Exploiting ING2 Epigenetic Modulation as a Therapeutic Opportunity for Non-Small Cell Lung Cancer
Abstract
1. Introduction
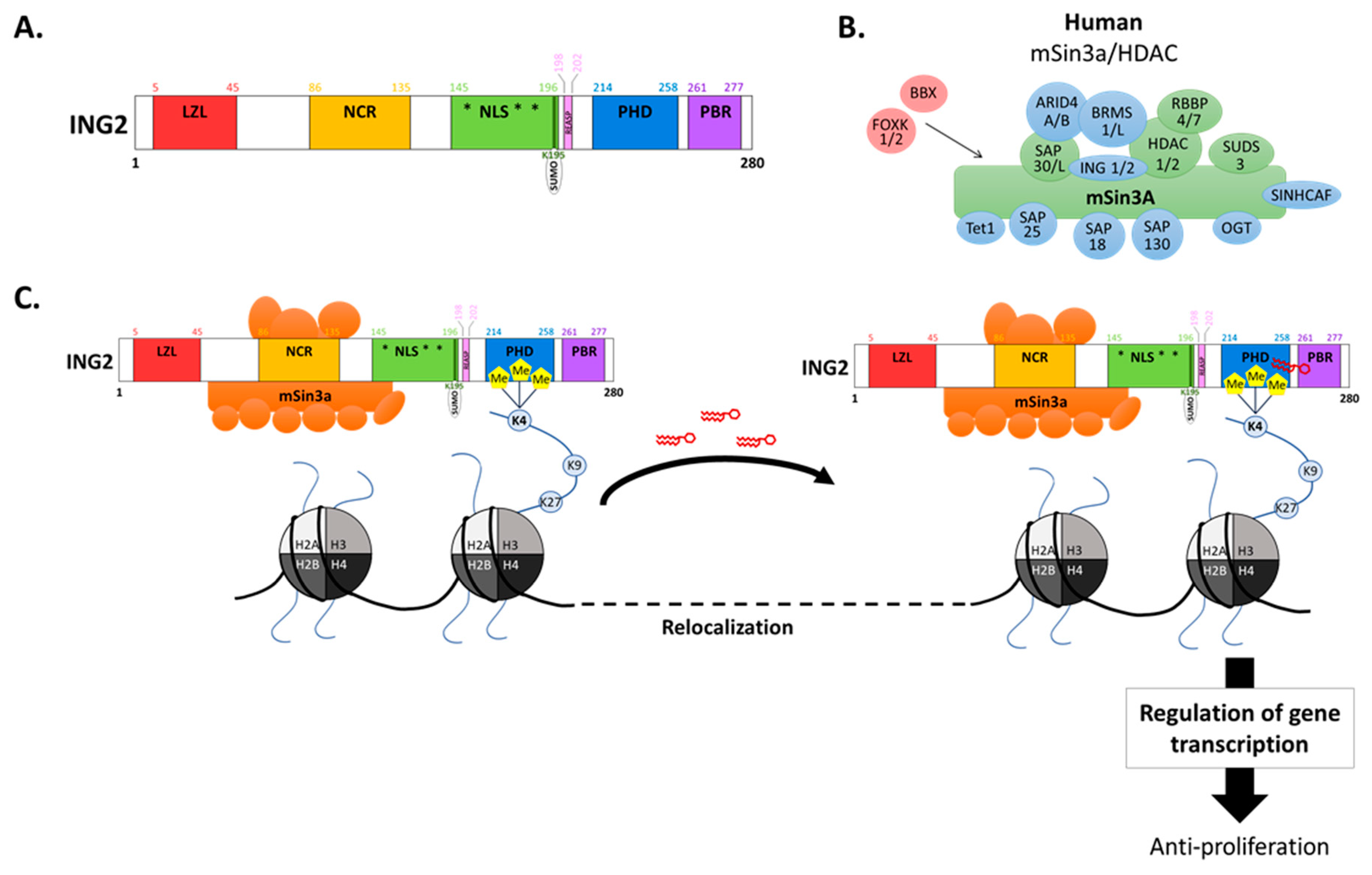
2. ING2 Modulates Transcriptional Activity Through Chromatin Remodeling
3. ING2 Status in Human Tumors
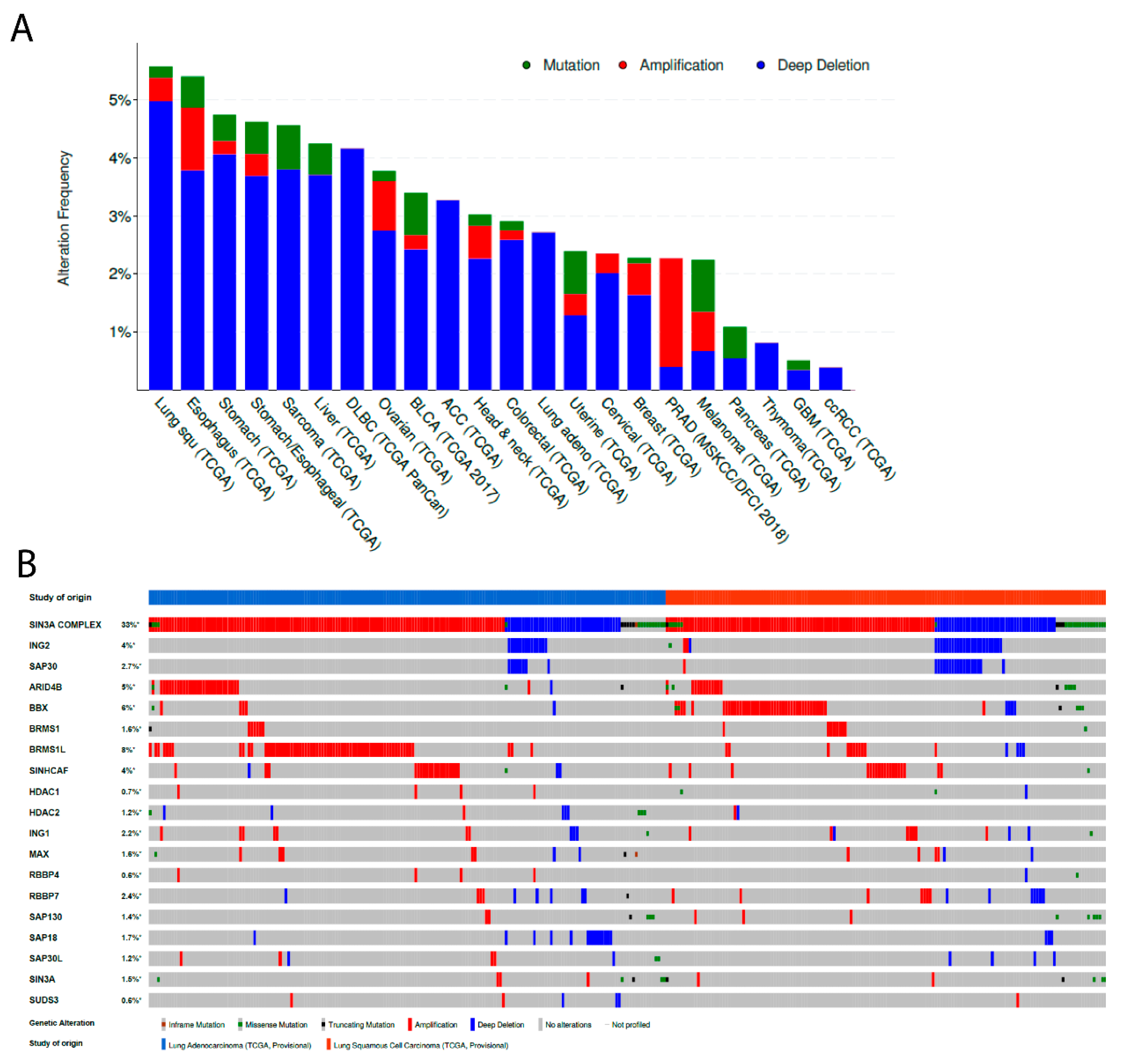
3.1. ING2 Alterations in Human Tumors
3.2. ING2 Alterations in Non-Small Cell Lung Cancer
4. Potential Role of the ING2 Epigenetic Modulation in Lung Cancer Treatment
4.1. Exploring ING2 Dependencies in Cancer Cells
4.2. Targeting SIN3A-Mediated ING2 Functions in Cancer Cells
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Molina, J.R.; Yang, P.; Cassivi, S.D.; Schild, S.E.; Adjei, A.A. Non-small cell lung cancer: Epidemiology, risk factors, treatment, and survivorship. Mayo Clin. Proc. 2008, 83, 584–594. [Google Scholar] [CrossRef]
- Shi, X.; Hong, T.; Walter, K.L.; Ewalt, M.; Michishita, E.; Hung, T.; Carney, D.; Peña, P.; Lan, F.; Kaadige, M.R.; et al. ING2 PHD domain links histone H3 lysine 4 methylation to active gene repression. Nature 2006, 442, 96–99. [Google Scholar] [CrossRef] [PubMed]
- Peña, P.V.; Davrazou, F.; Shi, X.; Walter, K.L.; Verkhusha, V.V.; Gozani, O.; Zhao, R.; Kutateladze, T.G. Molecular mechanism of histone H3K4me3 recognition by plant homeodomain of ING2. Nature 2006, 442, 100–103. [Google Scholar] [CrossRef] [PubMed]
- Doyon, Y.; Cayrou, C.; Ullah, M.; Landry, A.-J.; Côté, V.; Selleck, W.; Lane, W.S.; Tan, S.; Yang, X.-J.; Côté, J. ING tumor suppressor proteins are critical regulators of chromatin acetylation required for genome expression and perpetuation. Mol. Cell 2006, 21, 51–64. [Google Scholar] [CrossRef] [PubMed]
- Pile, L.A.; Spellman, P.T.; Katzenberger, R.J.; Wassarman, D.A. The SIN3 deacetylase complex represses genes encoding mitochondrial proteins: Implications for the regulation of energy metabolism. J. Biol. Chem. 2003, 278, 37840–37848. [Google Scholar] [CrossRef]
- Larrieu, D.; Ythier, D.; Binet, R.; Brambilla, C.; Brambilla, E.; Sengupta, S.; Pedeux, R. ING2 controls the progression of DNA replication forks to maintain genome stability. EMBO Rep. 2009, 10, 1168–1174. [Google Scholar] [CrossRef]
- Pedeux, R.; Sengupta, S.; Shen, J.C.; Demidov, O.N.; Saito, S.; Onogi, H.; Kumamoto, K.; Wincovitch, S.; Garfield, S.H.; McMenamin, M.; et al. ING2 regulates the onset of replicative senescence by induction of p300-dependent p53 acetylation. Mol. Cell. Biol. 2005, 25, 6639–6648. [Google Scholar] [CrossRef]
- Wang, J.; Chin, M.Y.; Li, G. The novel tumor suppressor p33ING2 enhances nucleotide excision repair via inducement of histone H4 acetylation and chromatin relaxation. Cancer Res. 2006, 66, 1906–1911. [Google Scholar] [CrossRef]
- Saito, M.; Kumamoto, K.; Robles, A.I.; Horikawa, I.; Furusato, B.; Okamura, S.; Goto, A.; Yamashita, T.; Nagashima, M.; Lee, T.-L.; et al. Targeted disruption of Ing2 results in defective spermatogenesis and development of soft-tissue sarcomas. PLoS ONE 2010, 5, e15541. [Google Scholar] [CrossRef]
