Comprehensive Volatilome and Metabolome Signatures of Colorectal Cancer in Urine: A Systematic Review and Meta-Analysis
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
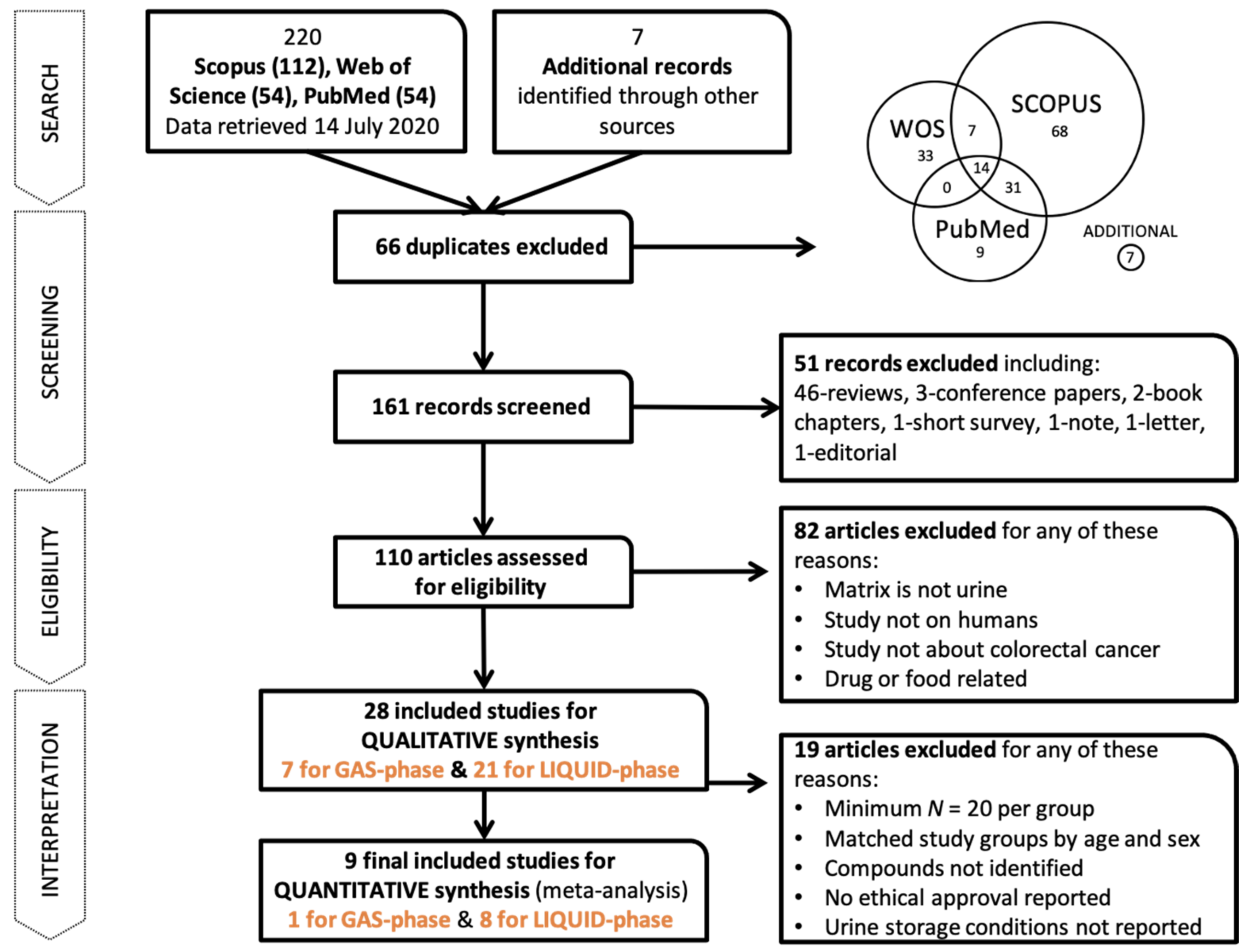
2.1. The PRISMA Method
2.2. Search Sentence
2.3. Inclusion and Exclusion Criteria
2.4. Meta-Analysis
3. Results
3.1. PRISMA Process
3.2. Characteristics of the Included Studies
| Ref. | Kind | Platform | Type of Study | Ethics Approval | Urine Collection | Urine Storage | Analytical Validation | ROC Curve (Training/Testing) |
|---|---|---|---|---|---|---|---|---|
| [28] | VOL | FAIMS + GC-MS | CRC/control | yes | ND | −80 °C | 1/2–1/2 repeated 5 times | - |
| * [35] | MET | GC + LC | CRC/control | yes | Fasting urine | −80 °C | 2/3–1/3 | 0.993 (7 compounds)/0.998 |
| [32] | MET | RRLC-TOF/MS | CRC/control | yes | First morning urine | −80 °C | 2/3–1/3 | - |
| * [36] | MET | NMR | CRC/control | yes | First morning urine | −80 °C | - | 0.823 taurine, 0.783 alanine, 0.842 3-aminoisobutyrate/ND |
| [33] | MET | LC-FAIMS-MS | CRC/control | yes with ID | ND | −80 °C | - | 0.71/ND |
| * [25] | MET | Targeted LC-MS/MS | CRC/control | yes with ID | Controls at 7–8 a.m., 11–12 a.m., and 5–6 p.m. CRC at 9 a.m. and 4 p.m. | −80 °C | bootstrapping with virtual datasets | 0.794/ND |
| [31] | MET | CE-MS | CRC/control (including stages) | yes | Morning urine | −80 °C | - | 0.906/ND |
| * [24] | MET | Targeted LC-MS/MS | CRC/control (including stages) | yes with ID | ND | −80 °C | yes | 0.903/0.872 |
| * [37] | MET | 1H-NMR | CRC/control (including stages + other cancers) | yes | Fasting morning urine | −80 °C | 80% training, 20% testing | 0.875 alanine, 0.913 glutamine, 0.933 aspartic acid/ND |
| [38] | MET | HPLC-ESI-MS/MS | CRC/control (+ other cancers) | yes | ND | −80 °C | - | - |
| [39] | MET | LC-MS/MS MRM | CRC/control (+ other cancers) | yes | First morning urine | −80 °C | - | - |
| * [29] | VOL | Needle trap + GC-MS | CRC/control (+ other cancer) | yes | First morning urine | −80 °C | 2/3–1/3 | - |
| [40] | VOL | GC-MS | CRC/control (+ other cancers) | yes | ND | ND | - | - |
| [41] | VOL | GC-MS | CRC/control (+ other cancers) | yes | Fasting morning urine | −80 °C | - | - |
| [30] | VOL | E-nose | CRC/control (+ other diseases) | yes | Fasting morning urine | −80 °C | - | - |
| [42] | MET | RP-HPLC | CRC/control (along time) | yes | Spontaneous urine samples 1 day before surgery and day 8 after | −20 °C | - | 0.896 1-methylguanosine, 0.816 pseudouridine/ND |
| [22] | MET | UPLC-QTOF-MS | CRC no-relapse/relapse | ND | ND | ND | 2-fold cross-validation with 10,000 validations | AUC: 0.9675 (positive charge) and 0.95 (negative charge)/ND |
| * [43] | MET | GC-MS | CRC/control (pre-/post-surgery) | yes | Fasting morning urine | −80 °C | 16/17–1/17 | - |
| * [44] | MET | 1H-NMR + GC-MS | CRC pre-/post-surgery and 6-/12-months follow-up AND intra-stages | yes | Pre-/post-surgery overnight fasting urine, 6-/12-months follow-up URINE spot | −80 °C | - | 0.89 (20 compounds)/ND |
| [45] | MET | UPLC-MS | CRC/control (pre-/post-surgery + along time) | yes | Fasting urine (7:00 a.m.) | −80 °C | - | - |
| [26] | MET | targeted HPLC/GC-MS | CRC/adenoma/control | yes | Spot sample before surgery | ND | - | 0.690 8-oxoGua, 0.635 8-oxoGuo, 0.669 5-hmUra/ND |
| [46] | MET | UPLC-MS/HPLC-MS | CRC/adenoma/control | yes | Morning fasting urine | −80 °C | 7-fold | 0.959 (12 compounds), 0.894 (7 nucleotides)/ND |
| [47] | MET | HPLC-MS/MS | CRC/adenoma/control | ND | Spot urine | −20 °C | - | - |
| [48] | VOL | FAIMS + GC-IMS | CRC/ adenoma/control (+ other diseases) | yes | ND | −80 °C | - | 0.98/ND |
| [49] | VOL | FAIMS | CRC/adenoma/control (+ other cancers and diseases) | yes | Spot urine | −80 °C | - | 0.9/ND |
| [23] | MET | NMR + targeted LC-MS/MS | Adenoma/control | yes with ID | Midstream urine | −80 °C ǂ | 2/3–1/3 | 0.687/0.692 |
| [27] | MET | 1D NMR | Adenoma/control | yes | Midstream urine | 4 h at 4 °C 24 h at −80 °C | Validation of Deng L 2017 | 0.717/ND |
