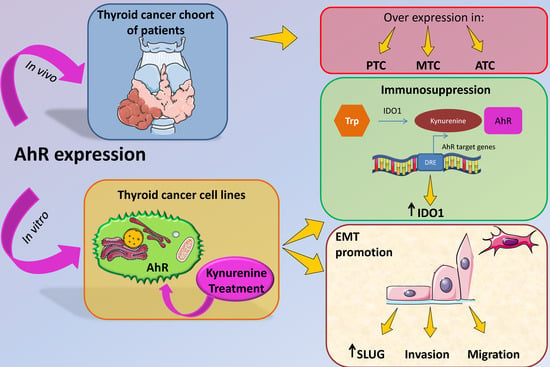
The Aryl Hydrocarbon Receptor Is Expressed in Thyroid Carcinoma and Appears to Mediate Epithelial-Mesenchymal-Transition
Abstract
1. Introduction
2. Results
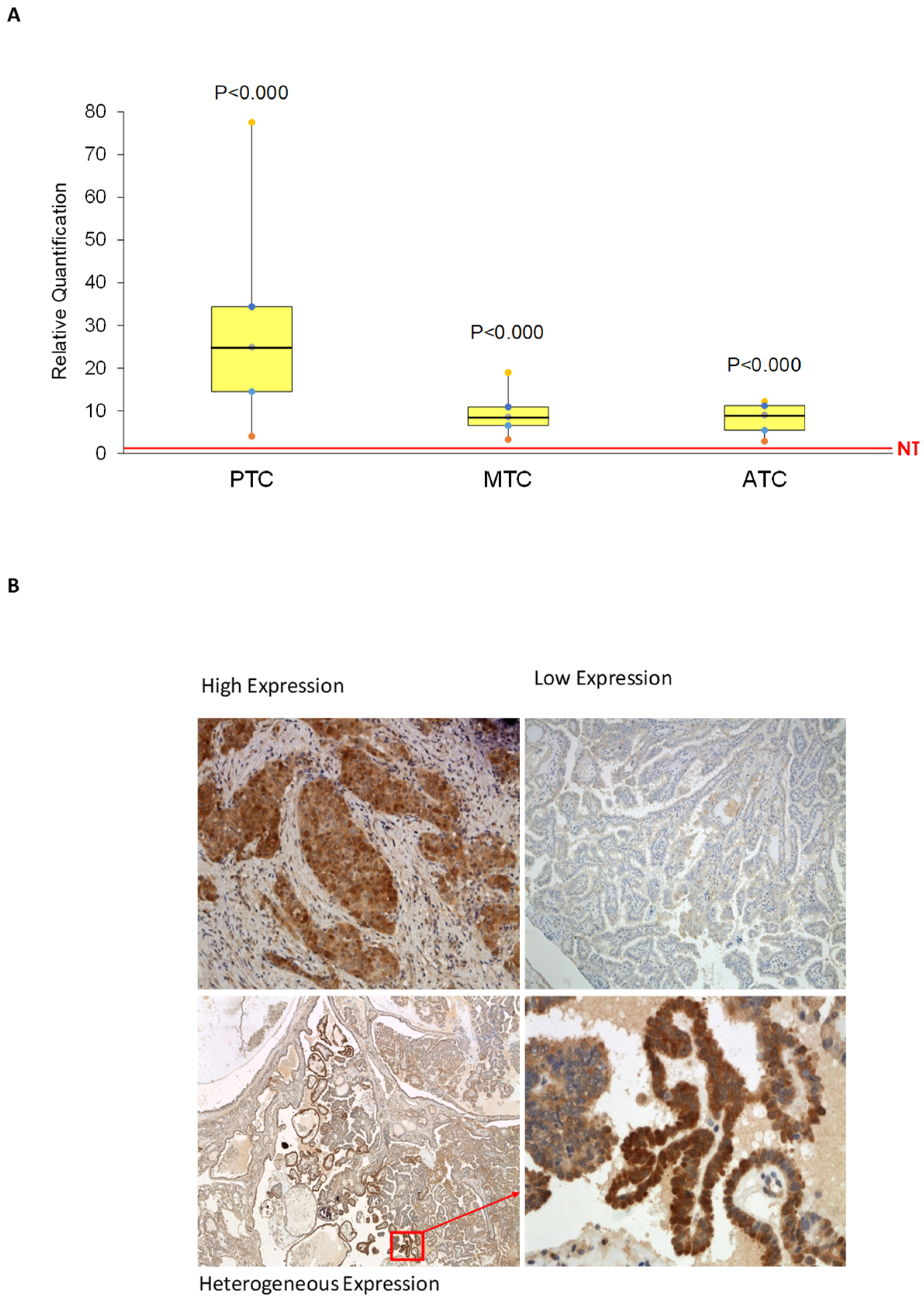
2.1. AhR Is Expressed and Functionally Activated in Thyroid Cancer Samples
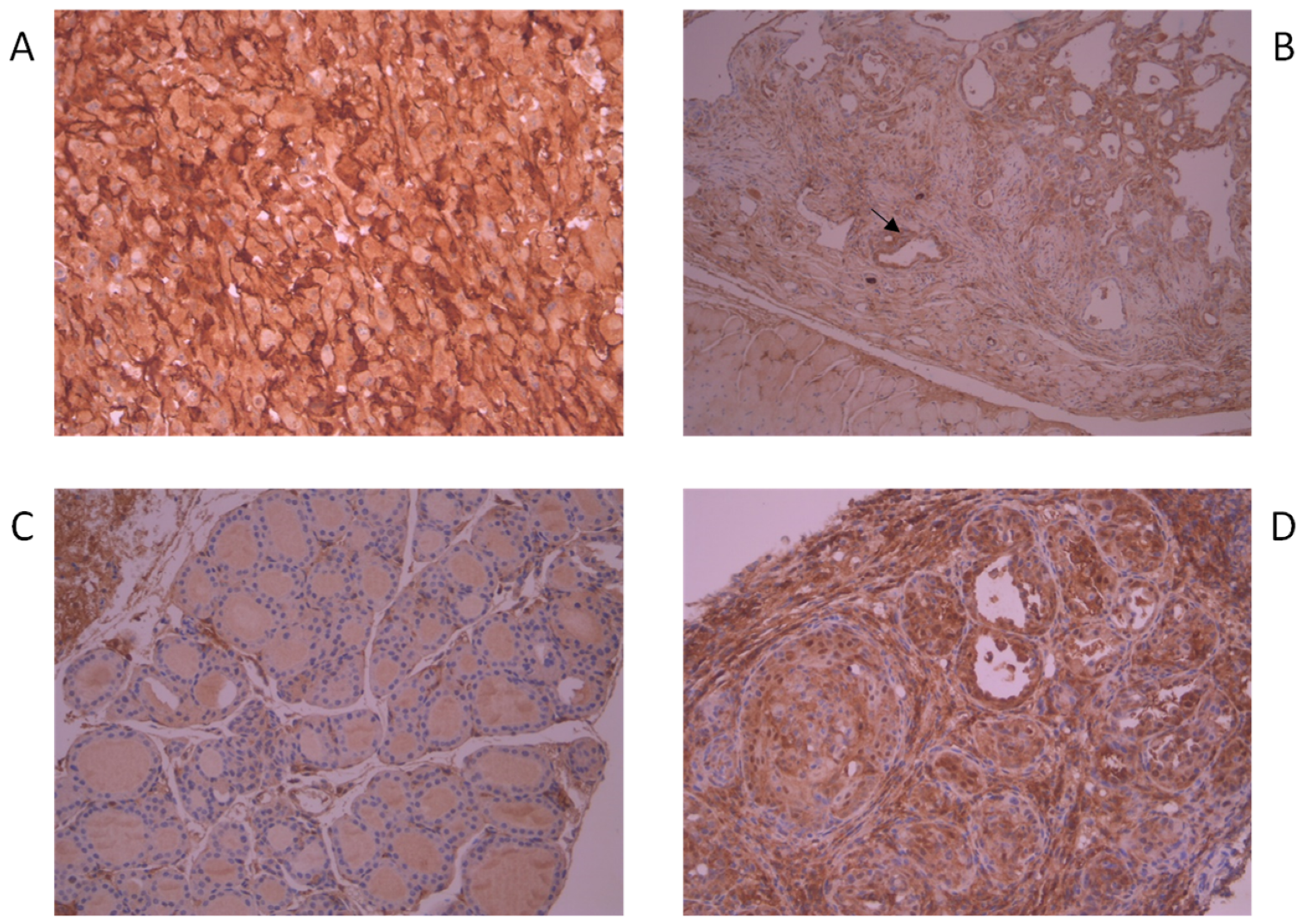
2.2. AhR Is Overexpressed in BRAFV600E-Harboring Murine Thyroid Cancer Tissue
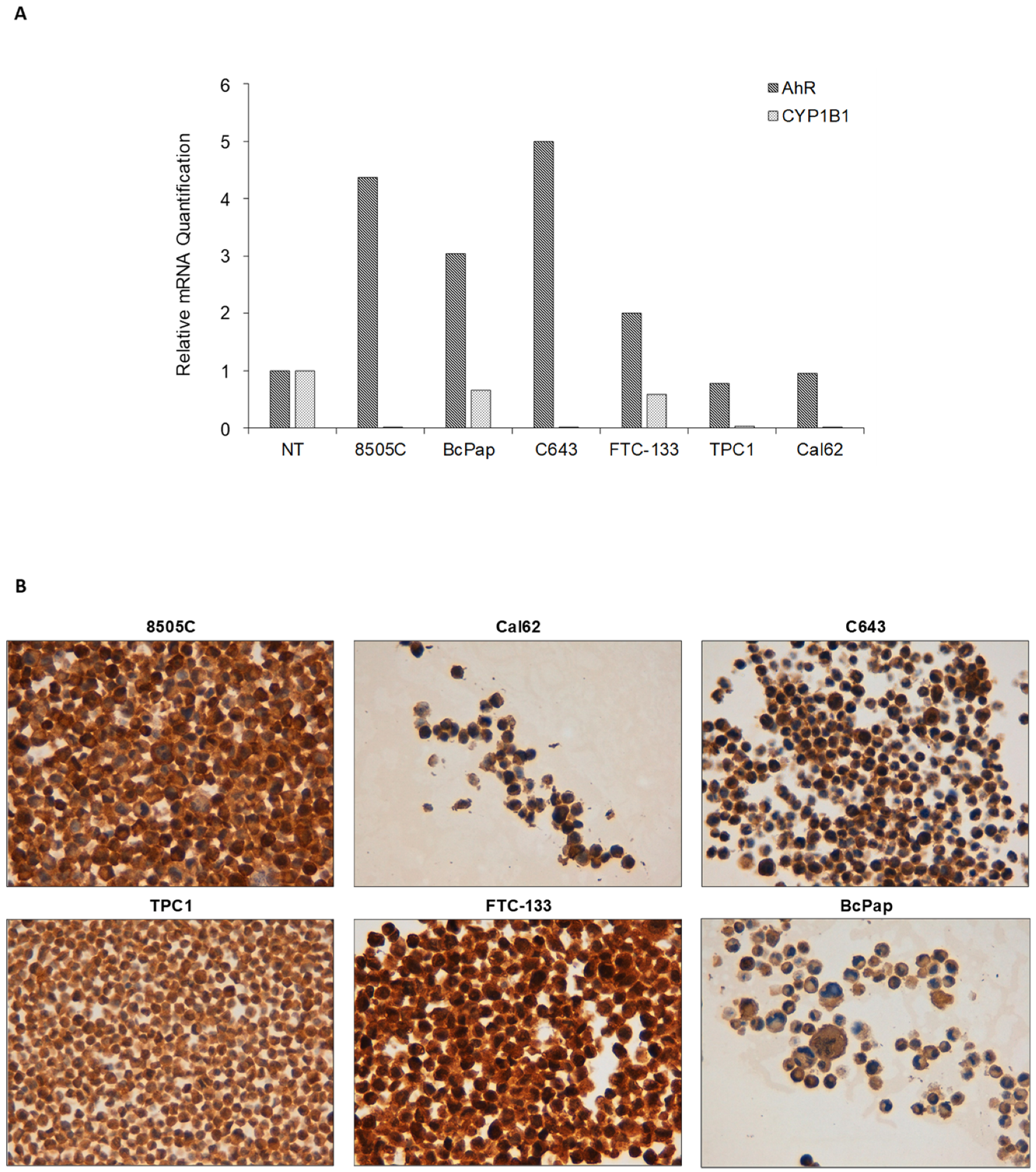
2.3. AhR Is Expressed and Functionally Activated in FTC-133 and Bcpap Cell Lines
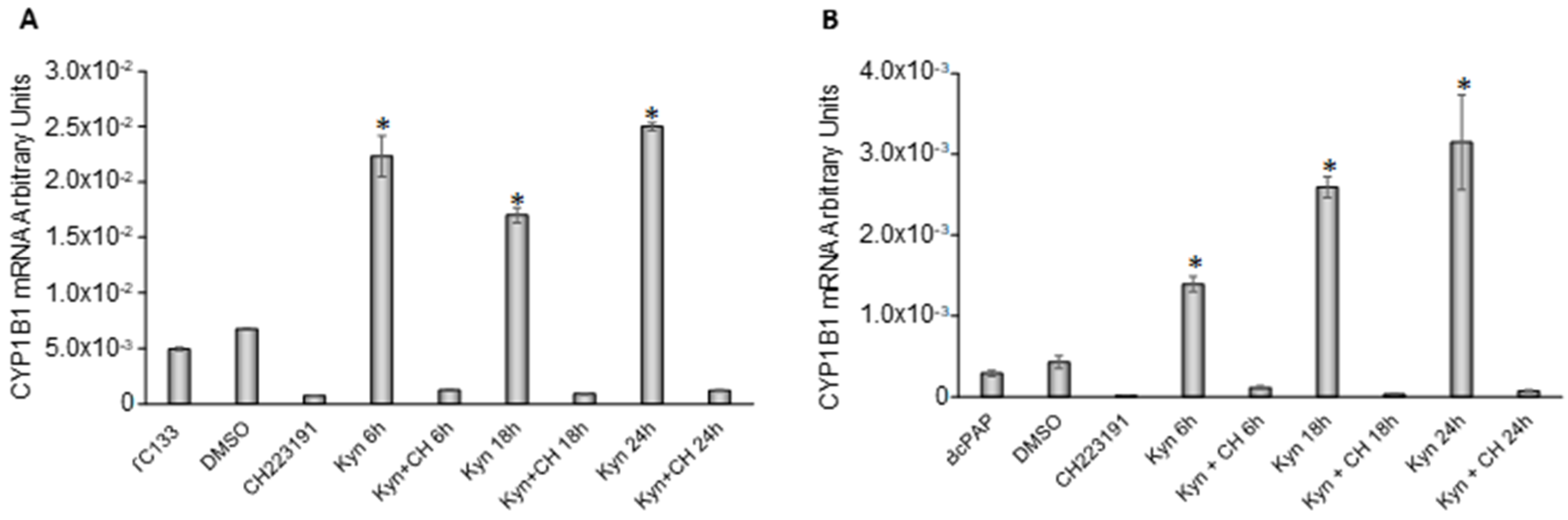
2.4. Kynurenine Activates AhR in Thyroid Cancer Cell Lines
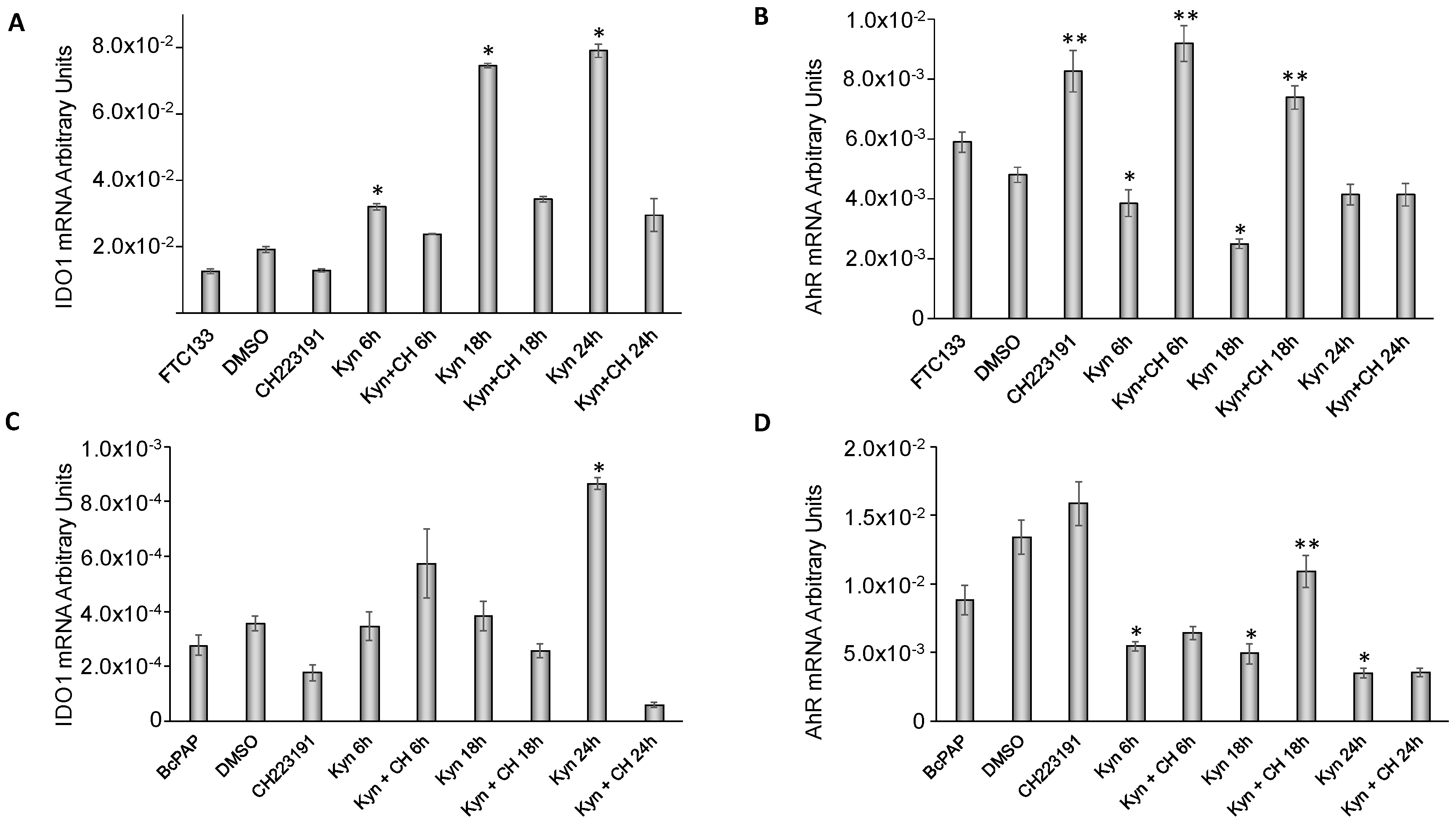
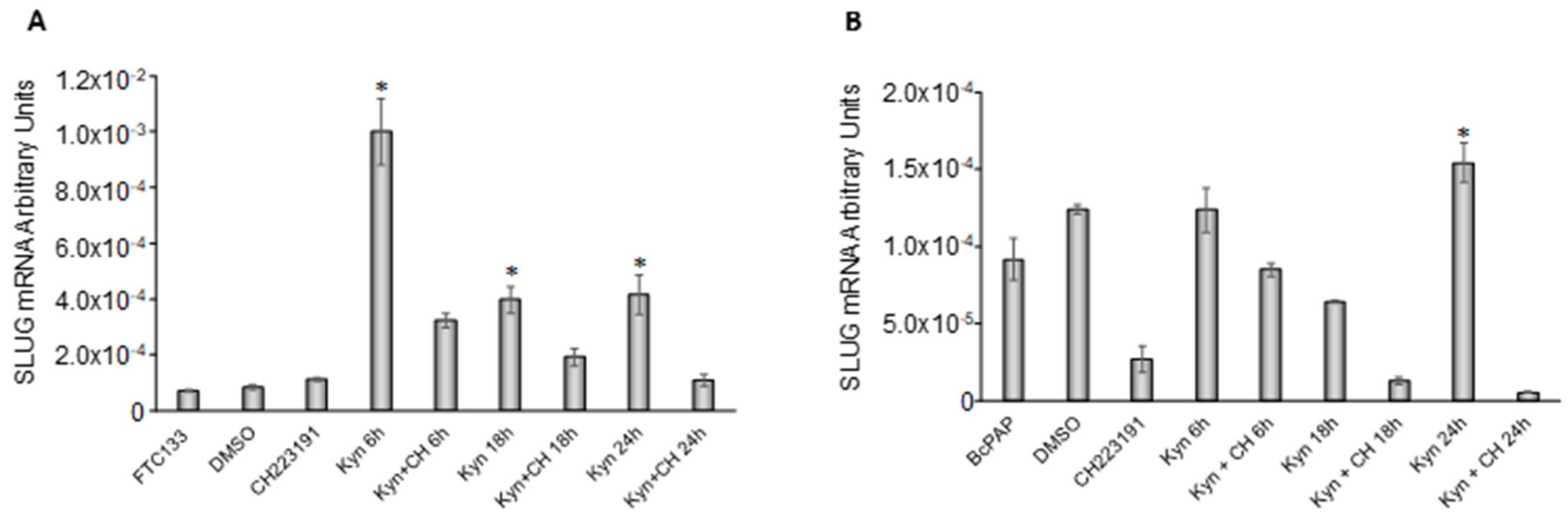
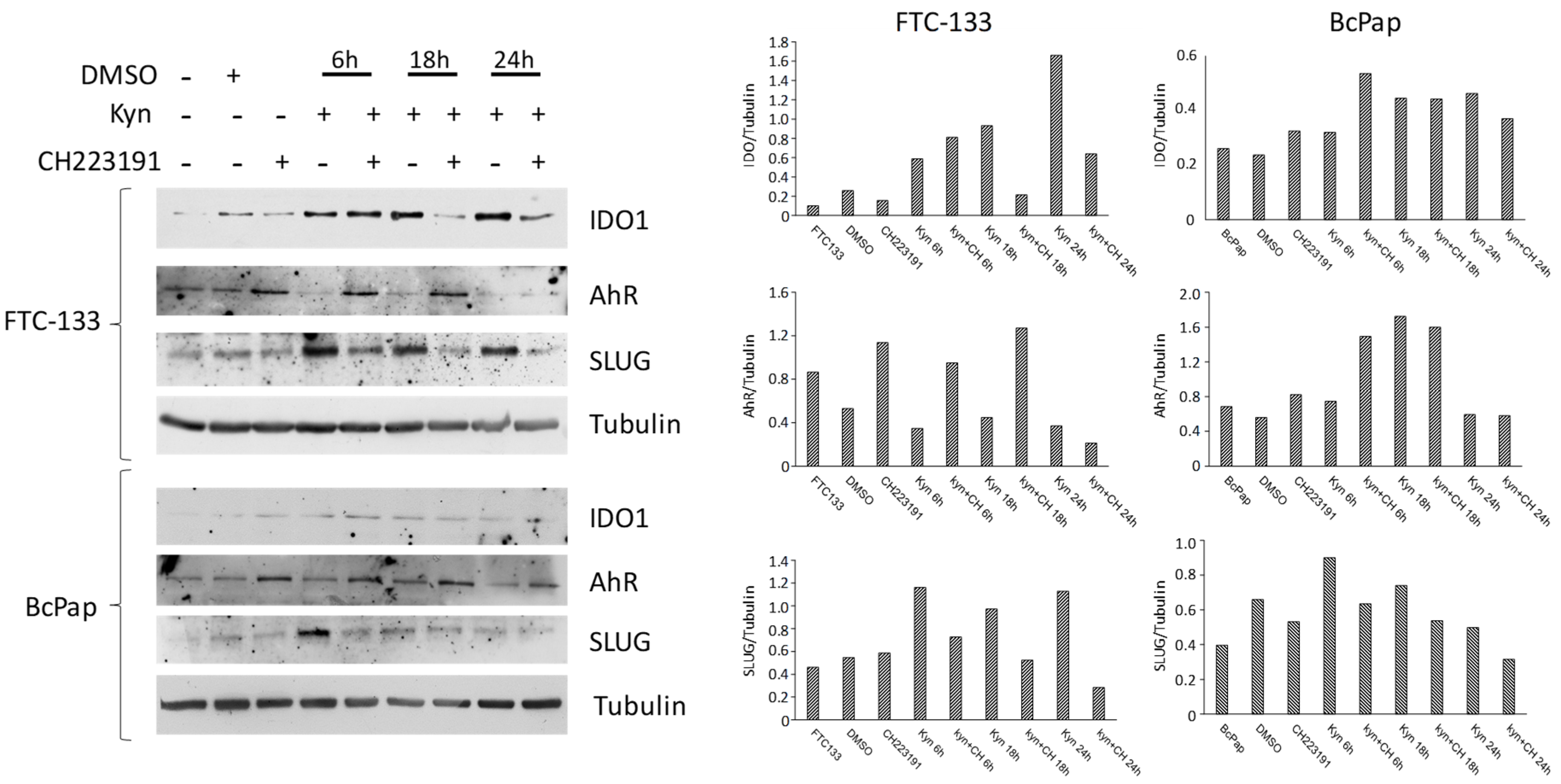
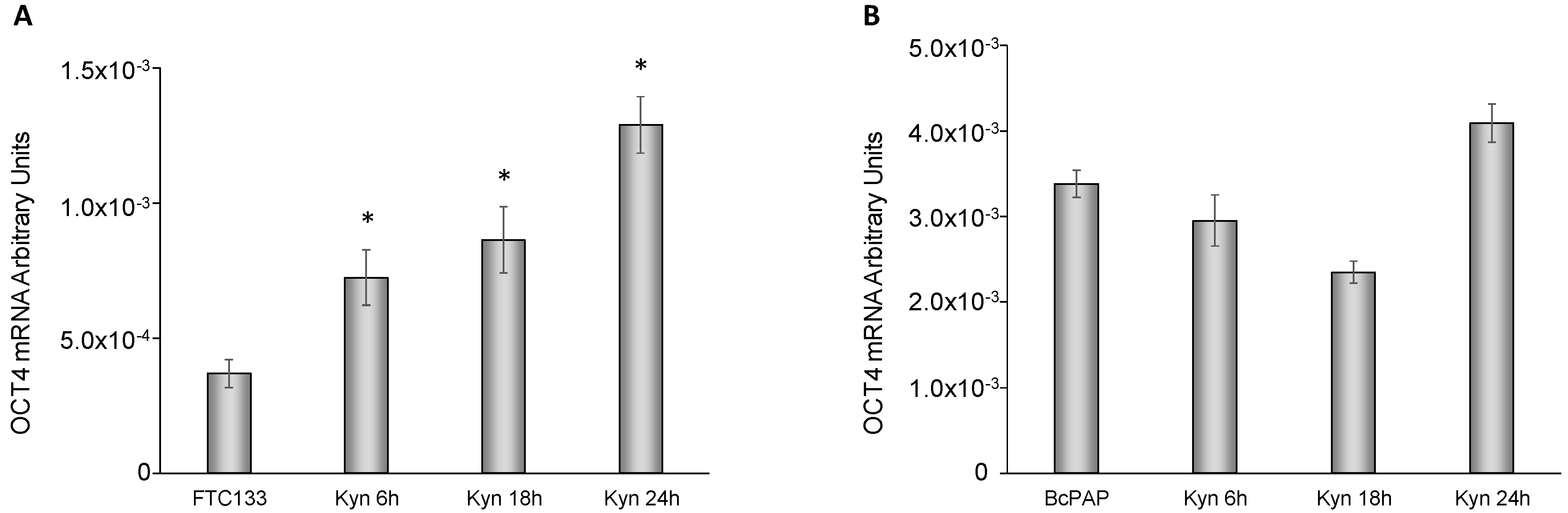
2.5. Kynurenine Induces IDO1 and SLUG Expression through AhR
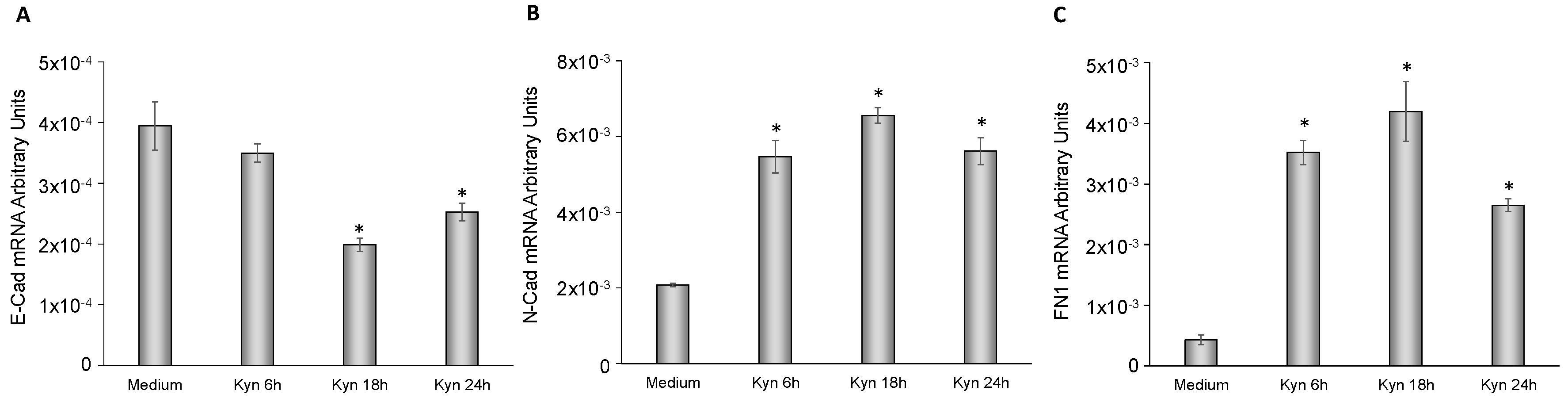
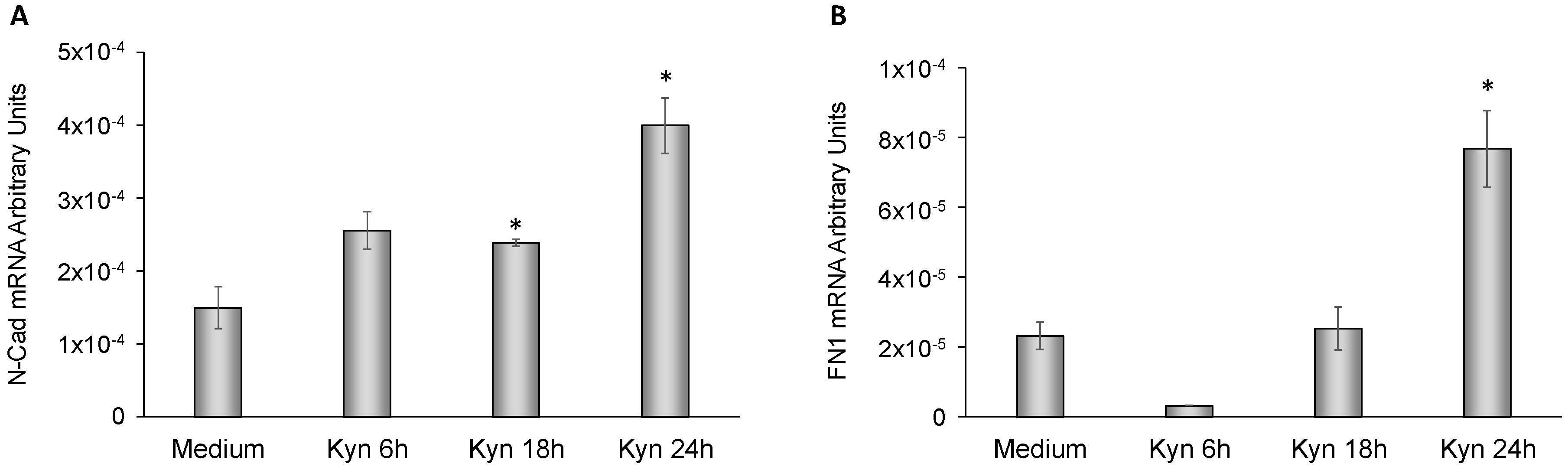
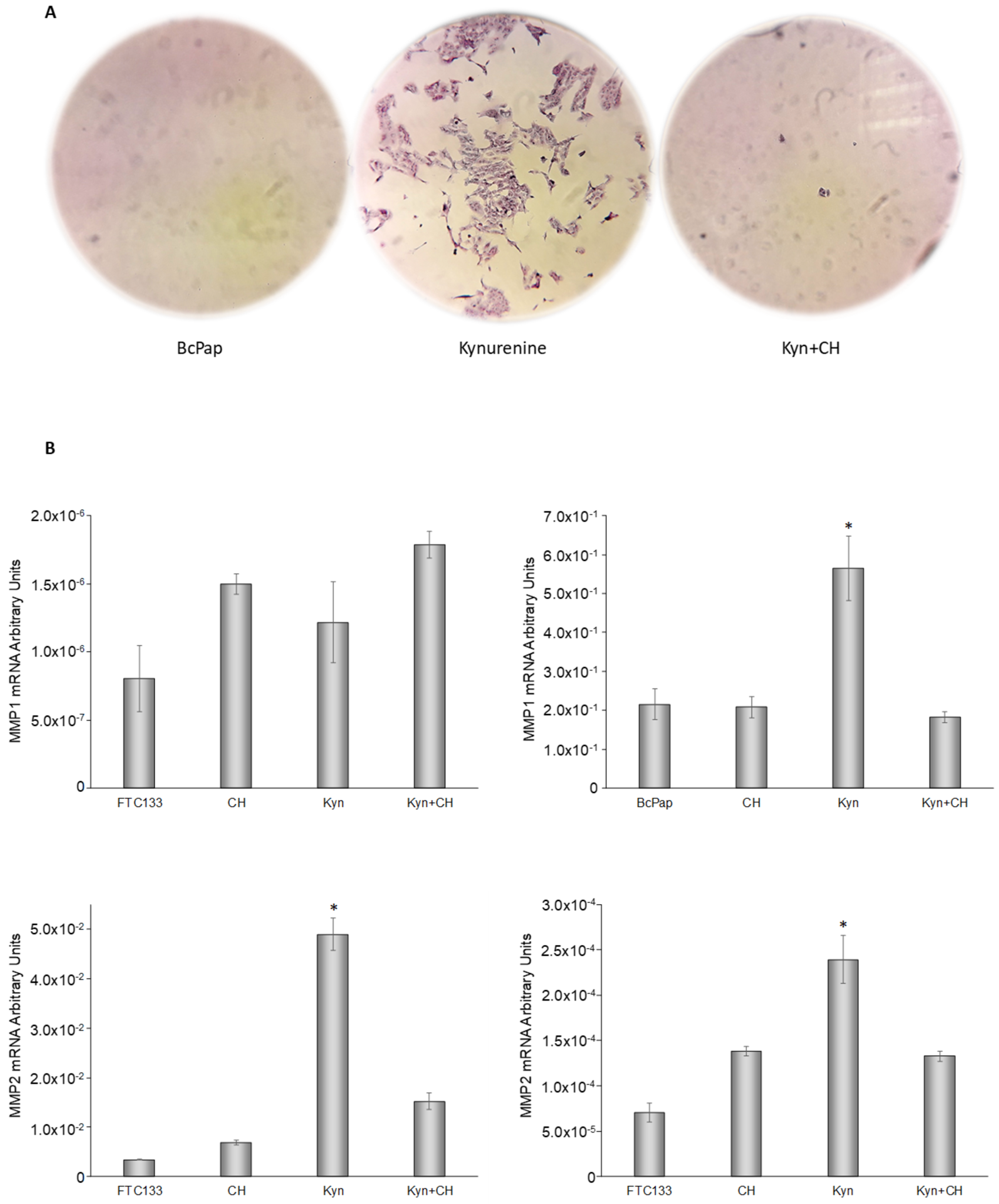
2.6. Kynurenine Mediates EMT
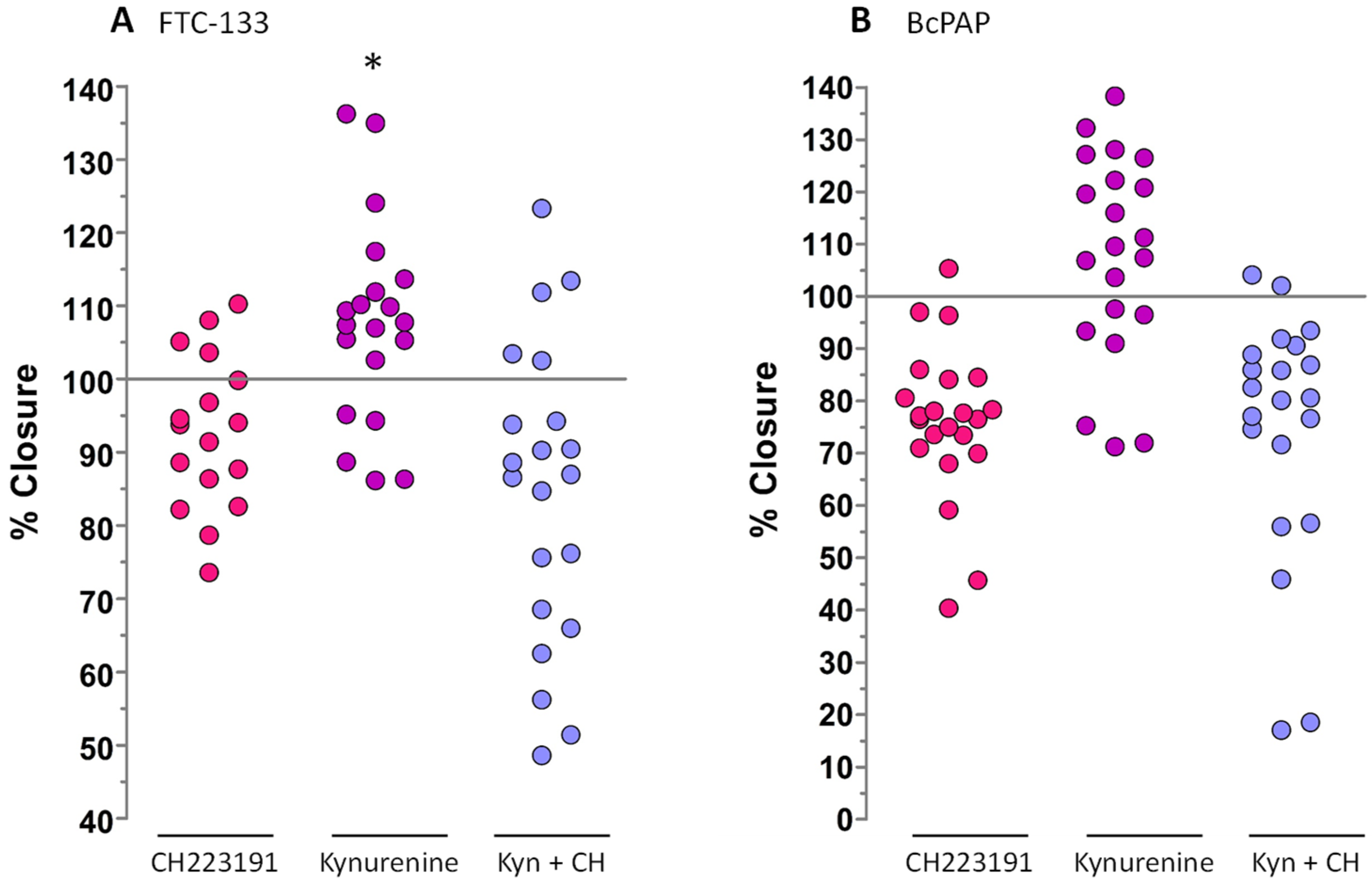
2.7. Cell Migration Assay
3. Discussion
4. Materials and Methods
4.1. Tissue Samples and Patients
4.2. Transgenic Mice Tissue Samples
4.3. Cell Cultures
4.4. Quantitative PCR
4.5. Immunohistochemistry
4.6. Immunoblotting Experiments
4.7. Evaluation of Cellular Migration and Invasiveness
4.8. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Knutson, J.C.; Poland, A. Response of murine epidermis to 2,3,7,8-tetrachlorodibenzo-p-dioxin: Interaction of the ah and hr loci. Cell 1982, 30, 225–234. [Google Scholar] [CrossRef]
