Multi-Level Characterization of Eggplant Accessions from Greek Islands and the Mainland Contributes to the Enhancement and Conservation of this Germplasm and Reveals a Large Diversity and Signatures of Differentiation between both Origins
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material and Cultivation Conditions
2.2. Morphological and Mineral Composition Characterization and Data Analysis
2.3. SSR Characterization and Data Analysis
3. Results
3.1. Morphological Characterization
3.2. Concentration of Minerals
3.3. SSR Characterization
3.4. SSR Molecular Analysis of Variance, Genetic Structure, and Multivariate Analysis
4. Discussion
4.1. Morphological Diversity
4.2. Mineral Composition
4.3. Molecular Diversity
4.4. Comparison of Morphological and Molecular Diversity
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Kougioumoutzis, K.; Valli, A.T.; Georgopoulou, E.; Simaiakis, S.M.; Triantis, K.A.; Trigas, P. Network biogeography of a complex island system: The Aegean Archipelago revisited. J. Biogeogr. 2017, 44, 651–660. [Google Scholar] [CrossRef]
- Graham, N.R.; Gruner, D.S.; Lim, J.Y.; Gillespie, R.G. Island ecology and evolution: Challenges in the Anthropocene. Environ. Conserv. 2017, 44, 323–335. [Google Scholar] [CrossRef]
- Price, J.P.; Otto, R.; Menezes de Sequeira, M.; Kueffer, C.; Schaefer, H.; Caujapé-Castells, J.; Fernández-Palacios, J.M. Colonization and diversification shape species–area relationships in three Macaronesian archipelagos. J. Biogeogr. 2018, 45, 2027–2039. [Google Scholar] [CrossRef]
- Warren, B.H.; Simberloff, D.; Ricklefs, R.E.; Aguilée, R.; Condamine, F.L.; Gravel, D.; Morlon, H.; Mouquet, N.; Rosindell, J.; Casquet, J.; et al. Islands as model systems in ecology and evolution: Prospects fifty years after MacArthur-Wilson. Ecol. Lett. 2015, 18, 200–217. [Google Scholar] [CrossRef]
- Douma, C.; Koutis, K.; Thanopoulos, R.; Tsigou, R.; Galanidis, A.; Bebeli, P.J. Diversity of agricultural plants on Lesvos Island (Northeast Aegean, Greece) with emphasis on fruit trees. Sci. Hortic. 2016, 210, 65–84. [Google Scholar] [CrossRef]
- Hagenblad, J.; Leino, M.W.; Hernández Afonso, G.; Afonso Morales, D. Morphological and genetic characterization of barley (Hordeum vulgare L.) landraces in the Canary Islands. Genet. Resour. Crop. Evol. 2019, 66, 465–480. [Google Scholar] [CrossRef]
- Médail, F. The specific vulnerability of plant biodiversity and vegetation on Mediterranean islands in the face of global change. Reg. Environ. Chang. 2017, 17, 1775–1790. [Google Scholar] [CrossRef]
- Hellenic Statistical Authority. 2019. Available online: http://www.statistics.gr/en/home/ (accessed on 1 August 2019).
- Sfenthourakis, S.; Triantis, K.A. The Aegean archipelago: A natural laboratory of evolution, ecology and civilisations. J. Biol. Res. 2017, 24, 4. [Google Scholar] [CrossRef]
- Strid, A. Phytogeographia Aegaea and the Flora Hellenica Database. Ann. Nat. Mus. Wien. 1996, 98, 279–289. [Google Scholar]
- Dimopoulos, P.; Raus, T.; Bergmeier, E.; Constantinidis, T.; Iatrou, G.; Kokkini, S.; Strid, A.; Tzanoudakis, D. Vascular plants of Greece: An annotated checklist. Willdenowia 2016, 46, 301–347. [Google Scholar] [CrossRef]
- Ninou, E.G.; Mylonas, I.G.; Tsivelikas, A.L.; Ralli, P.E. Phenotypic diversity of Greek dill (Anethum graveolens L.) landraces. Acta Agric. Scand. Sect. B Soil Plant. Sci. 2017, 67, 318–325. [Google Scholar]
- Tsanakas, G.F.; Mylona, P.V.; Koura, K.; Gleridou, A.; Polidoros, A.N. Genetic diversity analysis of the Greek lentil (Lens culinaris) landrace “Eglouvis” using morphological and molecular markers. Plant Genet. Resour. Characterisation Util. 2018, 16, 469–477. [Google Scholar] [CrossRef]
- Dwivedi, S.; Goldman, I.; Ortiz, R. Pursuing the potential of heirloom cultivars to improve adaptation, nutritional, and culinary features of food crops. Agronomy 2019, 9, 441. [Google Scholar] [CrossRef]
- Bota, J.; Conesa, M.À.; Ochogavia, J.M.; Medrano, H.; Francis, D.M.; Cifre, J. Characterization of a landrace collection for Tomàtiga de Ramellet (Solanum lycopersicum L.) from the Balearic Islands. Genet. Resour. Crop. Evol. 2014, 61, 1131–1146. [Google Scholar] [CrossRef]
- Carillo, P.; Kyriacou, M.C.; El-Nakhel, C.; Pannico, A.; Dell’Aversana, E.; D’Amelia, L.; Colla, G.; Caruso, G.; De Pascale, S.; Rouphael, Y. Sensory and functional quality characterization of protected designation of origin ‘Piennolo del Vesuvio’ cherry tomato landraces from Campania-Italy. Food Chem. 2019, 292, 166–175. [Google Scholar] [CrossRef]
- Schmidt, S.B.; George, T.S.; Brown, L.K.; Booth, A.; Wishart, J.; Hedley, P.E.; Martin, P.; Russell, J.; Husted, S. Ancient barley landraces adapted to marginal soils demonstrate exceptional tolerance to manganese limitation. Ann. Bot. 2019, 123, 831–843. [Google Scholar] [CrossRef]
- Missio, J.C.; Rivera, A.; Figàs, M.R.; Casanova, C.; Camí, B.; Soler, S.; Simó, J. A comparison of landraces vs. Modern varieties of lettuce in organic farming during the winter in the Mediterranean area: An approach considering the viewpoints of breeders, consumers, and farmers. Front. Plant Sci. 2018, 9, 1491. [Google Scholar] [CrossRef]
- Petropoulos, S.A.; Barros, L.; Ferreira, I.C.F.R. Editorial: Rediscovering local landraces: Shaping horticulture for the future. Front. Plant Sci. 2019, 10, 126. [Google Scholar] [CrossRef]
- Karanikolas, P.; Bebeli, P.J.; Thanopoulos, R. Farm economic sustainability and agrobiodiversity: Identifying viable farming alternatives during the economic crisis in Greece. J. Environ. Econ. Policy 2018, 7, 69–84. [Google Scholar] [CrossRef]
- McDorman, B.; Thomas, S. The Importance of Saving Seeds. In Promoting Biodiversity in Food Systems; Hawkins, I.W., Ed.; CRC PressI Llc: Boca Ratón, FL, USA, 2018; pp. 189–210. [Google Scholar]
- FAO STATISTICAL DATABASES. 2017. Available online: http://www.fao.org/faostat/ (accessed on 1 August 2019).
