Profiling of the Differential Abundance of Drought and Salt Stress-Responsive MicroRNAs Across Grass Crop and Genetic Model Plant Species
Abstract
1. Introduction
2. The Impact of Drought and Salt Stress on American and Australian Agriculture
3. The Role of Plant microRNAs in Response to Drought and Salt Stress
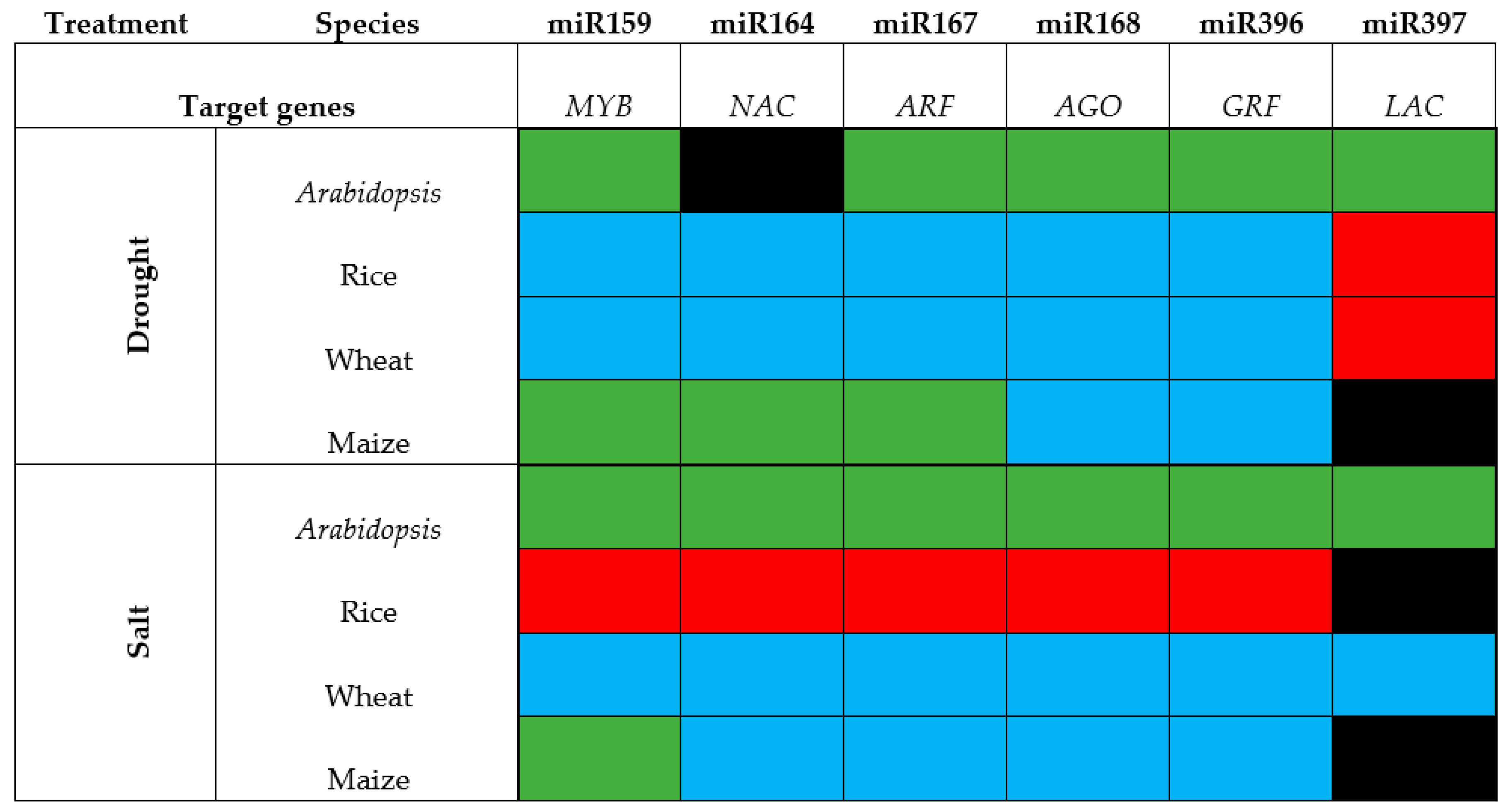
4. The Varying Responses of Six Highly Conserved microRNAs to Drought and Salt Stress
5. Investigation of the Transcription Factor Binding Site Landscapes of MICRORNA Gene Promoters
6. Timing, Treatment, Tissue and “Tolerance” to Drought and Salt Stress
7. Non-Conserved microRNAs Responsive to Drought or Salt Stress
8. Conclusions and Future Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- Foley, J.A.; Ramankutty, N.; Brauman, K.A.; Cassidy, E.S.; Gerber, J.S.; Johnston, M.; Mueller, N.D.; O’Connell, C.; Ray, D.K.; West, P.C.; et al. Solutions for a cultivated planet. Nature 2011, 478, 337–342. [Google Scholar] [CrossRef] [PubMed]
- Glover, J.D.; Reganold, J.P.; Bell, L.W.; Borevitz, J.; Brummer, E.C.; Buckler, E.S.; Cox, C.M.; Cox, T.S.; Crews, T.E.; Culman, S.W.; et al. Increased food and ecosystem security via perennial grains. Science 2010, 328, 1638–1639. [Google Scholar] [CrossRef] [PubMed]
- Sarwar, M.H.; Sarwar, M.F.; Sarwar, M.; Qadri, N.A.; Moghal, S. The importance of cereals (Poaceae: Gramineae) nutrition in human health: A review. J. Cereals Oilseeds 2013, 4, 32–35. [Google Scholar] [CrossRef]
- Mullet, J.; Morishige, D.; McCormick, R.; Truong, S.; Hilley, J.; McKinley, B.; Anderson, R.; Olson, S.N.; Rooney, W. Energy Sorghum—A genetic model for the design of C4 grass bioenergy crops. J. Exp. Bot. 2014, 65, 3479–3489. [Google Scholar] [CrossRef] [PubMed]
- Hoang, N.V.; Furtado, A.; Botha, F.C.; Simmons, B.A.; Henry, R.J. Potential for genetic improvement of sugarcane as a source of biomass for biofuels. Front. Bioeng. Biotechnol. 2015, 3, 182. [Google Scholar] [CrossRef] [PubMed]
- Baulcombe, D.; Crute, I.; Davies, B.; Dunwell, J.; Gale, M.; Jones, J.; Pretty, J.; Sutherland, W.; Toulmin, C. Reaping the Benefits: Science and the Sustainable Intensification of Global Agriculture; The Royal Society: London, UK, 2009; p. 72. [Google Scholar]
- Liu, H.; Searle, I.R.; Mather, D.E.; Able, A.J.; Able, J.A. Morphological, physiological and yield responses of durum wheat to pre-anthesis water-deficit stress are genotype-dependent. Crop Pasture Sci. 2015, 66, 1024–1038. [Google Scholar] [CrossRef]
- Earl, H.J.; Davis, R.F. Effect of drought stress on leaf and whole canopy radiation use efficiency and yield of maize. Agron. J. 2003, 95, 688–696. [Google Scholar] [CrossRef]
- Giunta, F.; Motzo, R.; Deidda, M. Effect of drought on yield and yield components of durum wheat and triticale in a Mediterranean environment. Field Crops Res. 1993, 33, 399–409. [Google Scholar] [CrossRef]
- Katerji, N.; Van Hoorn, J.W.; Hamdy, A.; Karam, F.; Mastrorilli, M. Effect of salinity on water stress, growth, and yield of maize and sunflower. Agric. Water Manag. 1996, 30, 237–249. [Google Scholar] [CrossRef]
