Precision Nutrition: A Review of Personalized Nutritional Approaches for the Prevention and Management of Metabolic Syndrome
Abstract
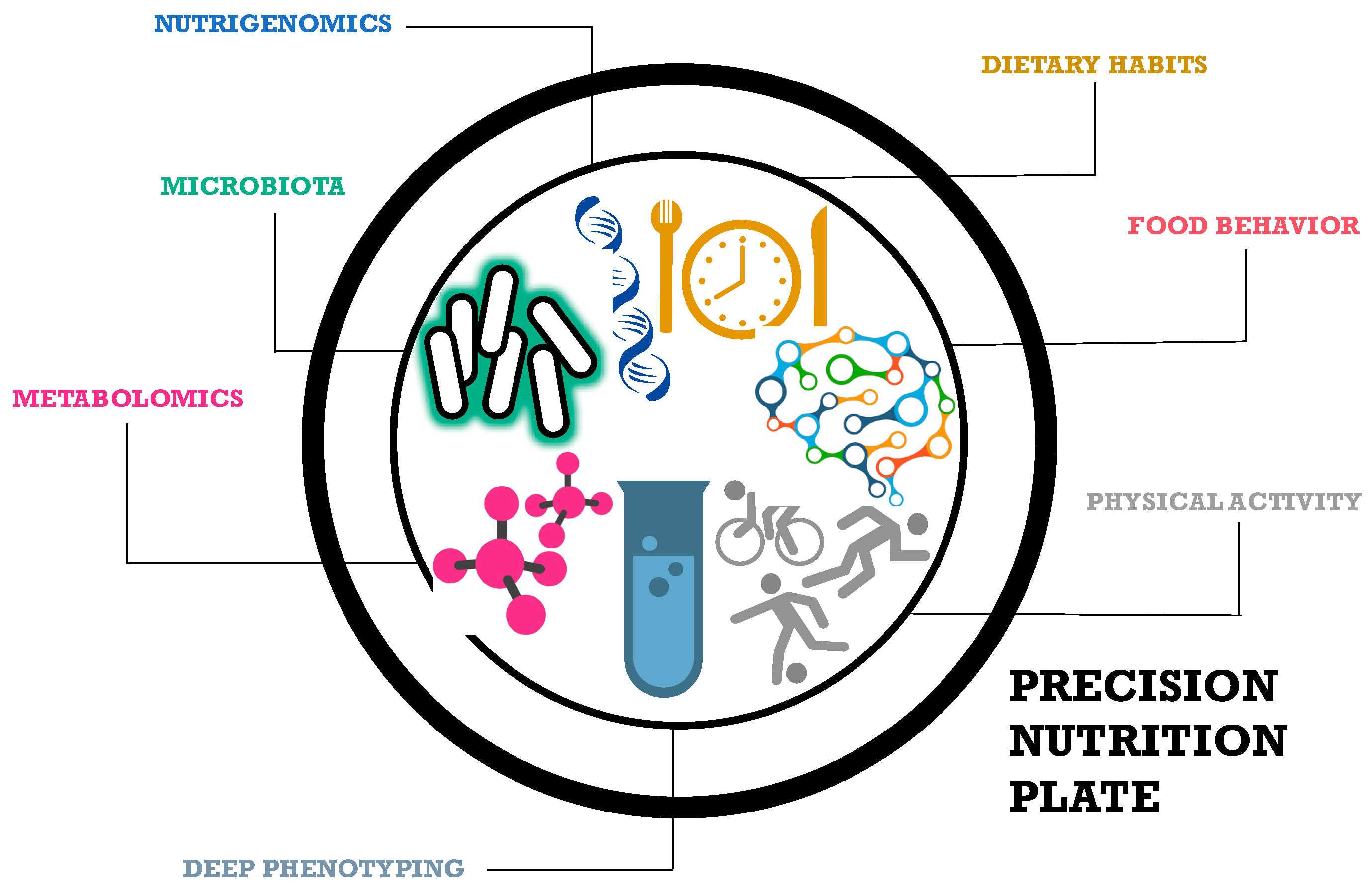
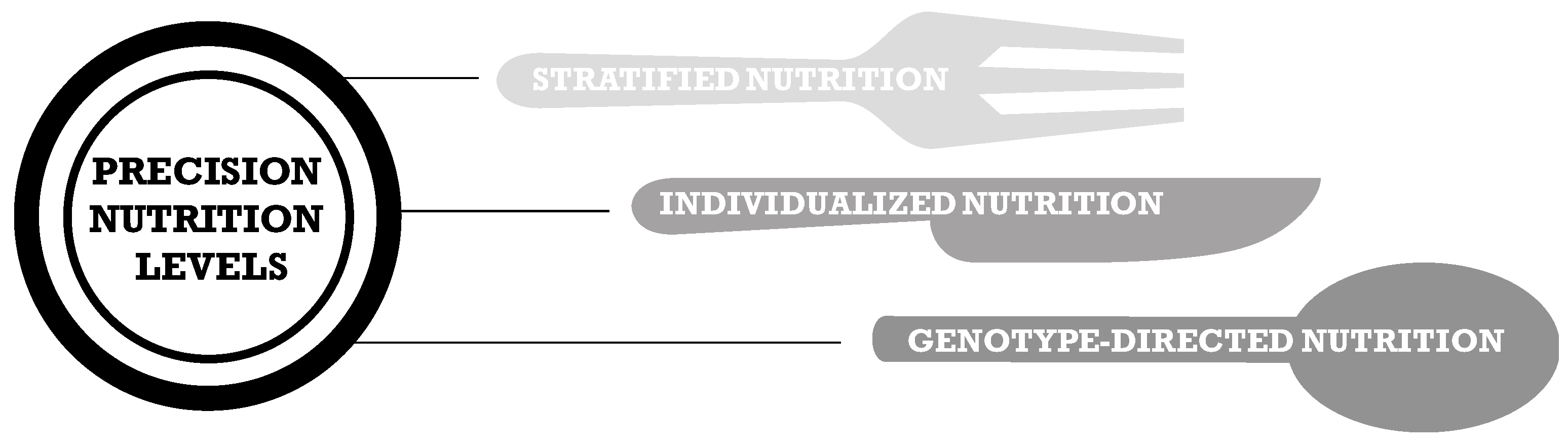
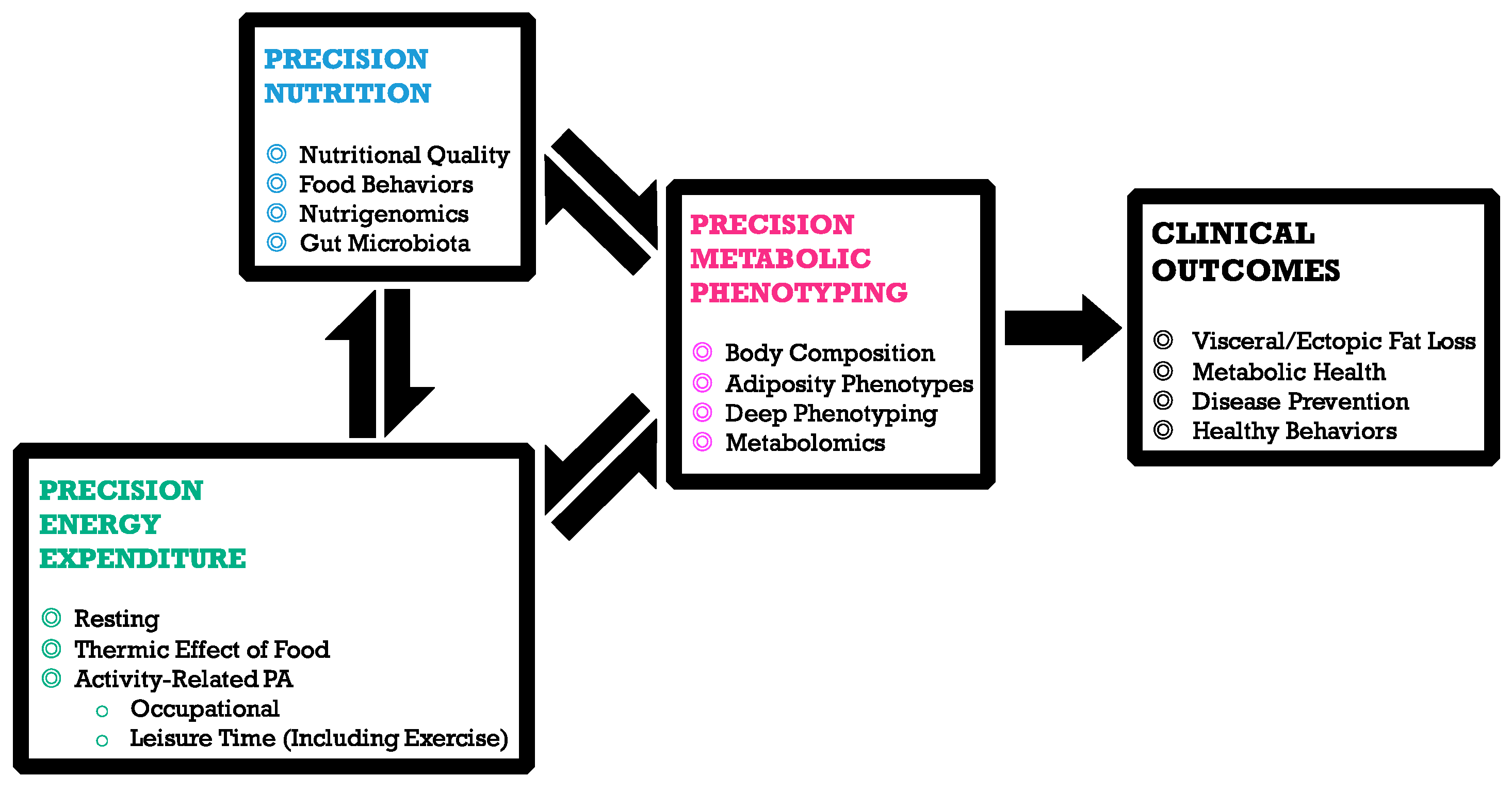
1. Precision Nutrition
The Road to Tailored Dietary Advices
2. Dietary Habits
Fine-Tuning Adherence
3. Food Behavior
Foodstyle Monitoring
4. Precision Physical Activity
Physical Activity: A Key Factor to Proper Precision Nutrition
5. Deep Phenotyping
High-Quality Phenotypes to Stratify Obesity
6. Metabolomics
Towards a Better Characterization of Eating
7. Microbiota Phenotyping
Diet-Gut Microbiome Interplay
8. Recent Advances in Precision Nutrition
8.1. From Nutrigenomics to Tailored Nutrition
8.2. PREDIMED
8.3. Food4Me
9. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Betts, J.A.; Gonzalez, J.T. Personalised nutrition: What makes you so special? Nutr. Bull. 2016, 41, 353–359. [Google Scholar] [CrossRef]
- McMahon, G.; Taylor, A.E.; Davey Smith, G.; Munafò, M.R. Phenotype refinement strengthens the association of AHR and CYP1A1 genotype with caffeine consumption. PLoS ONE 2014, 9, e103448. [Google Scholar] [CrossRef] [PubMed]
- Valleé Marcotte, B.; Cormier, H.; Guénard, F.; Rudkowska, I.; Lemieux, S.; Couture, P.; Vohl, M.C. Novel Genetic Loci Associated with the Plasma Triglyceride Response to an Omega-3 Fatty Acid Supplementation. J. Nutrigenet. Nutrigenomics 2016, 9, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Ouellette, C.; Rudkowska, I.; Lemieux, S.; Lamarche, B.; Couture, P.; Vohl, M.-C. Gene-diet interactions with polymorphisms of the MGLL gene on plasma low-density lipoprotein cholesterol and size following an omega-3 polyunsaturated fatty acid supplementation: A clinical trial. Lipids Health Dis. 2014, 13, 86. [Google Scholar] [CrossRef] [PubMed]
- Rudkowska, I.; Pérusse, L.; Bellis, C.; Blangero, J.; Després, J.-P.; Bouchard, C.; Vohl, M.-C. Interaction between Common Genetic Variants and Total Fat Intake on Low-Density Lipoprotein Peak Particle Diameter: A Genome-Wide Association Study. J. Nutrigenet. Nutrigenomics 2015, 8, 44–53. [Google Scholar] [CrossRef] [PubMed]
- Tremblay, B.L.; Cormier, H.; Rudkowska, I.; Lemieux, S.; Couture, P.; Vohl, M.-C. Association between polymorphisms in phospholipase A2 genes and the plasma triglyceride response to an n-3 PUFA supplementation: A clinical trial. Lipids Health Dis. 2015, 14, 12. [Google Scholar] [CrossRef] [PubMed]
- Palatini, P.; Ceolotto, G.; Ragazzo, F.; Dorigatti, F.; Saladini, F.; Papparella, I.; Mos, L.; Zanata, G.; Santonastaso, M. CYP1A2 genotype modifies the association between coffee intake and the risk of hypertension. J. Hypertens. 2009, 27, 1594–1601. [Google Scholar] [CrossRef] [PubMed]
- Ahmadi, K.R.; Andrew, T. Opportunism: A panacea for implementation of whole-genome sequencing studies in nutrigenomics research? Genes Nutr. 2014, 9, 387. [Google Scholar] [CrossRef] [PubMed]
- Özdemir, V.; Kolker, E. Precision Nutrition 4.0: A Big Data and Ethics Foresight Analysis—Convergence of Agrigenomics, Nutrigenomics, Nutriproteomics, and Nutrimetabolomics. OMIS J. Integr. Biol. 2016, 20, 69–75. [Google Scholar] [CrossRef] [PubMed]
- Rasinpera, H.; Savilahti, E.; Enattah, N.S.; Kuokkanen, M.; Tötterman, N.; Lindahl, H.; Järvelä, I.; Kolho, K.-L. A genetic test which can be used to diagnose adult-type hypolactasia in children. Gut 2004, 53, 1571–1576. [Google Scholar] [CrossRef] [PubMed]
- Ludvigsson, J.F.; Bai, J.C.; Biagi, F.; Card, T.R.; Ciacci, C.; Ciclitira, P.J.; Green, P.H.R.; Hadjivassiliou, M.; Holdoway, A.; van Heel, D.A.; et al. Diagnosis and management of adult coeliac disease: Guidelines from the British Society of Gastroenterology. Gut 2014, 63, 1210–1228. [Google Scholar] [CrossRef] [PubMed]
- DiLella, A.G.; Huang, W.M.; Woo, S.L. Screening for phenylketonuria mutations by DNA amplification with the polymerase chain reaction. Lancet 1988, 1, 497–499. [Google Scholar] [CrossRef]
- Cornelis, M.C.; El-Sohemy, A.; Campos, H. Genetic polymorphism of the adenosine A2A receptor is associated with habitual caffeine consumption. Am. J. Clin. Nutr. 2007, 86, 240–244. [Google Scholar] [PubMed]
- Cornelis, M.C.; El-Sohemy, A.; Kabagambe, E.K.; Campos, H. Coffee, CYP1A2 Genotype, and Risk of Myocardial Infarction. JAMA 2006, 295, 1135. [Google Scholar] [CrossRef] [PubMed]
- Corella, D.; Peloso, G.; Arnett, D.K.; Demissie, S.; Cupples, L.A.; Tucker, K.; Lai, C.-Q.; Parnell, L.D.; Coltell, O.; Lee, Y.-C.; et al. APOA2, Dietary Fat, and Body Mass Index. Arch. Intern. Med. 2009, 169, 1897. [Google Scholar] [CrossRef] [PubMed]
- Corella, D.; Tai, E.S.; Sorlí, J.V.; Chew, S.K.; Coltell, O.; Sotos-Prieto, M.; García-Rios, A.; Estruch, R.; Ordovas, J.M. Association between the APOA2 promoter polymorphism and body weight in Mediterranean and Asian populations: Replication of a gene-saturated fat interaction. Int. J. Obes. (Lond.) 2011, 35, 666–675. [Google Scholar] [CrossRef] [PubMed]
- Giner, V.; Poch, E.; Bragulat, E.; Oriola, J.; González, D.; Coca, A.; De La Sierra, A. Renin-angiotensin system genetic polymorphisms and salt sensitivity in essential hypertension. Hypertension 2000, 35, 512–517. [Google Scholar] [CrossRef] [PubMed]
- Poch, E.; González, D.; Giner, V.; Bragulat, E.; Coca, A.; de La Sierra, A. Molecular basis of salt sensitivity in human hypertension. Evaluation of renin-angiotensin-aldosterone system gene polymorphisms. Hypertension 2001, 38, 1204–1209. [Google Scholar] [CrossRef] [PubMed]
- Goni, L.; Cuervo, M.; Milagro, F.I.; Martínez, J.A. A genetic risk tool for obesity predisposition assessment and personalized nutrition implementation based on macronutrient intake. Genes Nutr. 2015, 10, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Rukh, G.; Sonestedt, E.; Melander, O.; Hedblad, B.; Wirfält, E.; Ericson, U.; Orho-Melander, M. Genetic susceptibility to obesity and diet intakes: Association and interaction analyses in the Malmö Diet and Cancer Study. Genes Nutr. 2013, 8, 535–547. [Google Scholar] [CrossRef] [PubMed]
- Olsen, N.J.; Ängquist, L.; Larsen, S.C.; Linneberg, A.; Skaaby, T.; Husemoen, L.L.N.; Toft, U.; Tjønneland, A.; Halkjær, J.; Hansen, T.; et al. Interactions between genetic variants associated with adiposity traits and soft drinks in relation to longitudinal changes in body weight and waist circumference. Am. J. Clin. Nutr. 2016, 104, 816–826. [Google Scholar] [CrossRef] [PubMed]
