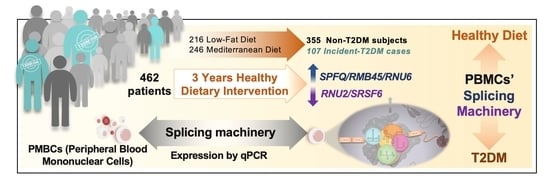
Dietary Intervention Modulates the Expression of Splicing Machinery in Cardiovascular Patients at High Risk of Type 2 Diabetes Development: From the CORDIOPREV Study
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Population
2.2. Study Diets
2.3. Metabolic Study Design
2.4. Blood Sampling and Processing to Isolate Peripheral Blood Mononuclear Cells (PBMCs)
2.5. RNA Extraction and Quantification
2.6. Analysis of Splicing Machinery Components by Microfluidic-Based Dynamic Quantitative Polymerase Chain Reaction (qPCR) Array
2.7. Statistical and Bioinformatical Analysis
3. Results
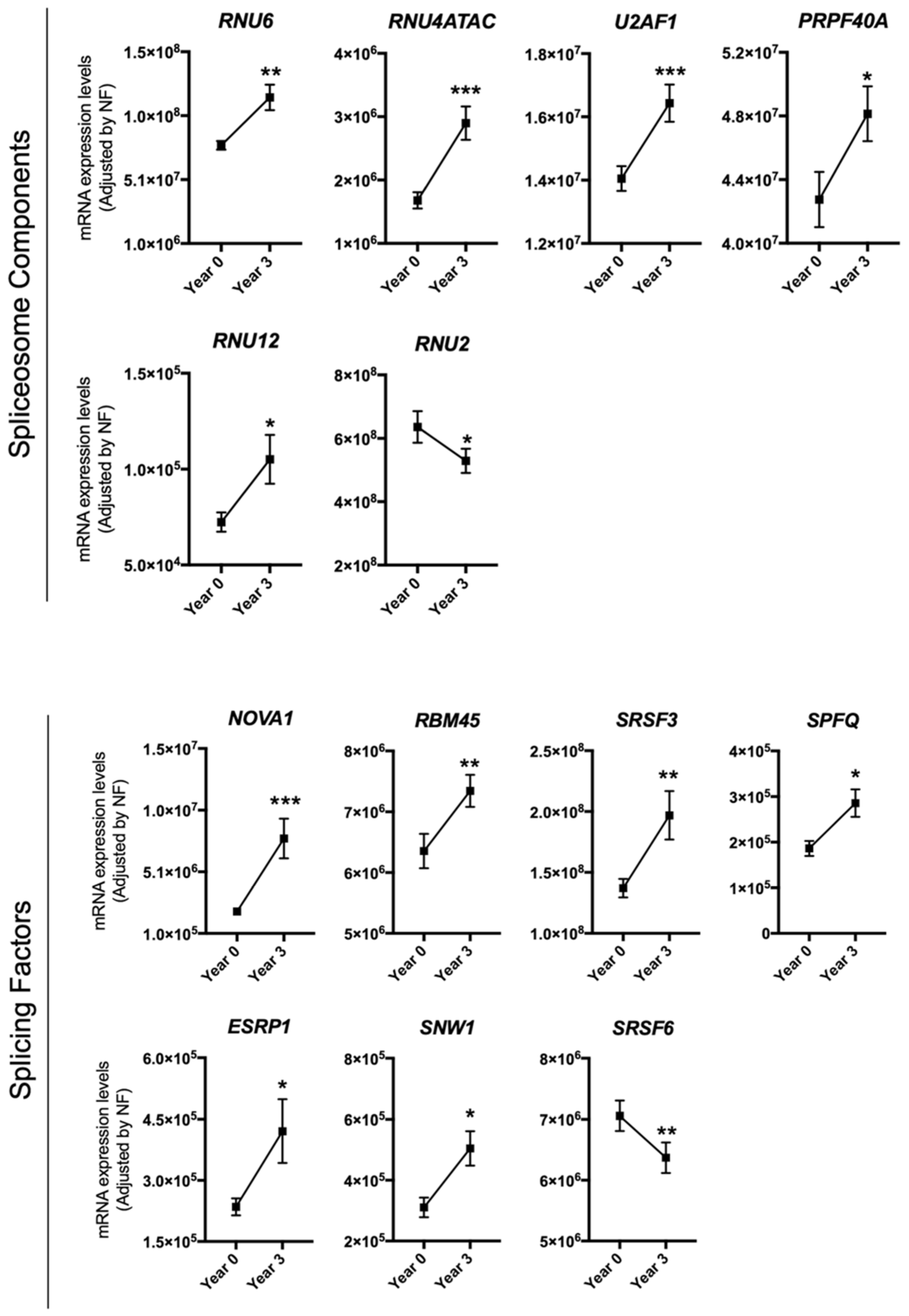
3.1. Dietary Intervention Modulated the Expression of Several Splicing Machinery Components
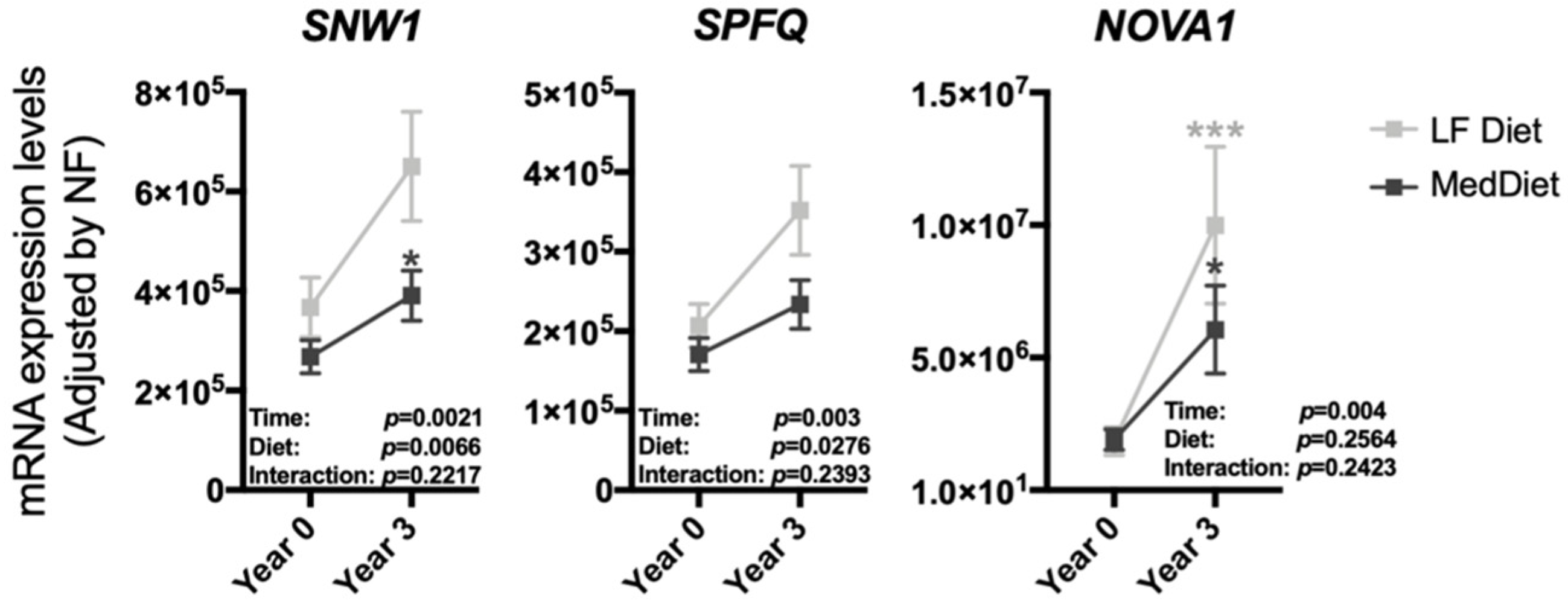
3.2. The Modulation of the Expression of Most Splicing Machinery Components Was Not Diet-Dependent
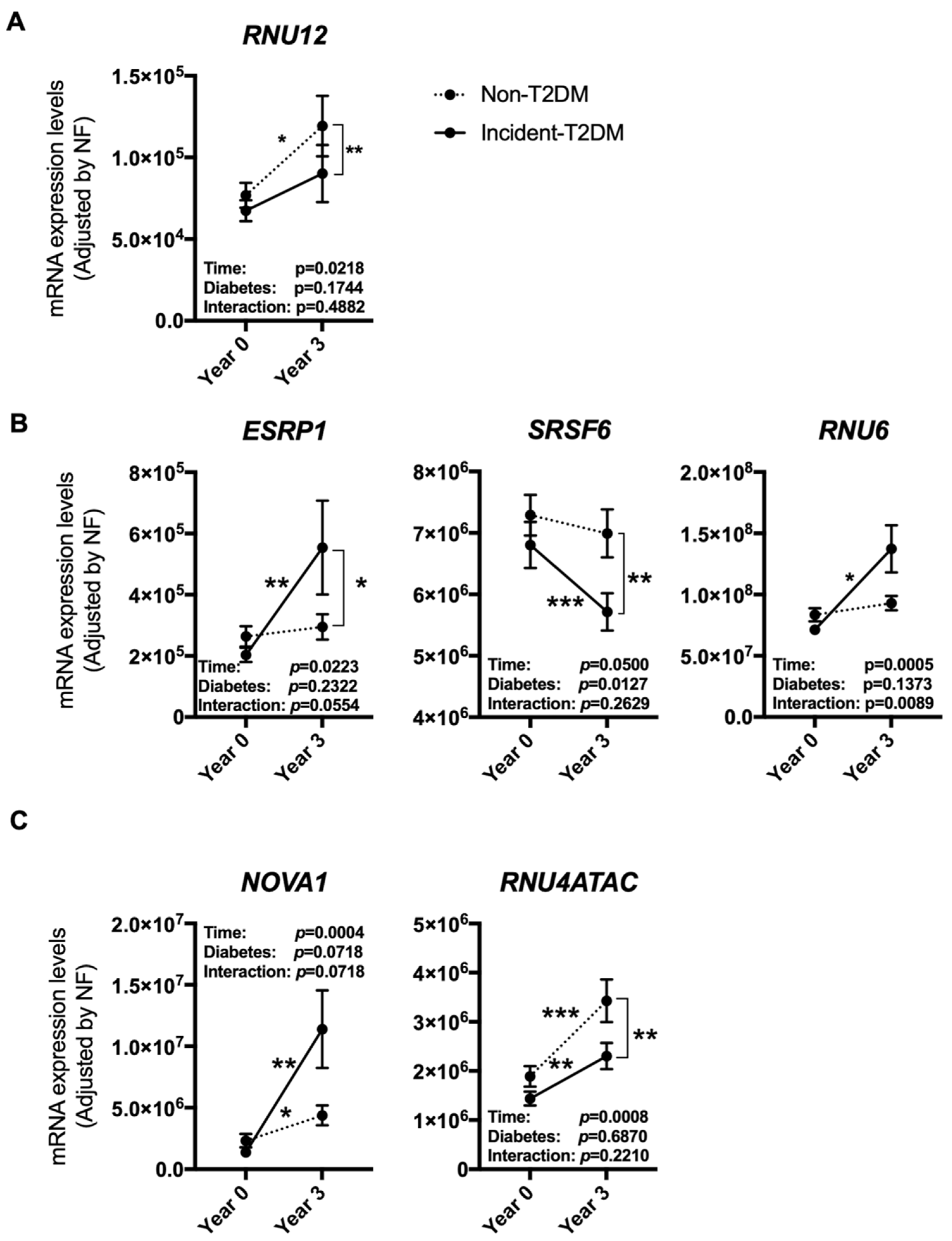
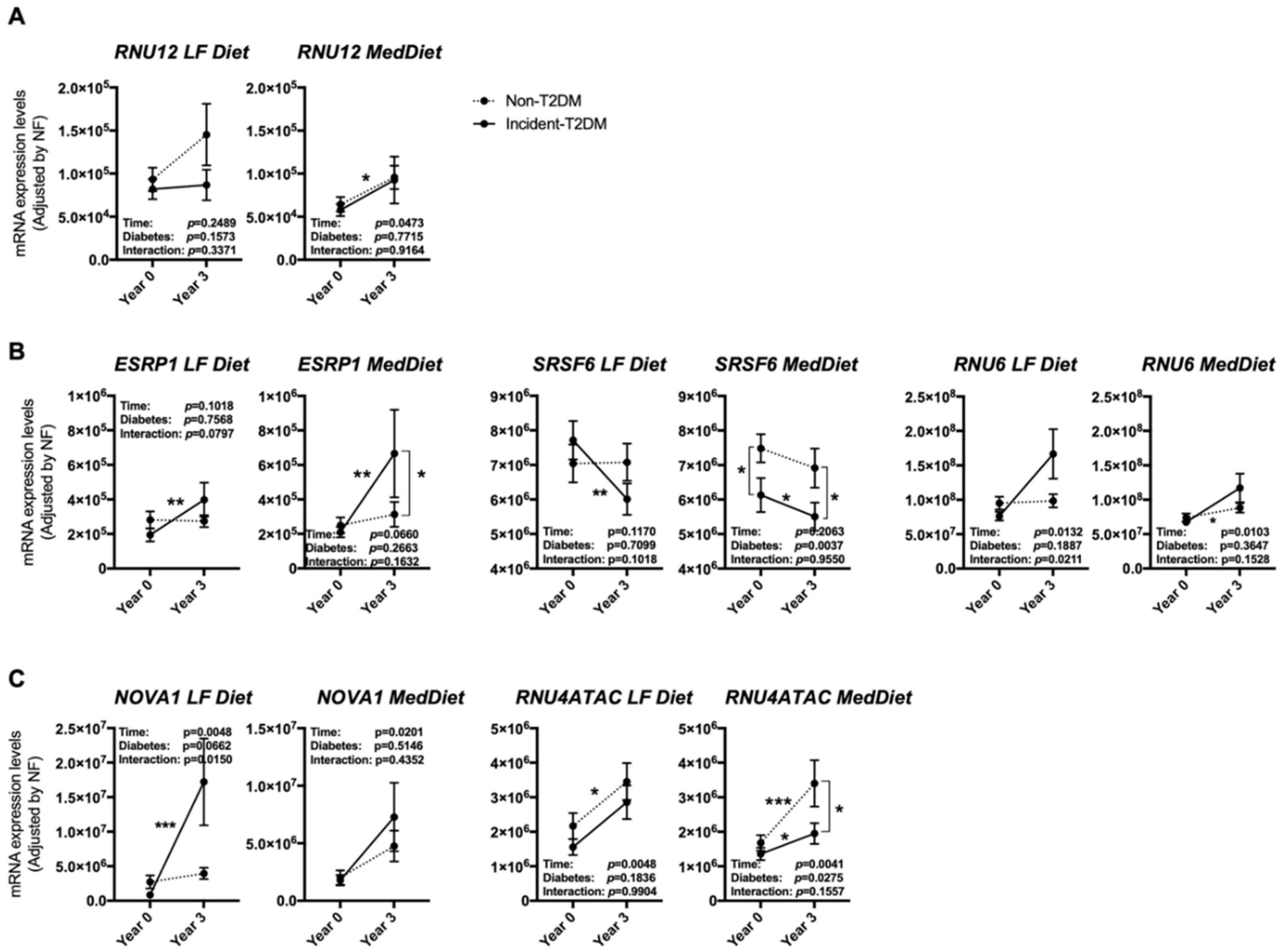
3.3. The Expression Pattern of Specific Splicing Machinery Components Was Differentially Modulated by Dietary Intervention in Incident-T2DM Cases and Non-T2DM Controls
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- World Health Organization. Global Status Report on Noncommunicable Diseases 2014; World Health Organization: Geneva, Switzerland, 2014. [Google Scholar]
- American Diabetes Association. 2. Classification and diagnosis of diabetes: Standards of medical care in diabetes-2019. Diabetes Care 2019, 42, S13–S28. [Google Scholar] [CrossRef]
- Garcia-Rios, A.; Ordovas, J.M.; Lopez-Miranda, J.; Perez-Martinez, P. New diet trials and cardiovascular risk. Curr. Opin. Cardiol. 2018, 33, 423–428. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Jiménez, F.; Ordovás, J.M.; García-Rios, A.; Delgado-Lista, J.; Delgado-Casado, N.; Cruz-Teno, C.; Camargo, A.; Yubero-Serrano, E.M.; Rodriguez, F.; Perezjimenez, F. Consumption of diets with different type of fat influences triacylglycerols-rich lipoproteins particle number and size during the postprandial state. Nutr. Metab. Cardiovasc. Dis. 2011, 21, 39–45. [Google Scholar] [CrossRef] [PubMed]
