Deep Sequencing of Early T Stage Colorectal Cancers Reveals Disruption of Homologous Recombination Repair in Microsatellite Stable Tumours with High Mutational Burdens
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Specimens
2.2. Next Generation Sequencing
2.3. Ion AmpliSeq™ Comprehensive Cancer Panel
2.4. Retrieval of TCGA datasets
3. Results
3.1. Robust Deep Targeted Sequencing of Early T Stage Colorectal Cancer
3.2. No Stage-Related Mutational Differences between Stage IIIA vs. Stage I in TMB-Low Group
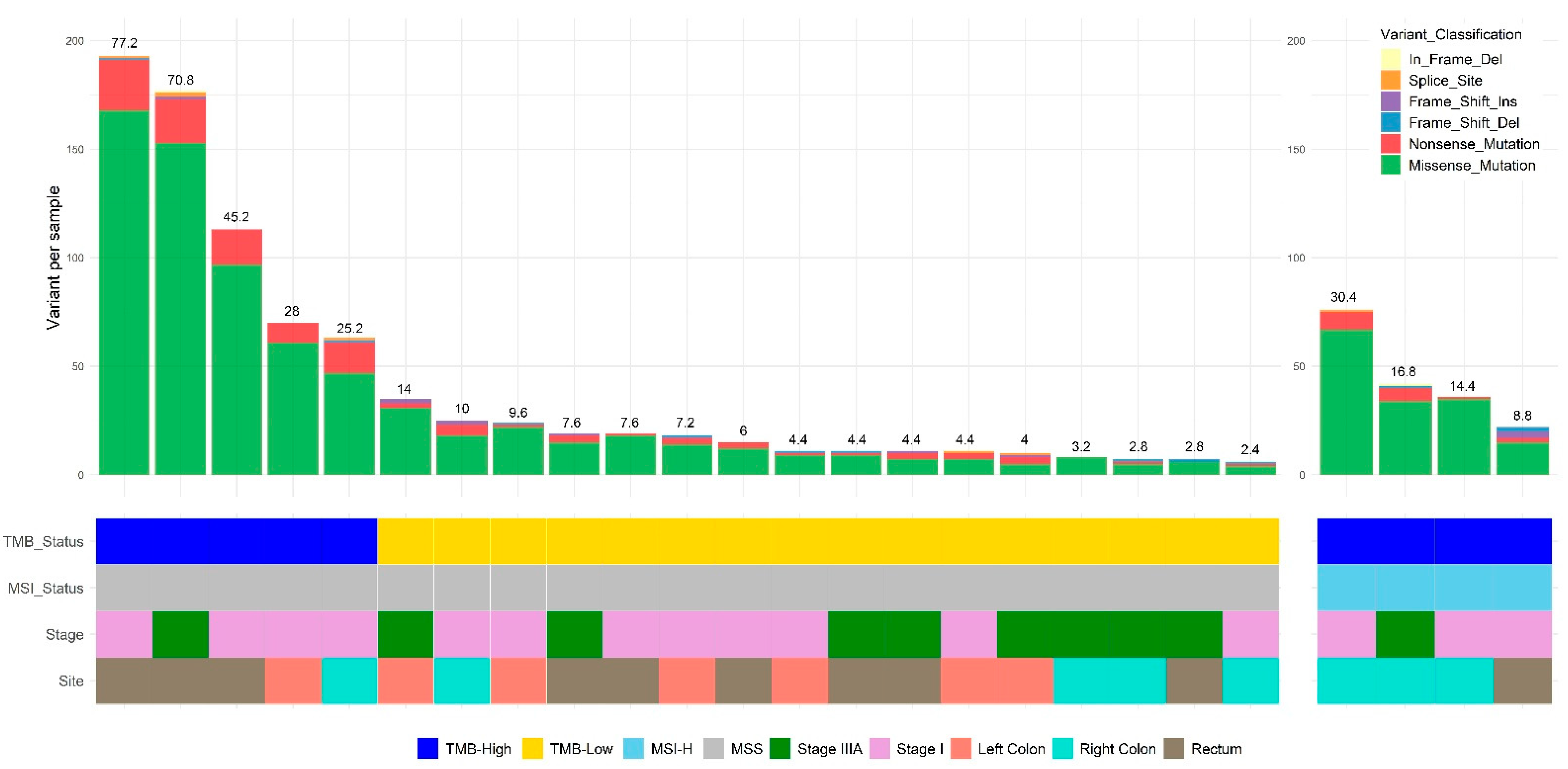
3.3. Mutational Spectrum of TMB-H and TMB-L CRC
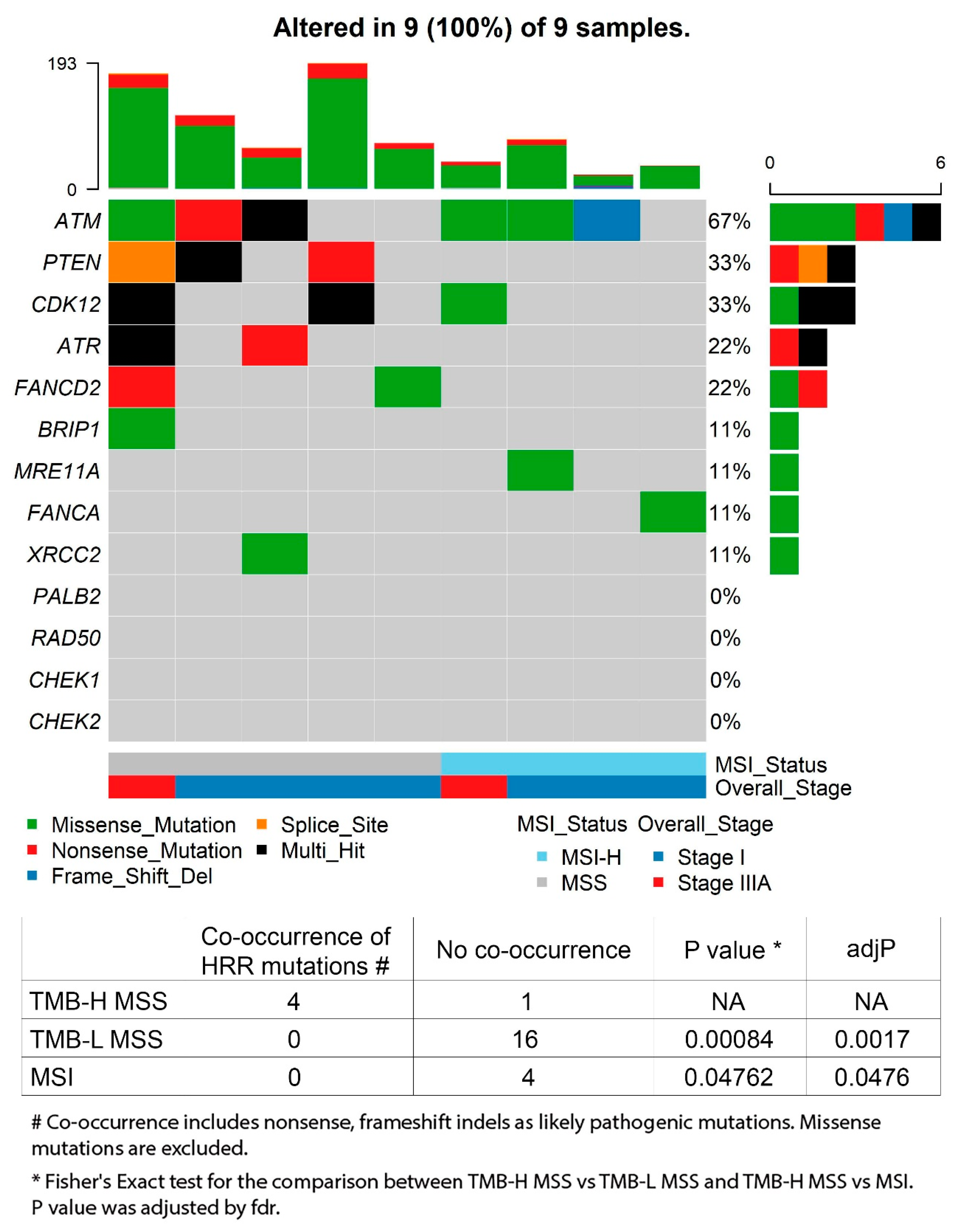
3.4. Deficiency of Homologous Recombination Repair in TMB-H CRC
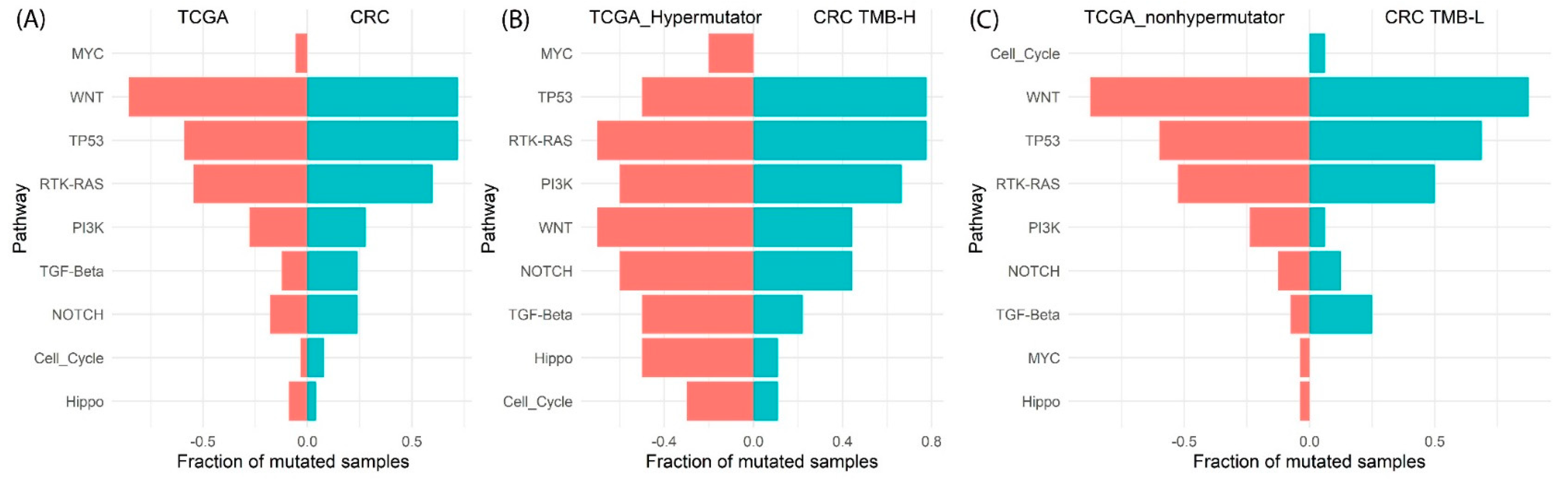
3.5. Oncogenic Pathway Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Xi, Y.; Xu, P. Global colorectal cancer burden in 2020 and projections to 2040. Transl. Oncol. 2021, 14, 101174. [Google Scholar] [CrossRef] [PubMed]
- Tse, B.C.Y.; Welham, Z.; Engel, A.F.; Molloy, M.P. Genomic, Microbial and Immunological Microenvironment of Colorectal Polyps. Cancers 2021, 13, 3382. [Google Scholar] [CrossRef] [PubMed]
- Lengauer, C.; Kinzler, K.W.; Vogelstein, B. Genetic instability in colorectal cancers. Nature 1997, 386, 623–627. [Google Scholar] [CrossRef]
- Leggett, B.; Whitehall, V. Role of the serrated pathway in colorectal cancer pathogenesis. Gastroenterology 2010, 138, 2088–2100. [Google Scholar] [CrossRef]
- Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature 2012, 487, 330–337. [Google Scholar] [CrossRef] [Green Version]
- Amin, M.B.; Edge, S.B.; Greene, F.L.; Byrd, D.R.; Brookland, R.K.; Washington, M.K.; Gershenwald, J.E.; Compton, C.C.; Hess, K.R.; Sullivan, D.C.; et al. (Eds.) AJCC Cancer Staging Manual, 8th ed.; Springer: New York, NY, USA, 2017. [Google Scholar]
