Uteroplacental Insufficiency Causes Microbiota Disruption and Lung Development Impairment in Growth-Restricted Newborn Rats
Abstract
1. Introduction
2. Materials and Methods
2.1. Animal Model
2.2. Tissue Collection and Processing
2.3. Lung Histology
2.4. Immunohistochemistry of Vascular Endothelial Growth Factor and von Willebrand Factor
2.5. Western Blot Analysis of VEGF
2.6. 16S rRNA Gene Sequencing and Analysis
2.7. Statistical Analysis
3. Results
3.1. UPI Impaired Lung Development in Growth Restricted Newborn Rats
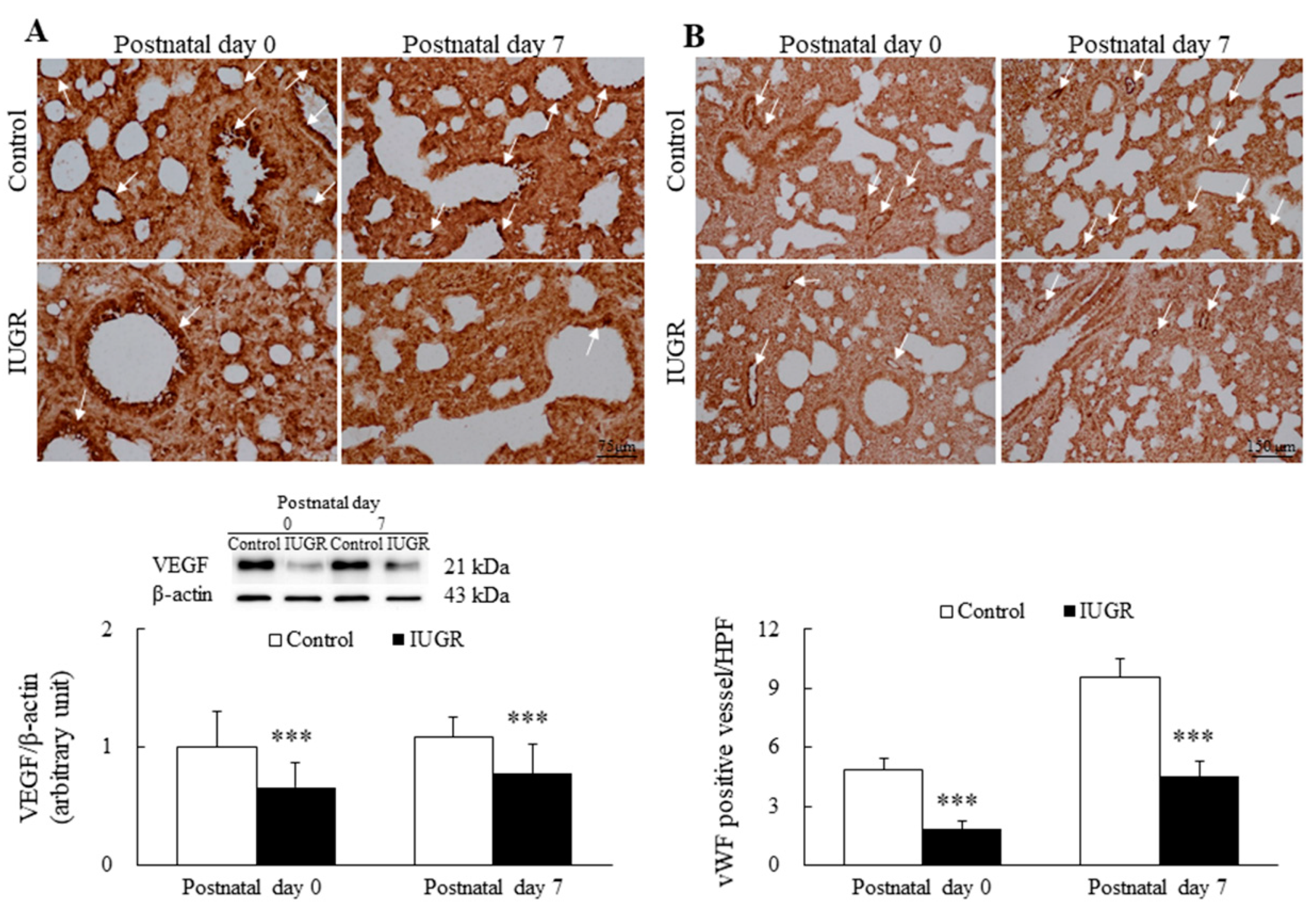
3.2. UPI Impaired Lung Angiogenesis in Growth-Restricted Newborn Rats
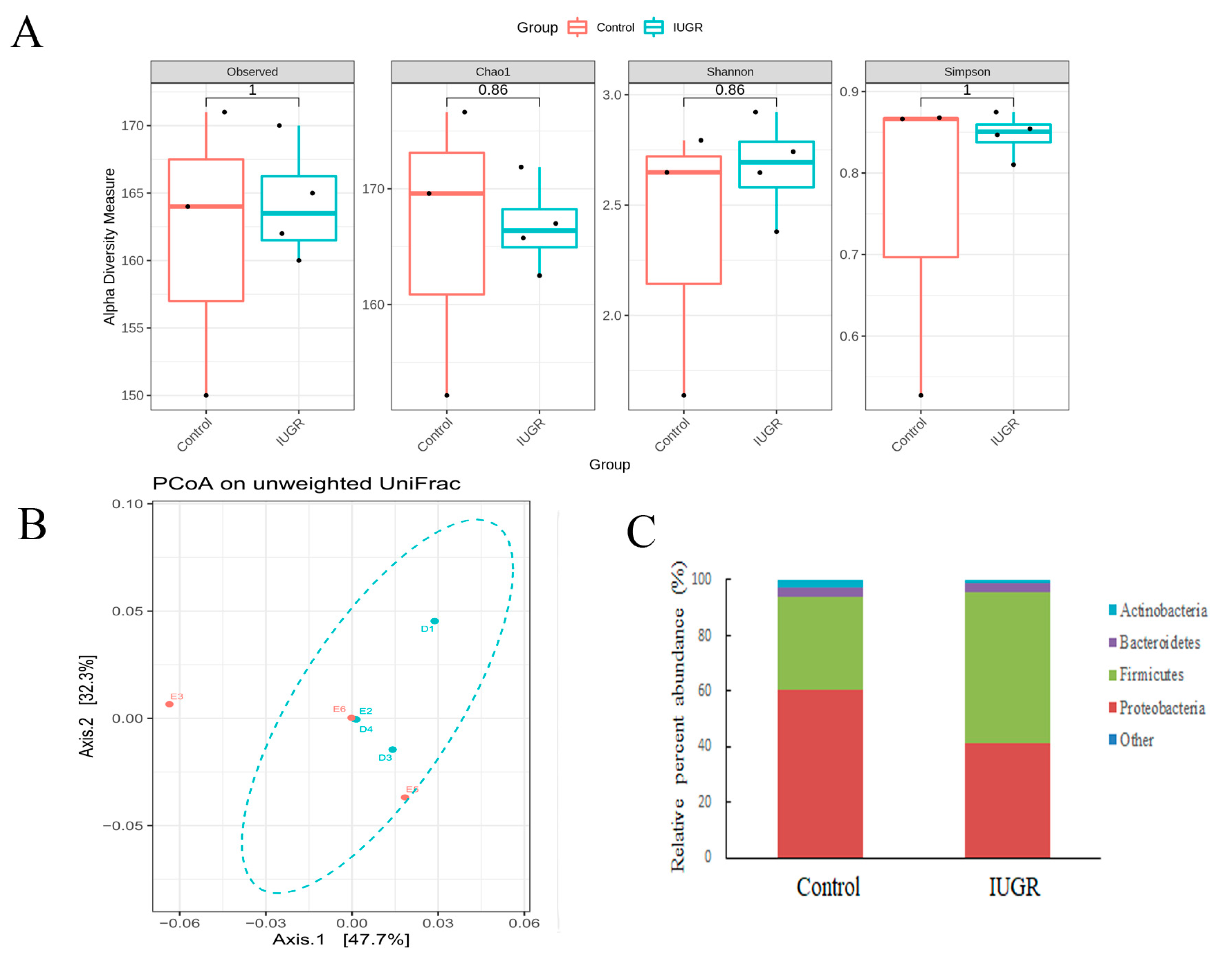
3.3. UPI Did Not Alter the Maternal Intestinal Microbiota
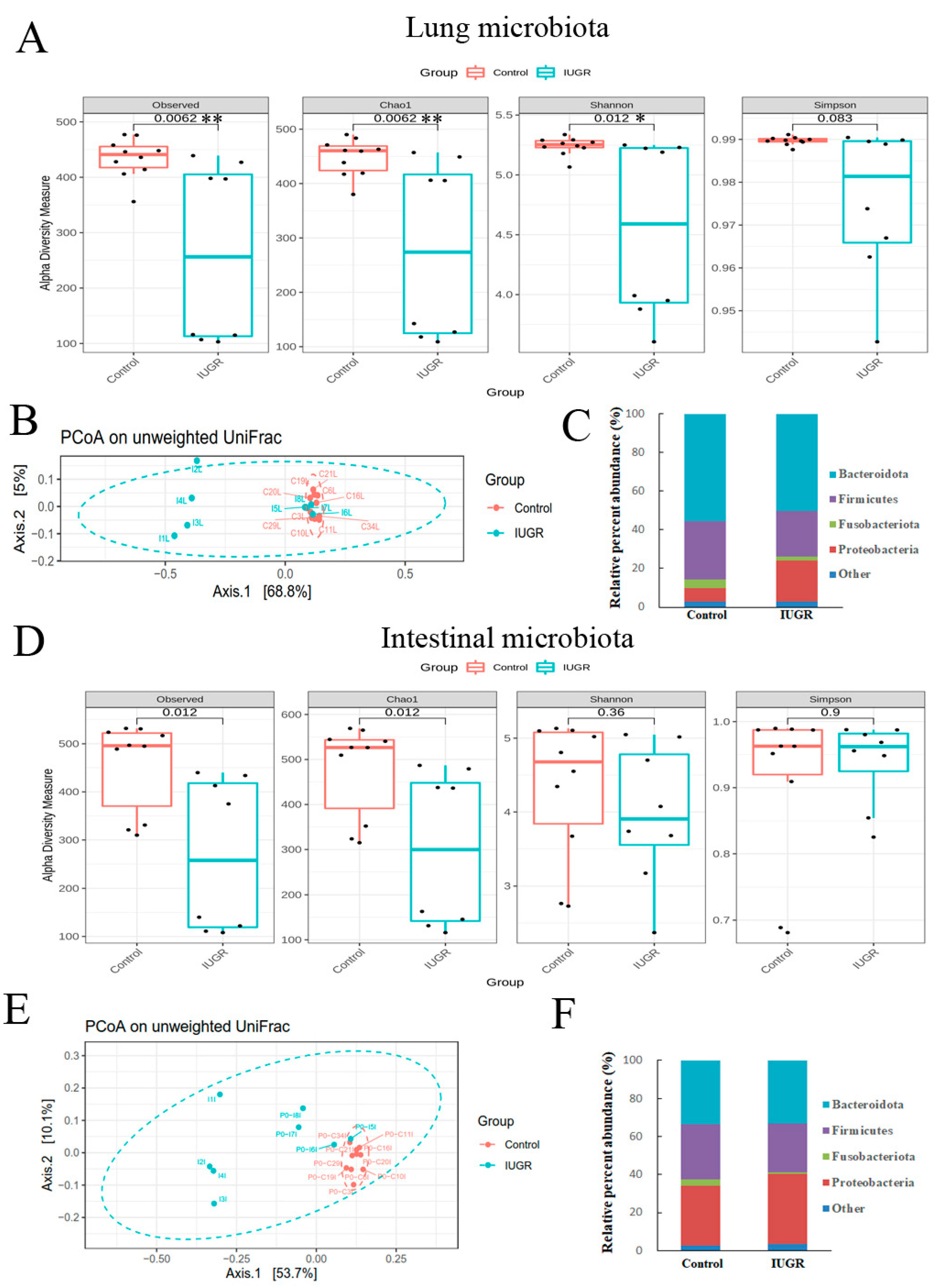
3.4. IUGR Altered the Lung and Intestinal Microbiota on Postnatal Day 0
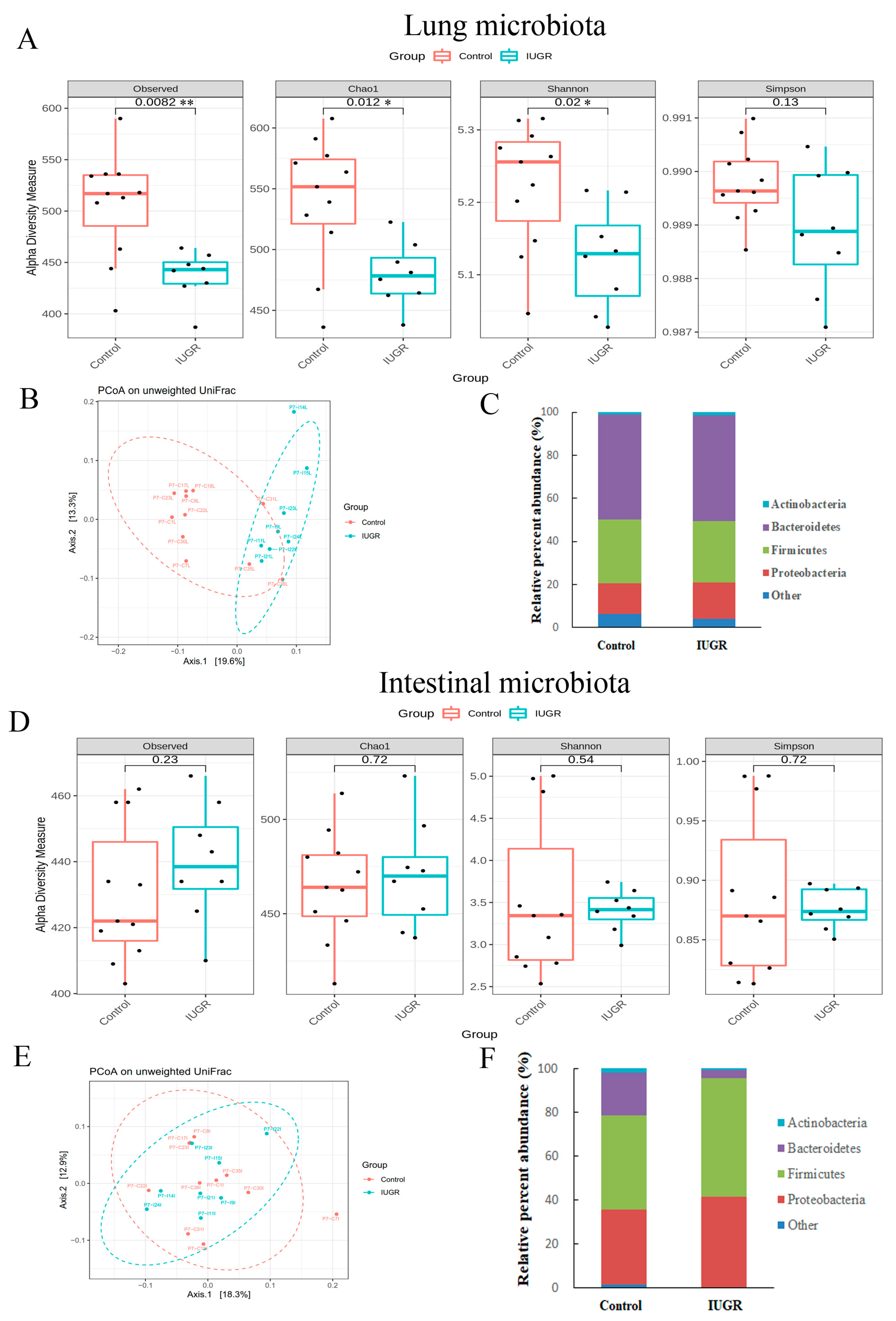
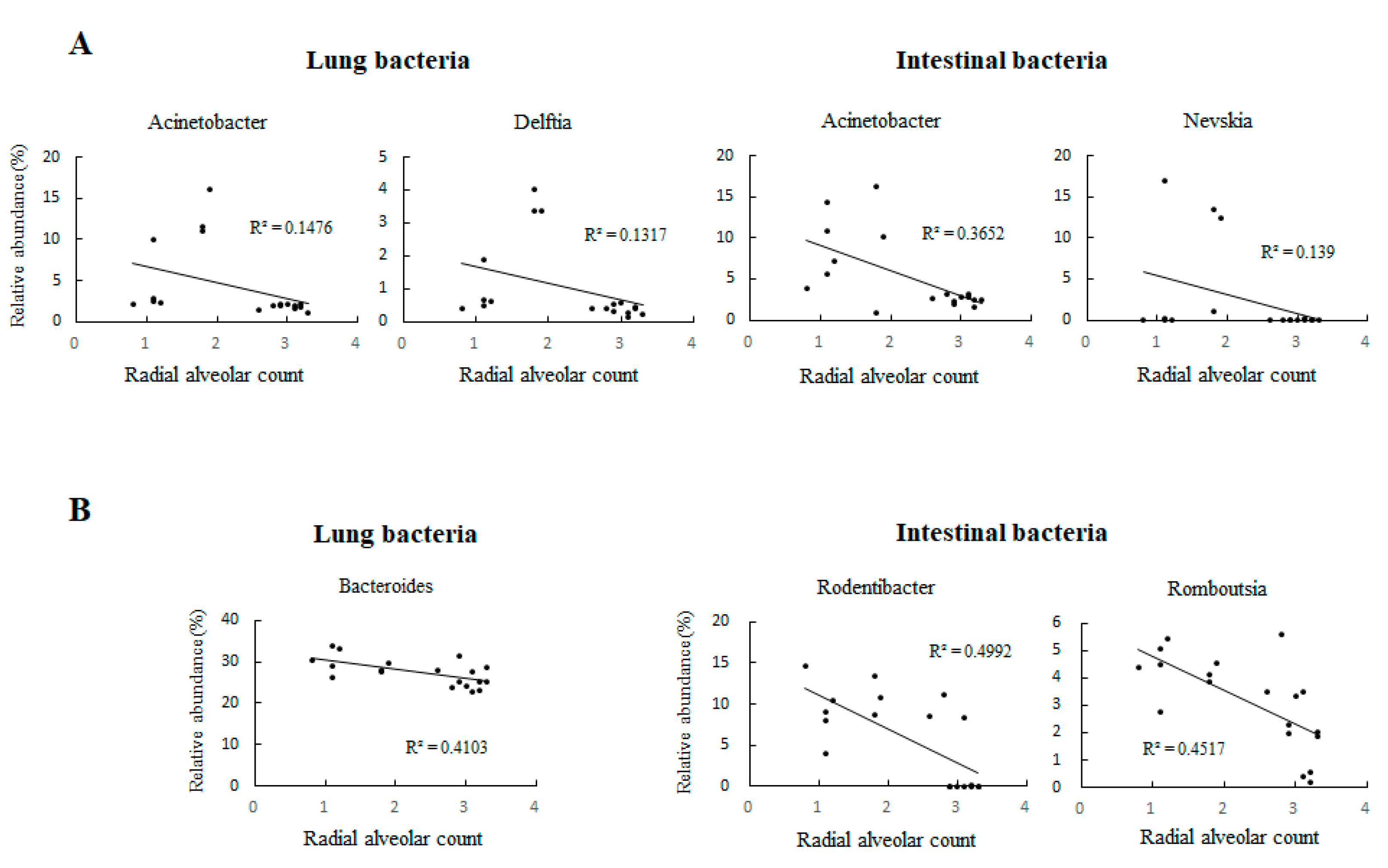
3.5. IUGR Altered the Lung Microbiota but Did Not Alter the Intestinal Microbiota on Postnatal Day 7
3.6. Lung Microbiota Compositions on Postnatal Days 0 and 7
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Harding, R.; Cock, M.L.; Louey, S.; Joyce, B.J.; Davey, M.G.; Albuquerque, C.A.; Hooper, S.B.; Maritz, G.S. The compromised intra-uterine environment: Implications for future lung health. Clin. Exp. Pharmacol. Physiol. 