- Qi, Q.; Chu, A.Y.; Kang, J.H.; Jensen, M.K.; Curhan, G.C.; Pasquale, L.R.; Ridker, P.M.; Hunter, D.J.; Willett, W.C.; Rimm, E.B.; et al. Sugar-Sweetened Beverages and Genetic Risk of Obesity. N. Engl. J. Med. 2012, 367, 1387–1396. [Google Scholar] [CrossRef] [PubMed]
- Brunkwall, L.; Chen, Y.; Hindy, G.; Rukh, G.; Ericson, U.; Barroso, I.; Johansson, I.; Franks, P.W.; Orho-Melander, M.; Renstrom, F. Sugar-sweetened beverage consumption and genetic predisposition to obesity in 2 Swedish cohorts. Am. J. Clin. Nutr. 2016, 104, 809–815. [Google Scholar] [CrossRef] [PubMed]
- Qi, Q.; Chu, A.Y.; Kang, J.H.; Huang, J.; Rose, L.M.; Jensen, M.K.; Liang, L.; Curhan, G.C.; Pasquale, L.R.; Wiggs, J.L.; et al. Fried food consumption, genetic risk, and body mass index: Gene-diet interaction analysis in three US cohort studies. BMJ 2014, 348, g1610. [Google Scholar] [CrossRef] [PubMed]
- Casas-Agustench, P.; Arnett, D.K.; Smith, C.E.; Lai, C.-Q.; Parnell, L.D.; Borecki, I.B.; Frazier-Wood, A.C.; Allison, M.; Chen, Y.-D.I.; Taylor, K.D.; et al. Saturated Fat Intake Modulates the Association between an Obesity Genetic Risk Score and Body Mass Index in Two US Populations. J. Acad. Nutr. Diet. 2014, 114, 1954–1966. [Google Scholar] [CrossRef] [PubMed]
- Koochakpoor, G.; Daneshpour, M.S.; Mirmiran, P.; Hosseini, S.A.; Hosseini-Esfahani, F.; Sedaghatikhayat, B.; Azizi, F. The effect of interaction between Melanocortin-4 receptor polymorphism and dietary factors on the risk of metabolic syndrome. Nutr. Metab. (Lond.) 2016, 13, 35. [Google Scholar] [CrossRef] [PubMed]
- Azizi, F.; Ghanbarian, A.; Momenan, A.A.; Hadaegh, F.; Mirmiran, P.; Hedayati, M.; Mehrabi, Y.; Zahedi-Asl, S. Tehran Lipid and Glucose Study Group. Prevention of non-communicable disease in a population in nutrition transition: Tehran Lipid and Glucose Study phase II. Trials 2009, 10, 5. [Google Scholar] [CrossRef] [PubMed]
- Nettleton, J.A.; Follis, J.L.; Ngwa, J.S.; Smith, C.E.; Ahmad, S.; Tanaka, T.; Wojczynski, M.K.; Voortman, T.; Lemaitre, R.N.; Kristiansson, K.; et al. Gene x dietary pattern interactions in obesity: Analysis of up to 68 317 adults of European ancestry. Hum. Mol. Genet. 2015, 24, 4728–4738. [Google Scholar] [CrossRef] [PubMed]
- Ferguson, L.R.; De Caterina, R.; Görman, U.; Allayee, H.; Kohlmeier, M.; Prasad, C.; Choi, M.S.; Curi, R.; de Luis, D.A.; Gil, Á.; et al. Guide and Position of the International Society of Nutrigenetics/Nutrigenomics on Personalised Nutrition: Part 1 - Fields of Precision Nutrition. J. Nutrigenet. Nutrigenomics 2016, 9, 12–27. [Google Scholar] [CrossRef] [PubMed]
- Allison, D.B.; Bassaganya-Riera, J.; Burlingame, B.; Brown, A.W.; le Coutre, J.; Dickson, S.L.; van Eden, W.; Garssen, J.; Hontecillas, R.; Khoo, C.S.H.; et al. Goals in Nutrition Science 2015–2020. Front. Nutr. 2015, 2, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Corella, D.; Coltell, O.; Mattingley, G.; Sorlí, J.V.; Ordovas, J.M. Utilizing nutritional genomics to tailor diets for the prevention of cardiovascular disease: A guide for upcoming studies and implementations. Expert Rev. Mol. Diagn. 2017, 17, 495–513. [Google Scholar] [CrossRef] [PubMed]
- Srinivasan, B.; Lee, S.; Erickson, D. Precision nutrition—Review of methods for point-of-care assessment of nutritional status. Curr. Opin. Biotechnol. 2017, 44, 103–108. [Google Scholar] [CrossRef] [PubMed]
- Zeevi, D.; Korem, T.; Zmora, N.; Israeli, D.; Rothschild, D.; Weinberger, A.; Ben-Yacov, O.; Lador, D.; Avnit-Sagi, T.; Lotan-Pompan, M.; et al. Personalized Nutrition by Prediction of Glycemic Responses. Cell 2015, 163, 1079–1095. [Google Scholar] [CrossRef] [PubMed]
- Wolever, T.M.S. Personalized nutrition by prediction of glycaemic responses: Fact or fantasy? Eur. J. Clin. Nutr. 2016, 70, 411–413. [Google Scholar] [CrossRef] [PubMed]
- Ohlhorst, S.D.; Russell, R.; Bier, D.; Klurfeld, D.M.; Li, Z.; Mein, J.R.; Milner, J.; Ross, A.C.; Stover, P.; Konopka, E. Nutrition research to affect food and a healthy lifespan. Adv. Nutr. Int. Rev. J. 2013, 4, 579–584. [Google Scholar] [CrossRef] [PubMed]
- Loos, R.J.F.; Janssens, A.C.J.W. Predicting Polygenic Obesity Using Genetic Information. Cell Metab. 2017, 25, 535–543. [Google Scholar] [CrossRef] [PubMed]
- Hebert, J.R.; Frongillo, E.A.; Adams, S.A.; Turner-McGrievy, G.M.; Hurley, T.G.; Miller, D.R.; Ockene, I.S. Perspective: Randomized Controlled Trials Are Not a Panacea for Diet-Related Research. Adv. Nutr. An Int. Rev. J. 2016, 7, 423–432. [Google Scholar] [CrossRef] [PubMed]
- Siebelink, E.; Geelen, A.; de Vries, J.H.M. Self-reported energy intake by FFQ compared with actual energy intake to maintain body weight in 516 adults. Br. J. Nutr. 2011, 106, 274–281. [Google Scholar] [CrossRef] [PubMed]
- Schaefer, E.J.; Augustin, J.L.; Schaefer, M.M.; Rasmussen, H.; Ordovas, J.M.; Dallal, G.E.; Dwyer, J.T. Lack of efficacy of a food-frequency questionnaire in assessing dietary macronutrient intakes in subjects consuming diets of known composition. Am. J. Clin. Nutr. 2000, 71, 746–751. [Google Scholar] [PubMed]
- Archer, E.; Pavela, G.; Lavie, C.J. The Inadmissibility of What We Eat in America and NHANES Dietary Data in Nutrition and Obesity Research and the Scientific Formulation of National Dietary Guidelines. Mayo Clin. Proc. 2015, 90, 911–926. [Google Scholar] [CrossRef] [PubMed]
- Shim, J.-S.; Oh, K.; Kim, H.C. Dietary assessment methods in epidemiologic studies. Epidemiol. Health 2014, 36, e2014009. [Google Scholar] [CrossRef] [PubMed]
- Schroder, H.; Fito, M.; Estruch, R.; Martinez-Gonzalez, M.A.; Corella, D.; Salas-Salvado, J.; Lamuela-Raventos, R.; Ros, E.; Salaverria, I.; Fiol, M.; et al. A Short Screener Is Valid for Assessing Mediterranean Diet Adherence among Older Spanish Men and Women. J. Nutr. 2011, 141, 1140–1145. [Google Scholar] [CrossRef] [PubMed]
- Martínez-González, M.A.; García-Arellano, A.; Toledo, E.; Salas-Salvadó, J.; Buil-Cosiales, P.; Corella, D.; Covas, M.I.; Schröder, H.; Arós, F.; Gómez-Gracia, E.; et al. A 14-item Mediterranean diet assessment tool and obesity indexes among high-risk subjects: The PREDIMED trial. PLoS ONE 2012, 7, e43134. [Google Scholar] [CrossRef] [PubMed]
