A Novel Non-Digestible, Carrot-Derived Polysaccharide (cRG-I) Selectively Modulates the Human Gut Microbiota while Promoting Gut Barrier Integrity: An Integrated In Vitro Approach
Abstract
1. Introduction
2. Materials and Methods
2.1. Products
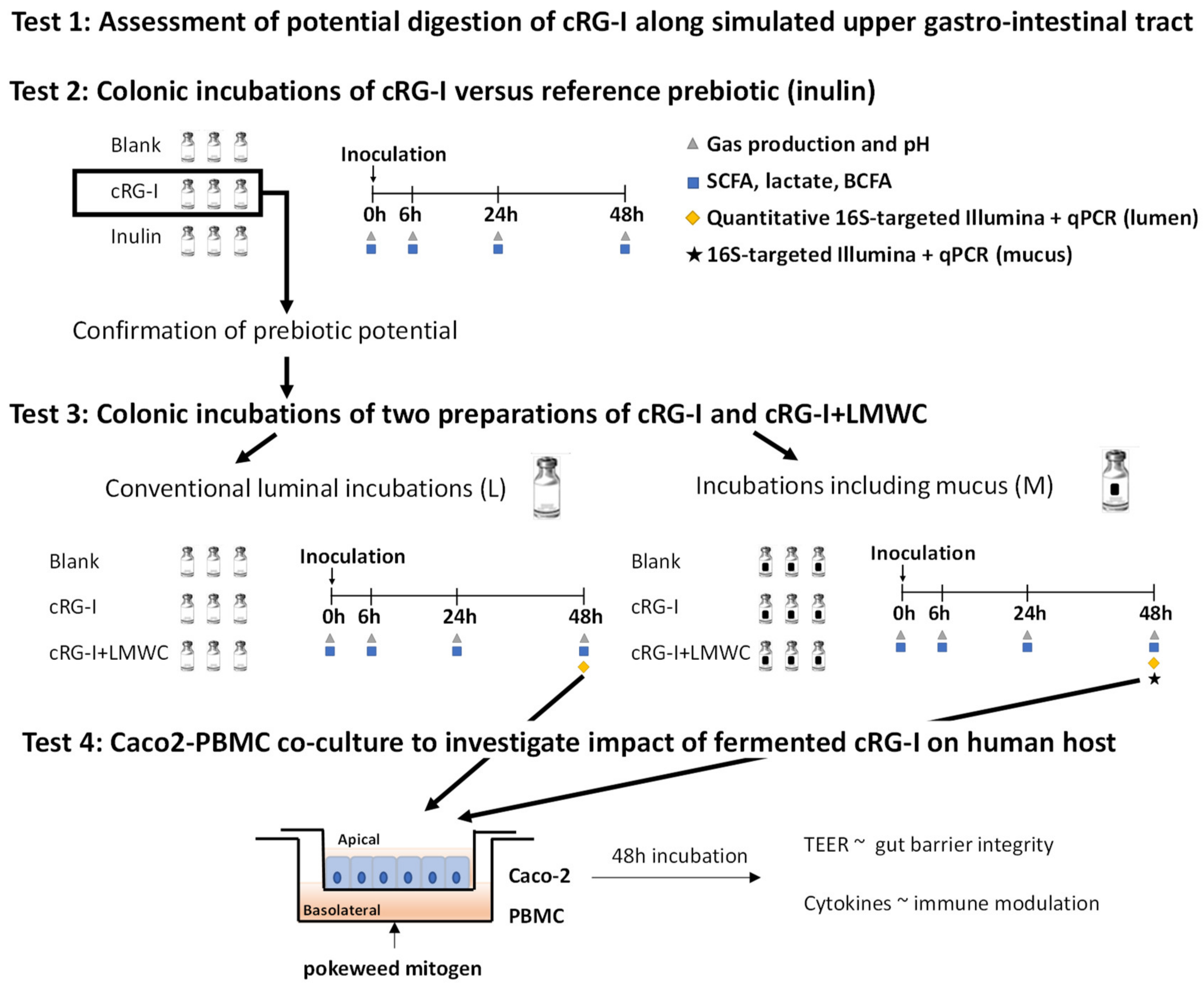
2.2. Digestion by Amylase and Brush Border Enzymes (Test 1)
2.3. Short-Term Colonic Batch Incubations (Tests 2 and 3)
2.4. Microbial Metabolic Activity (Tests 2 and 3)
2.5. Microbial Community Composition (Test 3)
2.6. Caco-2/PBMC Co-Culture Model (Test 4)
2.7. Statistics
3. Results
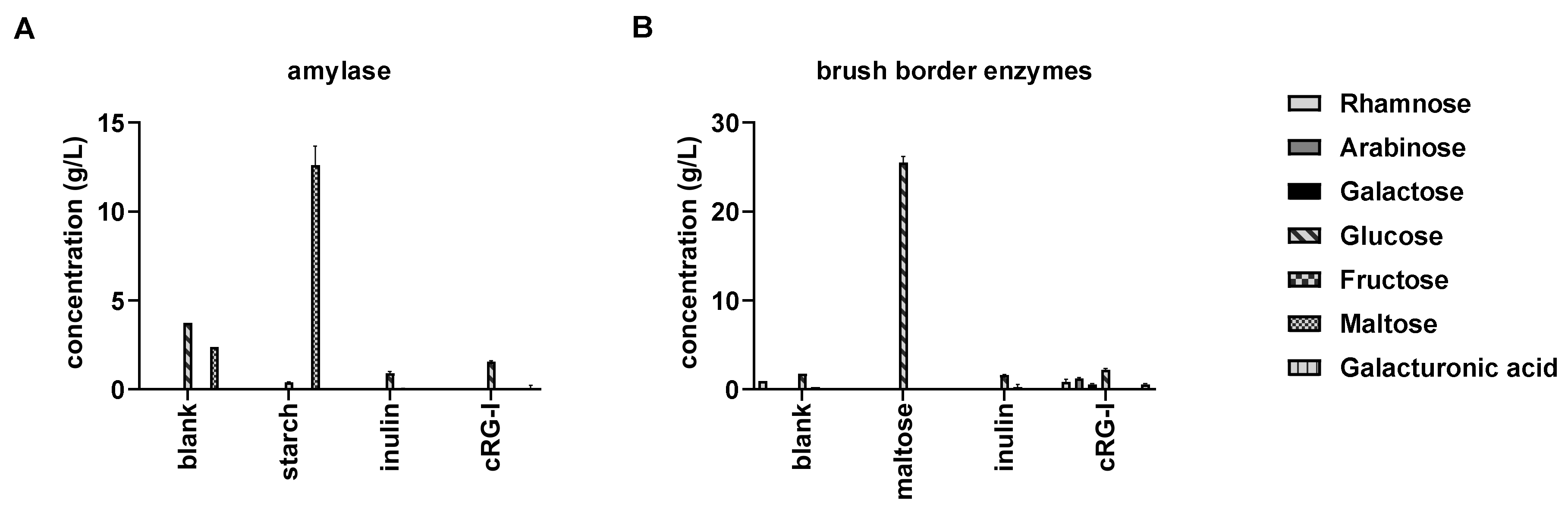
3.1. cRG-I is Resistant to Digestion in the Human Upper Gastro-Intestinal Tract (GIT) (Test 1)
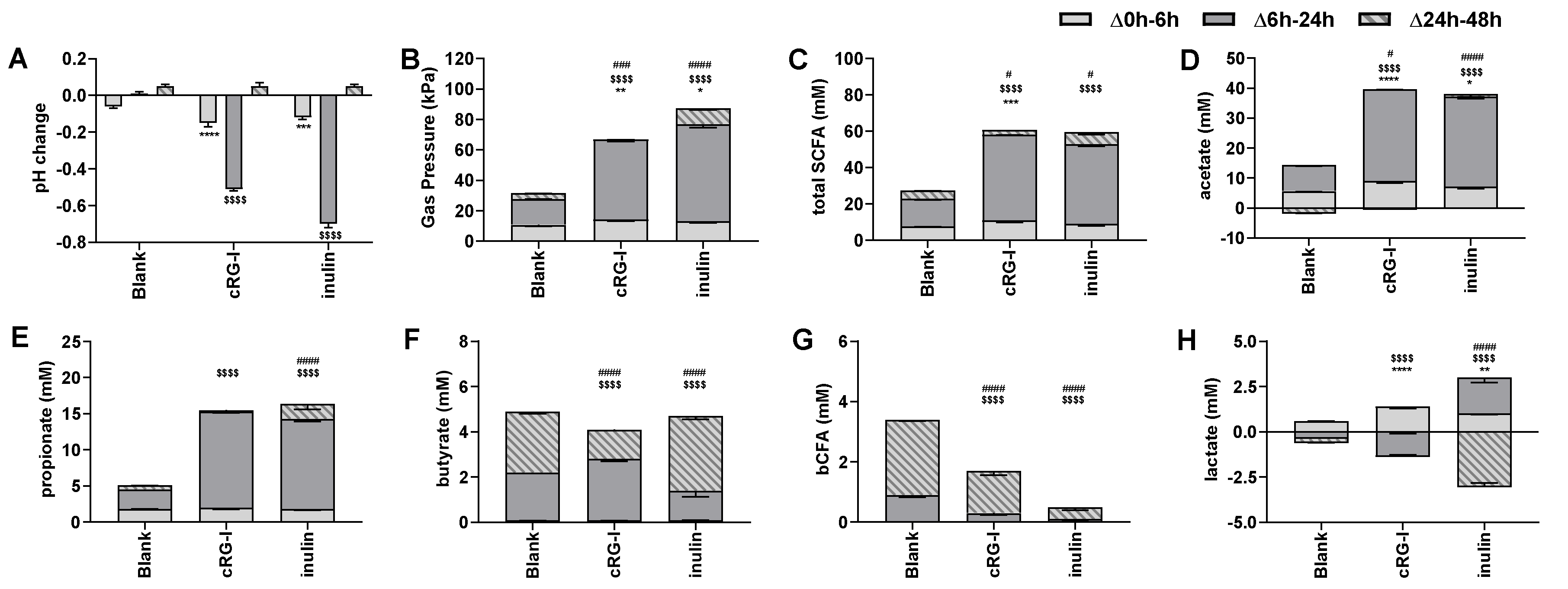
3.2. Effect of cRG-I on Microbial Metabolic Activity in Short-Term Colonic Incubations (Test 2)
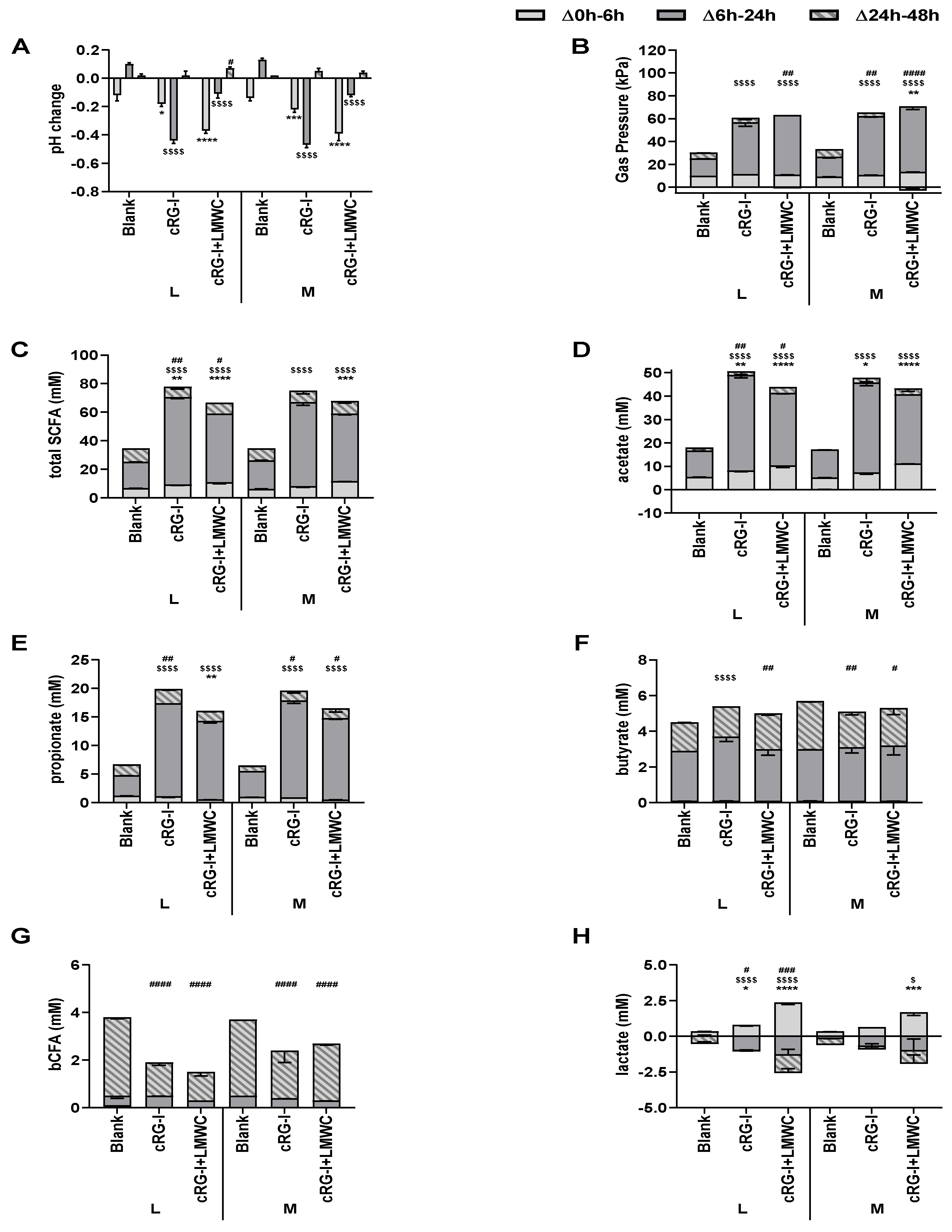
3.3. Effect of cRG-I and cRG-I +LMWC on Microbial Metabolic Activity in Short-Term Colonic Incubations with or without a Mucosal Compartment (Test 3)
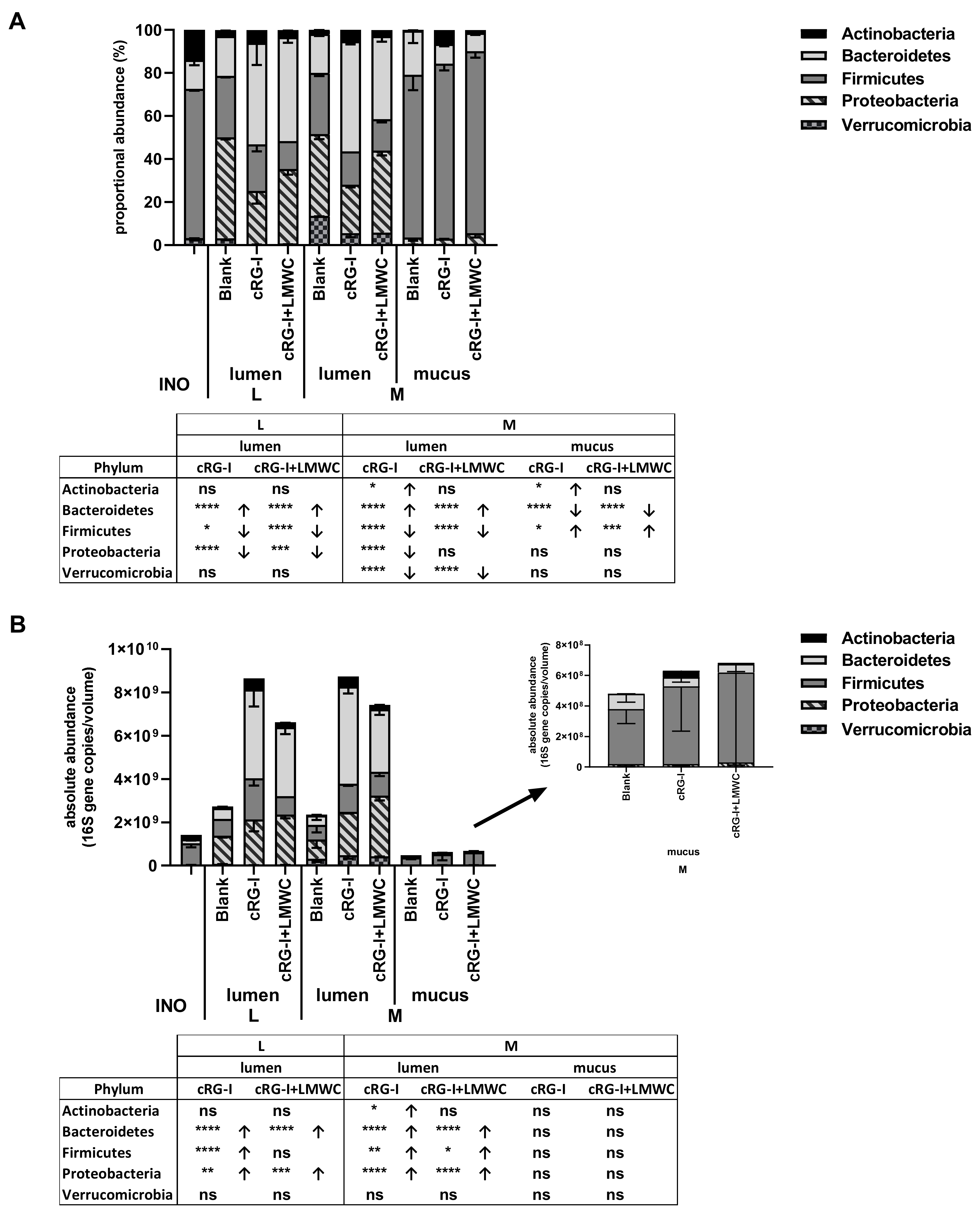
3.4. Effect of cRG-I and cRG-I +LMWC on Microbial Community Composition in Short-Term Colonic Incubations with or without a Mucosal Compartment (Test 3)
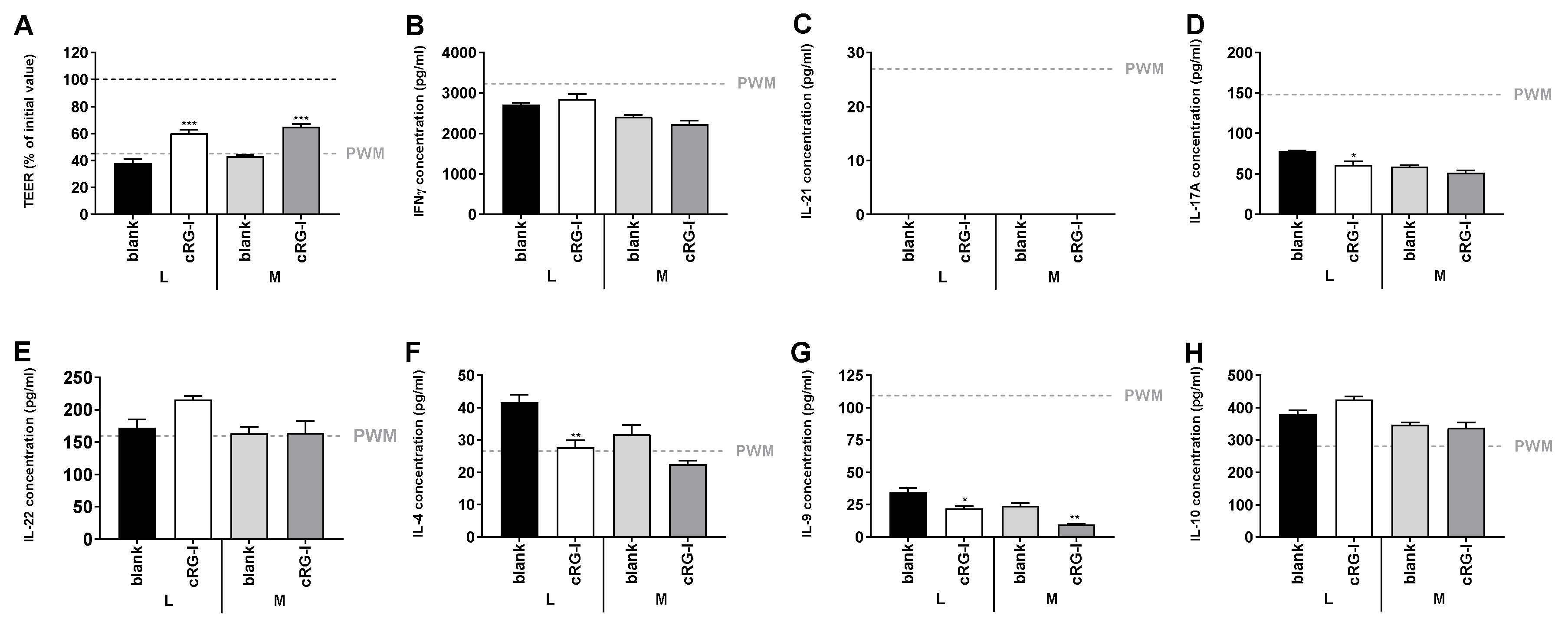
3.5. Effect of Fermented cRG-I on Intestinal Epithelial Barrier (Test 4)
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Forgie, A.J.; Fouhse, J.M.; Willing, B.P. Diet-Microbe-Host Interactions That Affect Gut Mucosal Integrity and Infection Resistance. Front. Immunol. 2019, 10, 1802. [Google Scholar] [CrossRef]
- Monteagudo-Mera, A.; Rastall, R.A.; Gibson, G.R.; Charalampopoulos, D.; Chatzifragkou, A. Adhesion mechanisms mediated by probiotics and prebiotics and their potential impact on human health. Appl. Microbiol. Biotechnol. 2019, 103, 6463–6472. [Google Scholar] [CrossRef]
- Kachrimanidou, M.; Tsintarakis, E. Insights into the Role of Human Gut Microbiota in Clostridioides difficile Infection. Microorganisms 2020, 8, 200. [Google Scholar] [CrossRef]
- Hooper, L.V.; Midtvedt, T.; Gordon, J.I. How host-microbial interactions shape the nutrient environment of the mammalian intestine. Annu. Rev. Nutr. 2002, 22, 283–307. [Google Scholar] [CrossRef]
- Liu, H.; Wang, J.; He, T.; Becker, S.; Zhang, G.; Li, D.; Ma, X. Butyrate: A Double-Edged Sword for Health? Adv. Nutr. 2018, 9, 21–29. [Google Scholar] [CrossRef] [PubMed]
- Delzenne, N.M.; Williams, C.M. Prebiotics and lipid metabolism. Curr. Opin. Lipidol. 2002, 13, 61–67. [Google Scholar] [CrossRef] [PubMed]
- Li, C.J.; Elsasser, T.H. Butyrate-induced apoptosis and cell cycle arrest in bovine kidney epithelial cells: Involvement of caspase and proteasome pathways. J Anim. Sci. 2005, 83, 89–97. [Google Scholar] [CrossRef]
- Jan, G.; Belzacq, A.S.; Haouzi, D.; Rouault, A.; Métivier, D.; Kroemer, G.; Brenner, C. Propionibacteria induce apoptosis of colorectal carcinoma cells via short-chain fatty acids acting on mitochondria. Cell Death Differ. 2002, 9, 179–188. [Google Scholar] [CrossRef]
- Lin, Y.G.; Vonk, R.J.; Slooff, M.J.H.; Kuipers, F.; Smit, M.J. Differences in propionate-induced inhibition of cholesterol and triacylglycerol synthesis between human and rat hepatocytes in primary culture. Br. J. Nutr. 1995, 74, 197–207. [Google Scholar] [CrossRef]