- Salas-Salvado, J.; Martinez-Gonzalez, M.A.; Bullo, M.; Ros, E. The role of diet in the prevention of type 2 diabetes. Nutr. Metab. Cardiovasc. Dis. 2011, 21 (Suppl. 2), B32–B48. [Google Scholar] [CrossRef] [PubMed]
- Delgado-Lista, J.; Perez-Martinez, P.; Garcia-Rios, A.; Díaz, J.F.A.; Perez-Caballero, A.I.; Delgado-Lista, J.; Fuentes, F.; Quintana-Navarro, G.; Lopez-Segura, F.; Ortiz-Morales, A.M.; et al. CORonary Diet Intervention with Olive oil and cardiovascular PREVention study (the CORDIOPREV study): Rationale, methods, and baseline characteristics: A clinical trial comparing the efficacy of a Mediterranean diet rich in olive oil versus a low-fat diet on cardiovascular disease in coronary patients. Am. Heart J. 2016, 177, 42–50. [Google Scholar] [PubMed]
- Blanco-Rojo, R.; Alcala-Diaz, J.F.; Wopereis, S.; Perez-Martinez, P.; Quintana-Navarro, G.M.; Marin, C.; Ordovas, J.M.; Van Ommen, B.; Perez-Jimenez, F.; Delgado-Lista, J.; et al. The insulin resistance phenotype (muscle or liver) interacts with the type of diet to determine changes in disposition index after 2 years of intervention: The CORDIOPREV-DIAB randomised clinical trial. Diabetologia 2015, 59, 67–76. [Google Scholar] [CrossRef]
- Camargo, A.; Jimenez-Lucena, R.; Alcala-Diaz, J.F.; Rangel-Zuñiga, O.A.; Garcia-Carpintero, S.; Lopez-Moreno, J.; Blanco-Rojo, R.; Delgado-Lista, J.; Perez-Martinez, P.; Van Ommen, B.; et al. Postprandial endotoxemia may influence the development of type 2 diabetes mellitus: From the CORDIOPREV study. Clin. Nutr. 2018, 38, 529–538. [Google Scholar] [CrossRef]