- Hahn, M.E.; Karchner, S.I.; Shapiro, M.A.; Perera, S.A. Molecular evolution of two vertebrate aryl hydrocarbon (dioxin) receptors (AHR1 and AHR2) and the PAS family. Proc. Natl. Acad. Sci. USA 1997, 94, 13743–13748. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez-Vázquez, C.; Quintana, F.J. Regulation of the Immune Response by the Aryl Hydrocarbon Receptor. Immunity 2018, 48, 19–33. [Google Scholar] [CrossRef] [PubMed]
- Xue, P.; Fu, J.; Zhou, Y. The Aryl Hydrocarbon Receptor and Tumor Immunity. Front. Immunol. 2018, 9, 286. [Google Scholar] [CrossRef]
- Murray, I.A.; Patterson, A.D.; Perdew, G.H. Aryl hydrocarbon receptor ligands in cancer: Friend and foe. Nat. Rev. Cancer 2014, 14, 801–814. [Google Scholar] [CrossRef]
- Feng, S.; Cao, Z.; Wang, X. Role of aryl hydrocarbon receptor in cancer. Biochim. Biophys. Acta 2013, 1836, 197–210. [Google Scholar] [CrossRef]
- Wang, K.; Li, Y.; Jiang, Y.Z.; Dai, C.F.; Patankar, M.S.; Song, J.S.; Zheng, J. An endogenous aryl hydrocarbon receptor ligand inhibits proliferation and migration of human ovarian cancer cells. Cancer Lett. 2013, 340, 63–71. [Google Scholar] [CrossRef]
- Opitz, C.A.; Litzenburger, U.M.; Sahm, F.; Ott, M.; Tritschler, I.; Trump, S.; Schumacher, T.; Jestaedt, L.; Schrenk, D.; Weller, M.; et al. An endogenous tumour-promoting ligand of the human aryl hydrocarbon receptor. Nature 2011, 478, 197–203. [Google Scholar] [CrossRef]
- Fan, Y.; Boivin, G.P.; Knudsen, E.S.; Nebert, D.W.; Xia, Y.; Puga, A. The aryl hydrocarbon receptor functions as a tumor suppressor of liver carcinogenesis. Cancer Res. 2010, 70, 212–220. [Google Scholar] [CrossRef]
- Hall, J.M.; Barhoover, M.A.; Kazmin, D.; McDonnell, D.P.; Greenlee, W.F.; Thomas, R.S. Activation of the aryl-hydrocarbon receptor inhibits invasive and metastatic features of human breast cancer cells and promotes breast cancer cell differentiation. Mol. Endocrinol. 2010, 24, 359–369. [Google Scholar] [CrossRef]
