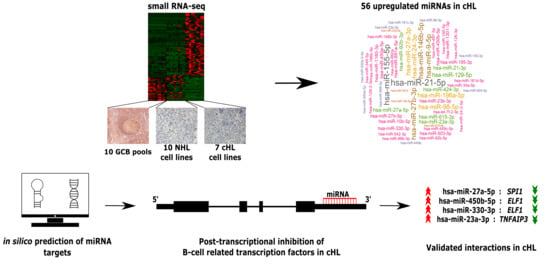
Deregulated miRNAs Contribute to Silencing of B-Cell Specific Transcription Factors and Activation of NF-κB in Classical Hodgkin Lymphoma
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Lymphoma Cell Lines, GCB Cells and Primary cHL Biopsies
2.2. NGS Based miRNA Profiling
2.3. Real-Time qPCR Based miRNA Expression Analysis
2.4. Identification of Putative Target Genes of the cHL Deregulated miRNAs
2.5. Validation of miRNA Target Genes
2.5.1. Vector Preparation
2.5.2. Luciferase Reporter Assay
- In total, 500 ng of vector containing the 3′UTR WT sequence + 50 µM of the analyzed miRNA mimic (mirVana® miRNA mimic, Invitrogen, Waltham, MA, USA);
- In total, 500 ng of vector containing the 3′UTR WT sequence + 50 µM of the mimic negative control (mirVana™ miRNA Mimic, Negative Control #1, Invitrogen, Waltham, MA, USA);
- In total, 500 ng of vector containing the 3′UTR MUT sequence + 50 µM of the analyzed miRNA mimic (mirVana® miRNA mimic, Invitrogen, Waltham, MA, USA);
- In total, 500 ng of vector containing the 3′UTR MUT sequence + 50 µM of the mimic negative control (mirVana™ miRNA Mimic, Negative Control #1, Invitrogen, Waltham, MA, USA).
2.6. MiRNA Overexpression
2.7. Western Blots
3. Results
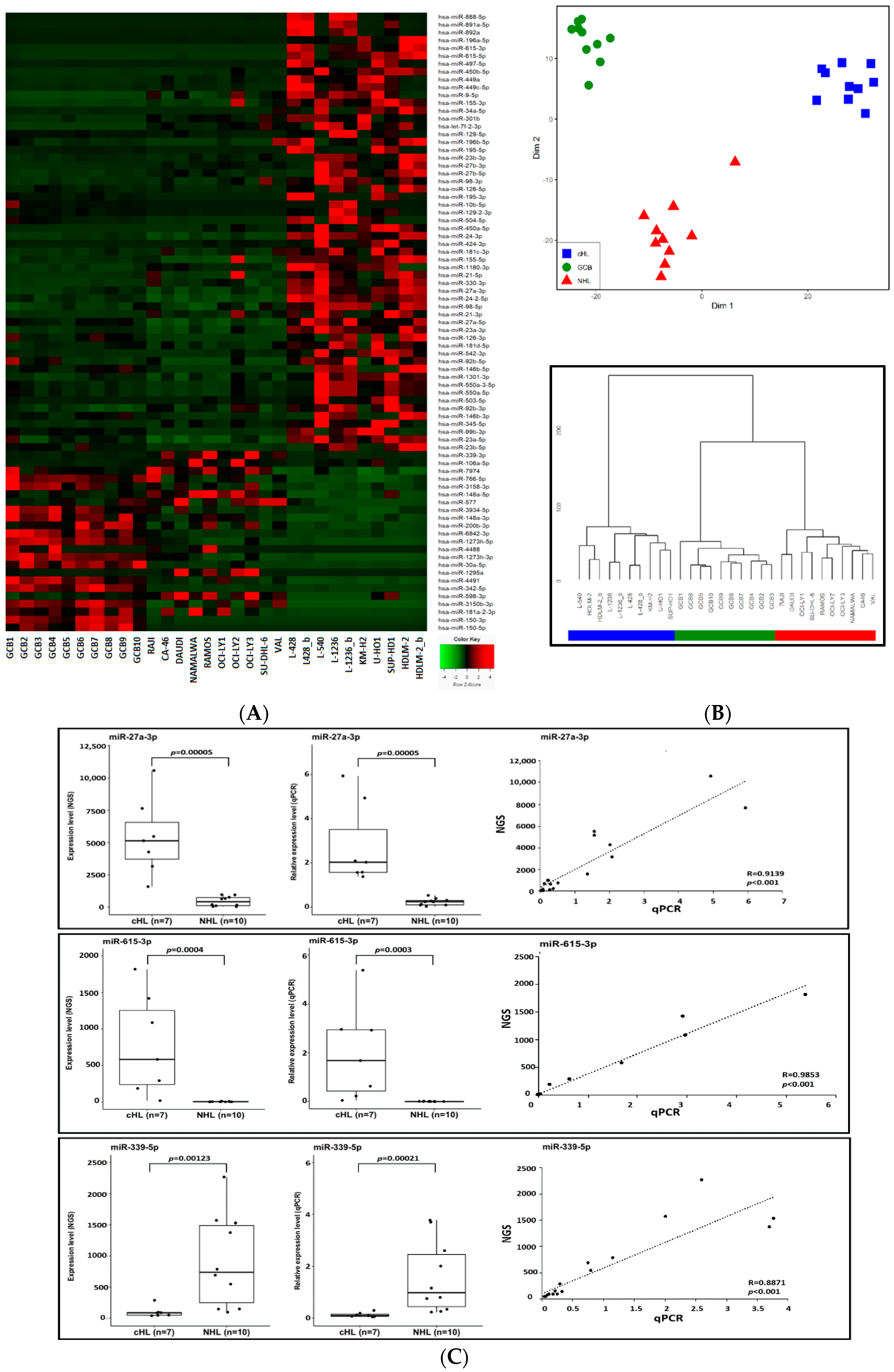
3.1. The cHL miRNome and miRNAs Deregulated in cHL
3.2. Biological Processes Deregulated by miRNAs in cHL
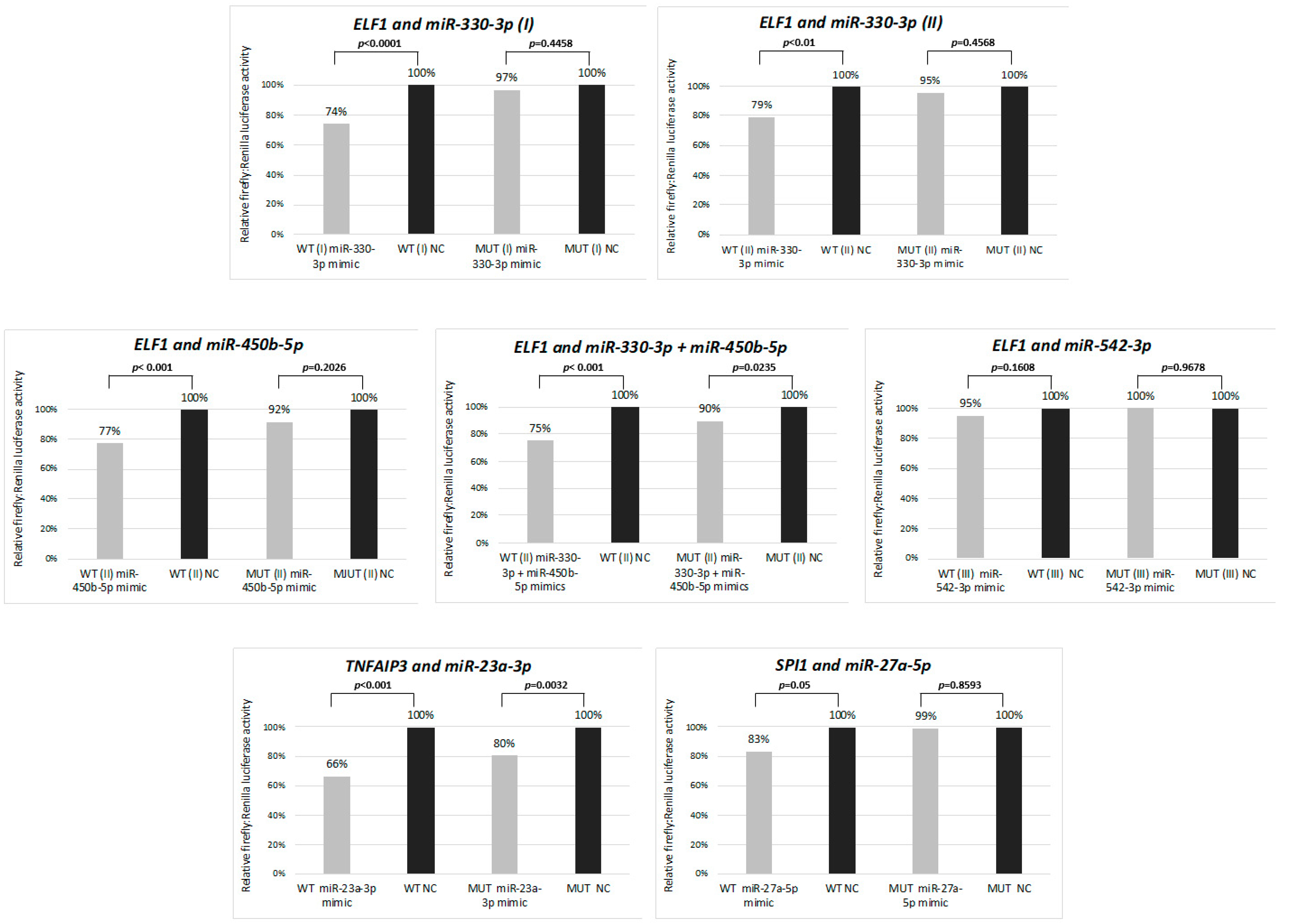
3.3. Functional Validation of miRNA-mRNA Interactions for Selected Candidates
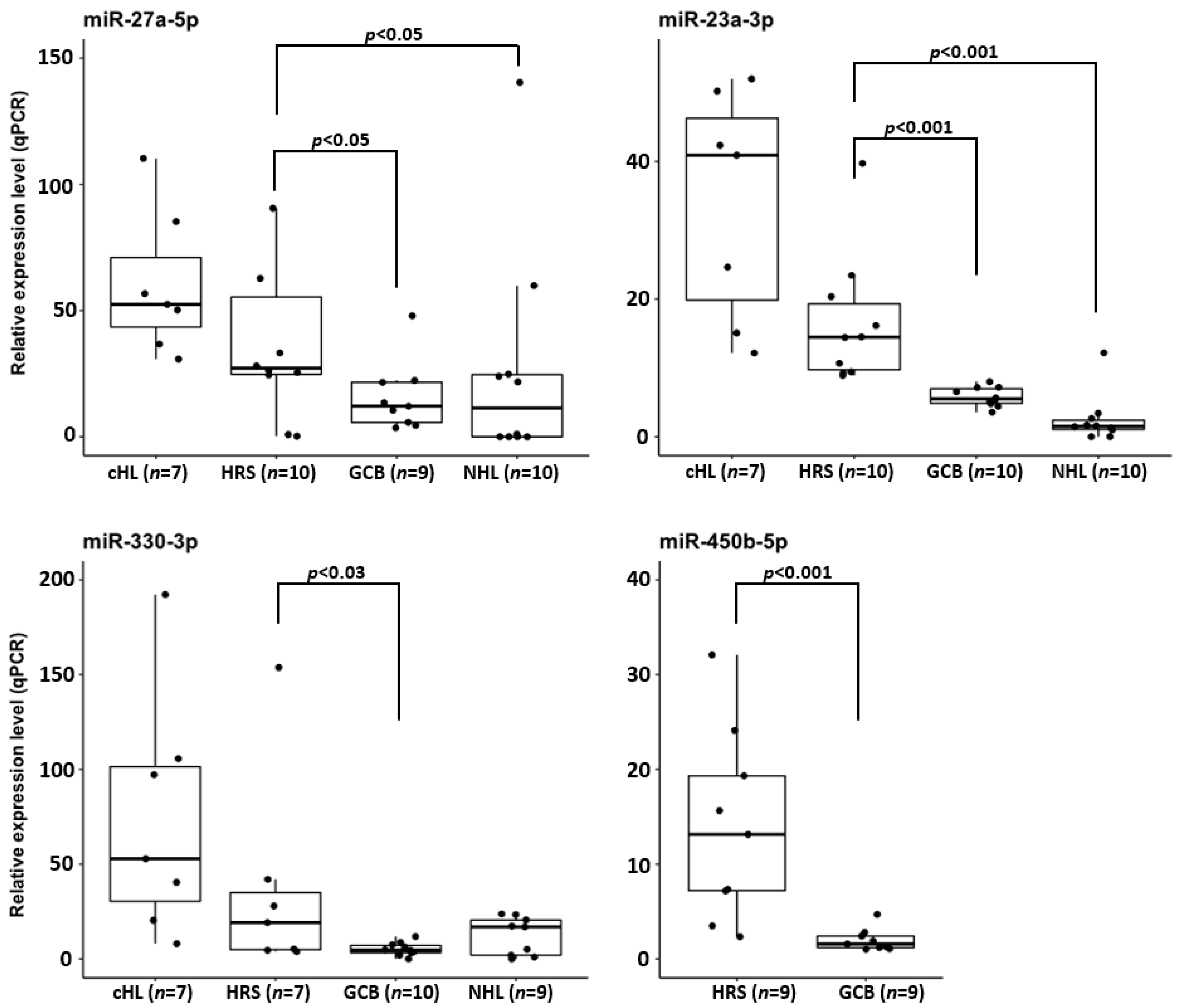
3.4. Validation of miRNA Expression in Microdissected Primary HRS Cells
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ammerpohl, O.; Haake, A.; Pellissery, S.; Giefing, M.; Richter, J.; Bálint, B.; Kulis, M.; Le, J.; Bibikova, M.; Drexler, H.G.; et al. Array-based DNA methylation analysis in classical Hodgkin lymphoma reveals new insights into the mechanisms underlying silencing of B cell-specific genes. Leukemia 2012, 26, 185–188. [Google Scholar] [CrossRef] [Green Version]