- Pessarakli, M.; Tucker, T.C.; Nakabayashi, K. Growth response of barley and wheat to salt stress. J. Plant Nutr. 1991, 14, 331–340. [Google Scholar] [CrossRef]
- Samarah, N.H. Effects of drought stress on growth and yield of barley. Agron. Sustain. Dev. 2005, 25, 145–149. [Google Scholar] [CrossRef]
- Venuprasad, R.; Lafitte, H.R.; Atlin, G.N. Response to direct selection for grain yield under drought stress in rice. J. Trop. Crop Sci. 2007, 47, 285–293. [Google Scholar] [CrossRef]
- Zeng, L.; Shannon, M.C.; Lesch, S.M. Timing of salinity stress affects rice growth and yield components. Agric. Water Manag. 2001, 48, 191–206. [Google Scholar] [CrossRef]
- Dai, A. Increasing drought under global warming in observations and models. Nat. Clim. Chang. 2013, 3, 52–58. [Google Scholar] [CrossRef]
- Lobell, D.B.; Field, C.B. Global scale climate–crop yield relationships and the impacts of recent warming. Environ. Res. Lett. 2007, 2, 014002. [Google Scholar] [CrossRef]
- Foley, J.A.; Asner, G.P.; Costa, M.H.; Coe, M.T.; DeFries, R.; Gibbs, H.K.; Howard, E.A.; Olson, S.; Patz, J.; Ramankutty, N.; et al. Amazonia revealed: Forest degradation and loss of ecosystem goods and services in the Amazon Basin. Front. Ecol. Environ. 2007, 5, 25–32. [Google Scholar] [CrossRef]
- Monfreda, C.; Ramankutty, N.; Foley, J.A. Farming the planet: 2. Geographic distribution of crop areas, yields, physiological types, and net primary production in the year 2000. Glob. Biogeochem. Cycles 2008, 22. [Google Scholar] [CrossRef]
- Ding, Y.; Tao, Y.; Zhu, C. Emerging roles of microRNAs in the mediation of drought stress response in plants. J. Exp. Bot. 2013, 64, 3077–3086. [Google Scholar] [CrossRef] [PubMed]
- Khraiwesh, B.; Zhu, J.K.; Zhu, J. Role of miRNAs and siRNAs in biotic and abiotic stress responses of plants. BBA Gene Regul. Mech. 2012, 1819, 137–148. [Google Scholar] [CrossRef] [PubMed]
- Kruszka, K.; Pieczynski, M.; Windels, D.; Bielewicz, D.; Jarmolowski, A.; Szweykowska-Kulinska, Z.; Vazquez, F. Role of microRNAs and other sRNAs of plants in their changing environments. J. Plant Physiol. 2012, 169, 1664–1672. [Google Scholar] [CrossRef] [PubMed]
- Budak, H.; Zhang, B. MicroRNAs in model and complex organisms. Funct. Integr. Genom. 2017, 17, 121–124. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Searle, I.R.; Watson-Haigh, N.S.; Baumann, U.; Mather, D.E.; Able, A.J.; Able, J.A. Genome-wide identification of microRNAs in leaves and the developing head of four durum genotypes during water deficit stress. PLoS ONE 2015, 10, e0142799. [Google Scholar] [CrossRef] [PubMed]
- Seki, M.; Narusaka, M.; Ishida, J.; Nanjo, T.; Fujita, M.; Oono, Y.; Shinozaki, K. Monitoring the expression profiles of 7000 Arabidopsis genes under drought, cold and high-salinity stresses using a full-length cDNA microarray. Plant J. 2002, 31, 279–292. [Google Scholar] [CrossRef] [PubMed]
- Sunkar, R. MicroRNAs with macro-effects on plant stress responses. Semin. Cell Dev. Biol. 2010, 21, 805–811. [Google Scholar] [CrossRef] [PubMed]