- Ikuta, T.; Kawajiri, K. Zinc finger transcription factor Slug is a novel target gene of aryl hydrocarbon receptor. Exp. Cell Res. 2006, 312, 3585–3594. [Google Scholar] [CrossRef] [PubMed]
- Bolós, V.; Peinado, H.; Pérez-Moreno, M.A.; Fraga, M.F.; Esteller, M.; Cano, A. The transcription factor Slug represses E-cadherin expression and induces epithelial to mesenchymal transitions: A comparison with Snail and E47 repressors. J. Cell Sci. 2003, 116, 499–511. [Google Scholar] [CrossRef] [PubMed]
- Gao, Z.; Bu, Y.; Liu, X.; Wang, X.; Zhang, G.; Wang, E.; Ding, S.; Liu, Y.; Shi, R.; Li, Q.; et al. TCDD promoted EMT of hFPECs via AhR, which involved the activation of EGFR/ERK signaling. Toxicol. Appl. Pharmacol. 2016, 298, 48–55. [Google Scholar] [CrossRef] [PubMed]
- Moretti, S.; Menicali, E.; Voce, P.; Morelli, S.; Cantarelli, S.; Sponziello, M.; Colella, R.; Fallarino, F.; Orabona, C.; Alunno, A.; et al. Indoleamine 2,3-dioxygenase 1 (IDO1) is up-regulated in thyroid carcinoma and drives the development of an immunosuppressant tumor microenvironment. J. Clin. Endocrinol. Metab. 2014, 99, E832–E840. [Google Scholar] [CrossRef]
- Mian, C.; Ceccato, F.; Barollo, S.; Watutantrige-Fernando, S.; Albiger, N.; Regazzo, D.; de Lazzari, P.; Pennelli, G.; Rotondi, S.; Nacamulli, D.; et al. AHR over-expression in papillary thyroid carcinoma: Clinical and molecular assessments in a series of Italian acromegalic patients with a long-term follow-up. PLoS ONE 2014, 9, e101560. [Google Scholar] [CrossRef]
