Abstract
Industries such as mining, cokemaking, (petro)chemical and electroplating produce effluents that contain free cyanide (fCN = HCN + CN−). Currently, fCN is mainly removed by (physico)chemical methods or by biotreatment with activated sludge. Cyanide hydratases (CynHs) (EC 4.2.1.66), which convert fCN to the much less toxic formamide, have been considered for a mild approach to wastewater decyanation. However, few data are available to evaluate the application potential of CynHs. In this study, we used a new CynH from Exidia glandulosa (protein KZV92691.1 designated NitEg by us), which was overproduced in Escherichia coli. The purified NitEg was highly active for fCN with 784 U/mg protein, kcat 927/s and kcat/KM 42/s/mM. It exhibited optimal activities at pH approximately 6–9 and 40–45 °C. It was quite stable in this pH range, and retained approximately 40% activity at 37 °C after 1 day. Silver and copper ions (1 mM) decreased its activity by 30–40%. The removal of 98–100% fCN was achieved for 0.6–100 mM fCN. Moreover, thiocyanate, sulfide, ammonia or phenol added in amounts typical of industrial effluents did not significantly reduce the fCN conversion, while electroplating effluents may need to be diluted due to high fCN and metal content. The ease of preparation of NitEg, its high specific activity, robustness and long shelf life make it a promising biocatalyst for the detoxification of fCN.
1. Introduction
Cyanide is widely used in industry due to its chelating and electrolytic properties. For example, it is used to leach gold and silver from their ores. These processes generate significant amounts of cyanide waste [1,2]. Cyanide wastewaters also come from, e.g., electroplating, jewelry industry, (petro)chemical industry and cokemaking [1,3,4,5,6,7].
Cyanide occurs as free cyanide (fCN), i.e., CN− and HCN, or as metal complexes. fCN is the most toxic; at pH ≤8.5, it occurs predominantly as volatile HCN, which is very hazardous [1]. The above industrial effluents also contain other chemicals such as sulfide, thiocyanate, ammonia, phenols or metals [4,5,6,7,8,9], which can complicate their treatment.
Organic pollutants have adverse effects on human health and the environment [10,11]. Cyanide is one of them, and is listed as toxic pollutant; according to the Environmental Protection Agency (EPA) effluent guidelines, the limit concentration of total cyanide is 1.30 mg/L for any one day in electroplating [12], while the limit for cokemaking is expressed in kg of total cyanide per ton of product (e.g., approximately 0.07 kg/t for any one day) [13].
The molecular mechanism of cyanide toxicity primarily consists in the formation of a complex with the iron cofactor in cytochrome c oxidase (EC 7.1.1.9), thereby affecting the respiratory chain. Consequently, multiple body systems (e.g., respiratory, cardiovascular, central nervous, sensory nervous, and endocrine) are severely affected by cyanide [14], with LD50 of 1–3 mg per kg of body weight (calculated for ingested HCN) [1]. In addition, aquatic ecosystems are extremely sensitive to low concentrations of fCN (≤0.01 mg/L and ≤0.15 mg/L for prolonged and acute exposure, respectively) [14].
Spontaneous processes based on volatilization, oxidation, precipitation, or microbial decomposition were traditionally used in the treatment of gold mine waste in “tailing ponds”. Later, the trend was toward more strictly controlled processes such as alkaline chlorination or oxidation with, e.g., SO2 + air (Inco process), or SO2 + air + H2O2 (CombinOx process) [1].
Nanomaterials are among the promising tools for eliminating or mitigating the harmful effects of environmental pollutants. For example, they were proposed for removing environmental contaminants such as industrial dyes [10,11,15], or nitrate ions [16]. Their uses for wastewater decyanation also emerged. These strategies used ZnO or Ga2O3/Pt nanoparticles acting as photocatalysts for the breakdown of fCN [17,18]. The current focus is on non-toxic nanoparticles produced through environmentally friendly methods, e.g., from biodegradable polymers [10,11].
Biological processes offer alternative ways to overcome the bottlenecks of the (physico)chemical approaches in terms of required space, time, energy or chemicals, or secondary waste. Microbial consortia (activated sludge) have already found full-scale applications in the treatment of cokemaking wastewaters [19].
In contrast, biocatalysts still have a very limited impact in this area, although cyanide hydratases (CynHs; EC 4.2.1.66) and cyanide dihydratases (CynDs; EC 3.5.5.1) seem to be promising tools for wastewater detoxification, as they catalyze the conversion of fCN to the less toxic formamide, and to a formic acid/ammonia mixture, respectively (Figure 1A). In particular, they are resistant to high fCN concentrations and do not require a cofactor. In addition, CynHs have the advantage of about 10-fold higher specific activity compared to CynDs [20]. CynHs also transform some nitriles such as 2-cyanopyridine (2CP), the products of which are the corresponding carboxylic acid and amide (Figure 1B).

Figure 1.
Transformation of (A) HCN by cyanide hydratase (CynH) or cyanide dihydratase (CynD), and (B) transformation of 2-cyanopyridine by CynH.
CynH has been considered a potential cyanide-degrading biocatalyst since it was first described in the fungus Stemphylium loti [21]. In the following years, other CynHs were obtained from Gloeocercospora sorghi [22], and genus Fusarium [23,24,25]. However, the CynH activities produced by the mycelia were mostly low: less than 0.05 U/mg dry weight in various fungal pathogens of plants [22] or 1.4 U/mg dry weight in Fusarium solani [24]. Only Fusarium lateritium showed a much higher activity of 102.5 U/mg dry weight [23] (Table S1). Specific activities typically ranged from 4.6 to 85 U/mg protein in cell-free extract (CFE) and increased to 128–1109 U/mg protein by purification [22,23,24,25] (Table S1).
The presence of a CynH was also suspected in the bacterium Serratia marcescens [26] after a reaction product corresponding to formamide was found in HPLC. The putative CynH sequence had a shorter amino acid chain than typical CynHs (326 residues). Similar proteins are widely distributed in the genus Serratia, other Proteobacteria, or Actinobacteria according to BLAST searching in the NCBI database [27].
In contrast, there are only a few characterized CynDs. They originate from Bacillus pumilus, Pseudomonas stutzeri and Alcaligenes xylosoxidans [28] and their homologues seem to occur only in bacteria.
In the last two decades, several fungal CynHs have been obtained, whose genes are expressed in Escherichia coli [3,29,30,31,32,33]. The activities of the whole E. coli cells (153–600 U/mg dry weight) and CFE (736 U/mg protein) [33] were substantially higher in comparison with the endogenous enzymes, whereas the activities of the purified enzymes from the homologous and heterologous producers were similar (100–1324 U/mg protein; Table S1) [29,31,33]. Further enhancement of CynH action on fCN may be possible by the specific mutations recently proposed in silico [34].
