Differential Gene Expression Analysis of Bovine Macrophages after Exposure to the Penicillium Mycotoxins Citrinin and/or Ochratoxin A
Abstract
1. Introduction
2. Results and Discussion
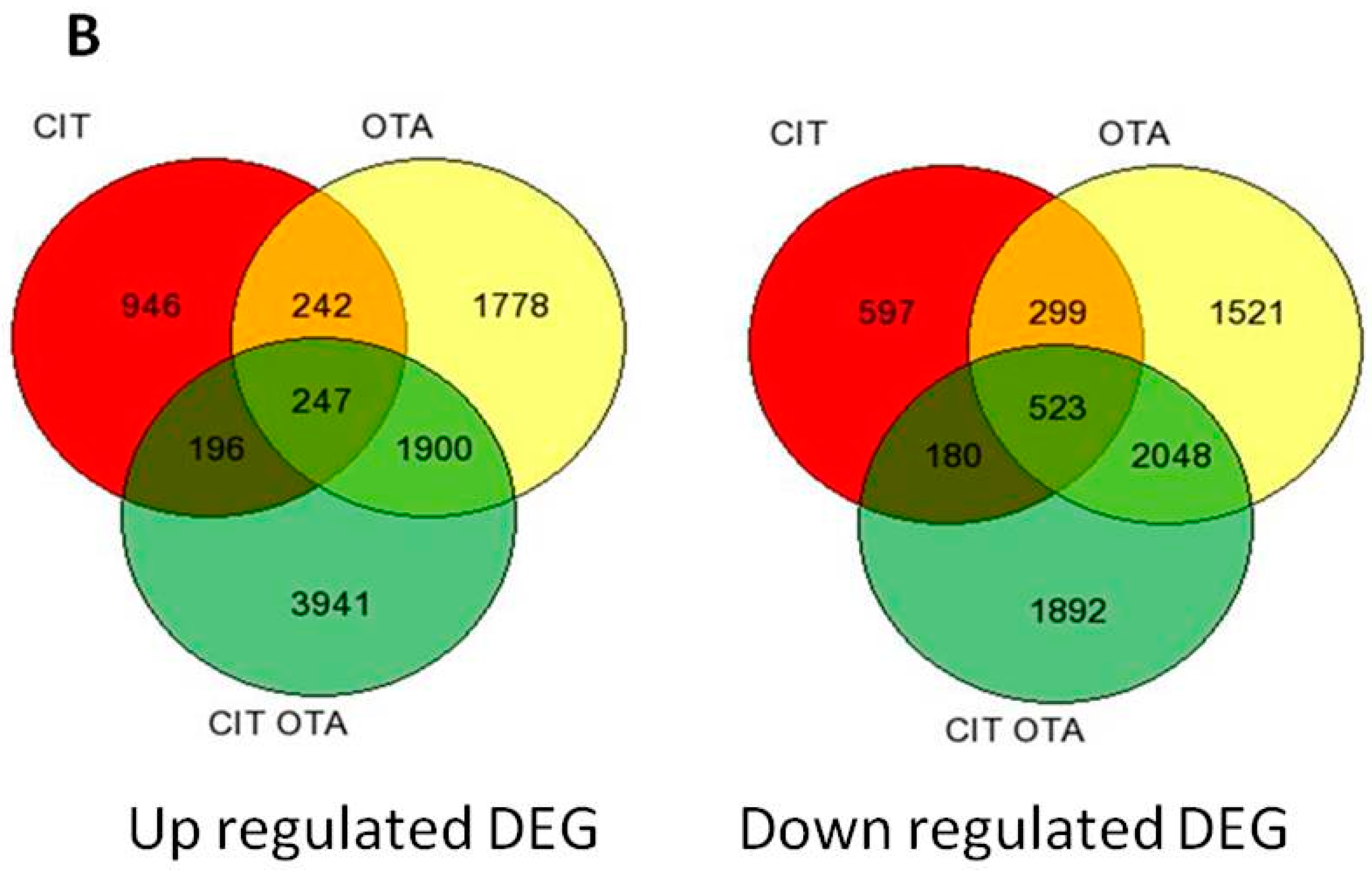
2.1. Summary of Differentially-Expressed Genes Identified from Microarray Analysis
2.2. Transcriptional Changes after 6 h of Exposure to PM
2.3. Transcriptional Changes after 24 h of Exposure to PM
3. Conclusions
4. Materials and Methods
4.1. Cell Culture and Mycotoxin Exposure
4.2. RNA Isolation and Microarray Analysis
4.3. Ingenuity Pathway Analysis (IPA)
4.4. Real-Time PCR
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Ostry, V.; Malir, F.; Ruprich, J. Producers and important dietary sources of ochratoxin A and citrinin. Toxins (Basel) 2013, 5, 1574–1586. [Google Scholar] [CrossRef] [PubMed]
- Bennett, J.W.; Klich, M. Mycotoxins. Clin. Microbiol. Rev. 2003, 16, 497–516. [Google Scholar] [CrossRef] [PubMed]
- Diekman, M.A.; Green, M.L. Mycotoxins and reproduction in domestic livestock. J. Anim. Sci. 1992, 70, 1615–1627. [Google Scholar] [CrossRef] [PubMed]
- Bennett, J.W. Mycotoxin research: 1989. Mycopathologia 1989, 107, 65–66. [Google Scholar] [CrossRef] [PubMed]
- Kuiper-Goodman, T.; Scott, P.M. Risk assessment of the mycotoxin ochratoxin A. Biomed. Environ. Sci. 1989, 2, 179–248. [Google Scholar] [PubMed]
- Razafimanjato, H.; Garmy, N.; Guo, X.J.; Varini, K.; Di Scala, C.; Di Pasquale, E.; Taieb, N.; Maresca, M. The food-associated fungal neurotoxin ochratoxin A inhibits the absorption of glutamate by astrocytes through a decrease in cell surface expression of the excitatory amino-acid transporters glast and GLT-1. Neurotoxicology 2010, 31, 475–484. [Google Scholar] [CrossRef] [PubMed]
- Maresca, M.; Yahi, N.; Younes-Sakr, L.; Boyron, M.; Caporiccio, B.; Fantini, J. Both direct and indirect effects account for the pro-inflammatory activity of enteropathogenic mycotoxins on the human intestinal epithelium: Stimulation of interleukin-8 secretion, potentiation of interleukin-1beta effect and increase in the transepithelial passage of commensal bacteria. Toxicol. Appl. Pharmacol. 2008, 228, 84–92. [Google Scholar] [PubMed]
- Marquardt, R.R.; Frohlich, A.A. A review of recent advances in understanding ochratoxicosis. J. Anim. Sci. 1992, 70, 3968–3988. [Google Scholar] [CrossRef] [PubMed]
- Meisner, H.; Meisner, P. Ochratoxin a, an in vivo inhibitor of renal phosphoenolpyruvate carboxykinase. Arch. Biochem. Biophys. 1981, 208, 146–153. [Google Scholar] [CrossRef]
- Rahimtula, A.D.; Bereziat, J.C.; Bussacchini-Griot, V.; Bartsch, H. Lipid peroxidation as a possible cause of ochratoxin A toxicity. Biochem. Pharmacol. 1988, 37, 4469–4477. [Google Scholar] [CrossRef]
