Foxd4l1.1 Negatively Regulates Chordin Transcription in Neuroectoderm of Xenopus Gastrula
Abstract
:1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Cloning of Chrd Genomic DNA
2.3. DNA and RNA Preparation
2.4. Promoter Constructs
2.5. Embryo Injection and Explants Culture
2.6. RT-PCR
2.7. Luciferase Assays
2.8. Smad2- and Smad3-Truncated Protein Construct
2.9. Co-Immunoprecipitation and Western Blotting
2.10. Site-Directed Mutagenesis
2.11. Chromatin Immunoprecipitation
2.12. ChIP-Sequencing Analysis
2.13. eGFP Fluorescence
2.14. Statistical Analysis
3. Results
3.1. Foxd4l1.1 Inhibits Chrd and Dorsal Target Gene Expression
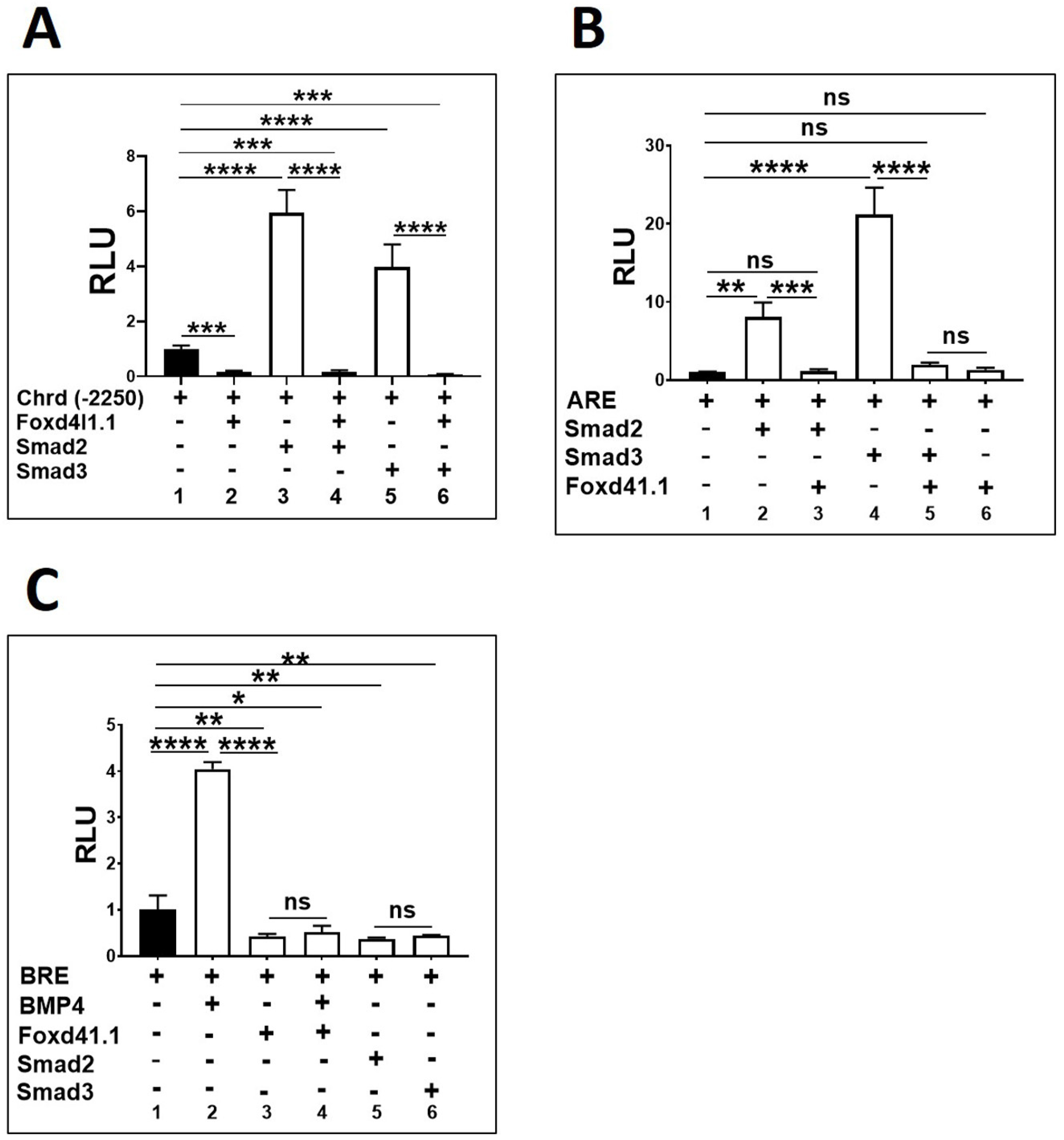
3.2. Foxd4l1.1 Abolishes Mesoderm and Ectoderm Specific Reporter Genes Expression
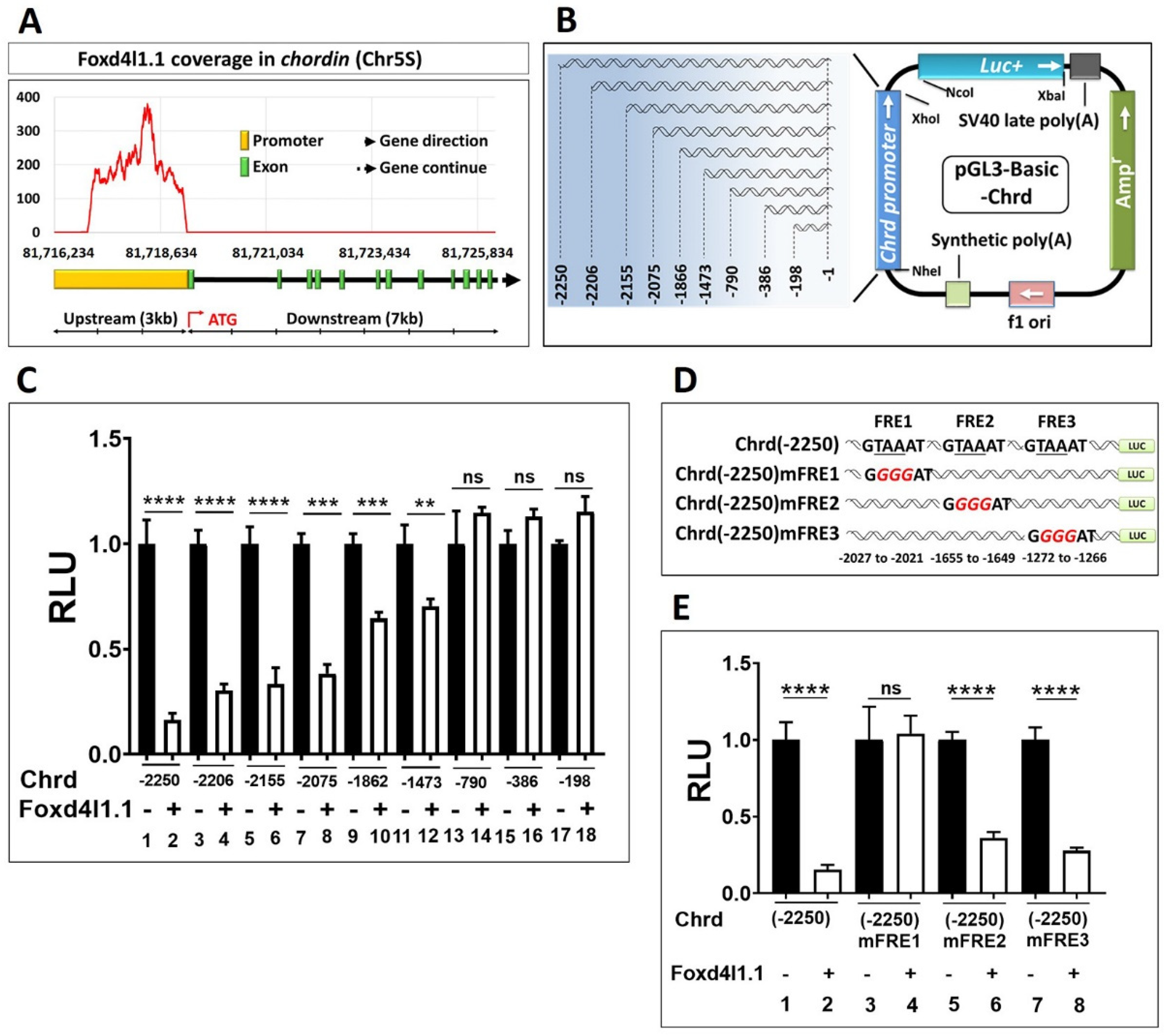
3.3. Chrd Promoter Contains Foxd4l1.1 Response Element (FRE)
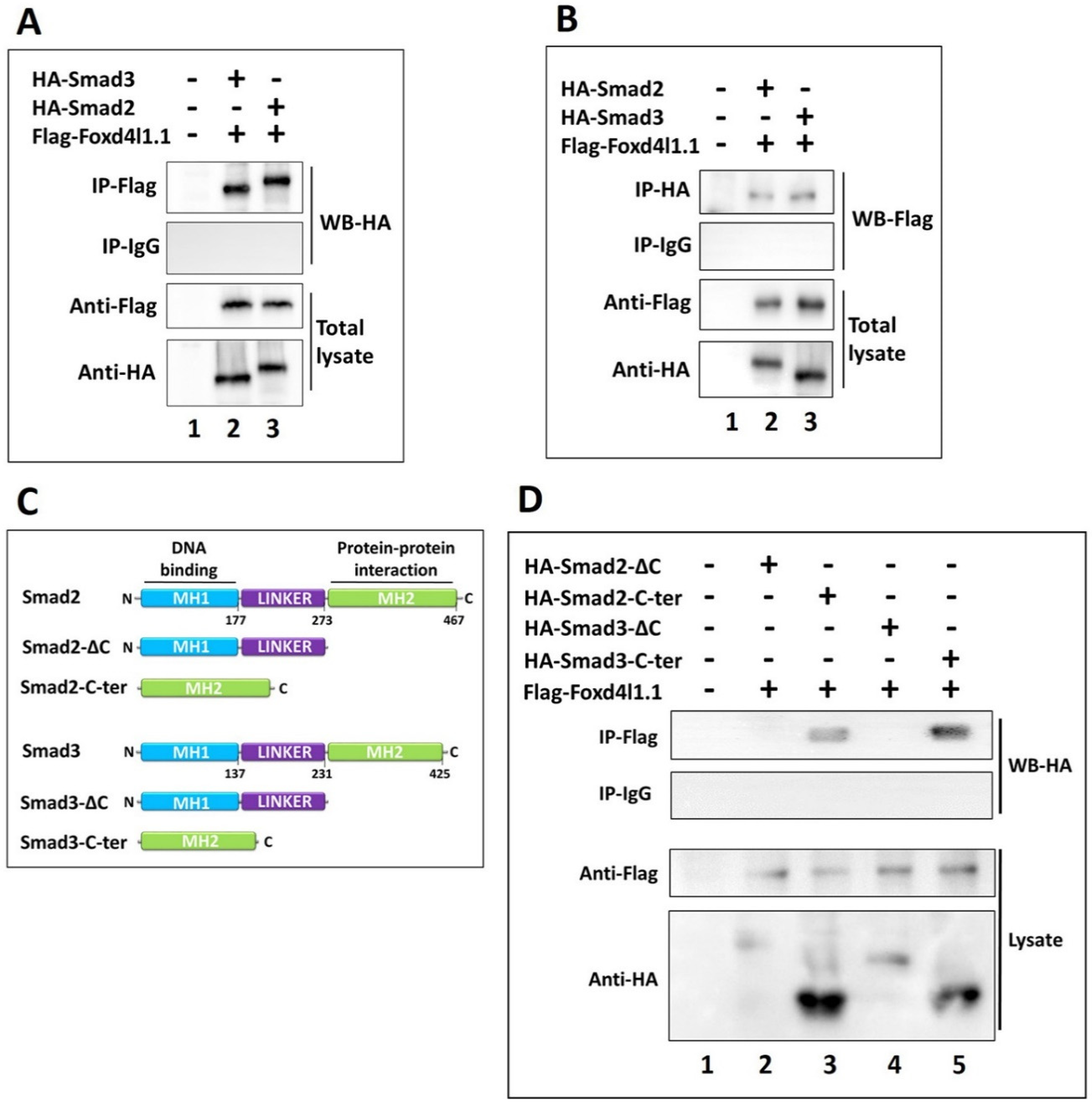
3.4. Foxd4l1.1 Physically Interacts with Smad2/3 via C-Ternimal MH2 Domain
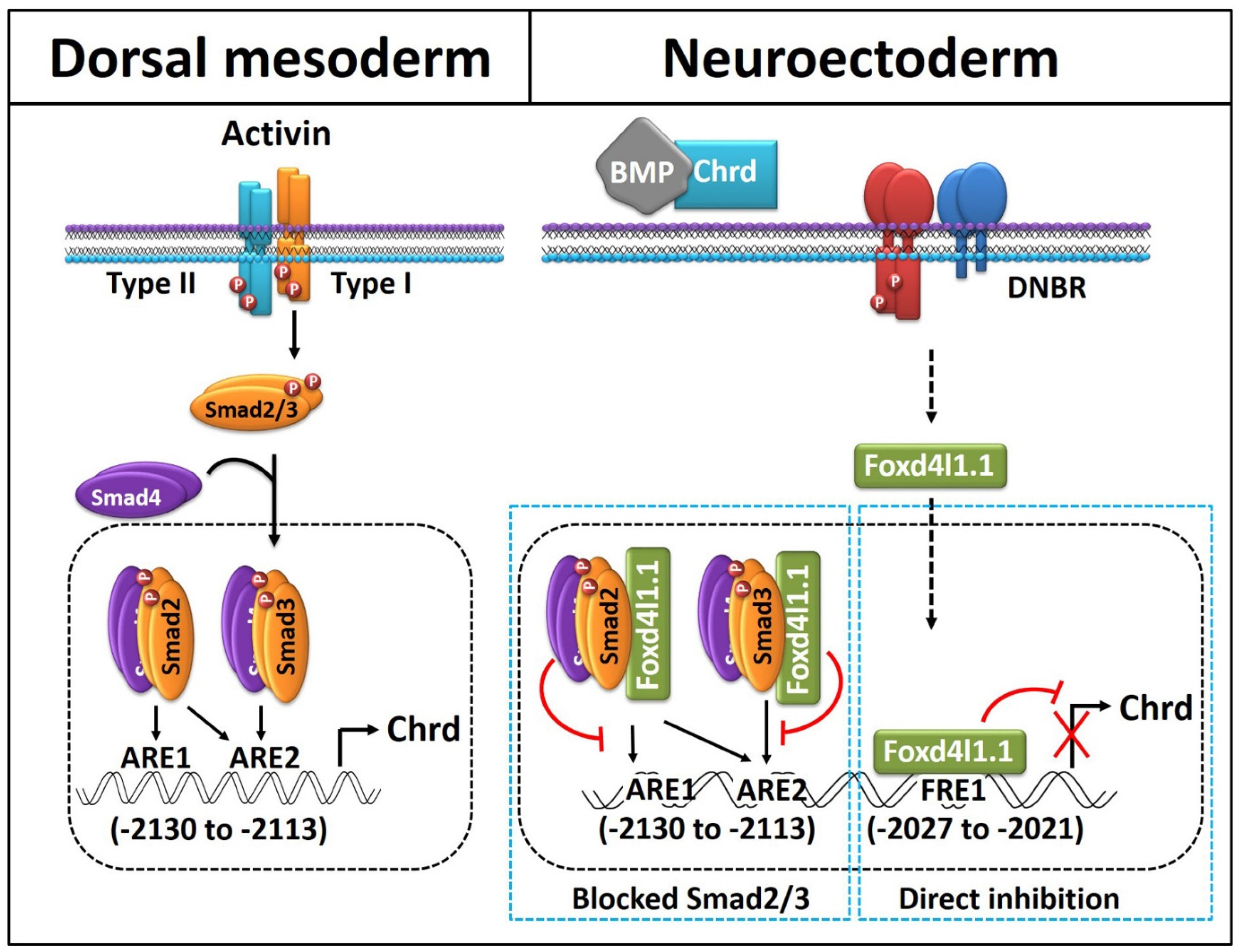
3.5. Foxd4l1.1 Binds to FRE1 and Blocks Smad2/3 Binding to AREs (ARE1 and ARE2) within the Chrd Promoter
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wessely, O.; Kim, J.I.; Geissert, D.; Tran, U.; De Robertis, E.M. Analysis of Spemann organizer formation in Xenopus embryos by cDNA macroarrays. Dev. Biol. 2004, 269, 552–566. [Google Scholar] [CrossRef] [Green Version]
- Cho, K.W.; Blumberg, B.; Steinbeisser, H.; De Robertis, E.M. Molecular nature of Spemann’s organizer: The role of the Xenopus homeobox gene goosecoid. Cell 1991, 67, 1111–1120. [Google Scholar] [CrossRef]
- Kumar, S.; Umair, Z.; Yoon, J.; Lee, U.; Kim, S.C.; Park, J.B.; Lee, J.Y.; Kim, J. Xbra and Smad-1 cooperate to activate the transcription of neural repressor ventx1.1 in Xenopus embryos. Sci. Rep. 2018, 8, 11391. [Google Scholar] [CrossRef] [Green Version]
- Hemmati-Brivanlou, A.; Thomsen, G.H. Ventral mesodermal patterning in Xenopus embryos: Expression patterns and activities of BMP-2 and BMP-4. Dev. Genet. 1995, 17, 78–89. [Google Scholar] [CrossRef] [PubMed]