- Garkavtsev, I.; Kazarov, A.; Gudkov, A.; Riabowol, K. Suppression of the novel growth inhibitor p33ING1 promotes neoplastic transformation. Nat. Genet. 1996, 14, 415–420. [Google Scholar] [CrossRef]
- Shimada, Y.; Saito, A.; Suzuki, M.; Takahashi, E.; Horie, M. Cloning of a novel gene (ING1L) homologous to ING1, a candidate tumor suppressor. Cytogenet. Cell Genet. 1998, 83, 232–235. [Google Scholar] [CrossRef] [PubMed]
- Nagashima, M.; Shiseki, M.; Miura, K.; Hagiwara, K.; Linke, S.P.; Pedeux, R.; Wang, X.W.; Yokota, J.; Riabowol, K.; Harris, C.C. DNA damage-inducible gene p33ING2 negatively regulates cell proliferation through acetylation of p53. Proc. Natl. Acad. Sci. USA 2001, 98, 9671–9676. [Google Scholar] [CrossRef] [PubMed]
- Nagashima, M.; Shiseki, M.; Pedeux, R.M.; Okamura, S.; Kitahama-Shiseki, M.; Miura, K.; Yokota, J.; Harris, C.C. A novel PHD-finger motif protein, p47ING3, modulates p53-mediated transcription, cell cycle control, and apoptosis. Oncogene 2003, 22, 343–350. [Google Scholar] [CrossRef] [PubMed]
- Shiseki, M.; Nagashima, M.; Pedeux, R.M.; Kitahama-Shiseki, M.; Miura, K.; Okamura, S.; Onogi, H.; Higashimoto, Y.; Appella, E.; Yokota, J.; et al. p29ING4 and p28ING5 bind to p53 and p300, and enhance p53 activity. Cancer Res. 2003, 63, 2373–2378. [Google Scholar]
- Unoki, M.; Kumamoto, K.; Robles, A.I.; Shen, J.C.; Zheng, Z.-M.; Harris, C.C. A novel ING2 isoform, ING2b, synergizes with ING2a to prevent cell cycle arrest and apoptosis. FEBS Lett. 2008, 582, 3868–3874. [Google Scholar] [CrossRef] [PubMed]
- Luger, K.; Mäder, A.W.; Richmond, R.K.; Sargent, D.F.; Richmond, T.J. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 1997, 389, 251–260. [Google Scholar] [CrossRef]
- Murray, K. The occurrence of epsilon-n-methyl lysine in histones. Biochemistry 1964, 3, 10–15. [Google Scholar] [CrossRef]
- Fischle, W.; Franz, H.; Jacobs, S.A.; Allis, C.D.; Khorasanizadeh, S. Specificity of the chromodomain Y chromosome family of chromodomains for lysine-methylated ARK(S/T) motifs. J. Biol. Chem. 2008, 283, 19626–19635. [Google Scholar] [CrossRef]
- Bernstein, B.E.; Kamal, M.; Lindblad-Toh, K.; Bekiranov, S.; Bailey, D.K.; Huebert, D.J.; McMahon, S.; Karlsson, E.K.; Kulbokas, E.J.; Gingeras, T.R.; et al. Genomic maps and comparative analysis of histone modifications in human and mouse. Cell 2005, 120, 169–181. [Google Scholar] [CrossRef]
- Santos-Rosa, H.; Schneider, R.; Bannister, A.J.; Sherriff, J.; Bernstein, B.E.; Emre, N.C.T.; Schreiber, S.L.; Mellor, J.; Kouzarides, T. Active genes are tri-methylated at K4 of histone H3. Nature 2002, 419, 407–411. [Google Scholar] [CrossRef]
- Gozani, O.; Karuman, P.; Jones, D.R.; Ivanov, D.; Cha, J.; Lugovskoy, A.A.; Baird, C.L.; Zhu, H.; Field, S.J.; Lessnick, S.L.; et al. The PHD finger of the chromatin-associated protein ING2 functions as a nuclear phosphoinositide receptor. Cell 2003, 114, 99–111. [Google Scholar] [CrossRef]
- Kaadige, M.R.; Ayer, D.E. The polybasic region that follows the plant homeodomain zinc finger 1 of Pf1 is necessary and sufficient for specific phosphoinositide binding. J. Biol. Chem. 2006, 281, 28831–28836. [Google Scholar] [CrossRef] [PubMed]