- Flajs, D.; Peraica, M. Toxicological properties of citrinin. Arh. Hig. Rada Toksikol. 2009, 60, 457–464. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, S.M.; Chagas, G.M.; Campello, A.P.; Kluppel, M.L. Mechanism of citrinin-induced dysfunction of mitochondria. V. Effect on the homeostasis of the reactive oxygen species. Cell Biochem. Funct. 1997, 15, 203–209. [Google Scholar] [CrossRef]
- Keblys, M.; Bernhoft, A.; Hofer, C.C.; Morrison, E.; Larsen, H.J.; Flaoyen, A. The effects of the penicillium mycotoxins citrinin, cyclopiazonic acid, ochratoxin A, patulin, penicillic acid, and roquefortine C on in vitro proliferation of porcine lymphocytes. Mycopathologia 2004, 158, 317–324. [Google Scholar] [CrossRef] [PubMed]
- Heussner, A.H.; Dietrich, D.R.; O'Brien, E. In vitro investigation of individual and combined cytotoxic effects of ochratoxin A and other selected mycotoxins on renal cells. Toxicol. In Vitro 2006, 20, 332–341. [Google Scholar] [CrossRef] [PubMed]
- Vesela, D.; Vesely, D.; Jelinek, R. Toxic effects of ochratoxin A and citrinin, alone and in combination, on chicken embryos. Appl. Environ. Microbiol. 1983, 45, 91–93. [Google Scholar] [PubMed]
- Oh, S.Y.; Boermans, H.J.; Swamy, H.V.L.N.; Sharma, B.S.; Karrow, N.A. Immunotoxicity of penicillium mycotoxins on viability and proliferation of bovine macrophage cell line (BOMACS). Mycol. J. 2012, 6, 11–16. [Google Scholar]
- Speijers, G.J.; Speijers, M.H. Combined toxic effects of mycotoxins. Toxicol. Lett. 2004, 153, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Muller, G.; Rosner, H.; Rohrmann, B.; Erler, W.; Geschwend, G.; Grafe, U.; Burkert, B.; Moller, U.; Diller, R.; Sachse, K.; et al. Effects of the mycotoxin ochratoxin A and some of its metabolites on the human cell line THP-1. Toxicology 2003, 184, 69–82. [Google Scholar] [CrossRef]
- Duhley, J.N. Effects of some indian herbs on macrophage functions in ochratoxin A treated mice. J. Ethnopharmacol. 1997, 58, 15–20. [Google Scholar] [CrossRef]
- Al-Anati, L.; Petzinger, E. Immunotoxic activity of ochratoxin A. J. Vet. Pharmacol. Ther. 2006, 29, 79–90. [Google Scholar] [CrossRef] [PubMed]
- Wichmann, G.; Herbarth, O.; Lehmann, I. The mycotoxins citrinin, gliotoxin, and patulin affect interferon-gamma rather than interleukin-4 production in human blood cells. Environ. Toxicol. 2002, 17, 211–218. [Google Scholar] [CrossRef] [PubMed]
- Marin-Kuan, M.; Cavin, C.; Delatour, T.; Schilter, B. Ochratoxin a carcinogenicity involves a complex network of epigenetic mechanisms. Toxicon 2008, 52, 195–202. [Google Scholar] [CrossRef] [PubMed]