However, the extent to which CynHs or CynDs can degrade fCN in different environments has scarcely been studied. Thus, fCN conversion by four recombinant CynHs was demonstrated for diluted Ag and Cu electroplating effluents containing 100 mM fCN [3]; it was almost complete in the presence of Ag but less than 70% in the presence of Cu. Another CynH was used with a spiked cokemaking effluent and degraded, e.g., 10 mM fCN to almost 100% [4]. In addition, an immobilized whole-cell CynD biocatalyst was used with diluted gold-mine wastewater and the maximum conversion was 98% starting from 17.6 mM fCN [2].
Recently, we have investigated Basidiomycota as a source of enzymes of the nitrilase superfamily, which include some putative CynHs [32,35]. The focus of this work is on a detailed study of one of these CynHs, which originates from the fungus Exidia glandulosa. The aim of this work was to determine its properties that are important for its future use: kinetic parameters, pH and temperature profiles, stability, and ability to remove cyanide under conditions relevant to wastewater treatment. Therefore, the degree of fCN conversion was studied at typical cyanide concentrations, alkaline pH (necessary to minimize the escape of highly toxic HCN), and the presence of contaminants accompanying cyanide in industrial effluents. Further steps toward the use of CynHs in wastewater treatment are proposed.
2. Results
2.1. Enzyme Preparation and Properties
The sequence of enzyme NitEg was taken from databases (GenBank: KZV92691; UniProtKB: A0A165HZS1), where this protein was classified as “cyanide hydratase”. The corresponding gene (nitEg) was synthesized, with codon usage bias optimized for E. coli (Figure S1).
2.1.1. Enzyme Overproduction
E. coli containing the optimized nitEg gene was grown to an optical density (determined at 600 nm; OD600) of 12.0, which corresponds to approximately 3.6 mg dry weight/mL. Total whole-cell activity was typically 318 U/mL culture medium (88.4 U/mg dry weight), as determined by the rate of fCN consumption. Cells from 200 mL of culture were sonicated to obtain CFE containing 9.3 mg protein/mL (214 mg protein in total) (Table S2). Analysis of the CFE by SDS-PAGE (Figure S2) revealed the major band to have an apparent molecular mass of approximately 43.85 kDa, which was essentially consistent with the theoretical molecular mass of NitEg carrying a His6-tag (41,800 Da). The CFE had a specific activity of 280 U/mg as determined by the picric acid method (Table S2).
2.1.2. Enzyme Purification and Activity
NitEg was purified in a single step. The amount of the purified enzyme was typically 12 mg from 200 mL of culture. The purification yield was 14.4% (Table S2) and the amount of NitEg accounted for approximately 40% of the total cell protein. Analysis of the purified NitEg by SDS-PAGE showed that the protein contained no significant impurities (Figure S2).
Formamide was the main product of fCN conversion by the purified enzyme (Figure 2). The concentration of formamide in the reaction mixture was about 22 mM after complete elimination of fCN (60 min). It was similar to the concentration of fCN in the control sample without enzyme after the same time. This suggests that the remainder of fCN (about 3 mM) was abiotically removed in both the reaction and the control.
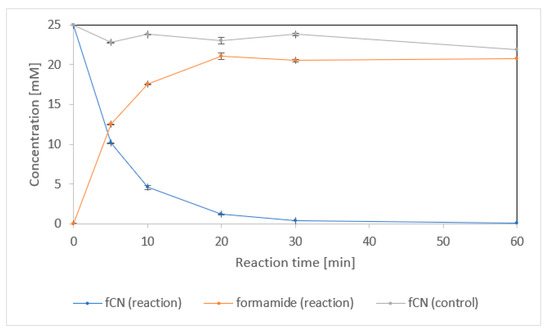
Figure 2.
Transformation of 25 mM free cyanide (fCN) to formamide by enzyme NitEg (4.0 µg protein/mL) in 100 mM glycine/NaOH buffer, pH 9.0. Residual fCN concentration was determined by picric acid method. The enzyme was omitted from the control.
The specific activities calculated from the rates of fCN consumption and formamide production were approximately 697 U/mg and 784 U/mg protein, respectively. The kinetic parameters were calculated from the rates of formamide production (Table 1). The values of Vmax and kcat were relatively high, but KM was also high. Nevertheless, the kcat/KM ratio indicates good catalytic efficiency of the enzyme.

Table 1.
Specific activity and kinetic parameters of enzyme NitEg.
2.1.3. Effect of pH and Temperature
The pH and temperature profiles of NitEg were determined using 2CP as substrate. This was because 2CP is less sensitive to pH and temperature than fCN. 2CP was generally a good substrate for the CynHs we previously studied [32,33]. NitEg also showed acceptable activity for 2CP. The activity was expressed as the sum of the reaction products pyridine-2-carboxylic acid and pyridine-2-carboxamide (Figure 1B), and was determined to be approximately 6.6 U/mg. NitEg was most active at 40–45 °C (Figure 3A) and exhibited a broad optimum at pH approximately 6–9 (Figure 3B). Its activity decreased significantly outside this pH range and at 50 °C. The stability of the enzyme was acceptable in a similar pH range (Figure 3B).
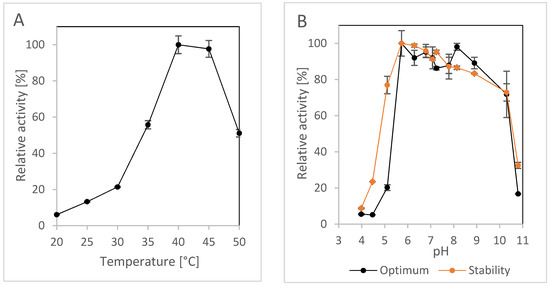
Figure 3.
Effect of (A) temperature on the activity of enzyme NitEg and (B) pH on the activity and stability of NitEg. 2-Cyanopyridine (25 mM) was used as substrate.
2.1.4. Effect of Silver and Copper
The effect of metal ions (Cu2+, Ag+) prevalent in some electroplating baths [3] on the activity of NitEg was studied by measuring the rate of fCN consumption at pH 9.0 and 30 °C (Table 2). Silver ions decreased the activity by about 1/3 at 0.1–1 mM and by 2/3 at 5 mM. No enzyme activity was detected at 10 mM concentration of silver ions. However, it should be noted that the concentration of fCN was also decreased in the controls (without enzyme) in the presence of silver (to about 15.7 mM and 5.7 mM at 5 mM and 10 mM Ag, respectively), apparently due to complex formation. A strong inhibition effect of Cu was observed at 5–10 mM.

Table 2.
Effect of copper and silver ions on the activity of enzyme NitEg.
2.1.5. Temperature stability and shelf life
At temperatures selected according to the previous work (27–50 °C) [3], the enzyme was pre-incubated for 24 h and the residual activities were determined using fCN as substrate. Under these conditions, the stability of the purified NitEg was acceptable at 27 °C or 37 °C, with about 60% and 40% of its initial specific activity, respectively, after 24 h. At 43 °C, it retained approximately 40% of its initial activity after 3 h, but only 10% of it after 24 h, and it lost more than 90% of its activity at 50 °C after 1 h (Figure 4).
