Epigenetic Regulation by lncRNAs: An Overview Focused on UCA1 in Colorectal Cancer
Abstract
1. Colorectal Cancer
2. Mechanisms of Regulation by Long Non-Coding RNAs in Cancer Cells
2.1. Interaction of lncRNAs with DNA
2.1.1. DNA Organization
2.1.2. DNA Transcription
2.2. Interaction of lncRNAs with RNA
2.3. Interaction of lncRNAs with Proteins
3. UCA1 Expression in Colorectal Cancer
3.1. Association of UCA1 Transcript Expression with Colorectal Cancer
3.2. Regulation of UCA1 Transcript Expression
4. UCA1-Mechanism of Regulation
4.1. UCA1-Regulated Transcription
4.2. UCA1 and miRNA-Mediated Decay
5. Role of UCA1-Mediated Regulation in Colorectal Cancer
5.1. UCA1-Mediated Regulation of the Cell Cycle
5.2. Association of UCA1 with Chemoresistance
5.3. UCA1 in Colorectal Cancer Diagnosis and Therapy
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- American Cancer Society. Cancer Facts and Figures. 2018. Available online: https://www.cancer.org (accessed on 1 June 2018).
- Malvezzi, M.; Carioli, G.; Bertuccio, P.; Boffetta, P.; Levi, F.; La Vecchia, C.; Negri, E. European cancer mortality predictions for the year 2018 with focus on colorectal cancer. Ann. Oncol. 2018, 29, 1016–1022. [Google Scholar] [CrossRef] [PubMed]
- WCRF/AICR. Continuous Update Project Report: Food, Nutrition, Physical Activity, and the Prevention of Colorectal Cancer. 2011. Available online: https://www.wcrf.org/sites/default/files/Colorectal-Cancer-2011-Report.pdf (accessed on 1 June 2018).
- Rank, K.M.; Shaukat, A. Stool Based Testing for Colorectal Cancer: An Overview of Available Evidence. Curr. Gastroenterol. Rep. 2017, 19. [Google Scholar] [CrossRef] [PubMed]
- Rahbari, N.N.; Carr, P.R.; Jansen, L.; Chang-Claude, J.; Weitz, J.; Hoffmeister, M.; Brenner, H. Time of Metastasis and Outcome in Colorectal Cancer. Ann. Surg. 2017, 1. [Google Scholar] [CrossRef] [PubMed]
- Akinyemiju, T.; Sakhuja, S.; Waterbor, J.; Pisu, M.; Altekruse, S.F. Racial/ethnic disparities in de novo metastases sites and survival outcomes for patients with primary breast, colorectal, and prostate cancer. Cancer Med. 2018, 7, 1183–1193. [Google Scholar] [CrossRef] [PubMed]
- Luo, Q.; O’Connell, D.L.; Kahn, C.; Yu, X.Q. Colorectal cancer metastatic disease progression in Australia: A population-based analysis. Cancer Epidemiol. 2017, 49, 92–100. [Google Scholar] [CrossRef] [PubMed]
- Benson, A.R.; Venook, A.; Cederquist, L.; Chan, E.; Chen, Y.; Cooper, H.; Deming, D.; Engstrom, P.; Enzinger, P.; Fichera, A.; et al. Clinical Practice Guidelines in Oncology. Natl. Compr. Cancer Netw. 2017, 3, 370–398. [Google Scholar] [CrossRef]
- Puccini, A.; Berger, M.D.; Naseem, M.; Tokunaga, R.; Battaglin, F.; Cao, S.; Hanna, D.L.; McSkane, M.; Soni, S.; Zhang, W.; et al. Colorectal cancer: Epigenetic alterations and their clinical implications. Biochim. Biophys. Acta Rev. Cancer 2017, 1868, 439–448. [Google Scholar] [CrossRef] [PubMed]
- Gupta, R.; Sinha, S.; Paul, R.N. The impact of microsatellite stability status in colorectal cancer. Curr. Probl. Cancer 2018. [Google Scholar] [CrossRef] [PubMed]
- Sadanandam, A.; Lyssiotis, C.A.; Homicsko, K.; Collisson, E.A.; Gibb, W.J.; Wullschleger, S.; Ostos, L.C.G.; Lannon, W.A.; Grotzinger, C.; Del Rio, M.; et al. A colorectal cancer classification system that associates cellular phenotype and responses to therapy. Nat. Med. 2013, 19, 619–625. [Google Scholar] [CrossRef] [PubMed]
- Guinney, J.; Dienstmann, R.; Wang, X.; de Reyniès, A.; Schlicker, A.; Soneson, C.; Marisa, L.; Roepman, P.; Nyamundanda, G.; Angelino, P.; et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 2015, 21, 1350–1356. [Google Scholar] [CrossRef] [PubMed]
- Merkel, S.; Schellerer, V.S.; Wein, A.; Semrau, S.; Geppert, C.; Göhl, J.; Hohenberger, W.; Weber, K.; Grützmann, R. The influence of tumour site on prognosis in metastatic colorectal carcinomas with primary tumour resection. Int. J. Color. Dis. 2018, 33, 1215–1223. [Google Scholar] [CrossRef] [PubMed]
- Cremolini, C.; Antoniotti, C.; Lonardi, S.; Bergamo, F.; Cortesi, E.; Tomasello, G.; Moretto, R.; Ronzoni, M.; Racca, P.; Loupakis, F.; et al. Primary Tumor Sidedness and Benefit from FOLFOXIRI plus Bevacizumab as Initial Therapy for Metastatic Colorectal Cancer. Ann. Oncol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Narayanan, S.; Gabriel, E.; Attwood, K.; Boland, P.; Nurkin, S. Association of Clinicopathologic and Molecular Markers on Stage-specific Survival of Right Versus Left Colon Cancer. Clin. Color. Cancer 2018, 5. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2018. CA Cancer J. Clin. 2018, 68, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Tao, Y.; Li, Y.; Zhao, J.; Zhang, L.; Zhang, X.; Dong, C.; Xie, Y.; Dai, X.; Zhang, X.; et al. The regulatory network analysis of long noncoding RNAs in human colorectal cancer. Funct. Integr. Genom. 2018, 18, 261–275. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Sun, Q.; Cheng, X.; Zhang, L.; Song, W.; Zhou, D.; Lin, J.; Wang, W. Genome-wide analysis of long noncoding RNA (lncRNA) expression in colorectal cancer tissues from patients with liver metastasis. Cancer Med. 2016, 5, 1629–1639. [Google Scholar] [CrossRef] [PubMed]
- De Bony, E.J.; Bizet, M.; Van Grembergen, O.; Hassabi, B.; Calonne, E.; Putmans, P.; Bontempi, G.; Fuks, F. Comprehensive identification of long noncoding RNAs in colorectal cancer. Oncotarget 2018, 9, 27605–27629. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Li, N.; Lao, Y.; Lin, W.; Jiang, G.; Wei, N.; Wang, C.; Liu, K.; Wu, J. Variable Levels of Long Noncoding RNA Expression in DNA Mismatch Repair-Proficient Early-Stage Colon Cancer. Dig. Dis. Sci. 2017, 62, 1235–1245. [Google Scholar] [CrossRef] [PubMed]
- Kawaguchi, T.; Hirose, T. Chromatin remodeling complexes in the assembly of long noncoding RNA-dependent nuclear bodies. Nucleus 2015, 6, 462–467. [Google Scholar] [CrossRef] [PubMed]
- Sawyer, I.A.; Dundr, M. Chromatin loops and causality loops: The influence of RNA upon spatial nuclear architecture. Chromosoma 2017, 126, 541–557. [Google Scholar] [CrossRef] [PubMed]
- Clemson, C.M.; Hutchinson, J.N.; Sara, S.A.; Ensminger, A.W.; Fox, A.H.; Chess, A.; Lawrence, J.B. An architectural role for a nuclear noncoding RNA: NEAT1 RNA is essential for the structure of paraspeckles. Mol. Cell. 2009, 33, 717–726. [Google Scholar] [CrossRef] [PubMed]
- Naganuma, T.; Hirose, T. Paraspeckle formation during the biogenesis of long non-coding RNAs. RNA Biol. 2013, 10, 456–461. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Chen, W.; He, F.; Tan, Z.; Zheng, J.; Wang, W.; Zhao, Q.; Li, J. NEAT expression is associated with tumor recurrence and unfavorable prognosis in colorectal cancer. Oncotarget 2015, 6, 27641–27650. [Google Scholar] [CrossRef] [PubMed]
- Quinn, J.J.; Chang, H.Y. Unique features of long non-coding RNA biogenesis and function. Nat. Rev. Genet. 2016, 17, 47–62. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Bao, C.; Gu, S.; Ye, D.; Jing, F.; Fan, C.; Jin, M.; Chen, K. Associations between novel genetic variants in the promoter region of MALAT1 and risk of colorectal cancer. Oncotarget 2017, 8, 92604–92614. [Google Scholar] [CrossRef] [PubMed]
- Khalil, A.M.; Guttman, M.; Huarte, M.; Garber, M.; Raj, A.; Rivea Morales, D.; Thomas, K.; Presser, A.; Bernstein, B.E.; van Oudenaarden, A.; et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc. Natl. Acad. Sci. USA 2009, 106, 11667–11672. [Google Scholar] [CrossRef] [PubMed]
- Davidovich, C.; Cech, T.R. The recruitment of chromatin modifiers by long noncoding RNAs: Lessons from PRC2. RNA 2015, 21, 2007–2022. [Google Scholar] [CrossRef] [PubMed]
- Ma, Z.; Peng, P.; Zhou, J.; Hui, B.; Ji, H.; Wang, J.; Wang, K. Long Non-Coding RNA SH3PXD2A-AS1 Promotes Cell Progression Partly Through Epigenetic Silencing P57 and KLF2 in Colorectal Cancer. Cell Physiol. Biochem. 2018, 46, 2197–2214. [Google Scholar] [CrossRef] [PubMed]
- Ding, J.; Xie, M.; Lian, Y.; Zhu, Y.; Peng, P.; Wang, J.; Wang, L.; Wang, K. Long noncoding RNA HOXA-AS2 represses P21 and KLF2 expression transcription by binding with EZH2, LSD1 in colorectal cancer. Oncogenesis 2017, 6, e288. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Cheng, C. Long noncoding RNA NEAT1 promotes the metastasis of osteosarcoma via interaction with the G9a-DNMT1-Snail complex. Am. J. Cancer Res. 2018, 8, 81–90. [Google Scholar] [PubMed]
- Tang, Y.; Wang, J.; Lian, Y.; Fan, C.; Zhang, P.; Wu, Y.; Li, X.; Xiong, F.; Li, X.; Li, G.; et al. Linking long non-coding RNAs and SWI/SNF complexes to chromatin remodeling in cancer. Mol. Cancer 2017, 16. [Google Scholar] [CrossRef] [PubMed]
- Prensner, J.R.; Iyer, M.K.; Sahu, A.; Asangani, I.A.; Cao, Q.; Patel, L.; Vergara, I.A.; Davicioni, E.; Erho, N.; Ghadessi, M.; et al. The long noncoding RNA SChLAP1 promotes aggressive prostate cancer and antagonizes the SWI/SNF complex. Nat. Genet. 2013, 45, 1392–1398. [Google Scholar] [CrossRef] [PubMed]
- Cheng, S.; Wang, L.; Deng, C.H.; Du, S.C.; Han, Z.G. ARID1A represses hepatocellular carcinoma cell proliferation and migration through lncRNA MVIH. Biochem. Biophys. Res. Commun. 2017, 491, 178–182. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.; Zhong, G.; Wu, J.; Chen, H.; Jia, Y. Long noncoding RNA AFAP1-AS1 facilitates tumor growth through enhancer of zeste homolog 2 in colorectal cancer. Am. J. Cancer Res. 2018, 8, 892–902. [Google Scholar] [PubMed]
- Su, J.; Zhang, E.; Han, L.; Yin, D.; Liu, Z.; He, X.; Zhang, Y.; Lin, F.; Lin, Q.; Mao, P.; et al. Long noncoding RNA BLACAT1 indicates a poor prognosis of colorectal cancer and affects cell proliferation by epigenetically silencing of p15. Cell Death Dis. 2017, 8, e2665. [Google Scholar] [CrossRef] [PubMed]
- Ding, J.; Li, J.; Wang, H.; Tian, Y.; Xie, M.; He, X.; Ji, H.; Ma, Z.; Hui, B.; Wang, K.; et al. Long noncoding RNA CRNDE promotes colorectal cancer cell proliferation via epigenetically silencing DUSP5/CDKN1A expression. Cell Death Dis. 2017, 8, e2997. [Google Scholar] [CrossRef] [PubMed]
- Kogo, R.; Shimamura, T.; Mimori, K.; Kawahara, K.; Imoto, S.; Sudo, T.; Tanaka, F.; Shibata, K.; Suzuki, A.; Komune, S.; et al. Long noncoding RNA HOTAIR regulates polycomb-dependent chromatin modification and is associated with poor prognosis in colorectal cancers. Cancer Res. 2011, 71, 6320–6326. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.J.; Huang, C.Q.; Peng, C.W.; Hou, J.X.; Liu, J.Y. Long noncoding RNA HULC promotes colorectal carcinoma progression through epigenetically repressing NKD2 expression. Gene 2016, 592, 172–178. [Google Scholar] [CrossRef] [PubMed]
- Marin-Bejar, O.; Marchese, F.P.; Athie, A.; Sanchez, Y.; Gonzalez, J.; Segura, V.; Huang, L.; Moreno, I.; Navarro, A.; Monzo, M.; et al. Pint lincRNA connects the p53 pathway with epigenetic silencing by the Polycomb repressive complex 2. Genome Biol. 2013, 14, R104. [Google Scholar] [CrossRef] [PubMed]
- Ma, Z.; Gu, S.; Song, M.; Yan, C.; Hui, B.; Ji, H.; Wang, J.; Zhang, J.; Wang, K.; Zhao, Q. Long non-coding RNA SNHG17 is an unfavourable prognostic factor and promotes cell proliferation by epigenetically silencing P57 in colorectal cancer. Mol. Biosyst. 2017, 13, 2350–2361. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.-Q.; Cai, Q.; Hu, L.; He, C.-Y.; Li, J.-F.; Quan, Z.-W.; Liu, B.-Y.; Li, C.; Zhu, Z.-G. Long noncoding RNA UCA1 induced by SP1 promotes cell proliferation via recruiting EZH2 and activating AKT pathway in gastric cancer. Cell Death Dis. 2017, 8, e2839. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.-J.; Song, W.; Zhang, S.-D.; Shen, X.-H.; Qiu, X.-M.; Wu, H.-Z.; Gong, P.-H.; Lu, S.; Zhao, Z.-J.; He, M.-L.; et al. HBx-upregulated lncRNA UCA1 promotes cell growth and tumorigenesis by recruiting EZH2 and repressing p27Kip1/CDK2 signaling. Sci. Rep. 2016, 6, 23521. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.C.; Yang, Y.W.; Liu, B.; Sanyal, A.; Corces-Zimmerman, R.; Chen, Y.; Lajoie, B.R.; Protacio, A.; Flynn, R.A.; Gupta, R.A.; et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 2011, 472, 120–124. [Google Scholar] [CrossRef] [PubMed]