- Nishina, P.M.; Freedland, R.A. Effects of propionate on lipid biosynthesis in isolated rat hepatocytes. J. Nutr. 1990, 120, 668–673. [Google Scholar] [CrossRef]
- Rinninella, E.; Raoul, P.; Cintoni, M.; Franceschi, F.; Miggiano, G.A.D.; Gasbarrini, A.; Mele, M.C. What is the Healthy Gut Microbiota Composition? A Changing Ecosystem across Age, Environment, Diet, and Diseases. Microorganisms 2019, 7, 14. [Google Scholar] [CrossRef] [PubMed]
- Van den Abbeele, P.; Belzer, C.; Goossens, M.; Kleerebezem, M.; De Vos, W.M.; Thas, O.; De Weirdt, R.; Kerckhof, F.M.; Van de Wiele, T. Butyrate-producing Clostridium cluster XIVa species specifically colonize mucins in an in vitro gut model. ISME J. 2013, 7, 949–961. [Google Scholar] [CrossRef] [PubMed]
- Eckburg, P.B.; Bik, E.M.; Bernstein, C.N.; Purdom, E.; Dethlefsen, L.; Sargent, M.; Gill, S.R.; Nelson, K.E.; Relman, D.A. Diversity of the human intestinal microbial flora. Science 2005, 308, 1635–1638. [Google Scholar] [CrossRef] [PubMed]
- Frank, D.N.; Amand, A.L.S.; Feldman, R.A.; Boedeker, E.C.; Harpaz, N.; Pace, N.R. Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc. Natl. Acad. Sci. USA 2007, 104, 13780–13785. [Google Scholar] [CrossRef]
- Shen, X.J.; Rawls, J.F.; Randall, T.A.; Burcall, L.; Mpande, C.; Jenkins, N.; Jovov, B.; Abdo, Z.; Sandler, R.S.; Keku, T.O. Molecular characterization of mucosal adherent bacteria and associations with colorectal adenomas. Gut Microbes 2010, 1, 138–147. [Google Scholar] [CrossRef]
- Wang, Y.; Antonopoulos, D.; Zhu, X.; Harrell, L.; Hanan, I.; Alverdy, J.; Meyer, F.; Musch, M.W.; Young, V.B.; Chang, E.B. Laser capture microdissection and metagenomic analysis of intact mucosa-associated microbial communities of human colon. Appl. Environ. Microbiol. 2010, 88, 1333–1342. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Willing, B.P.; Dicksved, J.; Halfvarson, J.; Andersson, A.F.; Lucio, M.; Zheng, Z.; Järnerot, G.; Tysk, C.; Jansson, J.K.; Engstrand, L. A pyrosequencing study in twins shows that gastrointestinal microbial profiles vary with inflammatory bowel disease phenotypes. Gastroenterology 2010, 139, 1844–1854. [Google Scholar] [CrossRef]
- Hong, P.-Y.; Croix, J.A.; Greenberg, E.; Gaskins, H.R.; Mackie, R.I. Pyrosequencing-based analysis of the mucosal microbiota in healthy individuals reveals ubiquitous bacterial groups and micro-heterogeneity. PLoS ONE 2011, 6, e25042. [Google Scholar] [CrossRef]
- Gibson, G.R.; Hutkins, R.; Sanders, M.E.; Prescott, S.L.; Reimer, R.A.; Salminen, S.J.; Scott, K.; Stanton, C.; Swanson, K.S.; Cani, P.D.; et al. Expert consensus document: The International Scientific Association for Probiotics and Prebiotics (ISAPP) consensus statement on the definition and scope of prebiotics. Nat. Rev. Gastroenterol. Hepatol. 2017, 14, 491–502. [Google Scholar] [CrossRef]
- Minekus, M.; Alminger, M.; Alvito, P.; Balance, S.; Bohn, T.; Bourlieu, C.; Carrière, F.; Boutrou, R.; Corredig, M.; Dupont, D.; et al. A standardised static in vitro digestion method suitable for food—An international consensus. Food Funct. 2014, 5, 1113–1124. [Google Scholar] [CrossRef]
- Hodoniczky, J.; Morris, C.A.; Rae, A.L. Oral and intestinal digestion of oligosaccharides as potential sweeteners: A systematic evaluation. J. Food Chem. 2012, 132, 1951–1958. [Google Scholar] [CrossRef]