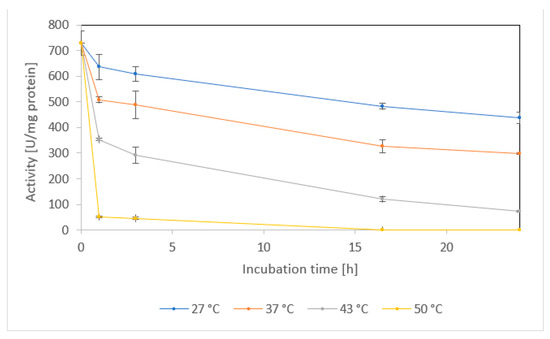
Figure 4.
Temperature stability of enzyme NitEg. The purified enzyme (10.3 mg protein/mL) was incubated in 50 mM Tris/HCl buffer, pH 8.0, with 150 mM NaCl, at different temperatures without shaking. Specific activities were determined by the formamide assay.
At 4 °C, the specific activity of purified NitEg (10.3 mg protein/mL) showed no statistically significant loss until day 69. Thereafter, the activity decreased, but was still almost 83% of the initial value after 98 days (Figure S3).
2.2. Enzyme Performance on Model Mixtures
The performance of purified NitEg was studied using model solutions that were prepared based on published wastewater analyses (see below). They mimicked the effluents of various industries in terms of concentrations of fCN and other contaminants, as well as pH.
2.2.1. Effect of Alkaline Media
First, the performance of NitEg was investigated with fCN at pH 9.0–10.5 (Figure 5). At pH 9.0, the removal of 25 mM fCN by 4 µg enzyme/mL was almost complete after 1 h. The abiotic removal accounted for approximately 10% within the same time. Higher pH resulted in less efficient fCN removal (approximately 89% and 54% at pH 9.5 and 10.0, respectively) under the same conditions. At pH 10.5, the enzyme was no more active in removing fCN (Figure 5A).
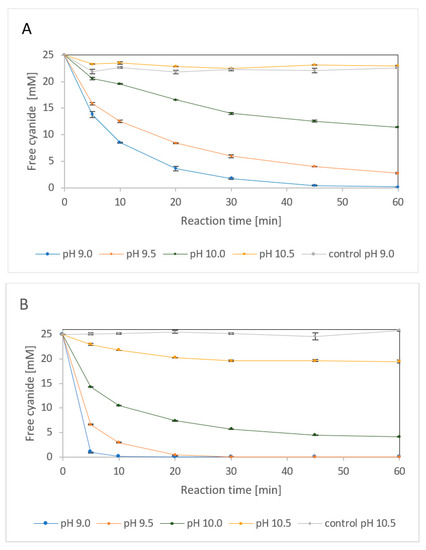
Figure 5.
Effect of pH on the biocatalyzed degradation of 25 mM free cyanide (fCN) in 100 mM glycine/NaOH buffer by (A) 4.0 µg enzyme/mL and (B) 20 µg enzyme/mL. Controls were carried out at (A) pH 9.0 and (B) pH 10.5 without enzyme. Residual fCN concentration was determined by picric acid method.
Increasing the enzyme concentration to 20 µg/mL allowed removal of nearly 25 mM fCN at pH 9.0 or 9.5 after 30 min and over 80% fCN at pH 10 after 1 h. Some decrease in fCN concentration was observed even at pH 10.5 (Figure 5B). This suggested that the enzymatic reaction must proceed quite rapidly to achieve a high degree of fCN removal. This is probably due to the limited stability of the enzyme under the reaction conditions as indicated by the pH profile of enzyme stability (Figure 3B).
2.2.2. Enzyme Performance in the Presence of Phenol and Inorganic Salts
Cokemaking effluents typically contain ≤ 0.6 mM fCN, but occasionally up to 10 mM fCN. Their common constituents also include phenols, thiocyanate, and ammonia. A model mixture was prepared according to the reported content [8] of fCN, SCN−, NH4+ and phenol in cokemaking effluents (Figure 6A). The upper limits of the concentrations were used, where the fCN concentration was 0.6 mM, and the pH was 9.1. Under these conditions, almost all of the fCN was degraded with 2.5 μg enzyme/mL after 90 min, with a negligible degree of abiotic elimination (Figure 6A).
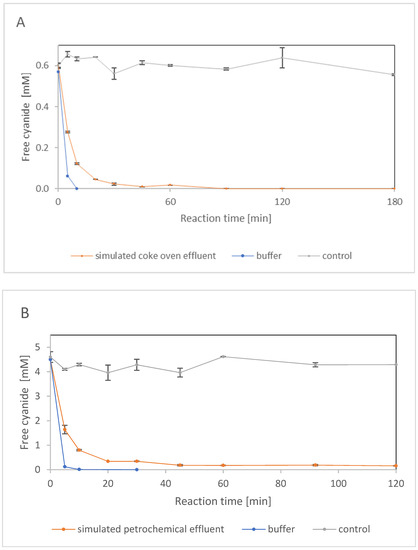
Figure 6.
Biocatalyzed degradation of free cyanide (fCN) by enzyme NitEg in 100 mM glycine/NaOH, pH 9.1, containing (A) 0.6 mM fCN, 8.6 mM SCN−, 10.7 mM NH4+ and 12.8 mM phenol (model coke plant effluent) [8], and (B) 4.6 mM fCN, 23.4 mM S2−, 2.5 mM NH4+ and 0.64 mM phenol (model petrochemical effluent) [5]. Reactions were also performed in the same buffer with (A) 0.6 mM fCN or (B) 4.6 mM fCN without other additives. Enzyme concentration was (A) 2.5 μg/mL, and (B) 5.0 μg/mL. The enzyme was omitted from the controls. Residual fCN concentration was determined by (A) picric acid method or (B) Spectroquant® kit.
A model petrochemical effluent was also prepared according to the literature [5]. Compared to the previous effluent, this contained almost eight times more fCN but less phenol or ammonia. It contained no SCN−, but a significant concentration of S2−; its pH was the same (9.1) (Figure 6B). The presence of S2− (interfering compound [36]) did not allow the use of the picric acid method to determine fCN. This interference caused an attenuation of the signal at 0.029 mM (1 ppm) and 0.29 mM H2S by 2.5% and 27.5%, respectively. In contrast, 2.9 mM and 29 mM H2S increased the signal more than twofold and twentyfold, respectively [36]. Therefore, the fCN concentration in the model effluent containing more than 20 mM S2− was monitored using the Spectroquant® kit, which is based on a different reagent (1,3-dimethylbarbituric acid). However, according to the manufacturer, this kit is not suitable for samples containing SCN−.
An enzyme concentration of 5 μg/mL was sufficient to remove approximately 96% fCN after 45 min. No further decrease in fCN concentration could be detected, which may have been caused by the background signal generated by one or more components of the mixture (Figure 6B).