- Sun, T.T.; He, J.; Liang, Q.; Ren, L.L.; Yan, T.T.; Yu, T.C.; Tang, J.Y.; Bao, Y.J.; Hu, Y.; Lin, Y.; et al. LncRNA GClnc1 Promotes Gastric Carcinogenesis and May Act as a Modular Scaffold of WDR5 and KAT2A Complexes to Specify the Histone Modification Pattern. Cancer Discov. 2016, 6, 784–801. [Google Scholar] [CrossRef] [PubMed]
- Gu, P.; Chen, X.; Xie, R.; Han, J.; Xie, W.; Wang, B.; Dong, W.; Chen, C.; Yang, M.; Jiang, J.; et al. lncRNA HOXD-AS1 Regulates Proliferation and Chemo-Resistance of Castration-Resistant Prostate Cancer via Recruiting WDR5. Mol. Ther. 2017. [Google Scholar] [CrossRef] [PubMed]
- Xia, M.; Yao, L.; Zhang, Q.; Wang, F.; Mei, H.; Guo, X.; Huang, W. Long noncoding RNA HOTAIR promotes metastasis of renal cell carcinoma by up-regulating histone H3K27 demethylase JMJD3. Oncotarget 2017, 8, 19795–19802. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Ge, D.; Chen, X.; Qiu, J.; Yin, Z.; Zheng, S.; Jiang, C. FOXP4-AS1 participates in the development and progression of osteosarcoma by downregulating LATS1 via binding to LSD1 and EZH2. Biochem. Biophys. Res. Commun. 2018, 502, 493–500. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; He, L.; Du, Y.; Zhu, P.; Huang, G.; Luo, J.; Yan, X.; Ye, B.; Li, C.; Xia, P.; et al. The long noncoding RNA lncTCF7 promotes self-renewal of human liver cancer stem cells through activation of Wnt signaling. Cell Stem Cell 2015, 16, 413–425. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Gao, Y.; Yao, L.; Liu, Y.; Huang, L.; Yan, Z.; Zhao, W.; Zhu, P.; Weng, H. LncFZD6 initiates Wnt/beta-catenin and liver TIC self-renewal through BRG1-mediated FZD6 transcriptional activation. Oncogene 2018, 37, 3098–3112. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Gong, Y.; Jin, B.O.; Wu, C.; Yang, J.; Wang, L.E.; Zhang, Z.; Mao, Z. Long non-coding RNA urothelial carcinoma associated 1 induces cell replication by inhibiting BRG1 in 5637 cells. Oncol. Rep. 2014, 32, 1281–1290. [Google Scholar] [CrossRef] [PubMed]
- Tsang, W.P.; Ng, E.K.; Ng, S.S.; Jin, H.; Yu, J.; Sung, J.J.; Kwok, T.T. Oncofetal H19-derived miR-675 regulates tumor suppressor RB in human colorectal cancer. Carcinogenesis 2010, 31, 350–358. [Google Scholar] [CrossRef] [PubMed]
- Matouk, I.J.; Raveh, E.; Abu-lail, R.; Mezan, S.; Gilon, M.; Gershtain, E.; Birman, T.; Gallula, J.; Schneider, T.; Barkali, M.; et al. Oncofetal H19 RNA promotes tumor metastasis. Biochim. Biophys. Acta 2014, 7, 1414–1426. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Li, T.; Qi, J.; Liu, J.; Qin, C. The miR-545/374a cluster encoded in the Ftx lncRNA is overexpressed in HBV-related hepatocellular carcinoma and promotes tumorigenesis and tumor progression. PLoS ONE 2014, 9, e109782. [Google Scholar] [CrossRef] [PubMed]
- Iio, A.; Takagi, T.; Miki, K.; Naoe, T.; Nakayama, A.; Akao, Y. DDX6 post-transcriptionally down-regulates miR-143/145 expression through host gene NCR143/145 in cancer cells. Biochim. Biophys. Acta 2013, 10, 1102–1110. [Google Scholar] [CrossRef] [PubMed]
- Augoff, K.; McCue, B.; Plow, E.F.; Sossey-Alaoui, K. miR-31 and its host gene lncRNA LOC554202 are regulated by promoter hypermethylation in triple-negative breast cancer. Mol. Cancer 2012, 11, 5. [Google Scholar] [CrossRef] [PubMed]
- Emmrich, S.; Streltsov, A.; Schmidt, F.; Thangapandi, V.R.; Reinhardt, D.; Klusmann, J.H. LincRNAs MONC and MIR100HG act as oncogenes in acute megakaryoblastic leukemia. Mol. Cancer 2014, 13, 1476–4598. [Google Scholar] [CrossRef] [PubMed]
- Sun, D.; Layer, R.; Mueller, A.C.; Cichewicz, M.A.; Negishi, M.; Paschal, B.M.; Dutta, A. Regulation of several androgen-induced genes through the repression of the miR-99a/let-7c/miR-125b-2 miRNA cluster in prostate cancer cells. Oncogene 2014, 33, 1448–1457. [Google Scholar] [CrossRef] [PubMed]
- Kotake, Y.; Kitagawa, K.; Ohhata, T.; Sakai, S.; Uchida, C.; Niida, H.; Naemura, M.; Kitagawa, M. Long Non-coding RNA, PANDA, Contributes to the Stabilization of p53 Tumor Suppressor Protein. Anticancer Res. 2016, 36, 1605–1611. [Google Scholar] [PubMed]
- Yan, C.; Chen, J.; Chen, N. Long noncoding RNA MALAT1 promotes hepatic steatosis and insulin resistance by increasing nuclear SREBP-1c protein stability. Sci. Rep. 2016, 6, 22640. [Google Scholar] [CrossRef] [PubMed]
- Jiang, H.; Li, T.; Qu, Y.; Wang, X.; Li, B.; Song, J.; Sun, X.; Tang, Y.; Wan, J.; Yu, Y.; et al. Long non-coding RNA SNHG15 interacts with and stabilizes transcription factor Slug and promotes colon cancer progression. Cancer Lett. 2018, 425, 78–87. [Google Scholar] [CrossRef] [PubMed]
- Nevers, A.; Doyen, A.; Malabat, C.; Neron, B.; Kergrohen, T.; Jacquier, A.; Badis, G. Antisense transcriptional interference mediates condition-specific gene repression in budding yeast. Nucleic Acids Res. 2018. [Google Scholar] [CrossRef] [PubMed]
- Muniz, L.; Deb, M.K.; Aguirrebengoa, M.; Lazorthes, S.; Trouche, D.; Nicolas, E. Control of Gene Expression in Senescence through Transcriptional Read-Through of Convergent Protein-Coding Genes. Cell Rep. 2017, 21, 2433–2446. [Google Scholar] [CrossRef] [PubMed]
- Pelechano, V.; Steinmetz, L.M. Gene regulation by antisense transcription. Nat. Rev. Genet. 2013, 14, 880–893. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Li, W.; Zhang, Y.; Zhang, X.; Liu, T.; Zhang, Y.; Yang, Y.; Wang, L.; Pan, H.; Ji, J.; et al. Increased expression of antisense lncRNA SPINT1-AS1 predicts a poor prognosis in colorectal cancer and is negatively correlated with its sense transcript. OncoTargets Ther. 2018, 11, 3969–3978. [Google Scholar] [CrossRef] [PubMed]
- Yin, J.; Luo, W.; Zeng, X.; Zeng, L.; Li, Z.; Deng, X.; Tan, X.; Hu, W. UXT-AS1-induced alternative splicing of UXT is associated with tumor progression in colorectal cancer. Am. J. Cancer Res. 2017, 7, 462–472. [Google Scholar] [PubMed]
- Engreitz, J.M.; Haines, J.E.; Perez, E.M.; Munson, G.; Chen, J.; Kane, M.; McDonel, P.E.; Guttman, M.; Lander, E.S. Local regulation of gene expression by lncRNA promoters, transcription and splicing. Nature 2016, 539, 452–455. [Google Scholar] [CrossRef] [PubMed]
- Jalali, S.; Singh, A.; Maiti, S.; Scaria, V. Genome-wide computational analysis of potential long noncoding RNA mediated DNA:DNA:RNA triplexes in the human genome. J. Transl. Med. 2017, 15, 186. [Google Scholar] [CrossRef] [PubMed]
- Amaral, P.P.; Leonardi, T.; Han, N.; Viré, E.; Gascoigne, D.K.; Arias-Carrasco, R.; Büscher, M.; Pandolfini, L.; Zhang, A.; Pluchino, S.; et al. Genomic positional conservation identifies topological anchor point RNAs linked to developmental loci. Genome Biol. 2018, 19, 32. [Google Scholar] [CrossRef] [PubMed]
- Xiang, J.-F.; Yin, Q.-F.; Chen, T.; Zhang, Y.; Zhang, X.-O.; Wu, Z.; Zhang, S.; Wang, H.-B.; Ge, J.; Lu, X.; et al. Human colorectal cancer-specific CCAT1-L lncRNA regulates long-range chromatin interactions at the MYC locus. Cell Res. 2014, 24, 513–531. [Google Scholar] [CrossRef] [PubMed]
- Gui, X.; Li, H.; Li, T.; Pu, H.; Lu, D. Long Noncoding RNA CUDR Regulates HULC and β-Catenin to Govern Human Liver Stem Cell Malignant Differentiation. Mol. Ther. 2015, 23, 1843–1853. [Google Scholar] [CrossRef] [PubMed]
- Mondal, T.; Subhash, S.; Vaid, R.; Enroth, S.; Uday, S.; Reinius, B.; Mitra, S.; Mohammed, A.; James, A.R.; Hoberg, E.; et al. MEG3 long noncoding RNA regulates the TGF-β pathway genes through formation of RNA–DNA triplex structures. Nat. Commun. 2015, 6, 7743. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Liu, Y.; Lin, L.; Huang, Q.; He, W.; Zhang, S.; Dong, S.; Wen, Z.; Rao, J.; Liao, W.; et al. The lncRNA MACC1-AS1 promotes gastric cancer cell metabolic plasticity via AMPK/Lin28 mediated mRNA stability of MACC1. Mol. Cancer 2018, 17, 69. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; Cheng, Y.; Yang, F.; Yao, Z.; Wang, O. Knockdown of LncRNA MAPT-AS1 inhibites proliferation and migration and sensitizes cancer cells to paclitaxel by regulating MAPT expression in ER-negative breast cancers. Cell Biosci. 2018, 8, 7. [Google Scholar] [CrossRef] [PubMed]
- Beltran, M.; Puig, I.; Pena, C.; Garcia, J.M.; Alvarez, A.B.; Pena, R.; Bonilla, F.; de Herreros, A.G. A natural antisense transcript regulates Zeb2/Sip1 gene expression during Snail1-induced epithelial-mesenchymal transition. Genes Dev. 2008, 22, 756–769. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, J.; Lu, Q.; Shen, B.; Huang, X.; Shen, L.; Zheng, X.; Huang, R.; Yan, J.; Guo, H. TGFbeta1 secreted by cancer-associated fibroblasts induces epithelial-mesenchymal transition of bladder cancer cells through lncRNA-ZEB2NAT. Sci. Rep. 2015, 5, 11924. [Google Scholar] [CrossRef] [PubMed]
- Fabian, M.R.; Sonenberg, N. The mechanics of miRNA-mediated gene silencing: A look under the hood of miRISC. Nat. Struct. Mol. Biol. 2012, 19, 586–593. [Google Scholar] [CrossRef] [PubMed]
- Tay, Y.; Rinn, J.; Pandolfi, P.P. The multilayered complexity of ceRNA crosstalk and competition. Nature 2014, 505, 344–352. [Google Scholar] [CrossRef] [PubMed]
- Yamamura, S.; Imai-Sumida, M.; Tanaka, Y.; Dahiya, R. Interaction and cross-talk between non-coding RNAs. Cell Mol. Life Sci. 2017, 75, 467–484. [Google Scholar] [CrossRef] [PubMed]
- Denzler, R.; McGeary, S.E.; Title, A.C.; Agarwal, V.; Bartel, D.P.; Stoffel, M. Impact of MicroRNA Levels, Target-Site Complementarity, and Cooperativity on Competing Endogenous RNA-Regulated Gene Expression. Mol. Cell 2016, 64, 565–579. [Google Scholar] [CrossRef] [PubMed]
- Li, J.-H.; Liu, S.; Zhou, H.; Qu, L.-H.; Yang, J.-H. starBase v2.0: Decoding miRNA-ceRNA, miRNA-ncRNA and protein–RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014, 42, D92–D97. [Google Scholar] [CrossRef] [PubMed]
- Fan, Q.; Liu, B. Comprehensive analysis of a long noncoding RNA-associated competing endogenous RNA network in colorectal cancer. OncoTargets Ther. 2018, 11, 2453–2466. [Google Scholar] [CrossRef] [PubMed]
- Schein, A.; Zucchelli, S.; Kauppinen, S.; Gustincich, S.; Carninci, P. Identification of antisense long noncoding RNAs that function as SINEUPs in human cells. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Hu, G.; Lou, Z.; Gupta, M. The Long Non-Coding RNA GAS5 Cooperates with the Eukaryotic Translation Initiation Factor 4E to Regulate c-Myc Translation. PLoS ONE 2014, 9, e107016. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Haider Ali, M.S.S.; Moran, M. The role of interactions of long non-coding RNAs and heterogeneous nuclear ribonucleoproteins in regulating cellular functions. Biochem. J. 2017, 474, 2925–2935. [Google Scholar] [CrossRef] [PubMed]
- Kong, J.; Sun, W.; Li, C.; Wan, L.; Wang, S.; Wu, Y.; Xu, E.; Zhang, H.; Lai, M. Long non-coding RNA LINC01133 inhibits epithelial–mesenchymal transition and metastasis in colorectal cancer by interacting with SRSF6. Cancer Lett. 2016, 380, 476–484. [Google Scholar] [CrossRef] [PubMed]
- Romero-Barrios, N.; Legascue, M.F.; Benhamed, M.; Ariel, F.; Crespi, M. Splicing regulation by long noncoding RNAs. Nucleic Acids Res. 2018, 46, 2169–2184. [Google Scholar] [CrossRef] [PubMed]