- Sonnenburg, E.D.; Zheng, H.; Joglekar, P.; Higginbottom, S.K.; Firbank, S.J.; Bolam, D.N.; Sonnenburg, J.L. Specificity of polysaccharide use in intestinal bacteroides species determines diet-induced microbiota alterations. Cell 2010, 141, 1241–1252. [Google Scholar] [CrossRef] [PubMed]
- Hiippala, K.; Kainulainen, V.; Suutarinen, M.; Heini, T.; Bowers, J.R.; Jasso-Selles, D.; Lemmer, D.; Valentine, M.; Barnes, R.; Engelthaler, D.M.; et al. Isolation of Anti-Inflammatory and Epithelium Reinforcing Bacteroides and Parabacteroides Spp. from A Healthy Fecal Donor. Nutrients 2020, 12, E935. [Google Scholar] [CrossRef] [PubMed]
- Coenen, G.J.; Bakx, E.; Verhoef, R.P.; Schols, H.A.; Voragen, A. Identification of the connecting linkage between homo- or xylogalacturonan and rhamnogalacturonan type I. Carbohydrate. Polymers 2007, 70, 224–235. [Google Scholar]
- Wu, D.; Zheng, J.; Mao, G.; Hu, W.; Ye, X.; Linhardt, R.J.; Chen, S. Rethinking the impact of RG-I mainly from fruits and vegetables on dietary health. Crit. Rev. Food Sci. Nutr. 2019, 1–23. [Google Scholar] [CrossRef]
- Chakaroun, R.M.; Massier, L.; Kovacs, P. Gut Microbiome, Intestinal Permeability, and Tissue Bacteria in Metabolic Disease: Perpetrators or Bystanders? Nutrients 2020, 12, E1082. [Google Scholar] [CrossRef]
- Marzorati, M.; Pinheiro, I.; Van den Abbeele, P.; Van de Wiele, T.; Possemiers, S. An in vitro technology platform to assess host microbiota interactions in the gastrointestinal tract. Agro Food Ind. Hi Tech 2013, 23, 8–11. [Google Scholar]
- Satsu, H.; Ishimoto, Y.; Nakano, T.; Mochizuki, T.; Iwanaga, T.; Shimizu, M. Induction by activated macrophage-like THP-1 cells of apoptotic and necrotic cell death in intestinal epithelial Caco-2 monolayers via tumor necrosis factor-alpha. Exp. Cell Res. 2006, 312, 3909–3919. [Google Scholar] [CrossRef]
- Van den Abbeele, P.; Taminiau, B.; Pinheiro, I.; Duysburgh, C.; Jacobs, H.; Pijls, L.; Marzorati, M. Arabinoxylo-Oligosaccharides and Inulin Impact Inter-Individual Variation on Microbial Metabolism and Composition, Which Immunomodulates Human Cells. J. Agric. Food Chem. 2018, 66, 1121–1130. [Google Scholar] [CrossRef] [PubMed]
- Daguet, D.; Pinheiro, I.; Verhelst, A.; Possemiers, S.; Marzorati, M. Arabinogalactan and fructooligosaccharides improve the gut barrier function in distinct areas of the colon in the Simulator of the Human Intestinal Microbial Ecosystem. J. Funct. Foods 2016, 20, 369–379. [Google Scholar] [CrossRef]
- Possemiers, S.; Pinheiro, I.; Verhelst, A.; Van den Abbeele, P.; Maignien, L.; Laukens, D.; Reeves, S.G.; Robinson, L.E.; Raas, T.; Schneider, Y.-J.; et al. A dried yeast fermentate selectively modulates both the luminal and mucosal gut microbiota and protects against inflammation, as studied in an integrated in vitro approach. J. Agric. Food Chem. 2013, 61, 9380–9392. [Google Scholar] [CrossRef] [PubMed]
- Kleiveland, C.R. Peripheral Blood Mononuclear Cells. In The Impact of Food Bioactives on Health: In vitro and Ex Vivo Models; Verhoeckx, K., Cotter, P., López-Expósito, I., Kleiveland, C., Lea, T., Mackie, A., Requena, T., Swiatecka, D., Wichers, H., Eds.; Springer Cham (CH): Cham, Switzerland, 2015; Chapter 15. [Google Scholar]
