Role of Probiotics in Stimulating the Immune System in Viral Respiratory Tract Infections: A Narrative Review
Abstract
1. Introduction
2. Etiology and Epidemiology of Viral Respiratory Tract Infections
3. Immune Responses against Respiratory Viruses
4. Probiotics and Immune Modulation in Viral Respiratory Infections
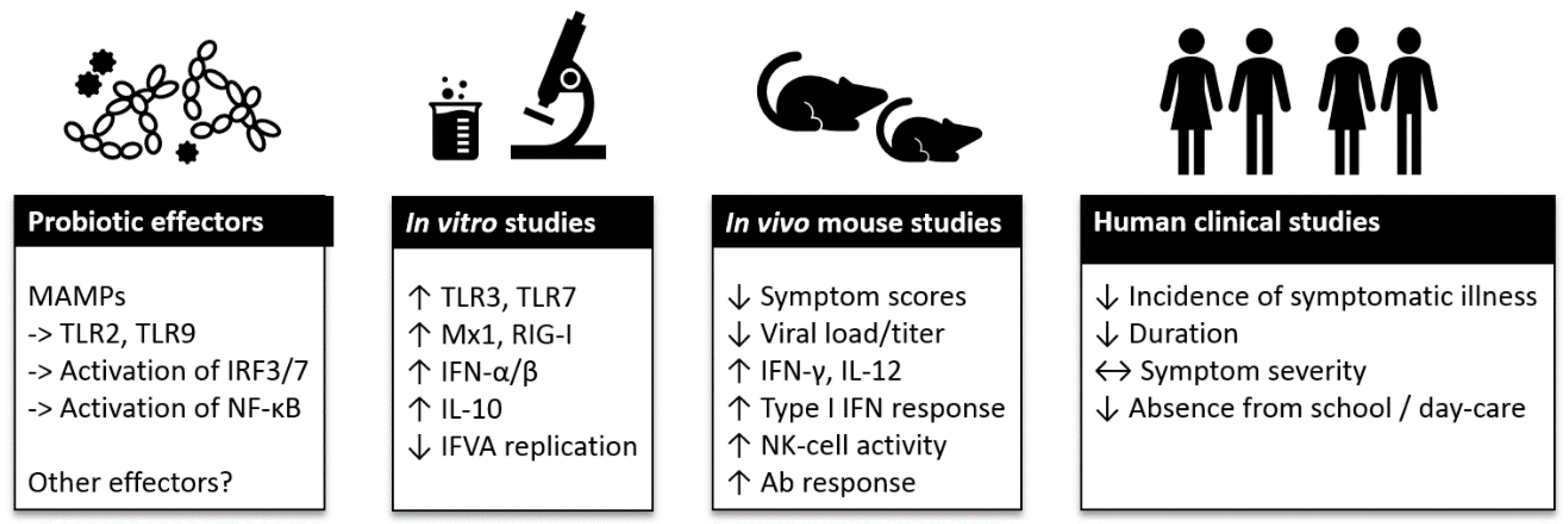
4.1. Overview of Probiotics and Immunomodulatory Mechanisms
4.2. Probiotic Immunostimulation and Inhibition of Viral Replication In Vitro
4.3. Antiviral Effects of Probiotics in Animal Studies
5. Clinical Evidence
6. Discussion and Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Heikkinen, T.; Järvinen, A. The common cold. Lancet 2003, 361, 51–59. [Google Scholar] [CrossRef]
- Jain, S. Epidemiology of Viral Pneumonia. Clin. Chest Med. 2017, 38, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.T.; Kim, C.M.; Ramakrishnan, V. Microbiome and disease in the upper airway. Curr. Opin. Allergy Clin. Immunol. 2019, 19, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Wypych, T.P.; Wickramasinghe, L.C.; Marsland, B.J. The influence of the microbiome on respiratory health. Nat. Immunol. 2019, 20, 1279–1290. [Google Scholar] [CrossRef] [PubMed]
- Hao, Q.; Dong, B.R.; Wu, T. Probiotics for preventing acute upper respiratory tract infections. Cochrane Database Syst. Rev. 2015, CD006895. [Google Scholar] [CrossRef] [PubMed]
- Hao, Q.; Lu, Z.; Dong, B.R.; Huang, C.Q.; Wu, T. Probiotics for preventing acute upper respiratory tract infections. Cochrane Database Syst. Rev. 2011, CD006895. [Google Scholar] [CrossRef]
- Kesson, A.M. Respiratory virus infections. Paediatr. Respir. Rev. 2007, 8, 240–248. [Google Scholar] [CrossRef]
- Dunn, J.J.; Miller, M.B. Emerging respiratory viruses other than influenza. Clin. Lab. Med. 2014, 34, 409–430. [Google Scholar] [CrossRef]
- Zou, L.; Ruan, F.; Huang, M.; Liang, L.; Huang, H.; Hong, Z.; Yu, J.; Kang, M.; Song, Y.; Xia, J.; et al. SARS-CoV-2 Viral Load in Upper Respiratory Specimens of Infected Patients. N. Engl. J. Med. 2020, 382, 1177–1179. [Google Scholar] [CrossRef]
- Monto, A.S. Epidemiology of viral respiratory infections. Am. J. Med. 2002, 112, 4s–12s. [Google Scholar] [CrossRef]
- Chen, W.-J.; Arnold, J.C.; Fairchok, M.P.; Danaher, P.J.; McDonough, E.A.; Blair, P.J.; Garcia, J.; Halsey, E.S.; Schofield, C.; Ottolini, M.; et al. Epidemiologic, clinical, and virologic characteristics of human rhinovirus infection among otherwise healthy children and adults: Rhinovirus among adults and children. J. Clin. Virol. 2015, 64, 74–82. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, J.L.; Turner, R.B.; Braciale, T.; Heymann, P.W.; Borish, L. Pathogenesis of rhinovirus infection. Curr. Opin. Virol. 2012, 2, 287–293. [Google Scholar] [CrossRef] [PubMed]
- Lehtinen, M.J.; Hibberd, A.A.; Männikkö, S.; Yeung, N.; Kauko, T.; Forssten, S.; Lehtoranta, L.; Lahtinen, S.J.; Stahl, B.; Lyra, A.; et al. Nasal microbiota clusters associate with inflammatory response, viral load, and symptom severity in experimental rhinovirus challenge. Sci. Rep. 2018, 8, 11411. [Google Scholar] [CrossRef]