- Bua, D.J.; Martin, G.M.; Binda, O.; Gozani, O. Nuclear phosphatidylinositol-5-phosphate regulates ING2 stability at discrete chromatin targets in response to DNA damage. Sci. Rep. 2013, 3, 2137. [Google Scholar] [CrossRef] [PubMed]
- Ohkouchi, C.; Kumamoto, K.; Saito, M.; Ishigame, T.; Suzuki, S.-I.; Takenoshita, S.; Harris, C.C. ING2, a tumor associated gene, enhances PAI-1 and HSPA1A expression with HDAC1 and mSin3A through the PHD domain and C-terminal. Mol. Med. Rep. 2017, 16, 7367–7374. [Google Scholar] [CrossRef][Green Version]
- Ythier, D.; Larrieu, D.; Binet, R.; Binda, O.; Brambilla, C.; Gazzeri, S.; Pedeux, R. Sumoylation of ING2 regulates the transcription mediated by Sin3A. Oncogene 2010, 29, 5946–5956. [Google Scholar] [CrossRef]
- Zhang, Y.; Iratni, R.; Erdjument-Bromage, H.; Tempst, P.; Reinberg, D. Histone deacetylases and SAP18, a novel polypeptide, are components of a human Sin3 complex. Cell 1997, 89, 357–364. [Google Scholar] [CrossRef]
- Parthun, M.R.; Widom, J.; Gottschling, D.E. The major cytoplasmic histone acetyltransferase in yeast: Links to chromatin replication and histone metabolism. Cell 1996, 87, 85–94. [Google Scholar] [CrossRef]
- Laherty, C.D.; Billin, A.N.; Lavinsky, R.M.; Yochum, G.S.; Bush, A.C.; Sun, J.M.; Mullen, T.M.; Davie, J.R.; Rose, D.W.; Glass, C.K.; et al. SAP30, a component of the mSin3 corepressor complex involved in N-CoR-mediated repression by specific transcription factors. Mol. Cell 1998, 2, 33–42. [Google Scholar] [CrossRef]
- Zhang, Y.; Sun, Z.W.; Iratni, R.; Erdjument-Bromage, H.; Tempst, P.; Hampsey, M.; Reinberg, D. SAP30, a novel protein conserved between human and yeast, is a component of a histone deacetylase complex. Mol. Cell 1998, 1, 1021–1031. [Google Scholar] [CrossRef]
- Alland, L.; David, G.; Shen-Li, H.; Potes, J.; Muhle, R.; Lee, H.-C.; Hou, H.; Chen, K.; DePinho, R.A. Identification of mammalian Sds3 as an integral component of the Sin3/histone deacetylase corepressor complex. Mol. Cell. Biol. 2002, 22, 2743–2750. [Google Scholar] [CrossRef]
- Yang, X.; Zhang, F.; Kudlow, J.E. Recruitment of O-GlcNAc transferase to promoters by corepressor mSin3A: Coupling protein O-GlcNAcylation to transcriptional repression. Cell 2002, 110, 69–80. [Google Scholar] [CrossRef]
- Muñoz, I.M.; MacArtney, T.; Sanchez-Pulido, L.; Ponting, C.P.; Rocha, S.; Rouse, J. Family with sequence similarity 60A (FAM60A) protein is a cell cycle-fluctuating regulator of the SIN3-HDAC1 histone deacetylase complex. J. Biol. Chem. 2012, 287, 32346–32353. [Google Scholar] [CrossRef] [PubMed]
- Fleischer, T.C.; Yun, U.J.; Ayer, D.E. Identification and characterization of three new components of the mSin3A corepressor complex. Mol. Cell. Biol. 2003, 23, 3456–3467. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Lin, Q.; Wang, W.; Wade, P.; Wong, J. Specific targeting and constitutive association of histone deacetylase complexes during transcriptional repression. Genes Dev. 2002, 16, 687–692. [Google Scholar] [CrossRef] [PubMed]