- Van den Abbeele, P.; Moens, F.; Pignataro, G.; Schnurr, J.; Ribecco, C.; Gramenzi, A.; Marzorati, M. Yeast-Derived Formulations Are Differentially Fermented by the Canine and Feline Microbiome As Assessed in a Novel In Vitro Colonic Fermentation Model. J. Agric. Food Chem. 2020. [Google Scholar] [CrossRef] [PubMed]
- Jonker, D.; Fowler, P.; Albers, R.; Tzoumaki, M.V.; van het Hof, K.H.; Aparicio-Vergara, M. Safety assessment of rhamnogalacturonan-enriched carrot pectin fraction: 90-day oral toxicity study in rats and in vitro genotoxicity studies. Food Chem. Toxicol. 2020, 139, 111243. [Google Scholar] [CrossRef]
- De Weirdt, R.; Possemiers, S.; Vermeulen, G.; Moerdijk-Poortvliet, T.C.W.; Boschker, H.T.S.; Verstraete, W.; Van de Wiele, T. Human faecal microbiota display variable patterns of glycerol metabolism. FEMS Microbiol. Ecol. 2010, 74, 601–611. [Google Scholar] [CrossRef] [PubMed]
- Boon, N.; Top, E.M.; Verstraete, W.; Siciliano, S.D. Bioaugmentation as a tool to protect the structure and function of an activated-sludge microbial community against a 3-chloroaniline shock load. Appl. Environ. Microbiol. 2003, 69, 1511–1520. [Google Scholar] [CrossRef]
- Furet, J.P.; Firmesse, O.; Gourmelon, M.; Bridonneau, C.; Tap, J.; Mondot, S.; Doré, J.; Corthier, G. Comparative assessment of human and farm animal faecal microbiota using real-time quantitative PCR. FEMS Microbiol. Ecol. 2009, 68, 351–362. [Google Scholar] [CrossRef]
- Rinttilä, T.; Kassinen, A.; Malinen, E.; Kroqius, L.; Palva, A. Development of an extensive set of 16S rDNA-targeted primers for quantification of pathogenic and indigenous bacteria in faecal samples by real-time PCR. J. Appl. Microbiol. 2004, 97, 1166–1177. [Google Scholar] [CrossRef]
- Collado, M.C.; Derrien, M.; Isolauri, E.; de Vos, W.M.; Salminen, S. Intestinal integrity and Akkermansia muciniphila, a mucin-degrading member of the intestinal microbiota present in infants, adults, and the elderly. Appl. Environ. Microbiol. 2007, 23, 7767–7770. [Google Scholar] [CrossRef]
- Guo, X.; Xia, X.; Tang, R.; Zhou, J.; Zhao, H.; Wang, K. Development of a real-time PCR method for Firmicutes and Bacteroidetes in faeces and its application to quantify intestinal population of obese and lean pigs. Lett. Appl. Microbiol. 2008, 47, 367–373. [Google Scholar] [CrossRef]
- Nakano, S.; Kobayashi, T.; Funabiki, K.; Matsumura, A.; Nagao, Y.; Yamada, T. Development of a PCR assay for detection of Enterobacteriaceae in foods. J. Food Prot. 2003, 66, 1798–1804. [Google Scholar] [CrossRef]
- Sokol, H.; Seksik, P.; Furet, J.P.; Firmesse, O.; Nion-Larmurier, I.; Beaugerie, L.; Cosnes, J.; Corthier, G.; Marteau, P.; Doré, J. Low counts of Faecalibacterium prausnitzii in colitis microbiota. Inflamm. Bowel Dis. 2009, 15, 1183–1189. [Google Scholar] [CrossRef] [PubMed]
- Ramirez-Farias, C.; Slezak, K.; Fuller, Z.; Duncan, A.; Holtrop, G.; Louis, P. Effect of inulin on the human gut microbiota: Stimulation of Bifidobacterium adolescentis and Faecalibacterium prausnitzii. Br. J. Nutr. 2009, 101, 541–550. [Google Scholar] [CrossRef] [PubMed]
- Props, R.; Kerckhof, F.-M.; Rubbens, P.; De Vrieze, J.; Hernandez Sanabria, E.; Waegeman, W.; Monsieurs, P.; Hammes, F.; Boon, N. Absolute quantification of microbial taxon abundances. ISME J. 2017, 11, 584–587. [Google Scholar] [CrossRef] [PubMed]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glöckner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef] [PubMed]