- Villamizar, O.; Chambers, C.B.; Riberdy, J.M.; Persons, D.A.; Wilber, A. Long noncoding RNA Saf and splicing factor 45 increase soluble Fas and resistance to apoptosis. Oncotarget 2016, 7, 13810–13826. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, V.; Ellis, J.D.; Shen, Z.; Song, D.Y.; Pan, Q.; Watt, A.T.; Freier, S.M.; Bennett, C.F.; Sharma, A.; Bubulya, P.A.; et al. The Nuclear-Retained Noncoding RNA MALAT1 Regulates Alternative Splicing by Modulating SR Splicing Factor Phosphorylation. Mol. Cell 2010, 39, 925–938. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Sun, L.; Liu, Q.; Gong, C.; Yao, Y.; Lv, X.; Lin, L.; Yao, H.; Su, F.; Li, D.; et al. A Cytoplasmic NF-κB Interacting Long Noncoding RNA Blocks IκB Phosphorylation and Suppresses Breast Cancer Metastasis. Cancer Cell 2015, 27, 370–381. [Google Scholar] [CrossRef] [PubMed]
- Lin, A.; Li, C.; Xing, Z.; Hu, Q.; Liang, K.; Han, L.; Wang, C.; Hawke, D.H.; Wang, S.; Zhang, Y.; et al. The LINK-A lncRNA activates normoxic HIF1α signalling in triple-negative breast cancer. Nat. Cell Biol. 2016, 18, 213–224. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Orera, J.; Messeguer, X.; Subirana, J.A.; Alba, M.M. Long non-coding RNAs as a source of new peptides. Elife 2014, 3, e03523. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, A.; Nakayama, K.I. Hidden Peptides Encoded by Putative Noncoding RNAs. Cell. Struct. Funct. 2018, 43, 75–83. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, A.; Pasut, A.; Matsumoto, M.; Yamashita, R.; Fung, J.; Monteleone, E.; Saghatelian, A.; Nakayama, K.I.; Clohessy, J.G.; Pandolfi, P.P. mTORC1 and muscle regeneration are regulated by the LINC00961-encoded SPAR polypeptide. Nature 2016, 541, 228–232. [Google Scholar] [CrossRef] [PubMed]
- D’Lima, N.G.; Ma, J.; Winkler, L.; Chu, Q.; Loh, K.H.; Corpuz, E.O.; Budnik, B.A.; Lykke-Andersen, J.; Saghatelian, A.; Slavoff, S.A. A human microprotein that interacts with the mRNA decapping complex. Nat. Chem. Biol. 2016, 13, 174–180. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.-Z.; Chen, M.; Chen, D.; Gao, X.-C.; Zhu, S.; Huang, H.; Hu, M.; Zhu, H.; Yan, G.-R. A Peptide Encoded by a Putative lncRNA HOXB-AS3 Suppresses Colon Cancer Growth. Mol. Cell 2017, 68, 171–184. [Google Scholar] [CrossRef] [PubMed]
- Carlevaro-Fita, J.; Rahim, A.; Guigó, R.; Vardy, L.A.; Johnson, R. Cytoplasmic long noncoding RNAs are frequently bound to and degraded at ribosomes in human cells. RNA 2016, 22, 867–882. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.; Hu, Z.; Feng, Y.; Hu, X.; Yuan, J.; Zhao, S.D.; Zhang, Y.; Yang, L.; Shan, W.; He, Q.; et al. Comprehensive Genomic Characterization of Long Non-coding RNAs across Human Cancers. Cancer Cell 2015, 28, 529–540. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Meng, H.; Bai, Y.; Wang, K. Regulation of lncRNA and Its Role in Cancer Metastasis. Oncol. Res. 2016, 23, 205–217. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.; Croce, C.M. Long noncoding RNAs: Undeciphered cellular codes encrypting keys of colorectal cancer pathogenesis. Cancer Lett. 2018, 417, 89–95. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.; Li, X.; Wang, Y.; Zhao, L.; Chen, W. Long non-coding RNA UCA1 regulated cell cycle distribution via CREB through PI3-K dependent pathway in bladder carcinoma cells. Gene 2012, 496, 8–16. [Google Scholar] [CrossRef] [PubMed]
- Tsang, W.P.; Wong, T.W.L.; Cheung, A.H.H.; Co, C.N.N.; Kwok, T.T. Induction of drug resistance and transformation in human cancer cells by the noncoding RNA CUDR. RNA 2007, 13, 890–898. [Google Scholar] [CrossRef] [PubMed]
- Xue, M.; Chen, W.; Li, X. Urothelial cancer associated 1: A long noncoding RNA with a crucial role in cancer. J. Cancer Res. Clin. Oncol. 2016, 142, 1407–1419. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.; Wan, D.; Zheng, D.; Zheng, Q.; Wu, F.; Zhi, Q. Long non-coding RNA UCA1 promotes the tumorigenesis in pancreatic cancer. Biomed. Pharmacother. 2016, 83, 1220–1226. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Gao, F.; Zhou, L.; Wang, H.; Shi, G.; Tan, X. UCA1 Regulates the Growth and Metastasis of Pancreatic Cancer by Sponging miR-135a. Oncol. Res. 2017, 25, 1529–1541. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Zhao, Y.; Zhang, Y.; Wang, D.; Gu, S.; Feng, W.; Peng, W.; Gong, A.; Xu, M. LncRNA UCA1 promotes migration and invasion in pancreatic cancer cells via the Hippo pathway. Biochim. Biophys. Acta Mol. Basis Dis. 2018, 1864, 1770–1782. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Yang, Y.-N.; Yuan, H.-H.; Zhang, T.-T.; Sui, H.; Wei, X.-L.; Liu, L.; Huang, P.; Zhang, W.-J.; Bai, Y.-X. UCA1, a long non-coding RNA up-regulated in colorectal cancer influences cell proliferation, apoptosis and cell cycle distribution. Pathology 2014, 46, 396–401. [Google Scholar] [CrossRef] [PubMed]
- Ni, B.; Yu, X.; Guo, X.; Fan, X.; Yang, Z.; Wu, P.; Yuan, Z.; Deng, Y.; Wang, J.; Chen, D.; et al. Increased urothelial cancer associated 1 is associated with tumor proliferation and metastasis and predicts poor prognosis in colorectal cancer. Int. J. Oncol. 2015, 47, 1329–1338. [Google Scholar] [CrossRef] [PubMed]
- Bian, Z.; Jin, L.; Zhang, J.; Yin, Y.; Quan, C.; Hu, Y.; Feng, Y.; Liu, H.; Fei, B.; Mao, Y.; et al. LncRNA—UCA1 enhances cell proliferation and 5-fluorouracil resistance in colorectal cancer by inhibiting miR-204-5p. Sci. Rep. 2016, 6, 23892. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Dong, Q.; Gao, H.; Zhu, Z. The prognostic significance of UCA1 for predicting clinical outcome in patients with digestive system malignancies. Oncotarget 2017, 8, 40620. [Google Scholar] [CrossRef] [PubMed]
- Renaud, F.; Vincent, A.; Mariette, C.; Crépin, M.; Stechly, L.; Truant, S.; Copin, M.-C.; Porchet, N.; Leteurtre, E.; Van Seuningen, I.; et al. MUC5AC hypomethylation is a predictor of microsatellite instability independently of clinical factors associated with colorectal cancer. Int. J. Cancer 2015, 136, 2811–2821. [Google Scholar] [CrossRef] [PubMed]
- Aguirre-Gamboa, R.; Gomez-Rueda, H.; Martinez-Ledesma, E.; Martinez-Torteya, A.; Chacolla-Huaringa, R.; Rodriguez-Barrientos, A.; Tamez-Pena, J.G.; Trevino, V. SurvExpress: An online biomarker validation tool and database for cancer gene expression data using survival analysis. PLoS ONE 2013, 8, e74250. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.; Zhang, S.; Li, X.; Xue, M.; Cao, S.; Chen, W. Ets-2 regulates cell apoptosis via the Akt pathway, through the regulation of urothelial cancer associated 1, a long non-coding RNA, in bladder cancer cells. PLoS ONE 2013, 8, e73920. [Google Scholar] [CrossRef] [PubMed]
- Li, G.Y.; Wang, W.; Sun, J.Y.; Xin, B.; Zhang, X.; Wang, T.; Zhang, Q.F.; Yao, L.B.; Han, H.; Fan, D.M.; et al. Long non-coding RNAs AC026904.1 and UCA1: A “one-two punch” for TGF-beta-induced SNAI2 activation and epithelial-mesenchymal transition in breast cancer. Theranostics 2018, 8, 2846–2861. [Google Scholar] [CrossRef] [PubMed]
- Hiemer, S.E.; Szymaniak, A.D.; Varelas, X. The Transcriptional Regulators TAZ and YAP Direct Transforming Growth Factor-induced Tumorigenic Phenotypes in Breast Cancer Cells. J. Biol. Chem. 2014, 289, 13461–13474. [Google Scholar] [CrossRef] [PubMed]
- Gou, L.; Liu, M.; Xia, J.; Wan, Q.; Jiang, Y.; Sun, S.; Tang, M.; Zhou, L.; He, T.; Zhang, Y. BMP9 Promotes the Proliferation and Migration of Bladder Cancer Cells through Up-Regulating lncRNA UCA1. Int. J. Mol. Sci. 2018, 19, 1116. [Google Scholar] [CrossRef] [PubMed]
- Xue, M.; Chen, W.; Xiang, A.; Wang, R.; Chen, H.; Pan, J.; Pang, H.; An, H.; Wang, X.; Hou, H.; et al. Hypoxic exosomes facilitate bladder tumor growth and development through transferring long non-coding RNA-UCA1. Mol. Cancer 2017, 16. [Google Scholar] [CrossRef] [PubMed]
- Xue, M.; Li, X.; Li, Z.; Chen, W. Urothelial carcinoma associated 1 is a hypoxia-inducible factor-1α-targeted long noncoding RNA that enhances hypoxic bladder cancer cell proliferation, migration, and invasion. Tumor Biol. 2014, 35, 6901–6912. [Google Scholar] [CrossRef] [PubMed]
- Xue, M.E.I.; Li, X.U.; Wu, W.; Zhang, S.; Wu, S.; Li, Z.; Chen, W.E.I. Upregulation of long non-coding RNA urothelial carcinoma associated 1 by CCAAT/enhancer binding protein α contributes to bladder cancer cell growth and reduced apoptosis. Oncol. Rep. 2014, 31, 1993–2000. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Zhang, Y.; Mayakonda, A.; Madan, V.; Ding, L.W.; Lin, L.H.; Zia, S.; Gery, S.; Tyner, J.W.; Zhou, W.; et al. ARID1A and CEBPalpha cooperatively inhibit UCA1 transcription in breast cancer. Oncogene 2018. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.-J.; Kim, M.; Kim, H.-P. Epigenetic regulation of long noncoding RNA UCA1 by SATB1 in breast cancer. BMB Rep. 2016, 49, 578–583. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.P.; Emechebe, U.; Smith, R.; Franklin, S.; Moore, B.; Yandell, M.; Lessnick, S.L.; Moon, A.M. Coordinated control of senescence by lncRNA and a novel T-box3 co-repressor complex. Elife 2014, 29, 02805. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Meng, X.; Chen, S.; Li, W.; Li, D.; Singer, R.; Gu, W. IMP1 regulates UCA1-mediated cell invasion through facilitating UCA1 decay and decreasing the sponge effect of UCA1 for miR-122-5p. Breast Cancer Res. 2018, 20. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Yuan, J.; Feng, N.; Li, Y.; Lin, Z.; Jiang, Z.; Gui, Y. Hsa-miR-1 downregulates long non-coding RNA urothelial cancer associated 1 in bladder cancer. Tumour Biol. 2014, 35, 10075–10084. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Zhou, N.; Watabe, K.; Lu, Z.; Wu, F.; Xu, M.; Mo, Y.Y. Long non-coding RNA UCA1 promotes breast tumor growth by suppression of p27 (Kip1). Cell Death Dis. 2014, 5, e1008. [Google Scholar] [CrossRef] [PubMed]
- Corvaisier, M.; Bauzone, M.; Corfiotti, F.; Renaud, F.; Amrani, M.E.; Monté, D.; Truant, S.; Leteurtre, E.; Formstecher, P.; Seuningen, I.V.; et al. Regulation of cellular quiescence by YAP/TAZ and Cyclin E1 in colon cancer cells: Implication in chemoresistance and cancer relapse. Oncotarget 2016, 7, 56699. [Google Scholar] [CrossRef] [PubMed]
- Jeggari, A.; Marks, D.S.; Larsson, E. miRcode: A map of putative microRNA target sites in the long non-coding transcriptome. Bioinformatics 2012, 28, 2062–2063. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Li, X.; Wu, S.; Xue, M.; Chen, W. Long non-coding RNA UCA1 promotes glycolysis by upregulating hexokinase 2 through the mTOR-STAT3/microRNA143 pathway. Cancer Sci. 2014, 105, 951–955. [Google Scholar] [CrossRef] [PubMed]
- Jahangiri, B.; Khalaj-kondori, M.; Asadollahi, E.; Sadeghizadeh, M. Cancer-associated fibroblasts enhance cell proliferation and metastasis of colorectal cancer SW480 cells by provoking long noncoding RNA UCA1. Cell Commun. Signal. 2018. [Google Scholar] [CrossRef] [PubMed]
- Vlachos, I.S.; Zagganas, K.; Paraskevopoulou, M.D.; Georgakilas, G.; Karagkouni, D.; Vergoulis, T.; Dalamagas, T.; Hatzigeorgiou, A.G. DIANA-miRPath v3.0: Deciphering microRNA function with experimental support. Nucleic Acids Res. 2015, 43, 14. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.; Yi, D.; Liu, Y.; Wang, M.; Zhu, Y.; Shi, H. Long nonding RNA UCA1 regulates neural stem cell differentiation by controlling miR-1/Hes1 expression. Am. J. Transl. Res. 2017, 9, 3696–3704. [Google Scholar] [PubMed]
