. Sci. 2021, 22, 6644. [Google Scholar] [CrossRef]
- Oh, D.; Choi, H.; Kim, M.; Jawad, A.; Lee, J.; Oh, B.C.; Hyun, S.-H. Interleukin-7 promotes porcine early embryogenesis in vitro and inner cell mass development through PI3K/AKT pathway after parthenogenetic activation. Sci. Rep. 2025, 15, 13850. [Google Scholar] [CrossRef]
- Whyte, J.J.; Prather, R.S. Genetic modifications of pigs for medicine and agriculture. Mol. Reprod. Dev. 2011, 78, 879–891. [Google Scholar] [CrossRef]
- Nichols, J.; Smith, A. Pluripotency in the embryo and in culture. Cold Spring Harb. Perspect. Biol. 2012, 4, a008128. [Google Scholar] [CrossRef] [PubMed]
- Boyer, L.A.; Lee, T.I.; Cole, M.F.; Johnstone, S.E.; Levine, S.S.; Zucker, J.P.; Guenther, M.G.; Kumar, R.M.; Murray, H.L.; Jenner, R.G. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 2005, 122, 947–956. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006, 126, 663–676. [Google Scholar] [CrossRef] [PubMed]
- Giorgetti, A.; Montserrat, N.; Aasen, T.; Gonzalez, F.; Rodríguez-Pizà, I.; Vassena, R.; Raya, A.; Boué, S.; Barrero, M.J.; Corbella, B.A.A. Generation of induced pluripotent stem cells from human cord blood using OCT4 and SOX2. Cell Stem Cell 2009, 5, 353–357. [Google Scholar] [CrossRef]
- Le Bin, G.C.; Muñoz-Descalzo, S.; Kurowski, A.; Leitch, H.; Lou, X.; Mansfield, W.; Etienne-Dumeau, C.; Grabole, N.; Mulas, C.; Niwa, H. Oct4 is required for lineage priming in the developing inner cell mass of the mouse blastocyst. Development 2014, 141, 1001–1010. [Google Scholar] [CrossRef]
- Wu, G.; Schöler, H.R. Role of Oct4 in the early embryo development. Cell Regen. 2014, 3, 7. [Google Scholar] [CrossRef]
- Okumura-Nakanishi, S.; Saito, M.; Niwa, H.; Ishikawa, F. Oct-3/4 and Sox2 Regulate Oct-3/4 Gene in Embryonic Stem Cells. J. Biol. Chem. 2005, 280, 5307–5317. [Google Scholar] [CrossRef]
- Bou, G.; Guo, J.; Liu, S.; Guo, S.; Davaakhuu, G.; Lv, Q.; Xue, B.; Qiao, S.; Lv, J.; Weng, X. OCT4 expression transactivated by GATA protein is essential for non-rodent trophectoderm early development. Cell Rep. 2022, 41, 111644. [Google Scholar] [CrossRef] [PubMed]
- Molkentin, J.D. The Zinc Finger-containing Transcription Factors GATA-4, -5, and -6 Ubiquitously Expressed Regulators of Tissue-Specific Gene Expression. J. Biol. Chem. 2000, 275, 38949–38952. [Google Scholar] [CrossRef]
- Masui, S.; Nakatake, Y.; Toyooka, Y.; Shimosato, D.; Yagi, R.; Takahashi, K.; Okochi, H.; Okuda, A.; Matoba, R.; Sharov, A.A. Pluripotency governed by Sox2 via regulation of Oct3/4 expression in mouse embryonic stem cells. Nat. Cell Biol. 2007, 9, 625–635. [Google Scholar] [CrossRef]
- Chakravarthy, H.; Boer, B.; Desler, M.; Mallanna, S.K.; Rizzino, A. Identification of DPPA4 and other genes as putative Sox2: Oct-3/4 target genes using a combination of in silico analysis and transcription-based assays. J. Cell. Physiol. 2010, 216, 651–662. [Google Scholar] [CrossRef]
- Liu, S.; Bou, G.; Sun, R.; Guo, S.; Xue, B.; Wei, R.; Cooney, A.J.; Liu, Z. Sox2 is the faithful marker for pluripotency in pig: Evidence from embryonic studies. Dev. Dyn. 2015, 244, 619–627. [Google Scholar] [CrossRef]