- Occhi, G.; Barollo, S.; Regazzo, D.; Bertazza, L.; Galuppini, F.; Guzzardo, V.; Jaffrain-Rea, M.L.; Vianello, F.; Ciato, D.; Ceccato, F.; et al. A constitutive active MAPK/ERK pathway due to BRAFV600E positively regulates AHR pathway in PTC. Oncotarget 2015, 6, 32104–32114. [Google Scholar] [CrossRef][Green Version]
- Landa, I.; Knauf, J.A. Mouse Models as a Tool for Understanding Progression in BrafV600E-Driven Thyroid Cancers. Endocrinol. Metab. 2019, 34, 11–22. [Google Scholar] [CrossRef]
- Kim, S.H.; Henry, E.C.; Kim, D.K.; Kim, Y.H.; Shin, K.J.; Han, M.S.; Lee, T.G.; Kang, J.K.; Gasiewicz, T.A.; Ryu, S.H.; et al. Novel compound 2-methyl-2H-pyrazole-3-carboxylic acid (2-methyl-4-o-tolylazo-phenyl)-amide (CH-223191) prevents 2,3,7,8-TCDD-induced toxicity by antagonizing the aryl hydrocarbon receptor. Mol. Pharmacol. 2006, 69, 1871–1878. [Google Scholar] [CrossRef]
- Androutsopoulos, V.P.; Tsatsakis, A.M.; Spandidos, D.A. Cytochrome P450 CYP1A1: Wider roles in cancer progression and prevention. BMC Cancer 2009, 9, 187. [Google Scholar] [CrossRef]
- Murray, G.I.; Taylor, M.C.; McFadyen, M.C.; McKay, J.A.; Greenlee, W.F.; Burke, M.D.; Melvin, W.T. Tumor-specific expression of cytochrome P450 CYP1B1. Cancer Res. 1997, 57, 3026–3031. [Google Scholar]
- Cheng, J.; Li, W.; Kang, B.; Zhou, Y.; Song, J.; Dan, S.; Yang, Y.; Zhang, X.; Li, J.; Yin, S.; et al. Tryptophan derivatives regulate the transcription of Oct4 in stem-like cancer cells. Nat. Commun. 2015, 6, 7209. [Google Scholar] [CrossRef] [PubMed]
- Ryu, B.; Moriarty, W.F.; Stine, M.J.; DeLuca, A.; Kim, D.S.; Meeker, A.K.; Grills, L.D.; Switzer, R.A.; Eller, M.S.; Alani, R.M. Global analysis of BRAFV600E target genes in human melanocytes identifies matrix metalloproteinase-1 as a critical mediator of melanoma growth. J. Investig. Dermatol. 2011, 131, 1579–1583. [Google Scholar] [CrossRef] [PubMed]