- D’Alò, F.; Leone, G.; Hohaus, S.; Teofili, L.; Bozzoli, V.; Tisi, M.C.; Rufini, V.; Calcagni, M.L.; Voso, M.T. Response to 5-azacytidine in a patient with relapsed Hodgkin lymphoma and a thera-py-related myelodysplastic syndrome. Br. J. Haematol. 2011, 154, 141–143. [Google Scholar] [CrossRef]
- Falchi, L.; Sawas, A.; Deng, C.; Amengual, J.E.; Colbourn, D.S.; Lichtenstein, E.A.; Khan, K.A.; Schwartz, L.H.; O’Connor, O.A. High rate of complete responses to immune checkpoint inhibitors in patients with relapsed or refractory Hodgkin lymphoma previously exposed to epigenetic therapy. J. Hematol. Oncol. 2016, 9, 1–4. [Google Scholar] [CrossRef] [Green Version]
- Yuki, H.; Ueno, S.; Tatetsu, H.; Niiro, H.; Iino, T.; Endo, S.; Kawano, Y.; Komohara, Y.; Takeya, M.; Hata, H.; et al. 1 is a potent tumor suppressor in classical Hodgkin lymphoma cells. Blood 2013, 121, 962–970. [Google Scholar] [CrossRef] [PubMed]
- Paczkowska, J.; Soloch, N.; Bodnar, M.; Kiwerska, K.; Janiszewska, J.; Vogt, J.; Domanowska, E.; Martin-Subero, J.I.; Ammerpohl, O.; Klapper, W.; et al. Expression of ELF1, a lymphoid ETS domain-containing transcription factor, is re-currently lost in classical Hodgkin lymphoma. Br. J. Haematol. 2019, 185, 79–88. [Google Scholar] [CrossRef] [PubMed]
- Coffre, M.; Koralov, S.B. miRNAs in B Cell Development and Lymphomagenesis. Trends Mol. Med. 2017, 23, 721–736. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; So, A.Y.-L.; Sookram, R.; Wong, S.; Wang, J.K.; Ouyang, Y.; He, P.; Su, Y.; Casellas, R.; Baltimore, D. Epigenetic silencing of miR-125b is required for normal B-cell development. Blood 2018, 131, 1920–1930. [Google Scholar] [CrossRef] [Green Version]
- Blume, J.; Ziętara, N.; Witzlau, K.; Liu, Y.; Sanchez, O.O.; Puchałka, J.; Winter, S.J.; Kunze-Schumacher, H.; Saran, N.; Düber, S.; et al. miR-191 modulates B-cell development and targets transcription factors E2A, Foxp1, and Egr1. Eur. J. Immunol. 2019, 49, 121–132. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arbore, G.; Henley, T.; Biggins, L.; Andrews, S.; Vigorito, E.; Turner, M.; Leyland, R. MicroRNA-155 is essential for the optimal proliferation and survival of plasmablast B cells. Life Sci. Alliance 2019, 2, e201800244. [Google Scholar] [CrossRef] [PubMed]