- Sunkar, R.; Chinnusamy, V.; Zhu, J.; Zhu, J.K. Small RNAs as big players in plant abiotic stress responses and nutrient deprivation. Trends Plant Sci. 2007, 12, 301–309. [Google Scholar] [CrossRef] [PubMed]
- Godfray, H.C.J.; Beddington, J.R.; Crute, I.R.; Haddad, L.; Lawrence, D.; Muir, J.F.; Toulmin, C. Food security: The challenge of feeding 9 billion people. Science 2010, 327, 812–818. [Google Scholar] [CrossRef] [PubMed]
- Tester, M.; Langridge, P. Breeding technologies to increase crop production in a changing world. Science 2010, 327, 818–822. [Google Scholar] [CrossRef] [PubMed]
- Soreng, R.J.; Peterson, P.M.; Romaschenko, K.; Davidse, G.; Zuloaga, F.O.; Judziewicz, E.J.; Filgueiras, T.S.; Davis, J.I.; Morrone, O. A worldwide phylogenetic classification of the Poaceae (Gramineae). J. Syst. Evol. 2015, 53, 117–137. [Google Scholar] [CrossRef]
- Mittler, R.; Blumwald, E. Genetic engineering for modern agriculture: Challenges and perspectives. Annu. Rev. Plant Biol. 2010, 61, 443–462. [Google Scholar] [CrossRef] [PubMed]
- Schlenker, W.; Lobell, D.B. Robust negative impacts of climate change on African agriculture. Environ. Res. Lett. 2010, 5, 014010. [Google Scholar] [CrossRef]
- Schlenker, W.; Roberts, M.J. Nonlinear temperature effects indicate severe damages to US crop yields under climate change. Proc. Natl. Acad. Sci. USA 2009, 106, 15594–15598. [Google Scholar] [CrossRef] [PubMed]
- McAlpine, C.A.; Syktus, J.; Ryan, J.G.; Deo, R.C.; McKeon, G.M.; McGowan, H.A.; Phinn, S.R. A continent under stress: Interactions, feedbacks and risks associated with impact of modified land cover on Australia’s climate. Glob. Chang. Biol. 2009, 15, 2206–2223. [Google Scholar] [CrossRef]
- World Food Programme. The State of Food Insecurity in the World 2013—The Multiple Dimensions of Food Security; FAO: Rome, Italy, 2013. [Google Scholar]
- Pinstrup-Andersen, P. Food security: Definition and measurement. Food Secur. 2009, 1, 5–7. [Google Scholar] [CrossRef]
- Schmidhuber, J.; Tubiello, F.N. Global food security under climate change. Proc. Natl. Acad. Sci. USA 2007, 104, 19703–19708. [Google Scholar] [CrossRef] [PubMed]
- Lal, R. Soils and food sufficiency. A review. Agron. Sustain. Dev. 2009, 29, 113–133. [Google Scholar] [CrossRef]
- Long, S.P.; Marshall-Colon, A.; Zhu, X.G. Meeting the global food demand of the future by engineering crop photosynthesis and yield potential. Cell 2015, 161, 56–66. [Google Scholar] [CrossRef] [PubMed]
- Ray, D.K.; Mueller, N.D.; West, P.C.; Foley, J.A. Yield trends are insufficient to double global crop production by 2050. PLoS ONE 2013, 8, e66428. [Google Scholar] [CrossRef] [PubMed]
- Tilman, D.; Balzer, C.; Hill, J.; Befort, B.L. Global food demand and the sustainable intensification of agriculture. Proc. Natl. Acad. Sci. USA 2011, 108, 20260–20264. [Google Scholar] [CrossRef] [PubMed]
- National Corn Growers Association. The World of Corn. Available online: http://www.worldofcorn.com/ (accessed on 12 December 2017).
- FAOSTAT. Food and Agriculture Organization of the United Nations Statistic Division; FAO: Rome, Italy, 2015.