2.2.3. Enzyme Performance in the Presence of Heavy Metal Salts
The fCN-containing effluents from electroplating and precious metal mining contain heavy metals. The two electroplating wastes used in a previous work [3] contained undefined amounts of Cu or Ag. Another one contained about 30 mM Cu, 83 mM Ni and 0.9 mM Zn [37]. The fCN concentrations in these effluents can be extremely high (approximately 0.5–2 M) [2,3,37]. The CynD was not sufficiently effective in the original gold-mine effluent (528 mM fCN) and only removed 43% of the fCN [2]. Therefore, it was necessary to dilute this solution to 17.6 mM fCN [2] before applying the enzyme. Similarly, the electroplating wastewater was diluted to 100 mM fCN [3]. This can also cause the metal concentration to drop to a level tolerated by CynH. The above experiments have shown that NitEg does not work well at Ag or Cu concentrations of about 5 mM or more (Section 2.1.4).
First, the potential of NitEg to degrade 100 mM fCN was investigated (Figure 7A). Here, the pH of the model mixture was adjusted to 9.0, at which the stability of the fCN solutions was higher than at pH 8 used previously [3]. Different amounts of enzyme (14, 20, or 30 μg/mL) eliminated more than 97% from 100 mM fCN within 1 h. However, more than 2% of fCN remained in the reaction mixture within the monitored period (3 h) regardless of enzyme loading. The abiotic loss of fCN was about 17%.
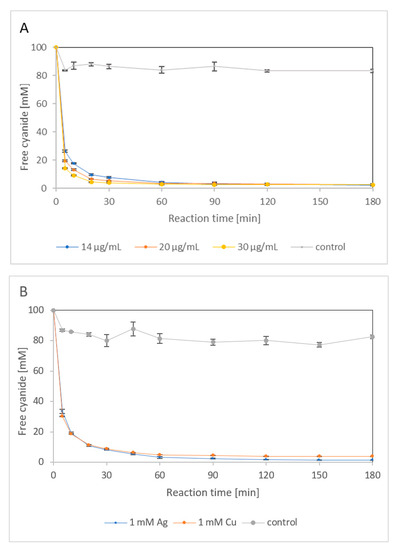
Figure 7.
Biocatalyzed degradation of 100 mM free cyanide (fCN) in 100 mM glycine/NaOH, pH 9.0, by (A) 14, 20 or 30 μg enzyme/mL, and (B) 14 μg enzyme/mL in the presence of 1 mM AgNO3 or 1 mM CuSO4. Residual fCN concentration was determined by picric acid method. The enzyme and the metal salts were omitted from the control.
The simulated electroplating effluents contained 100 mM fCN and 1 mM AgNO3 or 1 mM CuSO4. The reactions were catalyzed by 14 μg enzyme/mL, which was sufficient to remove more than 97% fCN in the metal-free medium after 1 h (Figure 7A). The metal ions slightly decreased the initial reaction rates (Figure 7B), which was consistent with their effects on NitEg activity (Section 2.1.4). However, the percentage of fCN removal (96–98%) was similar to that in the absence of both of these metals. A copper concentration of 10 mM completely inhibited the catalytic process (not shown), which is also consistent with the results of the inhibition experiments above. The reaction in the presence of 10 mM silver was not studied because a metal cyanide complex was formed, which reduced the amount of fCN available to the enzyme (Section 2.1.4).
3. Discussion
The presence of a large number of cynH genes in Ascomycota compared to Basidiomycota was previously demonstrated in our database searches [31,32]. The enzyme NitEg studied in this work is the first purified CynH from Basidiomycota. NitEg has an amino acid sequence identity of 86.29% (95% coverage) with CynH from Neurospora crassa (GenBank: XP_960160.2; UniProtKB: Q7RVT0) but NitEg is larger with its 366 amino acid residues (compared with 351 in N. crassa) (Figure S4). The latter enzyme was previously purified [3]. In this work, we compare the properties of NitEg and the CynH from N. crassa as far as the published data allow.
NitEg was produced in E. coli following the general protocol we developed for the production of nitrilases (EC 3.5.5.1) and the related CynHs [31]. The yield of purification of NitEg was about 14.4%, which is similar to the yield of CynH from Aspergillus niger (17%) [33]. However, the purification of NitEg (a one-step protocol) was simpler due to the presence of the His6-tag. We hypothesize that further optimization of the purification protocol can improve the yield of NitEg. Nevertheless, the amount of purified NitEg (approximately 12 mg) obtained from 200 mL of culture was sufficient for the aims of this work.
The CynH producing fungi F. solani and Fusarium oxysporum were cultivated at 10-L [24] or 45-L [25] scale, respectively. In contrast, heterologous production was performed at a small scale [3,33]. However, a related enzyme, nitrilase, was produced heterologously at 300-L scale. The growth medium was optimized and contained lactose as inducer. Under these conditions, the biomass yield was almost 13 g dry weight/L [38]. Therefore, we assume that a future scale-up of the production process of NitEg is very feasible using this method. It follows from the comparison of the specific activities of both enzymes (almost 500-fold higher in the CynH) that, after a suitable scale-up, the CynH production in the order of 106–107 U/L may be possible.
The use of whole cells of a recombinant organism for wastewater treatment could face regulatory obstacles, as it might be difficult to exclude cell escape. Enzyme purification would increase the cost of the catalyst but the use of CFE can be a suitable solution, as CFE has high specific activity and can be made free of residual whole cells.
Most CynHs are closely related with more than 60% sequence identity, but they differ in their specific activities (Table S1). It is not entirely clear to what extent these results are influenced by different experimental conditions. Differences in pH profiles were also found between CynHs studied by the same method [3]. The operation of CynHs in alkaline media is of paramount importance, since similar conditions occur in industrial wastewaters. In this work, we investigated whether NitEg was able to fully remove fCN from alkaline media. It was found that this was possible for 25 mM fCN at pH 9–9.5, while more than 80% was still removed at pH 10.0. This was consistent with the pH activity profile of the enzyme with an optimum at pH approximately 6–9.
Alternative biocatalysts such as CynDs were investigated for the degradation of fCN in alkaline media [2,28,39,40]. However, even the improved CynDs still lagged behind CynHs such as NitEg, which had activity greater than 600 U/mg at pH 9.0. Moreover, at alkaline pH, NitEg remained active long enough to almost completely remove high concentrations of fCN. Since immobilization of CynD was shown to have a stabilizing effect under alkaline conditions [2], we hypothesize that it may have a positive effect on the operation of CynHs as well.
The studies of two CynDs and the CynH from N. crassa showed the importance of the C-terminus for the thermostability of both types of enzymes [28,40]. The CynH was more thermostable than CynDs with almost 90% activity after 4 h at 42 °C [40], and about 50% activity after 1 day at 43 °C [3]. In contrast, the CynHs from G. sorghi and Aspergillus nidulans retained very low residual activities under the same conditions [3]. The latter has a very similar sequence to the CynH from A. niger [33]. The CynH from A. niger was also investigated for stability, but using a different method (1 h preincubation), and was found to be unstable above 35 °C. The stability of NitEg at 43 °C was intermediate, with the enzyme retaining 10% of its initial activity after 1 day. However, significantly higher residual activities were observed after the same time at 27 or 37 °C. The activity and stability of NitEg decreased sharply at 50 °C. Therefore, this enzyme probably cannot be used under these conditions.