| * [50] | MET | 1D NMR | Adenoma/control | yes | Midstream urine | 4 h at 4 °C 24 h at −80 °C | 2/3–1/3 | 0.752/ND |
| Ref. (Kind) | Group | N | Age (Error and Type) | Male/Female | Cancer Staging Classification (n) | Country |
|---|---|---|---|---|---|---|
| [28] | CRC | 83 | 60 (ND: 17) | 53/30 | ND | UK |
| (VOL) | Control | 50 | 47 (ND: 16) | 21/29 | - | |
| * [35] | CRC | 101 | 60 (R: 24–83) | 58/43 | 0 (0), I (24), II (45), III (27), IV (5) | CN |
| (MET) | Control | 103 | 58 (R: 31–76) | 31/72 | ||
| [32] | CRC | 29 | ND | - | ND | CN |
| (MET) | Control | 10 | ND | - | - | |
| * [36] | CRC | 92 | 60 (R: 32–85) | 62/30 | 0 (24), I (8), II (7), III (13), IV (4) | KR |
| (MET) | Control | 156 | 52 (R: 22–76) | 76/80 | ||
| [33] | CRC | 56 | 65.4 (SD: 11.5) | 33/23 | A (8), B (17), C1 (20), C2 (9) | UK |
| (MET) | Control (spouse) | 45 | 60.7 (SD: 12.1) | 15/30 | - | |
| Control (relative) | 37 | 50 (SD: 14.1) | 17/20 | - | ||
| * [25] | CRC-Malignant | 201 | 68.7 (ND: 0.8) | 114/87 | 0 (3), I/II (103), III (88), IV (7) | JP |
| (MET) | CRC-Benign | 14 | 65 (ND: 3.1) | 11/3 | - | |
| Control | 17 | 42.1 (ND: 2.8) | 13/4 | - | ||
| [31] | CRC | 20 | 73 (ND) | 10/10 | I/II (8), III/IV (12) | CN |
| (MET) | Control | 14 | 68 (ND) | 8/6 | - | |
| * [24] | CRC-CAD | 121 | 67.4 (ND: 10.9) | 68/59 | 0 (3); I (16), II (30), III (51), IV (21) | CA/US |
| (MET) | CRC-MSKCC | 50 | 63.8 (ND: 12.5) | 24/26 | 0 (0), I (14), II (20), III (6), IV (10) | |
| Control | 171 | 58.9 (ND: 5.6) | 100/71 | - | ||
| * [37] | CRC | 55 | 60 (ND) | 26/29 | I/II (23), III/IV (32) | CN |
| (MET) | Control | 40 | 59 (ND) | 19/21 | - | |
| EC | 18 | 61 (ND) | 8/10 | - | ||
| [38] | CRC | 26 | 65.3 (R: 33–88) | 12/24 | 0 (0), I (3), II (6), III (10), IV (7) | TW |
| (MET) | Control | 45 | ND | - | - | |
| LC | 27 | 60.8 (R: 42–81) | 16/11 | - | ||
| GC | 15 | 67.1 (R: 50–82) | 12/3 | - | ||
| BC | 36 | ND | - | - | ||
| [39] | CRC | 10 | 51.5 (SD: 6.6) | 5/5 | ND | CN |
| (MET) | Control | 10 | 48.7 (SD: 6.43) | 5/5 | - | |
| LC | 10 | 52.5 (SD: 7.47) | 5/5 | - | ||
| NpC | 10 | 49.3 (SD: 9.09) | 5/5 | - | ||
| * [29] | CRC | 30 | ND (R: 45–83) | 16/14 | ND | PT |
| (VOL) | Control | 30 | ND (R: 18–78) | 14/16 | - | |
| BC | 30 | ND (R: 38–83) | 0/30 | - | ||
| [40] | CRC | 8 | ND | - | ND | ND |
| (VOL) | Control | 35 | ND | - | - | |
| LC | 14 | ND | - | - | ||
| EC | 12 | ND | - | - | ||
| GC | 12 | ND | - | - | ||
| [41] | CRC | 11 | 62 (SD: 12.4 R: 49–78) | 8/3 | ND | PT |
| (VOL) | Control | 21 | 62 (SD: 10.3 R: 28–60) | 18/3 | - | |
| LeukC | 14 | 50.1 (SD: 12.4 R: 40–74) | 6/8 | - | ||
| LyC | 7 | 42 (SD: 19.1 R: 18–68) | 6/1 | - | ||
| [30] | CRC | 39 | 70 (ND) | 28/11 | ND | UK |
| (VOL) | Control | 18 | 41 (ND) | 13/5 | - | |
| IBS | 35 | 48 (ND) | 4/31 | - | ||
| [42] | CRC | 52 | 63 (R: 26–87) | 27/25 | A (5), B (22), C (18), D (7) | CN |
| (MET) | Control | 62 | 59 (R: 24–78) | 33/29 | - | |
| [22] | CRC non-relapse | 20 | ND | - | ND | |
| (MET) | CRC relapse | 20 | ND | - | ND | |
| * [43] | CRC | 60 | 58.8 (ND) | 34/26 | 0 (0), I (7), II (23), III (21), IV (9) | CN |
| (MET) | Control | 63 | 55.5 (ND) | 32/31 | - | |
| * [44] | CRC pre-S | 97 | 64.8 (SD: 12.9) | 59/38 | 0 (5), I (12), II (40), III (22), IV (18) | DE |
| (MET) | CRC post-S | 12 | 63.9 (SD: 12.5) | 10/2 | 0 (0), I (4), II (4), III (2), IV (2) | |
| CRC (6 m) | 52 | 60.1 (SD: 11) | 38/14 | 0 (0), I (12), II (17), III (15), IV (8) | ||
| CRC (12 m) | 38 | 61.5 (SD: 11.6) | 24/14 | 0 (0), I (7), II (13), III (14), IV (4) | ||
| [45] | CRC | 24 | 65.03 (SD: 10.43) | 13/11 | A (1), B (1), C (12), D (0) | CN |
| (MET) | Control | 80 | 64 (SD: 9.87) | 43/37 | - | |
| [26] | Adenoma | 15 | 66 (ND) | 8/7 | - | PL |
| (MET) | CRC | 72 | 54 (ND) | 31/41 | ND | |
| Control | 56 | 65 (ND) | 32/24 | - | ||
| [46] | CRC-Malignant | 94 | ND | - | ND | CN |
| (MET) | Control | 34 | ND | - | - | |
| [47] | Adenoma | 10 | ND | - | - | CN |
| (MET) | CRC | 52 | 60 (R: 26–87) | 29/23 | A (7), B (23), C (15), D (7) | |
| Control | 60 | 52 (R: 21–71) | 31/39 | - | ||
| [48] | Adenoma | 80 | 67 (ND)ǂ | 93/70ǂ | - | UK |