- Mejri, N.; Dridi, M.; Labidi, S.; El Benna, H.; Daoud, N.; Boussen, H. Annual hazard rate of relapse of stage II and III colorectal cancer after primary therapy. Clin. Transl. Oncol. 2017, 19, 1524–1530. [Google Scholar] [CrossRef]
- Osterman, E.; Glimelius, B. Recurrence Risk after Up-to-Date Colon Cancer Staging, Surgery, and Pathology: Analysis of the Entire Swedish Population. Dis. Colon Rectum 2018, 61, 1016–1025. [Google Scholar] [CrossRef]
- Kandimalla, R.; Ozawa, T.; Gao, F.; Wang, X.; Goel, A.; Group, T.C.C.S. Gene Expression Signature in Surgical Tissues and Endoscopic Biopsies Identifies High-Risk T1 Colorectal Cancers. Gastroenterology 2019, 156, 2338–2341.e3. [Google Scholar] [CrossRef]
- Ikematsu, H.; Yoda, Y.; Matsuda, T.; Yamaguchi, Y.; Hotta, K.; Kobayashi, N.; Fujii, T.; Oono, Y.; Sakamoto, T.; Nakajima, T.; et al. Long-term outcomes after resection for submucosal invasive colorectal cancers. Gastroenterology 2013, 144, 551–559. [Google Scholar] [CrossRef]
- Steffen, P.; Li, J.; Chandra, J.; Ahadi, M.S.; Gill, A.J.; Engel, A.F.; Molloy, M.P. Molecular Features of Lymph Node Metastasis in T1/2 Colorectal Cancer from Formalin-Fixed Paraffin-Embedded Archival Specimens. J. Proteome Res. 2021, 20, 1304–1312. [Google Scholar] [CrossRef]
- Mayakonda, A.; Lin, D.C.; Assenov, Y.; Plass, C.; Koeffler, H.P. Maftools: Efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 2018, 28, 1747–1756. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Campbell, B.B.; Light, N.; Fabrizio, D.; Zatzman, M.; Fuligni, F.; de Borja, R.; Davidson, S.; Edwards, M.; Elvin, J.A.; Hodel, K.P.; et al. Comprehensive Analysis of Hypermutation in Human Cancer. Cell 2017, 171, 1042–1056.e10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, M.; Eng, G.; Barbari, S.R.; Deshpande, V.; Shcherbakova, P.V.; Gala, M.K. Homologous Recombination Repair Truncations Predict Hypermutation in Microsatellite Stable Colorectal and Endometrial Tumors. Clin. Transl. Gastroenterol. 2020, 11, e00149. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Domingo, E.; Freeman-Mills, L.; Rayner, E.; Glaire, M.; Briggs, S.; Vermeulen, L.; Fessler, E.; Medema, J.P.; Boot, A.; Morreau, H.; et al. Somatic POLE proofreading domain mutation, immune response, and prognosis in colorectal cancer: A retrospective, pooled biomarker study. Lancet Gastroenterol. Hepatol. 2016, 1, 207–216. [Google Scholar] [CrossRef] [Green Version]
- Lawrence, M.S.; Stojanov, P.; Polak, P.; Kryukov, G.V.; Cibulskis, K.; Sivachenko, A.; Carter, S.L.; Stewart, C.; Mermel, C.H.; Roberts, S.A.; et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 2013, 499, 214–218. [Google Scholar] [CrossRef]
- Christie, M.; Jorissen, R.N.; Mouradov, D.; Sakthianandeswaren, A.; Li, S.; Day, F.; Tsui, C.; Lipton, L.; Desai, J.; Jones, I.T.; et al. Different APC genotypes in proximal and distal sporadic colorectal cancers suggest distinct WNT/beta-catenin signalling thresholds for tumourigenesis. Oncogene 2013, 32, 4675–4682. [Google Scholar] [CrossRef]
- Krajewska, M.; Fehrmann, R.S.; de Vries, E.G.; van Vugt, M.A. Regulators of homologous recombination repair as novel targets for cancer treatment. Front. Genet. 2015, 6, 96. [Google Scholar] [CrossRef] [Green Version]
- Sanchez-Vega, F.; Mina, M.; Armenia, J.; Chatila, W.K.; Luna, A.; La, K.C.; Dimitriadoy, S.; Liu, D.L.; Kantheti, H.S.; Saghafinia, S.; et al. Oncogenic Signaling Pathways in The Cancer Genome Atlas. Cell 2018, 173, 321–337.e10. [Google Scholar] [CrossRef] [Green Version]