- Sotos-Prieto, M.; Moreno-Franco, B.; Ordovás, J.; León, M.; Casasnovas, J.; Peñalvo, J. Design and development of an instrument to measure overall lifestyle habits for epidemiological research: The Mediterranean Lifestyle (MEDLIFE) index. Public Health Nutr. 2015, 18, 959–967. [Google Scholar] [CrossRef] [PubMed]
- Martínez-González, M.A.; Fernández-Jarne, E.; Serrano-Martínez, M.; Wright, M.; Gomez-Gracia, E. Development of a short dietary intake questionnaire for the quantitative estimation of adherence to a cardioprotective Mediterranean diet. Eur. J. Clin. Nutr. 2004, 58, 1550–1552. [Google Scholar] [CrossRef] [PubMed]
- Estruch, R.; Ros, E.; Salas-Salvadó, J.; Covas, M.-I.; Corella, D.; Arós, F.; Gómez-Gracia, E.; Ruiz-Gutiérrez, V.; Fiol, M.; Lapetra, J.; et al. Primary Prevention of Cardiovascular Disease with a Mediterranean Diet. N. Engl. J. Med. 2013, 368, 1279–1290. [Google Scholar] [CrossRef] [PubMed]
- McCullough, M.L.; Feskanich, D.; Stampfer, M.J.; Giovannucci, E.L.; Rimm, E.B.; Hu, F.B.; Spiegelman, D.; Hunter, D.J.; Colditz, G.A.; Willett, W.C. Diet quality and major chronic disease risk in men and women: Moving toward improved dietary guidance. Am. J. Clin. Nutr. 2002, 76, 1261–1271. [Google Scholar] [PubMed]
- Fung, T.T.; McCullough, M.L.; Newby, P.K.; Manson, J.E.; Meigs, J.B.; Rifai, N.; Willett, W.C.; Hu, F.B. Diet-quality scores and plasma concentrations of markers of inflammation and endothelial dysfunction. Am. J. Clin. Nutr. 2005, 82, 163–173. [Google Scholar] [PubMed]
- Sevilla-Villanueva, B.; Gibert, K.; Sanchez-Marre, M.; Fito, M.; Covas, M.-I. Evaluation of Adherence to Nutritional Intervention through Trajectory Analysis. IEEE J. Biomed. Heal. Inform. 2017, 21, 628–634. [Google Scholar] [CrossRef] [PubMed]
- Sevilla-Villanueva, B.; Gibert, K.; Sànchez-Marrè, M. Identifying Nutritional Patterns through Integrative Multiview Clustering. Artif. Intell. Res. Dev. 2015, 277, 185. [Google Scholar]
- Konstantinidou, V.; Covas, M.-I.; Munoz-Aguayo, D.; Khymenets, O.; de la Torre, R.; Saez, G.; Tormos, M.d.C.; Toledo, E.; Marti, A.; Ruiz-Gutierrez, V.; et al. In vivo nutrigenomic effects of virgin olive oil polyphenols within the frame of the Mediterranean diet: A randomized controlled trial. FASEB J. 2010, 24, 2546–2557. [Google Scholar] [CrossRef] [PubMed]
- Dhurandhar, N.V.; Schoeller, D.; Brown, A.W.; Heymsfield, S.B.; Thomas, D.; Sørensen, T.I.A.; Speakman, J.R.; Jeansonne, M.; Allison, D.B. Energy balance measurement: When something is not better than nothing. Int. J. Obes. 2015, 39, 1109–1113. [Google Scholar] [CrossRef] [PubMed]
- Nybacka, S.; Bertéus Forslund, H.; Wirfält, E.; Larsson, I.; Ericson, U.; Warensjö Lemming, E.; Bergström, G.; Hedblad, B.; Winkvist, A.; Lindroos, A.K. Comparison of a web-based food record tool and a food-frequency questionnaire and objective validation using the doubly labelled water technique in a Swedish middle-aged population. J. Nutr. Sci. 2016, 5, e39. [Google Scholar] [CrossRef] [PubMed]
- Bergström, G.; Berglund, G.; Blomberg, A.; Brandberg, J.; Engström, G.; Engvall, J.; Eriksson, M.; de Faire, U.; Flinck, A.; Hansson, M.G.; et al. The Swedish CArdioPulmonary BioImage Study: Objectives and design. J. Intern. Med. 2015, 278, 645–659. [Google Scholar] [CrossRef]
- Martin, C.K.; Han, H.; Coulon, S.M.; Allen, H.R.; Champagne, C.M.; Anton, S.D. A novel method to remotely measure food intake of free-living individuals in real time: The remote food photography method. Br. J. Nutr. 2009, 101, 446. [Google Scholar] [CrossRef] [PubMed]
- Martin, C.K.; Correa, J.B.; Han, H.; Allen, H.R.; Rood, J.C.; Champagne, C.M.; Gunturk, B.K.; Bray, G.A. Validity of the Remote Food Photography Method (RFPM) for Estimating Energy and Nutrient Intake in Near Real-Time. Obesity 2012, 20, 891–899. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Hoover, A.; Scisco, J.; Muth, E. A New Method for Measuring Meal Intake in Humans via Automated Wrist Motion Tracking. Appl. Psychophysiol. Biofeedback 2012, 37, 205–215. [Google Scholar] [CrossRef] [PubMed]
- Schoeller, D.A.; Thomas, D.; Archer, E.; Heymsfield, S.B.; Blair, S.N.; Goran, M.I.; Hill, J.O.; Atkinson, R.L.; Corkey, B.E.; Foreyt, J.; et al. Self-report-based estimates of energy intake offer an inadequate basis for scientific conclusions. Am. J. Clin. Nutr. 2013, 97, 1413–1415. [Google Scholar] [CrossRef] [PubMed]
- Mattfeld, R.S.; Muth, E.R.; Hoover, A. Measuring the consumption of individual solid and liquid bites using a table embedded scale during unrestricted eating. IEEE J. Biomed. Heal. Informatics 2016. [Google Scholar] [CrossRef] [PubMed]
- Fontana, J.M.; Farooq, M.; Sazonov, E. Automatic Ingestion Monitor: A Novel Wearable Device for Monitoring of Ingestive Behavior. IEEE Trans. Biomed. Eng. 2014, 61, 1772–1779. [Google Scholar] [CrossRef] [PubMed]
- Potter, G.D.M.; Cade, J.E.; Grant, P.J.; Hardie, L.J. Nutrition and the circadian system. Br. J. Nutr. 2016, 116, 434–442. [Google Scholar] [CrossRef] [PubMed]
- Garaulet, M.; Vera, B.; Bonnet-Rubio, G.; Gómez-Abellán, P.; Lee, Y.-C.; Ordovás, J.M. Lunch eating predicts weight-loss effectiveness in carriers of the common allele at PERILIPIN1: The ONTIME (Obesity, Nutrigenetics, Timing, Mediterranean) study. Am. J. Clin. Nutr. 2016, 104, 1160–1166. [Google Scholar] [CrossRef] [PubMed]
- Garaulet, M.; Corbalán-Tutau, M.D.; Madrid, J.A.; Baraza, J.C.; Parnell, L.D.; Lee, Y.-C.; Ordovas, J.M. PERIOD2 Variants Are Associated with Abdominal Obesity, Psycho-Behavioral Factors, and Attrition in the Dietary Treatment of Obesity. J. Am. Diet. Assoc. 2010, 110, 917–921. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Delgado, F.; Garcia-Rios, A.; Alcala-Diaz, J.F.; Rangel-Zuñiga, O.; Delgado-Lista, J.; Yubero-Serrano, E.M.; Lopez-Moreno, J.; Tinahones, F.J.; Ordovas, J.M.; Garaulet, M.; et al. Chronic consumption of a low-fat diet improves cardiometabolic risk factors according to the CLOCK gene in patients with coronary heart disease. Mol. Nutr. Food Res. 2015, 59, 2556–2564. [Google Scholar] [CrossRef] [PubMed]
- Dashti, H.S.; Smith, C.E.; Lee, Y.-C.; Parnell, L.D.; Lai, C.-Q.; Arnett, D.K.; Ordovás, J.M.; Garaulet, M. CRY1 circadian gene variant interacts with carbohydrate intake for insulin resistance in two independent populations: Mediterranean and North American. Chronobiol. Int. 2014, 31, 660–667. [Google Scholar] [CrossRef] [PubMed]
- Asher, G.; Sassone-Corsi, P. Time for Food: The Intimate Interplay between Nutrition, Metabolism, and the Circadian Clock. Cell 2015, 161, 84–92. [Google Scholar] [CrossRef] [PubMed]