- Schloss, P.D.; Westcott, S.L. Assessing and improving methods used in operational taxonomic unit-based approaches for 16S rRNA gene sequence analysis. Appl. Environ. Microbiol. 2011, 77, 3219–3226. [Google Scholar] [CrossRef]
- Kozich, J.J.; Westcott, S.L.; Baxter, N.T.; Highlander, S.K.; Schloss, P.D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 2013, 79, 5112–5120. [Google Scholar] [CrossRef]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef]
- Cole, J.R.; Wang, Q.; Cardenas, E.; Fish, J.; Chai, B.; Farris, R.J.; Kulam-Syed-Mohideen, A.S.; McGarrell, D.M.; Marsh, T.; Garrity, G.M.; et al. The ribosomal database project: Improved alignments and new tools for rRNA analysis. Nucleic Acids Res 2009, 37, D141–D145. [Google Scholar] [CrossRef]
- Louis, P.; Flint, H.J. Formation of propionate and butyrate by the human colonic microbiota. Environ. Microbiol. 2017, 19, 29–41. [Google Scholar] [CrossRef]
- Hamer, H.M.; De Preter, V.; Windey, K.; Verbeke, K. Functional analysis of colonic bacterial metabolism: Relevant to health. Am. J. Physiol. Liver Physiol. 2012, 302, G1–G9. [Google Scholar] [CrossRef]
- Scott, K.P.; Gratz, S.W.; Sheridan, P.O.; Flint, H.J.; Duncan, S.H. The influence of diet on the gut microbiota. Pharmacol. Res. 2013, 69, 52–60. [Google Scholar] [CrossRef] [PubMed]
- Kempski, J.; Brockmann, L.; Gagliani, N.; Huber, S. TH17 Cell and Epithelial Cell Crosstalk during Inflammatory Bowel Disease and Carcinogenesis. Front Immunol. 2017, 8, 1373. [Google Scholar] [CrossRef]
- Kurita-Ochiai, T.; Fukushima, K.; Ochiai, K. Lipopolysaccharide stimulates butyric acid-induced apoptosis in human peripheral blood mononuclear cells. Infect. Immun. 1999, 67, 22–29. [Google Scholar] [CrossRef] [PubMed]
- Weber, T.E.; Kerr, B.J. Butyrate differentially regulates cytokines and proliferation in porcine peripheral blood mononuclear cells. Vet. Immunol. Immunopathol. 2006, 113, 139–147. [Google Scholar] [CrossRef] [PubMed]
- Bonnema, A.L.; Kolberg, L.W.; Thomas, W.; Slavin, J.L. Gastrointestinal tolerance of chicory inulin products. J. Am. Diet. Assoc. 2010, 110, 865–868. [Google Scholar] [CrossRef]
- Timm, D.A.; Stewart, M.L.; Hospattankar, A.; Slavin, J.L. Wheat Dextrin, Psyllium, and Inulin Produce Distinct FermentationPatterns, GasVolumes, andShort-Chain Fatty AcidProfiles InVitro. J. Med. Food 2010, 13, 961–996. [Google Scholar] [CrossRef]
- Carlson, J.L.; Erickson, J.M.; Hess, J.M.; Gould, T.J.; Slavin, J.L. Prebiotic Dietary Fiber and Gut Health: Comparing the in Vitro Fermentations of Beta-Glucan, Inulin and Xylooligosaccharide. Nutrients 2017, 9, 1361. [Google Scholar] [CrossRef]
- Belenguer, A.; Duncan, S.H.; Calder, A.G.; Holtrop, G.; Louis, P.; Lobley, G.E.; Flint, H.J. Two Routes of Metabolic Cross-Feeding between Bifidobacterium adolescentis and Butyrate-Producing Anaerobes from the Human Gut. Appl. Environ. Microbiol. 2006, 72, 3593–3599. [Google Scholar] [CrossRef]
- De Vuyst, L.; Moens, F.; Selak, M.; Rivière, A.; Leroy, F. Summer Meeting 2013: Growth and physiology of bifidobacteria. J. Appl. Microbiol. 2014, 116, 477–491. [Google Scholar] [CrossRef]
- Milani, C.; Lugli, G.A.; Duranti, S.; Turroni, F.; Mancabelli, L.; Ferrario, C.; Mangifesta, M.; Hevia, A.; Viappiani, A.; Scholz, M.; et al. Bifidobacteria exhibit social behavior through carbohydrate resource sharing in the gut. Sci. Rep. 2015, 5, 15782. [Google Scholar] [CrossRef]