| (VOL) | CRC | 12 | ND | |||
| Control | 83 | - | ||||
| Other (DD, Hemorrhoids, etc.) | 33 | - | ||||
| [49] | Adenoma | 94 | 68 (R: 29–89) ǂ | 286/276 ǂ | - | UK |
| (VOL) | High risk adenoma | 27 | - | |||
| CRC | 35 | ND | ||||
| Control | 233 | - | ||||
| Others (DD, IBD, MC, etc.,) | 173 | - | ||||
| [23] | Adenoma | 155 | 59.9 (SD: 7.4) | 95/60 | ND | CA |
| (MET) | Control | 530 | 56.1 (SD: 8.2) | 222/308 | - | |
| [27] | Adenoma | 345 | 65.1 (SEM: 6.6) | 197/148 | ND | CN |
| (MET) | Control | 316 | 61.8 (SEM: 7.4) | 82/234 | - | |
| * [50] | Adenoma | 243 | 59.5 (SEM: 0.67) | 145/98 | ND | CA |
| (MET) | Control | 633 | 55.8 (SEM: 0.47) | 269/364 | - |
3.3. Systematic Review
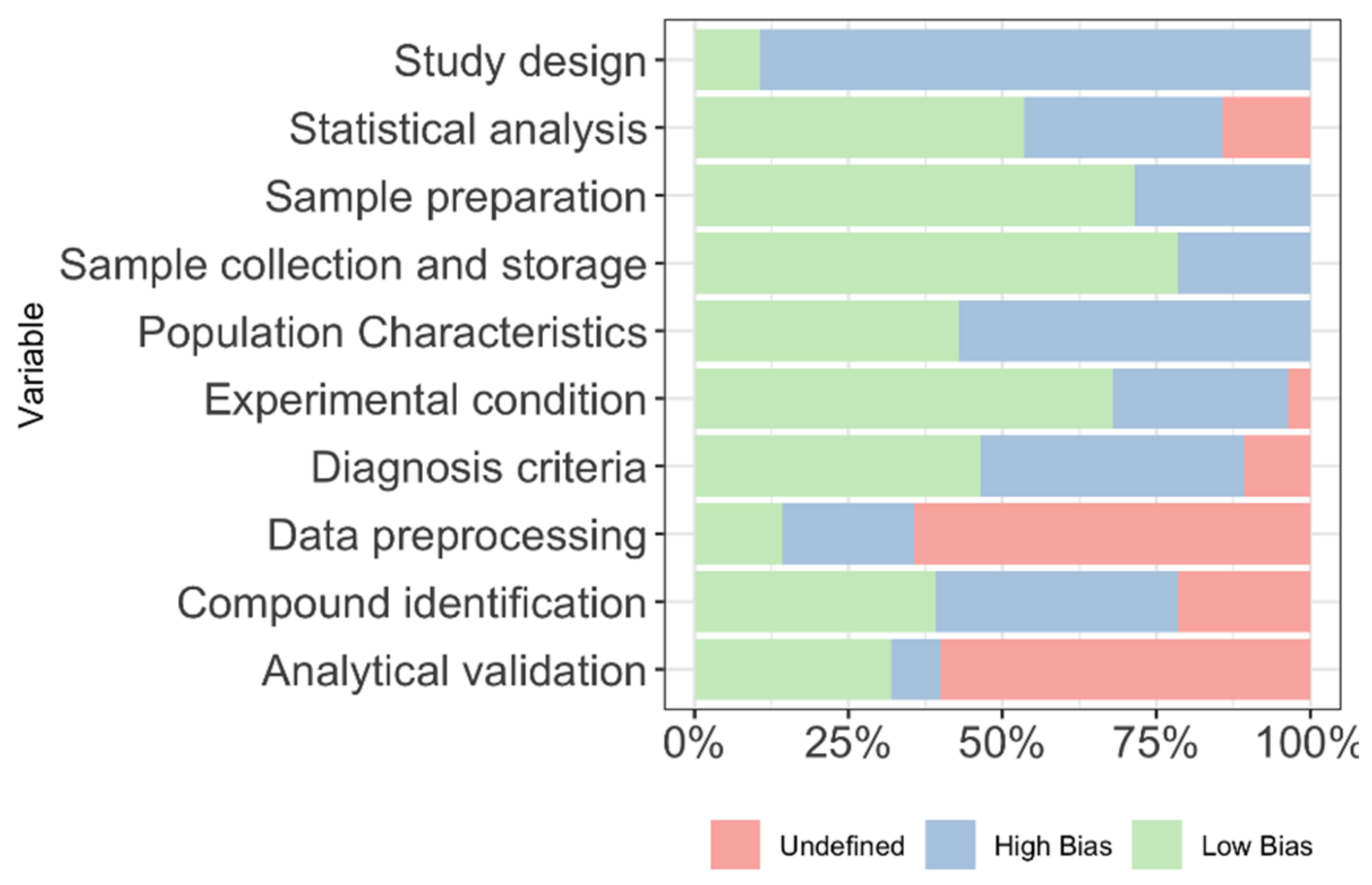
3.4. Quality Assurance
3.5. Meta-Analysis
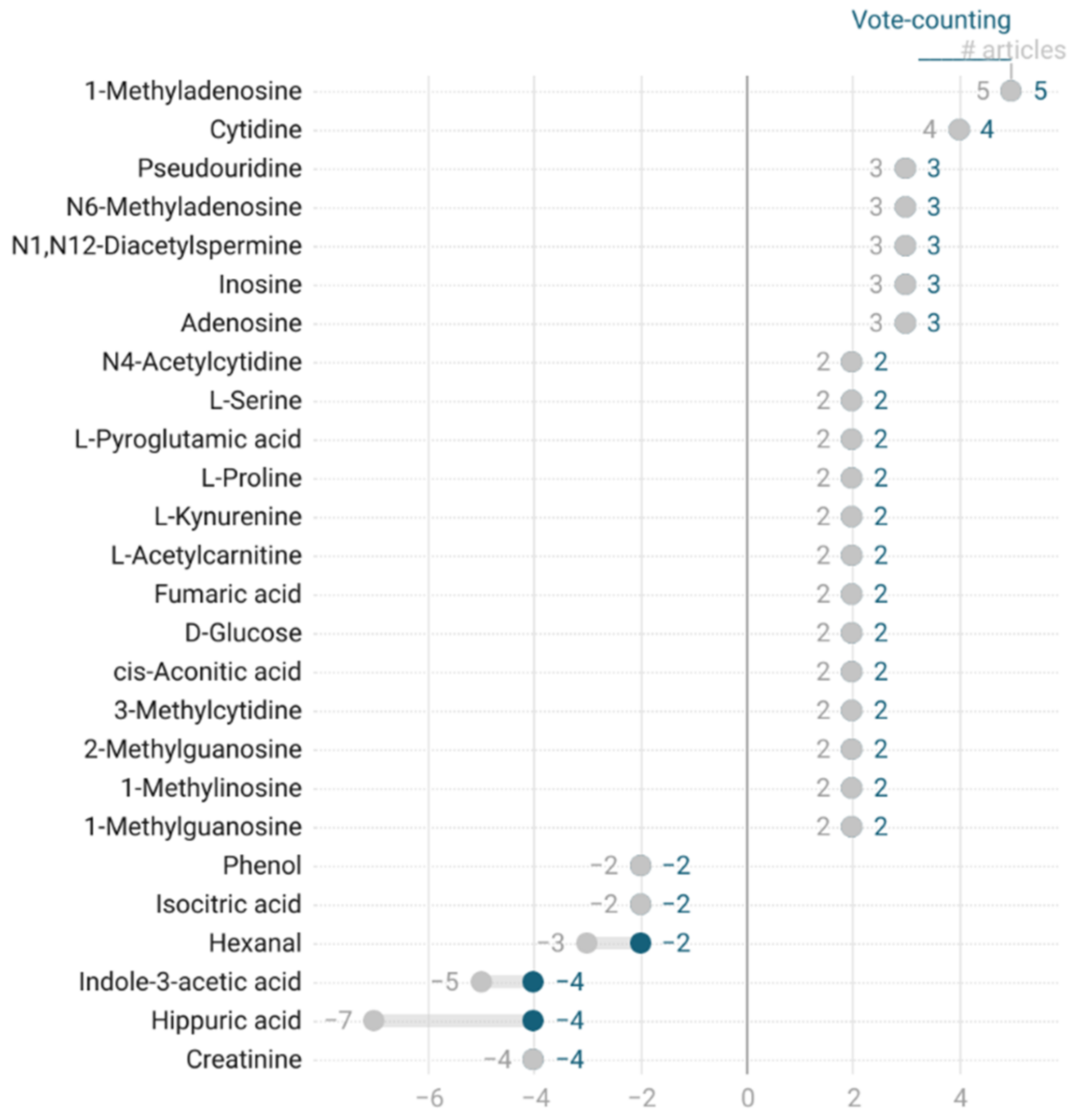
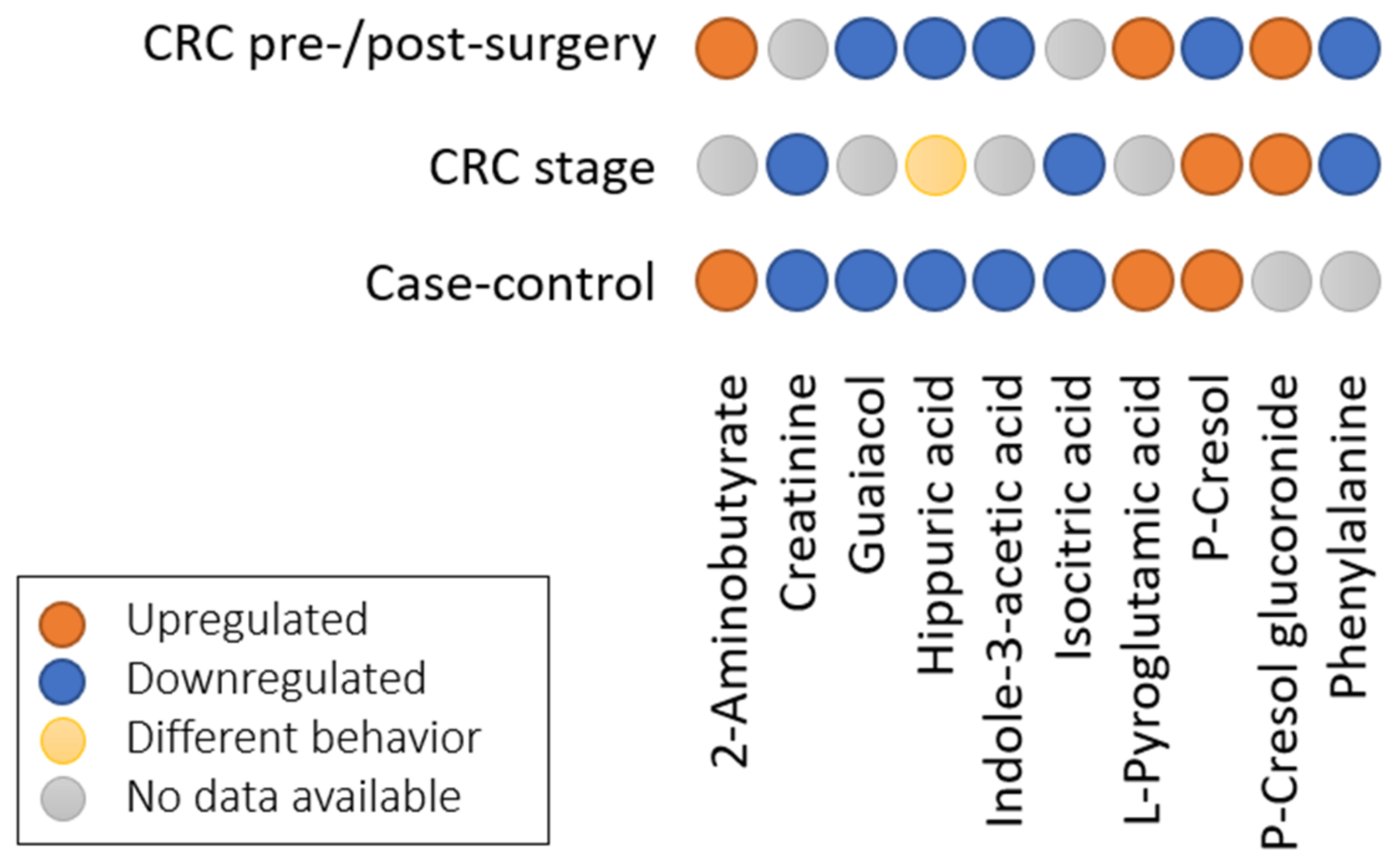
3.5.1. Vote-Counting Results by Group
| Common Name | No. of Cohorts | Behavior (Up-Down-Equal) | Vote-Counting | N | Reference |
|---|---|---|---|---|---|
| CRC and Advanced Adenoma vs. Control | |||||
| N1,N12-Diacetylspermine | 3 | 3–0–0 | 3 | 928 | [24,25] ɫ |
| D-Glucose | 2 | 2–0–0 | 2 | 696 | [24] ɫ |
| L-Kynurenine | 2 | 2–0–0 | 2 | 696 | [24] ɫ |
| L-Proline | 2 | 2–0–0 | 2 | 696 | [24] ɫ |
| Creatinine | 2 | 0–2–0 | −2 | 452 | [35,36] |
| Phenol | 2 | 0–2–0 | −2 | 294 | [29,35] |
| Putrescine | 3 | 2–0–1 | 2 | 900 | [24,35] ɫ |
| Hippuric acid | 4 | 0–3–1 | −3 | 1148 | [24,35,36] ɫ |
| Indole-3-acetic acid | 3 | 0–2–1 | −2 | 900 | [24,35] ɫ |
| Citric acid | 5 | 1–3–1 | −2 | 1271 | [24,35,36,43] ɫ |
| P-Cresol | 2 | 1–1–0 | 1 | 417 | [35,43] |
| Tetradecenoyl carnitine (C14:1) | 2 | 1–0–1 | 1 | 696 | [24] ɫ |
| 2-Aminohexanedioic acid | 2 | 0–1–1 | −1 | 696 | [24] ɫ |
| 3-(3-Hydroxyohenyl)-3-hydroxypropanoic acid | 2 | 0–1–1 | −1 | 696 | [24] ɫ |
| Aspartic acid | 2 | 0–1–1 | −1 | 696 | [24] ɫ |
| 3-Hidroxybutyric acid | 2 | 1–1–0 | 0 | 696 | [24] ɫ |
| Butyric acid | 2 | 1–1–0 | 0 | 696 | [24] ɫ |
| Hydroxyproline | 2 | 1–1–0 | 0 | 696 | [24] ɫ |
| L-Alanine | 2 | 1–1–0 | 0 | 452 | [35,36] |
| L-Dopa | 2 | 1–1–0 | 0 | 696 | [24] ɫ |
| L-Tryptophan | 2 | 1–1–0 | 0 | 327 | [35,43] |
| Urea | 2 | 1–1–0 | 0 | 452 | [35,36] |
| CRC Stage vs. Control | |||||
| Hippuric acid | 2 | 1–1–0 | 0 | 248 | [37,44] |
| Pre-surgery vs. Post-surgery | |||||
| Salicyluric acid | 2 | 0–2–0 | −2 | 258 | [43,44] |
| Asparagine | 2 | 1–1–0 | 0 | 258 | [43,44] |
| Citrate | 2 | 1–1–0 | 0 | 258 | [43,44] |
| Tyrosine | 2 | 1–1–0 | 0 | 246 | [43,44] |
3.5.2. Statistical Results by Group
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Kupfer, S.S.; Ellis, N.A. Hereditary colorectal cancer. Mol. Basis Hum. Cancer 2016, 381–400. [Google Scholar] [CrossRef]
- Fernandez-Villa, T.; Alvarez-Alvarez, L.; Rubin-Garcia, M.; Obon-Santacana, M.; Moreno, V. The role of dietary patterns in colorectal cancer: A 2019 update. Expert Rev. Gastroenterol. Hepatol. 2020, 14, 281–290. [Google Scholar] [CrossRef] [PubMed]
- Vernia, F.; Longo, S.; Stefanelli, G.; Viscido, A.; Latella, G. Dietary Factors Modulating Colorectal Carcinogenesis. Nutrients 2021, 13, 143. [Google Scholar] [CrossRef]
- Clarke, J.M.; Lockett, T. Primary prevention of colorectal cancer. Cancer Forum 2014, 38, 6–10. [Google Scholar] [CrossRef]
- Hamilton, W.; Round, A.; Sharp, D.; Peters, T.J. Clinical features of colorectal cancer before diagnosis: A population-based case-control study. Br. J. Cancer 2005, 93, 399–405. [Google Scholar] [CrossRef]
- Lin, J.S.; Piper, M.A.; Perdue, L.A.; Rutter, C.M.; Webber, E.M.; O’Connor, E.; Smith, N.; Whitlock, E.P. Screening for colorectal cancer: Updated evidence report and systematic review for the US preventive services task force. JAMA—J. Am. Med. Assoc. 2016, 315, 2576–2594. [Google Scholar] [CrossRef]
- Elsafi, S.H.; Alqahtani, N.I.; Zakary, N.Y.; Al Zahrani, E.M. The sensitivity, specificity, predictive values, and likelihood ratios of fecal occult blood test for the detection of colorectal cancer in hospital settings. Clin. Exp. Gastroenterol. 2015, 8, 279–284. [Google Scholar] [CrossRef]