- Knauf, J.A.; Luckett, K.A.; Chen, K.Y.; Voza, F.; Socci, N.D.; Ghossein, R.; Fagin, J.A. Hgf/Met activation mediates resistance to BRAF inhibition in murine anaplastic thyroid cancers. J. Clin. Investig. 2018, 128, 4086–4097. [Google Scholar] [CrossRef] [PubMed]
- Franco, A.T.; Malaguarnera, R.; Refetoff, S.; Liao, X.H.; Lundsmith, E.; Kimura, S.; Pritchard, C.; Marais, R.; Davies, T.F.; Weinstein, L.S.; et al. Thyrotrophin receptor signaling dependence of Braf-induced thyroid tumor initiation in mice. Proc. Natl. Acad. Sci. USA 2011, 108, 1615–1620. [Google Scholar] [CrossRef]
- Chakravarty, D.; Santos, E.; Ryder, M.; Knauf, J.A.; Liao, X.H.; West, B.L.; Bollag, G.; Kolesnick, R.; Thin, T.H.; Rosen, N.; et al. Small-molecule MAPK inhibitors restore radioiodine incorporation in mouse thyroid cancers with conditional BRAF activation. J. Clin. Investig. 2011, 121, 4700–4711. [Google Scholar] [CrossRef]
- Fabien, N.; Fusco, A.; Santoro, M.; Barbier, Y.; Dubois, P.M.; Paulin, C. Description of a human papillary thyroid carcinoma cell line. Morphologic study and expression of tumoral markers. Cancer 1994, 73, 2206–2212. [Google Scholar] [CrossRef]
- Tanaka, J.; Ogura, T.; Sato, H.; Hatano, M. Establishment and biological characterization of an in vitro human cytomegalovirus latency model. Virology 1987, 161, 62–72. [Google Scholar] [CrossRef]
- Schweppe, R.E.; Klopper, J.P.; Korch, C.; Pugazhenthi, U.; Benezra, M.; Knauf, J.A.; Fagin, J.A.; Marlow, L.A.; Copland, J.A.; Smallridge, R.C.; et al. Deoxyribonucleic acid profiling analysis of 40 human thyroid cancer cell lines reveals cross contamination resulting in cell line redundancy and misidentification. J. Clin. Endocrinol. Metab. 2008, 93, 4331–4341. [Google Scholar] [CrossRef]