- Moens, F.; Weckx, S.; De Vuyst, L. Bifidobacterial inulin-type fructan degradation capacity determines cross-feeding interactions between bifidobacteria and Faecalibacterium prausnitzii. Int. J. Food Microbiol. 2016, 231, 76–85. [Google Scholar] [CrossRef] [PubMed]
- Fukuda, S.; Toh, H.; Hase, K.; Oshima, K.; Nakanishi, Y.; Yoshimura, K.; Tobe, T.; Clarke, J.M.; Topping, D.L.; Suzuki, T.; et al. Bifidobacteria can protect from enteropathogenic infection through production of acetate. Nature 2011, 469, 543–547. [Google Scholar] [CrossRef] [PubMed]
- Srutkova, D.; Schwarzer, M.; Hudcovic, T.; Zakostelska, Z.; Drab, V.; Spanova, A.; Rittich, B.; Kozakova, H.; Schabussova, I. Bifidobacterium longum CCM 7952 Promotes Epithelial Barrier Function and Prevents Acute DSS-Induced Colitis in Strictly Strain-Specific Manner. PLoS ONE 2015, 10, e0134050. [Google Scholar] [CrossRef] [PubMed]
- Furrie, E.; Macfarlane, S.; Kennedy, A.; Cummings, J.H.; Walsh, S.V.; O’neil, D.A.; Macfarlane, G.T. Synbiotic therapy (Bifidobacterium longum/Synergy 1) initiates resolution of inflammation in patients with active ulcerative colitis: A randomised controlled pilot trial. Gut 2005, 54, 242–249. [Google Scholar] [CrossRef]
- Yu, R.; Zuo, F.; Ma, H.; Chen, S. Exopolysaccharide-Producing Bifidobacterium adolescentis Strains with Similar Adhesion Property Induce Differential Regulation of Inflammatory Immune Response in Treg/Th17 Axis of DSS-Colitis Mice. Nutrients 2019, 11, E782. [Google Scholar] [CrossRef] [PubMed]
- Hosseini, E.; Grootaert, C.; Verstraete, W.; Van de Wiele, T. Propionate as a health-promoting microbial metabolite in the human gut. Nutr. Rev. 2011, 69, 245–258. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, N.; Emoto, T.; Yamashita, T.; Watanabe, H.; Hayashi, T.; Tabata, T.; Hoshi, N.; Hatano, N.; Ozawa, G.; Sasaki, N.; et al. Bacteroides vulgatus and Bacteroides dorei Reduce Gut Microbial Lipopolysaccharide Production and Inhibit Atherosclerosis. Circulation 2018, 138, 2486–2498. [Google Scholar] [CrossRef]
- Ihekweazu, F.D.; Fofanova, T.Y.; Queliza, K.; Nagy-Szakal, D.; Stewart, C.J.; Engevik, M.A.; Hulten, K.G.; Tatevian, N.; Graham, D.Y.; Versalovic, J.; et al. Bacteroides ovatus ATCC 8483 monotherapy is superior to traditional fecal transplant and multi-strain bacteriotherapy in a murine colitis model. Gut Microbes 2019, 10, 504–520. [Google Scholar] [CrossRef] [PubMed]
- Morotomi, M.; Nagai, F.; Sakon, H.; Tanaka, R. Dialister succinatiphilus sp. nov. and Barnesiella intestinihominis sp. nov., isolated from human faeces. Int. J. Syst. Evol. Microbiol. 2008, 58, 2716–2720. [Google Scholar] [CrossRef]
- Sears, C.L. Enterotoxigenic Bacteroides fragilis: A rogue among symbiotes. Clin. Microbiol. Rev. 2009, 22, 349–366. [Google Scholar] [CrossRef]
- Wei, B.; Dalwadi, H.; Gordon, L.K.; Landers, C.; Bruckner, D.; Targan, S.R.; Braun, J. Molecular cloning of a Bacteroides caccae TonB-linked outer membrane protein identified by an inflammatory bowel disease marker antibody. Infect. Immun. 2001, 69, 6044–6054. [Google Scholar] [CrossRef] [PubMed]