The kinetic parameters Vmax and KM were previously reported for the CynHs from G. sorghi (His6-tagged) [41] and A. niger (without the tag) [33]. Both Vmax and KM were very high in these enzymes (Vmax = 4400 U/mg and 6800 U/mg, KM = 90 mM and 109 mM, respectively). In NitEg, both parameters were lower (Vmax = 1335 U/mg, KM = 22 mM). Thus, the Vmax/KM ratio (U/mg/mM) was 60 in NitEg and similar in A. niger CynH (62) but lower in G. sorghi CynH (49). In contrast, KM values were 2.6–7.3 mM in the CynDs, with Vmax of 88–100 U/mg [41] and Vmax/KM ratio of 14–34 U/mg/mM.
The ability of NitEg to remove low concentrations of fCN did not appear to be significantly affected by the relatively high KM. No residual fCN was found in the NitEg catalyzed degradation of 0.6 mM fCN. The reason for the incomplete (but still 96–98%) conversion of 100 mM fCN was probably the enzyme inactivation.
No entries for kcat or kcat/KM were found for CynHs in the BRENDA database [42]. Here, they were calculated for the CynHs with known Vmax (see above) and subunit molecular weights, which were similar in all these enzymes (approximately 40–42 kDa). Thus, the CynHs from G. sorghi and A. niger, and NitEg exhibit kcat (1/s) of approximately 3000, 4500 and 927, respectively, and kcat/KM (1/s/mM) of approximately 33, 41 and 42, respectively. For the related nitrilases, the kcat/KM (1/s/mM) reported in the BRENDA database ranged from approximately 0.001 to 205 [42]. The highest value was found for nitrilase from Pyrococcus sp. for benzonitrile [43], while all other values were lower than for NitEg.
There are few examples demonstrating the use of CynHs or CynDs in real or simulated industrial effluents (Table S3). Four CynHs were used to degrade 100 mM fCN in the presence of Ag or Cu, or their absence [3]. The concentrations of the CynHs were 7–13 µg/mL in the reaction mixtures with Ag or the control (without metals), but they were increased 10-fold in the mixtures with Cu. The enzymes showed large differences in their reaction rates. However, complete degradation of fCN was only achieved in buffer, whereas in the presence of Ag and Cu approximately 90% and 70% of fCN was degraded, respectively. It is difficult to compare the performance of these enzymes with that of NitEg because the composition of the electroplating effluents used in the previous work is largely unknown (not even the metal content), and the pH of the reaction mixture was lower (about 8) than for NitEg (9.0).
Nevertheless, it can be stated with some caution that the rapid degradation of 100 mM fCN in the absence of metals was similar for CynH from N. crassa (best of four CynHs tested) [3] and NitEg (98–100% conversions after 1–2 h). In contrast, the other enzymes required 12 to 48 h for similar conversions [3]. In addition, we characterized NitEg with respect to properties not reported for the N. crassa CynH: kinetic parameters, shelf life, and fCN conversion in different environments. The properties found for NitEg were favorable and suggest that this enzyme may be superior in terms of its ability to catalyze fCN degradation.
The evaluation of NitEg suggests that the enzyme is unlikely to be significantly inhibited by the major constituents of industrial effluents (phenol, ammonia, sulfide or thiocyanate). However, cokemaking wastewater, e.g., contains a large number of minor organic constituents, which can be expressed collectively as chemical oxygen demand (COD) or biological oxygen demand (BOD) [44]. This is difficult to simulate in model mixtures. NitEg is also likely to be functional at certain concentrations of heavy metals (e.g., 1 mM of Ag or Cu). Although electroplating or gold mining wastewaters may contain higher concentrations of metals, they can be suitably diluted for enzyme action. These types of effluent are also more complex than the model mixtures, also containing other metals and metal cyanide complexes. Therefore, the aim of the future research will be the evaluation of NitEg or other cyanide-degrading biocatalysts in real wastewaters. Importantly, detailed analysis must be available for these wastewaters to ensure reproducibility, reliability and comparability of the data but this has rarely been the case in previous studies.
4. Materials and Methods
4.1. Chemicals
Chemicals were obtained from standard suppliers and were of the highest purity: acetic acid (99%; Lach-ner, Neratovice, Czech Republic), acetonitrile (≥99.95%, HiPerSolv Chromanorm; VWR International, Radnor, PA, USA), ammonium chloride (≥99%; Lach-ner), boric acid (≥99.5%; Lach-ner), copper (II) sulphate (≥99%; Lach-ner), 2-cyanopyridine (99%; Sigma Aldrich, St. Louis, MO, USA), disodium hydrogen phosphate (≥99.6%; Lach-ner), ferric chloride (98%; Alfa Aesar, Haverhill, MA, USA), formamide (≥99%; VWR), glycine (≥99%; Reanal, Budapest, Hungary), hydrochloric acid (35%; Lach-ner), hydroxylamine hydrochloride (99%; Alfa Aesar), imidazole (research grade; Serva Electrophoresis GmbH, Heidelberg, Germany), isopropyl-β-D-thiogalactopyranoside (IPTG; research grade; Serva), methanol (100%; VWR), phenol (≥99%; VWR), phenylmethanesulfonyl fluoride (PMSF; Serva), picric acid (≥98%; Sigma Aldrich), phosphoric acid (≥85%; VWR), potassium cyanide (≥96%; Sigma Aldrich), potassium thiocyanate (≥99%; Lach-ner), 2-pyridinecarboxamide (98%; Sigma Aldrich), 2-pyridinecarboxylic acid (≥98%; Sigma Aldrich), silver nitrate (p.a.; Lach-ner), sodium carbonate (≥99%; VWR), sodium chloride (≥98%; Lach-ner), sodium dihydrogen phosphate (≥99%; Lach-ner), sodium dodecyl sulfate (SDS; ≥99%; Carl Roth GmbH + Co. KG, Karlsruhe, Germany), sodium hydroxide (≥98 %; Lach-ner), sodium sulfide (Alfa Aesar), and tris(hydroxymethyl)aminomethane (Tris; ≥99%; Merck KGaA, Darmstadt, Germany).
4.2. Strain
The gene encoding protein KZV92691.1 (NitEg) was optimized (Figure S1), synthesized and ligated into vector pET22b(+) by GeneArt (ThermoFisher Scientific, Waltham, MA, USA). The gene was fused to C-terminal His6-tag coding sequence. The resulting construct was used to transform E. coli Origami B(DE3) cells (Merck).