- Yang, Z.; Shi, X.; Li, C.; Wang, X.; Hou, K.; Li, Z.; Zhang, X.; Fan, Y.; Qu, X.; Che, X.; et al. Long non-coding RNA UCA1 upregulation promotes the migration of hypoxia-resistant gastric cancer cells through the miR-7-5p/EGFR axis. Exp. Cell Res. 2018, 368, 194–201. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Liu, Y.; Xie, P.; Li, F.; Li, G. PAX6, a novel target of microRNA-7, promotes cellular proliferation and invasion in human colorectal cancer cells. Dig. Dis. Sci. 2014, 59, 598–606. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.L.; Zhang, Q.; Yuan, X.; Jin, L.; Wang, L.L.; Fang, T.T.; Wang, W.B. Long noncoding RNA RP4 functions as a competing endogenous RNA through miR-7-5p sponge activity in colorectal cancer. World J. Gastroenterol. 2018, 24, 1004–1012. [Google Scholar] [CrossRef] [PubMed]
- Nagano, Y.; Toiyama, Y.; Okugawa, Y.; Imaoka, H.; Fujikawa, H.; Yasuda, H.; Yoshiyama, S.; Hiro, J.; Kobayashi, M.; Ohi, M.; et al. MicroRNA-7 Is Associated with Malignant Potential and Poor Prognosis in Human Colorectal Cancer. Anticancer Res. 2016, 36, 6521–6526. [Google Scholar] [CrossRef] [PubMed]
- Pothoulakis, C.; Torre-Rojas, M.; Duran-Padilla, M.A.; Gevorkian, J.; Zoras, O.; Chrysos, E.; Chalkiadakis, G.; Baritaki, S. CRHR2/Ucn2 signaling is a novel regulator of miR-7/YY1/Fas circuitry contributing to reversal of colorectal cancer cell resistance to Fas-mediated apoptosis. Int. J. Cancer 2018, 142, 334–346. [Google Scholar] [CrossRef] [PubMed]
- Suto, T.; Yokobori, T.; Yajima, R.; Morita, H.; Fujii, T.; Yamaguchi, S.; Altan, B.; Tsutsumi, S.; Asao, T.; Kuwano, H. MicroRNA-7 expression in colorectal cancer is associated with poor prognosis and regulates cetuximab sensitivity via EGFR regulation. Carcinogenesis 2015, 36, 338–345. [Google Scholar] [CrossRef] [PubMed]
- Xu, K.; Chen, Z.; Qin, C.; Song, X. miR-7 inhibits colorectal cancer cell proliferation and induces apoptosis by targeting XRCC2. OncoTargets Ther. 2014, 7, 325–332. [Google Scholar] [CrossRef] [PubMed]
- Zeng, C.Y.; Zhan, Y.S.; Huang, J.; Chen, Y.X. MicroRNA7 suppresses human colon cancer invasion and proliferation by targeting the expression of focal adhesion kinase. Mol. Med. Rep. 2016, 13, 1297–1303. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Li, X.; Wu, C.W.; Dong, Y.; Cai, M.; Mok, M.T.; Wang, H.; Chen, J.; Ng, S.S.; Chen, M.; et al. microRNA-7 is a novel inhibitor of YY1 contributing to colorectal tumorigenesis. Oncogene 2013, 32, 5078–5088. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Jiao, C.; Lin, Y.; Chen, M.; Zhang, J.; Wang, J.; Zhang, Z. lncRNA UCA1 Contributes to Imatinib Resistance by Acting as a ceRNA Against miR-16 in Chronic Myeloid Leukemia Cells. DNA Cell Biol. 2017, 36, 18–25. [Google Scholar] [CrossRef] [PubMed]
- Hasakova, K.; Bezakova, J.; Vician, M.; Reis, R.; Zeman, M.; Herichova, I. Gender-dependent expression of leading and passenger strand of miR-21 and miR-16 in human colorectal cancer and adjacent colonic tissues. Physiol. Res. 2017, 66, S575–S582. [Google Scholar] [PubMed]
- Li, H.J.; Li, X.; Pang, H.; Pan, J.J.; Xie, X.J.; Chen, W. Long non-coding RNA UCA1 promotes glutamine metabolism by targeting miR-16 in human bladder cancer. Jpn. J. Clin. Oncol. 2015, 45, 1055–1063. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.-Y.; Bai, W.-D.; Ye, X.-M.; Yang, A.-G.; Jia, L.-T. Long non-coding RNA UCA1 desensitizes breast cancer cells to trastuzumab by impeding miR-18a repression of Yes-associated protein 1. Biochem. Biophys. Res. Commun. 2018, 496, 1308–1313. [Google Scholar] [CrossRef] [PubMed]
- Yue, J.; Humphreys, K.J.; McKinnon, R.A.; Michael, M.Z. miR-18a Inhibits CDC42 and Plays a Tumour Suppressor Role in Colorectal Cancer Cells. PLoS ONE 2014, 9, e112288. [Google Scholar] [CrossRef]
- Li, X.; Wu, Y.; Liu, A.; Tang, X. Long non-coding RNA UCA1 enhances tamoxifen resistance in breast cancer cells through a miR-18a-HIF1alpha feedback regulatory loop. Tumour Biol. 2016, 37, 14733–14743. [Google Scholar] [CrossRef] [PubMed]
- Fang, Q.; Chen, X.; Zhi, X. Long Non-Coding RNA (LncRNA) Urothelial Carcinoma Associated 1 (UCA1) Increases Multi-Drug Resistance of Gastric Cancer via Downregulating miR-27b. Med. Sci. Monit. 2016, 22, 3506–3513. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.; Yu, S.Y.; Chen, J.J.; Qin, J.; Qiu, Y.E.; Zhong, M.; Chen, M. MiR-27b directly targets Rab3D to inhibit the malignant phenotype in colorectal cancer. Oncotarget 2018, 9, 3830–3841. [Google Scholar] [CrossRef] [PubMed]
- Iino, I.; Kikuchi, H.; Miyazaki, S.; Hiramatsu, Y.; Ohta, M.; Kamiya, K.; Kusama, Y.; Baba, S.; Setou, M.; Konno, H. Effect of miR-122 and its target gene cationic amino acid transporter 1 on colorectal liver metastasis. Cancer Sci. 2013, 104, 624–630. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Liu, Y.; Xu, X. Knockdown of LncRNA-UCA1 suppresses chemoresistance of pediatric AML by inhibiting glycolysis through the microRNA-125a/hexokinase 2 pathway. J. Cell. Biochem. 2018, 119, 6296–6308. [Google Scholar] [CrossRef] [PubMed]
- Sun, M.D.; Zheng, Y.Q.; Wang, L.P.; Zhao, H.T.; Yang, S. Long noncoding RNA UCA1 promotes cell proliferation, migration and invasion of human leukemia cells via sponging miR-126. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 2233–2245. [Google Scholar] [PubMed]
- Liu, Q.; Li, Y.; Lv, W.; Zhang, G.; Tian, X.; Li, X.; Cheng, H.; Zhu, C. UCA1 promotes cell proliferation and invasion and inhibits apoptosis through regulation of the miR129-SOX4 pathway in renal cell carcinoma. OncoTargets Ther. 2018, 11, 2475–2487. [Google Scholar] [CrossRef] [PubMed]
- Fesler, A.; Zhai, H.; Ju, J. miR-129 as a novel therapeutic target and biomarker in gastrointestinal cancer. OncoTargets Ther. 2014, 7, 1481–1485. [Google Scholar]
- Karaayvaz, M.; Zhai, H.; Ju, J. miR-129 promotes apoptosis and enhances chemosensitivity to 5-fluorouracil in colorectal cancer. Cell Death Dis. 2013, 6, 193. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Shi, Z.; Yu, Z.; He, Z. LncRNA HIF1A-AS2 positively affects the progression and EMT formation of colorectal cancer through regulating miR-129-5p and DNMT3A. Biomed. Pharmacother. 2018, 98, 433–439. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Qian, J.; Li, C.; Kwok, L.; Cheng, F.; Liu, P.; Perdomo, C.; Kotton, D.; Vaziri, C.; Anderlind, C.; et al. miR-129 regulates cell proliferation by downregulating Cdk6 expression. Cell Cycle 2010, 9, 1809–1818. [Google Scholar] [CrossRef] [PubMed]
- Wu, N.; Fesler, A.; Liu, H.; Ju, J. Development of novel miR-129 mimics with enhanced efficacy to eliminate chemoresistant colon cancer stem cells. Oncotarget 2017, 9, 8887–8897. [Google Scholar] [CrossRef] [PubMed]
- Ya, G.; Wang, H.; Ma, Y.; Hu, A.; Hu, J.; Yu, Y. Serum miR-129 functions as a biomarker for colorectal cancer by targeting estrogen receptor (ER) beta. Pharmazie 2017, 72, 107–112. [Google Scholar] [PubMed]
- Wang, J.; Ye, C.; Liu, J.; Hu, Y. UCA1 confers paclitaxel resistance to ovarian cancer through miR-129/ABCB1 axis. Biochem. Biophys. Res. Commun. 2018, 501, 1034–1040. [Google Scholar] [CrossRef] [PubMed]
- Nagel, R.; le Sage, C.; Diosdado, B.; van der Waal, M.; Oude Vrielink, J.A.F.; Bolijn, A.; Meijer, G.A.; Agami, R. Regulation of the Adenomatous Polyposis Coli Gene by the miR-135 Family in Colorectal Cancer. Cancer Res. 2008, 68, 5795–5802. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Hou, Z.; Li, D. Long noncoding RNA UCA1 promotes anaplastic thyroid cancer cell proliferation via miR-135a mediated cMYC activation. Mol. Med. Rep. 2018, 18, 3068–3076. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, A.; Gomes, S.E.; Pereira, D.M.; Roma-Rodrigues, C.; Fernandes, A.R.; Borralho, P.M.; Rodrigues, C.M.P. Convergence of miR-143 overexpression, oxidative stress and cell death in HCT116 human colon cancer cells. PLoS ONE 2018, 13, e0191607. [Google Scholar] [CrossRef]
- Tuo, Y.-L.; Li, X.-M.; Luo, J. Long noncoding RNA UCA1 modulates breast cancer cell growth and apoptosis through decreasing tumor suppressive miR-143. Eur. Rev. Med. Pharm. Sci. 2015, 19, 3403–3411. [Google Scholar]
- Tian, S.; Zhao, W.; Yang, D.; Yu, Y.; Zou, J.; Liu, Z.; Du, Z. Atorvastatin inhibits miR-143 expression: A protective mechanism against oxidative stress in cardiomyocytes. Int. J. Cardiol. 2016, 211, 115–118. [Google Scholar] [CrossRef] [PubMed]
- Luo, J.; Chen, J.; Li, H.; Yang, Y.; Yun, H.; Yang, S.; Mao, X. LncRNA UCA1 promotes the invasion and EMT of bladder cancer cells by regulating the miR-143/HMGB1 pathway. Oncol. Lett. 2017, 14, 5556–5562. [Google Scholar] [CrossRef] [PubMed]
- Xue, M.; Pang, H.; Li, X.; Li, H.; Pan, J.; Chen, W. Long non-coding RNA urothelial cancer-associated 1 promotes bladder cancer cell migration and invasion by way of the hsa-miR-145-ZEB1/2-FSCN1 pathway. Cancer Sci. 2016, 107, 18–27. [Google Scholar] [CrossRef] [PubMed]
- Yuan, F.; Sun, R.; Li, L.; Jin, B.; Wang, Y.; Liang, Y.; Che, G.; Gao, L.; Zhang, L. A functional variant rs353292 in the flanking region of miR-143/145 contributes to the risk of colorectal cancer. Sci. Rep. 2016, 6, 30195. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Wu, X.; Xu, Y.; Wu, S.; Li, Z.; Chen, R.; Huang, N.; Zhu, Z.; Xu, X. miR-145 suppresses colorectal cancer cell migration and invasion by targeting an ETS-related gene. Oncol. Rep. 2016, 36, 1917–1926. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; Wang, Y.; Huang, G.; Wang, Q.; Zhao, D.; Chen, L. The lncRNA UCA1 interacts with miR-182 to modulate glioma proliferation and migration by targeting iASPP. Arch. Biochem. Biophys. 2017, 623–624, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.U.; Wang, X.; Wang, Z.; Tang, H.U.I.; Fan, H.; Guo, Q. miR-182 promotes cell growth and invasion by targeting forkhead box F2 transcription factor in colorectal cancer. Oncol. Rep. 2015, 33, 2592–2598. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; You, C.; Zhao, D. Long non-coding RNA UCA1/miR-182/PFKFB2 axis modulates glioblastoma-associated stromal cells-mediated glycolysis and invasion of glioma cells. Biochem. Biophys. Res. Commun. 2018, 500, 569–576. [Google Scholar] [CrossRef] [PubMed]
- Fang, Z.; Zhao, J.; Xie, W.; Sun, Q.; Wang, H.; Qiao, B. LncRNA UCA1 promotes proliferation and cisplatin resistance of oral squamous cell carcinoma by surpressing miR-184 expression. Cancer Med. 2017, 6, 2897–2908. [Google Scholar] [CrossRef] [PubMed]
- Wu, G.; Liu, J.; Wu, Z.; Wu, X.; Yao, X. MicroRNA-184 inhibits cell proliferation and metastasis in human colorectal cancer by directly targeting IGF-1R. Oncol. Lett. 2017, 14, 3215–3222. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Zhou, C. Long non-coding RNA UCA1 promotes lung cancer cell proliferation and migration via microRNA-193a/HMGB1 axis. Biochem. Biophys. Res. Commun. 2018, 496, 738–745. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Ji, D.B.; Han, H.B.; Shi, Y.F.; Du, C.Z.; Gu, J. Downregulation of miR-193a-5p correlates with lymph node metastasis and poor prognosis in colorectal cancer. World J. Gastroenterol. 2014, 20, 12241–12248. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, H.; Takahashi, M.; Ohnuma, S.; Unno, M.; Yoshino, Y.; Ouchi, K.; Takahashi, S.; Yamada, Y.; Shimodaira, H.; Ishioka, C. microRNA-193a-3p is specifically down-regulated and acts as a tumor suppressor in BRAF-mutated colorectal cancer. BMC Cancer 2017, 17, 723. [Google Scholar] [CrossRef] [PubMed]
- Pekow, J.; Meckel, K.; Dougherty, U.; Huang, Y.; Chen, X.; Almoghrabi, A.; Mustafi, R.; Ayaloglu-Butun, F.; Deng, Z.; Haider, H.I.; et al. miR-193a-3p is a Key Tumor Suppressor in Ulcerative Colitis-Associated Colon Cancer and Promotes Carcinogenesis through Upregulation of IL17RD. Clin. Cancer Res. 2017, 23, 5281–5291. [Google Scholar] [CrossRef] [PubMed]