- Brune, V.; Tiacci, E.; Pfeil, I.; Doring, C.; Eckerle, S.; van Noesel, C.J.; Klapper, W.; Falini, B.; von Heydebreck, A.; Metzler, D.; et al. Origin and pathogenesis of nodular lymphocyte-predominant Hodgkin lymphoma as re-vealed by global gene expression analysis. J. Exp. Med. 2008, 205, 2251–2268. [Google Scholar] [CrossRef]
- Chomczynski, P. A reagent for the single-step simultaneous isolation of RNA, DNA and proteins from cell and tissue samples. Biotechniques 1993, 15, 532–534. [Google Scholar]
- Mets, E.; Van Peer, G.; Van der Meulen, J.; Boice, M.; Taghon, T.; Goossens, S.; Mestdagh, P.; Benoit, Y.; De Moerloose, B.; Van Roy, N.; et al. MicroRNA-128-3p is a novel oncomiR targeting PHF6 in T-cell acute lympho-blastic leukemia. Haematologica 2014, 99, 1326–1333. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kluiver, J.; Slezak-Prochazka, I.; van den Berg, A. Studying microRNAs in lymphoma. Methods Mol. Biol. 2013, 971, 265–276. [Google Scholar] [PubMed]
- Bednarek, K.; Kiwerska, K.; Szaumkessel, M.; Bodnar, M.; Kostrzewska-Poczekaj, M.; Marszalek, A.; Janiszewska, J.; Bartochowska, A.; Jackowska, J.; Wierzbicka, M.; et al. Recurrent CDK1 overexpression in laryngeal squamous cell carcinoma. Tumor Biol. 2016, 37, 11115–11126. [Google Scholar] [CrossRef] [Green Version]
- Paczkowska, J.; Janiszewska, J.; Bein, J.; Schneider, M.; Bednarek, K.; Ustaszewski, A.; Hartmann, S.; Hansmann, M.L.; Giefing, M. The Tumor Suppressive mir-148a Is Epigenetically Inactivated in Classical Hodg-kin Lymphoma. Cells 2020, 9, 2292. [Google Scholar] [CrossRef] [PubMed]
- Torlakovic, E.; Tierens, A.; Dang, H.D.; Delabie, J. The Transcription Factor PU.1, Necessary for B-Cell Development Is Expressed in Lymphocyte Predominance, But Not Classical Hodgkin’s Disease. Am. J. Pathol. 2001, 159, 1807–1814. [Google Scholar] [CrossRef]