2000, 27, 965–974. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.M.; Wang, L.F.; Su, B. Effects of maternal undernutrition during late gestation on the lung surfactant system and morphometry in rats. Pediatr. Res. 2004, 56, 329–335. [Google Scholar] [CrossRef]
- Huang, L.T.; Chou, H.C.; Lin, C.M.; Chen, C.M. Uteroplacental Insufficiency Alters the Retinoid Pathway and Lung Development in Newborn Rats. Pediatr. Neonatol. 2016, 57, 508–514. [Google Scholar] [CrossRef] [PubMed]
- Arigliani, M.; Stocco, C.; Valentini, E.; De Pieri, C.; Castriotta, L.; Ferrari, M.E.; Canciani, C.; Driul, L.; Orsaria, M.; Cattarossi, L.; et al. Lung function between 8 and 15 years of age in very preterm infants with fetal growth restriction. Pediatr. Res. 2021, 90, 657–663. [Google Scholar] [CrossRef] [PubMed]
- den Dekker, H.T.; Jaddoe, V.W.V.; Reiss, I.K.; de Jongste, J.C.; Duijts, L. Fetal and Infant Growth Patterns and Risk of Lower Lung Function and Asthma. The Generation R Study. Am. J. Respir. Crit. Care Med. 2018, 197, 183–192. [Google Scholar] [CrossRef] [PubMed]
- Jordan, B.K.; McEvoy, C.T. Trajectories of Lung Function in Infants and Children: Setting a Course for Lifelong Lung Health. Pediatrics 2020, 146, e20200417. [Google Scholar] [CrossRef]