- Patterson, A.M.; Mulder, I.E.; Travis, A.J.; Lan, A.; Cerf-Bensussan, N.; Gaboriau-Routhiau, V.; Garden, K.; Logan, E.; Delday, M.I.; Coutts, A.G.P.; et al. Human Gut Symbiont Roseburia hominis Promotes and Regulates Innate Immunity. Front. Immunol. 2017, 8, 1166. [Google Scholar] [CrossRef]
- Machiels, K.; Joossens, M.; Sabino, J.; De Preter, V.; Arijs, I.; Eeckhaut, V.; Ballet, V.; Claes, K.; Van Immerseel, F.; Verbeke, K.; et al. A decrease of the butyrate-producing species Roseburia hominis and Faecalibacterium prausnitzii defines dysbiosis in patients with ulcerative colitis. Gut 2014, 63, 1275–1283. [Google Scholar] [CrossRef] [PubMed]
- Tilg, H.; Danese, S. Roseburia hominis: A novel guilty player in ulcerative colitis pathogenesis? Gut 2014, 63, 1204–1205. [Google Scholar] [CrossRef] [PubMed]
- Qiu, X.; Zhang, M.; Yang, X.; Hong, N.; Yu, C. Faecalibacterium prausnitzii upregulates regulatory T cells and anti-inflammatory cytokines in treating TNBS-induced colitis. J. Crohns Colitis 2013, 7, e558–e568. [Google Scholar] [CrossRef] [PubMed]
- Udayappan, S.; Manneras-Holm, L.; Chaplin-Scott, A.; Belzer, C.; Herrema, H.; Dallinga-Thie, G.M.; Duncan, S.H.; Stroes, E.S.G.; Groen, A.K.; Flint, H.J.; et al. Oral treatment with Eubacterium hallii improves insulin sensitivity in db/db mice. NPJ Biofilms Microbiomes 2016, 2, 16009. [Google Scholar] [CrossRef]
- Murai, M.; Turovskaya, O.; Kim, G.; Madan, R.; Karp, C.L.; Cheroutre, H.; Kronenberg, M. Interleukin 10 acts on regulatory T cells to maintain expression of the transcription factor Foxp3 and suppressive function in mice with colitis. Nat. Immunol. 2009, 10, 1178–1184. [Google Scholar] [CrossRef]
- Kumar, P.; Monin, L.; Castillo, P.; Elsegeiny, W.; Horne, W.; Eddens, T.; Vikram, A.; Good, M.; Schoenborn, A.A.; Bibby, K.; et al. Intestinal Interleukin-17 Receptor Signaling Mediates Reciprocal Control of the Gut Microbiota and Autoimmune Inflammation. Immunity. 2016, 44, 659–671. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.L.; Sundrud, M.S. Cytokine Networks and T-Cell Subsets in Inflammatory Bowel Diseases. Inflamm. Bowel Dis. 2016, 22, 1157–1167. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.H. Developing food allergy: A potential immunologic pathway linking skin barrier to gut. F1000Research 2016, 5, F1000. [Google Scholar] [CrossRef] [PubMed]
- Clark, D.P. The fermentation pathways of Escherichia coli. FEMS Microbiol. Rev. 1989, 5, 223–234. [Google Scholar] [CrossRef]
- Rey, F.E.; Gonzalez, M.D.; Cheng, J.; Wu, M.; Ahern, P.P.; Gordon, J.I. Metabolic niche of a prominent sulfate-reducing human gut bacterium. Proc. Natl. Acad. Sci. USA 2013, 110, 13582–13587. [Google Scholar] [CrossRef] [PubMed]
| Absolute Abundance (16S Gene Copies/mL) | Proportional Abundance (%) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Phylum | Family | Inoculum | L | M | |||||||
| lumen | lumen | mucus | |||||||||
| blank | cRG-I | cRG-I+LMWC | blank | cRG-I | cRG-I+LMWC | blank | cRG-I | cRG-I+LMWC | |||
| Actinobacteria | Bifidobacteriaceae | 7.28 | 7.18 | 8.68 | 8.28 | 6.89 | 8.63 | 8.26 | 0.2% | 6.4% | 1.2% |
| Coriobacteriaceae | 8.22 | 6.60 | 6.39 | 6.88 | 6.25 | 6.36 | 6.80 | 0.0% | 0.0% | 0.0% | |
| Eggerthellaceae | 6.56 | 7.72 | 7.40 | 7.22 | 7.49 | 7.29 | 7.09 | 0.1% | 0.0% | 0.0% | |
| Bacteroidetes | Bacteroidaceae | 7.94 | 8.63 | 9.60 | 9.50 | 8.52 | 9.65 | 9.45 | 20.1% | 9.1% | 8.5% |
| Marinifilaceae | 6.23 | 6.12 | 6.36 | 6.04 | 6.46 | 6.43 | 6.25 | 0.1% | 0.0% | 0.0% | |