- Nielson, C.M.; Petrik, A.F.; Jacob, L.; Vollmer, W.M.; Keast, E.M.; Schneider, J.L.; Rivelli, J.S.; Kapka, T.J.; Meenan, R.T.; Mummadi, R.R.; et al. Positive predictive values of fecal immunochemical tests used in the STOP CRC pragmatic trial. Cancer Med. 2018, 7, 4781–4790. [Google Scholar] [CrossRef]
- Robertson, D.J.; Lee, J.K.; Boland, C.R.; Dominitz, J.A.; Giardiello, F.M.; Johnson, D.A.; Kaltenbach, T.; Lieberman, D.; Levin, T.R.; Rex, D.K. Recommendations on fecal immunochemical testing to screen for colorectal neoplasia: A consensus statement by the US Multi-Society Task Force on colorectal cancer. Gastrointest. Endosc. 2017, 85, 2–21.e3. [Google Scholar] [CrossRef]
- Saraceni, A.F.; Azevedo, R.; Almeida, C.M.G.; Baraviera, A.C.; Kiss, D.R.; Almeida, M.G. Association of fecal occult blood tests results with colonoscopic findings in a general hospital and validation of the screening test. J. Coloproctology 2019, 39, 121–126. [Google Scholar] [CrossRef]
- Davis, C.E.; Pleil, J.D.; Beauchamp, J.D. (Eds.) Breathborne Biomarkers and the Human Volatilome; Elsevier: Amsterdam, The Netherlands, 2020. [Google Scholar]
- Buszewski, B.; Rudnicka, J.; Ligor, T.; Walczak, M.; Jezierski, T.; Amann, A. Analytical and unconventional methods of cancer detection using odor. TrAC—Trends Anal. Chem. 2012, 38, 1–12. [Google Scholar] [CrossRef]
- Amann, A.; Costello, B.D.L.; Miekisch, W.; Schubert, J.; Buszewski, B.; Pleil, J.; Ratcliffe, N.; Risby, T. The human volatilome: Volatile organic compounds (VOCs) in exhaled breath, skin emanations, urine, feces and saliva. J. Breath Res. 2014, 8, 034001. [Google Scholar] [CrossRef]
- Bouatra, S.; Aziat, F.; Mandal, R.; Guo, A.C.; Wilson, M.R.; Knox, C.; Bjorndahl, T.C.; Krishnamurthy, R.; Saleem, F.; Liu, P.; et al. The Human Urine Metabolome. PLoS ONE 2013, 8, e73076. [Google Scholar] [CrossRef] [PubMed]
- Moher, D.; Liberati, A.; Tetzlaff, J.; Altman, D.G.; Altman, D.; Antes, G.; Atkins, D.; Barbour, V.; Barrowman, N.; Berlin, J.A.; et al. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA statement. PLoS Med. 2009, 6. [Google Scholar] [CrossRef]
- Edwards, P.; Clarke, M.; DiGuiseppi, C.; Pratap, S.; Roberts, I.; Wentz, R. Identification of randomized controlled trials in systematic reviews: Accuracy and reliability of screening records. Stat. Med. 2002, 21, 1635–1640. [Google Scholar] [CrossRef]
- Billoir, E.; Navratil, V.; Blaise, B.J. Sample size calculation in metabolic phenotyping studies. Brief. Bioinform. 2014, 16, 813–819. [Google Scholar] [CrossRef]
- Gika, H.G.; Theodoridis, G.A.; Wilson, I.D. Liquid chromatography and ultra-performance liquid chromatography-mass spectrometry fingerprinting of human urine. Sample stability under different handling and storage conditions for metabonomics studies. J. Chromatogr. A 2008, 1189, 314–322. [Google Scholar] [CrossRef] [PubMed]
- Llambrich, M.; Correig, E.; Gumà, J.; Brezmes, J.; Cumeras, R. Amanida: An R package for meta-analysis of metabolomics non-integral data. Bioinformatics 2021. Submitted. [Google Scholar]
- Morgan, A.A.; Khatri, P.; Jones, R.H.; Sarwal, M.M.; Butte, A.J. Comparison of multiplex meta analysis techniques for understanding the acute rejection of solid organ transplants. BMC Bioinform. 2010, 11 (Suppl. 9), S6. [Google Scholar] [CrossRef]
- Madhavan, S.; Gusev, Y.; Natarajan, T.G.; Song, L.; Bhuvaneshwar, K.; Gauba, R.; Pandey, A.; Haddad, B.R.; Goerlitz, D.; Cheema, A.K.; et al. Genome-wide multi-omics profiling of colorectal cancer identifies immune determinants strongly associated with relapse. Front. Genet. 2013, 4, 236. [Google Scholar] [CrossRef] [PubMed]
- Deng, L.; Chang, D.; Foshaug, R.R.; Eisner, R.; Tso, V.K.; Wishart, D.S.; Fedorak, R.N. Development and validation of a high-throughput mass spectrometry based urine metabolomic test for the detection of colonic adenomatous polyps. Metabolites 2017, 7, 32. [Google Scholar] [CrossRef] [PubMed]
- Deng, L.; Ismond, K.; Liu, Z.; Constable, J.; Wang, H.; Alatise, O.I.; Weiser, M.R.; Kingham, T.P.; Chang, D. Urinary metabolomics to identify a unique biomarker panel for detecting colorectal cancer: A multicenter study. Cancer Epidemiol. Biomark. Prev. 2019, 28, 1283–1292. [Google Scholar] [CrossRef] [PubMed]