- Sanjuán, R.; Domingo-Calap, P. Mechanisms of viral mutation. Cell Mol. Life Sci. 2016, 73, 4433–4448. [Google Scholar] [CrossRef] [PubMed]
- Christiaansen, A.; Varga, S.M.; Spencer, J.V. Viral manipulation of the host immune response. Curr. Opin. Immunol. 2015, 36, 54–60. [Google Scholar] [CrossRef]
- Kutter, J.S.; Spronken, M.I.; Fraaij, P.L.; Fouchier, R.A.; Herfst, S. Transmission routes of respiratory viruses among humans. Curr. Opin. Virol. 2018, 28, 142–151. [Google Scholar] [CrossRef]
- Proud, D.; Leigh, R. Epithelial cells and airway diseases. Immunol. Rev. 2011, 242, 186–204. [Google Scholar] [CrossRef]
- Newton, A.H.; Cardani, A.; Braciale, T.J. The host immune response in respiratory virus infection: Balancing virus clearance and immunopathology. Semin. Immunopathol. 2016, 38, 471–482. [Google Scholar] [CrossRef]
- Norrby, E. The morphology of virus particles. Classification of viruses. In Textbook of Medical Virology; Lycke, E., Norrby, E., Eds.; Butterworth-Heinemann: Oxford, UK, 1983; pp. 4–16. [Google Scholar] [CrossRef]
- Kreijtz, J.H.; Fouchier, R.A.; Rimmelzwaan, G.F. Immune responses to influenza virus infection. Virus Res. 2011, 162, 19–30. [Google Scholar] [CrossRef]
- Totura, A.L.; Baric, R.S. SARS coronavirus pathogenesis: Host innate immune responses and viral antagonism of interferon. Curr. Opin. Virol. 2012, 2, 264–275. [Google Scholar] [CrossRef]
- Kuiken, T.; Riteau, B.; Fouchier, R.A.; Rimmelzwaan, G.F. Pathogenesis of influenza virus infections: The good, the bad and the ugly. Curr. Opin. Virol. 2012, 2, 276–286. [Google Scholar] [CrossRef] [PubMed]
- Lupfer, C.; Malik, A.; Kanneganti, T.D. Inflammasome control of viral infection. Curr. Opin. Virol. 2015, 12, 38–46. [Google Scholar] [CrossRef] [PubMed]
- Swanson, K.V.; Deng, M.; Ting, J.P. The NLRP3 inflammasome: Molecular activation and regulation to therapeutics. Nat. Rev. Immunol. 2019, 19, 477–489. [Google Scholar] [CrossRef] [PubMed]
- Mortha, A.; Burrows, K. Cytokine Networks between Innate Lymphoid Cells and Myeloid Cells. Front. Immunol. 2018, 9, 191. [Google Scholar] [CrossRef] [PubMed]
- Iwasaki, A.; Foxman, E.F.; Molony, R.D. Early local immune defences in the respiratory tract. Nat. Rev. Immunol. 2017, 17, 7–20. [Google Scholar] [CrossRef] [PubMed]
- Gorski, S.A.; Hahn, Y.S.; Braciale, T.J. Group 2 Innate Lymphoid Cell Production of IL-5 Is Regulated by NKT Cells during Influenza Virus Infection. PLoS Pathog. 2013, 9, e1003615. [Google Scholar] [CrossRef]
- Openshaw, P.J.M.; Chiu, C.; Culley, F.J.; Johansson, C. Protective and Harmful Immunity to RSV Infection. Annu. Rev. Immunol. 2017, 35, 501–532. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, M.E.; Varga, S.M. The CD8 T Cell Response to Respiratory Virus Infections. Front. Immunol. 2018, 9, 678. [Google Scholar] [CrossRef]