- Daryanto, S.; Wang, L.; Jacinthe, P.A. Global synthesis of drought effects on maize and wheat production. PLoS ONE 2016, 11, e0156362. [Google Scholar] [CrossRef] [PubMed]
- Brown, D.; Taylor, J.; Bell, M. The demography of desert Australia. Rangel. J. 2008, 30, 29–43. [Google Scholar] [CrossRef]
- Turner, D.; Ostendorf, B.; Lewis, M. An introduction to patterns of fire in arid and semi-arid Australia, 1998–2004. Rangel. J. 2008, 30, 95–107. [Google Scholar] [CrossRef]
- Connor, J.; Schwabe, K.; King, D.; Kaczan, D.; Kirby, M. Impacts of climate change on lower Murray irrigation. Aust. J. Agric. Resour. Econ. 2009, 53, 437–456. [Google Scholar] [CrossRef]
- Qureshi, M.E.; Hanjra, M.A.; Ward, J. Impact of water scarcity in Australia on global food security in an era of climate change. Food Policy 2013, 38, 136–145. [Google Scholar] [CrossRef]
- Cline, W.R. Global Warming and Agriculture: Impact Estimates by Country; Peterson Institute: Washington, DC, USA, 2007. [Google Scholar]
- Gunasekera, D.; Kim, Y.; Tulloh, C.; Ford, M. Climate change-impacts on Australian agriculture. Aust. Commod. Forecasts Issues 2007, 14, 657–676. [Google Scholar]
- McJannet, D.L.; Webster, I.T.; Stenson, M.P.; Sherman, B.S. Estimating Open Water Evaporation for the Murray-Darling Basin: A Report to the Australian Government from the CSIRO Murray-Darling Basin Sustainable Yields Project; CSIRO: Canberra, Australia, 2008. [Google Scholar]
- Rengasamy, P. Soil processes affecting crop production in salt-affected soils. Funct. Plant Biol. 2010, 37, 613–620. [Google Scholar] [CrossRef]
- Rengasamy, P. World salinization with emphasis on Australia. J. Exp. Bot. 2006, 57, 1017–1023. [Google Scholar] [CrossRef] [PubMed]
- Scanlon, B.R.; Jolly, I.; Sophocleous, M.; Zhang, L. Global impacts of conversions from natural to agricultural ecosystems on water resources: Quantity versus quality. Water Resour. Res. 2007, 43. [Google Scholar] [CrossRef]
- Holburt, M. The lower Colorado—A salty river. Calif. Agric. 1984, 38, 6–8. [Google Scholar]
- Borda, C. Economic Impacts from Salinity in the Lower Colorado River Basin; Technical Memorandum Number EC-04-02; US Department of the Interior, Bureau of Reclamation: Washington, DC, USA, 2004.
- Hütsch, B.W.; Jung, S.; Schubert, S. Comparison of Salt and Drought-Stress Effects on Maize Growth and Yield Formation with Regard to Acid Invertase Activity in the Kernels. J. Agron. Crop Sci. 2015, 201, 353–367. [Google Scholar] [CrossRef]
- Maas, E.V.; Grieve, C.M. Spike and leaf development of salt-stressed wheat. Crop Sci. 1990, 30, 1309–1313. [Google Scholar] [CrossRef]
- Maas, E.V.; Lesch, S.M.; Francois, L.E.; Grieve, C.M. Tiller development in salt-stressed wheat. Crop Sci. 1994, 34, 1594–1603. [Google Scholar] [CrossRef]
- Khatun, S.; Flowers, T.J. Effects of salinity on seed set in rice. Plant Cell Environ. 1995, 18, 61–67. [Google Scholar] [CrossRef]
- Axelrod, J. Water Crisis in the Murray-Darling Basin: Australia Attempts to Balance Agricultural Need with Environmental Reality. Sustain. Dev. Law Policy 2012, 12, 13. [Google Scholar]
- Goss, K.F. Environmental flows, river salinity and biodiversity conservation: Managing trade-offs in the Murray–Darling basin. Aust. J. Bot. 2003, 51, 619–625. [Google Scholar] [CrossRef]
- Mullen, J.D.; Crean, J. Productivity Growth in Australian Agriculture: Trends, Sources, Performance; This Report Was Prepared for the Australian Farm Institute, the Rural Industries Research and Development Corporation, the Grains Research and Development Corporation, and Meat and Livestock Australia; Australian Farm Institute: Surry Hills, Australia, 2007. [Google Scholar]
- Wilson, S.M. Dryland and Urban Salinity Costs across the Murray-Darling Basin. An Overview and Guidelines for Identifying and Valuing the Impacts; Murray-Darling Basin Commission: Canberra, Australia, 2004. [Google Scholar]
- Connor, J.D.; Schwabe, K.; King, D.; Knapp, K. Irrigated agriculture and climate change: The influence of water supply variability and salinity on adaptation. Ecol. Econ. 2012, 77, 149–157. [Google Scholar] [CrossRef]
- Mickelbart, M.V.; Hasegawa, P.M.; Bailey-Serres, J. Genetic mechanisms of abiotic stress tolerance that translate to crop yield stability. Nat. Rev. Genet. 2015, 16, 237–251. [Google Scholar] [CrossRef] [PubMed]