- Augustinos, A.A.; Petropoulos, C.; Karasoulou, V.; Bletsos, F.; Papasotiropoulos, V. Assessing diversity among traditional Greek and foreign eggplant cultivars using molecular markers and morphometrical descriptors. Span. J. Agric. Res. 2016, 14, e0710. [Google Scholar] [CrossRef]
- Thomas, K.; Thanopoulos, R.; Knüpffer, H.; Bebeli, P.J. Plant genetic resources of Lemnos (Greece), an isolated island in the Northern Aegean Sea, with emphasis on landraces. Genet Resour. Crop. Evol. 2012, 59, 1417–1440. [Google Scholar] [CrossRef]
- García-Verdugo, C.; Sajeva, M.; La Mantia, T.; Harrouni, C.; Msanda, F.; Caujapé-Castells, J. Do island plant populations really have lower genetic variation than mainland populations? Effects of selection and distribution range on genetic diversity estimates. Mol. Ecol. 2015, 24, 726–741. [Google Scholar] [CrossRef] [PubMed]
- Hiraoka, Y.; Tamaki, I.; Watanabe, A. The origin of wild populations of Toxicodendron succedaneum on mainland Japan revealed by genetic variation in chloroplast and nuclear DNA. J. Plant Res. 2018, 131, 225–238. [Google Scholar] [CrossRef] [PubMed]
- Jiménez, A.; Weigelt, B.; Santos-Guerra, A.; Caujapé-Castells, J.; Fernández-Palacios, J.M.; Conti, E. Surviving in isolation: Genetic variation, bottlenecks and reproductive strategies in the Canarian endemic Limonium macrophyllum (Plumbaginaceae). Genetica 2017, 145, 91–104. [Google Scholar] [CrossRef]
- Wheelwright, N.T.; Begin, E.; Ellwanger, C.; Taylor, S.H.; Stone, J.L. Minimal loss of genetic diversity and no inbreeding depression in blueflag iris (Iris versicolor) on islands in the Bay of Fundy. Botany 2016, 94, 543–554. [Google Scholar] [CrossRef]
- Hedrén, M.; Olofsson, S.N.; Paun, O. Orchid colonization: Multiple parallel dispersal events and mosaic genetic structure in Dactylorhiza majalis ssp. lapponica on the Baltic island of Gotland. Ann. Bot. 2018, 122, 1019–1032. [Google Scholar]
- Hufford, K.M.; Mazer, S.J.; Hodges, S.A. Genetic variation among mainland and island populations of a native perennial grass used in restoration. AoB Plants 2014, 6. [Google Scholar] [CrossRef]
- Mcglaughlin, M.E.; Wallace, L.E.; Wheeler, G.L.; Bresowar, G.; Riley, L.; Britten, N.R.; Helenurm, K. Do the island biogeography predictions of MacArthur and Wilson hold when examining genetic diversity on the near mainland California Channel Islands? Examples from endemic Acmispon (Fabaceae). Bot. J. Linn. Soc. 2014, 174, 289–304. [Google Scholar] [CrossRef]
- Idrissi, O.; Piergiovanni, A.R.; Toklu, F.; Houasli, C.; Udupa, S.M.; De Keyser, E.; Van Damme, P.; De Riek, J. Molecular variance and population structure of lentil (Lens culinaris Medik.) landraces from Mediterranean countries as revealed by simple sequence repeat DNA markers: Implications for conservation and use. Plant Genet. Resour. Characterisation Util. 2018, 16, 249–259. [Google Scholar] [CrossRef]
- Acquadro, A.; Barchi, L.; Gramazio, P.; Portis, E.; Vilanova, S.; Comino, C.; Plazas, M.; Prohens, J.; Lanteri, S. Coding SNPs analysis highlights genetic relationships and evolution pattern in eggplant complexes. PLoS ONE 2017, 12, e0180774. [Google Scholar] [CrossRef]
- Cericola, F.; Portis, E.; Toppino, L.; Barchi, L.; Acciarri, N.; Ciriaci, T.; Sala, T.; Rotino, G.L.; Lanteri, S. The population structure and diversity of eggplant from Asia and the Mediterranean Basin. PLoS ONE 2013, 8, e73702. [Google Scholar] [CrossRef]
- Hurtado, M.; Vilanova, S.; Plazas, M.; Gramazio, P.; Fonseka, H.H.; Fonseka, R.; Prohens, J. Diversity and relationships of eggplants from three geographically distant secondary centers of diversity. PLoS ONE 2012, 7, e41748. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Yang, Y.; Zhou, X.; Bao, S.; Zhuang, Y. Genetic diversity and population structure of worldwide eggplant (Solanum melongena L.) germplasm using SSR markers. Genet. Resour. Crop. Evol. 2018, 65, 1663–1670. [Google Scholar] [CrossRef]