- Kotecha, S.J.; Watkins, W.J.; Heron, J.; Henderson, J.; Dunstan, F.D.; Kotecha, S. Spirometric lung function in school-age children: Effect of intrauterine growth retardation and catch-up growth. Am. J. Respir. Crit. Care Med. 2010, 181, 969–974. [Google Scholar] [CrossRef]
- Pike, K.; Jane Pillow, J.; Lucas, J.S. Long term respiratory consequences of intrauterine growth restriction. Semin. Fetal Neonatal Med. 2012, 17, 92–98. [Google Scholar] [CrossRef]
- Henriksen, T.; Clausen, T. The fetal origins hypothesis: Placental insufficiency and inheritance versus maternal malnutrition in well-nourished populations. Acta Obstet. Gynecol. Scand. 2002, 81, 112–114. [Google Scholar] [CrossRef]
- Wu, G.; Bazer, F.W.; Wallace, J.M.; Spencer, T.E. Board-invited review: Intrauterine growth retardation: Implications for the animal sciences. J. Anim. Sci. 2006, 84, 2316–2337. [Google Scholar] [CrossRef]
- Pierro, M.; Villamor-Martinez, E.; van Westering-Kroon, E.; Alvarez-Fuente, M.; Abman, S.H.; Villamor, E. Association of the dysfunctional placentation endotype of prematurity with bronchopulmonary dysplasia: A systematic review, meta-analysis and meta-regression. Thorax 2022, 77, 268–275. [Google Scholar] [CrossRef] [PubMed]