- Peinado, H.; Ballestar, E.; Esteller, M.; Cano, A. Snail mediates E-cadherin repression by the recruitment of the Sin3A/histone deacetylase 1 (HDAC1)/HDAC2 complex. Mol. Cell. Biol. 2004, 24, 306–319. [Google Scholar] [CrossRef]
- Kumamoto, K.; Fujita, K.; Kurotani, R.; Saito, M.; Unoki, M.; Hagiwara, N.; Shiga, H.; Bowman, E.D.; Yanaihara, N.; Okamura, S.; et al. ING2 is upregulated in colon cancer and increases invasion by enhanced MMP13 expression. Int. J. Cancer 2009, 125, 1306–1315. [Google Scholar] [CrossRef]
- Sardiu, M.E.; Smith, K.T.; Groppe, B.D.; Gilmore, J.M.; Saraf, A.; Egidy, R.; Peak, A.; Seidel, C.W.; Florens, L.; Workman, J.L.; et al. Suberoylanilide hydroxamic acid (SAHA)-induced dynamics of a human histone deacetylase protein interaction network. Mol. Cell Proteomics 2014, 13, 3114–3125. [Google Scholar] [CrossRef]
- Kumamoto, K.; Spillare, E.A.; Fujita, K.; Horikawa, I.; Yamashita, T.; Appella, E.; Nagashima, M.; Takenoshita, S.; Yokota, J.; Harris, C.C. Nutlin-3a activates p53 to both down-regulate inhibitor of growth 2 and up-regulate mir-34a, mir-34b, and mir-34c expression, and induce senescence. Cancer Res. 2008, 68, 3193–3203. [Google Scholar] [CrossRef]
- Serrano, M.; Lin, A.W.; McCurrach, M.E.; Beach, D.; Lowe, S.W. Oncogenic ras provokes premature cell senescence associated with accumulation of p53 and p16INK4a. Cell 1997, 88, 593–602. [Google Scholar] [CrossRef]
- Lotem, J.; Sachs, L. Hematopoietic cells from mice deficient in wild-type p53 are more resistant to induction of apoptosis by some agents. Blood 1993, 82, 1092–1096. [Google Scholar] [CrossRef]
- Yonish-Rouach, E.; Resnitzky, D.; Lotem, J.; Sachs, L.; Kimchi, A.; Oren, M. Wild-type p53 induces apoptosis of myeloid leukaemic cells that is inhibited by interleukin-6. Nature 1991, 352, 345–347. [Google Scholar] [CrossRef] [PubMed]
- Di Leonardo, A.; Linke, S.P.; Clarkin, K.; Wahl, G.M. DNA damage triggers a prolonged p53-dependent G1 arrest and long-term induction of Cip1 in normal human fibroblasts. Genes Dev. 1994, 8, 2540–2551. [Google Scholar] [CrossRef] [PubMed]
- Lu, F.; Dai, D.L.; Martinka, M.; Ho, V.; Li, G. Nuclear ING2 expression is reduced in human cutaneous melanomas. Br. J. Cancer 2006, 95, 80–86. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Pan, K.; Wang, H.; Weng, D.; Song, H.; Zhou, J.; Huang, W.; Li, J.; Chen, M.; Xia, J. Decreased expression of ING2 gene and its clinicopathological significance in hepatocellular carcinoma. Cancer Lett. 2008, 261, 183–192. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Yang, X.-F.; Gou, W.-F.; Lu, H.; Li, H.; Zhu, Z.-T.; Sun, H.-Z.; Zheng, H.-C. Expression profiles of inhibitor of growth protein 2 in normal and cancer tissues: An immunohistochemical screening analysis. Mol. Med. Rep. 2016, 13, 1881–1887. [Google Scholar] [CrossRef][Green Version]
- Han, X.-R.; Bai, X.-Z.; Sun, Y.; Yang, Y. Nuclear ING2 expression is reduced in osteosarcoma. Oncol. Rep. 2014, 32, 1967–1972. [Google Scholar] [CrossRef]