- Nakajima, T.; Katsumata, K.; Kuwabara, H.; Soya, R.; Enomoto, M.; Ishizaki, T.; Tsuchida, A.; Mori, M.; Hiwatari, K.; Soga, T.; et al. Urinary polyamine biomarker panels with machine-learning differentiated colorectal cancers, benign disease, and healthy controls. Int. J. Mol. Sci. 2018, 19, 756. [Google Scholar] [CrossRef]
- Rozalski, R.; Gackowski, D.; Siomek-Gorecka, A.; Starczak, M.; Modrzejewska, M.; Banaszkiewicz, Z.; Olinski, R. Urinary 5-hydroxymethyluracil and 8-oxo-7,8-dihydroguanine as potential biomarkers in patients with colorectal cancer. Biomarkers 2015, 20, 287–291. [Google Scholar] [CrossRef]
- Deng, L.; Fang, H.; Tso, V.K.; Sun, Y.; Foshaug, R.R.; Krahn, S.C.; Zhang, F.; Yan, Y.; Xu, H.; Chang, D.; et al. Clinical validation of a novel urine-based metabolomic test for the detection of colonic polyps on Chinese population. Int. J. Colorectal Dis. 2017, 32, 741–743. [Google Scholar] [CrossRef]
- Arasaradnam, R.P.; Mcfarlane, M.J.; Ryan-Fisher, C.; Westenbrink, E.; Hodges, P.; Thomas, M.G.; Chambers, S.; O’Connell, N.; Bailey, C.; Harmston, C.; et al. Detection of colorectal cancer (CRC) by urinary volatile organic compound analysis. PLoS ONE 2014, 9. [Google Scholar] [CrossRef]
- Porto-Figueira, P.; Pereira, J.A.M.; Câmara, J.S. Exploring the potential of needle trap microextraction combined with chromatographic and statistical data to discriminate different types of cancer based on urinary volatomic biosignature. Anal. Chim. Acta 2018, 1023, 53–63. [Google Scholar] [CrossRef]
- Westenbrink, E.; Arasaradnam, R.P.; O’Connell, N.; Bailey, C.; Nwokolo, C.; Bardhan, K.D.; Covington, J.A. Development and application of a new electronic nose instrument for the detection of colorectal cancer. Biosens. Bioelectron. 2015, 67, 733–738. [Google Scholar] [CrossRef]
- Chen, J.L.; Fan, J.; Yan, L.S.; Guo, H.Q.; Xiong, J.J.; Ren, Y.; Hu, J.D. Urine metabolite profiling of human colorectal cancer by capillary electrophoresis mass spectrometry based on MRB. Gastroenterol. Res. Pract. 2012, 2012. [Google Scholar] [CrossRef]
- Yue, H.; Wang, Y.; Zhang, Y.; Ren, H.; Wu, J.; Ma, L.; Liu, S. A metabonomics study of colorectal cancer by RRLC-QTOF/MS. J. Liq. Chromatogr. Relat. Technol. 2013, 36, 428–438. [Google Scholar] [CrossRef]
- McFarlane, M.; Millard, A.; Hall, H.; Savage, R.; Constantinidou, C.; Arasaradnam, R.; Nwokolo, C. Urinary volatile organic compounds and faecal microbiome profiles in colorectal cancer. Color. Dis. 2019, 21, 1259–1269. [Google Scholar] [CrossRef]
- Young, G.P.; Rabeneck, L.; Winawer, S.J. The Global Paradigm Shift in Screening for Colorectal Cancer. Gastroenterology 2019, 156, 843–851.e2. [Google Scholar] [CrossRef]
- Cheng, Y.; Xie, G.; Chen, T.; Qiu, Y.; Zou, X.; Zheng, M.; Tan, B.; Feng, B.; Dong, T.; He, P.; et al. Distinct urinary metabolic profile of human colorectal cancer. J. Proteome Res. 2012, 11, 1354–1363. [Google Scholar] [CrossRef]
- Kim, E.R.; Kwon, H.N.; Nam, H.; Kim, J.J.; Park, S.; Kim, Y.H. Urine-NMR metabolomics for screening of advanced colorectal adenoma and early stage colorectal cancer. Sci. Rep. 2019, 9, 4786. [Google Scholar] [CrossRef]
- Wang, Z.; Lin, Y.; Liang, J.; Huang, Y.; Ma, C.; Liu, X.; Yang, J. NMR-based metabolomic techniques identify potential urinary biomarkers for early colorectal cancer detection. Oncotarget 2017, 8, 105819–105831. [Google Scholar] [CrossRef]
- Hsu, W.Y.; Chen, C.J.; Huang, Y.C.; Tsai, F.J.; Jeng, L.B.; Lai, C.C. Urinary nucleosides as biomarkers of breast, colon, lung, and gastric cancer in Taiwanese. PLoS ONE 2013, 8, e81701. [Google Scholar] [CrossRef]
- Jiang, H.P.; Qi, C.B.; Chu, J.M.; Yuan, B.F.; Feng, Y.Q. Profiling of cis-diol-containing nucleosides and ribosylated metabolites by boronate-affinity organic-silica hybrid monolithic capillary liquid chromatography/mass spectrometry. Sci. Rep. 2015, 5, 7785. [Google Scholar] [CrossRef]