- Oike, H.; Oishi, K.; Kobori, M. Nutrients, Clock Genes, and Chrononutrition. Curr. Nutr. Rep. 2014, 3, 204–212. [Google Scholar] [CrossRef] [PubMed]
- Hill, J.O.; Wyatt, H.R.; Peters, J.C. Energy balance and obesity. Circulation 2012, 126, 126–132. [Google Scholar] [CrossRef] [PubMed]
- Bouchard, C.; Blair, S.N.; Church, T.S.; Earnest, C.P.; Hagberg, J.M.; H?kkinen, K.; Jenkins, N.T.; Karavirta, L.; Kraus, W.E.; Leon, A.S.; et al. Adverse Metabolic Response to Regular Exercise: Is It a Rare or Common Occurrence? PLoS ONE 2012, 7, e37887. [Google Scholar] [CrossRef] [PubMed]
- de Lannoy, L.; Clarke, J.; Stotz, P.J.; Ross, R.; Senn, S.; Meyer, T. Effects of intensity and amount of exercise on measures of insulin and glucose: Analysis of inter-individual variability. PLoS ONE 2017, 12, e0177095. [Google Scholar] [CrossRef] [PubMed]
- Atkinson, G.; Batterham, A.M. True and false interindividual differences in the physiological response to an intervention. Exp. Physiol. 2015, 100, 577–588. [Google Scholar] [CrossRef] [PubMed]
- Qi, Q.; Li, Y.; Chomistek, A.K.; Kang, J.H.; Curhan, G.C.; Pasquale, L.R.; Willett, W.C.; Rimm, E.B.; Hu, F.B.; Qi, L. Television Watching, Leisure Time Physical Activity, and the Genetic Predisposition in Relation to Body Mass Index in Women and Men. Circulation 2012, 126, 1821–1827. [Google Scholar] [CrossRef] [PubMed]
- Loos, R.J.F.; Yeo, G.S.H. The bigger picture of FTO: The first GWAS-identified obesity gene. Nat. Rev. Endocrinol. 2014, 10, 51–61. [Google Scholar] [CrossRef] [PubMed]
- Frayling, T.M.; Timpson, N.J.; Weedon, M.N.; Zeggini, E.; Freathy, R.M.; Lindgren, C.M.; Perry, J.R.B.; Elliott, K.S.; Lango, H.; Rayner, N.W.; et al. A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 2007, 316, 889–894. [Google Scholar] [CrossRef] [PubMed]
- Vimaleswaran, K.S.; Li, S.; Zhao, J.H.; Luan, J.; Bingham, S.A.; Khaw, K.-T.; Ekelund, U.; Wareham, N.J.; Loos, R.J. Physical activity attenuates the body mass index-increasing influence of genetic variation in the FTO gene. Am. J. Clin. Nutr. 2009, 90, 425–428. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Zhao, J.H.; Luan, J.; Ekelund, U.; Luben, R.N.; Khaw, K.T.; Wareham, N.J.; Loos, R.J.F. Physical activity attenuates the genetic predisposition to obesity in 20,000 men and women from EPIC-Norfolk prospective population study. PLoS Med. 2010, 7, e1000332. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, S.; Rukh, G.; Varga, T. V.; Ali, A.; Kurbasic, A.; Shungin, D.; Ericson, U.; Koivula, R.W.; Chu, A.Y.; Rose, L.M.; et al. Gene × Physical Activity Interactions in Obesity: Combined Analysis of 111,421 Individuals of European Ancestry. PLoS Genet. 2013, 9, e1003607. [Google Scholar] [CrossRef] [PubMed]
- Graff, M.; Scott, R.A.; Justice, A.E.; Young, K.L.; Feitosa, M.F.; Barata, L.; Winkler, T.W.; Chu, A.Y.; Mahajan, A.; Hadley, D.; et al. Genome-wide physical activity interactions in adiposity―A meta-analysis of 200,452 adults. PLoS Genet. 2017, 13, e1006528. [Google Scholar] [CrossRef] [PubMed]
- Scott, R.A.; Chu, A.Y.; Grarup, N.; Manning, A.K.; Hivert, M.-F.; Shungin, D.; Tönjes, A.; Yesupriya, A.; Barnes, D.; Bouatia-Naji, N.; et al. No Interactions Between Previously Associated 2-h Glucose Gene Variants and Physical Activity or BMI on 2-Hour Glucose Levels. Diabetes 2012, 61, 1291–1296. [Google Scholar] [CrossRef] [PubMed]
- Moon, J.-Y.; Wang, T.; Sofer, T.; North, K.E.; Isasi, C.R.; Cai, J.; Gellman, M.D.; Moncrieft, A.E.; Sotres-Alvarez, D.; Argos, M.; et al. Gene-environment Interaction Analysis Reveals Evidence for Independent Influences of Physical Activity and Sedentary Behavior on Obesity: Results From the Hispanic Community Health Study/study of Latinos (HCHS/SOL). Circulation 2017, 135, AMP027. [Google Scholar]
- Celis-Morales, C.; Marsaux, C.F.M.; Livingstone, K.M.; Navas-Carretero, S.; San-Cristobal, R.; O’donovan, C.B.; Forster, H.; Woolhead, C.; Fallaize, R.; Macready, A.L.; et al. Physical activity attenuates the effect of the FTO genotype on obesity traits in European adults: The Food4Me study. Obesity 2016, 24, 962–969. [Google Scholar] [CrossRef] [PubMed]
- Andreasen, C.H.; Stender-Petersen, K.L.; Mogensen, M.S.; Torekov, S.S.; Wegner, L.; Andersen, G.; Nielsen, A.L.; Albrechtsen, A.; Borch-Johnsen, K.; Rasmussen, S.S.; et al. Low physical activity acentuates the effect of rs9939609 polymorphism. Diabetes 2008, 57, 95–101. [Google Scholar] [CrossRef] [PubMed]
- Goodman, E.; Evans, W.D.; DiPietro, L. Preliminary Evidence for School-Based Physical Activity Policy Needs in Washington, DC. J. Phys. Act. Heal. 2012, 9, 124–128. [Google Scholar] [CrossRef]
- Cadenas-Sanchez, C.; Ruiz, J.R.; Labayen, I.; Huybrechts, I.; Manios, Y.; Gonz??lez-Gross, M.; Breidenassel, C.; Kafatos, A.; De Henauw, S.; Vanhelst, J.; et al. Prevalence of Metabolically Healthy but Overweight/Obese Phenotype and Its Association With Sedentary Time, Physical Activity, and Fitness. J. Adolesc. Heal. 2016. [Google Scholar] [CrossRef] [PubMed]
- Cameron, N.; Godino, J.; Nichols, J.F.; Wing, D.; Hill, L.; Patrick, K. Associations between physical activity and BMI, body fatness, and visceral adiposity in overweight or obese Latino and non-Latino adults. Int. J. Obes. 2017. [Google Scholar] [CrossRef] [PubMed]
- Jeran, S.; Steinbrecher, A.; Pischon, T. Prediction of activity-related energy expenditure using accelerometer-derived physical activity under free-living conditions: A systematic review. Int. J. Obes. 2016, 40, 1187–1197. [Google Scholar] [CrossRef] [PubMed]
- Westerterp, K.R. Reliable assessment of physical activity in disease. Curr. Opin. Clin. Nutr. Metab. Care 2014, 17, 401–406. [Google Scholar] [CrossRef] [PubMed]
- Ross, R.; Hudson, R.; Stotz, P.J.; Lam, M. Effects of Exercise Amount and Intensity on Abdominal Obesity and Glucose Tolerance in Obese Adults. Ann. Intern. Med. 2015, 162, 325. [Google Scholar] [CrossRef] [PubMed]
- Livingstone, K.M.; Celis-Morales, C.; Navas-Carretero, S.; San-Cristoba, R.; MacReady, A.L.; Fallaize, R.; Forster, H.; Woolhead, C.; O’Donovan, C.B.; Marsaux, C.F.M.; et al. Effect of an Internet-based, personalized nutrition randomized trial on dietary changes associated with the Mediterranean diet: The Food4Me Study. Am. J. Clin. Nutr. 2016, 104, 288–297. [Google Scholar] [CrossRef] [PubMed]