- Rodriguez-Jimenez, J.; Amaya-Guerra, C.; Baez-Gonzalez, J.; Aguilera-Gonzalez, C.; Urias-Orona, V.; Nino-Medina, G. Physicochemical, functional, and nutraceutical properties of eggplant flours obtained by different drying methods. Molecules 2018, 23, 3210. [Google Scholar] [CrossRef] [PubMed]
- Arivalagan, M.; Bhardwaj, R.; Gangopadhyay, K.K.; Prasad, T.V.; Sarkar, S.K. Mineral composition and their genetic variability analysis in eggplant (Solanum melongena L.) germplasm. J. Appl. Bot. Food Qual. 2013, 86, 99–103. [Google Scholar]
- Raigón, M.D.; Prohens, J.; Muñoz-Falcón, J.E.; Nuez, F. Comparison of eggplant landraces and commercial varieties for fruit content of phenolics, minerals, dry matter and protein. J. Food Compos. Anal. 2008, 21, 370–376. [Google Scholar] [CrossRef]
- Arivalagan, M.; Gangopadhyay, K.K.; Kumar, G.; Bhardwaj, R.; Prasad, T.V.; Sarkar, S.K.; Roy, A. Variability in mineral composition of Indian eggplant (Solanum melongena L.) genotypes. J. Food Compos. Anal. 2012, 26, 173–176. [Google Scholar] [CrossRef]
- Ranil, R.H.G.; Niran, H.M.L.; Plazas, M.; Fonseka, R.M.; Fonseka, H.H.; Vilanova, S.; Andújar, I.; Gramazio, P.; Fita, A.; Prohens, J. Improving seed germination of the eggplant rootstock Solanum torvum by testing multiple factors using an orthogonal array design. Sci. Hortic. 2015, 193, 174–181. [Google Scholar] [CrossRef]
- van der Weerden, G.M.; Barendse, G.W.M. A web-based searchable database developed for the EGGNET project and applied to the radboud university Solanaceae database. Acta Hortic. 2007, 745, 503–506. [Google Scholar] [CrossRef]
- Kaushik, P.; Prohens, J.; Vilanova, S.; Gramazio, P.; Plazas, M. Phenotyping of eggplant wild relatives and interspecific hybrids with conventional and phenomics descriptors provides insight for their potential utilization in breeding. Front. Plant Sci. 2016, 7, 677. [Google Scholar] [CrossRef]
- IBPGR. Descriptors for Eggplant; International Board for Plant Genetic Resources: Rome, Italy, 1990. [Google Scholar]
- Kumar, G.; Meena, B.L.; Kar, R.; Tiwari, S.K.; Gangopadhyay, K.K.; Bisht, I.S.; Mahajan, R.K. Morphological diversity in brinjal (Solanum melongena L.) germplasm accessions. Plant Genet. Res. 2008, 6, 232–236. [Google Scholar] [CrossRef]
- Campbell, C.R.; Plank, C.O. Preparation of Plant Tissue for Laboratory Analysis. In Handbook of Reference Methods for Plant Analysis; Kalra, Y.P., Ed.; CRC Press: Boca Ratón, FL, USA, 1998; pp. 37–50. [Google Scholar]
- Ihaka, R.; Gentleman, R. R: A Language for Data Analysis and Graphics. J. Comput. Graph. Stat. 1996, 5, 299–314. [Google Scholar]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016; ISBN 978-3-319-24277-4. [Google Scholar]
- Alonso, D.; Gramazio, P.; Plazas, M.; Villanueva, G.; García-Fortea, E.; Díez, M.J.; Prohens, J.; Vilanova, S. Protocolo innovador para la obtención de ADN genómico vegetal de alta calidad. In Proceedings of the Congress “I Congreso de Jóvenes Investigadores en Ciencias Agroalimentarias”, Almeria, Spain, 20 December 2018. [Google Scholar]
- Vilanova, S.; Manzur, J.P.; Prohens, J. Development and characterization of genomic simple sequence repeat markers in eggplant and their application to the study of diversity and relationships in a collection of different cultivar types and origins. Mol. Breed. 2012, 30, 647–660. [Google Scholar] [CrossRef]
- Nunome, T.; Negoro, S.; Kono, I.; Kanamori, H.; Miyatake, K.; Yamaguchi, H.; Ohyama, A.; Fukuoka, H. Development of SSR markers derived from SSR-enriched genomic library of eggplant (Solanum melongena L.). Theor. Appl. Genet. 2009, 119, 1143–1153. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Muse, S.V. PowerMaker: An integrated analysis environment for genetic maker analysis. Bioinformatics 2005, 21, 2128–2129. [Google Scholar] [CrossRef] [PubMed]