- Message, S.D.; Johnston, S.L. Host defense function of the airway epithelium in health and disease: Clinical background. J. Leukoc. Biol. 2004, 75, 5–17. [Google Scholar] [CrossRef]
- Damjanovic, D.; Small, C.-L.; Jeyananthan, M.; McCormick, S.; Xing, Z. Immunopathology in influenza virus infection: Uncoupling the friend from foe. Clin. Immunol. 2012, 144, 57–69. [Google Scholar] [CrossRef]
- Tay, M.Z.; Poh, C.M.; Rénia, L.; MacAry, P.A.; Ng, L.F.P. The trinity of COVID-19: Immunity, inflammation and intervention. Nat. Rev. Immunol. 2020, 20, 363–374. [Google Scholar] [CrossRef] [PubMed]
- Gern, J.E.; Vrtis, R.; Grindle, K.A.; Swenson, C.; Busse, W.W. Relationship of upper and lower airway cytokines to outcome of experimental rhinovirus infection. Am. J. Respir. Crit. Care Med. 2000, 162, 2226–2231. [Google Scholar] [CrossRef] [PubMed]
- Teran, L.M.; Johnston, S.L.; Schröder, J.M.; Church, M.K.; Holgate, S.T. Role of nasal interleukin-8 in neutrophil recruitment and activation in children with virus-induced asthma. Am. J. Respir Crit Care Med. 1997, 155, 1362–1366. [Google Scholar] [CrossRef] [PubMed]
- Röseler, S.; Holtappels, G.; Wagenmann, M.; Bachert, C. Elevated levels of interleukins IL-1 beta, IL-6 and IL-8 in naturally acquired viral rhinitis. Eur. Arch. Otorhinolaryngol. 1995, 252, S61–63. [Google Scholar] [CrossRef]
- Kaiser, L.; Fritz, R.S.; Straus, S.E.; Gubareva, L.; Hayden, F.G. Symptom pathogenesis during acute influenza: Interleukin-6 and other cytokine responses. J. Med. Virol. 2001, 64, 262–268. [Google Scholar] [CrossRef]
- Arpaia, N.; Green, J.A.; Moltedo, B.; Arvey, A.; Hemmers, S.; Yuan, S.; Treuting, P.M.; Rudensky, A.Y. A Distinct Function of Regulatory T Cells in Tissue Protection. Cell 2015, 162, 1078–1089. [Google Scholar] [CrossRef]
- Sun, J.; Madan, R.; Karp, C.L.; Braciale, T.J. Effector T cells control lung inflammation during acute influenza virus infection by producing IL-10. Nat. Med. 2009, 15, 277–284. [Google Scholar] [CrossRef]
- Russell, C.D.; Schwarze, J. The role of pro-resolution lipid mediators in infectious disease. Immunology 2014, 141, 166–173. [Google Scholar] [CrossRef]
- Hill, C.; Guarner, F.; Reid, G.; Gibson, G.R.; Merenstein, D.J.; Pot, B.; Morelli, L.; Canani, R.B.; Flint, H.J.; Salminen, S.; et al. Expert consensus document. The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat. Rev. Gastroenterol. Hepatol. 2014, 11, 506–514. [Google Scholar] [CrossRef]
- Zheng, J.; Wittouck, S.; Salvetti, E.; Franz, C.; Harris, H.M.B.; Mattarelli, P.; O’Toole, P.W.; Pot, B.; Vandamme, P.; Walter, J.; et al. A taxonomic note on the genus Lactobacillus: Description of 23 novel genera, emended description of the genus Lactobacillus Beijerinck 1901, and union of Lactobacillaceae and Leuconostocaceae. Int. J. Syst. Evol. Microbiol. 2020, 70, 2782–2858. [Google Scholar] [CrossRef]
- Tapiovaara, L.; Lehtoranta, L.; Poussa, T.; Mäkivuokko, H.; Korpela, R.; Pitkäranta, A. Absence of adverse events in healthy individuals using probiotics—Analysis of six randomised studies by one study group. Benef. Microbes 2016, 7, 161–169. [Google Scholar] [CrossRef] [PubMed]