- Abbasi, A.; Peelen, L.M.; Corpeleijn, E.; van der Schouw, Y.T.; Stolk, R.P.; Spijkerman, A.M.; Moons, K.G.; Navis, G.; Bakker, S.J.; Beulens, J.W. Prediction models for risk of developing type 2 diabetes: Systematic literature search and independent external validation study. BMJ 2012, 345, e5900. [Google Scholar] [CrossRef]
- Gahete, M.D.; del Rio-Moreno, M.; Camargo, A.; Alcala-Diaz, J.F.; Alors-Perez, E.; Delgado-Lista, J.; Reyes, O.; Ventura, S.; Perez-Martínez, P.; Castaño, J.P.; et al. Changes in splicing machinery components influence, precede, and early predict the development of type 2 diabetes: From the CORDIOPREV study. EBioMedicine 2018, 37, 356–365. [Google Scholar] [CrossRef]
- Burczynski, M.E.; Dorner, A.J. Transcriptional profiling of peripheral blood cells in clinical pharmacogenomic studies. Pharmacogenomics 2006, 7, 187–202. [Google Scholar] [CrossRef]
- Cohen, R.M.; Haggerty, S.; Herman, W.H. HbA1c for the diagnosis of diabetes and prediabetes: Is it time for a mid-course correction? J. Clin. Endocrinol. Metab. 2010, 95, 5203–5206. [Google Scholar] [CrossRef] [PubMed]
- Matera, A.G.; Wang, Z. A day in the life of the spliceosome. Nat. Rev. Mol. Cell Biol. 2014, 15, 108–121. [Google Scholar] [CrossRef] [PubMed]
- Scotti, M.M.; Swanson, M.S. RNA mis-splicing in disease. Nat. Rev. Genet. 2016, 17, 19–32. [Google Scholar] [CrossRef] [PubMed]
- Dlamini, Z.; Mokoena, F.; Hull, R. Abnormalities in alternative splicing in diabetes: Therapeutic targets. J. Mol. Endocrinol. 2017, 59, R93–R107. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.C.; Abdel-Wahab, O. Therapeutic targeting of splicing in cancer. Nat. Med. 2016, 22, 976–986. [Google Scholar] [CrossRef]
- Mercader, J.M.; Liao, R.G.; Bell, A.D.; Dymek, Z.; Estrada, K.; Tukiainen, T.; Huerta-Chagoya, A.; Moreno-Macías, H.; Jablonski, K.A.; Hanson, R.L.; et al. A loss-of-function splice acceptor variant in IGF2 is protective for type 2 diabetes. Diabetes 2017, 66, 2903–2914. [Google Scholar] [CrossRef]