- Peakall, P.; Smouse, R. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research—An update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef]
- Botstein, D.; White, R.L.; Skolnick, M.; Davis, R.W. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am. J. Hum. Genet. 1980, 32, 314–331. [Google Scholar]
- Nei, M. Genetic distance between populations. Am. Nat. 1972, 106, 283–292. [Google Scholar] [CrossRef]
- Wright, S. The interpretation of population structure by F-statistics with special regard to systems of mating. Evolution 1965, 19, 395–420. [Google Scholar] [CrossRef]
- Pritchard, J.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef] [PubMed]
- Earl, D.A.; VonHoldt, B.M. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 2012, 4, 359–361. [Google Scholar] [CrossRef]
- Remington, D.L.; Thornsberry, J.M.; Matsuoka, Y.; Wilson, L.M.; Whitt, S.R.; Doebley, J.; Kresovich, S.; Goodman, M.M.; Buckler, I.V.E.S. Structure of linkage disequilibrium and phenotypic associations in the maize genome. Proc. Natl. Acad. Sci. USA 2001, 98, 11479–11484. [Google Scholar] [CrossRef]
- Beerling, D.J.; Leake, J.R.; Long, S.P.; Scholes, J.D.; Ton, J.; Nelson, P.N.; Bird, M.; Kantzas, E.; Taylor, L.L.; Sarkar, B.; et al. Farming with crops and rocks to address global climate, food and soil security. Nat. Plants 2018, 4, 138–147. [Google Scholar] [CrossRef] [PubMed]
- Marconi, G.; Ferradini, N.; Russi, L.; Concezzi, L.; Veronesi, F.; Albertini, E. Genetic characterization of the apple germplasm collection in central Italy: The value of local varieties. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Li, J.; Zhao, J.; He, C. Evolutionary developmental genetics of fruit morphological variation within the Solanaceae. Front. Plant Sci. 2015, 6, 248. [Google Scholar] [CrossRef] [PubMed]
- Prohens, J.; Blanca, J.M.; Nuez, F. Morphological and molecular variation in a collection of eggplants from a secondary center of diversity: Implications for conservation and breeding. J. Am. Soc. Hortic. Sci. 2005, 130, 54–63. [Google Scholar] [CrossRef]
- Boyaci, H.F.; Topcu, V.; Tepe, A.; Yildirim, I.K.; Oten, M.; Aktas, A. Morphological and molecular characterization and relationships of Turkish local eggplant heirlooms. Not. Bot. Horti Agrobot. Cluj Napoca 2015, 43, 100–107. [Google Scholar] [CrossRef]
- Tümbilen, Y.; Frary, A.; Mutlu, S.; Doganlar, S. Genetic diversity in Turkish eggplant (Solanum melongena) varieties as determined by morphological and molecular analyses. Int. Res. J. Biotechnol. 2011, 2, 16–25. [Google Scholar]
- Muñoz-Falcón, J.E.; Prohens, J.; Rodríguez-Burruezo, A.; Nuez, F. Potential of local varieties and their hybrids for the improvement of eggplant production in the open field and greenhouse cultivation. J. Food Agric. Environ. 2008, 6, 83–88. [Google Scholar]
- Muñoz-Falcón, J.E.; Prohens, J.; Vilanova, S.; Nuez, F. Characterization, diversity, and relationships of the Spanish striped (Listada) eggplants: A model for the enhancement and protection of local heirlooms. Euphytica 2008, 164, 405–419. [Google Scholar] [CrossRef]
- Polignano, G.; Uggenti, P.; Bisignano, V.; Gatta, C. Della Genetic divergence analysis in eggplant (Solanum melongena L.) and allied species. Genet. Resour. Crop. Evol. 2010, 57, 171–181. [Google Scholar] [CrossRef]
- Russo, V.M. Cultural methods and mineral content of eggplant Solanum melongena fruit. J. Sci. Food Agric. 1996, 71, 119–123. [Google Scholar] [CrossRef]
- Raigón, M.D.; Rodríguez-Burruezo, A.; Prohens, J. Effects of organic and conventional cultivation methods on composition of eggplant fruits. J. Agric. Food Chem. 2010, 58, 6833–6840. [Google Scholar] [CrossRef] [PubMed]