- Jiang, J.; Liu, Y.; Zhao, Y.; Tian, F.; Wang, G. miR-153-3p Suppresses Inhibitor of Growth Protein 2 Expression to Function as Tumor Suppressor in Acute Lymphoblastic Leukemia. Technol. Cancer Res. Treat. 2019, 18, 1533033819852990. [Google Scholar] [CrossRef]
- Temel, M.; Turkmen, A.; Dokuyucu, R.; Cevik, C.; Oztuzcu, S.; Cengiz, B.; Mutaf, M. A novel tumor suppressor gene in basal cell carcinoma: Inhibition of growth factor-2. Tumour Biol. 2015, 36, 4611–4616. [Google Scholar] [CrossRef]
- Okano, T.; Gemma, A.; Hosoya, Y.; Hosomi, Y.; Nara, M.; Kokubo, Y.; Yoshimura, A.; Shibuya, M.; Nagashima, M.; Harris, C.C.; et al. Alterations in novel candidate tumor suppressor genes, ING1 and ING2 in human lung cancer. Oncol. Rep. 2006, 15, 545–549. [Google Scholar] [CrossRef][Green Version]
- Walzak, A.A.; Veldhoen, N.; Feng, X.; Riabowol, K.; Helbing, C.C. Expression profiles of mRNA transcript variants encoding the human inhibitor of growth tumor suppressor gene family in normal and neoplastic tissues. Exp. Cell Res. 2008, 314, 273–285. [Google Scholar] [CrossRef]
- Gao, Y.; Ma, H.; Gao, C.; Lv, Y.; Chen, X.; Xu, R.; Sun, M.; Liu, X.; Lu, X.; Pei, X.; et al. Tumor-promoting properties of miR-8084 in breast cancer through enhancing proliferation, suppressing apoptosis and inducing epithelial-mesenchymal transition. J. Transl. Med. 2018, 16, 38. [Google Scholar] [CrossRef] [PubMed]
- Gournay, M.; Paineau, M.; Archambeau, J.; Pedeux, R. Regulat-INGs in tumors and diseases: Focus on ncRNAs. Cancer Lett. 2019, 447, 66–74. [Google Scholar] [CrossRef] [PubMed]
- Sironi, E.; Cerri, A.; Tomasini, D.; Sirchia, S.M.; Porta, G.; Rossella, F.; Grati, F.R.; Simoni, G. Loss of heterozygosity on chromosome 4q32-35 in sporadic basal cell carcinomas: Evidence for the involvement of p33ING2/ING1L and SAP30 genes. J. Cutan. Pathol. 2004, 31, 318–322. [Google Scholar] [CrossRef] [PubMed]
- Borkosky, S.S.; Gunduz, M.; Nagatsuka, H.; Beder, L.B.; Gunduz, E.; Ali, M.A.L.S.; Rodriguez, A.P.; Cilek, M.Z.; Tominaga, S.; Yamanaka, N.; et al. Frequent deletion of ING2 locus at 4q35.1 associates with advanced tumor stage in head and neck squamous cell carcinoma. J. Cancer Res. Clin. Oncol. 2009, 135, 703–713. [Google Scholar] [CrossRef]
- Zhang, H.; Ma, H.; Wang, Q.; Chen, M.; Weng, D.; Wang, H.; Zhou, J.; Li, Y.; Sun, J.; Chen, Y.; et al. Analysis of loss of heterozygosity on chromosome 4q in hepatocellular carcinoma using high-throughput SNP array. Oncol. Rep. 2010, 23, 445–455. [Google Scholar] [PubMed]
- Borkosky, S.S.; Gunduz, M.; Beder, L.; Tsujigiwa, H.; Tamamura, R.; Gunduz, E.; Katase, N.; Rodriguez, A.P.; Sasaki, A.; Nagai, N.; et al. Allelic loss of the ING gene family loci is a frequent event in ameloblastoma. Oncol. Res. 2010, 18, 509–518. [Google Scholar] [CrossRef] [PubMed]
- Cetin, E.; Cengiz, B.; Gunduz, E.; Gunduz, M.; Nagatsuka, H.; Bekir-Beder, L.; Fukushima, K.; Pehlivan, D.; Nishizaki, K.; Shimizu, K.; et al. Deletion mapping of chromosome 4q22-35 and identification of four frequently deleted regions in head and neck cancers. Neoplasma 2008, 55, 299–304. [Google Scholar]