- Prince, S.A.; Adamo, K.B.; Hamel, M.; Hardt, J.; Connor Gorber, S.; Tremblay, M.; O’Brien, W.; Bassett, D.; Schmitz, K.; Emplaincourt, P. A comparison of direct versus self-report measures for assessing physical activity in adults: A systematic review. Int. J. Behav. Nutr. Phys. Act. 2008, 5, 56. [Google Scholar] [CrossRef] [PubMed]
- Thompson, D.; Peacock, O.; Western, M.; Batterham, A.M. Multidimensional physical activity: An opportunity, not a problem. Exerc. Sport Sci. Rev. 2015, 43, 67–74. [Google Scholar] [CrossRef] [PubMed]
- Tracy, R.P. “Deep phenotyping”: Characterizing populations in the era of genomics and systems biology. Curr. Opin. Lipidol. 2008, 19, 151–157. [Google Scholar] [CrossRef] [PubMed]
- Kramer, C.K.; Zinman, B.; Retnakaran, R. Are metabolically healthy overweight and obesity benign conditions?: A systematic review and meta-analysis. Ann. Intern. Med. 2013, 159, 758–769. [Google Scholar] [CrossRef] [PubMed]
- Tchernof, A.; Després, J.-P. Pathophysiology of human visceral obesity: An update. Physiol. Rev. 2013, 93, 359–404. [Google Scholar] [CrossRef] [PubMed]
- Sam, S.; Haffner, S.; Davidson, M.H.E. Al Predicts Increased Visceral Fat in Subjects With Type 2 Diabetes. Diabetes 2009, 32, 1916–1920. [Google Scholar] [CrossRef]
- Lemieux, I.; Poirier, P.; Bergeron, J.; Alméras, N.; Lamarche, B.; Cantin, B.; Dagenais, G.R.; Després, J.-P. Hypertriglyceridemic waist: A useful screening phenotype in preventive cardiology? Can. J. Cardiol. 2007, 23 Suppl B, 23B–31B. [Google Scholar] [CrossRef]
- Arsenault, B.J.; Lemieux, I.; Despres, J.-P.; Wareham, N.J.; Kastelein, J.J.P.; Khaw, K.-T.; Boekholdt, S.M. The hypertriglyceridemic-waist phenotype and the risk of coronary artery disease: Results from the EPIC-Norfolk Prospective Population Study. Can. Med. Assoc. J. 2010, 182, 1427–1432. [Google Scholar] [CrossRef] [PubMed]
- Guénard, F.; Tchernof, A.; Deshaies, Y.; Pérusse, L.; Biron, S.; Lescelleur, O.; Biertho, L.; Marceau, S.; Vohl, M.-C. Differential methylation in visceral adipose tissue of obese men discordant for metabolic disturbances. Physiol. Genomics 2014, 46, 216–222. [Google Scholar] [CrossRef]
- Macartney-Coxson, D.; Benton, M.C.; Blick, R.; Stubbs, R.S.; Hagan, R.D.; Langston, M.A. Genome-wide DNA methylation analysis reveals loci that distinguish different types of adipose tissue in obese individuals. Clin. Epigenetics 2017, 9, 48. [Google Scholar] [CrossRef] [PubMed]
- Guénard, F.; Deshaies, Y.; Hould, F.-S.; Lebel, S.; Tchernof, A.; Marceau, P.; Vohl, M.-C. Use of Blood as a Surrogate Model for the Assessment of Visceral Adipose Tissue Methylation Profiles Associated with the Metabolic Syndrome in Men. J. Mol. Genet. Med. 2016, 10, 1–8. [Google Scholar] [CrossRef]
- Rönn, T.; Volkov, P.; Gillberg, L.; Kokosar, M.; Perfilyev, A.; Jacobsen, A.L.; Jørgensen, S.W.; Brøns, C.; Jansson, P.-A.; Eriksson, K.-F.; et al. Impact of age, BMI and HbA1c levels on the genome-wide DNA methylation and mRNA expression patterns in human adipose tissue and identification of epigenetic biomarkers in blood. Hum. Mol. Genet. 2015, 24, 3792–3813. [Google Scholar] [CrossRef]
- Moleres, A.; Campion, J.; Milagro, F.I.; Marcos, A.; Campoy, C.; Garagorri, J.M.; Gomez-Martinez, S.; Martinez, J.A.; Azcona-Sanjulian, M.C.; Marti, A.; et al. Differential DNA methylation patterns between high and low responders to a weight loss intervention in overweight or obese adolescents: The EVASYON study. FASEB J. 2013, 27, 2504–2512. [Google Scholar] [CrossRef] [PubMed]
- Milagro, F.I.; Campión, J.; Cordero, P.; Goyenechea, E.; Gómez-Uriz, A.M.; Abete, I.; Zulet, M.A.; Martínez, J.A. A dual epigenomic approach for the search of obesity biomarkers: DNA methylation in relation to diet-induced weight loss. FASEB J. 2011, 25, 1378–1389. [Google Scholar] [CrossRef] [PubMed]
- Bouchard, L.; Rabasa-Lhoret, R.; Faraj, M.; Lavoie, M.-E.; Mill, J.; Pérusse, L.; Vohl, M.-C. Differential epigenomic and transcriptomic responses in subcutaneous adipose tissue between low and high responders to caloric restriction. Am. J. Clin. Nutr. 2010, 91, 309–320. [Google Scholar] [CrossRef] [PubMed]
- Nicoletti, C.F.; Nonino, C.B.; de Oliveira, B.A.P.; Pinhel, M.A.; de Souza Pinhel, M.A.; Mansego, M.L.; Milagro, F.I.; Zulet, M.A.; Martinez, J.A. DNA Methylation and Hydroxymethylation Levels in Relation to Two Weight Loss Strategies: Energy-Restricted Diet or Bariatric Surgery. Obes. Surg. 2016, 26, 603–611. [Google Scholar] [CrossRef] [PubMed]
- Robinson, P.N. Deep phenotyping for precision medicine. Hum. Mutat. 2012, 33, 777–780. [Google Scholar] [CrossRef] [PubMed]
- Delude, C.M. Deep phenotyping: The details of disease. Nature 2015, 527, S14–S15. [Google Scholar] [CrossRef] [PubMed]
- Schram, M.T.; Sep, S.J.S.; van der Kallen, C.J.; Dagnelie, P.C.; Koster, A.; Schaper, N.; Henry, R.M.A.; Stehouwer, C.D.A. The Maastricht Study: An extensive phenotyping study on determinants of type 2 diabetes, its complications and its comorbidities. Eur. J. Epidemiol. 2014, 29, 439–451. [Google Scholar] [CrossRef] [PubMed]
- Lanktree, M.B.; Hassell, R.G.; Lahiry, P.; Hegele, R.A. Phenomics: Expanding the role of clinical evaluation in genomic studies. J. Investig. Med. 2010, 58, 700–706. [Google Scholar] [CrossRef] [PubMed]
- Sébédio, J.L. Metabolomics, Nutrition, and Potential Biomarkers of Food Quality, Intake, and Health Status. Adv. Food Nutr. Res. 2017, 82, 83–116. [Google Scholar] [CrossRef] [PubMed]
- Trabado, S.; Al-Salameh, A.; Croixmarie, V.; Masson, P.; Corruble, E.; Fève, B.; Colle, R.; Ripoll, L.; Walther, B.; Boursier-Neyret, C.; et al. The human plasma-metabolome: Reference values in 800 French healthy volunteers; impact of cholesterol, gender and age. PLoS ONE 2017, 12, e0173615. [Google Scholar] [CrossRef] [PubMed]
- Edmands, W.M.; Ferrari, P.; Rothwell, J.A.; Rinaldi, S.; Slimani, N.; Barupal, D.K.; Biessy, C.; Jenab, M.; Clavel-Chapelon, F.; Fagherazzi, G.; et al. Polyphenol metabolome in human urine and its association with intake of polyphenol-rich foods across European countries. Am. J. Clin. Nutr. 2015, 102, 905–913. [Google Scholar] [CrossRef] [PubMed]
- Garg, R.; Brennan, L.; Price, R.K.; Wallace, J.M.W.; Strain, J.J.; Gibney, M.J.; Shewry, P.R.; Ward, J.L.; Garg, L.; Welch, R.W. Using NMR-Based Metabolomics to Evaluate Postprandial Urinary Responses Following Consumption of Minimally Processed Wheat Bran or Wheat Aleurone by Men and Women. Nutrients 2016, 8, 96. [Google Scholar] [CrossRef] [PubMed]