- Oh, S.Y.; Balch, C.G.; Cliff, R.L.; Sharma, B.S.; Boermans, H.J.; Swamy, H.V.; Quinton, V.M.; Karrow, N.A. Exposure to penicillium mycotoxins alters gene expression of enzymes involved in the epigenetic regulation of bovine macrophages (BOMACS). Mycotoxin Res. 2013, 29, 235–243. [Google Scholar] [CrossRef] [PubMed]
- Sansing, G.A.; Lillehoj, E.B.; Detroy, R.W.; Miller, M.A. Synergistic toxic effects of citrinin, ochratoxin A and penicillic acid in mice. Toxicon 1976, 14, 213–220. [Google Scholar] [CrossRef]
- Baek, J.Y.; Han, S.H.; Sung, S.H.; Lee, H.E.; Kim, Y.M.; Noh, Y.H.; Bae, S.H.; Rhee, S.G.; Chang, T.S. Sulfiredoxin protein is critical for redox balance and survival of cells exposed to low steady-state levels of H2O2. J. Biol. Chem. 2012, 287, 81–89. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Li, Y.; Chen, L.; Chan, T.H.; Song, Y.; Fu, L.; Zeng, T.T.; Dai, Y.D.; Zhu, Y.H.; Chen, J.; et al. Allele-specific imbalance of oxidative stress-induced growth inhibitor 1 associates with progression of hepatocellular carcinoma. Gastroenterology 2014, 146, 1084–1096. [Google Scholar] [CrossRef] [PubMed]
- Raphael, A.R.; Couthouis, J.; Sakamuri, S.; Siskind, C.; Vogel, H.; Day, J.W.; Gitler, A.D. Congenital muscular dystrophy and generalized epilepsy caused by gmppb mutations. Brain Res. 2014, 1575, 66–71. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Floyd-Smith, G.; Slattery, E.; Lengyel, P. Interferon action: Rna cleavage pattern of a (2′-5′)oligoadenylate—Dependent endonuclease. Science 1981, 212, 1030–1032. [Google Scholar] [CrossRef] [PubMed]
- Zabielska, M.A.; Borkowski, T.; Slominska, E.M.; Smolenski, R.T. Inhibition of amp deaminase as therapeutic target in cardiovascular pathology. Pharmacol. Rep. 2015, 67, 682–688. [Google Scholar] [CrossRef] [PubMed]
- Holness, C.L.; Simmons, D.L. Molecular cloning of cd68, a human macrophage marker related to lysosomal glycoproteins. Blood 1993, 81, 1607–1613. [Google Scholar] [PubMed]
- Song, L.; Lee, C.; Schindler, C. Deletion of the murine scavenger receptor CD68. J. Lipid Res. 2011, 52, 1542–1550. [Google Scholar] [CrossRef] [PubMed]
- Mungrue, I.N.; Pagnon, J.; Kohannim, O.; Gargalovic, P.S.; Lusis, A.J. CHAC1/MGC4504 is a novel proapoptotic component of the unfolded protein response, downstream of the ATF4-ATF3-CHOP cascade. J. Immunol. 2009, 182, 466–476. [Google Scholar] [CrossRef] [PubMed]
- Crawford, R.R.; Prescott, E.T.; Sylvester, C.F.; Higdon, A.N.; Shan, J.; Kilberg, M.S.; Mungrue, I.N. Human CHAC1 protein degrades glutathione, and mRNA induction is regulated by the transcription factors ATF4 and ATF3 and a bipartite atf/cre regulatory element. J. Biol. Chem. 2015, 290, 15878–15891. [Google Scholar] [CrossRef] [PubMed]