- Mittal, K.R.; Chen, F.; Wei, J.J.; Rijhvani, K.; Kurvathi, R.; Streck, D.; Dermody, J.; Toruner, G.A. Molecular and immunohistochemical evidence for the origin of uterine leiomyosarcomas from associated leiomyoma and symplastic leiomyoma-like areas. Mod. Pathol. 2009, 22, 1303–1311. [Google Scholar] [CrossRef]
- Wang, H.-Q.; Yang, C.-Y.; Wang, S.-Y.; Wang, T.; Han, J.-L.; Wei, K.; Liu, F.-C.; Xu, J.; Peng, X.-Z.; Wang, J.-M. Cell-free plasma hypermethylated CASZ1, CDH13 and ING2 are promising biomarkers of esophageal cancer. J. Biomed. Res. 2018, 32, 424–433. [Google Scholar]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012, 2, 401–404. [Google Scholar] [CrossRef]
- Gao, J.; Aksoy, B.A.; Dogrusoz, U.; Dresdner, G.; Gross, B.; Sumer, S.O.; Sun, Y.; Jacobsen, A.; Sinha, R.; Larsson, E.; et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal 2013, 6, pl1. [Google Scholar] [CrossRef] [PubMed]
- Ythier, D.; Brambilla, E.; Binet, R.; Nissou, D.; Vesin, A.; de Fraipont, F.; Moro-Sibilot, D.; Lantuejoul, S.; Brambilla, C.; Gazzeri, S.; et al. Expression of candidate tumor suppressor gene ING2 is lost in non-small cell lung carcinoma. Lung Cancer 2010, 69, 180–186. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.Q.; Zhang, X.; Xu, D.P.; Bao, W.G.; Lin, A.F.; Xu, H.H.; Yan, W.H. Decreased expression of ING2 gene and its clinicopathological significance in Chinese NSCLC patients. Neoplasma 2014, 61, 468–475. [Google Scholar] [CrossRef] [PubMed]
- Al Zeyadi, M.; Dimova, I.; Ranchich, V.; Rukova, B.; Nesheva, D.; Hamude, Z.; Georgiev, S.; Petrov, D.; Toncheva, D. Whole genome microarray analysis in non-small cell lung cancer. Biotechnol. Biotechnol. Equip. 2015, 29, 111–118. [Google Scholar] [CrossRef] [PubMed]
- Nie, J.; Liu, L.; Wu, M.; Xing, G.; He, S.; Yin, Y.; Tian, C.; He, F.; Zhang, L. HECT ubiquitin ligase Smurf1 targets the tumor suppressor ING2 for ubiquitination and degradation. FEBS Lett. 2010, 584, 3005–3012. [Google Scholar] [CrossRef]
- Li, H.; Xiao, N.; Wang, Y.; Wang, R.; Chen, Y.; Pan, W.; Liu, D.; Li, S.; Sun, J.; Zhang, K.; et al. Smurf1 regulates lung cancer cell growth and migration through interaction with and ubiquitination of PIPKIγ. Oncogene 2017, 36, 5668–5680. [Google Scholar] [CrossRef]
- Tan, W.-L.; Jain, A.; Takano, A.; Newell, E.W.; Iyer, N.G.; Lim, W.-T.; Tan, E.-H.; Zhai, W.; Hillmer, A.M.; Tam, W.-L.; et al. Novel therapeutic targets on the horizon for lung cancer. Lancet Oncol. 2016, 17, e347–e362. [Google Scholar] [CrossRef]
- Smolle, E.; Fink-Neuboeck, N.; Lindenmann, J.; Smolle-Juettner, F.; Pichler, M. The Biological and Clinical Relevance of Inhibitor of Growth (ING) Genes in Non-Small Cell Lung Cancer. Cancers 2019, 11, 1118. [Google Scholar] [CrossRef]
- Tsherniak, A.; Vazquez, F.; Montgomery, P.G.; Weir, B.A.; Kryukov, G.; Cowley, G.S.; Gill, S.; Harrington, W.F.; Pantel, S.; Krill-Burger, J.M.; et al. Defining a Cancer Dependency Map. Cell 2017, 170, 564–576. [Google Scholar] [CrossRef]