- Pierik, R.; Testerink, C. The art of being flexible: How to escape from shade, salt, and drought. Plant Physiol. 2014, 166, 5–22. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Chen, Y.Q. A new mechanism in plant engineering: The potential roles of microRNAs in molecular breeding for crop improvement. Biotechnol. Adv. 2010, 28, 301–307. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B. MicroRNA: A new target for improving plant tolerance to abiotic stress. J. Exp. Bot. 2015, 66, 1749–1761. [Google Scholar] [CrossRef] [PubMed]
- Baumberger, N.; Baulcombe, D.C. Arabidopsis ARGONAUTE1 is an RNA Slicer that selectively recruits microRNAs and short interfering RNAs. Proc. Natl. Acad. Sci. USA 2005, 102, 11928–11933. [Google Scholar] [CrossRef] [PubMed]
- Jones-Rhoades, M.W.; Bartel, D.P.; Bartel, B. MicroRNAs and their regulatory roles in plants. Annu. Rev. Plant Biol. 2006, 57, 19–53. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R. Role of microRNAs in biotic and abiotic stress responses in crop plants. Appl. Biochem. Biotechnol. 2014, 174, 93–115. [Google Scholar] [CrossRef] [PubMed]
- Franck, A.; Guilley, H.; Jonard, G.; Richards, K.; Hirth, L. Nucleotide sequence of cauliflower mosaic virus DNA. Cell 1980, 21, 285–294. [Google Scholar] [CrossRef]
- Baker, C.C.; Sieber, P.; Wellmer, F.; Meyerowitz, E.M. The early extra petals1 mutant uncovers a role for microRNA miR164c in regulating petal number in Arabidopsis. Curr. Biol. 2005, 15, 303–315. [Google Scholar] [CrossRef] [PubMed]
- Fujii, H.; Chiou, T.J.; Lin, S.I.; Aung, K.; Zhu, J.K. A miRNA involved in phosphate-starvation response in Arabidopsis. Curr. Biol. 2005, 15, 2038–2043. [Google Scholar] [CrossRef] [PubMed]
- Guo, H.S.; Xie, Q.; Fei, J.F.; Chua, N.H. MicroRNA directs mRNA cleavage of the transcription factor NAC1 to downregulate auxin signals for Arabidopsis lateral root development. Plant Cell 2005, 17, 1376–1386. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.F.; Tian, Q.; Reed, J.W. Arabidopsis microRNA167 controls patterns of ARF6 and ARF8 expression, and regulates both female and male reproduction. Development 2006, 133, 4211–4218. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.C.; Yu, Y.; Wang, C.Y.; Li, Z.Y.; Liu, Q.; Xu, J.; Liao, J.Y.; Wang, X.J.; Qu, L.H.; Chen, F.; et al. Overexpression of microRNA OsmiR397 improves rice yield by increasing grain size and promoting panicle branching. Nat. Biotechnol. 2013, 31, 848–852. [Google Scholar] [CrossRef] [PubMed]
- Jian, C.; Han, R.; Chi, Q.; Wang, S.; Ma, M.; Liu, X.; Zhao, H. Virus-Based MicroRNA Silencing and Overexpressing in Common Wheat (Triticum aestivum L.). Front. Plant Sci. 2017, 8, 500. [Google Scholar] [CrossRef] [PubMed]
- Franco-Zorrilla, J.M.; Valli, A.; Todesco, M.; Mateos, I.; Puga, M.I.; Rubio-Somoza, I.; Leyva, A.; Weigel, D.; García, J.A.; Paz-Ares, J. Target mimicry provides a new mechanism for regulation of microRNA activity. Nat. Genet. 2007, 39, 1033–1037. [Google Scholar] [CrossRef] [PubMed]
- Todesco, M.; Rubio-Somoza, I.; Paz-Ares, J.; Weigel, D. A collection of target mimics for comprehensive analysis of microRNA function in Arabidopsis thaliana. PLoS Genet. 2010, 6, e1001031. [Google Scholar] [CrossRef] [PubMed]
- Vaistij, F.E.; Elias, L.; George, G.L.; Jones, L. Suppression of microRNA accumulation via RNA interference in Arabidopsis thaliana. Plant Mol. Biol. 2010, 73, 391–397. [Google Scholar] [CrossRef] [PubMed]
- Eamens, A.L.; Agius, C.; Smith, N.A.; Waterhouse, P.M.; Wang, M.B. Efficient silencing of endogenous microRNAs using artificial microRNAs in Arabidopsis thaliana. Mol. Plant 2011, 4, 157–170. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Gu, Y.; Jia, X.; Kang, W.; Pan, S.; Tang, X.; Chen, X.; Tang, G. Effective small RNA destruction by the expression of a short tandem target mimic in Arabidopsis. Plant Cell 2012, 24, 415–427. [Google Scholar] [CrossRef] [PubMed]
- Reichel, M.; Li, Y.; Li, J.; Millar, A.A. Inhibiting plant microRNA activity: Molecular SPONGEs, target MIMICs and STTMs all display variable efficacies against target microRNAs. Plant Biotechnol. J. 2015, 13, 915–926. [Google Scholar] [CrossRef] [PubMed]
- Wei, L.; Zhang, D.; Xiang, F.; Zhang, Z. Differentially expressed miRNAs potentially involved in the regulation of defense mechanism to drought stress in maize seedlings. Int. J. Plant Sci. 2009, 170, 979–989. [Google Scholar] [CrossRef]