- Gustavsson, B.; Hermansson, A.; Andersson, A.C.; Grimelius, L.; Bergh, J.; Westermark, B.; Heldin, N.E. Decreased growth rate and tumour formation of human anaplastic thyroid carcinoma cells transfected with a human thyrotropin receptor cDNA in NMRI nude mice treated with propylthiouracil. Mol. Cell Endocrinol. 1996, 121, 143–151. [Google Scholar] [CrossRef]
- Ito, T.; Seyama, T.; Hayashi, Y.; Hayashi, T.; Dohi, K.; Mizuno, T.; Iwamoto, K.S.; Tsuyama, N.; Nakamura, N.; Akiyama, M. Establishment of 2 human thyroid-carcinoma cell-lines (8305c, 8505c) bearing p53 gene-mutations. Int. J. Oncol. 1994, 4, 583–586. [Google Scholar] [CrossRef]
- Goretzki, P.E.; Frilling, A.; Simon, D.; Roeher, H.D. Growth regulation of normal thyroids and thyroid tumors in man. Recent Results Cancer Res. 1990, 118, 48–63. [Google Scholar] [PubMed]
- Gioanni, J.; Zanghellini, E.; Mazeau, C.; Zhang, D.; Courdi, A.; Farges, M.; Lambert, J.C.; Duplay, H.; Schneider, M. Characterization of a human cell line from an anaplastic carcinoma of the thyroid gland. Bull. Cancer 1991, 78, 1053–1062. [Google Scholar] [PubMed]
- Durante, C.; Puxeddu, E.; Ferretti, E.; Morisi, R.; Moretti, S.; Bruno, R.; Barbi, F.; Avenia, N.; Scipioni, A.; Verrienti, A.; et al. BRAFmutations in papillary thyroid carcinomas inhibit genes involved in iodine metabolism. J. Clin. Endocrinol. Metab. 2007, 92, 2840–2843. [Google Scholar] [CrossRef] [PubMed]
- Puxeddu, E.; Knauf, J.A.; Sartor, M.A.; Mitsutake, N.; Smith, E.P.; Medvedovic, M.; Tomlinson, C.R.; Moretti, S.; Fagin, J.A. RET/PTC-induced gene expression in thyroid PCCL3 cells reveals early activation of genes involved in regulation of the immune response. Endocr. Rel. Cancer 2005, 12, 319–334. [Google Scholar] [CrossRef] [PubMed]