- Lim, D.A.; Tramontin, A.D.; Trevejo, J.M.; Herrera, D.G.; Garcia-Verdugo, J.M.; Alvarez-Buylla, A. Noggin antagonizes BMP signaling to create a niche for adult neurogenesis. Neuron 2000, 28, 713–726. [Google Scholar] [CrossRef] [Green Version]
- Piccolo, S.; Sasai, Y.; Lu, B.; De Robertis, E.M. Dorsoventral patterning in Xenopus: Inhibition of ventral signals by direct binding of chordin to BMP-4. Cell 1996, 86, 589–598. [Google Scholar] [CrossRef] [Green Version]
- Umair, Z.; Kumar, S.; Kim, D.H.; Rafiq, K.; Kumar, V.; Kim, S.; Park, J.B.; Lee, J.Y.; Lee, U.; Kim, J. Ventx1.1 as a Direct Repressor of Early Neural Gene zic3 in Xenopus laevis. Mol. Cells 2018, 41, 1061–1071. [Google Scholar] [CrossRef]
- Hwang, Y.S.; Lee, H.S.; Roh, D.H.; Cha, S.; Lee, S.Y.; Seo, J.J.; Kim, J.; Park, M.J. Active repression of organizer genes by C-terminal domain of PV.1. Biochem. Biophys. Res. Commun. 2003, 308, 79–86. [Google Scholar] [CrossRef]
- Hwang, Y.S.; Seo, J.J.; Cha, S.W.; Lee, H.S.; Lee, S.Y.; Roh, D.H.; Kung Hf, H.F.; Kim, J.; Ja Park, M. Antimorphic PV.1 causes secondary axis by inducing ectopic organizer. Biochem. Biophys. Res. Commun. 2002, 292, 1081–1086. [Google Scholar] [CrossRef] [PubMed]
- Ault, K.T.; Dirksen, M.L.; Jamrich, M. A novel homeobox gene PV.1 mediates induction of ventral mesoderm in Xenopus embryos. Proc. Natl. Acad. Sci. USA 1996, 93, 6415–6420. [Google Scholar] [CrossRef] [Green Version]
- Yan, B.; Neilson, K.M.; Moody, S.A. foxD5 plays a critical upstream role in regulating neural ectodermal fate and the onset of neural differentiation. Dev. Biol. 2009, 329, 80–95. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sullivan, S.A.; Akers, L.; Moody, S.A. foxD5a, a Xenopus winged helix gene, maintains an immature neural ectoderm via transcriptional repression that is dependent on the C-terminal domain. Dev. Biol. 2001, 232, 439–457. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kumar, S.; Umair, Z.; Kumar, V.; Kumar, S.; Lee, U.; Kim, J. Foxd4l1.1 negatively regulates transcription of neural repressor ventx1.1 during neuroectoderm formation in Xenopus embryos. Sci. Rep. 2020, 10, 16780. [Google Scholar] [CrossRef] [PubMed]
- Yoon, J.; Kim, J.H.; Kim, S.C.; Park, J.B.; Lee, J.Y.; Kim, J. PV. 1 suppresses the expression of FoxD5b during neural induction in Xenopus embryos. Mol. Cells. 2014, 37, 220–225. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yoon, J.; Kim, J.H.; Lee, O.J.; Lee, S.Y.; Lee, S.H.; Park, J.B.; Lee, J.Y.; Kim, S.C.; Kim, J. AP-1(c-Jun/FosB) mediates xFoxD5b expression in Xenopus early developmental neurogenesis. Int. J. Dev. Biol. 2013, 57, 865–872. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rogers, C.D.; Moody, S.A.; Casey, E.S. Neural induction and factors that stabilize a neural fate. Birth Defects Res. Part C Embryo Today Rev. 2009, 87, 249–262. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Robertis, E.M.; Kuroda, H. Dorsal-ventral patterning and neural induction in Xenopus embryos. Annu. Rev. Cell Dev. Biol. 2004, 20, 285–308. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mizuseki, K.; Kishi, M.; Matsui, M.; Nakanishi, S.; Sasai, Y. Xenopus Zic-related-1 and Sox-2, two factors induced by chordin, have distinct activities in the initiation of neural induction. Development 1998, 125, 579–587. [Google Scholar] [CrossRef]
- Kumar, V.; Umair, Z.; Kumar, S.; Lee, U.; Kim, J. Smad2 and Smad3 differentially modulate chordin transcription via direct binding on the distal elements in gastrula Xenopus embryos. Biochem. Biophys. Res. Commun. 2021, 559, 168–175. [Google Scholar] [CrossRef]
- Lee, H.S.; Lee, S.Y.; Lee, H.; Hwang, Y.S.; Cha, S.W.; Park, S.; Lee, J.Y.; Park, J.B.; Kim, S.; Park, M.J.; et al. Direct response elements of BMP within the PV.1A promoter are essential for its transcriptional regulation during early Xenopus development. PLoS ONE 2011, 6, e22621. [Google Scholar] [CrossRef] [Green Version]