- Didari, T.; Solki, S.; Mozaffari, S.; Nikfar, S.; Abdollahi, M. A systematic review of the safety of probiotics. Expert Opin. Drug Saf. 2014, 13, 227–239. [Google Scholar] [CrossRef]
- Sanders, M.E.; Akkermans, L.M.A.; Haller, D.; Hammerman, C.; Heimbach, J.; Hörmannsperger, G.; Huys, G.; Levy, D.D.; Lutgendorff, F.; Mack, D.; et al. Safety assessment of probiotics for human use. Gut Microbes 2010, 1, 164–185. [Google Scholar] [CrossRef] [PubMed]
- King, S.; Glanville, J.; Sanders, M.E.; Fitzgerald, A.; Varley, D. Effectiveness of probiotics on the duration of illness in healthy children and adults who develop common acute respiratory infectious conditions: A systematic review and meta-analysis. Br. J. Nutr. 2014, 112, 41–54. [Google Scholar] [CrossRef]
- Pot, B.; Foligné, B.; Daniel, C.; Grangette, C. Understanding immunomodulatory effects of probiotics. Nestle Nutr. Inst. Workshop Ser. 2013, 77, 75–90. [Google Scholar] [CrossRef] [PubMed]
- Kumpu, M.; Swanljung, E.; Tynkkynen, S.; Hatakka, K.; Kekkonen, R.A.; Järvenpää, S.; Korpela, R.; Pitkäranta, A. Recovery of probiotic Lactobacillus rhamnosus GG in tonsil tissue after oral administration: Randomised, placebo-controlled, double-blind clinical trial. Br. J. Nutr. 2013, 109, 2240–2246. [Google Scholar] [CrossRef] [PubMed]
- Swanljung, E.; Tapiovaara, L.; Lehtoranta, L.; Mäkivuokko, H.; Roivainen, M.; Korpela, R.; Pitkäranta, A. Lactobacillus rhamnosus GG in adenoid tissue: Double-blind, placebo-controlled, randomized clinical trial. Acta Oto-Laryngologica 2015, 135, 824–830. [Google Scholar] [CrossRef]
- Troost, F.J.; van Baarlen, P.; Lindsey, P.; Kodde, A.; de Vos, W.M.; Kleerebezem, M.; Brummer, R.-J.M. Identification of the transcriptional response of human intestinal mucosa to Lactobacillus plantarum WCFS1 in vivo. BMC Genom. 2008, 9, 374. [Google Scholar] [CrossRef]
- van Baarlen, P.; Wells, J.M.; Kleerebezem, M. Regulation of intestinal homeostasis and immunity with probiotic lactobacilli. Trends Immunol. 2013, 34, 208–215. [Google Scholar] [CrossRef]
- Brandtzaeg, P. Mucosal immunity: Induction, dissemination, and effector functions. Scand. J. Immunol. 2009, 70, 505–515. [Google Scholar] [CrossRef]
- Venegas, D.P.; De la Fuente, M.K.; Landskron, G.; González, M.J.; Quera, R.; Dijkstra, G.; Harmsen, H.J.M.; Faber, K.N.; Hermoso, M.A. Short Chain Fatty Acids (SCFAs)-Mediated Gut Epithelial and Immune Regulation and Its Relevance for Inflammatory Bowel Diseases. Front. Immunol. 2019, 10, 277. [Google Scholar] [CrossRef] [PubMed]
- Bermudez-Brito, M.; Plaza-Díaz, J.; Muñoz-Quezada, S.; Gómez-Llorente, C.; Gil, A. Probiotic mechanisms of action. Ann. Nutr. Metab. 2012, 61, 160–174. [Google Scholar] [CrossRef] [PubMed]
- Enaud, R.; Prevel, R.; Ciarlo, E.; Beaufils, F.; Wieërs, G.; Guery, B.; Delhaes, L. The Gut-Lung Axis in Health and Respiratory Diseases: A Place for Inter-Organ and Inter-Kingdom Crosstalks. Front. Cell Infect. Microbiol. 2020, 10, 9. [Google Scholar] [CrossRef]