- Gibbons, H.; McNulty, B.A.; Nugent, A.P.; Walton, J.; Flynn, A.; Gibney, M.J.; Brennan, L. A metabolomics approach to the identification of biomarkers of sugar-sweetened beverage intake. Am. J. Clin. Nutr. 2015, 101, 471–477. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Aloy, M.; Llorach, R.; Urpi-Sarda, M.; Tulipani, S.; Estruch, R.; Martínez-González, M.A.; Corella, D.; Fitó, M.; Ros, E.; Salas-Salvadó, J.; et al. Novel Multimetabolite Prediction of Walnut Consumption by a Urinary Biomarker Model in a Free-Living Population: The PREDIMED Study. J. Proteome Res. 2014, 13, 3476–3483. [Google Scholar] [CrossRef] [PubMed]
- Playdon, M.C.; Moore, S.C.; Derkach, A.; Reedy, J.; Subar, A.F.; Sampson, J.N.; Albanes, D.; Gu, F.; Kontto, J.; Lassale, C.; et al. Identifying biomarkers of dietary patterns by using metabolomics. Am. J. Clin. Nutr. 2017, 105, 450–465. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Perez, I.; Posma, J.M.; Gibson, R.; Chambers, E.S.; Hansen, T.H.; Vestergaard, H.; Hansen, T.; Beckmann, M.; Pedersen, O.; Elliott, P.; et al. Objective assessment of dietary patterns by use of metabolic phenotyping: A randomised, controlled, crossover trial. Lancet Diabetes Endocrinol. 2017, 5, 184–195. [Google Scholar] [CrossRef]
- Bhupathiraju, S.N.; Hu, F.B. One (small) step towards precision nutrition by use of metabolomics. Lancet Diabetes Endocrinol. 2017, 5, 154–155. [Google Scholar] [CrossRef]
- Rådjursöga, M.; Karlsson, G.B.; Lindqvist, H.M.; Pedersen, A.; Persson, C.; Pinto, R.C.; Ellegård, L.; Winkvist, A. Metabolic profiles from two different breakfast meals characterized by 1H NMR-based metabolomics. Food Chem. 2017, 231, 267–274. [Google Scholar] [CrossRef] [PubMed]
- Brennan, L. Metabolomics in nutrition research–a powerful window into nutritional metabolism. Essays Biochem. 2016, 60, 451–458. [Google Scholar] [CrossRef] [PubMed]
- Allam-Ndoul, B.; Guénard, F.; Garneau, V.; Cormier, H.; Barbier, O.; Pérusse, L.; Vohl, M.-C. Association between Metabolite Profiles, Metabolic Syndrome and Obesity Status. Nutrients 2016, 8, 324. [Google Scholar] [CrossRef] [PubMed]
- Bakker, G.C.; van Erk, M.J.; Pellis, L.; Wopereis, S.; Rubingh, C.M.; Cnubben, N.H.; Kooistra, T.; van Ommen, B.; Hendriks, H.F. An antiinflammatory dietary mix modulates inflammation and oxidative and metabolic stress in overweight men: A nutrigenomics approach. Am. J. Clin. Nutr. 2010, 91, 1044–1059. [Google Scholar] [CrossRef] [PubMed]
- Paquette, M.; Medina Larqué, A.S.; Weisnagel, S.J.; Desjardins, Y.; Marois, J.; Pilon, G.; Dudonné, S.; Marette, A.; Jacques, H. Strawberry and cranberry polyphenols improve insulin sensitivity in insulin-resistant, non-diabetic adults: A parallel, double-blind, controlled and randomised clinical trial. Br. J. Nutr. 2017, 117, 519–531. [Google Scholar] [CrossRef] [PubMed]
- Riedl, A.; Gieger, C.; Hauner, H.; Daniel, H.; Linseisen, J. Metabotyping and its application in targeted nutrition: An overview. Br. J. Nutr. 2017, 117, 1631–1644. [Google Scholar] [CrossRef] [PubMed]
- Connaugton, R.M.; Mcmorrow, A.M.; Healy, M.L.; Mcgillicuddy, F.C.; Lithander, F.E.; Roche, H.M. An anti-inflammatory nutritional intervention selectively improves insulin sensitivity in overweight and obese adolescents wherein baseline metabotype predicts response. Proc. Nutr. Soc. 2017, 73, E84. [Google Scholar] [CrossRef]
- Kang, J.X. Gut microbiota and personalized nutrition. J. Nutrigenet. Nutrigenomics 2013, 6, 6–7. [Google Scholar] [CrossRef] [PubMed]
- Ridaura, V.K.; Faith, J.J.; Rey, F.E.; Cheng, J.; Alexis, E.; Kau, A.L.; Griffin, N.W.; Lombard, V.; Henrissat, B.; Bain, J.R.; et al. Cultured gut microbiota from twins discordant for obesity modulate adiposity and metabolic phenotypes in mice. Science 2014, 341, 1241214. [Google Scholar] [CrossRef] [PubMed]
- Famodu, O.A.; Cuff, C.F.; Cockburn, A.; Downes, M.T.; Murray, P.J.; McFadden, J.W.; Colby, S.E.; Morrell, J.S.; Olfert, I.M. Impact of free-living nutrition intervention on microbiome in college students at risk for Disease: FRUVEDomic pilot study. FASEB J. 2016, 30, 146–147. [Google Scholar]
- Olfert, M.D.; Cuff, C.; Cockburn, A.; Olfert, M.; McFadden, J.W.; Downes, M.; Murray, P.J.; Holaskova, I.; Barr, M.L.; Colby, S.E.; et al. Nutrition Intervention to Profile Microbiome and Behaviors in Young Adults at Risk for Metabolic Syndrome: FRUVEDomic Pilot Study. J. Nutr. Educ. Behav. 2016, 48, S145. [Google Scholar] [CrossRef]
- Mathews, A.T.; Famodu, O.A.; Olfert, M.D.; Murray, P.J.; Cuff, C.F.; Downes, M.T.; Haughey, N.J.; Colby, S.E.; Olfert, I.M.; McFadden, J.W. Fruit and Vegetable Intervention Lowers Circulating Ceramide Levels and Improves Estimated Insulin Sensitivity in Young Adults at Risk of Developing Metabolic Syndrome: A FRUVEDomic Pilot Study. FASEB J. 2016, 30, 1260.3. [Google Scholar]
- Bonder, M.J.; Kurilshikov, A.; Tigchelaar, E.F.; Mujagic, Z.; Imhann, F.; Vila, A.V.; Deelen, P.; Vatanen, T.; Schirmer, M.; Smeekens, S.P.; et al. The effect of host genetics on the gut microbiome. Nat. Genet. 2016, 48, 1407–1412. [Google Scholar] [CrossRef] [PubMed]
- Corella, D.; Arregui, M.; Coltell, O.; Portolés, O.; Guillem-Sáiz, P.; Carrasco, P.; Sorlí, J.V.; Ortega-Azorín, C.; González, J.I.; Ordovás, J.M. Association of the LCT-13910C>T Polymorphism With Obesity and Its Modulation by Dairy Products in a Mediterranean Population. Obesity 2011, 19, 1707–1714. [Google Scholar] [CrossRef] [PubMed]
- Heianza, Y.; Qi, L. Gene-diet interaction and precision nutrition in obesity. Int. J. Mol. Sci. 2017, 18, 787. [Google Scholar] [CrossRef] [PubMed]
- Koeth, R.A.; Wang, Z.; Levison, B.S.; Buffa, J.A.; Org, E.; Sheehy, B.T.; Britt, E.B.; Fu, X.; Wu, Y.; Li, L.; et al. Intestinal microbiota metabolism of L-carnitine, a nutrient in red meat, promotes atherosclerosis. Nat. Med. 2013, 19, 576–585. [Google Scholar] [CrossRef] [PubMed]
- Tang, W.H.W.; Wang, Z.; Levison, B.S.; Koeth, R.A.; Britt, E.B.; Fu, X.; Wu, Y.; Hazen, S.L. Intestinal Microbial Metabolism of Phosphatidylcholine and Cardiovascular Risk. N. Engl. J. Med. 2013, 368, 1575–1584. [Google Scholar] [CrossRef] [PubMed]