- Schmitz, R.; Hansmann, M.-L.; Bohle, V.; Martin-Subero, J.I.; Hartmann, S.; Mechtersheimer, G.; Klapper, W.; Vater, I.; Giefing, M.; Gesk, S.; et al. TNFAIP3(A20) is a tumor suppressor gene in Hodgkin lymphoma and primary mediastinal B cell lymphoma. J. Cell Biol. 2009, 185, i4. [Google Scholar] [CrossRef] [Green Version]
- Kluiver, J.; Poppema, S.; de Jong, D.; Blokzijl, T.; Harms, G.; Jacobs, S.; Kroesen, B.J.; van den Berg, A. BIC and miR-155 are highly expressed in Hodgkin, primary mediastinal and diffuse large B cell lymphomas. J. Pathol. 2005, 207, 243–249. [Google Scholar] [CrossRef]
- Navarro, A.; Gaya, A.; Martinez, A.; Urbano-Ispizua, A.; Pons, A.; Balagué, O.; Gel, B.; Abrisqueta, P.; Lopez-Guillermo, A.; Artells, R.; et al. MicroRNA expression profiling in classic Hodgkin lymphoma. Blood 2008, 111, 2825–2832. [Google Scholar] [CrossRef] [Green Version]
- Van Vlierberghe, P.; De Weer, A.N.; Mestdagh, P.; Feys, T.; De Preter, K.; De Paepe, P.; Lambein, K.; Vandesompele, J.; Van Roy, N.; Verhasselt, B.; et al. Comparison of miRNA profiles of microdissected Hodg-kin/Reed-Sternberg cells and Hodgkin cell lines versus CD77+ B-cells reveals a distinct subset of differentially expressed miRNAs. Br. J. Haematol. 2009, 147, 686–690. [Google Scholar] [CrossRef]
- Gibcus, J.H.; Tan, L.P.; Harms, G.; Schakel, R.N.; de Jong, D.; Blokzijl, T.; Möller, P.; Poppema, S.; Kroesen, B.J.; van den Berg, A. Hodgkin lymphoma cell lines are characterized by a specific miRNA expression profile. Neoplasia 2009, 11, 167–176. [Google Scholar] [CrossRef] [Green Version]
- Sánchez-Espiridión, B.; Martín-Moreno, A.M.; Montalbán, C.; Figueroa, V.; Vega, F.; Younes, A.; Medeiros, L.J.; Alvés, F.J.; Canales, M.; Estévez, M.; et al. MicroRNA signatures and treatment response in patients with advanced classical Hodgkin lymphoma. Br. J. Haematol. 2013, 162, 336–347. [Google Scholar] [CrossRef] [Green Version]
- Paczkowska, J.; Giefing, M. MicroRNA signature in classical Hodgkin lymphoma. J. Appl. Genet. 2021, 62, 281–288. [Google Scholar] [CrossRef]