- Mamoori, A.; Wahab, R.; Islam, F.; Lee, K.; Vider, J.; Lu, C.T.; Gopalan, V.; Lam, A.K. Clinical and biological significance of miR-193a-3p targeted KRAS in colorectal cancer pathogenesis. Hum. Pathol. 2018, 71, 145–156. [Google Scholar] [CrossRef] [PubMed]
- Lin, M.; Duan, B.; Hu, J.; Yu, H.; Sheng, H.; Gao, H.; Huang, J. Decreased expression of miR-193a-3p is associated with poor prognosis in colorectal cancer. Oncol. Lett. 2017, 14, 1061–1067. [Google Scholar] [CrossRef] [PubMed]
- Nie, W.; Ge, H.J.; Yang, X.Q.; Sun, X.; Huang, H.; Tao, X.; Chen, W.S.; Li, B. LncRNA-UCA1 exerts oncogenic functions in non-small cell lung cancer by targeting miR-193a-3p. Cancer Lett. 2016, 371, 99–106. [Google Scholar] [CrossRef] [PubMed]
- Li, H.-J.; Sun, X.-M.; Li, Z.-K.; Yin, Q.-W.; Pang, H.; Pan, J.-J.; Li, X.; Chen, W. LncRNA UCA1 Promotes Mitochondrial Function of Bladder Cancer via the MiR-195/ARL2 Signaling Pathway. Cell. Physiol. Biochem. 2017, 43, 2548–2561. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Chen, J.; Zhou, Z.; He, Z. miR-195 enhances the radiosensitivity of colorectal cancer cells by suppressing CARM1. OncoTargets Ther. 2017, 10, 1027–1038. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Xu, J.; Jiang, T.; Liu, G.; Wang, D.; Lu, Y. MicroRNA-195 suppresses colorectal cancer cells proliferation via targeting FGF2 and regulating Wnt/beta-catenin pathway. Am. J. Cancer Res. 2016, 6, 2631–2640. [Google Scholar] [PubMed]
- Ye, S.; Song, W.; Xu, X.; Zhao, X.; Yang, L. IGF2BP2 promotes colorectal cancer cell proliferation and survival through interfering with RAF-1 degradation by miR-195. FEBS Lett. 2016, 590, 1641–1650. [Google Scholar] [CrossRef] [PubMed]
- Yang, I.P.; Tsai, H.L.; Miao, Z.F.; Huang, C.W.; Kuo, C.H.; Wu, J.Y.; Wang, W.M.; Juo, S.H.; Wang, J.Y. Development of a deregulating microRNA panel for the detection of early relapse in postoperative colorectal cancer patients. J. Transl. Med. 2016, 14, 108. [Google Scholar] [CrossRef] [PubMed]
- Yang, B.; Tan, Z.; Song, Y. Study on the molecular regulatory mechanism of MicroRNA-195 in the invasion and metastasis of colorectal carcinoma. Int. J. Clin. Exp. Med. 2015, 8, 3793–3800. [Google Scholar] [PubMed]
- Kim, C.; Hong, Y.; Lee, H.; Kang, H.; Lee, E.K. MicroRNA-195 desensitizes HCT116 human colon cancer cells to 5-fluorouracil. Cancer Lett. 2018, 412, 264–271. [Google Scholar] [CrossRef] [PubMed]
- Pan, J.; Li, X.; Wu, W.; Xue, M.; Hou, H.; Zhai, W.; Chen, W. Long non-coding RNA UCA1 promotes cisplatin/gemcitabine resistance through CREB modulating miR-196a-5p in bladder cancer cells. Cancer Lett. 2016, 382, 64–76. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Su, Y.L.; Yu, H.; Qian, B.Y. Meta-Analysis of the Association between Mir-196a-2 Polymorphism and Cancer Susceptibility. Cancer Biol. Med. 2012, 9, 63–72. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Zhang, C.; Zhao, D.; Liu, K.; Li, T.; Tian, H. Mir-196a-2 C>T polymorphism as a susceptibility factor for colorectal cancer. Int. J. Clin. Exp. Med. 2015, 8, 2600–2606. [Google Scholar] [PubMed]
- Ge, J.; Chen, Z.; Li, R.; Lu, T.; Xiao, G. Upregulation of microRNA-196a and microRNA-196b cooperatively correlate with aggressive progression and unfavorable prognosis in patients with colorectal cancer. Cancer Cell Int. 2014, 14, 128. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Du, P.; She, J.; Cao, L.; Li, Y.; Xia, H. Loss of ZG16 is regulated by miR-196a and contributes to stemness and progression of colorectal cancer. Oncotarget 2016, 7, 86695–86703. [Google Scholar] [CrossRef] [PubMed]
- Xiao, J.N.; Yan, T.H.; Yu, R.M.; Gao, Y.; Zeng, W.L.; Lu, S.W.; Que, H.X.; Liu, Z.P.; Jiang, J.H. Long non-coding RNA UCA1 regulates the expression of Snail2 by miR-203 to promote hepatocellular carcinoma progression. J. Cancer Res. Clin. Oncol. 2017, 143, 981–990. [Google Scholar] [CrossRef] [PubMed]
- Ye, H.; Hao, H.; Wang, J.; Chen, R.; Huang, Z. miR-203 as a novel biomarker for the diagnosis and prognosis of colorectal cancer: A systematic review and meta-analysis. OncoTargets Ther. 2017, 10, 3685–3696. [Google Scholar] [CrossRef] [PubMed]
- Xu, Q.; Liu, M.; Zhang, J.; Xue, L.; Zhang, G.; Hu, C.; Wang, Z.; He, S.; Chen, L.; Ma, K.; et al. Overexpression of KLF4 promotes cell senescence through microRNA-203-survivin-p21 pathway. Oncotarget 2016, 7, 60290–60302. [Google Scholar] [CrossRef] [PubMed]
- Takano, Y.; Masuda, T.; Iinuma, H.; Yamaguchi, R.; Sato, K.; Tobo, T.; Hirata, H.; Kuroda, Y.; Nambara, S.; Hayashi, N.; et al. Circulating exosomal microRNA-203 is associated with metastasis possibly via inducing tumor-associated macrophages in colorectal cancer. Oncotarget 2017, 8, 78598–78613. [Google Scholar] [CrossRef] [PubMed]
- Shen, B.; Yuan, Y.; Zhang, Y.; Yu, S.; Peng, W.; Huang, X.; Feng, J. Long non-coding RNA FBXL19-AS1 plays oncogenic role in colorectal cancer by sponging miR-203. Biochem. Biophys. Res. Commun. 2017, 488, 67–73. [Google Scholar] [CrossRef] [PubMed]
- Kingham, T.P.; Nguyen, H.C.B.; Zheng, J.; Konstantinidis, I.T.; Sadot, E.; Shia, J.; Kuk, D.; Zhang, S.; Saltz, L.; D’Angelica, M.I.; et al. MicroRNA-203 predicts human survival after resection of colorectal liver metastasis. Oncotarget 2017, 8, 18821–18831. [Google Scholar] [CrossRef] [PubMed]
- Fu, Q.; Zhang, J.; Xu, X.; Qian, F.; Feng, K.; Ma, J. miR-203 is a predictive biomarker for colorectal cancer and its expression is associated with BIRC5. Tumour Biol. 2016. [Google Scholar] [CrossRef] [PubMed]
- Yin, Y.; Zhang, B.; Wang, W.; Fei, B.; Quan, C.; Zhang, J.; Song, M.; Bian, Z.; Wang, Q.; Ni, S.; et al. miR-204-5p inhibits proliferation and invasion and enhances chemotherapeutic sensitivity of colorectal cancer cells by downregulating RAB22A. Clin. Cancer Res. 2014, 20, 6187–6199. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Liang, Y.; Shen, L. MicroRNA-204 modulates colorectal cancer cell sensitivity in response to 5-fluorouracil-based treatment by targeting high mobility group protein A2. Biol. Open 2016, 5, 563–570. [Google Scholar] [CrossRef] [PubMed]
- Sumbul, A.T.; Gogebakan, B.; Ergun, S.; Yengil, E.; Batmaci, C.Y.; Tonyali, O.; Yaldiz, M. miR-204-5p expression in colorectal cancer: An autophagy-associated gene. Tumour Biol. 2014, 35, 12713–12719. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Bu, X.; Zhang, Y.; Zhao, X.; Kong, Y.; Ma, L.; Niu, S.; Wu, B.; Meng, C. LncRNA-UCA1 enhances MMP-13 expression by inhibiting miR-204-5p in human chondrocytes. Oncotarget 2017, 8, 91281–91290. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Yang, B.; Ma, B. The UCA1/miR-204/Sirt1 axis modulates docetaxel sensitivity of prostate cancer cells. Cancer Chemother. Pharmacol. 2016, 78, 1025–1031. [Google Scholar] [CrossRef] [PubMed]
- Jiao, C.; Song, Z.; Chen, J.; Zhong, J.; Cai, W.; Tian, S.; Chen, S.; Yi, Y.; Xiao, Y. lncRNA-UCA1 enhances cell proliferation through functioning as a ceRNA of Sox4 in esophageal cancer. Oncol. Rep. 2016, 36, 2960–2966. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Cui, C.; Chen, J.; Hu, Z.; Wang, Y.; Hu, D. Long noncoding RNA UCA1 promotes papillary thyroid cancer cell proliferation via miR204mediated BRD4 activation. Mol. Med. Rep. 2018, 18, 3059–3067. [Google Scholar] [CrossRef] [PubMed]
- Yan, Q.; Tian, Y.; Hao, F. Downregulation of lncRNA UCA1 inhibits proliferation and invasion of cervical cancer cells through miR-206 expression. Oncol. Res. 2018. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Ying, H.Q.; He, B.S.; Pan, Y.Q.; Deng, Q.W.; Sun, H.L.; Chen, J.; Liu, X.; Wang, S.K. Upregulated lncRNA-UCA1 contributes to progression of hepatocellular carcinoma through inhibition of miR-216b and activation of FGFR1/ERK signaling pathway. Oncotarget 2015, 6, 7899–7917. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Liu, X.; He, B.; Pan, Y.; Sun, H.; Xu, T.; Hu, X.; Wang, S. MiR-216b functions as a tumor suppressor by targeting HMGB1-mediated JAK2/STAT3 signaling way in colorectal cancer. Am. J. Cancer Res. 2017, 7, 2051–2069. [Google Scholar] [PubMed]
- Kim, S.Y.; Lee, Y.H.; Bae, Y.S. MiR-186, miR-216b, miR-337-3p, and miR-760 cooperatively induce cellular senescence by targeting alpha subunit of protein kinase CKII in human colorectal cancer cells. Biochem. Biophys. Res. Commun. 2012, 429, 173–179. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Li, Q.; Wang, H. MiR-216b suppresses colorectal cancer proliferation, migration, and invasion by targeting SRPK1. OncoTargets Ther. 2018, 11, 1671–1681. [Google Scholar] [CrossRef] [PubMed]
- Zou, J.; Kuang, W.; Hu, J.; Rao, H. miR-216b promotes cell growth and enhances chemosensitivity of colorectal cancer by suppressing PDZ-binding kinase. Biochem. Biophys. Res. Commun. 2017, 488, 247–252. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Jiang, Y.; Wan, Y.; Zhang, L.; Qiu, J.; Zhou, S.; Cheng, W. UCA1 functions as a competing endogenous RNA to suppress epithelial ovarian cancer metastasis. Tumour Biol. 2016, 37, 10633–10641. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Liu, W.-G.; Lu, J.-H.; Liu, Z.J.; Li, H.-B.; Liu, G.-J.; She, H.-Y.; Li, G.-Y.; Shi, X.-H. LncRNA UCA1 promotes renal cell carcinoma proliferation through epigenetically repressing p21 expression and negatively regulating miR-495. Tumor Biol. 2017, 39, 101042831770163. [Google Scholar] [CrossRef] [PubMed]
- Bai, Z.; Wang, J.; Wang, T.; Li, Y.; Zhao, X.; Wu, G.; Yang, Y.; Deng, W.; Zhang, Z. The MiR-495/Annexin A3/P53 Axis Inhibits the Invasion and EMT of Colorectal Cancer Cells. Cell Physiol. Biochem. 2017, 44, 1882–1895. [Google Scholar] [CrossRef] [PubMed]
- Chuang, A.Y.; Chuang, J.C.; Zhai, Z.; Wu, F.; Kwon, J.H. NOD2 expression is regulated by microRNAs in colonic epithelial HCT116 cells. Inflamm. Bowel Dis. 2014, 20, 126–135. [Google Scholar] [CrossRef] [PubMed]
- Guo, S.; Yang, P.; Jiang, X.; Li, X.; Wang, Y.; Zhang, X.; Sun, B.; Zhang, Y.; Jia, Y. Genetic and epigenetic silencing of mircoRNA-506-3p enhances COTL1 oncogene expression to foster non-small lung cancer progression. Oncotarget 2017, 8, 644–657. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Liu, S.; Tian, L.; Wu, M.; Ai, F.; Tang, W.; Zhao, L.; Ding, J.; Zhang, L.; Tang, A. miR-124 and miR-506 inhibit colorectal cancer progression by targeting DNMT3B and DNMT1. Oncotarget 2015, 6, 38139–38150. [Google Scholar] [CrossRef] [PubMed]
- Krawczyk, P.; Powrozek, T.; Olesinski, T.; Dmitruk, A.; Dziwota, J.; Kowalski, D.; Milanowski, J. Evaluation of miR-506 and miR-4316 expression in early and non-invasive diagnosis of colorectal cancer. Int. J. Colorectal Dis. 2017, 32, 1057–1060. [Google Scholar] [CrossRef] [PubMed]
- Tong, J.L.; Zhang, C.P.; Nie, F.; Xu, X.T.; Zhu, M.M.; Xiao, S.D.; Ran, Z.H. MicroRNA 506 regulates expression of PPAR alpha in hydroxycamptothecin-resistant human colon cancer cells. FEBS Lett. 2011, 585, 3560–3568. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Zhang, Y.; Tang, A.; Tian, L. miR-506 inhibits cell proliferation and invasion by targeting TET family in colorectal cancer. Iran J. Basic Med. Sci. 2016, 19, 316–322. [Google Scholar] [PubMed]
- Zhou, H.; Lin, C.; Zhang, Y.; Zhang, X.; Zhang, C.; Zhang, P.; Xie, X.; Ren, Z. miR-506 enhances the sensitivity of human colorectal cancer cells to oxaliplatin by suppressing MDR1/P-gp expression. Cell Prolif. 2017, 50, e12341. [Google Scholar] [CrossRef] [PubMed]
- Zu, C.; Liu, T.; Zhang, G. MicroRNA-506 Inhibits Malignancy of Colorectal Carcinoma Cells by Targeting LAMC1. Ann. Clin. Lab. Sci. 2016, 46, 666–674. [Google Scholar] [PubMed]