- Ge, H.; Liu, Y.; Jiang, M.; Zhang, J.; Han, H.; Chen, H. Analysis of genetic diversity and structure of eggplant populations (Solanum melongena L.) in China using simple sequence repeat markers. Sci. Hortic. 2013, 162, 71–75. [Google Scholar] [CrossRef]
- Muñoz-Falcón, J.E.; Vilanova, S.; Plazas, M.; Prohens, J. Diversity, relationships, and genetic fingerprinting of the Listada de Gandía eggplant landrace using genomic SSRS and EST-SSRs. Sci. Hortic. 2011, 129, 238–246. [Google Scholar] [CrossRef]
- Demir, K.; Bakir, M.; Sarikamiş, G.; Acunalp, S. Genetic diversity of eggplant (Solanum melongena) germplasm from Turkey assessed by SSR and RAPD markers. Genet. Mol. Res. 2010, 9, 1568–1576. [Google Scholar] [CrossRef]
- Vilanova, S.; Hurtado, M.; Cardona, A.; Plazas, M.; Gramazio, P.; Herraiz, F.J.; Andújar, I.; Prohens, J. Genetic diversity and relationships in local varieties of eggplant from different cultivar groups as assessed by genomic SSR markers. Not. Bot. Horti Agrobot. Cluj Napoca 2014, 42, 59–65. [Google Scholar] [CrossRef][Green Version]
- Pessarakli, M.; Dris, R. Pollination and breeding of eggplants. J. Food Agric. Environ. 2004, 2, 218–219. [Google Scholar]
- Muñoz-Falcón, J.E.; Prohens, J.; Vilanova, S.; Nuez, F. Diversity in commercial varieties and landraces of black eggplants and implications for broadening the breeders’ gene pool. Ann. Appl. Biol. 2009, 154, 453–465. [Google Scholar] [CrossRef]
- Frankham, R. Do island populations have less genetic variation than mainland populations? Heredity 1997, 78, 311–327. [Google Scholar] [CrossRef] [PubMed]
- Whittaker, R.; Fernández-Palacios, J. Island Biogeography: Ecology, Evolution, and Conservation; Oxford University Press: Oxford, UK, 2007; ISBN 978-0198566120. [Google Scholar]
- Hargreaves, S.; Maxted, N.; Hirano, R.; Abberton, M.; Skøt, L.; Ford-Lloyd, B.V. Islands as refugia of Trifolium repens genetic diversity. Conserv. Genet. 2010, 11, 1317–1326. [Google Scholar] [CrossRef]
- Lawson, D.A.; Rands, S.A. The effects of rainfall on plant–pollinator interactions. Arthropod Plant Interact. 2019, 13, 561–569. [Google Scholar] [CrossRef]
- Kyriakopoulou, O.G.; Arens, P.; Pelgrom, K.T.B.; Karapanos, I.; Bebeli, P.; Passam, H.C. Genetic and morphological diversity of okra (Abelmoschus esculentus [L.] Moench.) genotypes and their possible relationships, with particular reference to Greek landraces. Sci. Hortic. 2014, 171, 58–70. [Google Scholar] [CrossRef]
- Caguiat, X.G.I.; Hautea, D.M. Genetic diversity analysis of eggplant (Solanum melongena L.) and related wild species in the Philippines using morphological and SSR markers. Sabrao J. Breed. Genet. 2014, 46, 183–201. [Google Scholar]
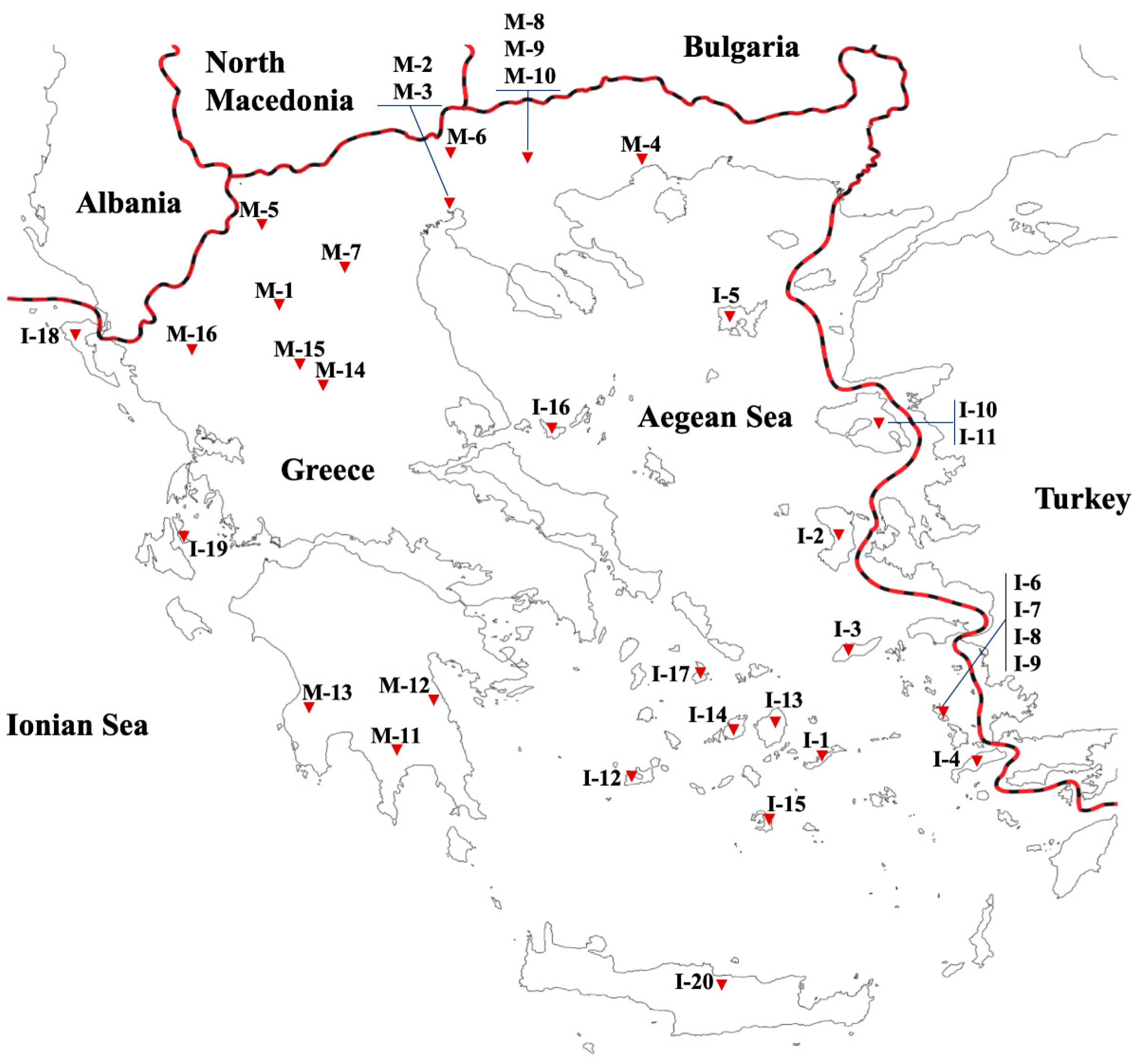


| Accession Code | Study Code | Territory | Geographic Region | Collection Site | Longitude | Latitude | Altitude (m) | Status |
|---|---|---|---|---|---|---|---|---|
| ANP-025/07 | I-1 | Island | Aegean Sea | Amorgos, Aegiali | 36°90′ Ν | 25°99′ Ε | 298 | Landrace |
| X-034/06 | I-2 | Island | Aegean Sea | Chios, Ag. Georgios Sikousis | 38°19′ N | 26°03′ E | 369 | Landrace |
| IS-031/07 | I-3 | Island | Aegean Sea | Ikaria, Droutsoulas | 37°36′ N | 26°11′ E | 410 | Landrace |
| LKK-094/07 | I-4 | Island | Aegean Sea | Kos, Antimacheia | 36°48′ N | 27°05′ E | 133 | Landrace |
| X-116/06 | I-5 | Island | Aegean Sea | Lemnos, Kontopouli | 39°55′ N | 25°20′ E | 40 | Landrace |
| White Leros | I-6 | Island | Aegean Sea | Leros Island | 37°09′ N | 26°49′ E | 32 | Cultivar |
| Wide Purple | I-7 | Island | Aegean Sea | Leros Island | 37°08′ N | 26°51′ E | 6 | Landrace |
| Long Purple | I-8 | Island | Aegean Sea | Leros Island | 37°08′ N | 26°51′ E | 6 | Landrace |
| LKK-008/07 | I-9 | Island | Aegean Sea | Leros, Kamara | 37°09′ N | 26°49′ E | 29 | Landrace |