- Laherty, C.D.; Yang, W.M.; Sun, J.M.; Davie, J.R.; Seto, E.; Eisenman, R.N. Histone deacetylases associated with the mSin3 corepressor mediate mad transcriptional repression. Cell 1997, 89, 349–356. [Google Scholar] [CrossRef]
- Cowley, S.M.; Iritani, B.M.; Mendrysa, S.M.; Xu, T.; Cheng, P.F.; Yada, J.; Liggitt, H.D.; Eisenman, R.N. The mSin3A chromatin-modifying complex is essential for embryogenesis and T-cell development. Mol. Cell. Biol. 2005, 25, 6990–7004. [Google Scholar] [CrossRef]
- Dannenberg, J.-H.; David, G.; Zhong, S.; van der Torre, J.; Wong, W.H.; Depinho, R.A. mSin3A corepressor regulates diverse transcriptional networks governing normal and neoplastic growth and survival. Genes Dev. 2005, 19, 1581–1595. [Google Scholar] [CrossRef] [PubMed]
- Yao, C.; Carraro, G.; Konda, B.; Guan, X.; Mizuno, T.; Chiba, N.; Kostelny, M.; Kurkciyan, A.; David, G.; McQualter, J.L.; et al. Sin3a regulates epithelial progenitor cell fate during lung development. Development 2017, 144, 2618–2628. [Google Scholar] [CrossRef] [PubMed]
- Bansal, N.; David, G.; Farias, E.; Waxman, S. Emerging Roles of Epigenetic Regulator Sin3 in Cancer. Adv. Cancer Res. 2016, 130, 113–135. [Google Scholar] [PubMed]
- Kwon, Y.-J.; Petrie, K.; Leibovitch, B.A.; Zeng, L.; Mezei, M.; Howell, L.; Gil, V.; Christova, R.; Bansal, N.; Yang, S.; et al. Selective Inhibition of SIN3 Corepressor with Avermectins as a Novel Therapeutic Strategy in Triple-Negative Breast Cancer. Mol. Cancer Ther. 2015, 14, 1824–1836. [Google Scholar] [CrossRef] [PubMed]
- McDonel, P.; Demmers, J.; Tan, D.W.M.; Watt, F.; Hendrich, B.D. Sin3a is essential for the genome integrity and viability of pluripotent cells. Dev. Biol. 2012, 363, 62–73. [Google Scholar] [CrossRef]
- Smith, K.T.; Martin-Brown, S.A.; Florens, L.; Washburn, M.P.; Workman, J.L. Deacetylase inhibitors dissociate the histone-targeting ING2 subunit from the Sin3 complex. Chem. Biol. 2010, 17, 65–74. [Google Scholar] [CrossRef]
- Richon, V.M.; Sandhoff, T.W.; Rifkind, R.A.; Marks, P.A. Histone deacetylase inhibitor selectively induces p21WAF1 expression and gene-associated histone acetylation. Proc. Natl. Acad. Sci. USA 2000, 97, 10014–10019. [Google Scholar] [CrossRef]
- Larrieu, D.; Ythier, D.; Brambilla, C.; Pedeux, R. ING2 controls the G1 to S-phase transition by regulating p21 expression. Cell Cycle 2010, 9, 3984–3990. [Google Scholar] [CrossRef]
- Jones, P.A.; Issa, J.-P.J.; Baylin, S. Targeting the cancer epigenome for therapy. Nat. Rev. Genet. 2016, 17, 630–641. [Google Scholar] [CrossRef]
- Loewith, R.; Meijer, M.; Lees-Miller, S.P.; Riabowol, K.; Young, D. Three yeast proteins related to the human candidate tumor suppressor p33(ING1) are associated with histone acetyltransferase activities. Mol. Cell. Biol. 2000, 20, 3807–3816. [Google Scholar] [CrossRef] [PubMed]
- Kichina, J.V.; Zeremski, M.; Aris, L.; Gurova, K.V.; Walker, E.; Franks, R.; Nikitin, A.Y.; Kiyokawa, H.; Gudkov, A.V. Targeted disruption of the mouse ing1 locus results in reduced body size, hypersensitivity to radiation and elevated incidence of lymphomas. Oncogene 2006, 25, 857–866. [Google Scholar] [CrossRef] [PubMed]