- Lee, S.Y.; Lee, H.S.; Moon, J.S.; Kim, J.I.; Park, J.B.; Lee, J.Y.; Park, M.J.; Kim, J. Transcriptional regulation of Zic3 by heterodimeric AP-1(c-Jun/c-Fos) during Xenopus development. Exp. Mol. Med. 2004, 36, 468–475. [Google Scholar] [CrossRef]
- Kumar, S.; Umair, Z.; Kumar, V.; Lee, U.; Choi, S.C.; Kim, J. Ventx1.1 competes with a transcriptional activator Xcad2 to regulate negatively its own expression. BMB Rep. 2019, 52, 403–408. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Umair, Z.; Kumar, S.; Rafiq, K.; Kumar, V.; Reman, Z.U.; Lee, S.H.; Kim, S.; Lee, J.Y.; Lee, U.; Kim, J. Dusp1 modulates activin/smad2 mediated germ layer specification via FGF signal inhibition in Xenopus embryos. Anim. Cells Syst. 2020, 24, 359–370. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Feng, X.; We, R.; Derynck, R. Receptor-associated Mad homologues synergize as effectors of the TGF-beta response. Nature 1996, 383, 168–172. [Google Scholar] [CrossRef] [PubMed]
- Blythe, S.A.; Reid, C.D.; Kessler, D.S.; Klein, P.S. Chromatin immunoprecipitation in early Xenopus laevis embryos. Dev. Dyn. 2009, 238, 1422–1432. [Google Scholar] [CrossRef] [Green Version]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R.; Genome Project Data Processing, S. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.; Liu, T.; Meyer, C.A.; Eeckhoute, J.; Johnson, D.S.; Bernstein, B.E.; Nusbaum, C.; Myers, R.M.; Brown, M.; Li, W.; et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 2008, 9, R137. [Google Scholar] [CrossRef] [Green Version]
- Li, J.; Dai, S.; Chen, X.; Liang, X.; Qu, L.; Jiang, L.; Guo, M.; Zhou, Z.; Wei, H.; Zhang, H.; et al. Mechanism of forkhead transcription factors binding to a novel palindromic DNA site. Nucleic Acids Res. 2021, 49, 3573–3583. [Google Scholar] [CrossRef] [PubMed]