- Meng, Q.; Jia, Z.; Wang, W.; Li, B.; Ma, K.; Zhou, C. Inhibitor of DNA binding 1 (Id1) induces differentiation and proliferation of mouse embryonic carcinoma P19CL6 cells. Biochem. Biophys. Res. Commun. 2011, 412, 253–259. [Google Scholar] [CrossRef] [PubMed]
- Ho, D.V.; Chan, J.Y. Induction of herpud1 expression by er stress is regulated by NRF1. FEBS Lett. 2015, 589, 615–620. [Google Scholar] [CrossRef] [PubMed]
- Chan, W.H. Citrinin induces apoptosis via a mitochondria-dependent pathway and inhibition of survival signals in embryonic stem cells, and causes developmental injury in blastocysts. Biochem. J. 2007, 404, 317–326. [Google Scholar] [CrossRef] [PubMed]
- Yu, F.Y.; Liao, Y.C.; Chang, C.H.; Liu, B.H. Citrinin induces apoptosis in HL-60 cells via activation of the mitochondrial pathway. Toxicol. Lett. 2006, 161, 143–151. [Google Scholar] [CrossRef] [PubMed]
- Alberts, B.J.A.; Lewis, J.; Raff, M.; Roberts, K.; Walker, P. Molecular Biology of the Cell, 5th ed.; Garland Science: New York, NY, USA, 2007. [Google Scholar]
- Kulms, D.; Dussmann, H.; Poppelmann, B.; Stander, S.; Schwarz, A.; Schwarz, T. Apoptosis induced by disruption of the actin cytoskeleton is mediated via activation of CD95 (FAS/APO-1). Cell Death Differ. 2002, 9, 598–608. [Google Scholar] [CrossRef] [PubMed]
- Luhe, A.; Hildebrand, H.; Bach, U.; Dingermann, T.; Ahr, H.J. A new approach to studying ochratoxin A (OTA)-induced nephrotoxicity: Expression profiling in vivo and in vitro employing cdna microarrays. Toxicol. Sci. 2003, 73, 315–328. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.; Ansari, K.M.; Chaudhari, B.P.; Dhawan, A.; Dwivedi, P.D.; Jain, S.K.; Das, M. Topical application of ochratoxin A causes DNA damage and tumor initiation in mouse skin. PLoS ONE 2012, 7, e47280. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.; Dwivedi, P.D.; Dhawan, A.; Das, M.; Ansari, K.M. Citrinin-generated reactive oxygen species cause cell cycle arrest leading to apoptosis via the intrinsic mitochondrial pathway in mouse skin. Toxicol. Sci. 2011, 122, 557–566. [Google Scholar] [CrossRef] [PubMed]
- Apel, K.; Hirt, H. Reactive oxygen species: Metabolism, oxidative stress, and signal transduction. Annu. Rev. Plant Biol. 2004, 55, 373–399. [Google Scholar] [CrossRef] [PubMed]
- Donepudi, M.; Grutter, M.G. Structure and zymogen activation of caspases. Biophys. Chem. 2002, 101–102, 145–153. [Google Scholar] [CrossRef]
- Charoenpornsook, K.; Fitzpatrick, J.L.; Smith, J.E. The effects of four mycotoxins on the mitogen stimulated proliferation of bovine peripheral blood mononuclear cells in vitro. Mycopathologia 1998, 143, 105–111. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Wang, Y.; Cui, J.; Xing, L.; Shen, H.; Wu, S.; Lian, H.; Wang, J.; Yan, X.; Zhang, X. Ochratoxin a induces oxidative DNA damage and g1 phase arrest in human peripheral blood mononuclear cells in vitro. Toxicol. Lett. 2012, 211, 164–171. [Google Scholar] [CrossRef] [PubMed]