- Zmora, N.; Zeevi, D.; Korem, T.; Segal, E.; Elinav, E. Taking it Personally: Personalized Utilization of the Human Microbiome in Health and Disease. Cell Host Microbe 2016, 19, 12–20. [Google Scholar] [CrossRef] [PubMed]
- Rohrmann, S.; Overvad, K.; Bueno-de-Mesquita, H.B.; Jakobsen, M.U.; Egeberg, R.; Tjønneland, A.; Nailler, L.; Boutron-Ruault, M.-C.; Clavel-Chapelon, F.; Krogh, V.; et al. Meat consumption and mortality - results from the European Prospective Investigation into Cancer and Nutrition. BMC Med. 2013, 11, 63. [Google Scholar] [CrossRef] [PubMed]
- Suez, J.; Korem, T.; Zeevi, D.; Zilberman-Schapira, G.; Thaiss, C.A.; Maza, O.; Israeli, D.; Zmora, N.; Gilad, S.; Weinberger, A.; et al. Artificial sweeteners induce glucose intolerance by altering the gut microbiota. Nature 2014, 514, 181–186. [Google Scholar] [CrossRef] [PubMed]
- Bokulich, N.A.; Blaser, M.J. A Bitter Aftertaste: Unintended Effects of Artificial Sweeteners on the Gut Microbiome. Cell Metab. 2014, 20, 701–703. [Google Scholar] [CrossRef] [PubMed]
- Feehley, T.; Nagler, C.R. Health: The weighty costs of non-caloric sweeteners. Nature 2014, 514, 176–177. [Google Scholar] [CrossRef] [PubMed]
- Nettleton, J.E.; Reimer, R.A.; Shearer, J. Reshaping the gut microbiota: Impact of low calorie sweeteners and the link to insulin resistance? Physiol. Behav. 2016, 164, 488–493. [Google Scholar] [CrossRef] [PubMed]
- Frankenfeld, C.L.; Sikaroodi, M.; Lamb, E.; Shoemaker, S.; Gillevet, P.M. High-intensity sweetener consumption and gut microbiome content and predicted gene function in a cross-sectional study of adults in the United States. Ann. Epidemiol. 2015, 25, 736–742. [Google Scholar] [CrossRef] [PubMed]
- Locke, A.E.; Kahali, B.; Berndt, S.I.; Justice, A.E.; Pers, T.H.; Day, F.R.; Powell, C.; Vedantam, S.; Buchkovich, M.L.; Yang, J.; et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 2015, 518, 197–206. [Google Scholar] [CrossRef] [PubMed]
- Fall, T.; Ingelsson, E. Genome-wide association studies of obesity and metabolic syndrome. Mol. Cell. Endocrinol. 2014, 382, 740–757. [Google Scholar] [CrossRef] [PubMed]
- Go, M.J.; Hwang, J.-Y.; Park, T.-J.; Kim, Y.J.; Oh, J.H.; Kim, Y.-J.; Han, B.-G.; Kim, B.-J. Genome-wide association study identifies two novel Loci with sex-specific effects for type 2 diabetes mellitus and glycemic traits in a korean population. Diabetes Metab. J. 2014, 38, 375–387. [Google Scholar] [CrossRef] [PubMed]
- Winkler, T.W.; Justice, A.E.; Graff, M.; Barata, L.; Feitosa, M.F.; Chu, S.; Czajkowski, J.; Esko, T.; Fall, T.; Kilpeläinen, T.O.; et al. The Influence of Age and Sex on Genetic Associations with Adult Body Size and Shape: A Large-Scale Genome-Wide Interaction Study. PLoS Genet. 2015, 11, e1005378. [Google Scholar] [CrossRef] [PubMed]
- Loos, R.J.F.; Lindgren, C.M.; Li, S.; Wheeler, E.; Zhao, J.H.; Prokopenko, I.; Inouye, M.; Freathy, R.M.; Attwood, A.P.; Beckmann, J.S.; et al. Common variants near MC4R are associated with fat mass, weight and risk of obesity. Nat. Genet. 2008, 40, 768–775. [Google Scholar] [CrossRef] [PubMed]
- Schierding, W.; O’Sullivan, J.M. Connecting SNPs in Diabetes: A Spatial Analysis of Meta-GWAS Loci. Front. Endocrinol. (Lausanne) 2015, 6, 102. [Google Scholar] [CrossRef] [PubMed]
- Belsky, D.W.; Moffitt, T.E.; Sugden, K.; Williams, B.; Houts, R.; McCarthy, J.; Caspi, A. Development and Evaluation of a Genetic Risk Score for Obesity. Biodemography Soc. Biol. 2013, 59, 85–100. [Google Scholar] [CrossRef] [PubMed]
- Welter, D.; MacArthur, J.; Morales, J.; Burdett, T.; Hall, P.; Junkins, H.; Klemm, A.; Flicek, P.; Manolio, T.; Hindorff, L.; et al. The NHGRI GWAS Catalog, a curated resource of SNP-trait associations. Nucleic Acids Res. 2014, 42, D1001–D1006. [Google Scholar] [CrossRef] [PubMed]
- Claussnitzer, M.; Dankel, S.N.; Kim, K.-H.; Quon, G.; Meuleman, W.; Haugen, C.; Glunk, V.; Sousa, I.S.; Beaudry, J.L.; Puviindran, V.; et al. FTO Obesity Variant Circuitry and Adipocyte Browning in Humans. N. Engl. J. Med. 2015, 373, 895–907. [Google Scholar] [CrossRef] [PubMed]
- Sofi, F.; Abbate, R.; Gensini, G.F.; Casini, A. Accruing evidence on benefits of adherence to the Mediterranean diet on health: An updated systematic review and meta-analysis. Am. J. Clin. Nutr. 2010, 92, 1189–1196. [Google Scholar] [CrossRef] [PubMed]
- Mente, A.; de Koning, L.; Shannon, H.S.; Anand, S.S. A Systematic Review of the Evidence Supporting a Causal Link Between Dietary Factors and Coronary Heart Disease. Arch. Intern. Med. 2009, 169, 659. [Google Scholar] [CrossRef] [PubMed]
- Martínez-González, M.A.; García-Arellano, A.; Toledo, E.; Salas-Salvadó, J.; Buil-Cosiales, P.; Corella, D.; Covas, M.I.; Schröder, H.; Arós, F.; Gómez-Gracia, E.; et al. A 14-Item Mediterranean Diet Assessment Tool and Obesity Indexes among High-Risk Subjects: The PREDIMED Trial. PLoS ONE 2012, 7, e43134. [Google Scholar] [CrossRef] [PubMed]
- Fito, M.; Melander, O.; Martinez, J.A.; Toledo, E.; Carpene, C.; Corella, D. Advances in integrating traditional and omic biomarkers when analyzing the effects of the mediterranean diet intervention in cardiovascular prevention. Int. J. Mol. Sci. 2016, 17, 1469. [Google Scholar] [CrossRef] [PubMed]
- Ortega-Azorin, C.; Sorli, J. V.; Estruch, R.; Asensio, E.M.; Coltell, O.; Gonzalez, J.I.; Martinez-Gonzalez, M.A.; Ros, E.; Salas-Salvado, J.; Fito, M.; et al. Amino Acid Change in the Carbohydrate Response Element Binding Protein Is Associated With Lower Triglycerides and Myocardial Infarction Incidence Depending on Level of Adherence to the Mediterranean Diet in the PREDIMED Trial. Circ. Cardiovasc. Genet. 2014, 7, 49–58. [Google Scholar] [CrossRef] [PubMed]
- Rudkowska, I.; Guenard, F.; Julien, P.; Couture, P.; Lemieux, S.; Barbier, O.; Calder, P.C.; Minihane, A.M.; Vohl, M.-C. Genome-wide association study of the plasma triglyceride response to an n-3 polyunsaturated fatty acid supplementation. J. Lipid Res. 2014, 55, 1245–1253. [Google Scholar] [CrossRef] [PubMed]
- Ortega-Azorín, C.; Sorlí, J.V.; Asensio, E.M.; Coltell, O.; Martínez-González, M.; Salas-Salvadó, J.; Covas, M.-I.; Arós, F.; Lapetra, J.; Serra-Majem, L.; et al. Associations of the FTO rs9939609 and the MC4R rs17782313 polymorphisms with type 2 diabetes are modulated by diet, being higher when adherence to the Mediterranean diet pattern is low. Cardiovasc. Diabetol. 2012, 11, 137. [Google Scholar] [CrossRef]