- Webster, N.J.G. Alternative RNA splicing in the pathogenesis of liver disease. Front. Endocrinol. (Lausanne) 2017, 8, 133. [Google Scholar] [CrossRef]
- Gallego-Paez, L.M.; Bordone, M.C.; Leote, A.C.; Saraiva-Agostinho, N.; Ascensao-Ferreira, M.; Barbosa-Morais, N.L. Alternative splicing: The pledge, the turn, and the prestige: The key role of alternative splicing in human biological systems. Hum. Genet. 2017, 136, 1015–1042. [Google Scholar] [CrossRef]
- Vázquez-Borrego, M.C.; Fuentes-Fayos, A.C.; Venegas-Moreno, E.; Rivero-Cortés, E.; Dios, E.; Moreno-Moreno, P.; Madrazo-Atutxa, A.; Remón, P.; Solivera, J.; Wildemberg, L.E.; et al. Splicing machinery is dysregulated in pituitary neuroendocrine tumors and is associated with aggressiveness features. Cancers 2019, 11, 1439. [Google Scholar]
- Jiménez-Vacas, J.M.; Herrero-Aguayo, V.; Gómez-Gómez, E.; León-González, A.J.; Sáez-Martínez, P.; Alors-Pérez, E.; Fuentes-Fayos, A.C.; Martínez-López, A.; Sánchez-Sánchez, R.; González-Serrano, T.; et al. Spliceosome component SF3B1 as novel prognostic biomarker and therapeutic target for prostate cancer. Transl. Res. 2019, 212, 89–103. [Google Scholar]
- Stumvoll, M.; Goldstein, B.J.; van Haeften, T.W. Type 2 diabetes: Pathogenesis and treatment. Lancet 2008, 371, 2153–2156. [Google Scholar] [CrossRef]
- Juan-Mateu, J.; Villate, O.; Eizirik, D.L. Mechanisms in endocrinology: Alternative splicing: The new frontier in diabetes research. Eur. J. Endocrinol. 2016, 174, R225–R238. [Google Scholar] [CrossRef] [PubMed]
- Ravi, S.; Schilder, R.J.; Kimball, S.R. Role of precursor mRNA splicing in nutrient-induced alterations in gene expression and metabolism. J. Nutr. 2015, 145, 841–846. [Google Scholar] [CrossRef] [PubMed]