| Muribaculaceae | 6.10 | 7.02 | 6.73 | 6.15 | 6.78 | 6.56 | 6.03 | 0.1% | 0.0% | 0.0% | |
| Prevotellaceae | 6.74 | <LOQ | 7.04 | 6.04 | 5.48 | 6.55 | 6.15 | 0.0% | 0.0% | 0.0% | |
| Rikenellaceae | 7.94 | 6.85 | 7.02 | 6.60 | 6.57 | 6.83 | 6.68 | 0.1% | 0.0% | 0.1% | |
| Tannerellaceae | 6.19 | 7.70 | 7.35 | 7.05 | 7.58 | 7.42 | 7.24 | 0.1% | 0.0% | 0.0% | |
| Firmicutes | Christensenellaceae | 7.58 | 7.06 | 7.14 | 6.66 | 6.82 | 6.94 | 6.75 | 0.0% | 0.0% | 0.0% |
| Clostridiaceae cluster I | 6.15 | 5.96 | <LOQ | 5.95 | 6.40 | 5.94 | 6.17 | 41.7% | 2.1% | 1.8% | |
| Erysipelotrichaceae | 7.41 | 6.93 | 7.32 | 7.33 | 7.07 | 7.55 | 7.81 | 1.1% | 1.0% | 1.4% | |
| Clostridiaceae cluster XI | <LOQ | 6.27 | 6.58 | 6.39 | 6.36 | 6.48 | 6.69 | 0.0% | 0.0% | 0.0% | |
| Clostridiaceae cluster XIII | 6.43 | 6.45 | 6.14 | <LOQ | 6.73 | 6.16 | 6.09 | 0.0% | 0.0% | 0.0% | |
| Lachnospiraceae | 8.59 | 8.65 | 8.93 | 8.57 | 8.51 | 8.73 | 8.58 | 28.3% | 74.0% | 79.1% | |
| Peptococcaceae | 6.53 | 6.36 | 6.07 | 5.85 | 6.16 | <LOQ | 5.69 | 0.0% | 0.0% | 0.0% | |
| Peptostreptococcaceae | 6.45 | 6.15 | 6.70 | 6.43 | 5.95 | 7.16 | 7.30 | 0.0% | 0.0% | 0.0% | |
| Ruminococcaceae | 8.64 | 8.38 | 8.83 | 8.30 | 8.34 | 8.72 | 8.49 | 4.1% | 3.0% | 1.5% | |
| Streptococcaceae | 6.37 | <LOQ | 6.70 | 8.14 | 5.26 | 5.80 | 8.17 | 0.1% | 1.0% | 0.5% | |
| Veillonellaceae | 7.77 | 7.67 | 8.45 | 8.01 | 7.51 | 8.31 | 8.16 | 0.2% | 0.2% | 0.2% | |
| Proteobacteria | Burkholderiaceae | 5.18 | 6.78 | 6.83 | 6.32 | 6.74 | 6.60 | 6.41 | 0.0% | 0.0% | 0.0% |
| Desulfovibrionaceae | 6.19 | 6.26 | 6.73 | 6.75 | 6.15 | 6.80 | 6.93 | 0.0% | 0.0% | 0.0% | |
| Enterobacteriaceae | 6.03 | 9.10 | 9.32 | 9.35 | 8.90 | 9.29 | 9.45 | 2.8% | 2.8% | 5.1% | |
| Verrucomicrobia | Akkermansiaceae | 7.63 | 7.91 | 7.55 | 7.71 | 8.46 | 8.67 | 8.64 | 0.6% | 0.2% | 0.4% |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Van den Abbeele, P.; Verstrepen, L.; Ghyselinck, J.; Albers, R.; Marzorati, M.; Mercenier, A. A Novel Non-Digestible, Carrot-Derived Polysaccharide (cRG-I) Selectively Modulates the Human Gut Microbiota while Promoting Gut Barrier Integrity: An Integrated In Vitro Approach. Nutrients 2020, 12, 1917. https://doi.org/10.3390/nu12071917
Van den Abbeele P, Verstrepen L, Ghyselinck J, Albers R, Marzorati M, Mercenier A. A Novel Non-Digestible, Carrot-Derived Polysaccharide (cRG-I) Selectively Modulates the Human Gut Microbiota while Promoting Gut Barrier Integrity: An Integrated In Vitro Approach. Nutrients. 2020; 12(7):1917. https://doi.org/10.3390/nu12071917
Chicago/Turabian StyleVan den Abbeele, Pieter, Lynn Verstrepen, Jonas Ghyselinck, Ruud Albers, Massimo Marzorati, and Annick Mercenier. 2020. "A Novel Non-Digestible, Carrot-Derived Polysaccharide (cRG-I) Selectively Modulates the Human Gut Microbiota while Promoting Gut Barrier Integrity: An Integrated In Vitro Approach" Nutrients 12, no. 7: 1917. https://doi.org/10.3390/nu12071917
APA StyleVan den Abbeele, P., Verstrepen, L., Ghyselinck, J., Albers, R., Marzorati, M., & Mercenier, A. (2020). A Novel Non-Digestible, Carrot-Derived Polysaccharide (cRG-I) Selectively Modulates the Human Gut Microbiota while Promoting Gut Barrier Integrity: An Integrated In Vitro Approach. Nutrients, 12(7), 1917. https://doi.org/10.3390/nu12071917