- Anand, S.; Mande, S.S. Diet, Microbiota and Gut-Lung Connection. Front. Microbiol. 2018, 9, 2147. [Google Scholar] [CrossRef] [PubMed]
- Christensen, H.R.; Frøkiær, H.; Pestka, J.J. Lactobacilli Differentially Modulate Expression of Cytokines and Maturation Surface Markers in Murine Dendritic Cells. J. Immunol. 2002, 168, 171. [Google Scholar] [CrossRef] [PubMed]
- Fink, L.N.; Zeuthen, L.H.; Ferlazzo, G.; Frøkiær, H. Human antigen-presenting cells respond differently to gut-derived probiotic bacteria but mediate similar strain-dependent NK and T cell activation. FEMS Immunol. Med. Microbiol. 2007, 51, 535–546. [Google Scholar] [CrossRef] [PubMed]
- Baaten, B.J.; Cooper, A.M.; Swain, S.L.; Bradley, L.M. Location, location, location: The impact of migratory heterogeneity on T cell function. Front. Immunol. 2013, 4, 311. [Google Scholar] [CrossRef] [PubMed]
- Weiss, G.; Rasmussen, S.; Zeuthen, L.H.; Nielsen, B.N.; Jarmer, H.; Jespersen, L.; Frokiaer, H. Lactobacillus acidophilus induces virus immune defence genes in murine dendritic cells by a Toll-like receptor-2-dependent mechanism. Immunology 2010, 131, 268–281. [Google Scholar] [CrossRef]
- Nakayama, Y.; Moriya, T.; Sakai, F.; Ikeda, N.; Shiozaki, T.; Hosoya, T.; Nakagawa, H.; Miyazaki, T. Oral administration of Lactobacillus gasseri SBT2055 is effective for preventing influenza in mice. Sci. Rep. 2014, 4, 4638. [Google Scholar] [CrossRef] [PubMed]
- Miettinen, M.; Pietila, T.E.; Kekkonen, R.A.; Kankainen, M.; Latvala, S.; Pirhonen, J.; Osterlund, P.; Korpela, R.; Julkunen, I. Nonpathogenic Lactobacillus rhamnosus activates the inflammasome and antiviral responses in human macrophages. Gut Microbes 2012, 3, 510–522. [Google Scholar] [CrossRef]
- Gao, X.; Huang, L.; Zhu, L.; Mou, C.; Hou, Q.; Yu, Q. Inhibition of H9N2 virus invasion into dendritic cells by the S-Layer protein from L. acidophilus ATCC 4356. Front. Cell Infect. Microbiol. 2016, 6, 137. [Google Scholar] [CrossRef]
- Huang, C.C.; Duffy, K.E.; San Mateo, L.R.; Amegadzie, B.Y.; Sarisky, R.T.; Mbow, M.L. A pathway analysis of poly(I:C)-induced global gene expression change in human peripheral blood mononuclear cells. Physiol. Genom. 2006, 26, 125–133. [Google Scholar] [CrossRef] [PubMed]
- Vareille, M.; Kieninger, E.; Edwards, M.R.; Regamey, N. The airway epithelium: Soldier in the fight against respiratory viruses. Clin. Microbiol. Rev. 2011, 24, 210–229. [Google Scholar] [CrossRef] [PubMed]
- Aeffner, F.; Traylor, Z.P.; Yu, E.N.; Davis, I.C. Double-stranded RNA induces similar pulmonary dysfunction to respiratory syncytial virus in BALB/c mice. Am. J. Physiol. Lung Cell Mol. Physiol. 2011, 301, L99–L109. [Google Scholar] [CrossRef] [PubMed]
- Vintiñi, E.O.; Medina, M.S. Non-Viable Lactobacillus Casei Beneficially Modulates Poly I: C Immune Response in Co-Cultures of Human Cells. Iran. J. Immunol. 2017, 14, 325–339. [Google Scholar] [PubMed]