- Sacchi, C.; Marino, C.; Nosarti, C.; Vieno, A.; Visentin, S.; Simonelli, A. Association of Intrauterine Growth Restriction and Small for Gestational Age Status With Childhood Cognitive Outcomes: A Systematic Review and Meta-analysis. JAMA Pediatr. 2020, 174, 772–781. [Google Scholar] [CrossRef] [PubMed]
- Amon, P.; Sanderson, I. What is the microbiome? Arch. Dis. Child. Educ. Pract. Ed. 2017, 102, 257–260. [Google Scholar] [CrossRef] [PubMed]
- Cryan, J.F.; Dinan, T.G. Gut microbiota: Microbiota and neuroimmune signalling-Metchnikoff to microglia. Nat. Rev. Gastroenterol. Hepatol. 2015, 12, 494–496. [Google Scholar] [CrossRef] [PubMed]
- Tamburini, S.; Shen, N.; Wu, H.C.; Clemente, J.C. The microbiome in early life: Implications for health outcomes. Nat. Med. 2016, 22, 713–722. [Google Scholar] [CrossRef]
- Fanca-Berthon, P.; Hoebler, C.; Mouzet, E.; David, A.; Michel, C. Intrauterine growth restriction not only modifies the cecocolonic microbiota in neonatal rats but also affects its activity in young adult rats. J. Pediatr. Gastroenterol. Nutr. 2010, 51, 402–413. [Google Scholar] [CrossRef]
- Huang, S.; Li, N.; Liu, C.; Li, T.; Wang, W.; Jiang, L.; Li, Z.; Han, D.; Tao, S.; Wang, J. Characteristics of the gut microbiota colonization, inflammatory profile, and plasma metabolome in intrauterine growth restricted piglets during the first 12 hours after birth. J. Microbiol. 2019, 57, 748–758. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Ma, C.; Xie, P.; Zhu, Q.; Wang, X.; Yin, Y.; Kong, X. Gut microbiota of newborn piglets with intrauterine growth restriction have lower diversity and different taxonomic abundances. J. Appl. Microbiol. 2019, 127, 354–369. [Google Scholar] [CrossRef]
- Cooney, T.P.; Thurlbeck, W.M. The radial alveolar count method of Emery and Mithal: A reappraisal 2--intrauterine and early postnatal lung growth. Thorax 1982, 37, 580–583. [Google Scholar] [CrossRef]
- Irwin, D.; Helm, K.; Campbell, N.; Imamura, M.; Fagan, K.; Harral, J.; Carr, M.; Young, K.A.; Klemm, D.; Gebb, S.; et al. Neonatal lung side population cells demonstrate endothelial potential and are altered in response to hyperoxia-induced lung simplification. Am. J. Physiol. Lung Cell. Mol. Physiol. 2007, 293, L941–L951. [Google Scholar] [CrossRef]
- Yang, Y.S.H.; Chang, H.W.; Lin, I.H.; Chien, L.N.; Wu, M.J.; Liu, Y.R.; Chu, P.G.; Xie, G.; Dong, F.; Jia, W.; et al. Long-term Proton Pump Inhibitor Administration Caused Physiological and Microbiota Changes in Rats. Sci. Rep. 2020, 10, 866. [Google Scholar] [CrossRef] [PubMed]
- Callahan, B.J.; Sankaran, K.; Fukuyama, J.A.; McMurdie, P.J.; Holmes, S.P. Bioconductor Workflow for Microbiome Data Analysis: From raw reads to community analyses. F1000Res 2016, 5, 1492. [Google Scholar] [CrossRef] [PubMed]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glockner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, D590–D596. [Google Scholar] [CrossRef] [PubMed]