- Rozhentsov, A.A.; Koptina, A.V.; Mitrakov, N.N.; Sharipova, T.; Tsapaev, I.; Ryzhkov, V.L.; Lychagin, K.A.; Furina, R.R.; Mitrakova, N.N. A new method to diagnose cancer based on image analysis of mass chromatograms of volatile organic compounds in urine. Sovrem. Tehnol. Med. 2014, 6, 151–157. [Google Scholar]
- Silva, C.L.; Passos, M.; Câmara, J.S. Investigation of urinary volatile organic metabolites as potential cancer biomarkers by solid-phase microextraction in combination with gas chromatography-mass spectrometry. Br. J. Cancer 2011, 105, 1894–1904. [Google Scholar] [CrossRef]
- Feng, B.; Zheng, M.H.; Zheng, Y.F.; Lu, A.G.; Li, J.W.; Wang, M.L.; Ma, J.J.; Xu, G.W.; Liu, B.Y.; Zhu, Z.G. Normal and modified urinary nucleosides represent novel biomarkers for colorectal cancer diagnosis and surgery monitoring. J. Gastroenterol. Hepatol. 2005, 20, 1913–1919. [Google Scholar] [CrossRef]
- Qiu, Y.; Cai, G.; Su, M.; Chen, T.; Liu, Y.; Xu, Y.; Ni, Y.; Zhao, A.; Cai, S.; Xu, L.X.; et al. Urinary metabonomic study on colorectal cancer. J. Proteome Res. 2010, 9, 1627–1634. [Google Scholar] [CrossRef] [PubMed]
- Liesenfeld, D.B.; Habermann, N.; Toth, R.; Owen, R.W.; Frei, E.; Böhm, J.; Schrotz-King, P.; Klika, K.D.; Ulrich, C.M. Changes in urinary metabolic profiles of colorectal cancer patients enrolled in a prospective cohort study (ColoCare). Metabolomics 2015, 11, 998–1012. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.L.; Qin, H.L.; Liu, W.J.; Peng, J.Y.; Huang, L.; Zhao, X.P.; Cheng, Y.Y. Ultra-high performance liquid chromatography-mass spectrometry for the metabolomic analysis of urine in colorectal cancer. Dig. Dis. Sci. 2009, 54, 2655–2662. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Feng, B.; Li, X.; Yin, P.; Gao, P.; Zhao, X.; Lu, X.; Zheng, M.; Xu, G. Urinary metabolic profiling of colorectal carcinoma based on online affinity solid phase extraction-high performance liquid chromatography and ultra performance liquid chromatography-mass spectrometry. Mol. Biosyst. 2010, 6, 1947–1955. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.F.; Yang, J.; Zhao, X.J.; Feng, B.; Kong, H.W.; Chen, Y.J.; Lv, S.; Zheng, M.H.; Xu, G.W. Urinary nucleosides as biological markers for patients with colorectal cancer. World J. Gastroenterol. 2005, 11, 3871–3876. [Google Scholar] [CrossRef] [PubMed]
- Mozdiak, E.; Wicaksono, A.N.; Covington, J.A.; Arasaradnam, R.P. Colorectal cancer and adenoma screening using urinary volatile organic compound (VOC) detection: Early results from a single-centre bowel screening population (UK BCSP). Tech. Coloproctol. 2019, 23, 343–351. [Google Scholar] [CrossRef] [PubMed]
- Widlak, M.M.; Neal, M.; Daulton, E.; Thomas, C.L.; Tomkins, C.; Singh, B.; Harmston, C.; Wicaksono, A.; Evans, C.; Smith, S.; et al. Risk stratification of symptomatic patients suspected of colorectal cancer using faecal and urinary markers. Color. Dis. 2018, 20, O335–O342. [Google Scholar] [CrossRef]
- Wang, H.; Tso, V.; Wong, C.; Sadowski, D.; Fedorak, R.N. Development and validation of a highly sensitive urine-based test to identify patients with colonic adenomatous polyps. Clin. Transl. Gastroenterol. 2014, 5, e54. [Google Scholar] [CrossRef]
- Wohlgemuth, G.; Haldiya, P.K.; Willighagen, E.; Kind, T.; Fiehn, O. The chemical translation service-a web-based tool to improve standardization of metabolomic reports. Bioinformatics 2010, 26, 2647–2648. [Google Scholar] [CrossRef]
- Moriya, Y.; Itoh, M.; Okuda, S.; Yoshizawa, A.C.; Kanehisa, M. KAAS: An automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 2007, 35, 182–185. [Google Scholar] [CrossRef]
- Topaz, N.; Mojib, N.; Chande, A.T.; Kubanek, J.; Jordan, I.K. RampDB: A web application and database for the exploration and prediction of receptor activity modifying protein interactions. Database 2017, 2017, bax067. [Google Scholar] [CrossRef]
- Wishart, D.S.; Bartok, B.; Oler, E.; Liang, K.Y.H.; Budinski, Z.; Berjanskii, M.; Guo, A.; Cao, X.; Wilson, M. MarkerDB: An online database of molecular biomarkers. Nucleic Acids Res. 2020, 49, 1259–1267. [Google Scholar] [CrossRef]