- Wei, Y.H.; Lu, C.Y.; Lin, T.N.; Wei, R.D. Effect of ochratoxin A on rat liver mitochondrial respiration and oxidative phosphorylation. Toxicology 1985, 36, 119–130. [Google Scholar] [CrossRef]
- Shao, J.; Berger, L.F.; Hendriksen, P.J.; Peijnenburg, A.A.; van Loveren, H.; Volger, O.L. Transcriptome-based functional classifiers for direct immunotoxicity. Arch. Toxicol. 2014, 88, 673–689. [Google Scholar] [CrossRef] [PubMed]
- Friggeri, A.; Banerjee, S.; Xie, N.; Cui, H.; De Freitas, A.; Zerfaoui, M.; Dupont, H.; Abraham, E.; Liu, G. Extracellular histones inhibit efferocytosis. Mol. Med. 2012, 18, 825–833. [Google Scholar] [CrossRef] [PubMed]
- So, M.Y.; Tian, Z.; Phoon, Y.S.; Sha, S.; Antoniou, M.N.; Zhang, J.; Wu, R.S.; Tan-Un, K.C. Gene expression profile and toxic effects in human bronchial epithelial cells exposed to zearalenone. PLoS ONE 2014, 9, e96404. [Google Scholar] [CrossRef] [PubMed]
- Kirmes, I.; Szczurek, A.; Prakash, K.; Charapitsa, I.; Heiser, C.; Musheev, M.; Schock, F.; Fornalczyk, K.; Ma, D.; Birk, U.; et al. A transient ischemic environment induces reversible compaction of chromatin. Genome Biol. 2015, 16, 246. [Google Scholar] [CrossRef] [PubMed]
- Muharram, G.; Sahgal, P.; Korpela, T.; De Franceschi, N.; Kaukonen, R.; Clark, K.; Tulasne, D.; Carpen, O.; Ivaska, J. Tensin-4-dependent met stabilization is essential for survival and proliferation in carcinoma cells. Dev. Cell 2014, 29, 421–436. [Google Scholar] [CrossRef] [PubMed]
- Xie, J.; Yuan, Y.; Liu, Z.; Xiao, Y.; Zhang, X.; Qin, C.; Sheng, Z.; Xu, T.; Wang, X. CMTM3 is frequently reduced in clear cell renal cell carcinoma and exhibits tumor suppressor activities. Clin. Transl. Oncol. 2014, 16, 402–409. [Google Scholar] [CrossRef] [PubMed]
- Gayathri, L.; Dhivya, R.; Dhanasekaran, D.; Periasamy, V.S.; Alshatwi, A.A.; Akbarsha, M.A. Hepatotoxic effect of ochratoxin A and citrinin, alone and in combination, and protective effect of vitamine: In vitro study in HepG2 cell. Food Chem. Toxicol. 2015, 83, 151–163. [Google Scholar] [CrossRef] [PubMed]
- Anninou, N.; Chatzaki, E.; Papachristou, F.; Pitiakoudis, M.; Simopoulos, C. Mycotoxins′ activity at toxic and sub-toxic concentrations: Differential cytotoxic and genotoxic effects of single and combined administration of sterigmatocystin, ochratoxin A and citrinin on the hepatocellular cancer cell line HeP3B. Int. J. Environ. Res. Public Health 2014, 11, 1855–1872. [Google Scholar] [CrossRef] [PubMed]