- Gilbert, L.A.; Horlbeck, M.A.; Adamson, B.; Villalta, J.E.; Chen, Y.; Whitehead, E.H.; Guimaraes, C.; Panning, B.; Ploegh, H.L.; Bassik, M.C. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 2014, 159, 647–661. [Google Scholar] [CrossRef]
- Yamagata, T.; Raveau, M.; Kobayashi, K.; Miyamoto, H.; Tatsukawa, T.; Ogiwara, I.; Itohara, S.; Hensch, T.K.; Yamakawa, K. CRISPR/dCas9-based Scn1a gene activation in inhibitory neurons ameliorates epileptic and behavioral phenotypes of Dravet syndrome model mice. Neurobiol. Dis. 2020, 141, 104954. [Google Scholar] [CrossRef]
- Cai, R.; Lv, R.; Shi, X.e.; Yang, G.; Jin, J. CRISPR/dCas9 tools: Epigenetic mechanism and application in gene transcriptional regulation. Int. J. Mol. Sci. 2023, 24, 14865. [Google Scholar] [CrossRef]
- Balboa, D.; Weltner, J.; Eurola, S.; Trokovic, R.; Wartiovaara, K.; Otonkoski, T. Conditionally Stabilized dCas9 Activator for Controlling Gene Expression in Human Cell Reprogramming and Differentiation. Stem Cell Rep. 2015, 5, 448–459. [Google Scholar] [CrossRef] [PubMed]
- Zhou1, Y.; Zhu1, S.; Cai, C.; Yuan, P.; Li, C.; Huang, Y.; Wei, W. High-throughput screening of a CRISPR/Cas9 library for functional genomics in human cells. Nature 2014, 509, 487–491. [Google Scholar] [CrossRef] [PubMed]
- Zalatan, J.G.; Lee, M.E.; Almeida, R.; Gilbert, L.A.; Whitehead, E.H.; La Russa, M.; Tsai, J.C.; Weissman, J.S.; Dueber, J.E.; Qi, L.S. Engineering complex synthetic transcriptional programs with CRISPR RNA scaffolds. Cell 2015, 160, 339–350. [Google Scholar] [CrossRef]
- Barber, H.M.; Pater, A.A.; Gagnon, K.T.; Damha, M.J.; O’Reilly, D. Chemical engineering of CRISPR–Cas systems for therapeutic application. Nat. Rev. Drug Discov. 2024, 24, 209–230. [Google Scholar] [CrossRef]
- Tanenbaum, M.E.; Gilbert, L.A.; Qi, L.S.; Weissman, J.S.; Vale, R.D. A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell 2014, 159, 635–646. [Google Scholar] [CrossRef]
- Dominguez, A.A.; Chavez, M.G.; Urke, A.; Gao, Y.; Wang, L.; Qi, L.S. CRISPR-Mediated Synergistic Epigenetic and Transcriptional Control. Cris. J. 2022, 5, 264–275. [Google Scholar] [CrossRef]
- Ding, X.; Yu, L.; Chen, L.; Li, Y.; Zhang, J.; Sheng, H.; Ren, Z.; Li, Y.; Yu, X.; Jin, S. Recent progress and future prospect of CRISPR/Cas-derived transcription activation (CRISPRa) system in plants. Cells 2022, 11, 3045. [Google Scholar] [CrossRef]
- Li, K.; Liu, Y.; Cao, H.; Zhang, Y.; Gu, Z.; Liu, X.; Yu, A.; Kaphle, P.; Dickerson, K.E.; Ni, M. Interrogation of enhancer function by enhancer-targeting CRISPR epigenetic editing. Nat. Commun. 2020, 11, 485. [Google Scholar] [CrossRef]
- Chavez, A.; Tuttle, M.; Pruitt, B.W.; Ewen-Campen, B.; Chari, R.; Ter-Ovanesyan, D.; Haque, S.J.; Cecchi, R.J.; Kowal, E.J.; Buchthal, J. Comparison of Cas9 activators in multiple species. Nat. Methods 2016, 13, 563–567. [Google Scholar] [CrossRef] [PubMed]
- Jiang, J.; Sun, Y.; Xiao, R.; Wai, K.; Ahmad, M.J.; Khan, F.A.; Zhou, H.; Li, Z.; Zhang, Y.; Zhou, A. Porcine antiviral activity is increased by CRISPRa-SAM system. Biosci. Rep. 2019, 39, BSR20191496. [Google Scholar] [CrossRef] [PubMed]
- Didovyk, A.; Borek, B.; Tsimring, L.; Hasty, J. Transcriptional regulation with CRISPR-Cas9: Principles, advances, and applications. Curr. Opin. Biotechnol. 2016, 40, 177–184. [Google Scholar] [CrossRef]