- Yuan, Y.; Niu, F.; Nolte, I.M.; Koerts, J.; De Jong, D.; Rutgers, B.; Osinga, J.; Azkanaz, M.; Terpstra, M.; Bystrykh, L.; et al. MicroRNA High Throughput Loss-of-Function Screening Reveals an Oncogenic Role for miR-21-5p in Hodgkin Lymphoma. Cell. Physiol. Biochem. 2018, 49, 144–159. [Google Scholar] [CrossRef]
- Reed, S.M.; Hagen, J.; Muniz, V.P.; Rosean, T.R.; Borcherding, N.; Sciegienka, S.; Goeken, J.A.; Naumann, P.W.; Zhang, W.; Tompkins, V.S.; et al. NIAM-deficient mice are predisposed to the development of proliferative lesions in-cluding B-cell lymphomas. PLoS ONE 2014, 9, e112126. [Google Scholar] [CrossRef] [Green Version]
- Slezak-Prochazka, I.; Kluiver, J.; De Jong, D.; Smigielska-Czepiel, K.; Kortman, G.; Winkle, M.; Rutgers, B.; Koerts, J.; Visser, L.; Diepstra, A.; et al. Inhibition of the miR-155 target NIAM phenocopies the growth promoting effect of miR-155 in B-cell lymphoma. Oncotarget 2015, 7, 2391–2400. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tie, Y.; Chen, C.; Yang, Y.; Qian, Z.; Yuan, H.; Wang, H.; Tang, H.; Peng, Y.; Du, X.; Liu, B. Upregulation of let-7f-5p promotes chemotherapeutic resistance in colorectal cancer by directly repressing several pro-apoptotic proteins. Oncol. Lett. 2018, 15, 8695–8702. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.; Kluiver, J.; Koerts, J.; De Jong, D.; Rutgers, B.; Razak, F.R.A.; Terpstra, M.; Plaat, B.E.; Nolte, I.M.; Diepstra, A.; et al. miR-24-3p Is Overexpressed in Hodgkin Lymphoma and Protects Hodgkin and Reed-Sternberg Cells from Apoptosis. Am. J. Pathol. 2017, 187, 1343–1355. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, X.; Zhou, X.; Wang, Z.; Li, F.; Liu, F.; Zhong, L.; Li, X.; Han, X.; Wu, Z.; Chen, S.; et al. CD99 triggers upregulation of miR-9-modulated PRDM1/BLIMP1 in Hodg-kin/Reed-Sternberg cells and induces redifferentiation. Int. J. Cancer 2012, 131, E382–E394. [Google Scholar] [CrossRef]