- Endo, A.; Kuroda, M. Citrinin, an inhibitor of cholesterol synthesis. J. Antibiot. (Tokyo) 1976, 29, 841–843. [Google Scholar] [CrossRef] [PubMed]
- Kuroda, M.; Hazama-Shimada, Y.; Endo, A. Inhibition of sterol synthesis by citrinin in a cell-free system from rat liver and yeast. Biochim. Biophys. Acta 1977, 486, 254–259. [Google Scholar] [PubMed]
- Idzko, M.; Ferrari, D.; Eltzschig, H.K. Nucleotide signalling during inflammation. Nature 2014, 509, 310–317. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.; Jang, H.J.; Kim, J.K.; Kim, J.M.; Park, E.H.; Yang, J.H.; Kim, Y.H.; Yea, K.; Ryu, S.H.; Suh, P.G. Ochratoxin a inhibits adipogenesis through the extracellular signal-related kinases-peroxisome proliferator-activated receptor-gamma pathway in human adipose tissue-derived mesenchymal stem cells. Stem Cells Dev. 2011, 20, 415–426. [Google Scholar] [CrossRef] [PubMed]
- Zurich, M.G.; Lengacher, S.; Braissant, O.; Monnet-Tschudi, F.; Pellerin, L.; Honegger, P. Unusual astrocyte reactivity caused by the food mycotoxin ochratoxin A in aggregating rat brain cell cultures. Neuroscience 2005, 134, 771–782. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Pascual, G.; Glass, C.K. Peroxisome proliferator-activated receptor gamma-dependent repression of the inducible nitric oxide synthase gene. Mol. Cell. Biol. 2000, 20, 4699–4707. [Google Scholar] [CrossRef] [PubMed]
- Shrestha, N.; Bahnan, W.; Wiley, D.J.; Barber, G.; Fields, K.A.; Schesser, K. Eukaryotic initiation factor 2 (eIF2) signaling regulates proinflammatory cytokine expression and bacterial invasion. J. Biol. Chem. 2012, 287, 28738–28744. [Google Scholar] [CrossRef] [PubMed]
- Woltman, A.M.; van der Kooij, S.W.; Coffer, P.J.; Offringa, R.; Daha, M.R.; van Kooten, C. Rapamycin specifically interferes with GM-CSF signaling in human dendritic cells, leading to apoptosis via increased p27KIP1 expression. Blood 2003, 101, 1439–1445. [Google Scholar] [CrossRef] [PubMed]
- Mobashar, M.; Hummel, J.; Blank, R.; Sudekum, K.H. Ochratoxin A in ruminants—A review on its degradation by gut microbes and effects on animals. Toxins (Basel) 2010, 2, 809–839. [Google Scholar] [CrossRef] [PubMed]
- Sreemannarayana, O.; Frohlich, A.A.; Vitti, T.G.; Marquardt, R.R.; Abramson, D. Studies of the tolerance and disposition of ochratoxin A in young calves. J. Anim. Sci. 1988, 66, 1703–1711. [Google Scholar] [CrossRef] [PubMed]
- Xiao, H.; Marquardt, R.R.; Frohlich, A.A.; Phillips, G.D.; Vitti, T.G. Effect of a hay and a grain diet on the bioavailability of ochratoxin A in the rumen of sheep. J. Anim. Sci. 1991, 69, 3715–3723. [Google Scholar] [CrossRef] [PubMed]