- Ganesh, K.; Stadler, Z.K.; Cercek, A.; Mendelsohn, R.B.; Shia, J.; Segal, N.H.; Diaz, L.A., Jr. Immunotherapy in colorectal cancer: Rationale, challenges and potential. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 361–375. [Google Scholar] [CrossRef]
- Hu, W.; Yang, Y.; Ge, W.; Zheng, S. Deciphering molecular properties of hypermutated gastrointestinal cancer. J. Cell Mol. Med. 2019, 23, 370–379. [Google Scholar] [CrossRef] [PubMed]
- Lord, C.J.; Ashworth, A. PARP inhibitors: Synthetic lethality in the clinic. Science 2017, 355, 1152–1158. [Google Scholar] [CrossRef] [PubMed]
- Lord, C.J.; Ashworth, A. BRCAness revisited. Nat. Rev. Cancer 2016, 16, 110–120. [Google Scholar] [CrossRef] [PubMed]
- Ray-Coquard, I.; Pautier, P.; Pignata, S.; Perol, D.; Gonzalez-Martin, A.; Berger, R.; Fujiwara, K.; Vergote, I.; Colombo, N.; Maenpaa, J.; et al. Olaparib plus Bevacizumab as First-Line Maintenance in Ovarian Cancer. N. Engl. J. Med. 2019, 381, 2416–2428. [Google Scholar] [CrossRef] [PubMed]


| Stage I (n = 16) | Stage IIIA (n = 10) | p-Value * | |
|---|---|---|---|
| Year | |||
| Range | 2007–2017 | 2008–2018 | |
| Sex | p = 0.692 | ||
| Female | 5 (31.25%) | 4 (40.00%) | |
| Male | 11 (68.75%) | 6 (60.00%) | |
| Age | p= 0.038 | ||
| Range | 49–87 | 31–85 | |
| Mean(sd) | 71.31 (11.07) | 57.40 (21.36) | |
| Tumour size | p = 0.065 | ||
| Range | 15–30 | 13–55 | |
| Mean(sd) | 22.56 (4.91) | 30.50 (15.36) | |
| Site | p = 0.806 | ||
| Right | 6 (37.50%) | 3 (30.00%) | |
| Left | 5 (31.25%) | 2 (20.00%) | |
| Rectal | 5 (31.25%) | 5 (50.00%) | |
| Total nodes | p = 0.091 | ||