- Kantar, M.; Lucas, S.J.; Budak, H. miRNA expression patterns of Triticum dicoccoides in response to shock drought stress. Planta 2011, 233, 471–484. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Liu, Y.; Liu, Z.; Kong, D.; Duan, M.; Luo, L. Genome-wide identification and analysis of drought-responsive microRNAs in Oryza sativa. J. Exp. Bot. 2010, 61, 4157–4168. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.H.; Tian, X.; Li, Y.J.; Wu, C.A.; Zheng, C.C. Microarray-based analysis of stress-regulated microRNAs in Arabidopsis thaliana. RNA 2008, 14, 836–843. [Google Scholar] [CrossRef] [PubMed]
- Ding, D.; Zhang, L.; Wang, H.; Liu, Z.; Zhang, Z.; Zheng, Y. Differential expression of miRNAs in response to salt stress in maize roots. Ann. Bot. 2009, 103, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Eren, H.; Pekmezci, M.Y.; Okay, S.; Turktas, M.; Inal, B.; Ilhan, E.; Atak, M.; Erayman, M.; Unver, T. Hexaploid wheat (Triticum aestivum) root miRNome analysis in response to salt stress. Ann. Appl. Biol. 2015, 167, 208–216. [Google Scholar] [CrossRef]
- Shen, J.; Xie, K.; Xiong, L. Global expression profiling of rice microRNAs by one-tube stem-loop reverse transcription quantitative PCR revealed important roles of microRNAs in abiotic stress responses. Mol. Genet. Genom. 2010, 284, 477–488. [Google Scholar] [CrossRef] [PubMed]
- Bakhshi, B.; Fard, E.M.; Gharechahi, J.; Safarzadeh, M.; Nikpay, N.; Fotovat, R.; Azimi, M.R.; Salekdeh, G.H. The contrasting microRNA content of a drought tolerant and a drought susceptible wheat cultivar. Plant Physiol. 2017, 216, 35–43. [Google Scholar] [CrossRef] [PubMed]
- Barrera-Figueroa, B.E.; Gao, L.; Wu, Z.; Zhou, X.; Zhu, J.; Jin, H.; Liu, R.; Zhu, J.K. High throughput sequencing reveals novel and abiotic stress-regulated microRNAs in the inflorescences of rice. BMC Plant Biol. 2012, 12, 132. [Google Scholar] [CrossRef] [PubMed]
- Noman, A.; Fahad, S.; Aqeel, M.; Ali, U.; Anwar, S.; Baloch, S.K.; Zainab, M. miRNAs: Major modulators for crop growth and development under abiotic stresses. Biotechnol. Lett. 2017, 39, 685–700. [Google Scholar] [CrossRef] [PubMed]
- Shriram, V.; Kumar, V.; Devarumath, R.M.; Khare, T.S.; Wani, S.H. MicroRNAs as potential targets for abiotic stress tolerance in plants. Front. Plant Sci. 2016, 7, 817. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Able, A.J.; Able, J.A. SMARTER de-stressed cereal breeding. Trends Plant Sci. 2016, 21, 909–925. [Google Scholar] [CrossRef] [PubMed]
- Alptekin, B.; Langridge, P.; Budak, H. Abiotic stress miRNomes in the Triticeae. Funct. Integr. Genom. 2017, 17, 145–170. [Google Scholar] [CrossRef] [PubMed]
- Aravind, J.; Rinku, S.; Pooja, B.; Shikha, M.; Kaliyugam, S.; Mallikarjuna, M.G.; Kumar, A.; Rao, A.R.; Nepolean, T. Identification, characterization, and functional validation of drought-responsive microRNAs in subtropical maize inbreds. Front. Plant Sci. 2017, 8, 941. [Google Scholar] [CrossRef] [PubMed]
- Bakhshi, B.; Fard, E.M.; Nikpay, N.; Ebrahimi, M.A.; Bihamta, M.R.; Mardi, M.; Salekdeh, G.H. MicroRNA signatures of drought signaling in rice root. PLoS ONE 2016, 11, e0156814. [Google Scholar] [CrossRef] [PubMed]
- Mittal, D.; Sharma, N.; Sharma, V.; Sopory, S.K.; Sanan-Mishra, N. Role of microRNAs in rice plant under salt stress. Ann. Appl. Biol. 2016, 168, 2–18. [Google Scholar] [CrossRef]
- Li, J.S.; Fu, F.L.; Ming, A.N.; Zhou, S.F.; She, Y.H.; Li, W.C. Differential expression of microRNAs in response to drought stress in maize. J. Integr. Agric. 2013, 12, 1414–1422. [Google Scholar] [CrossRef]
- Sunkar, R.; Zhou, X.; Zheng, Y.; Zhang, W.; Zhu, J.K. Identification of novel and candidate miRNAs in rice by high throughput sequencing. BMC Plant Biol. 2008, 8, 25. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Sun, Y.F.; Song, N.; Wei, J.P.; Wang, X.J.; Feng, H.; Yin, Z.Y.; Kang, Z.S. MicroRNAs involving in cold, wounding and salt stresses in Triticum aestivum L. Plant Physiol. Biochem. 2014, 80, 90–96. [Google Scholar] [CrossRef] [PubMed]
- Lescot, M.; Déhais, P.; Moreau, Y.; De Moor, B.; Rouzé, P.; Rombauts, S. PlantCARE: A database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef] [PubMed]
- Dolferus, R.; Klok, E.J.; Ismond, K.; Delessert, C.; Wilson, S.; Good, A.; Peacock, J.; Dennis, L. Molecular basis of the anaerobic response in plants. IUBMB Life 2001, 51, 79–82. [Google Scholar] [CrossRef] [PubMed]