- Yubero-Serrano, E.M.; Delgado-Casado, N.; Gomez-Delgado, F.; Perez-Martinez, P.; Tasset-Cuevas, I.; Santos-Gonzalez, M.; Caballero-Villarraso, J.; Garcia-Rios, A.; Marín, C.; Gutierrez-Mariscal, F.M.; et al. Postprandial antioxidant effect of the Mediterranean diet supplemented with coenzyme Q10 in elderly men and women. Age 2011, 33, 579–590. [Google Scholar] [CrossRef] [PubMed]
- Yubero-Serrano, E.M.; Gonzalez-Guardia, L.; Rangelzuniga, O.A.; Delgado-Casado, N.; Delgado-Lista, J.; Perez-Martinez, P.; Garcia-Rios, A.; Caballero, J.; Marin, C.; Gutierrez-Mariscal, F.M.; et al. Postprandial antioxidant gene expression is modified by Mediterranean diet supplemented with coenzyme Q(10) in elderly men and women. Age 2013, 35, 159–170. [Google Scholar] [CrossRef]
- de Mello, V.D.; Kolehmanien, M.; Schwab, U.; Pulkkinen, L.; Uusitupa, M. Gene expression of peripheral blood mononuclear cells as a tool in dietary intervention studies: What do we know so far? Mol. Nutr. Food Res. 2012, 56, 1160–1172. [Google Scholar] [CrossRef]
- Gahete, M.D.; Luque, R.M.; Yubero-Serrano, E.M.; Cruz-Teno, C.; Ibañez-Costa, A.; Delgado-Lista, J.; Gracia-Navarro, F.; Perez-Jimenez, F.; Castaño, J.P.; Lopez-Miranda, J. Dietary fat alters the expression of cortistatin and ghrelin systems in the PBMCs of elderly subjects: Putative implications in the postprandial inflammatory response. Mol. Nutr. Food Res. 2014, 58, 1897–1906. [Google Scholar] [CrossRef]
- Fuchs, D.; Piller, R.; Linseisen, J.; Daniel, H.; Wenzel, U. The human peripheral blood mononuclear cell proteome responds to a dietary flaxseed-intervention and proteins identified suggest a protective effect in atherosclerosis. Proteomics 2007, 7, 3278–3288. [Google Scholar] [CrossRef]
- Rendo-Urteaga, T.; García-Calzón, S.; González-Muniesa, P.; Milagro, F.I.; Chueca, M.; Oyarzabal, M.; Azcona-Sanjulian, M.C.; Martínez, J.A.; Marti, A. Peripheral blood mononuclear cell gene expression profile in obese boys who followed a moderate energy-restricted diet: Differences between high and low responders at baseline and after the intervention. Br. J. Nutr. 2015, 113, 331–342. [Google Scholar] [CrossRef]