- Schliep, K.P. phangorn: Phylogenetic analysis in R. Bioinformatics 2011, 27, 592–593. [Google Scholar] [CrossRef]
- McMurdie, P.J.; Holmes, S. phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef]
- Chen, J.; Bittinger, K.; Charlson, E.S.; Hoffmann, C.; Lewis, J.; Wu, G.D.; Collman, R.G.; Bushman, F.D.; Li, H. Associating microbiome composition with environmental covariates using generalized UniFrac distances. Bioinformatics 2012, 28, 2106–2113. [Google Scholar] [CrossRef]
- Hu, J.; Robinson, J.L. Systematic review of invasive Acinetobacter infections in children. Can. J. Infect. Dis. Med. Microbiol. 2010, 21, 83–88. [Google Scholar] [CrossRef]
- Ertan, K.; Yilmaz, R. Delftia Acidovorans sepsis in an immunodeficient child with congenital heart disease. Pediatr. Pract. Res. 2020, 8, 46–49. [Google Scholar]
- Bilgin, H.; Sarmis, A.; Tigen, E.; Soyletir, G.; Mulazimoglu, L. Delftia acidovorans: A rare pathogen in immunocompetent and immunocompromised patients. Can. J. Infect. Dis. Med. Microbiol. 2015, 26, 277–279. [Google Scholar] [CrossRef]