- Blank, R.; Rolfs, J.P.; Sudekum, K.H.; Frohlich, A.A.; Marquardt, R.R.; Wolffram, S. Effects of chronic ingestion of ochratoxin A on blood levels and excretion of the mycotoxin in sheep. J. Agric. Food Chem. 2003, 51, 6899–6905. [Google Scholar] [CrossRef] [PubMed]
- Hohler, D.; Sudekum, K.H.; Wolffram, S.; Frohlich, A.A.; Marquardt, R.R. Metabolism and excretion of ochratoxin A fed to sheep. J. Anim. Sci. 1999, 77, 1217–1223. [Google Scholar] [CrossRef] [PubMed]
- Galtier, P.; Alvinerie, M.; Charpenteau, J.L. The pharmacokinetic profiles of ochratoxin A in pigs, rabbits and chickens. Food Cosmet. Toxicol 1981, 19, 735–738. [Google Scholar] [CrossRef]
- Prelusky, D.B.; Hamilton, R.M.; Trenholm, H.L.; Miller, J.D. Tissue distribution and excretion of radioactivity following administration of 14C-labeled deoxynivalenol to white Leghorn hens. Fundam. Appl. Toxicol. 1986, 7, 635–645. [Google Scholar] [CrossRef]
- Stabel, J.R.; Stabel, T.J. Immortalization and characterization of bovine peritoneal macrophages transfected with SV40 plasmid DNA. Vet. Immunol. Immunopathol. 1995, 45, 211–220. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]

| Bio-Function-Associated Genes 1 | Number of Altered Genes |
|---|---|
| Citrinin | |
| Cell death and survival | 339 |
| Cellular compromise | 55 |
| Cellular function and maintenance | 104 |
| Cellular movement | 56 |
| Cell cycle | 22 |
| Ochratoxin A | |
| Gene expression | 759 |
| Cell cycle | 254 |
| Cellular assembly and organization | 55 |
| DNA replication, recombination and repair | 166 |
| Energy production | 13 |
| Citrinin + Ochratoxin A | |
| DNA replication, recombination and repair | 290 |
| Gene expression | 928 |
| Cell cycle | 566 |
| Post-translational modification | 217 |
| Cellular development | 866 |
| Most-Altered Canonical Pathways 1 | Ratio 2 | p Value |
|---|---|---|
| Citrinin | ||
| NRF2-mediated oxidative stress | 25/130 (0.192) | *** |
| Endoplasmic reticulum stress pathway | 7/16 (0.438) | *** |
| Induction of apoptosis by HIV | 11/51 (0.216) | *** |
| Unfolded protein response | 9/41 (0.22) | *** |
| Glucocorticoid receptor signaling | 26/204 (0.127) | *** |
| Ochratoxin A | ||
| Role of RIG1-like receptors in antiviral innate immunity | 14/25 (0.56) | * |
| Eicosanoid signaling | 17/34 (0.50) | * |
| Oxidative phosphorylation | 30/78 (0.385) | * |
| April mediated signaling | 14/31 (0.452) | * |
| Phospholipases | 14/31 (0.452) | * |
| Citrinin + Ochratoxin A | ||
| Role of PKR in interferon induction and antiviral response | 20/34 (0.588) | *** |
| NF-κB signaling | 55/129 (0.426) | *** |
| TGF-B signaling | 29/60 (0.483) | ** |
| P53 signaling | 31/69 (0.449) | ** |
| Glioma signaling | 32/72 (0.444) | ** |
| Bio-Function-Associated Genes 1 | Number of Altered Genes |
|---|---|
| Citrinin | |
| Cell death and survival | 814 |