- Quintana-Navarro, G.M.; Alcala-Diaz, J.F.; Lopez-Moreno, J.; Perez-Corral, I.; Leon-Acuña, A.; Torres-Peña, J.D.; Rangel-Zuñiga, O.A.; De Larriva, A.P.A.; Corina, A.; Camargo, A.; et al. Long-term dietary adherence and changes in dietary intake in coronary patients after intervention with a Mediterranean diet or a low-fat diet: The CORDIOPREV randomized trial. Eur. J. Nutr. 2019, 59, 2099–2110. [Google Scholar] [CrossRef]
- Roncero-Ramos, I.; Jimenez-Lucena, R.; Alcala-Diaz, J.F.; Vals-Delgado, C.; Arenas-Larriva, A.P.; Rangel-Zúñiga, O.A.; Leon-Acuña, A.; Malagon, M.M.; Delgado-Lista, J.; Perez-Martinez, P.; et al. Alpha cell function interacts with diet to modulate prediabetes and Type 2 diabetes. J. Nutr. Biochem. 2018, 62, 247–256. [Google Scholar] [CrossRef] [PubMed]
- del Río-Moreno, M.; Alors-Pérez, E.; González-Rubio, S.; Ferrín, G.; Reyes, O.; Rodríguez-Perálvarez, M.; Sánchez-Frías, M.E.; Sánchez-Sánchez, R.; Ventura, S.; López-Miranda, J.; et al. Dysregulation of the splicing machinery is associated to the development of non-alcoholic fatty liver disease. J. Clin. Endocrinol. Metab. 2019, 104, 3389–3402. [Google Scholar]
- Vernia, S.; Edwards, Y.J.; Han, M.S.; Cavanagh-Kyros, J.; Barrett, T.; Kim, J.K.; Davis, R.J. An alternative splicing program promotes adipose tissue thermogenesis. Elife 2016, 5, e17672. [Google Scholar] [CrossRef] [PubMed]
- Kaminska, D.; Hämäläinen, M.; Cederberg, H.; Käkelä, P.; Venesmaa, S.; Miettinen, P.; Ilves, I.; Herzig, K.-H.; Kolehmainen, M.; Karhunen, L.; et al. Adipose tissue INSR splicing in humans associates with fasting insulin level and is regulated by weight loss. Diabetologia 2014, 57, 347–351. [Google Scholar] [CrossRef]