- Roca, M.S.; Di Gennaro, E.; Budillon, A. Implication for Cancer Stem Cells in Solid Cancer Chemo-Resistance: Promising Therapeutic Strategies Based on the Use of HDAC Inhibitors. J. Clin. Med. 2019, 8, 912. [Google Scholar] [CrossRef] [PubMed]
- Patra, S.; Panigrahi, D.P.; Praharaj, P.P.; Bhol, C.S.; Mahapatra, K.K.; Mishra, S.R.; Behera, B.P.; Jena, M.; Bhutia, S.K. Dysregulation of histone deacetylases in carcinogenesis and tumor progression: A possible link to apoptosis and autophagy. Cell. Mol. Life Sci. 2019, 76, 3263–3282. [Google Scholar] [CrossRef]
- Eckschlager, T.; Plch, J.; Stiborova, M.; Hrabeta, J. Histone Deacetylase Inhibitors as Anticancer Drugs. Int. J. Mol. Sci. 2017, 18, 1414. [Google Scholar] [CrossRef]

| Tissue Type | Origin | Mutation Type/Expression Change | Methods | Position | Coding | Frequency | Ref. |
|---|---|---|---|---|---|---|---|
| Lung cancer | Cell lines | Downregulation | RT-QPCR | 7/8 | [49] | ||
| Patient | Substitution | PCR-SSCP, Sequencing | LZL (13) | Ala -> Ala | 6/31 | ||
| Patient | Substitution | PCR-SSCP, Sequencing | 6bp downstream exon 1 | 6/31 | |||
| Lung cancer | Cell lines | Downregulation | Q-PCR | 2/2 | [50] | ||
| Lung cancer | Patient | Downregulation | IHC | 70/120 | [62] | ||
| Patient | No LOH | MM | 0/12 | ||||
| Patient | Substitution | Sequencing | 39 | Ala -> Ala | 21/22 | ||
| Patient | Downregulation | Q-PCR | 15/22 | ||||
| No change | Q-PCR | 6/22 | |||||
| Upregulation | Q-PCR | 1/22 | |||||
| NSCLC | Patient | Downregulation, aberrantly localization | IHC, RT-PCR, WB | 21/64 (32.8%) | [63] | ||
| Adenocarcinoma | Patient | Downregulation, aberrantly localization | IHC, RT-PCR, WB | 11/24 (45.8%) | |||
| Squamous cell carcinoma | Patient | Downregulation, aberrantly localization | IHC, RT-PCR, WB | 10/38 (26.3%) | |||
| NSCLC | Patient | Chromosomal deletion | cDNA Microarray | 4q34.2–q35.1 | 2/10 (20%) | [64] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Blondel, A.; Benberghout, A.; Pedeux, R.; Ricordel, C. Exploiting ING2 Epigenetic Modulation as a Therapeutic Opportunity for Non-Small Cell Lung Cancer. Cancers 2019, 11, 1601. https://doi.org/10.3390/cancers11101601
Blondel A, Benberghout A, Pedeux R, Ricordel C. Exploiting ING2 Epigenetic Modulation as a Therapeutic Opportunity for Non-Small Cell Lung Cancer. Cancers. 2019; 11(10):1601. https://doi.org/10.3390/cancers11101601
Chicago/Turabian StyleBlondel, Alice, Amine Benberghout, Rémy Pedeux, and Charles Ricordel. 2019. "Exploiting ING2 Epigenetic Modulation as a Therapeutic Opportunity for Non-Small Cell Lung Cancer" Cancers 11, no. 10: 1601. https://doi.org/10.3390/cancers11101601
APA StyleBlondel, A., Benberghout, A., Pedeux, R., & Ricordel, C. (2019). Exploiting ING2 Epigenetic Modulation as a Therapeutic Opportunity for Non-Small Cell Lung Cancer. Cancers, 11(10), 1601. https://doi.org/10.3390/cancers11101601