- Leucci, E.; Zriwil, A.; Gregersen, L.H.; Jensen, K.T.; Obad, S.; Bellan, C.; Leoncini, L.; Kauppinen, S.; Lund, A. Inhibition of miR-9 de-represses HuR and DICER1 and impairs Hodgkin lymphoma tumour outgrowth in vivo. Oncogene 2012, 31, 5081–5089. [Google Scholar] [CrossRef] [Green Version]
- Xin, H.; Wang, C.; Liu, Z. miR-196a-5p promotes metastasis of colorectal cancer via targeting IκBα. BMC Cancer 2019, 19, 30. [Google Scholar] [CrossRef]
- Danger, R.; Braza, F.; Giral, M.; Soulillou, J.-P.; Brouard, S. MicroRNAs, Major Players in B Cells Homeostasis and Function. Front. Immunol. 2014, 5. [Google Scholar] [CrossRef] [Green Version]
- Robaina, M.C.; Mazzoccoli, L.; Klumb, C.E.; Klumb, E. Germinal Centre B Cell Functions and Lymphomagenesis: Circuits Involving MYC and MicroRNAs. Cells 2019, 8, 1365. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Batista, C.R.; Li, S.K.H.; Xu, L.S.; Solomon, L.A.; DeKoter, R.P. PU.1 Regulates Ig Light Chain Transcription and Rearrangement in Pre-B Cells during B Cell Development. J. Immunol. 2017, 198, 1565–1574. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oettgen, P.; Akbarali, Y.; Boltax, J.; Best, J.; Kunsch, C.; Libermann, T.A. Characterization of NERF, a novel transcription factor re-lated to the Ets factor ELF-1. Mol. Cell Biol. 1996, 16, 5091–5106. [Google Scholar] [CrossRef] [Green Version]
- Wertz, I.E.; O’rourke, K.M.; Zhou, H.; Eby, M.; Aravind, L.; Seshagiri, S.; Wu, P.; Wiesmann, C.; Baker, R.; Boone, D.L.; et al. De-ubiquitination and ubiquitin ligase domains of A20 downregulate NF-kappaB signalling. Nature 2004, 430, 694–699. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Mei, H.; Sheng, Z.; Meng, Q.; Véniant, M.M.; Yin, H. Hsa-miRNA-23a-3p promotes atherogenesis in a novel mouse model of atherosclerosis. J. Lipid Res. 2020, 61, 1764–1775. [Google Scholar] [CrossRef]
- Wei, C.; Zhang, R.; Cai, Q.; Gao, X.; Tong, F.; Dong, J.; Hu, Y.; Wu, G.; Dong, X. MicroRNA-330-3p promotes brain metastasis and epithelial-mesenchymal transition via GRIA3 in non-small cell lung cancer. Aging 2019, 11, 6734–6761. [Google Scholar] [CrossRef]