- Muller, P.Y.; Janovjak, H.; Miserez, A.R.; Dobbie, Z. Processing of gene expression data generated by quantitative real-time RT-PCR. Biotechniques 2002, 32, 1372–1374, 1376, 1378–1379. [Google Scholar]
- Colella, R.; Mameli, M.G.; Bellezza, G.; Sordo, R.D.; Cavaliere, A.; Sidoni, A. Endometriosis-associated skeletal muscle regeneration: A hitherto undescribed entity and a potential diagnostic pitfall. Am. J. Surg. Pathol. 2010, 34, 10–17. [Google Scholar] [CrossRef]
| Mice Model: TetOn-BRAF-P53 | |||||
|---|---|---|---|---|---|
| ID | Sex | Analyzed Tissue | Dox Treatment (Weeks) | Dox Withdrawal (Weeks) | Age at Sacrifice (Weeks) |
| B101 | F | Thyroid Cancer | 6 | 0 | 10 |
| B101L | F | Lymph node metastasis | 6 | 0 | 10 |
| A3858 | F | Thyroid Cancer | 7 | 7 | 20 |
| A3624 | M | Thyroid Cancer | 5 | 0 | 11 |
| A8717 | M | Thyroid Cancer | 8 | 3 | 17 |
| A8742L | M | Lymph node metastasis | 7 | 4 | 17 |
| A2492 | M | Thyroid Cancer | 6 | 0 | 13 |
| B246 | M | Thyroid Cancer | 10 | 1 | 16 |
| Mice Model: BRAF-Lox/TPO-Cre | ||||
|---|---|---|---|---|
| ID | Sex | BRAF Genotyping | Histotype | Age at Sacrifice (Weeks) |
| 6135 | M | HET | PTC | 10 |
| 2790 | M | WT | NT | 33 |
| 2731 | M | HET | PTC | 35 |
| 4826 | F | WT | NT | 19 |
| 4816 | F | HET | PTC | 21 |
| 4820 | F | HET | PTC | 20 |
| 2811 | F | WT | NT | 31 |
| 2786 | F | HET | PTC | 36 |
| 2787 | F | HET | PTC | 36 |
| Primer Sequence (5′-3′) | |
|---|---|
| Β-Actin | GCACAGAGCCTCGCCTTTGCC ATGCCCACCATCACGCCCTGG |
| CYP1B1 | GGCTGGATTTGGAGAACGTA CTCGAGTCTGCACATCAGGA |
| E-Cadherin | CCCAGGAGCCAGACACATTT TTAGGGCTGTGTACGTGCTG |