| Cellular movement | 520 |
| Lipid metabolism | 229 |
| Small molecule biochemistry | 397 |
| Cellular growth and proliferation | 846 |
| Ochratoxin A | |
| DNA replication, recombination and repair | 272 |
| Gene Expression | 97 |
| RNA post-transcriptional modification | 109 |
| Cellular development | 72 |
| Cell signaling | 85 |
| Citrinin + Ochratoxin A | |
| Cell cycle | 603 |
| Cellular growth and proliferation | 1368 |
| Gene expression | 858 |
| Post-translational modification | 539 |
| Cellular movement | 762 |
| Most-Altered Canonical Pathways 1 | Ratio 2 | p Value 3 |
|---|---|---|
| Citrinin | ||
| Superpathway of cholesterol biosynthesis | 17/23 (0.739) | *** |
| Cholesterol biosynthesis I | 8/10 (0.80) | *** |
| Cholesterol biosynthesis II (via 24, 25-dihydrolanosterol) | 8/10 (0.80) | *** |
| Cholesterol biosynthesis III (via desmosterol) | 8/10 (0.80) | *** |
| tRNA charging | 16/32 (0.5) | *** |
| Ochratoxin A | ||
| P2Y purigenic receptor signaling pathway | 43/93 | ** |
| Breast cancer regulation by Stathmin1 | 58/137 | ** |
| PPAR signaling | 33/69 (0.478) | ** |
| CREB signaling in neurons | 50/117 (0.427) | ** |
| GADD45 Signaling | 11/17 (0.647) | ** |
| Citrinin + Ochratoxin A | ||
| EIF2 signaling | 85/182 (0.443) | *** |
| Regulation of eIF4 and p70S6K signaling | 57/146 (0.39) | *** |
| Molecular mechanisms of cancer | 114/365 (0.312) | *** |
| mTOR signaling | 67/188 (0.356) | *** |
| Germ cell-Sertoli cell junction signaling | 56/160 (0.35) | *** |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Brennan, K.M.; Oh, S.-Y.; Yiannikouris, A.; Graugnard, D.E.; Karrow, N.A. Differential Gene Expression Analysis of Bovine Macrophages after Exposure to the Penicillium Mycotoxins Citrinin and/or Ochratoxin A. Toxins 2017, 9, 366. https://doi.org/10.3390/toxins9110366
Brennan KM, Oh S-Y, Yiannikouris A, Graugnard DE, Karrow NA. Differential Gene Expression Analysis of Bovine Macrophages after Exposure to the Penicillium Mycotoxins Citrinin and/or Ochratoxin A. Toxins. 2017; 9(11):366. https://doi.org/10.3390/toxins9110366
Chicago/Turabian StyleBrennan, Kristen M., Se-Young Oh, Alexandros Yiannikouris, Daniel E. Graugnard, and Niel A. Karrow. 2017. "Differential Gene Expression Analysis of Bovine Macrophages after Exposure to the Penicillium Mycotoxins Citrinin and/or Ochratoxin A" Toxins 9, no. 11: 366. https://doi.org/10.3390/toxins9110366
APA StyleBrennan, K. M., Oh, S.-Y., Yiannikouris, A., Graugnard, D. E., & Karrow, N. A. (2017). Differential Gene Expression Analysis of Bovine Macrophages after Exposure to the Penicillium Mycotoxins Citrinin and/or Ochratoxin A. Toxins, 9(11), 366. https://doi.org/10.3390/toxins9110366