- Dickson, R.P.; Singer, B.H.; Newstead, M.W.; Falkowski, N.R.; Erb-Downward, J.R.; Standiford, T.J.; Huffnagle, G.B. Enrichment of the lung microbiome with gut bacteria in sepsis and the acute respiratory distress syndrome. Nat. Microbiol. 2016, 1, 16113. [Google Scholar] [CrossRef]
- Tanaka, M.; Nakayama, J. Development of the gut microbiota in infancy and its impact on health in later life. Allergol. Int. 2017, 66, 515–522. [Google Scholar] [CrossRef] [PubMed]
- Jimenez, E.; Marin, M.L.; Martin, R.; Odriozola, J.M.; Olivares, M.; Xaus, J.; Fernandez, L.; Rodriguez, J.M. Is meconium from healthy newborns actually sterile? Res. Microbiol. 2008, 159, 187–193. [Google Scholar] [CrossRef] [PubMed]
- Zakosek Pipan, M.; Kajdic, L.; Kalin, A.; Plavec, T.; Zdovc, I. Do newborn puppies have their own microbiota at birth? Influence of type of birth on newborn puppy microbiota. Theriogenology 2020, 152, 18–28. [Google Scholar] [CrossRef] [PubMed]
- Peters, K.G.; De Vries, C.; Williams, L.T. Vascular endothelial growth factor receptor expression during embryogenesis and tissue repair suggests a role in endothelial differentiation and blood vessel growth. Proc. Natl. Acad. Sci. USA 1993, 90, 8915–8919. [Google Scholar] [CrossRef] [PubMed]
- Jakkula, M.; Le Cras, T.D.; Gebb, S.; Hirth, K.P.; Tuder, R.M.; Voelkel, N.F.; Abman, S.H. Inhibition of angiogenesis decreases alveolarization in the developing rat lung. Am. J. Physiol. Lung Cell. Mol. Physiol. 2000, 279, L600–L607. [Google Scholar] [CrossRef]