- Mesci, A.; Huang, X.; Taeb, S.; Jahangiri, S.; Kim, Y.; Fokas, E.; Bruce, J.; Leong, H.; Liu, S.K. Targeting of CCBE1 by miR-330-3p in human breast cancer promotes metastasis. Br. J. Cancer 2017, 116, 1350–1357. [Google Scholar] [CrossRef]
- Ye, Y.-P.; Wu, P.; Gu, C.-C.; Deng, D.-L.; Jiao, H.-L.; Li, T.-T.; Wang, S.-Y.; Wang, Y.-X.; Xiao, Z.-Y.; Wei, W.-T.; et al. miR-450b-5p induced by oncogenic KRAS is required for colorectal cancer progression. Oncotarget 2016, 7, 61312–61324. [Google Scholar] [CrossRef] [Green Version]

| GO Analysis of Target Genes of miRNAs Upregulated in cHL | |||
|---|---|---|---|
| Top 3 GO Terms (p < 0.05) | PANTHER Database | STRING Database | DAVID Database |
| 1 | DNA-binding transcription factor activity, RNA polymerase II-specific (GO:0000981) | DNA-binding transcription factor activity (GO:0003700) | DNA-binding transcription factor activity (GO:0003700) |
| 2 | DNA-binding transcription factor activity (GO:0003700) | molecular function regulator (GO:0098772) | protein binding (GO:0005515) |
| 3 | molecular function regulator (GO:0098772) | transcription regulator activity (GO:0140110) | regulation of transcription, DNA-templated (GO:0006355) |
| GO analysis of target genes of miRNAs downregulated in cHL | |||
| Top 3 GO terms (p < 0.05) | PANTHER database | STRING database | DAVID database |
| 1 | transcription regulatory region sequence-specific DNA binding (GO:0000976) | DNA-binding transcription factor activity, RNA polymerase II-specific (GO:0000981) | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding (GO:0001077) |
| 2 | RNA polymerase II regulatory region DNA binding (GO:0001012) | transcription regulatory region sequence-specific DNA binding (GO:0000976) | neuron migration (GO:0001764) |
| 3 | transcription regulatory region DNA binding (GO:0044212) | RNA polymerase II regulatory region sequence-specific DNA binding (GO:0000977) | RNA polymerase II core promoter proximal region sequence-specific DNA binding (GO:0000978) |