- López-Ibáñez, J.; Pazos, F.; Chagoyen, M. MBROLE 2.0-functional enrichment of chemical compounds. Nucleic Acids Res. 2016, 44, W201–W204. [Google Scholar] [CrossRef]
- Icard, P.; Poulain, L.; Lincet, H. Understanding the central role of citrate in the metabolism of cancer cells. Biochim. Biophys. Acta Rev. Cancer 2012, 1825, 111–116. [Google Scholar] [CrossRef]
- La Vecchia, S.; Sebastián, C. Metabolic pathways regulating colorectal cancer initiation and progression. Semin. Cell Dev. Biol. 2020, 98, 63–70. [Google Scholar] [CrossRef]
- Hiramatsu, K.; Takahashi, K.; Yamaguchi, T.; Matsumoto, H.; Miyamoto, H.; Tanaka, S.; Tanaka, C.; Tamamori, Y.; Imajo, M.; Kawaguchi, M.; et al. N 1,N 12-diacetylspermine as a sensitive and specific novel marker for early- and late-stage colorectal and breast cancers. Clin. Cancer Res. 2005, 11, 2986–2990. [Google Scholar] [CrossRef]
- Nakayama, Y.; Torigoe, T.; Minagawa, N.; Yamaguchi, K. The clinical usefulness of urinary N 1,N 12-diacetylspermine(DiAcSpm) levels as a tumor marker in patients withcolorectal cancer. Oncol. Lett. 2012, 3, 970–974. [Google Scholar] [CrossRef]
- Umemori, Y.; Ohe, Y.; Kuribayashi, K.; Tsuji, N.; Nishidate, T.; Kameshima, H.; Hirata, K.; Watanabe, N. Evaluating the utility of N1,N12-diacetylspermine and N1,N8-diacetylspermidine in urine as tumor markers for breast and colorectal cancers. Clin. Chim. Acta 2010, 411, 1894–1899. [Google Scholar] [CrossRef]
- Pang, Y.; Kartsonaki, C.; Guo, Y.; Chen, Y.; Yang, L.; Bian, Z.; Bragg, F.; Millwood, I.Y.; Shen, L.; Zhou, S.; et al. Diabetes, plasma glucose and incidence of colorectal cancer in Chinese adults: A prospective study of 0.5 million people. J. Epidemiol. Community Health 2018, 72, 919–925. [Google Scholar] [CrossRef]
- Vulcan, A.; Manjer, J.; Ohlsson, B. High blood glucose levels are associated with higher risk of colon cancer in men: A cohort study. BMC Cancer 2017, 17. [Google Scholar] [CrossRef] [PubMed]
- Venkateswaran, N.; Conacci-Sorrell, M. Kynurenine: An oncometabolite in colon cancer. Cell Stress 2020, 4, 24–26. [Google Scholar] [CrossRef] [PubMed]
- Barberini, L.; Restivo, A.; Noto, A.; Deidda, S.; Fattuoni, C.; Fanos, V.; Saba, L.; Zorcolo, L.; Mussap, M. A gas chromatography-mass spectrometry (GC-MS) metabolomic approach in human colorectal cancer (CRC): The emerging role of monosaccharides and amino acids. Ann. Transl. Med. 2019, 7. [Google Scholar] [CrossRef] [PubMed]
- Ge, S.; Zhang, Q.; Tian, Y.; Hao, L.; Duan, J.; Zhang, B. Cell metabolic profiling of colorectal cancer via 1H NMR. Clin. Chim. Acta 2020, 510, 291–297. [Google Scholar] [CrossRef]
- Ni, Y.; Xie, G.; Jia, W. Metabonomics of human colorectal cancer: New approaches for early diagnosis and biomarker discovery. J. Proteome Res. 2014, 13, 3857–3870. [Google Scholar] [CrossRef]
- Brown, D.G.; Rao, S.; Weir, T.L.; O’Malia, J.; Bazan, M.; Brown, R.J.; Ryan, E.P. Metabolomics and metabolic pathway networks from human colorectal cancers, adjacent mucosa, and stool. Cancer Metab. 2016, 4, 11. [Google Scholar] [CrossRef]
- Loke, M.F.; Chua, E.G.; Gan, H.M.; Thulasi, K.; Wanyiri, J.W.; Thevambiga, I.; Goh, K.L.; Wong, W.F.; Vadivelu, J. Metabolomics and 16S rRNA sequencing of human colorectal cancers and adjacent mucosa. PLoS ONE 2018, 13, e0208584. [Google Scholar] [CrossRef]
- Corral, M.; Wallace, H.M. Upregulation of Polyamine Transport in Human Colorectal Cancer Cells. Biomolecules 2020, 10, 499. [Google Scholar] [CrossRef]
- Alam, M.N.; Almoyad, M.; Huq, F. Polyphenols in Colorectal Cancer: Current State of Knowledge including Clinical Trials and Molecular Mechanism of Action. BioMed Res. Int. 2018, 2018, 4154185. [Google Scholar] [CrossRef]
- Bashir, A.I.J.; Kankipati, C.S.; Jones, S.; Newman, R.M.; Safrany, S.T.; Perry, C.J.; Nicholl, I.D. A novel mechanism for the anticancer activity of aspirin and salicylates. Int. J. Oncol. 2019, 54, 1256–1270. [Google Scholar] [CrossRef]