4.3. Enzyme Induction and Purification
The transformed cells were grown in 2xYT medium (g/L: Tryptone (ThermoFisher Scientific) 16, yeast extract (ThermoFisher Scientific) 10, NaCl 5; pH 7.0) at 37 °C. After the OD600 reached 1.0 mM IPTG was added to a final concentration of 0.02 mM, and cultivation continued for 20 h at 20 °C. Harvested cells were sonicated and NitEg was purified from the CFE by cobalt ion affinity chromatography on TALON® Metal Affinity Resin (Clontech Laboratories, Inc., Mountain View, CA, USA) as previously described for nitrilases [35], but with minor modifications: NitEg was eluted with 20 mM sodium phosphate buffer, pH 7.6, containing 300 mM NaCl, 0.1mM PMSF and 200 mM imidazole. The enzyme solution was concentrated using Amicon Ultra-30 K filter (Merck, Millipore, Burlington, MA, USA) and the buffer was replaced with 50 mM Tris/HCl buffer, pH 8.0, supplemented with 150 mM NaCl.
4.4. Enzyme Assays
The standard activity assay was performed using Eppendorf ThermoMixer Comfort (Eppendorf, Hamburg, Germany) at 30 °C and 850 rpm. Reaction mixtures (0.5 mL) in 1.5 mL Eppendorf tubes contained 100 mM glycine/NaOH buffer, pH 9.0, and an appropriate amount of whole cells, CFE, or purified enzyme. After 5 min of pre-incubation, the reaction was started by adding KCN at a final concentration of 25 mM. After 1–2 min of incubation, the reaction was stopped by adding 1 mL of methanol. Whole cells were removed by centrifugation (MiniSpin plus, Eppendorf; 14,500 rpm, 4 min). Centrifugation was omitted for samples containing CFE or purified enzyme. The residual fCN was determined spectrophotometrically by the picric acid method [36] with some modifications. Samples (0.01 mL) were mixed with 0.09 mL of 0.2 M Tris/HCl buffer, pH 8.0, and 0.2 mL of 0.5% picric acid in 0.25 M Na2CO3, and incubated at 100 °C in a water bath for 5 min. The reaction was stopped by placing the test tube in an ice bath. After addition of 0.7 mL of distilled water, absorbance was determined at 520 nm against the control prepared in the same way but without fCN.
Formamide was determined spectrophotometrically [23] with some modifications. Samples (0.2 mL) were mixed with 0.4 mL of 2.3 M hydroxylamine/3.5 M NaOH (1/1) and incubated at 60 °C for 10 min. After the addition of 0.2 mL of 4 M HCl and 0.2 mL of 1.23 M FeCl3, absorbance was determined at 540 nm against the control prepared in the same way but without formamide.
One unit of CynH activity was defined as the amount of enzyme that consumed 1 µmol of fCN/min or produced 1 µmol of formamide/min under the above conditions.
Enzyme kinetics was investigated using 2.5–25 mM KCN, and activities were determined using the formamide assay.
The thermostability of the purified enzyme was studied according to the literature [3] with some modifications. The enzyme was incubated at 27, 37, 43 or 50 °C without shaking, and its specific activity was determined after 1, 3, 16.5 and 24 h using the formamide assay. The shelf life of the purified enzyme at 4 °C was investigated within 98 days by periodic determination of its specific activity using the picric acid method.
The activity for 2-cyanopyridine (2CP) was determined analogously as that for fCN but using 50 mM Tris/HCl buffer, pH 8.0, with 150 mM NaCl, and the reaction was stopped after 10 min by adding 0.05 mL of 2M HCl per 0.5 mL of sample. The reaction mixtures were centrifuged and supernatants were used for HPLC analysis. The temperature optimum was determined at 20–50 °C. The pH optimum was determined using Britton–Robinson (CH3COOH/H3BO3/H3PO4/NaOH) buffers, pH 4.0–10.8, at 30 °C. The pH stability was determined by pre-incubating the enzyme in the same buffers for 2 h at 30 °C followed by determination of the residual activity at pH 8.0 and 30 °C. The concentrations of substrate and reaction products (2-pyridinecarboxylic acid and 2-pyridinecarboxamide) were determined by reversed-phase HPLC (column ACE C8, 5 µm, 250 mm × 4 mm (Advanced Chromatography Technologies Ltd, Aberdeen, UK); mobile phase 10% acetonitrile in 5 mM sodium phosphate buffer, pH 7.2, flow rate 0.9 mL/min).
4.5. General Protocol for the Biocatalyzed Degradation of fCN
Reaction mixtures (1 mL) in 2-mL Eppendorf tubes were incubated in Eppendorf ThermoMixer Comfort (30 °C, 850 rpm). Simulated effluents were prepared according to published data [5,8]. The fCN concentrations were 0.6, 4.6, 25 or 100 mM fCN. Optionally the reaction mixtures were supplemented with additives (Na2S, KSCN, NH4Cl, phenol, CuSO4, AgNO3; see Figure 6 and Figure 7 for details). The buffer was 100 mM glycine/NaOH (pH 9.0, 9.5, 10.0, or 10.5). Samples (0.1 mL) were withdrawn at various intervals (5–180 min) and the reaction was stopped with 0.2 mL methanol. The fCN concentrations were determined using the picric acid method (Section 4.4) or using the Spectroquant® kit (Merck).
5. Conclusions
In this work, proof-of-concept was provided for the performance of NitEg, a unique CynH from the Basidiomycota division, in fCN solutions of up to 100 mM concentrations at alkaline conditions. This CynH was studied as a potential biocatalyst using model mixtures prepared according to the literature. This ensures that the results obtained are reproducible and comparable with other biocatalysts used in the same way in the future. However, further investigation of NitEg or other CyHs in real wastewater will be necessary in the future, preferably with effluents that have been analyzed in detail. This work has shown that the enzyme is highly active, resistant to alkaline pH, tolerates high concentrations of fCN, and other industrial contaminants, and can be stored for extended periods under standard conditions. The enzyme is formed at high expression levels and its production is likely to be readily scalable. Immobilization can be useful to increase its robustness and enable its reuse. Other potential applications of NitEg or CynHs in general could be cyanide biosensors, some of which have been based on CynDs [45]. Importantly, NitEg is unique with respect to its origin, as CynHs are rare in Basidiomycota. Therefore, it may be of interest to study its natural role and potential impact on the interaction between E. glandulosa and its plant hosts.
Supplementary Materials
The following are available online at https://www.mdpi.com/article/10.3390/catal11111410/s1, Figure S1. Optimized sequence of the gene encoding NitEg. Figure S2. (A) SDS-PAGE of purified NitEg. (B) Determination of enzyme molecular mass. Figure S3. Shelf life of NitEg at pH 8.0 and 4 °C. Figure S4. Multiple sequence alignment of NitEg (UniProtKB: A0A165HZS1) and its closest characterized homologue from Neurospora crassa (UniProtKB: Q7RVT0). Table S1. Specific activities of cyanide hydratases. Table S2. Purification of NitEg from 200 mL of culture. Table S3. Performance of cyanide hydratase (CynH) and cyanide dihydratase (CynD) in model mixtures and real effluents.
Author Contributions
A.S.: Investigation; Data curation; Writing-review and editing. L.R.: Investigation; Data curation; Writing-review and editing. P.B.: Writing-review and editing. M.G.: Investigation; Data curation. P.N.: Investigation; Data curation. B.K.: Investigation; Data curation; Writing-review and editing. M.P.: Writing-review and editing; Funding acquisition; Project administration. L.M.: Conceptualization; Experimental idea; Paper writing—original draft; Funding acquisition; Project administration. All authors have read and agreed to the published version of the manuscript.