- Wei, Y.; Sun, Q.; Zhao, L.; Wu, J.; Chen, X.; Wang, Y.; Zang, W.; Zhao, G. LncRNA UCA1-miR-507-FOXM1 axis is involved in cell proliferation, invasion and G0/G1 cell cycle arrest in melanoma. Med. Oncol. 2016, 33. [Google Scholar] [CrossRef] [PubMed]
- Gu, L.; Lu, L.-S.; Zhou, D.-L.; Liu, Z.-C. UCA1 promotes cell proliferation and invasion of gastric cancer by targeting CREB1 sponging to miR-590-3p. Cancer Med. 2018. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.W.; Oh, E.T.; Kim, J.M.; Park, J.S.; Lee, D.H.; Lee, J.S.; Kim, K.K.; Park, H.J. Hypoxia-induced microRNA-590-5p promotes colorectal cancer progression by modulating matrix metalloproteinase activity. Cancer Lett. 2018, 416, 31–41. [Google Scholar] [CrossRef] [PubMed]
- Ou, C.; Sun, Z.; Li, X.; Ren, W.; Qin, Z.; Zhang, X.; Yuan, W.; Wang, J.; Yu, W.; Zhang, S.; et al. MiR-590-5p, a density-sensitive microRNA, inhibits tumorigenesis by targeting YAP1 in colorectal cancer. Cancer Lett. 2017, 399, 53–63. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.Q.; Shi, K.; Zhou, Q.B.; Zeng, X.Y.; Liu, J.; Yang, S.X.; Wang, Q.S.; Li, Z.; Wang, G.X.; Song, J.M.; et al. MiR-590-3p promotes proliferation and metastasis of colorectal cancer via Hippo pathway. Oncotarget 2017, 8, 58061–58071. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Q.; Zhu, Y.; Wei, X.; Zhou, J.; Chang, L.; Sui, H.; Han, Y.; Piao, D.; Sha, R.; Bai, Y. MiR-590-5p inhibits colorectal cancer angiogenesis and metastasis by regulating nuclear factor 90/vascular endothelial growth factor A axis. Cell Death Dis. 2016, 7, e2413. [Google Scholar] [CrossRef] [PubMed]
- Alvarez-Diaz, S.; Valle, N.; Ferrer-Mayorga, G.; Lombardia, L.; Herrera, M.; Dominguez, O.; Segura, M.F.; Bonilla, F.; Hernando, E.; Munoz, A. MicroRNA-22 is induced by vitamin D and contributes to its antiproliferative, antimigratory and gene regulatory effects in colon cancer cells. Hum. Mol. Genet. 2012, 21, 2157–2165. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Song, Y.; Liu, T.J.; Cui, Y.B.; Jiang, Y.; Xie, Z.S.; Xie, S.L. miRNA-22 suppresses colon cancer cell migration and invasion by inhibiting the expression of T-cell lymphoma invasion and metastasis 1 and matrix metalloproteinases 2 and 9. Oncol. Rep. 2013, 29, 1932–1938. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Sun, H.; Zhang, H. The predicted target gene validation, function, and prognosis studies of miRNA-22 in colorectal cancer tissue. Tumour Biol. 2017, 39, 1010428317692257. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Zhang, Y.; Zhao, J.; Kong, F.; Chen, Y. Overexpression of miR-22 reverses paclitaxel-induced chemoresistance through activation of PTEN signaling in p53-mutated colon cancer cells. Mol. Cell. Biochem. 2011, 357, 31–38. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Chen, X.; Cheng, R.; Yang, F.; Yu, M.; Wang, C.; Cui, S.; Hong, Y.; Liang, H.; Liu, M.; et al. The Jun/miR-22/HuR regulatory axis contributes to tumourigenesis in colorectal cancer. Mol. Cancer 2018, 17, 11. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; Liu, X.; Su, B.; Zhang, Z.; Zeng, X.; Lei, Y.; Shan, J.; Wu, Y.; Tang, H.; Su, Q. microRNA-22 acts as a metastasis suppressor by targeting metadherin in gastric cancer. Mol. Med. Rep. 2015, 11, 454–460. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Tsuchiya, N.; Izumiya, M.; Ogata-Kawata, H.; Okamoto, K.; Fujiwara, Y.; Nakai, M.; Okabe, A.; Schetter, A.J.; Bowman, E.D.; Midorikawa, Y.; et al. Tumor suppressor miR-22 determines p53-dependent cellular fate through post-transcriptional regulation of p21. Cancer Res. 2011, 71, 4628–4639. [Google Scholar] [CrossRef] [PubMed]
- Yamakuchi, M.; Yagi, S.; Ito, T.; Lowenstein, C.J. MicroRNA-22 regulates hypoxia signaling in colon cancer cells. PLoS ONE 2011, 6, e20291. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Hu, Y.; Liu, H.X.; Wan, Y.J. MiR-22-silenced cyclin A expression in colon and liver cancer cells is regulated by bile acid receptor. J. Biol. Chem. 2015, 290, 6507–6515. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Xia, S.; Tian, H.; Liu, Z.; Zhou, T. Clinical significance of miR-22 expression in patients with colorectal cancer. Med. Oncol. 2012, 29, 3108–3112. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Tang, J.; Li, C.; Kong, J.; Wang, J.; Wu, Y.; Xu, E.; Lai, M. MiR-22 regulates 5-FU sensitivity by inhibiting autophagy and promoting apoptosis in colorectal cancer cells. Cancer Lett. 2015, 356, 781–790. [Google Scholar] [CrossRef] [PubMed]
- Jahid, S.; Sun, J.; Edwards, R.A.; Dizon, D.; Panarelli, N.C.; Milsom, J.W.; Sikandar, S.S.; Gumus, Z.H.; Lipkin, S.M. miR-23a promotes the transition from indolent to invasive colorectal cancer. Cancer Discov. 2012, 2, 540–553. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Liao, D.; Wang, X.; Wu, Z.; Nie, J.; Bai, M.; Fu, X.; Mei, Q.; Han, W. Elevated microRNA-23a Expression Enhances the Chemoresistance of Colorectal Cancer Cells with Microsatellite Instability to 5-Fluorouracil by Directly Targeting ABCF1. Curr. Protein Pept. Sci. 2015, 16, 301–309. [Google Scholar] [CrossRef] [PubMed]
- Shang, J.; Yang, F.; Wang, Y.; Xue, G.; Mei, Q.; Wang, F.; Sun, S. MicroRNA-23a antisense enhances 5-fluorouracil chemosensitivity through APAF-1/caspase-9 apoptotic pathway in colorectal cancer cells. J. Cell. Biochem. 2014, 115, 772–784. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wei, W.; Sarkar, F.H. miR-23a, a critical regulator of “migR”ation and metastasis in colorectal cancer. Cancer Discov. 2012, 2, 489–491. [Google Scholar] [CrossRef] [PubMed]
- Yong, F.L.; Law, C.W.; Wang, C.W. Potentiality of a triple microRNA classifier: miR-193a-3p, miR-23a and miR-338-5p for early detection of colorectal cancer. BMC Cancer 2013, 13, 280. [Google Scholar] [CrossRef] [PubMed]
- Yong, F.L.; Wang, C.W.; Roslani, A.C.; Law, C.W. The involvement of miR-23a/APAF1 regulation axis in colorectal cancer. Int. J. Mol. Sci. 2014, 15, 11713–11729. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.; Liu, Y.; Jin, X.; Lu, W.; Liu, J.; Xia, Z.; Yuan, Q.; Zhao, X.; Xu, N.; Liang, S. MicroRNA-26a regulates glucose metabolism by direct targeting PDHX in colorectal cancer cells. BMC Cancer 2014, 14, 1471–2407. [Google Scholar] [CrossRef] [PubMed]
- Fu, X.; Meng, Z.; Liang, W.; Tian, Y.; Wang, X.; Han, W.; Lou, G.; Lou, F.; Yen, Y.; Yu, H.; et al. miR-26a enhances miRNA biogenesis by targeting Lin28B and Zcchc11 to suppress tumor growth and metastasis. Oncogene 2014, 33, 4296–4306. [Google Scholar] [CrossRef] [PubMed]
- Ghanbari, R.; Mosakhani, N.; Asadi, J.; Nouraee, N.; Mowla, S.J.; Yazdani, Y.; Mohamadkhani, A.; Poustchi, H.; Knuutila, S.; Malekzadeh, R. Downregulation of Plasma MiR-142-3p and MiR-26a-5p in Patients With Colorectal Carcinoma. Iran J. Cancer Prev. 2015, 8, 22. [Google Scholar] [CrossRef] [PubMed]
- Ghanbari, R.; Rezasoltani, S.; Hashemi, J.; Mohamadkhani, A.; Tahmasebifar, A.; Arefian, E.; Mobarra, N.; Asadi, J.; Nazemalhosseini Mojarad, E.; Yazdani, Y.; et al. Expression Analysis of Previously Verified Fecal and Plasma Dow-regulated MicroRNAs (miR-4478, 1295-3p, 142-3p and 26a-5p), in FFPE Tissue Samples of CRC Patients. Arch. Iran Med. 2017, 20, 92–95. [Google Scholar] [PubMed]
- Konishi, H.; Fujiya, M.; Ueno, N.; Moriichi, K.; Sasajima, J.; Ikuta, K.; Tanabe, H.; Tanaka, H.; Kohgo, Y. microRNA-26a and -584 inhibit the colorectal cancer progression through inhibition of the binding of hnRNP A1-CDK6 mRNA. Biochem. Biophys. Res. Commun. 2015, 467, 847–852. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Sun, Z.; Liu, B.; Shan, Y.; Zhao, L.; Jia, L. Tumor-suppressive miR-26a and miR-26b inhibit cell aggressiveness by regulating FUT4 in colorectal cancer. Cell Death Dis. 2017, 8, 281. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Urrutia, E.; Coronel-Hernandez, J.; Garcia-Castillo, V.; Contreras-Romero, C.; Martinez-Gutierrez, A.; Estrada-Galicia, D.; Terrazas, L.I.; Lopez-Camarillo, C.; Maldonado-Martinez, H.; Jacobo-Herrera, N.; et al. MiR-26a downregulates retinoblastoma in colorectal cancer. Tumour Biol. 2017, 39, 1010428317695945. [Google Scholar] [CrossRef] [PubMed]
- Ying, H.Q.; Peng, H.X.; He, B.S.; Pan, Y.Q.; Wang, F.; Sun, H.L.; Liu, X.; Chen, J.; Lin, K.; Wang, S.K. MiR-608, pre-miR-124-1 and pre-miR26a-1 polymorphisms modify susceptibility and recurrence-free survival in surgically resected CRC individuals. Oncotarget 2016, 7, 75865–75873. [Google Scholar] [CrossRef] [PubMed]
- Zeitels, L.R.; Acharya, A.; Shi, G.; Chivukula, D.; Chivukula, R.R.; Anandam, J.L.; Abdelnaby, A.A.; Balch, G.C.; Mansour, J.C.; Yopp, A.C.; et al. Tumor suppression by miR-26 overrides potential oncogenic activity in intestinal tumorigenesis. Genes Dev. 2014, 28, 2585–2590. [Google Scholar] [CrossRef] [PubMed]
- Fasihi, A.; Soltani, B.; Atashi, A.; Nasiri, S. Introduction of hsa-miR-103a and hsa-miR-1827 and hsa-miR-137 as new regulators of Wnt signaling pathway and their relation to colorectal carcinoma. J. Cell. Biochem. 2017, 119, 5104–5117. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.Y.; Lin, Y.M.; Chung, H.C.; Lang, Y.D.; Lin, C.J.; Huang, J.; Wang, W.C.; Lin, F.M.; Chen, Z.; Huang, H.D.; et al. miR-103/107 promote metastasis of colorectal cancer by targeting the metastasis suppressors DAPK and KLF4. Cancer Res. 2012, 72, 3631–3641. [Google Scholar] [CrossRef] [PubMed]
- Molina-Pinelo, S.; Carnero, A.; Rivera, F.; Estevez-Garcia, P.; Bozada, J.M.; Limon, M.L.; Benavent, M.; Gomez, J.; Pastor, M.D.; Chaves, M.; et al. MiR-107 and miR-99a-3p predict chemotherapy response in patients with advanced colorectal cancer. BMC Cancer 2014, 14, 1471–2407. [Google Scholar] [CrossRef] [PubMed]
- Deng, G.; Kakar, S.; Kim, Y.S. MicroRNA-124a and microRNA-34b/c are frequently methylated in all histological types of colorectal cancer and polyps, and in the adjacent normal mucosa. Oncol. Lett. 2011, 2, 175–180. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.R.; Wang, H.P.; Zhang, S.L.; Wang, M.X.; Zhu, Z.S. Pri-miR-124 rs531564 polymorphism and colorectal cancer risk. Sci. Rep. 2015, 5, 14818. [Google Scholar] [CrossRef] [PubMed]
- Jinushi, T.; Shibayama, Y.; Kinoshita, I.; Oizumi, S.; Jinushi, M.; Aota, T.; Takahashi, T.; Horita, S.; Dosaka-Akita, H.; Iseki, K. Low expression levels of microRNA-124-5p correlated with poor prognosis in colorectal cancer via targeting of SMC4. Cancer Med. 2014, 3, 1544–1552. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Chen, W.; Lei, S.; Xiong, L.; Zhao, H.; Liang, D.; Lei, Z.; Zhou, N.; Yao, H.; Liang, Y. Wild-type and mutant p53 differentially modulate miR-124/iASPP feedback following pohotodynamic therapy in human colon cancer cell line. Cell Death Dis. 2017, 8, e3096. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Yao, H.; Lei, S.; Xiong, L.; Qi, H.; Qian, K.; Liu, J.; Wang, P.; Zhao, H. The miR-124-p63 feedback loop modulates colorectal cancer growth. Oncotarget 2017, 8, 29101–29115. [Google Scholar] [CrossRef] [PubMed]
- Park, S.Y.; Kim, H.; Yoon, S.; Bae, J.A.; Choi, S.Y.; Jung, Y.D.; Kim, K.K. KITENIN-targeting microRNA-124 suppresses colorectal cancer cell motility and tumorigenesis. Mol. Ther. 2014, 22, 1653–1664. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Zhao, X.; Luo, M.; Zhou, Y.; Ren, W.; Wu, K.; Li, X.; Shen, J.; Hu, Y. The pro-apoptotic role of the regulatory feedback loop between miR-124 and PKM1/HNF4alpha in colorectal cancer cells. Int. J. Mol. Sci. 2014, 15, 4318–4332. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Zhao, X.; Zhou, Y.; Hu, Y. miR-124, miR-137 and miR-340 regulate colorectal cancer growth via inhibition of the Warburg effect. Oncol. Rep. 2012, 28, 1346–1352. [Google Scholar] [CrossRef] [PubMed]