- Villate, O.; Turatsinze, J.-V.; Mascali, L.G.; Grieco, F.A.; Nogueira, T.C.; Cunha, D.A.; Nardelli, T.R.; Sammeth, M.; Salunkhe, V.A.; Esguerra, J.L.S.; et al. Nova1 is a master regulator of alternative splicing in pancreatic beta cells. Nucleic Acids Res. 2014, 42, 11818–11830. [Google Scholar] [CrossRef]
- Reardon, H.T.; Hsieh, A.T.; Park, W.J.; Kothapalli, K.S.; Anthony, J.C.; Nathanielsz, P.W.; Brenna, J.T. Dietary long-chain polyunsaturated fatty acids upregulate expression of FADS3 transcripts. Prostaglandins Leukot. Essent. Fat. Acids 2013, 88, 15–19. [Google Scholar] [CrossRef]
- Walsh, C.M.; Suchanek, A.L.; Cyphert, T.J.; Kohan, A.B.; Szeszel-Fedorowicz, W.; Salati, L.M. Serine arginine splicing factor 3 is involved in enhanced splicing of glucose-6-phosphate dehydrogenase RNA in response to nutrients and hormones in liver. J. Biol. Chem. 2013, 288, 2816–2828. [Google Scholar] [CrossRef]
- Malakar, P.; Chartarifsky, L.; Hija, A.; Leibowitz, G.; Glaser, B.; Dor, Y.; Karni, R. Insulin receptor alternative splicing is regulated by insulin signaling and modulates beta cell survival. Sci. Rep. 2016, 6, 31222. [Google Scholar] [CrossRef]
- Malatesta, M.; Bertoni-Freddari, C.; Fattoretti, P.; Baldelli, B.; Fakan, S.; Gazzanelli, G. Aging and vitamin E deficiency are responsible for altered RNA pathways. Ann. N. Y. Acad. Sci. 2004, 1019, 379–382. [Google Scholar] [CrossRef]
- Verma, B.; Akinyi, M.V.; Norppa, A.J.; Frilander, M.J. Minor spliceosome and disease. Semin. Cell Dev. Biol. 2018, 79, 103–112. [Google Scholar] [CrossRef]
| Fold Change during Follow-Up | ||||
|---|---|---|---|---|
| Study Population (n = 215) Incident-T2DM + Non-T2DM Controls | Incident-T2DM (n = 107) | |||
| SPFQ | SRSF6 | |||
| Fold change during follow-up | HOMA-IR | ρ (rho) | −0.157 | |
| p | 0.03 * | |||
| HIRI | ρ (rho) | −0.176 | ||
| p | 0.018 * | |||
| HbA1c (%) | ρ (rho) | 0.227 | ||
| p | 0.047 * | |||