Funding
Research funding by Czech Science Foundation, grant number 18-00184S, Czech Technical University in Prague—Faculty of Biomedical Engineering, grant number SGS21/181/OHK4/3T/17, and Czech Academy of Sciences, grant number RVO61388971, is gratefully acknowledged.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Kuyucak, N.; Akcil, A. Cyanide and removal options from effluents in gold mining and metallurgical processes. Miner. Eng. 2013, 50–51, 13–29. [Google Scholar] [CrossRef]
- Carmona-Orozco, M.L.; Panay, A.J. Immobilization of E. coli expressing Bacillus pumilus CynD in three organic polymer matrices. Appl. Microbiol. Biotechnol. 2019, 103, 5401–5410. [Google Scholar] [CrossRef]
- Basile, L.J.; Willson, R.C.; Sewell, B.T.; Benedik, M.J. Genome mining of cyanide-degrading nitrilases from filamentous fungi. Appl. Microbiol. Biotechnol. 2008, 80, 427–435. [Google Scholar] [CrossRef]
- Martínková, L.; Chmátal, M. The integration of cyanide hydratase and tyrosinase catalysts enables effective degradation of cyanide and phenol in coking wastewaters. Water Res. 2016, 102, 90–95. [Google Scholar] [CrossRef]
- Jarrah, N.; Mu’azu, N.D. Simultaneous electro-oxidation of phenol, CN−, S2− and NH4+ in synthetic wastewater using boron doped diamond anode. J. Environ. Chem. Eng. 2016, 4, 2656–2664. [Google Scholar] [CrossRef]
- Ibáñez, M.I.; Cabello, P.; Luque-Almagro, V.M.; Sáez, L.P.; Olaya, A.; Sánchez de Medina, V.; Luque de Castro, M.D.; Moreno-Vivián, C.; Roldán, M.D. Quantitative proteomic analysis of Pseudomonas pseudoalcaligenes CECT5344 in response to industrial cyanide-containing wastewaters using Liquid Chromatography-Mass Spectrometry/Mass Spectrometry (LC-MS/MS). PLoS ONE 2017, 12, e0172908. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.L.; Liu, G.S.; Chen, Y.H.; Miao, D.T.; Wei, Q.P.; Li, H.C.; Ma, L.; Zhou, K.C.; Liu, L.B.; Yu, Z.M. Persulfate enhanced electrochemical oxidation of highly toxic cyanide-containing organic wastewater using boron-doped diamond anode. Chemosphere 2020, 252, 126499. [Google Scholar] [CrossRef] [PubMed]
- Papadimitriou, C.A.; Samaras, P.; Sakellaropoulos, G.P. Comparative study of phenol and cyanide containing wastewater in CSTR and SBR activated sludge reactors. Bioresour. Technol. 2009, 100, 31–37. [Google Scholar] [CrossRef]
- Yu, X.B.; Xu, R.H.; Wei, C.H.; Wu, H.Z. Removal of cyanide compounds from coking wastewater by ferrous sulfate: Improvement of biodegradability. J. Hazard. Mater. 2016, 302, 468–474. [Google Scholar] [CrossRef]
- Makhado, E.; Pandey, S.; Nomngongo, P.N.; Ramontja, J. Preparation and characterization of xanthan gum-cl-poly(acrylic acid)/o-MWCNTs hydrogel nanocomposite as highly effective reusable adsorbent for removal of methylene blue from aqueous solutions. J. Colloid Interface Sci. 2018, 513, 700–714. [Google Scholar] [CrossRef]
- Pandey, S.; Do, J.Y.; Kim, J.; Kang, M. Fast and highly efficient catalytic degradation of dyes using κ-carrageenan stabilized silver nanoparticles nanocatalyst. Carbohydr. Polym. 2020, 230, 115597. [Google Scholar] [CrossRef]
- Electroplating and metal finishing point source categories; Effluent limitations guidelines, pretreatment, standards and new source performance standards. Fed. Regist. 1983, 48, 32487. Available online: https://www.epa.gov/sites/default/files/2015-10/documents/electroplating-and-metal-finishing_proposed-rule_08-31-1982_47-fr-38462.pdf (accessed on 12 November 2021).
- Code of Federal Regulations. 40, Subpart A-Cokemaking Subcategory. Available online: https://www.ecfr.gov/current/title-40/chapter-I/subchapter-N/part-420/subpart-A (accessed on 12 November 2021).
- Cosmos, A.; Erdenekhuyag, B.O.; Yao, G.; Li, H.J.; Zhao, J.G.; Wang, L.J.; Lyu, X.J. Principles and methods of bio detoxification of cyanide contaminants. J Mater. Cycles Waste Manag. 2020, 22, 939–954. [Google Scholar] [CrossRef]
- Rahdar, S.; Rahdar, A.; Sattari, M.; Hafshejani, L.D.; Tolkou, A.K.; Kyzas, G.Z. Barium/cobalt@polyethylene glycol nanocomposites for dye removal from aqueous solutions. Polymers 2021, 13, 1161. [Google Scholar] [CrossRef]
- Rahdar, S.; Pal, K.; Mohammadi, L.; Rahdar, A.; Goharniya, Y.; Samani, S.; Kyzas, G.Z. Response surface methodology for the removal of nitrate ions by adsorption onto copper oxide nanoparticles. J. Mol. Struct. 2021, 1231, 129686. [Google Scholar] [CrossRef]
- Baeissa, E.S.; Mohamed, R.M. Enhancement of photocatalytic properties of Ga2O3-SiO2 nanoparticles by Pt deposition. Chin. J. Catal. 2013, 34, 1167–1172. [Google Scholar] [CrossRef]
- Bagabas, A.; Alshammari, A.; Aboud, M.F.A.; Kosslick, H. Room-temperature synthesis of zinc oxide nanoparticles in different media and their application in cyanide photodegradation. Nanoscale Res. Lett. 2013, 8, 516. [Google Scholar] [CrossRef]
- Fan, L.; Yao, H.; Deng, S.; Jia, F.; Cai, W.; Hu, Z.; Guo, J.; Li, H. Performance and microbial community dynamics relationship within a step-feed anoxic/oxic/anoxic/oxic process (SF-A/O/A/O) for coking wastewater treatment. Sci. Total Environ. 2021, 792, 148263. [Google Scholar] [CrossRef]
- Martínková, L.; Veselá, A.B.; Rinágelová, A.; Chmátal, M. Cyanide hydratases and cyanide dihydratases: Emerging tools in the biodegradation and biodetection of cyanide. Appl. Microbiol. Biotechnol. 2015, 99, 8875–8882. [Google Scholar] [CrossRef]
- Fry, W.E.; Millar, R.L. Cyanide degradation by an enzyme from Stemphylium loti. Arch. Biochem. Biophys. 1972, 151, 468–474. [Google Scholar] [CrossRef]
- Wang, P.; VanEtten, H.D. Cloning and properties of a cyanide hydratase gene from the phytopathogenic fungus Gloeocercospora sorghi. Biochem. Biophys. Res. Commun. 1992, 187, 1048–1054. [Google Scholar] [CrossRef]
- Cluness, M.J.; Turner, P.D.; Clements, E.; Brown, D.T.; O’Reilly, C. Purification and properties of cyanide hydratase from Fusarium lateritium and analysis of the corresponding chy1 gene. J. Gen. Microbiol. 1993, 139, 1807–1815. [Google Scholar] [CrossRef][Green Version]
- Barclay, M.; Tett, V.A.; Knowles, C.J. Metabolism and enzymology of cyanide/metallocyanide biodegradation by Fusarium solani under neutral and acidic conditions. Enzym. Microb. Technol. 1998, 23, 321–330. [Google Scholar] [CrossRef]
- Yanase, H.; Sakamoto, A.; Okamoto, K.; Kita, K.; Sato, Y. Degradation of the metal-cyano complex tetracyanonickelate (II) by Fusarium oxysporum N-10. Appl. Microbiol. Biotechnol. 2000, 53, 328–334. [Google Scholar] [CrossRef]
- Kushwaha, M.; Kumar, V.; Mahajan, R.; Bhalla, T.C.; Chatterjee, S.; Akhter, Y. Molecular insights into the activity and mechanism of cyanide hydratase enzyme associated with cyanide biodegradation by Serratia marcescens. Arch. Microbiol. 2018, 200, 971–977. [Google Scholar] [CrossRef]
- Basic Local Alignment Search Tool. Available online: https://blast.ncbi.nlm.nih.gov (accessed on 12 November 2021).