| Fibronectin 1 | GAGATGAAACCTGAAGCTGA ACCAATCTTGTAGGACTGAC |
| MMP1 | AGTCCAGAAATACCTGGAAAAATAC CCACATCAGGCACTCCACAT |
| MMP2 | GCTACGATGGAGGCGCTAAT TCAGGTATTGCACTGCCAACT |
| MMP9 | CTGGAGGTTCGACGTGAAGG AGCGGTCCTGGCAGAAATAG |
| N-Cadherin | GGACCGAGAATCACCAAATG CTTGAGGTAACACTTGAGGG |
| OCT4 | GAGTGAGAGGCAACCTGGAG ACACTCGGACCACATCCTTC |
| SLUG | GACCCTGGTTGCTTCAAGGA GAATGGGTCTGCAGATGAGC |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Moretti, S.; Nucci, N.; Menicali, E.; Morelli, S.; Bini, V.; Colella, R.; Mandarano, M.; Sidoni, A.; Puxeddu, E. The Aryl Hydrocarbon Receptor Is Expressed in Thyroid Carcinoma and Appears to Mediate Epithelial-Mesenchymal-Transition. Cancers 2020, 12, 145. https://doi.org/10.3390/cancers12010145
Moretti S, Nucci N, Menicali E, Morelli S, Bini V, Colella R, Mandarano M, Sidoni A, Puxeddu E. The Aryl Hydrocarbon Receptor Is Expressed in Thyroid Carcinoma and Appears to Mediate Epithelial-Mesenchymal-Transition. Cancers. 2020; 12(1):145. https://doi.org/10.3390/cancers12010145
Chicago/Turabian StyleMoretti, Sonia, Nicole Nucci, Elisa Menicali, Silvia Morelli, Vittorio Bini, Renato Colella, Martina Mandarano, Angelo Sidoni, and Efisio Puxeddu. 2020. "The Aryl Hydrocarbon Receptor Is Expressed in Thyroid Carcinoma and Appears to Mediate Epithelial-Mesenchymal-Transition" Cancers 12, no. 1: 145. https://doi.org/10.3390/cancers12010145
APA StyleMoretti, S., Nucci, N., Menicali, E., Morelli, S., Bini, V., Colella, R., Mandarano, M., Sidoni, A., & Puxeddu, E. (2020). The Aryl Hydrocarbon Receptor Is Expressed in Thyroid Carcinoma and Appears to Mediate Epithelial-Mesenchymal-Transition. Cancers, 12(1), 145. https://doi.org/10.3390/cancers12010145