- Dachineni, R.; Kumar, D.R.; Callegari, E.; Kesharwani, S.S.; Sankaranarayanan, R.; Seefeldt, T.; Tummala, H.; Bhat, G.J. Salicylic acid metabolites and derivatives inhibit CDK activity: Novel insights into aspirin’s chemopreventive effects against colorectal cancer. Int. J. Oncol. 2017, 51, 1661–1673. [Google Scholar] [CrossRef]
- Barupal, D.K.; Haldiya, P.K.; Wohlgemuth, G.; Kind, T.; Kothari, S.L.; Pinkerton, K.E.; Fiehn, O. MetaMapp: Mapping and visualizing metabolomic data by integrating information from biochemical pathways and chemical and mass spectral similarity. BMC Bioinform. 2012, 13. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- Goveia, J.; Pircher, A.; Conradi, L.-C.; Kalucka, J.; Lagani, V.; Dewerchin, M.; Eelen, G.; DeBerardinis, R.J.; Wilson, I.D.; Carmeliet, P. Meta-analysis of clinical metabolic profiling studies in cancer: Challenges and opportunities. EMBO Mol. Med. 2016, 8, 1134–1142. [Google Scholar] [CrossRef]
- Krupp, D.; Doberstein, N.; Shi, L.; Remer, T. Hippuric Acid in 24-Hour Urine Collections Is a Potential Biomarker for Fruit and Vegetable Consumption in Healthy Children and Adolescents. J. Nutr. Nutr. Epidemiol. J. Nutr 2012, 142, 1314–1320. [Google Scholar] [CrossRef]
- Dinges, S.S.; Hohm, A.; Vandergrift, L.A.; Nowak, J.; Habbel, P.; Kaltashov, I.A.; Cheng, L.L. Cancer metabolomic markers in urine: Evidence, techniques and recommendations. Nat. Rev. Urol. 2019, 16, 339–362. [Google Scholar] [CrossRef]
- Brial, F.; Chilloux, J.; Nielsen, T.; Vieira-Silva, S.; Falony, G.; Andrikopoulos, P.; Olanipekun, M.; Hoyles, L.; Djouadi, F.; Neves, A.L.; et al. Human and preclinical studies of the host–gut microbiome co-metabolite hippurate as a marker and mediator of metabolic health. Gut 2021, 1–10. [Google Scholar] [CrossRef]
- Iwasaki, H.; Shimura, T.; Kataoka, H. Current status of urinary diagnostic biomarkers for colorectal cancer. Clin. Chim. Acta 2019, 498, 76–83. [Google Scholar] [CrossRef]
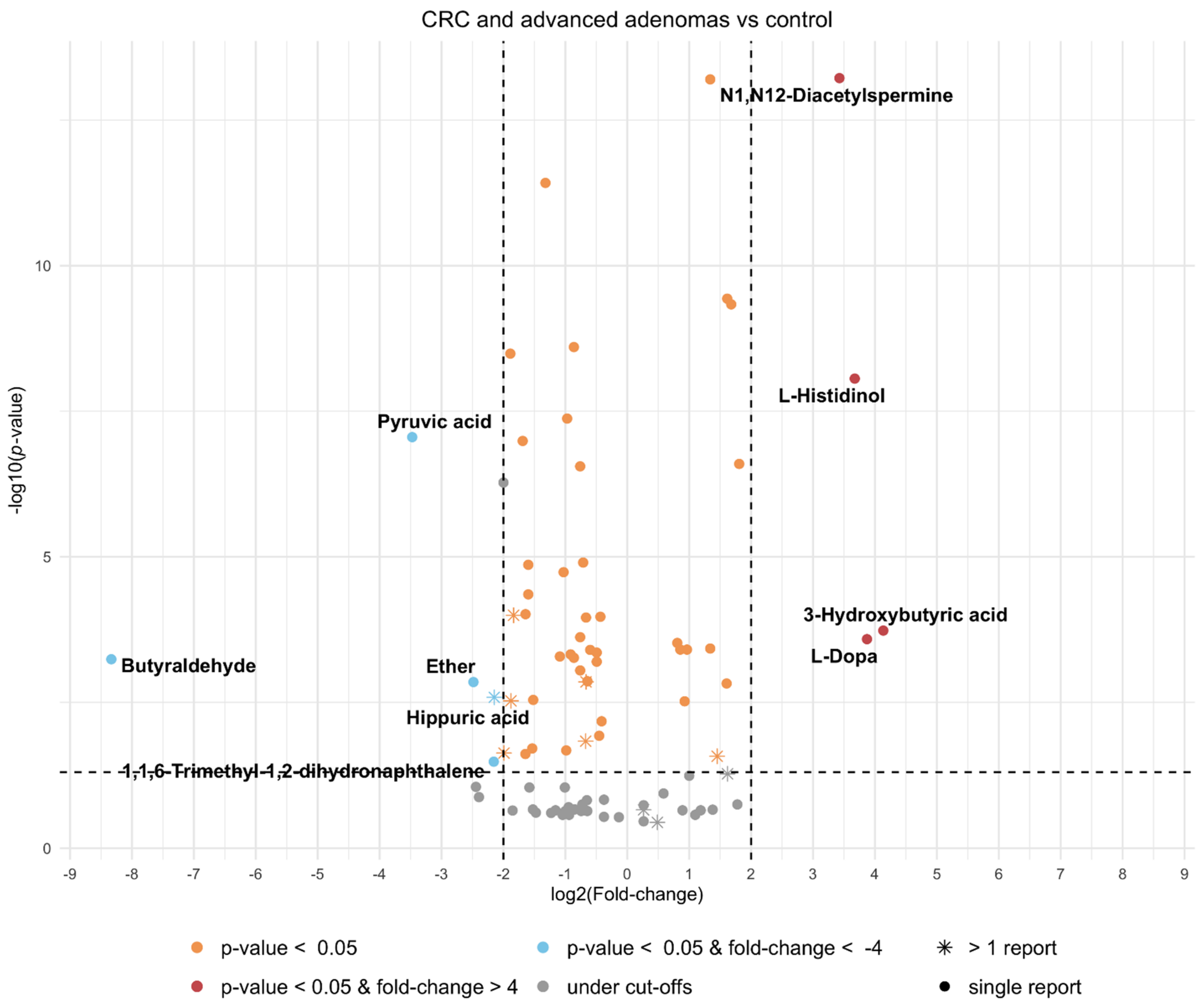
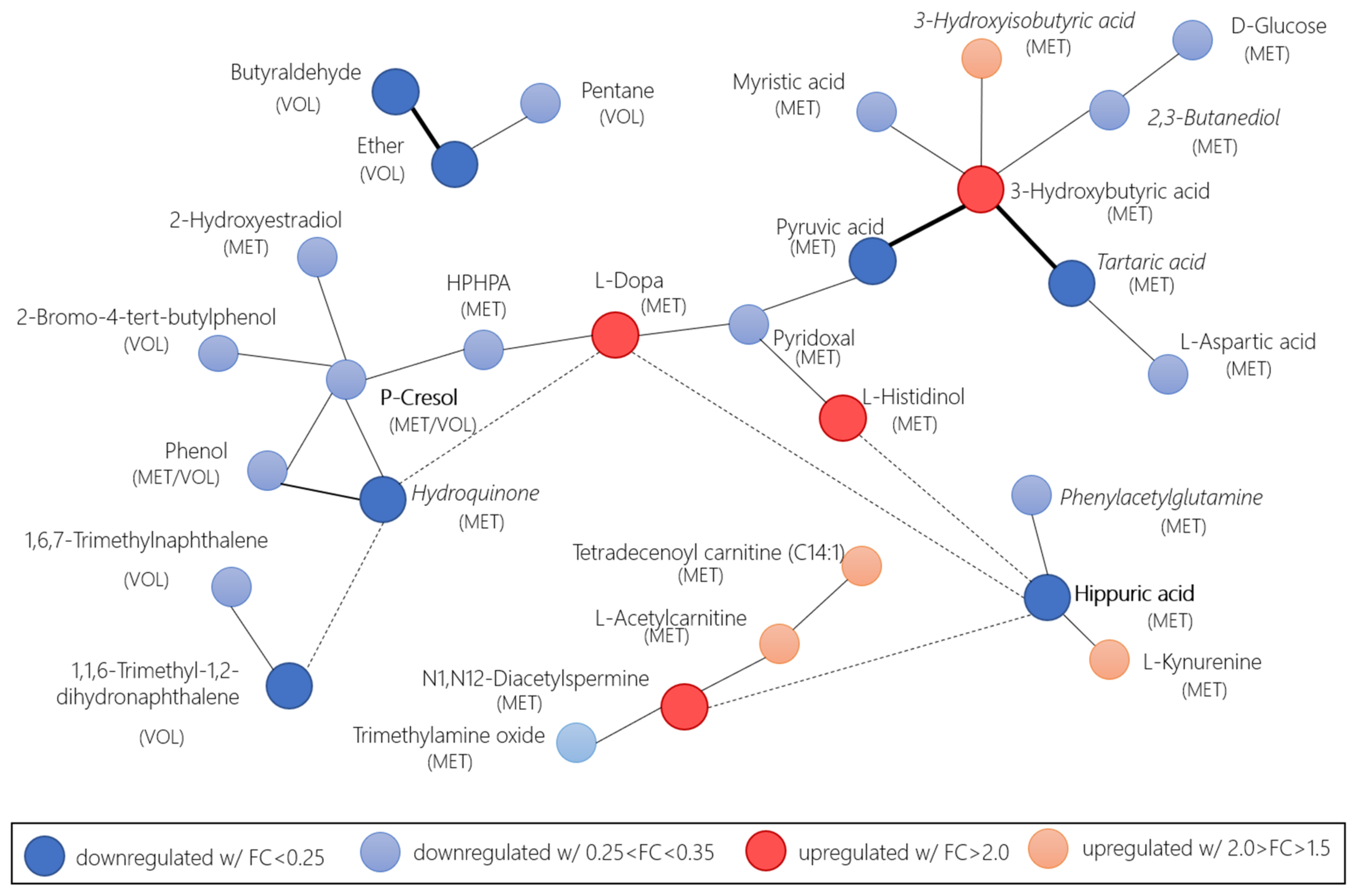
| Compound Name | Combined and Weighted p-Value | Combined and Weighted Fold-Change | N Total | HMDB ID | Healthy Normal Urine Concentration (Adult > 18 y) (µmol/mmol Creatinine) | CRC Projected Urine Concentration (Adult > 18 y) (µmol/mmol Creatinine) |
|---|---|---|---|---|---|---|
| 3-Hidroxybutyric acid | 1.85 × 10−4 | 17.56 | 342 | HMDB0000357 | 1.4–2.7 | 5.8–11.2 |
| L-Dopa | 2.60 × 10−4 | 14.63 | 342 | HMDB0000181 | 0.01–0.04 | 0.04–0.15 |
| L-Histidinol | 8.71 × 10−9 | 12.76 | 204 | HMDB0003431 | NQ | - |
| N1,N12-Diacetylspermine | 6.00 × 10−14 | 10.75 | 342 | HMDB0002172 | 0–0.0260 | 0–0.280 |
| 1,1,6-Trimethyl-1,2-dihydronaphthalene | 3.31 × 10−2 | 0.22 | 60 | HMDB0040284 | NQ | - |
| Hippuric acid | 2.59 × 10−3 | 0.23 | 546 | HMDB0000714 | 28–610 | 6–140 |
| Ether | 1.42 × 10−3 | 0.18 | 60 | - | NQ | - |
| Pyruvic acid | 8.82 × 10−8 | 0.09 | 204 | HMDB0000243 | 1–3.7 | 0.09–0.33 |
| Butyraldehyde | 5.76 × 10−4 | 0.003 | 60 | HMDB0003543 | NQ | - |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mallafré-Muro, C.; Llambrich, M.; Cumeras, R.; Pardo, A.; Brezmes, J.; Marco, S.; Gumà, J. Comprehensive Volatilome and Metabolome Signatures of Colorectal Cancer in Urine: A Systematic Review and Meta-Analysis. Cancers 2021, 13, 2534. https://doi.org/10.3390/cancers13112534
Mallafré-Muro C, Llambrich M, Cumeras R, Pardo A, Brezmes J, Marco S, Gumà J. Comprehensive Volatilome and Metabolome Signatures of Colorectal Cancer in Urine: A Systematic Review and Meta-Analysis. Cancers. 2021; 13(11):2534. https://doi.org/10.3390/cancers13112534
Chicago/Turabian StyleMallafré-Muro, Celia, Maria Llambrich, Raquel Cumeras, Antonio Pardo, Jesús Brezmes, Santiago Marco, and Josep Gumà. 2021. "Comprehensive Volatilome and Metabolome Signatures of Colorectal Cancer in Urine: A Systematic Review and Meta-Analysis" Cancers 13, no. 11: 2534. https://doi.org/10.3390/cancers13112534
APA StyleMallafré-Muro, C., Llambrich, M., Cumeras, R., Pardo, A., Brezmes, J., Marco, S., & Gumà, J. (2021). Comprehensive Volatilome and Metabolome Signatures of Colorectal Cancer in Urine: A Systematic Review and Meta-Analysis. Cancers, 13(11), 2534. https://doi.org/10.3390/cancers13112534