- Park, J.M.; Sewell, B.T.; Benedik, M.J. Cyanide bioremediation: The potential of engineered nitrilases. Appl. Microbiol. Biotechnol. 2017, 101, 3029–3042. [Google Scholar] [CrossRef]
- Nolan, L.M.; Harnedy, P.A.; Turner, P.; Hearne, A.B.; O’Reilly, C. The cyanide hydratase enzyme of Fusarium lateritium also has nitrilase activity. FEMS Microbiol. Lett. 2003, 221, 161–165. [Google Scholar] [CrossRef]
- Kaplan, O.; Veselá, A.B.; Petříčková, A.; Pasquarelli, F.; Pičmanová, M.; Rinágelová, A.; Bhalla, T.C.; Pátek, M.; Martínková, L. A comparative study of nitrilases identified by genome mining. Mol. Biotechnol. 2013, 54, 996–1003. [Google Scholar] [CrossRef] [PubMed]
- Veselá, A.B.; Rucká, L.; Kaplan, O.; Pelantová, H.; Nešvera, J.; Pátek, M.; Martínková, L. Bringing nitrilase sequences from databases to life: The search for novel substrate specificities with a focus on dinitriles. Appl. Microbiol. Biotechnol. 2016, 100, 2193–2202. [Google Scholar] [CrossRef]
- Rucká, L.; Chmátal, M.; Kulik, N.; Petrásková, L.; Pelantová, H.; Novotný, P.; Příhodová, R.; Pátek, M.; Martínková, L. Genetic and functional diversity of nitrilases in Agaricomycotina. Int. J. Mol. Sci. 2019, 20, 5990. [Google Scholar] [CrossRef]
- Rinágelová, A.; Kaplan, O.; Veselá, A.B.; Chmátal, M.; Křenková, A.; Plíhal, O.; Pasquarelli, F.; Cantarella, M.; Martínková, L. Cyanide hydratase from Aspergillus niger K10: Overproduction in Escherichia coli, purification, characterization and use in continuous cyanide degradation. Proc. Biochem. 2014, 49, 445–450. [Google Scholar] [CrossRef]
- Malmir, N.; Fard, N.A.; Mgwatyu, Y.; Mekuto, L. Cyanide hydratase modification using computational design and docking analysis for improved binding affinity in cyanide detoxification. Molecules 2021, 26, 1799. [Google Scholar] [CrossRef]
- Rucká, L.; Kulik, N.; Novotný, P.; Sedova, A.; Petrásková, L.; Příhodová, R.; Křístková, B.; Halada, P.; Pátek, M.; Martínková, L. Plant nitrilase homologues in fungi: Phylogenetic and functional analysis with focus on nitrilases in Trametes versicolor and Agaricus bisporus. Molecules 2020, 25, 3861. [Google Scholar] [CrossRef]
- Fisher, F.B.; Brown, J.S. Colorimetric determination of cyanide in stack gas and waste water. Anal. Chem. 1952, 24, 1440–1444. [Google Scholar] [CrossRef]
- Pérez-Cid, B.; Calvar, S.; Belén Moldes, A.; Cruz, J.M. Effective removal of cyanide and heavy metals from an industrial electroplating stream using calcium alginate hydrogels. Molecules 2020, 25, 5183. [Google Scholar] [CrossRef]
- Xue, Y.P.; Wang, Y.P.; Xu, Z.; Liu, Z.Q.; Shu, X.R.; Jia, D.X.; Zheng, Y.G.; Shen, Y.C. Chemoenzymatic synthesis of gabapentin by combining nitrilase-mediated hydrolysis with hydrogenation over Raney-nickel. Catal. Commun. 2015, 66, 121–125. [Google Scholar] [CrossRef]
- Wang, L.; Watermeyer, J.M.; Mulelu, A.E.; Sewell, B.T.; Benedik, M.J. Engineering pH-tolerant mutants of a cyanide dihydratase. Appl. Microbiol. Biotechnol. 2012, 94, 131–140. [Google Scholar] [CrossRef]
- Crum, M.A.; Park, J.M.; Mulelu, A.E.; Sewell, B.T.; Benedik, M.J. Probing C-terminal interactions of the Pseudomonas stutzeri cyanide-degrading CynD protein. Appl. Microbiol. Biotechnol. 2015, 99, 3093–3102. [Google Scholar] [CrossRef]
- Jandhyala, D.M.; Willson, R.C.; Sewell, B.T.; Benedik, M.J. Comparison of cyanide-degrading nitrilases. Appl. Microbiol. Biotechnol. 2005, 68, 327–335. [Google Scholar] [CrossRef] [PubMed]
- BRENDA. The Comprehensive Enzyme Information System. Available online: https://brenda-enzymes.org (accessed on 2 November 2021).
- Dennett, G.V.; Blamey, J.M. A new thermophilic nitrilase from an antarctic hyperthermophilic microorganism. Front. Bioeng. Biotechnol. 2016, 4, 5. [Google Scholar] [CrossRef]
- Mondal, A.; Sarkar, S.; Nair, U.G. Comparative characterization of cyanide-containing steel industrial wastewater. Water Sci. Technol. 2021, 83, 322–330. [Google Scholar] [CrossRef] [PubMed]
- Ketterer, L.; Keusgen, M. Amperometric sensor for cyanide utilizing cyanidase and formate dehydrogenase. Anal. Chim. Acta 2010, 673, 54–59. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).