| M-039/06 | I-10 | Island | Aegean Sea | Lesvos, Paleokipos | 39°05′ N | 26°45′ E | 58 | Landrace |
| M-069/06 | I-11 | Island | Aegean Sea | Lesvos, Keramia | 39°12′ N | 26°42′ E | 14 | Landrace |
| MFS-030/07 | I-12 | Island | Aegean Sea | Milos, Mitakas | 36°44′ N | 24°29′ E | 50 | landrace |
| ANP-180/07 | I-13 | Island | Aegean Sea | Naxos, Agia Anna | 37°04′ Ν | 25°21′ Ε | 4 | Landrace |
| ANP-215/07 | I-14 | Island | Aegean Sea | Paros, Lefkes | 37°07′ Ν | 25°12′ Ε | 210 | Landrace |
| HL-027/07 | I-15 | Island | Aegean Sea | Santorini, Vourvoulos | 36°26′ N | 25°26′ E | 116 | Landrace |
| SAS-078/07 | I-16 | Island | Aegean Sea | Skopelos, Chora | 39°07′ N | 23°43′ E | 41 | Landrace |
| ATS-110/06 | I-17 | Island | Aegean Sea | Syros Island, Chrousa | 37°24′ N | 24°55′ E | 320 | Landrace |
| GRC-002/08 | I-18 | Island | Ionian Sea | Corfu, Skripero | 39°42′ N | 19°46′ E | 139 | Landrace |
| HL-237/07 | I-19 | Island | Crete | Iraklion, Moni Savvathianon | 35°37′ N | 25°00′ E | 467 | Landrace |
| IK-082/06 | I-20 | Island | Ionian Sea | Ithaki Island, Perachori | 38°34′ N | 20°71′ E | 343 | Landrace |
| K-153/06 | M-1 | Mainland | Macedonia | Grevena, Pontini | 40°04′ N | 21°40′ E | 819 | Landrace |
| EMI | M-2 | Mainland | Macedonia | IPGRB/HAO DEMETER | 40°32′ N | 22°59′ Ε | 19 | Cultivar |
| Lagkada | M-3 | Mainland | Macedonia | IPGRB/HAO DEMETER | 40°32′ N | 22°59′ Ε | 19 | Cultivar |
| KD-053/07 | M-4 | Mainland | Macedonia | Kavala, Platanotopos | 40°50′ N | 24°03′ E | 244 | Landrace |
| F-154/06 | M-5 | Mainland | Macedonia | Kastoria, Ampelokipoi | 40°46′ N | 21°31′ E | 638 | Landrace |
| SK-044/066 | M-6 | Mainland | Macedonia | Kilkis, Eptalofos | 41°00′ N | 23°08′ E | 451 | Landrace |
| K-054/06 | M-7 | Mainland | Macedonia | Kozani, Anarachi | 40°29′ N | 21°34′ E | 727 | Landrace |
| VG-011/083 | M-8 | Mainland | Macedonia | Serres | 41°05′ N | 23°35′ E | 45 | Landrace |
| SK-056/06 | M-9 | Mainland | Macedonia | Serres, Platanakia | 41°17′ N | 22°56′ E | 313 | Landrace |
| Scoutari | M-10 | Mainland | Macedonia | Serres/Skoutari | 41°01′ N | 23°31′ E | 14 | Cultivar |
| P-175/06 | M-11 | Mainland | Peloponnese | Lakonia, Lyra | 36°38′ N | 22°57′ E | 400 | Landrace |
| Tsakoniki | M-12 | Mainland | Peloponnese | Leonidio | 37°10′ N | 22°51′ E | 40 | Cultivar |
| P-084/06 | M-13 | Mainland | Peloponnese | Messinia, Kakana | 37°18′ Ν | 21°44′ E | 136 | Landrace |
| T-099/06 | M-14 | Mainland | Thessaly | Karditsa, Neo Ikonio | 39°27′ N | 22°21′ E | 107 | Landrace |
| T-527/06 | M-15 | Mainland | Thessaly | Trikala, Megarxis | 39°36′ N | 21°45′ E | 142 | Landrace |
| GRC 1430/04 | M-16 | Mainland | Epirus | Ioannina, Pogoni Vasiliko | 40°00′ N | 20°35′ E | 805 | Landrace |
| Descriptor Code | Descriptor Name | Descriptor Scale/Unit |
|---|---|---|
| Plant descriptors | ||
| PGH | Plant Growth Habit | 3–7 (3 = upright; 7 = prostrate) |
| PH | Plant Height | cm |
| NOLFF | Number of Leaves to First Flower | number |
| DSFS | Days Since Fruit Set | number |
| Leaf descriptors | ||
| LPL | Leaf Petiole Length | cm |
| LBLe | Leaf Blade Length | cm |
| LBW | Leaf Blade Width | cm |
| LBLo | Leaf Blade Lobing | 1–9 (1 = very weak; 9 = very strong) |
| LSS | Leaf Surface Shape | 1–9 (1 = flat; 9 = very convex or bullate) |
| LP | Leaf Prickles | 0–9 (0 = none; 9 = more than 20) |
| Flower descriptors | ||
| NOFPI | Number of Flowers Per Inflorescence | number |
| Fruit descriptors | ||
| FCP | Fruit Calyx Prickles | 0–9 (0 = none; 9 = more than 30) |
| FPL | Fruit Pedicel Length | cm |
| FL | Fruit Length | cm |
| FB | Fruit Breadth | mm |
| FLBR | Fruit Length to Breadth Ratio | 1–9 (1 = broader than long; 9 = several times longer than broad) |
| FS | Fruit Shape | 3–7 (Position of widest part of fruit: 3 = ¼ way from base to tip; 5 = ½ way from base to tip; 7 = ¾ way from base to tip) |