| Levels at Year 3 (n = 215) Incident-T2DM + Non-T2DM Controls | |||
|---|---|---|---|
| RBM45 | |||
| Levels at year 3 | HOMA-IR | ρ (rho) | −0.166 |
| p | 0.018 * | ||
| HIRI | ρ (rho) | −0.166 | |
| p | 0.018 * |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
del Río-Moreno, M.; Luque, R.M.; Rangel-Zúñiga, O.A.; Alors-Pérez, E.; Alcalá-Diaz, J.F.; Roncero-Ramos, I.; Camargo, A.; Gahete, M.D.; López-Miranda, J.; Castaño, J.P. Dietary Intervention Modulates the Expression of Splicing Machinery in Cardiovascular Patients at High Risk of Type 2 Diabetes Development: From the CORDIOPREV Study. Nutrients 2020, 12, 3528. https://doi.org/10.3390/nu12113528
del Río-Moreno M, Luque RM, Rangel-Zúñiga OA, Alors-Pérez E, Alcalá-Diaz JF, Roncero-Ramos I, Camargo A, Gahete MD, López-Miranda J, Castaño JP. Dietary Intervention Modulates the Expression of Splicing Machinery in Cardiovascular Patients at High Risk of Type 2 Diabetes Development: From the CORDIOPREV Study. Nutrients. 2020; 12(11):3528. https://doi.org/10.3390/nu12113528
Chicago/Turabian Styledel Río-Moreno, Mercedes, Raúl M. Luque, Oriol A. Rangel-Zúñiga, Emilia Alors-Pérez, Juan F. Alcalá-Diaz, Irene Roncero-Ramos, Antonio Camargo, Manuel D. Gahete, José López-Miranda, and Justo P. Castaño. 2020. "Dietary Intervention Modulates the Expression of Splicing Machinery in Cardiovascular Patients at High Risk of Type 2 Diabetes Development: From the CORDIOPREV Study" Nutrients 12, no. 11: 3528. https://doi.org/10.3390/nu12113528
APA Styledel Río-Moreno, M., Luque, R. M., Rangel-Zúñiga, O. A., Alors-Pérez, E., Alcalá-Diaz, J. F., Roncero-Ramos, I., Camargo, A., Gahete, M. D., López-Miranda, J., & Castaño, J. P. (2020). Dietary Intervention Modulates the Expression of Splicing Machinery in Cardiovascular Patients at High Risk of Type 2 Diabetes Development: From the CORDIOPREV Study. Nutrients, 12(11), 3528. https://doi.org/10.3390/nu12113528