- Taniguchi, K.; Sugito, N.; Kumazaki, M.; Shinohara, H.; Yamada, N.; Matsuhashi, N.; Futamura, M.; Ito, Y.; Otsuki, Y.; Yoshida, K.; et al. Positive feedback of DDX6/c-Myc/PTB1 regulated by miR-124 contributes to maintenance of the Warburg effect in colon cancer cells. Biochim. Biophys. Acta 2015, 1852, 1971–1980. [Google Scholar] [CrossRef] [PubMed]
- Taniguchi, K.; Sugito, N.; Kumazaki, M.; Shinohara, H.; Yamada, N.; Nakagawa, Y.; Ito, Y.; Otsuki, Y.; Uno, B.; Uchiyama, K.; et al. MicroRNA-124 inhibits cancer cell growth through PTB1/PKM1/PKM2 feedback cascade in colorectal cancer. Cancer Lett. 2015, 363, 17–27. [Google Scholar] [CrossRef] [PubMed]
- Ueda, Y.; Ando, T.; Nanjo, S.; Ushijima, T.; Sugiyama, T. DNA methylation of microRNA-124a is a potential risk marker of colitis-associated cancer in patients with ulcerative colitis. Dig. Dis. Sci. 2014, 59, 2444–2451. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.J.; Li, Y.; Wang, R.; Wang, C.; Yu, Y.Y.; Yang, L.; Zhang, Y.; Zhou, B.; Zhou, Z.G.; Sun, X.F. Downregulation of microRNA-124 is an independent prognostic factor in patients with colorectal cancer. Int. J. Color. Dis. 2013, 28, 183–189. [Google Scholar] [CrossRef] [PubMed]
- Xi, Z.W.; Xin, S.Y.; Zhou, L.Q.; Yuan, H.X.; Wang, Q.; Chen, K.X. Downregulation of rho-associated protein kinase 1 by miR-124 in colorectal cancer. World J. Gastroenterol. 2015, 21, 5454–5464. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Lu, Y.; Yue, X.; Li, H.; Luo, X.; Wang, Y.; Wang, K.; Wan, J. MiR-124 suppresses growth of human colorectal cancer by inhibiting STAT3. PLoS ONE 2013, 8, e70300. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Xu, Z.; Ren, X.; Chen, K.; Xin, S. MicroRNA-124 (MiR-124) Inhibits Cell Proliferation, Metastasis and Invasion in Colorectal Cancer by Downregulating Rho-Associated Protein Kinase 1(ROCK1). Cell Physiol. Biochem. 2016, 38, 1785–1795. [Google Scholar] [CrossRef] [PubMed]
- Cristobal, I.; Torrejon, B.; Gonzalez-Alonso, P.; Manso, R.; Rojo, F.; Garcia-Foncillas, J. Downregulation of miR-138 as a Contributing Mechanism to Lcn-2 Overexpression in Colorectal Cancer with Liver Metastasis. World J. Surg. 2016, 40, 1021–1022. [Google Scholar] [CrossRef] [PubMed]
- Long, L.; Huang, G.; Zhu, H.; Guo, Y.; Liu, Y.; Huo, J. Down-regulation of miR-138 promotes colorectal cancer metastasis via directly targeting TWIST2. J. Transl. Med. 2013, 11, 1479–5876. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.L.; Xu, L.L.; Wang, F. Hsa_circ_0020397 regulates colorectal cancer cell viability, apoptosis and invasion by promoting the expression of the miR-138 targets TERT and PD-L1. Cell Biol. Int. 2017, 41, 1056–1064. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Yu, H.; Yi, S.; Peng, X.; Su, P.; Xiao, Z.; Liu, R.; Tang, A.; Li, X.; Liu, F.; et al. The tumor suppressor miR-138-5p targets PD-L1 in colorectal cancer. Oncotarget 2016, 7, 45370–45384. [Google Scholar] [CrossRef] [PubMed]
- Chandrasekaran, K.S.; Sathyanarayanan, A.; Karunagaran, D. MicroRNA-214 suppresses growth, migration and invasion through a novel target, high mobility group AT-hook 1, in human cervical and colorectal cancer cells. Br. J. Cancer 2016, 115, 741–751. [Google Scholar] [CrossRef] [PubMed]
- Cristobal, I.; Carames, C.; Madoz-Gurpide, J.; Rojo, F.; Aguilera, O.; Garcia-Foncillas, J. Downregulation of miR-214 is specific of liver metastasis in colorectal cancer and could play a role determining the metastatic niche. Int. J. Color. Dis. 2014, 29, 885. [Google Scholar] [CrossRef] [PubMed]
- He, G.Y.; Hu, J.L.; Zhou, L.; Zhu, X.H.; Xin, S.N.; Zhang, D.; Lu, G.F.; Liao, W.T.; Ding, Y.Q.; Liang, L. The FOXD3/miR-214/MED19 axis suppresses tumour growth and metastasis in human colorectal cancer. Br. J. Cancer 2016, 115, 1367–1378. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.L.; He, G.Y.; Lan, X.L.; Zeng, Z.C.; Guan, J.; Ding, Y.; Qian, X.L.; Liao, W.T.; Ding, Y.Q.; Liang, L. Inhibition of ATG12-mediated autophagy by miR-214 enhances radiosensitivity in colorectal cancer. Oncogenesis 2018, 7, 16. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Hong, L. Prediction value of miR-483 and miR-214 in prognosis and multidrug resistance of esophageal squamous cell carcinoma. Genet. Test. Mol. Biomarkers 2013, 17, 470–474. [Google Scholar] [CrossRef] [PubMed]
- Chai, J.; Guo, D.; Ma, W.; Han, D.; Dong, W.; Guo, H.; Zhang, Y. A feedback loop consisting of RUNX2/LncRNA-PVT1/miR-455 is involved in the progression of colorectal cancer. Am. J. Cancer Res. 2018, 8, 538–550. [Google Scholar] [PubMed]
- Chai, J.; Wang, S.; Han, D.; Dong, W.; Xie, C.; Guo, H. MicroRNA-455 inhibits proliferation and invasion of colorectal cancer by targeting RAF proto-oncogene serine/threonine-protein kinase. Tumour Biol. 2015, 36, 1313–1321. [Google Scholar] [CrossRef] [PubMed]
- Yunqi, H.; Fangrui, Y.; Yongyan, Y.; Yunjian, J.; Wenhui, Z.; Kun, C.; Min, L.; Xianfeng, L.; Caixia, B. MiR-455 functions as a tumor suppressor through targeting GATA6 in colorectal cancer. Oncol. Res. 2018, 3. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.; Lin, Z.; Zhang, L.; Chen, H. MicroRNA-455-3p Inhibits Tumor Cell Proliferation and Induces Apoptosis in HCT116 Human Colon Cancer Cells. Med. Sci. Monit. 2016, 22, 4431–4437. [Google Scholar] [CrossRef] [PubMed]
- Su, E.C.-Y.; Chen, Y.-S.; Tien, Y.-C.; Liu, J.; Ho, B.-C.; Yu, S.-L.; Singh, S. ChemiRs: A web application for microRNAs and chemicals. BMC Bioinform. 2016, 17. [Google Scholar] [CrossRef] [PubMed]
- Mi, H.; Huang, X.; Muruganujan, A.; Tang, H.; Mills, C.; Kang, D.; Thomas, P.D. PANTHER version 11: Expanded annotation data from Gene Ontology and Reactome pathways, and data analysis tool enhancements. Nucleic Acids Res. 2017, 45, D183–D189. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Shen, B.; Tan, M.; Mu, X.; Qin, Y.; Zhang, F.; Liu, Y. Long non-coding RNA UCA1 increases chemoresistance of bladder cancer cells by regulating Wnt signaling. FEBS J. 2014, 281, 1750–1758. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Wang, G.; Yang, L.; Qu, J.; Yang, Z.; Zhou, X. Knockdown of Long Non-Coding RNA UCA1 Increases the Tamoxifen Sensitivity of Breast Cancer Cells through Inhibition of Wnt/beta-Catenin Pathway. PLoS ONE 2016, 11, e0168406. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.T.; Wang, Y.F.; Lai, J.Y.; Shen, S.Y.; Wang, F.; Kong, J.; Zhang, W.; Yang, H.Y. Long non-coding RNA UCA1 contributes to the progression of oral squamous cell carcinoma by regulating the WNT/beta-catenin signaling pathway. Cancer Sci. 2016, 107, 1581–1589. [Google Scholar] [CrossRef] [PubMed]
- Xiao, C.; Wu, C.H.; Hu, H.Z. LncRNA UCA1 promotes epithelial-mesenchymal transition (EMT) of breast cancer cells via enhancing Wnt/beta-catenin signaling pathway. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 2819–2824. [Google Scholar] [PubMed]
- He, L.; Zhu, H.; Zhou, S.; Wu, T.; Wu, H.; Yang, H.; Mao, H.; SekharKathera, C.; Janardhan, A.; Edick, A.M.; et al. Wnt pathway is involved in 5-FU drug resistance of colorectal cancer cells. Exp. Mol. Med. 2018, 50. [Google Scholar] [CrossRef] [PubMed]
- Bian, Z.; Zhang, J.; Li, M.; Feng, Y.; Yao, S.; Song, M.; Qi, X.; Fei, B.; Yin, Y.; Hua, D.; et al. Long non-coding RNA LINC00152 promotes cell proliferation, metastasis, and confers 5-FU resistance in colorectal cancer by inhibiting miR-139-5p. Oncogenesis 2017, 6. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Liang, G.; Yang, S.; Sui, J.; Yao, W.; Shen, X.; Zhang, Y.; Peng, H.; Hong, W.; Xu, S.; et al. Dysregulated lncRNA-UCA1 contributes to the progression of gastric cancer through regulation of the PI3K-Akt-mTOR signaling pathway. Oncotarget 2017, 8, 93476. [Google Scholar] [PubMed]
- Xu, Y.; Yao, Y.; Leng, K.; Li, Z.; Qin, W.; Zhong, X.; Kang, P.; Wan, M.; Jiang, X.; Cui, Y. Long non-coding RNA UCA1 indicates an unfavorable prognosis and promotes tumorigenesis via regulating AKT/GSK-3beta signaling pathway in cholangiocarcinoma. Oncotarget 2017, 8, 96203–96214. [Google Scholar] [PubMed]
- Wang, Z.-Q.; He, C.-Y.; Hu, L.; Shi, H.-P.; Li, J.-F.; Gu, Q.-L.; Su, L.-P.; Liu, B.-Y.; Li, C.; Zhu, Z. Long noncoding RNA UCA1 promotes tumour metastasis by inducing GRK2 degradation in gastric cancer. Cancer Lett. 2017, 408, 10–21. [Google Scholar] [CrossRef] [PubMed]
- Zhen, S.; Hua, L.; Liu, Y.-H.; Sun, X.-M.; Jiang, M.-M.; Chen, W.; Zhao, L.; Li, X. Inhibition of long non-coding RNA UCA1 by CRISPR/Cas9 attenuated malignant phenotypes of bladder cancer. Oncotarget 2017, 8, 9634–9646. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.; Sun, C.; Cui, Z. A long noncoding RNA UCA1 promotes proliferation and predicts poor prognosis in glioma. Clin. Transl. Oncol. 2017, 19, 735–741. [Google Scholar] [CrossRef] [PubMed]
- Zhengyuan, X.; Hu, X.; Qiang, W.; Nanxiang, L.; Junbin, C.; Wangming, Z. Silencing of Urothelial Carcinoma Associated 1 Inhibits the Proliferation and Migration of Medulloblastoma Cells. Med. Sci. Monit. 2017, 23, 4454–4461. [Google Scholar] [CrossRef] [PubMed]
- Pu, H.; Zheng, Q.; Li, H.; Wu, M.; An, J.; Gui, X.; Li, T.; Lu, D. CUDR promotes liver cancer stem cell growth through upregulating TERT and C-Myc. Oncotarget 2015, 6, 40775–40798. [Google Scholar] [CrossRef] [PubMed]
- Li, L.Q.; Pan, D.; Chen, Q.; Zhang, S.W.; Xie, D.Y.; Zheng, X.L.; Chen, H. Sensitization of Gastric Cancer Cells to 5-FU by MicroRNA-204 Through Targeting the TGFBR2-Mediated Epithelial to Mesenchymal Transition. Cell. Physiol. Biochem. 2018, 47, 1533–1545. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.A.; Kim, I.; Yoon, S.K.; Lee, E.K.; Kuh, H.J. Indirect modulation of sensitivity to 5-fluorouracil by microRNA-96 in human colorectal cancer cells. Arch. Pharm. Res. 2015, 38, 239–248. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Li, X.; Wu, Z.; Nie, J.; Guo, M.; Mei, Q.; Han, W. DNA methylation-mediated repression of miR-181a/135a/302c expression promotes the microsatellite-unstable colorectal cancer development and 5-FU resistance via targeting PLAG1. J. Genet. Genom. 2018, 45, 205–214. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Liu, Y.; Zhao, L.; Pan, Y.; Shan, Y.; Li, Y.; Jia, L. Upregulation of microRNA-135b and microRNA-182 promotes chemoresistance of colorectal cancer by targeting ST6GALNAC2 via PI3K/AKT pathway. Mol. Carcinog. 2017, 56, 2669–2680. [Google Scholar] [CrossRef] [PubMed]
- Pagliuca, A.; Valvo, C.; Fabrizi, E.; di Martino, S.; Biffoni, M.; Runci, D.; Forte, S.; De Maria, R.; Ricci-Vitiani, L. Analysis of the combined action of miR-143 and miR-145 on oncogenic pathways in colorectal cancer cells reveals a coordinate program of gene repression. Oncogene 2013, 32, 4806–4813. [Google Scholar] [CrossRef] [PubMed]
- Feng, C.; Zhang, L.; Sun, Y.; Li, X.; Zhan, L.; Lou, Y.; Wang, Y.; Liu, L.; Zhang, Y. GDPD5, a target of miR-195-5p, is associated with metastasis and chemoresistance in colorectal cancer. Biomed. Pharmacother. 2018, 101, 945–952. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.; Wang, M.; Hu, H.; Huang, Q.; Chen, Y.; Wang, G. Overcoming stemness and chemoresistance in colorectal cancer through miR-195-5p-modulated inhibition of notch signaling. Int. J. Biol. Macromol. 2018, 117, 445–453. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Wan, G.; Spizzo, R.; Ivan, C.; Mathur, R.; Hu, X.; Ye, X.; Lu, J.; Fan, F.; Xia, L.; et al. miR-203 induces oxaliplatin resistance in colorectal cancer cells by negatively regulating ATM kinase. Mol. Oncol. 2014, 8, 83–92. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Gao, F.; Zhang, X.P. miR-203 enhances chemosensitivity to 5-fluorouracil by targeting thymidylate synthase in colorectal cancer. Oncol. Rep. 2015, 33, 607–614. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Gao, S.; Chen, X.; Liu, M.; Mao, C.; Fang, X. Overexpression of miR-203 sensitizes paclitaxel (Taxol)-resistant colorectal cancer cells through targeting the salt-inducible kinase 2 (SIK2). Tumour Biol. 2016, 37, 12231–12239. [Google Scholar] [CrossRef] [PubMed]