- Garcia-Aloy, M.; Llorach, R.; Urpi-Sarda, M.; Jáuregui, O.; Corella, D.; Ruiz-Canela, M.; Salas-Salvadó, J.; Fitó, M.; Ros, E.; Estruch, R.; et al. A metabolomics-driven approach to predict cocoa product consumption by designing a multimetabolite biomarker model in free-living subjects from the PREDIMED study. Mol. Nutr. Food Res. 2015, 59, 212–220. [Google Scholar] [CrossRef] [PubMed]
- Ryan, N.M.; O’Donovan, C.B.; Forster, H.; Woolhead, C.; Walsh, M.C. New tools for personalised nutrition: The Food4Me project. Nutr. Bull. 2015, 40, 134–139. [Google Scholar] [CrossRef]
- Livingstone, K.M.; Celis-Morales, C.; Mathers, J. Who Benefits Most from Personalized Nutrition? Findings from the Pan-European Food4Me Randomized Controlled Trial. FASEB J. 2017, 31, 963.4. [Google Scholar]
- Celis-Morales, C.; Livingstone, K.M.; Marsaux, C.F.M.; Forster, H.; O’Donovan, C.B.; Woolhead, C.; Macready, A.L.; Fallaize, R.; Navas-Carretero, S.; San-Cristobal, R.; et al. Design and baseline characteristics of the Food4Me study: A web-based randomised controlled trial of personalised nutrition in seven European countries. Genes Nutr. 2015, 10, 450. [Google Scholar] [CrossRef] [PubMed]
- Fallaize, R.; Forster, H.; Macready, A.L.; Walsh, M.C.; Mathers, J.C.; Brennan, L.; Gibney, E.R.; Gibney, M.J.; Lovegrove, J.A. Online dietary intake estimation: Reproducibility and validity of the Food4Me food frequency questionnaire against a 4-day weighed food record. J. Med. Internet Res. 2014, 16, e190. [Google Scholar] [CrossRef] [PubMed]
- Forster, H.; Fallaize, R.; Gallagher, C.; O’Donovan, C.B.; Woolhead, C.; Walsh, M.C.; Macready, A.L.; Lovegrove, J.A.; Mathers, J.C.; Gibney, M.J.; et al. Online Dietary Intake Estimation: The Food4Me Food Frequency Questionnaire. J. Med. Internet Res. 2014, 16, e150. [Google Scholar] [CrossRef] [PubMed]
- Baecke, J.A.H.; Burema, J.; Frijters, J.E.R. A short questionnaire for the measurement of habitual physical activity in epidemiological studies. Am. J. Clin. Nutr. 1982, 36, 936–942. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, D.E.; El-Sohemy, A. Disclosure of Genetic Information and Change in Dietary Intake: A Randomized Controlled Trial. PLoS ONE 2014, 9, e112665. [Google Scholar] [CrossRef] [PubMed]
- Celis-Morales, C.; Livingstone, K.M.; Marsaux, C.F.M.; Macready, A.L.; Fallaize, R.; O’Donovan, C.B.; Woolhead, C.; Forster, H.; Walsh, M.C.; Navas-Carretero, S.; et al. Effect of personalized nutrition on health-related behaviour change: Evidence from the Food4me European randomized controlled trial. Int. J. Epidemiol. 2016. [Google Scholar] [CrossRef] [PubMed]
- Guenther, P.M.; Casavale, K.O.; Reedy, J.; Kirkpatrick, S.I.; Hiza, H.A.B.; Kuczynski, K.J.; Kahle, L.L.; Krebs-Smith, S.M. Update of the Healthy Eating Index: HEI-2010. J. Acad. Nutr. Diet. 2013, 113, 569–580. [Google Scholar] [CrossRef] [PubMed]
- Kirwan, L.; Walsh, M.C.; Celis-Morales, C.; Marsaux, C.F.M.; Livingstone, K.M.; Navas-Carretero, S.; Fallaize, R.; O’Donovan, C.B.; Woolhead, C.; Forster, H.; et al. Phenotypic factors influencing the variation in response of circulating cholesterol level to personalised dietary advice in the Food4Me study. Br. J. Nutr. 2016, 116, 2011–2019. [Google Scholar] [CrossRef] [PubMed]
- Celis-Morales, C.; Marsaux, C.F.; Livingstone, K.M.; Navas-Carretero, S.; San-Cristobal, R.; Fallaize, R.; Macready, A.L.; O’Donovan, C.; Woolhead, C.; Forster, H.; et al. Can genetic-based advice help you lose weight? Findings from the Food4Me European randomized controlled trial. Am. J. Clin. Nutr. 2017, 105, 1204–1213. [Google Scholar] [CrossRef] [PubMed]
- Livingstone, K.M.; Celis-Morales, C.; Papandonatos, G.D.; Erar, B.; Florez, J.C.; Jablonski, K.A.; Razquin, C.; Marti, A.; Heianza, Y.; Huang, T.; et al. FTO genotype and weight loss: Systematic review and meta-analysis of 9563 individual participant data from eight randomised controlled trials. BMJ 2016, 354, i4707. [Google Scholar] [CrossRef] [PubMed]
- Ramos-Lopez, O.; Milagro, F.I.; Allayee, H.; Chmurzynska, A.; Choi, M.S.; Curi, R.; De Caterina, R.; Ferguson, L.R.; Goni, L.; Kang, J.X.; et al. Guide for Current Nutrigenetic, Nutrigenomic, and Nutriepigenetic Approaches for Precision Nutrition Involving the Prevention and Management of Chronic Diseases Associated with Obesity. J. Nutrigenet. Nutrigenomics 2017, 10, 43–62. [Google Scholar] [CrossRef] [PubMed]
- Abrahams, M.; Frewer, L.J.; Bryant, E.; Stewart-Knox, B. Factors determining the integration of nutritional genomics into clinical practice by registered dietitians. Trends Food Sci. Technol. 2017, 59, 139–147. [Google Scholar] [CrossRef]
- Cormier, H.; Tremblay, B.L.; Paradis, A.-M.; Garneau, V.; Desroches, S.; Robitaille, J.; Vohl, M.-C. Nutrigenomics-perspectives from registered dietitians: A report from the Quebec-wide e-consultation on nutrigenomics among registered dietitians. J. Hum. Nutr. Diet. 2014, 27, 391–400. [Google Scholar] [CrossRef] [PubMed]
- Kohlmeier, M.; De Caterina, R.; Ferguson, L.R.; Görman, U.; Allayee, H.; Prasad, C.; Kang, J.X.; Nicoletti, C.F.; Martinez, J.A. Guide and Position of the International Society of Nutrigenetics/Nutrigenomics on Personalized Nutrition: Part 2-Ethics, Challenges and Endeavors of Precision Nutrition. J. Nutrigenet. Nutrigenomics 2016, 9, 28–46. [Google Scholar] [CrossRef] [PubMed]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
De Toro-Martín, J.; Arsenault, B.J.; Després, J.-P.; Vohl, M.-C. Precision Nutrition: A Review of Personalized Nutritional Approaches for the Prevention and Management of Metabolic Syndrome. Nutrients 2017, 9, 913. https://doi.org/10.3390/nu9080913
De Toro-Martín J, Arsenault BJ, Després J-P, Vohl M-C. Precision Nutrition: A Review of Personalized Nutritional Approaches for the Prevention and Management of Metabolic Syndrome. Nutrients. 2017; 9(8):913. https://doi.org/10.3390/nu9080913
Chicago/Turabian StyleDe Toro-Martín, Juan, Benoit J. Arsenault, Jean-Pierre Després, and Marie-Claude Vohl. 2017. "Precision Nutrition: A Review of Personalized Nutritional Approaches for the Prevention and Management of Metabolic Syndrome" Nutrients 9, no. 8: 913. https://doi.org/10.3390/nu9080913
APA StyleDe Toro-Martín, J., Arsenault, B. J., Després, J.-P., & Vohl, M.-C. (2017). Precision Nutrition: A Review of Personalized Nutritional Approaches for the Prevention and Management of Metabolic Syndrome. Nutrients, 9(8), 913. https://doi.org/10.3390/nu9080913