- Nakagawa, S.; Gisselbrecht, S.S.; Rogers, J.M.; Hartl, D.L.; Bulyk, M.L. DNA-binding specificity changes in the evolution of forkhead transcription factors. Proc. Natl. Acad. Sci. USA 2013, 110, 12349–12354. [Google Scholar] [CrossRef] [Green Version]
- Schiffer, M.; von Gersdorff, G.; Bitzer, M.; Susztak, K.; Bottinger, E.P. Smad proteins and transforming growth factor-beta signaling. Kidney Int. 2000, 77, S45–S52. [Google Scholar] [CrossRef] [Green Version]
- Pohl, B.S.; Knochel, W. Of Fox and Frogs: Fox (fork head/winged helix) transcription factors in Xenopus development. Gene 2005, 344, 21–32. [Google Scholar] [CrossRef] [PubMed]
- Troilo, H.; Zuk, A.V.; Tunnicliffe, R.B.; Wohl, A.P.; Berry, R.; Collins, R.F.; Jowitt, T.A.; Sengle, G.; Baldock, C. Nanoscale structure of the BMP antagonist chordin supports cooperative BMP binding. Proc. Natl. Acad. Sci. USA 2014, 111, 13063–13068. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, A.; Niswander, L.A. Bone morphogenetic protein signalling and vertebrate nervous system development. Nat. Rev. Neurosci. 2005, 6, 945–954. [Google Scholar] [CrossRef]
- Balemans, W.; Van Hul, W. Extracellular regulation of BMP signaling in vertebrates: A cocktail of modulators. Dev. Biol 2002, 250, 231–250. [Google Scholar] [CrossRef] [PubMed]
- Larrain, J.; Bachiller, D.; Lu, B.; Agius, E.; Piccolo, S.; De Robertis, E.M. BMP-binding modules in chordin: A model for signalling regulation in the extracellular space. Development 2000, 127, 821–830. [Google Scholar] [CrossRef]
- Fetka, I.; Doederlein, G.; Bouwmeester, T. Neuroectodermal specification and regionalization of the Spemann organizer in Xenopus. Mech. Dev. 2000, 93, 49–58. [Google Scholar] [CrossRef]
- Christian, J.L.; Moon, R.T. Interactions between Xwnt-8 and Spemann organizer signaling pathways generate dorsoventral pattern in the embryonic mesoderm of Xenopus. Genes Dev. 1993, 7, 13–28. [Google Scholar] [CrossRef] [Green Version]