- Meng, X.; Fu, R. miR-206 regulates 5-FU resistance by targeting Bcl-2 in colon cancer cells. OncoTargets Ther. 2018, 11, 1757–1765. [Google Scholar] [CrossRef] [PubMed]
- Hon, K.W.; Abu, N.; Ab Mutalib, N.-S.; Jamal, R. miRNAs and lncRNAs as Predictive Biomarkers of Response to FOLFOX Therapy in Colorectal Cancer. Front. Pharmacol. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Dong, L.; Lin, W.; Qi, P.; Xu, M.D.; Wu, X.; Ni, S.; Huang, D.; Weng, W.W.; Tan, C.; Sheng, W.; et al. Circulating Long RNAs in Serum Extracellular Vesicles: Their Characterization and Potential Application as Biomarkers for Diagnosis of Colorectal Cancer. Cancer Epidemiol. Biomarkers Prev. 2016, 25, 1158–1166. [Google Scholar] [CrossRef] [PubMed]
- Tang, S.; Tan, G.; Jiang, X.; Han, P.; Zhai, B.; Dong, X.; Qiao, H.; Jiang, H.; Sun, X. An artificial lncRNA targeting multiple miRNAs overcomes sorafenib resistance in hepatocellular carcinoma cells. Oncotarget 2016, 7, 73257–73269. [Google Scholar] [CrossRef] [PubMed]
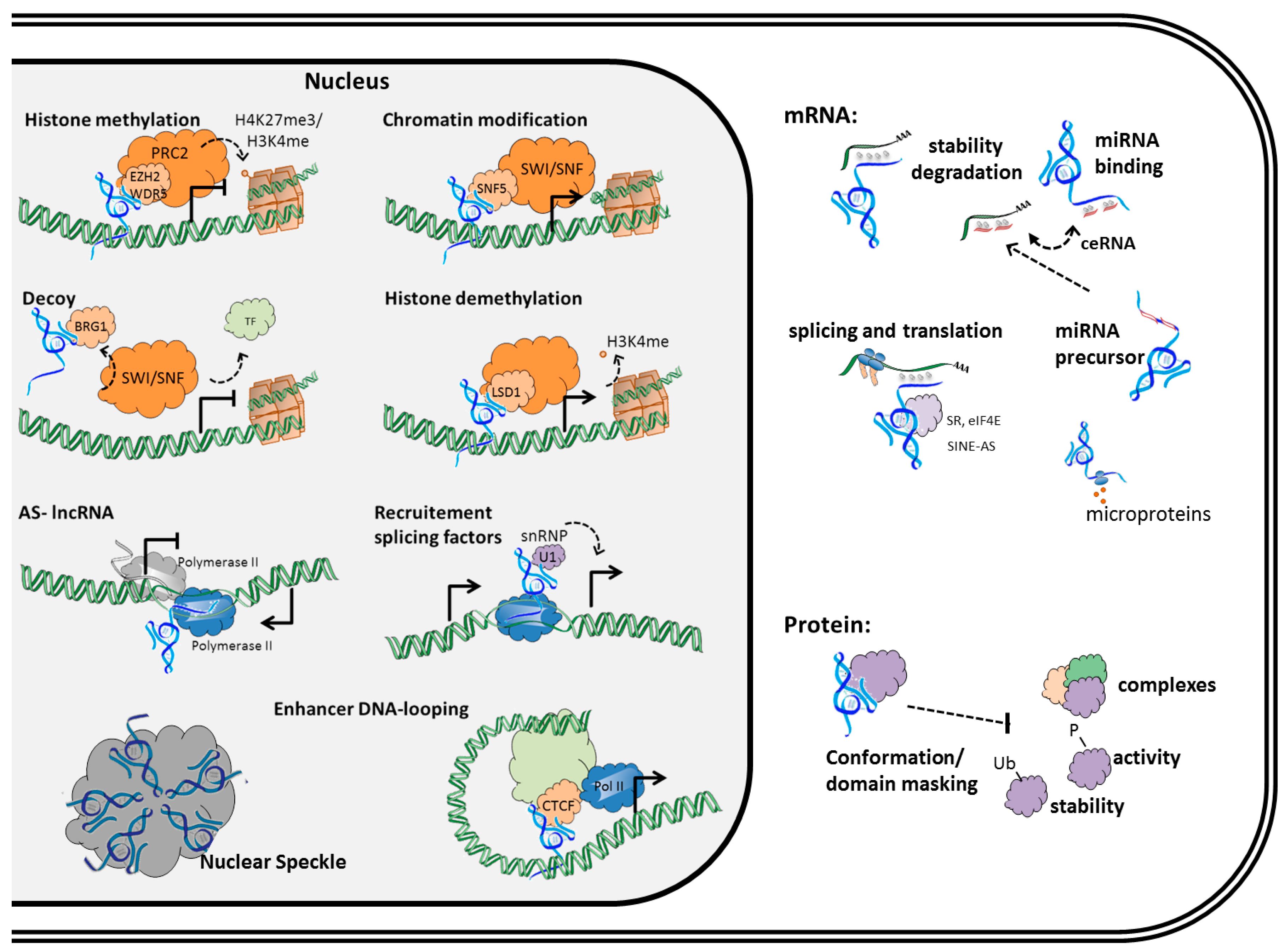
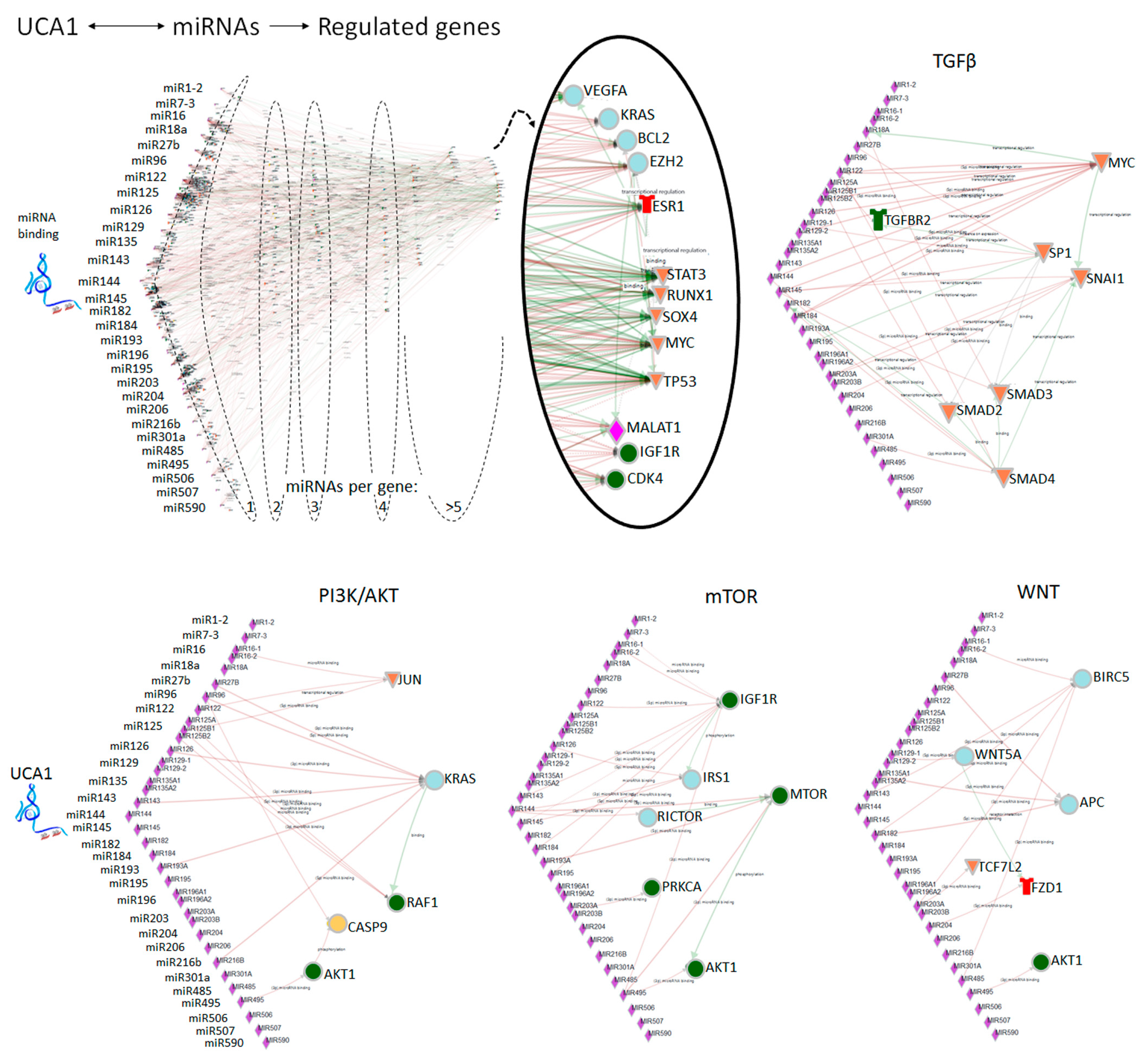
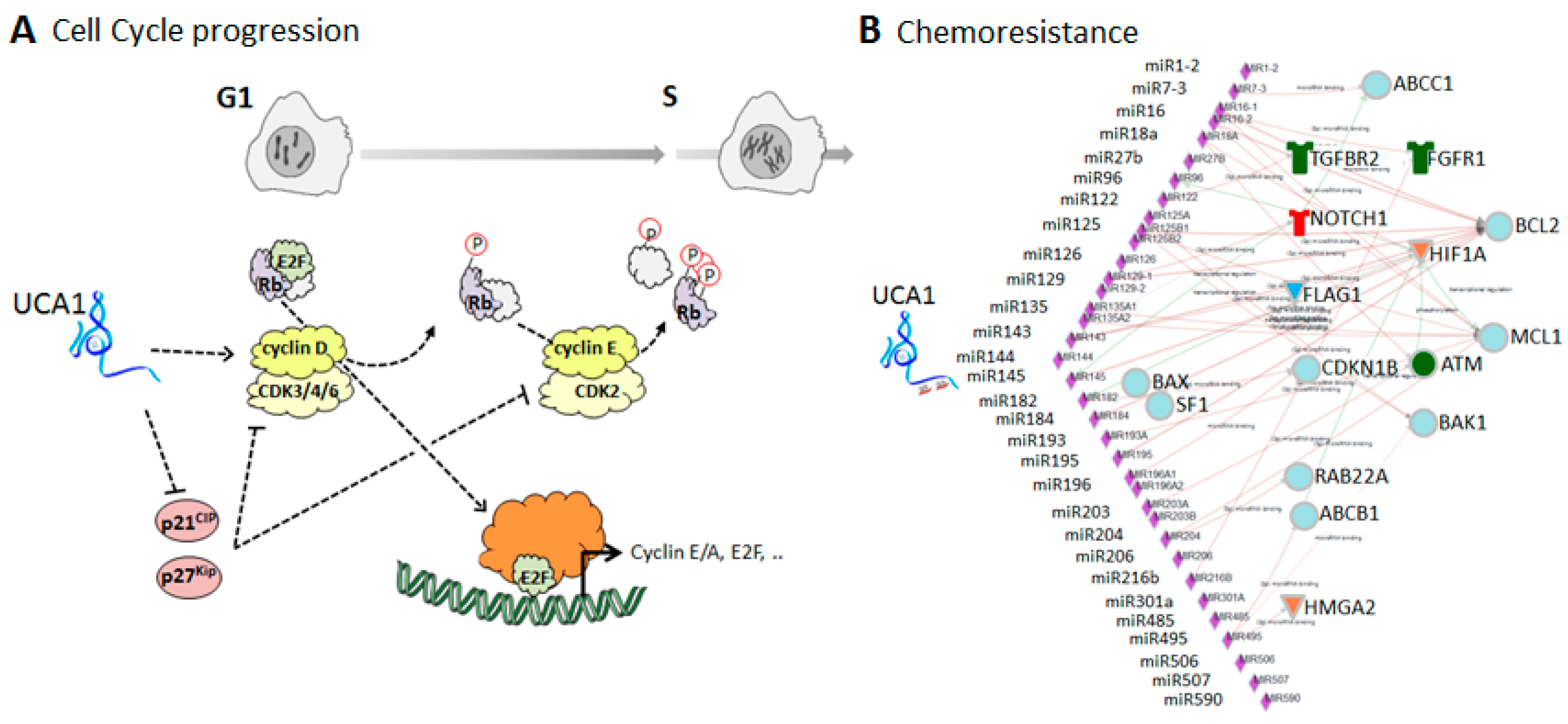
| lncRNA | References | |
|---|---|---|
| DNA organization | ||
| Speckle formation | ||
| NEAT1 | [23,24,25] | |
| MALAT1 | [25,27] | |
| Histone modification | ||
| EZH2 | AFAP1-AS1 | [36] |
| BLACAT1 | [37] | |
| CRNDE | [38] | |
| HOTAIR | [39] | |
| HOXA-AS2 | [31] | |
| HULC | [40] | |
| PINT | [41] | |
| SH3PXD2A-AS1 | [30] | |
| SNHG17 | [42] | |
| UCA1 | [43,44] | |
| WDR5 | HOTTIP | [45] |
| GClnc1 | [46] | |
| HOXD-AS | [47] | |
| LSD1 | HOTTAIR | [48] |
| FOXP4-AS1 | [49] | |
| HOXA-AS2 | [31] | |
| SWI/SNF chromatin modulation | ||
| BRG1 | lncTCF7 | [50] |
| lncFDZ6 | [51] | |
| NEAT1 | [21] | |
| UCA1 | [52] | |
| SNF5 | SChLAP1 | [34] |
| BAF200a | MVIH | [35] |
| RNA interaction | ||
| miRNA-precursors | ||
| miR-675 | H19 | [53,54] |
| miR-545/374a | Ftx | [55] |
| miR-143, -145 | NCR143/145 | [56] |
| miR-31 | LOC554202 | [57] |
| miR-125b-2, miR-99a and let-7c | MONC | [58] |
| miR-100, miR-125b-1 and let-7a-2 | MIR100HG | [58] |
| Let-7c, miR99a and miR125b | LINC00478 | [59] |
| Protein interaction | ||
| Protein stability | ||
| P53 | PANDA | [60] |
| SREBP-1c | MALAT1 | [61] |
| SLUG | SNHG15 | [62] |
| TCGA Cancer Classification | Total Patients Number (N); N in Low vs. High Risk Group | Log Rank Equal Curves | Hazard Ratio (95% CI) | p Value |
|---|---|---|---|---|
| Acute Myeloid Leukemia | N = 149; 138 vs. 11 | p = 0.77 | 1.12 (CI 0.52; 2.43) | p = 0.77 |
| Bile Duct Cholangiocarcinoma | N = 35; 16 vs. 19 | p = 0.15 | 2.06 (CI 0.75; 5.64) | p = 0.16 |
| Bladder—Urothelial Carcinoma | N = 389; 112 vs. 277 | p = 0.0034 | 1.75 (CI 1.2; 2.56) | p = 0.0038 |
| Breast invasive carcinoma—July 2016 | N = 962; 844 vs. 118 | p = 0.29 | 1.31 (CI 0.79; 2.15) | p = 0.29 |
| Cervical squamous cell carcinoma and endocervical adenocarcinoma | N = 191; 121 vs. 70 | p = 0.051 | 1.82 (CI 0.99; 3.35) | p = 0.054 |
| Colon and Rectum adenocarcinoma: | N = 422; 151 vs. 371 | p = 0.66 | 1.1 (CI 0.72; 1.69) | p = 0.66 |
| Colon | N = 350; 197 vs. 153 | p = 0.51 | 0.86 (CI 0.54; 1.36) | p = 0.51 |
| Rectum | N = 57; 39 vs. 18 | p = 0.0075 | 4.54 (CI 1.35; 15.27) | p = 0.014 |
| Esophageal carcinoma | N = 184; 148 vs. 36 | p = 0.29 | 0.72 (CI 0.38; 1.33) | p = 0.29 |
| Head and Neck squamous cell carcinoma | N = 506; 304 vs. 198 | p = 0.45 | 1.11 (CI 0.85; 1.46) | p = 0.45 |
| Kidney PAN cancer | N = 892; 715 vs. 77 | p = 0.67 | 1.11 (CI 0.68; 1.83) | p = 0.67 |
| Liver hepatocellular carcinoma | N = 361; 318 vs. 43 | p = 0.025 | 1.68 (CI 1.06; 2.66) | p = 0.027 |
| Lung adenocarcinoma | N = 475; 384 vs. 91 | p = 0.0041 | 1.69 (CI 1.18;2.44) | p = 0.0046 |
| Lung squamous cell carcinoma | N = 175; 123 vs. 52 | p = 0.93 | 0.98 (CI 0.61; 1.58) | p = 0.93 |
| Ovarian serous cystadenocarcinoma | N = 247; 25 vs. 222 | p = 0.21 | 0.72 (CI 0.43; 1.21) | p = 0.21 |
| Pancreatic adenocarcinoma | N = 176; 154 vs. 22 | p = 1.766 × 10-0.5 | 2.94 (CI 1.75; 4.92) | p = 4.249 × 10-0.5 |
| Stomach and Esophagous adenocarcinoma | N = 440; 220 vs. 220 | p = 0.90 | 0.98 (CI 0.72; 1.33) | p = 0.90 |
| Stomach adenocarcinoma | N = 352; 135 vs. 217 | p = 0.70 | 1.07 (CI 0.75; 1.52) | p = 0.70 |
| Testicular Germ Cell Tumors | N = 133; 105 vs. 28 | p = 0.19 | 3.39 (CI 0.48; 24.1) | p = 0.22 |
| Uterine Corpus Endometrial Carcinoma | N = 247; 130 vs. 117 | p = 0.084 | 1.85 (CI 0.91; 3.75) | p = 0.089 |
| Kaplan-Meier survival curve statistics are reported from the TCGA cohort data using the SurvExpress portal [113]. | ||||
| Interaction | Complex | Target | Cells | Ref. |
|---|---|---|---|---|
| Enhancer of zeste homolog 2 (EZH2) | polycomb repressive complex-2 | Cycline D1 | gastric cancer | [43] |
| p27Kip1 | hepatocarcinoma | [44] | ||
| CCCTC-binding factor (CTCF) | chromatin looping with RNA polII and P300 | HULC | Embryonic hepatocyte-like | [72] |
| Brahma related gene 1 (BRG1) | SWI/SNF chromatin remodeling | p21 | bladder cancer | [52] |
| MOB1, Lats1, and YAP | YAP-TEAD transcription complex | Hippo pathway | Pancreatic cancer | [107] |
| UCA1-Mediated miR Regulation (Sponges/Competing Endogenous RNA) | miR-Mediated Regulation | |||||
|---|---|---|---|---|---|---|
| miRNA * | miR-Target | Type of Cells | Targets ** | Biological Process *** | CRC | |
| miR-1 | Hes1 | neural stem cell | [132] | 915 | ||
| bladder cancer cells | [125] | |||||
| Slug | Breast cancer | [115] | ||||
| miR-7 | EGFR | gastric cancer | [133] | 884 | transmembrane receptor protein tyrosine kinase signaling pathway | [134,135,136,137,138,139,140,141] |
| miR-16 | MDR1 | chronic myeloid leukemia | [142] | 1646 | protein folding, rRNA metabolic process, tRNA aminoacylation for protein translation, protein acetylation, regulation of sequence-specific DNA binding transcription factor activity, nuclear transport, nucleobase-containing compound transport, tRNA metabolic process, RNA localization, protein targeting, cellular component biogenesis | [143] |
| GLS2 | bladder cancer | [144] | ||||
| miR-18a | YAP | breast cancer | [145] | N.A. | N.A. | [146] |
| HIF1α | breast cancer | [147] | ||||
| miR-27b | gastric cancer | [148] | 447 | regulation of cell cycle, intracellular protein transport | [149] | |
| miR-122 | breast cancer | [124] | 580 | N.S. | [150] | |
| miR-125 | HK2 | acute myeloid leukemia | [151] | 899 | ||
| miR-126 | RAC1 | human myelogenous leukemia | [152] | 152 | N.S. | - |
| miR-129 | SOX4 | renal cell carcinoma | [153] | 499 | N.S. | [154,155,156,157,158,159] |
| ABCB1 | ovarian cancer | [160] | ||||
| miR-135a | pancreatic cancer | [106] | 121 | N.S. | [161] | |
| cMYC | thyroid cancer | [162] | ||||
| miR-143 | mTOR (cyclin D1, p27) | colorectal cancer | [130] | 478 | N.S. | [130,163] |
| ERBB3 BCL-2 | breast cancer | [164] | ||||
| FOXO1 | cardiomyocyte | [165] | ||||
| HMGB1 | bladder cancer | [166] | ||||
| HK2 | bladder cancer | [129] | ||||
| miR-145 | FSCN1 ZEB1/2 | bladder cancer | [167] | 263 | cell proliferation, cytokinesis, negative regulation of apoptotic process, anatomical structure morphogenesis, regulation of cell cycle, MAPK cascade | [168,169] |
| miR-182 | p53 (iASPP) | Glioma | [170] | 189 | N.S. | [171] |
| PFKFB2 | glioblastoma-associated stromal cells | [172] | ||||
| miR-184 | SF1 | oral squamous cell carcinoma | [173] | 29 | N.S. | [174] |
| miR-193a | HMGB1 | lung cancer | [175] | 144 | N.S. | [176,177,178,179,180] |
| ERBB4 | non-small cell lung cancer | [181] | ||||
| miR-195 | ARL2 | bladder Cancer | [182] | 692 | angiogenesis, cell proliferation, cytokinesis, anatomical structure morphogenesis, mitosis, regulation of transcription from RNA polymerase II promoter | [183,184,185,186,187,188] |
| miR-196a | CREB | bladder cancer | [189] | 450 | RNA splicing, via transesterification reactions, response to stress, organelle organization | [190,191,192,193] |
| miR-203 | Snail2 | hepatocellular carcinoma | [194] | 528 | N.S. | [195,196,197,198,199,200] |
| Slug | Breast cancer | [115] | ||||
| miR-204 | CREB1, BCL2, RAB22A | colorectal cancer | [110] | 488 | N.S. | [110,201,202,203] |
| MMP-13 | chondrocytes | [204] | ||||
| Sirt1 | prostate cancer | [205] | ||||
| Sox4 | esophageal cancer | [206] | ||||
| BRD4 | thyroid cancer | [207] | ||||
| miR-206 | VEGF | cervical cancer | [208] | 95 | pentose-phosphate shunt, chromatin assembly, chromatin remodeling, negative regulation of apoptotic process, chromatin organization, regulation of phosphate, metabolic process | - |
| miR-216b | FGFR1 | hepatocellular carcinoma | [209] | 235 | protein targeting | [209,210,211,212,213] |
| miR-485 | MMP14 | epithelial ovarian | [214] | 505 | N.S. | - |
| miR-495 | p21 | renal cell carcinoma | [215] | 241 | N.S. | [216,217] |
| miR-506 | COTL1 | non-small cell lung cancer | [218] | 180 | N.S. | [219,220,221,222,223,224] |
| miR-507 | FOXM1 | melanoma | [225] | 169 | N.S. | - |
| miR-590 | CREB | gastric cancer | [226] | 419 | N.S. | [227,228,229,230] |
| miR-22 | - | - | - | 221 | negative regulation of apoptotic process, regulation of transcription from RNA polymerase II promoter | [231,232,233,234,235,236,237,238,239,240,241] |
| miR-23a | - | - | - | 353 | N.S. | [242,243,244,245,246,247] |
| miR-26a | - | - | - | 531 | Mitosis, regulation of cell cycle, phosphate-containing compound, metabolic process | [248,249,250,251,252,253,254,255,256] |
| miR-103a/107/107ab | - | - | - | 857 | cytoskeleton organization, cell cycle | [257,258,259] |
| miR-124 | - | - | - | 1520 | regulation of binding, cytokinesis, transmembrane receptor, protein tyrosine kinase signaling pathway, regulation of cell cycle | [219,255,260,261,262,263,264,265,266,267,268,269,270,271,272,273,274] |
| miR-138 | - | - | - | 239 | N.S. | [275,276,277,278] |
| miR-190 | - | - | - | 770 | RNA splicing, via transesterification reactions | - |
| miR-214 | - | - | - | 352 | N.S. | [279,280,281,282,283] |
| miR-455 | - | - | - | 557 | N.S. | [284,285,286,287] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Neve, B.; Jonckheere, N.; Vincent, A.; Van Seuningen, I. Epigenetic Regulation by lncRNAs: An Overview Focused on UCA1 in Colorectal Cancer. Cancers 2018, 10, 440. https://doi.org/10.3390/cancers10110440
Neve B, Jonckheere N, Vincent A, Van Seuningen I. Epigenetic Regulation by lncRNAs: An Overview Focused on UCA1 in Colorectal Cancer. Cancers. 2018; 10(11):440. https://doi.org/10.3390/cancers10110440
Chicago/Turabian StyleNeve, Bernadette, Nicolas Jonckheere, Audrey Vincent, and Isabelle Van Seuningen. 2018. "Epigenetic Regulation by lncRNAs: An Overview Focused on UCA1 in Colorectal Cancer" Cancers 10, no. 11: 440. https://doi.org/10.3390/cancers10110440
APA StyleNeve, B., Jonckheere, N., Vincent, A., & Van Seuningen, I. (2018). Epigenetic Regulation by lncRNAs: An Overview Focused on UCA1 in Colorectal Cancer. Cancers, 10(11), 440. https://doi.org/10.3390/cancers10110440