- Kim, S.-H.; Choi, K.-H.; Lee, D.-K.; Lee, M.; Cho, M.-H.; Oh, J.-N.; Lee, C.-K. Functional analysis of porcine OCT4 transcriptional regulatory region-based reporter system. Reprod. Fertil. Dev. 2019, 31, 187–188. [Google Scholar] [CrossRef]
- Xie, Z.; Pang, D.; Wang, K.; Li, M.; Guo, N.; Yuan, H.; Li, J.; Zou, X.; Jiao, H.; Ouyang, H. Optimization of a CRISPR/Cas9-mediated Knock-in Strategy at the Porcine Rosa26 Locus in Porcine Foetal Fibroblasts. Sci. Rep. 2017, 7, 3036. [Google Scholar] [CrossRef]
- Kong, Q.; Hai, T.; Ma, J.; Huang, T.; Liu, Z. Rosa26 Locus Supports Tissue-Specific Promoter Driving Transgene Expression Specifically in Pig. PLoS ONE 2014, 9, e107945. [Google Scholar] [CrossRef]
- Li, Y.Q. Networks of Transcription Factors for Oct4 Expression in Mice. Dna Cell Biol. 2017, 36, 725. [Google Scholar] [CrossRef] [PubMed]
- Ko, L.J.; Engel, J.D. DNA-binding specificities of the GATA transcription factor family. Mol. Cell. Biol. 1993, 13, 4011–4022. [Google Scholar] [CrossRef] [PubMed]
- Merika, M.; Orkin, S.H. DNA-binding specificity of GATA family transcription factors. Mol. Cell. Biol. 1993, 13, 3999–4010. [Google Scholar] [CrossRef] [PubMed]
- Pu, W.T.; Ishiwata, T.; Juraszek, A.L.; Ma, Q.; Izumo, S. GATA4 is a dosage-sensitive regulator of cardiac morphogenesis. Dev. Biol. 2004, 275, 235–244. [Google Scholar] [CrossRef] [PubMed]
- Romano, O.; Miccio, A. GATA factor transcriptional activity: Insights from genome-wide binding profiles. IUBMB Life 2020, 72, 10–26. [Google Scholar] [CrossRef]
- Takaya, T.; Kawamura, T.; Morimoto, T.; Ono, K.; Kita, T.; Shimatsu, A.; Hasegawa, K. Identification of p300-targeted acetylated residues in GATA4 during hypertrophic responses in cardiac myocytes. J. Biol. Chem. 2008, 13, S28. [Google Scholar] [CrossRef]
- Chlon, T.M.; Crispino, J.D. Combinatorial regulation of tissue specification by GATA and FOG factors. Development 2012, 139, 3905–3916. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yao, X.; Feng, M.; Sun, C.; Yang, S.; Yuan, Z.; Liu, X.; Li, Q.; Jiang, C.; Weng, X.; Song, J.; et al. Establishment of a CRISPR/dCas9 Activation Library for Screening Transcription Factors Co-Regulating OCT4 with GATA4 in Pig Cells. Cells 2025, 14, 1330. https://doi.org/10.3390/cells14171330
Yao X, Feng M, Sun C, Yang S, Yuan Z, Liu X, Li Q, Jiang C, Weng X, Song J, et al. Establishment of a CRISPR/dCas9 Activation Library for Screening Transcription Factors Co-Regulating OCT4 with GATA4 in Pig Cells. Cells. 2025; 14(17):1330. https://doi.org/10.3390/cells14171330
Chicago/Turabian StyleYao, Xiaoxia, Mingjie Feng, Chengbo Sun, Sijia Yang, Zhongyu Yuan, Xueqing Liu, Qinjian Li, Chaoqian Jiang, Xiaogang Weng, Jun Song, and et al. 2025. "Establishment of a CRISPR/dCas9 Activation Library for Screening Transcription Factors Co-Regulating OCT4 with GATA4 in Pig Cells" Cells 14, no. 17: 1330. https://doi.org/10.3390/cells14171330
APA StyleYao, X., Feng, M., Sun, C., Yang, S., Yuan, Z., Liu, X., Li, Q., Jiang, C., Weng, X., Song, J., & Mu, Y. (2025). Establishment of a CRISPR/dCas9 Activation Library for Screening Transcription Factors Co-Regulating OCT4 with GATA4 in Pig Cells. Cells, 14(17), 1330. https://doi.org/10.3390/cells14171330