- Moody, S.A.; Klein, S.L.; Karpinski, B.A.; Maynard, T.M.; Lamantia, A.S. On becoming neural: What the embryo can tell us about differentiating neural stem cells. Am. J. Stem. Cells 2013, 2, 74–94. [Google Scholar] [PubMed]
- Hemmati-Brivanlou, A.; Melton, D. Vertebrate embryonic cells will become nerve cells unless told otherwise. Cell 1997, 88, 13–17. [Google Scholar] [CrossRef] [Green Version]
- Wilson, P.A.; Hemmati-Brivanlou, A. Induction of epidermis and inhibition of neural fate by Bmp-4. Nature 1995, 376, 331–333. [Google Scholar] [CrossRef]
- Sasai, Y.; Lu, B.; Steinbeisser, H.; Geissert, D.; Gont, L.K.; De Robertis, E.M. Xenopus chordin: A novel dorsalizing factor activated by organizer-specific homeobox genes. Cell 1994, 79, 779–790. [Google Scholar] [CrossRef]
- Kumar, V.; Goutam, R.S.; Park, S.; Lee, U.; Kim, J. Functional Roles of FGF Signaling in Early Development of Vertebrate Embryos. Cells 2021, 10, 2148. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, C.H.; Majumdar, H.D.; Neilson, K.M.; Moody, S.A. Six1 and Irx1 have reciprocal interactions during cranial placode and otic vesicle formation. Dev. Biol. 2019, 446, 68–79. [Google Scholar] [CrossRef] [PubMed]
- Sander, V.; Reversade, B.; De Robertis, E.M. The opposing homeobox genes Goosecoid and Vent1/2 self-regulate Xenopus patterning. EMBO J. 2007, 26, 2955–2965. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Derynck, R.; Budi, E.H. Specificity, versatility, and control of TGF-beta family signaling. Sci. Signal. 2019, 12. [Google Scholar] [CrossRef] [Green Version]
- Beich-Frandsen, M.; Aragon, E.; Llimargas, M.; Benach, J.; Riera, A.; Pous, J.; Macias, M.J. Structure of the N-terminal domain of the protein Expansion: An ‘Expansion’ to the Smad MH2 fold. Acta Crystallogr. Sect. D Biol. Crystallogr. 2015, 71, 844–853. [Google Scholar] [CrossRef] [Green Version]


| Constructs | Primer Name | Sequence (5′ to 3′) |
|---|---|---|
| Upstream primer | Chrd (-2250)_F | GGGGCTAGCGAACGATACTTCAAGGACAAT |
| Chrd (-2206)_F | GGGGCTAGCCCACTATCCCCACTAAGATGA | |