- Knight, H.; Knight, M.R. Abiotic stress signalling pathways: Specificity and cross-talk. Trends Plant Sci. 2001, 6, 262–267. [Google Scholar] [CrossRef]
- Robert-Seilaniantz, A.; Grant, M.; Jones, J.D. Hormone crosstalk in plant disease and defense: More than just jasmonate-salicylate antagonism. Annu. Rev. Phytopathol. 2011, 49, 317–343. [Google Scholar] [CrossRef] [PubMed]
- Peleg, Z.; Blumwald, E. Hormone balance and abiotic stress tolerance in crop plants. Curr. Opin. Plant Biol. 2011, 14, 290–295. [Google Scholar] [CrossRef] [PubMed]
- Verma, V.; Ravindran, P.; Kumar, P.P. Plant hormone-mediated regulation of stress responses. BMC Plant Biol. 2016, 16, 86. [Google Scholar] [CrossRef] [PubMed]
- Siré, C.; Moreno, A.B.; Garcia-Chapa, M.; López-Moya, J.J.; Segundo, B.S. Diurnal oscillation in the accumulation of Arabidopsis microRNAs, miR167, miR168, miR171 and miR398. FEBS Lett. 2009, 583, 1039–1044. [Google Scholar] [CrossRef] [PubMed]
- Hazen, S.P.; Naef, F.; Quisel, T.; Gendron, J.M.; Chen, H.; Ecker, J.R.; Borevitz, J.O.; Kay, S.A. Exploring the transcriptional landscape of plant circadian rhythms using genome tiling arrays. Genome Biol. 2009, 10, 17. [Google Scholar] [CrossRef] [PubMed]
- Dong, Z.; Han, M.H.; Fedoroff, N. The RNA-binding proteins HYL1 and SE promote accurate in vitro processing of pri-miRNA by DCL1. Proc. Natl. Acad. Sci. USA 2008, 105, 9970–9975. [Google Scholar] [CrossRef] [PubMed]
- Han, M.H.; Goud, S.; Song, L.; Fedoroff, N. The Arabidopsis double-stranded RNA-binding protein HYL1 plays a role in microRNA-mediated gene regulation. Proc. Natl. Acad. Sci. USA 2004, 101, 1093–1098. [Google Scholar] [CrossRef] [PubMed]
- Park, W.; Li, J.; Song, R.; Messing, J.; Chen, X. CARPEL FACTORY, a Dicer homolog, and HEN1, a novel protein, act in microRNA metabolism in Arabidopsis thaliana. Curr. Biol. 2002, 12, 1484–1495. [Google Scholar] [CrossRef]
- Yang, L.; Liu, Z.; Lu, F.; Dong, A.; Huang, H. SERRATE is a novel nuclear regulator in primary microRNA processing in Arabidopsis. Plant J. 2006, 47, 841–850. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.K.; Chaabane, S.B.; Shah, P.; Poulsen, C.P.; Yang, S.W. COP1 E3 ligase protects HYL1 to retain microRNA biogenesis. Nat. Commun. 2014, 5, 5867. [Google Scholar] [CrossRef] [PubMed]
- Koornneef, M.; Alonso-Blanco, C.; Peeters, A.J.; Soppe, W. Genetic control of flowering time in Arabidopsis. Annu. Rev. Plant Biol. 1998, 49, 345–370. [Google Scholar] [CrossRef] [PubMed]
- Araki, T. Transition from vegetative to reproductive phase. Curr. Opin. Plant Biol. 2001, 4, 63–68. [Google Scholar] [CrossRef]
- Wang, J.W. The Multifaceted Roles of miR156-targeted SPL Transcription Factors in Plant Developmental Transitions. Plant Transcr. Factors 2015, 281–293. [Google Scholar]
- Fan, T.; Li, X.; Yang, W.; Xia, K.; Ouyang, J.; Zhang, M. Rice osa-miR171c mediates phase change from vegetative to reproductive development and shoot apical meristem maintenance by repressing four OsHAM transcription factors. PLoS ONE 2015, 10, e0125833. [Google Scholar] [CrossRef] [PubMed]
- Jung, C.; Pillen, K.; Staiger, D.; Coupland, G.; von Korff, M. Recent Advances in Flowering Time Control. Front. Plant Sci. 2017, 7, 2011. [Google Scholar] [CrossRef] [PubMed]
- He, H.; Serraj, R. Involvement of peduncle elongation, anther dehiscence and spikelet sterility in upland rice response to reproductive-stage drought stress. Environ. Exp. Bot. 2012, 75, 120–127. [Google Scholar] [CrossRef]
- Fujita, Y.; Fujita, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. ABA-mediated transcriptional regulation in response to osmotic stress in plants. J. Plant Res. 2011, 124, 509–525. [Google Scholar] [CrossRef] [PubMed]
- Harb, A.; Krishnan, A.; Ambavaram, M.M.; Pereira, A. Molecular and physiological analysis of drought stress in Arabidopsis reveals early responses leading to acclimation in plant growth. Plant Physiol. 2010, 154, 1254–1271. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi-Shinozaki, K.; Shinozaki, K. Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu. Rev. Plant Biol. 2006, 57, 781–803. [Google Scholar] [CrossRef] [PubMed]
- Roitsch, T. Source-sink regulation by sugar and stress. Curr. Opin. Plant Biol. 1999, 2, 198–206. [Google Scholar] [CrossRef]