| Range | 13–22 | 12–49 | |
| Mean(sd) | 17.31 (3.11) | 23.50 (13.66) | |
| pT.7th.Ed | p< 0.001 | ||
| T1 | 16 (100.00%) | 6 (60.00%) | |
| T2 | 0 (0.00%) | 4 (40.00%) | |
| pN.7th.Ed | |||
| N0 | 16 (100.00%) | 0 (0.00%) | By definition |
| N1a | 0 (0.00%) | 5 (50.00%) | |
| N1b | 0 (0.00%) | 3 (30.00%) | |
| N1c | 0 (0.00%) | 2 (20.00%) | |
| Thin walled vessel invasion | p = 0.055 | ||
| 0 (Absent) | 15 (93.75%) | 6 (60.00%) | |
| 1 (Present) | 1 (6.25%) | 4 (40.00%) | |
| Combined.BRAF.MMR.status | p = 1.000 | ||
| BRAF V600E +/MSS | 1 (6.25%) | 1 (10.00%) | |
| BRAF V600E −/MSH | 0 (0.00%) | 0 (0.00%) | |
| BRAF V600E +/MSH | 3 (18.75%) | 1 (10.00%) | |
| BRAF V600E −/MSS | 12 (75.00%) | 8 (80.00%) |
| Gene Symbol | Stage IIIA (freq%) | Stage I (freq%) | Pval | Odds Ratio | Confidence Interval. Upper | Confidence Interval. Lower | AdjPval |
|---|---|---|---|---|---|---|---|
| KRAS | 5 (62.5%) | 1 (12.5%) | 0.12 | 9.76 | 625.22 | 0.68 | 0.24 |
| TP53 | 7 (87.5%) | 4 (50%) | 0.28 | 6.16 | 391.63 | 0.42 | 0.38 |
| APC | 7 (87.5%) | 7 (87.5%) | 1 | 1 | 89.51 | 0.01 | 1 |
| Gene | TMB-H (n = 9) | TMB-L (n = 16) | Pval | AdjPval | TCGA COAD/READ Hypermutated (n = 10) | TCGA COAD/READ Nonhypermutated (n = 81) | Pval | AdjPval |
|---|---|---|---|---|---|---|---|---|
| ATM | 67% | 0% | 0.0005 | 0.0090 | 60% | 9% | 0.0004 | 0.0006 |
| ALK | 56% | 0% | 0.0024 | 0.0090 | 20% | 2% | 0.0583 | 0.1486 |
| NSD1 | 56% | 0% | 0.0024 | 0.0090 | 20% | 2% | 0.0583 | 0.1486 |
| UBR5 | 56% | 0% | 0.0024 | 0.0090 | 30% | 0% | 0.0010 | 0.0114 |
| TRRAP | 67% | 13% | 0.0099 | 0.0188 | 50% | 2% | 0.0001 | 0.0002 |
| BCL9 | 56% | 6% | 0.0119 | 0.0188 | 20% | 1% | 0.0310 | 0.0850 |
| CARD11 | 56% | 6% | 0.0119 | 0.0188 | 0% | 9% | 1.0000 | 1.0000 |