| FC | Fruit Curvature | 1–9 (1 = none, fruit straight; 9 = U shaped) |
| FW | Fruit Weight | g |
| FPC | Fruit Predominant Color | 1–9 (1 = milk white; 9 = black) |
| FAC | Fruit Additional Color | 1–9 (1 = milk white; 9 = black) |
| FFC | Fruit Flesh Color | 3–7 (3 = white; 7 = green) |
| SSR Locus | Motif | Forward Primer and Reverse Primer (5′→3′) | Size Range (bp) | Tº Annealing | Dye | Source |
|---|---|---|---|---|---|---|
| csm4 | (GA)15 | GCGTACCAATTCTAACCACAAG | 238–254 | 60 | PET | Vilanova et al. [50] |
| GTAATCCGCTTCCCATTTCTC | ||||||
| csm27 | (GA)23 | TGTTTGGAGGTGAGGGAAAG | 193–210 | 60 | VIC | Vilanova et al. [50] |
| TCCAACTCACCGGAAAAATC | ||||||
| csm32 | (AG)23 | TCGAAAGTACAGCGGAGAAAG | 248–254 | 60 | NED | Vilanova et al. [50] |
| GGGGGTTTGATTTTCATTTTC | ||||||
| emi02c21 | (AC)13A(TA)4 | TGTGAGGAGAAGAATCAGAGGATCA | 126–136 | 60 | VIC | Nunome et al. [51] |
| CGCGACTAAGTTTTGTTCCTGAAA | ||||||
| eme11f04 | (TC)16 | ACCCCCAAATCAAATCATTTACCC | 88–100 | 60 | FAM | Nunome et al. [51] |
| GGCATGGTTAGGGTTTTTAGCGTT |
| Mainland | Islands | p-Value | |||
|---|---|---|---|---|---|
| Mean | CV (%) | Mean | CV (%) | ||
| (Range) | (Range) | ||||
| Plant descriptors | |||||
| PGH | 4.36 | 23.70 | 5.09 | 21.50 | 0.052 |
| (3.00–6.50) | (3.00–7.00) | ||||
| PH | 80.59 | 16.30 | 71.72 | 21.90 | 0.083 |
| (56.83–106.00) | (46.50–106.17) | ||||
| NOLFF | 4.97 | 13.30 | 5.41 | 12.50 | 0.060 |
| (4.00–6.67) | (4.20–6.83) | ||||
| DSFS | 50.51 | 8.20 | 48.26 | 9.10 | 0.129 |
| (42.83–58.33) | (41.00–55.67) | ||||
| Leaf descriptors | |||||
| LPL | 9.83 | 17.60 | 10.22 | 19.40 | 0.541 |
| (6.79–12.29) | (6.53–14.63) | ||||
| LBLe | 31.51 | 8.40 | 31.4 | 10.50 | 0.918 |
| (27.50–36.97) | (25.57–37.48) | ||||
| LBW | 20.43 | 11.50 | 22.14 | 9.80 | 0.032 * |
| (16.35–25.62) | (18.16–25.63) | ||||
| LBLo | 3.38 | 23.80 | 4.37 | 21.90 | 0.004 * |
| (3.00–5.00) | (3.00–5.00) | ||||
| LSS | 4.88 | 27.90 | 5.42 | 19.80 | 0.203 |
| (3.00–7.00) | (3.00–7.00) | ||||
| LP | 0.18 | 126.40 | 0.21 | 125.70 | 0.892 |
| (0.00–0.92) | (0.00–0.92) | ||||
| Flower descriptors | |||||
| NOFPI | 1.15 | 11.10 | 1.22 | 16.90 | 0.388 |
| (1.00–1.40) | (1.00–1.60) | ||||
| Fruit descriptors | |||||
| FCP | 1.51 | 83.90 | 3.32 | 57.90 | 0.003 * |
| (0.00–4.00) | (0.33–7.00) | ||||
| FPL | 6.33 | 16.70 | 6.24 | 32.10 | 0.840 |
| (4.91–8.22) | (3.58–9.72) | ||||
| FL | 18.46 | 13.00 | 18.4 | 42.80 | 0.246 |
| (14.50–22.32) | (11.2–43.85) | ||||
| FB | 61.93 | 16.20 | 85.1 | 20.30 | 0.000 * |
| (50.91–87.78) | (52.75–110.76) | ||||
| FLBR | 7.49 | 11.50 | 5.2 | 44.40 | 0.002 * |
| (5.00–8.50) | (1.33–8.33) | ||||
| FS | 4.69 | 7.00 | 4.36 | 12.20 | 0.106 |
| (4.00–5.33) | (3.33–5.00) | ||||
| FC | 3.3 | 32.30 | 1.81 | 53.70 | 0.000 * |
| (1.00–5.00) | (1.00–4.00) | ||||
| FW | 246.33 | 19.70 | 329.64 | 20.80 | 0.000 * |
| (177.32–368.40) | (215.66–500.67) | ||||
| FPC | 6.52 | 22.10 | 5.83 | 40.50 | 0.550 |
| (2.00–8.00) | (1.00–8.17) | ||||
| FAC | 4.31 | 48.00 | 4.78 | 48.20 | 0.527 |
| (1.00–7.50) | (1.00–8.17) | ||||
| FFC | 4.73 | 15.10 | 4.45 | 20.60 | 0.218 |
| (3.00–5.33) | (3.00–6.00) | ||||
| Minerals | |||||
| Κ (mg/g DW) | 25.05 | 13.21 | 23.14 | 11.58 | 0.073 |
| (20.07–31.50) | (19.80–29.00) | ||||
| Mg (mg/g DW) | 2.22 | 6.75 | 2.05 | 12.19 | 0.015 * |
| (1.96–2.46) | (1.63–2.43) | ||||
| Cu (mg/Kg DW) | 9.56 | 47.91 | 7.65 | 69.67 | 0.278 |
| (2.91–18.14) | (2.35–26.93) | ||||
| Fe (mg/Kg DW) | 31.8 | 14.68 | 31.48 | 15.78 | 0.850 |
| (25.14–43.94) | (22.02–41.84) | ||||
| Mn (mg/Kg DW) | 12.25 | 8.82 | 11.74 | 9.71 | 0.198 |
| (9.61–13.67) | (9.79–13.84) | ||||
| Zn (mg/Kg DW) | 49.75 | 26.85 | 49.39 | 31.14 | 0.942 |
| (30.30–77.26) | (29.53–76.90) | ||||
| SSR Locus | Na | f | Ne | Ng | PIC | Ho | He | F |