- White, A.C.; Rogers, A.; Rees, M.; Osborne, C.P. How can we make plants grow faster? A source–sink perspective on growth rate. J. Exp. Bot. 2015, 67, 31–45. [Google Scholar] [CrossRef] [PubMed]
- Yu, S.M.; Lo, S.F.; Ho, T.H.D. Source–sink communication: Regulated by hormone, nutrient, and stress cross-signalling. Trends Plant Sci. 2015, 20, 844–857. [Google Scholar] [CrossRef] [PubMed]
- Kong, L.; Guo, H.; Sun, M. Signal transduction during wheat grain development. Planta 2015, 241, 789–801. [Google Scholar] [CrossRef] [PubMed]
- Austin, R.S.; Hiu, S.; Waese, J.; Ierullo, M.; Pasha, A.; Wang, T.T.; Fan, J.; Foong, C.; Breit, R.; Desveaux, D.; et al. New BAR tools for mining expression data and exploring Cis-elements in Arabidopsis thaliana. Plant J. 2016, 88, 490–504. [Google Scholar] [CrossRef] [PubMed]
- Landi, S.; Hausman, J.F.; Guerriero, G.; Esposito, S. Poaceae vs. Abiotic Stress: Focus on Drought and Salt Stress, Recent Insights and Perspectives. Front. Plant Sci. 2017, 8, 1214. [Google Scholar] [CrossRef] [PubMed]
- Lenka, S.K.; Katiyar, A.; Chinnusamy, V.; Bansal, K.C. Comparative analysis of drought-responsive transcriptome in Indica rice genotypes with contrasting drought tolerance. Plant Biotechnol. J. 2011, 9, 315–327. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Able, A.J.; Able, J.A. Genotypic water-deficit stress responses in durum wheat: Association between physiological traits, microRNA regulatory modules and yield components. Funct. Plant Biol. 2017, 44, 538–551. [Google Scholar] [CrossRef]
- Jiao, Y.; Song, W.; Zhang, M.; Lai, J. Identification of novel maize miRNAs by measuring the precision of precursor processing. BMC Plant Biol. 2011, 11, 141. [Google Scholar] [CrossRef] [PubMed]
- Fahlgren, N.; Feldman, M.; Gehan, M.A.; Wilson, M.S.; Shyu, C.; Bryant, D.W.; Hill, S.T.; McEntee, C.J.; Warnasooriya, S.N.; Kumar, I.; et al. A versatile phenotyping system and analytics platform reveals diverse temporal responses to water availability in Setaria. Mol. Plant 2015, 8, 1520–1535. [Google Scholar] [CrossRef] [PubMed]
- Furbank, R.T.; Tester, M. Phenomics–technologies to relieve the phenotyping bottleneck. Trends Plant Sci. 2011, 16, 635–644. [Google Scholar] [CrossRef] [PubMed]
| Mature miRNA | Number of Pre-miRNAs | Number of cis-Regulatory Elements in Promoter Region of PRE-MIRNA Encoding Sequence | Total | ||
|---|---|---|---|---|---|
| Hormone Related | Circadian Rhythm-Related | Abiotic Stress Related | |||
| Ath-miR159 | 3 (A-C) | 20 | 4 | 16 | 40 |
| Osa-miR159 | 6 (A-F) | 40 | 8 | 47 | 95 |
| Zma-miR159 | 8 (A-H) | 66 | 15 | 56 | 137 |
| Ath-miR164 | 3 (A-C) | 25 | 4 | 25 | 54 |
| Osa-miR164 | 6 (A-F) | 54 | 5 | 65 | 124 |
| Zma-miR164 | 8 (A-H) | 88 | 11 | 76 | 175 |
| Ath-miR167 | 4 (A-D) | 20 | 4 | 31 | 55 |
| Osa-miR167 | 10 (A-J) | 95 | 28 | 82 | 205 |
| Zma-miR167 | 4 (A-D) | 33 | 6 | 31 | 43 |
| Ath-miR168 | 2 (A-B) | 11 | 2 | 10 | 23 |
| Osa-miR168 | 1 (A) | 6 | 1 | 14 | 21 |
| Zma-miR168 | 2 (A-B) | 16 | 6 | 11 | 33 |
| Ath-miR396 | 2 (A-B) | 18 | 2 | 25 | 45 |
| Osa-miR396 | 3 (A-C) | 26 | 5 | 16 | 47 |
| Zma-miR396 | 2 (A-B) | 12 | 2 | 20 | 34 |
| Ath-miR397 | 2 (A-B) | 10 | 3 | 17 | 30 |
| Osa-miR397 | 2 (A-B) | 22 | 4 | 9 | 35 |
| Zma-miR397 | 2 (A-B) | 17 | 0 | 9 | 26 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pegler, J.L.; Grof, C.P.L.; Eamens, A.L. Profiling of the Differential Abundance of Drought and Salt Stress-Responsive MicroRNAs Across Grass Crop and Genetic Model Plant Species. Agronomy 2018, 8, 118. https://doi.org/10.3390/agronomy8070118
Pegler JL, Grof CPL, Eamens AL. Profiling of the Differential Abundance of Drought and Salt Stress-Responsive MicroRNAs Across Grass Crop and Genetic Model Plant Species. Agronomy. 2018; 8(7):118. https://doi.org/10.3390/agronomy8070118
Chicago/Turabian StylePegler, Joseph L., Christopher P. L. Grof, and Andrew L. Eamens. 2018. "Profiling of the Differential Abundance of Drought and Salt Stress-Responsive MicroRNAs Across Grass Crop and Genetic Model Plant Species" Agronomy 8, no. 7: 118. https://doi.org/10.3390/agronomy8070118
APA StylePegler, J. L., Grof, C. P. L., & Eamens, A. L. (2018). Profiling of the Differential Abundance of Drought and Salt Stress-Responsive MicroRNAs Across Grass Crop and Genetic Model Plant Species. Agronomy, 8(7), 118. https://doi.org/10.3390/agronomy8070118