| KDM5C | 56% | 6% | 0.0119 | 0.0188 | 10% | 1% | 0.2088 | 0.2491 |
| MN1 | 56% | 6% | 0.0119 | 0.0188 | 10% | 0% | 0.1099 | 0.1513 |
| PIK3CA | 56% | 6% | 0.0119 | 0.0188 | 40% | 20% | 0.2178 | 0.2341 |
| PTPRT | 56% | 6% | 0.0119 | 0.0188 | 30% | 6% | 0.0405 | 0.0475 |
| APC | 44% | 88% | 0.0581 | 0.0736 | 40% | 85% | 0.0035 | 0.0048 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, J.; Steffen, P.; Tse, B.C.Y.; Ahadi, M.S.; Gill, A.J.; Engel, A.F.; Molloy, M.P. Deep Sequencing of Early T Stage Colorectal Cancers Reveals Disruption of Homologous Recombination Repair in Microsatellite Stable Tumours with High Mutational Burdens. Cancers 2022, 14, 2933. https://doi.org/10.3390/cancers14122933
Li J, Steffen P, Tse BCY, Ahadi MS, Gill AJ, Engel AF, Molloy MP. Deep Sequencing of Early T Stage Colorectal Cancers Reveals Disruption of Homologous Recombination Repair in Microsatellite Stable Tumours with High Mutational Burdens. Cancers. 2022; 14(12):2933. https://doi.org/10.3390/cancers14122933
Chicago/Turabian StyleLi, Jun, Pascal Steffen, Benita C. Y. Tse, Mahsa S. Ahadi, Anthony J. Gill, Alexander F. Engel, and Mark P. Molloy. 2022. "Deep Sequencing of Early T Stage Colorectal Cancers Reveals Disruption of Homologous Recombination Repair in Microsatellite Stable Tumours with High Mutational Burdens" Cancers 14, no. 12: 2933. https://doi.org/10.3390/cancers14122933
APA StyleLi, J., Steffen, P., Tse, B. C. Y., Ahadi, M. S., Gill, A. J., Engel, A. F., & Molloy, M. P. (2022). Deep Sequencing of Early T Stage Colorectal Cancers Reveals Disruption of Homologous Recombination Repair in Microsatellite Stable Tumours with High Mutational Burdens. Cancers, 14(12), 2933. https://doi.org/10.3390/cancers14122933