| Chrd (-2155)_F | GGGGCTAGCAGGCATACTTTGGTTTGTGTGT | |
| Chrd (-2075)_F | GGGGCTAGCTGCAAGTCGAGATCATTGTGT | |
| Chrd (-1862)_F | GGGGCTAGCAAGAACACAGTGCCAGGCACT | |
| Chrd (-1473)_F | GGGGCTAGCCAGTAGGTTAGATGAACTACT | |
| Chrd (-790)_F | GGGGCTAGCACACTCTCTACCCCAATTCT | |
| Chrd (-386)_F | GGGGCTAGCCTTGACGGCTTTGTTTGCTT | |
| Chrd (-198)_F | GGGGCTAGCGTGTGGGTACAGAGCAACAA | |
| Downstream primer | Chrd (-2250)_R | GGGCTCGAGTTTTGTGGTTCCAAACGTTCT |
| Gene | Primer Name | Sequence (5′ to 3′) | Cycles |
|---|---|---|---|
| Chrd | Chrd_F | TTAGAGAGGAGAGCAACTCGGGCAAT | 25 |
| Chrd_R | GTGCTCCTGTTGCGAAACTCTACAGA | ||
| Noggin | Noggin_F | AGTTGCAGATGTGGCTCT | 27 |
| Noggin_R | AGTCCAAGAGTCTGAGCA | ||
| Gsc | Gsc_F | GCTGATTCCACCAGTGCCTCACCAG | 30 |
| Gsc_R | GGTCCTGTGCCTCCTCCTCCTCCTG | ||
| Xbra | Xbra_F | GGATCGTTATCACCTCTG | 25 |
| Xbra_R | GTGTAGTCTGTAGCAGCA | ||
| Ventx1.1 | Ventx1.1_F | CCTTCAGCATGGTTCAACAG | 28 |
| Ventx1.1_R | CATCCTTCTTCCTTGGCATCTCCT | ||
| Bmp4 | BMP4_F | GCATGTACGGATAAGTCGATC | 25 |
| BMP4_R | GATCTCAGACTCAACGGCAC | ||
| ODC | ODC_F | GTCAATGATGGAGTGTATGGATC | 25 |
| ODC_R | TCCATTCCGCTCTCCTGAGCAC |
| Construct | Primer Name | Sequence (5′ to 3′) | Cycles |
|---|---|---|---|
| Smad2-ΔC | hSmad2_(N-terminal)-F | CCCGAATTCCATGTCGTCCATCTTGCCATT | 25 |
| hSmad2_(Linker)-R | GGGCTCGAGTCAAAATGCAGGTTCTGAGT | ||
| Smad2-C-ter | hSmad2_(C-terminal)-F | GGTCTCGAGTTATGACATGCTTGAGCAACGC | 25 |
| hSmad2_(C-terminal)-R | GGTCTCGAGTTATGACATGCTTGAGCAACGC | ||
| Smad3-ΔC | hSmad3_(N-terminal)-F | CCCGAATTCCATGTCGTCCATCCTGCCTTTC | 25 |
| hSmad3_(Linker)-R | AAGTCTAGACCGGCTCGCAGTAGGTAAC | ||
| Smad3-C-ter | hSmad3_(C-terminal)-F | CTTGAATTCCTACTCAGAACCTGCATTTTGG | 25 |
| hSmad3_(C-terminal)-R | CCATCTAGACTAAGACACACTGGAACAGC |
| Mutated Site | Primer Name | Sequence (5′ to 3′) | Cycles |
|---|---|---|---|
| FRE1 | Chrd(−2250)mFRE1_F | TTAAACAGTATAAGGGGATGCTAAAAACACAG | 20 |
| Chrd(−2250)mFRE1_R | CTGTGTTTTTAGCATCCCCTTATACTGTTTAA | ||