| MiRNA in cHL | MiRNA | Target Gene | UTR miRNA Binding Site [Based on TargetScan Database] | Interaction Confirmed in the Luciferase Assay | Gene Function | ||
|---|---|---|---|---|---|---|---|
| Binding Sites Conservation | Number of Binding Sites | Characteristic of Binding Sites | |||||
| UPREGULATED | hsa-miR-27a-5p | SPI1 (PU.1) | poorly conserved | 1 | 7mer-m8 | YES | B-cell related transcription factor |
| hsa-miR-330-3p | ELF-1 | poorly conserved | 2 | 7mer-m8, 7mer-m8 | YES | B-cell related transcription factor | |
| hsa-miR-542-3p | ELF-1 | poorly conserved | 2 | 7mer-A1, 7mer-m8 | NO | B-cell related transcription factor | |
| hsa-miR-450b-5p | ELF-1 | poorly conserved | 1 | 7mer-A1 | YES | B-cell related transcription factor | |
| hsa-miR-23a-3p | TNFAIP3 | conserved | 1 | 8mer | YES | negative regulator of NF-kappaB | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Paczkowska, J.; Janiszewska, J.; Ustaszewski, A.; Bein, J.; Skalski, M.; Dzikiewicz-Krawczyk, A.; Rozwadowska, N.; Hansmann, M.-L.; Hartmann, S.; Giefing, M. Deregulated miRNAs Contribute to Silencing of B-Cell Specific Transcription Factors and Activation of NF-κB in Classical Hodgkin Lymphoma. Cancers 2021, 13, 3131. https://doi.org/10.3390/cancers13133131
Paczkowska J, Janiszewska J, Ustaszewski A, Bein J, Skalski M, Dzikiewicz-Krawczyk A, Rozwadowska N, Hansmann M-L, Hartmann S, Giefing M. Deregulated miRNAs Contribute to Silencing of B-Cell Specific Transcription Factors and Activation of NF-κB in Classical Hodgkin Lymphoma. Cancers. 2021; 13(13):3131. https://doi.org/10.3390/cancers13133131
Chicago/Turabian StylePaczkowska, Julia, Joanna Janiszewska, Adam Ustaszewski, Julia Bein, Marcin Skalski, Agnieszka Dzikiewicz-Krawczyk, Natalia Rozwadowska, Martin-Leo Hansmann, Sylvia Hartmann, and Maciej Giefing. 2021. "Deregulated miRNAs Contribute to Silencing of B-Cell Specific Transcription Factors and Activation of NF-κB in Classical Hodgkin Lymphoma" Cancers 13, no. 13: 3131. https://doi.org/10.3390/cancers13133131