- Liu, X.; Lin, Y.; Tian, B.; Miao, J.; Xi, C.; Liu, C. Maternal protein restriction alters VEGF signaling and decreases pulmonary alveolar in fetal rats. Int. J. Clin. Exp. Pathol. 2014, 7, 3101–3111. [Google Scholar]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, Y.-C.S.H.; Chou, H.-C.; Liu, Y.-R.; Chen, C.-M. Uteroplacental Insufficiency Causes Microbiota Disruption and Lung Development Impairment in Growth-Restricted Newborn Rats. Nutrients 2022, 14, 4388. https://doi.org/10.3390/nu14204388
Yang Y-CSH, Chou H-C, Liu Y-R, Chen C-M. Uteroplacental Insufficiency Causes Microbiota Disruption and Lung Development Impairment in Growth-Restricted Newborn Rats. Nutrients. 2022; 14(20):4388. https://doi.org/10.3390/nu14204388
Chicago/Turabian StyleYang, Yu-Chen S. H., Hsiu-Chu Chou, Yun-Ru Liu, and Chung-Ming Chen. 2022. "Uteroplacental Insufficiency Causes Microbiota Disruption and Lung Development Impairment in Growth-Restricted Newborn Rats" Nutrients 14, no. 20: 4388. https://doi.org/10.3390/nu14204388
APA StyleYang, Y.-C. S. H., Chou, H.-C., Liu, Y.-R., & Chen, C.-M. (2022). Uteroplacental Insufficiency Causes Microbiota Disruption and Lung Development Impairment in Growth-Restricted Newborn Rats. Nutrients, 14(20), 4388. https://doi.org/10.3390/nu14204388