|---|---|---|---|---|---|---|---|---|
| All accessions (n = 36) | ||||||||
| csm4 | 7 | 0.32 | 4.41 | 10.00 | 0.76 | 0.14 | 0.77 | 0.83 |
| csm27 | 4 | 0.50 | 2.63 | 7.00 | 0.58 | 0.17 | 0.62 | 0.75 |
| csm32 | 5 | 0.53 | 2.57 | 7.00 | 0.55 | 0.17 | 0.61 | 0.73 |
| emi02c21 | 4 | 0.61 | 2.19 | 4.00 | 0.48 | 0.00 | 0.54 | 1.00 |
| eme11f04 | 3 | 0.65 | 1.92 | 5.00 | 0.40 | 0.17 | 0.48 | 0.66 |
| Mean ± SE | 4.6 ± 0.7 | 0.522 ± 0.05 | 2.746 ± 0.43 | 6.6 ± 1.00 | 0.552 ± 0.05 | 0.130 ± 0.03 | 0.605 ± 0.04 | 0.796 ± 0.05 |
| Islands accessions (n = 20) | ||||||||
| csm4 | 6 | 0.33 | 4.65 | 8.00 | 0.75 | 0.20 | 0.79 | 0.76 |
| csm27 | 4 | 0.53 | 2.28 | 5.00 | 0.47 | 0.25 | 0.56 | 0.57 |
| csm32 | 5 | 0.58 | 2.53 | 7.00 | 0.56 | 0.25 | 0.61 | 0.60 |
| emi02c21 | 3 | 0.50 | 2.20 | 3.00 | 0.44 | 0.00 | 0.55 | 1.00 |
| eme11f04 | 3 | 0.50 | 2.29 | 5.00 | 0.47 | 0.30 | 0.56 | 0.49 |
| Mean ± SE | 4.2 ± 0.6 | 0.485 ± 0.04 | 2.79 ± 0.46 | 5.6 ± 0.90 | 0.538 ± 0.05 | 0.200 ± 0.05 | 0.612 ± 0.04 | 0.686 ± 0.09 |
| Mainland accessions (n = 16) | ||||||||
| csm4 | 4 | 0.37 | 3.44 | 5.00 | 0.66 | 0.07 | 0.71 | 0.91 |
| csm27 | 3 | 0.50 | 2.57 | 4.00 | 0.54 | 0.07 | 0.61 | 0.90 |
| csm32 | 3 | 0.47 | 2.26 | 4.00 | 0.46 | 0.06 | 0.56 | 0.89 |
| emi02c21 | 4 | 0.75 | 1.71 | 4.00 | 0.39 | 0.00 | 0.41 | 1.00 |
| eme11f04 | 2 | 0.94 | 1.13 | 2.00 | 0.11 | 0.00 | 0.12 | 1.00 |
| Mean ± SE | 3.2 ± 0.4 | 0.604 ± 0.10 | 2.22 ± 0.39 | 3.8 ± 0.50 | 0.429 ± 0.09 | 0.039 ± 0.01 | 0.482 ± 0.10 | 0.923 ± 0.02 |
| Source | df | S.S. | M.S. | Variance Components | Percentage of Variation | F-Statistics | p-Value |
|---|---|---|---|---|---|---|---|
| Between groups | 1 | 9.19 | 9.19 | 0.19 | 11.12 | Fst 0.111 | 0.001 |
| Among accessions | 34 | 89.42 | 2.63 | 1.16 | 69.62 | Fis 0.783 | 0.001 |
| Within accessions | 36 | 11.50 | 0.32 | 0.32 | 19.25 | Fit 0.807 | 0.001 |
| Total | 71 | 110.11 | 1.66 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gramazio, P.; Chatziefstratiou, E.; Petropoulos, C.; Chioti, V.; Mylona, P.; Kapotis, G.; Vilanova, S.; Prohens, J.; Papasotiropoulos, V. Multi-Level Characterization of Eggplant Accessions from Greek Islands and the Mainland Contributes to the Enhancement and Conservation of this Germplasm and Reveals a Large Diversity and Signatures of Differentiation between both Origins. Agronomy 2019, 9, 887. https://doi.org/10.3390/agronomy9120887
Gramazio P, Chatziefstratiou E, Petropoulos C, Chioti V, Mylona P, Kapotis G, Vilanova S, Prohens J, Papasotiropoulos V. Multi-Level Characterization of Eggplant Accessions from Greek Islands and the Mainland Contributes to the Enhancement and Conservation of this Germplasm and Reveals a Large Diversity and Signatures of Differentiation between both Origins. Agronomy. 2019; 9(12):887. https://doi.org/10.3390/agronomy9120887
Chicago/Turabian StyleGramazio, Pietro, Eleni Chatziefstratiou, Constantinos Petropoulos, Vasileia Chioti, Photini Mylona, George Kapotis, Santiago Vilanova, Jaime Prohens, and Vasileios Papasotiropoulos. 2019. "Multi-Level Characterization of Eggplant Accessions from Greek Islands and the Mainland Contributes to the Enhancement and Conservation of this Germplasm and Reveals a Large Diversity and Signatures of Differentiation between both Origins" Agronomy 9, no. 12: 887. https://doi.org/10.3390/agronomy9120887
APA StyleGramazio, P., Chatziefstratiou, E., Petropoulos, C., Chioti, V., Mylona, P., Kapotis, G., Vilanova, S., Prohens, J., & Papasotiropoulos, V. (2019). Multi-Level Characterization of Eggplant Accessions from Greek Islands and the Mainland Contributes to the Enhancement and Conservation of this Germplasm and Reveals a Large Diversity and Signatures of Differentiation between both Origins. Agronomy, 9(12), 887. https://doi.org/10.3390/agronomy9120887