| FRE2 | Chrd(−2250)mFRE2_F | GTCCTGGCATATGAGGGGATTCAGAGCTATCCT | 20 |
| Chrd(−2250)mFRE2_R | AGGATAGCTCTGAATCCCCTCATATGCCAGGAC | ||
| FRE3 | Chrd(−2250)mFRE3_F | TTCAATCCTTAGCAGGGAATTCCCTCATCTTTC | 20 |
| Chrd(−2250)mFRE3_R | GAAAGATGAGGGAATTCCCTGCTAAGGATTGAA |
| Primer Name | Sequence (5′ to 3′) | Cycles | |
|---|---|---|---|
| Chrd (Foxd4l1.1_ChIP) | Chrd(FRE)_F | GTTGCTTCTGTTTTCCACCT | 25 |
| Chrd(FRE)_R | GTCTGGCATATCTAGCAGGTC | ||
| Chrd (Internal control 1) | Chrd(C1)_F | TGCGCCGACTAAGTTTCCT | 25(qPCR 40) |
| Chrd(C1)_R | ATTAGTGACCCATGGCAGG | ||
| Chrd (Internal control 2) | Chrd(C2)_F | ACACTCTCTACCCCAATTCT | 25(qPCR 40) |
| Chrd(C2)_R | CAGAATGGCATGTGGGAAGA | ||
| Chrd (Smad2 ChIP) | Chrd(AREs)_F | CGATACTTCAAGGACAATTG | 24 |
| Chrd(AREs)_R | AGGTGGAAAACAGAAGCAAC | ||
| Chrd (Smad3 ChIP) | Chrd(AREs)_F | CGATACTTCAAGGACAATTG | 24 |
| Chrd(AREs)_R | AGGTGGAAAACAGAAGCAAC | ||
| Ventx2.1 (External control) | Ventx2.1_F | CTACAGCACTAGCACTGACT | 28 |
| Ventx2.1_R | AGAAAGCTGGAGTTTGGCTGC |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kumar, V.; Goutam, R.S.; Umair, Z.; Park, S.; Lee, U.; Kim, J. Foxd4l1.1 Negatively Regulates Chordin Transcription in Neuroectoderm of Xenopus Gastrula. Cells 2021, 10, 2779. https://doi.org/10.3390/cells10102779
Kumar V, Goutam RS, Umair Z, Park S, Lee U, Kim J. Foxd4l1.1 Negatively Regulates Chordin Transcription in Neuroectoderm of Xenopus Gastrula. Cells. 2021; 10(10):2779. https://doi.org/10.3390/cells10102779
Chicago/Turabian StyleKumar, Vijay, Ravi Shankar Goutam, Zobia Umair, Soochul Park, Unjoo Lee, and Jaebong Kim. 2021. "Foxd4l1.1 Negatively Regulates Chordin Transcription in Neuroectoderm of Xenopus Gastrula" Cells 10, no. 10: 2779. https://doi.org/10.3390/cells10102779
APA StyleKumar, V., Goutam, R. S., Umair, Z., Park, S., Lee, U., & Kim, J. (2021). Foxd4l1.1 Negatively Regulates Chordin Transcription in Neuroectoderm of Xenopus Gastrula. Cells, 10(10), 2779. https://doi.org/10.3390/cells10102779