- Kanmani, P.; Kim, H. Immunobiotic strains modulate Toll-like receptor 3 agonist induced innate antiviral immune response in human intestinal epithelial cells by modulating IFN regulatory factor 3 and NF-κB signaling. Front. Immunol. 2019, 10, 1536. [Google Scholar] [CrossRef]
- Wang, Y.; Xie, J.; Wang, N.; Li, Y.; Sun, X.; Zhang, Y.; Zhang, H. Lactobacillus casei Zhang modulate cytokine and Toll-like receptor expression and beneficially regulate poly I: C-induced immune responses in RAW264. 7 macrophages. Microbiol. Immunol. 2013, 57, 54–62. [Google Scholar] [CrossRef]
- Hosoya, S.; Villena, J.; Shimazu, T.; Tohno, M.; Fujie, H.; Chiba, E.; Shimosato, T.; Aso, H.; Suda, Y.; Kawai, Y. Immunobiotic lactic acid bacteria beneficially regulate immune response triggered by poly (I: C) in porcine intestinal epithelial cells. Veter. Res. 2011, 42, 111. [Google Scholar] [CrossRef]
- Ishizuka, T.; Kanmani, P.; Kobayashi, H.; Miyazaki, A.; Soma, J.; Suda, Y.; Aso, H.; Nochi, T.; Iwabuchi, N.; Xiao, J.Z.; et al. Immunobiotic Bifidobacteria Strains Modulate Rotavirus Immune Response in Porcine Intestinal Epitheliocytes via Pattern Recognition Receptor Signaling. PLoS ONE 2016, 11, e0152416. [Google Scholar] [CrossRef]
- Kanmani, P.; Albarracin, L.; Kobayashi, H.; Iida, H.; Komatsu, R.; Humayun Kober, A.K.M.; Ikeda-Ohtsubo, W.; Suda, Y.; Aso, H.; Makino, S.; et al. Exopolysaccharides from Lactobacillus delbrueckii OLL1073R-1 modulate innate antiviral immune response in porcine intestinal epithelial cells. Mol. Immunol. 2018, 93, 253–265. [Google Scholar] [CrossRef]
- Villena, J.; Chiba, E.; Vizoso-Pinto, M.G.; Tomosada, Y.; Takahashi, T.; Ishizuka, T.; Aso, H.; Salva, S.; Alvarez, S.; Kitazawa, H. Immunobiotic Lactobacillus rhamnosus strains differentially modulate antiviral immune response in porcine intestinal epithelial and antigen presenting cells. BMC Microbiol. 2014, 14, 126. [Google Scholar] [CrossRef] [PubMed]
- Arai, S.; Iwabuchi, N.; Takahashi, S.; Xiao, J.Z.; Abe, F.; Hachimura, S. Orally administered heat-killed Lactobacillus paracasei MCC1849 enhances antigen-specific IgA secretion and induces follicular helper T cells in mice. PLoS ONE 2018, 13, e0199018. [Google Scholar] [CrossRef] [PubMed]
- Belkacem, N.; Serafini, N.; Wheeler, R.; Derrien, M.; Boucinha, L.; Couesnon, A.; Cerf-Bensussan, N.; Gomperts Boneca, I.; Di Santo, J.P.; Taha, M.K.; et al. Lactobacillus paracasei feeding improves immune control of influenza infection in mice. PLoS ONE 2017, 12, e0184976. [Google Scholar] [CrossRef] [PubMed]
- Kawahara, T.; Takahashi, T.; Oishi, K.; Tanaka, H.; Masuda, M.; Takahashi, S.; Takano, M.; Kawakami, T.; Fukushima, K.; Kanazawa, H.; et al. Consecutive oral administration of Bifidobacterium longum MM-2 improves the defense system against influenza virus infection by enhancing natural killer cell activity in a murine model. Microbiol. Immunol. 2015, 59, 1–12. [Google Scholar] [CrossRef]
- Mahooti, M.; Abdolalipour, E.; Salehzadeh, A.; Mohebbi, S.R.; Gorji, A.; Ghaemi, A. Immunomodulatory and prophylactic effects of Bifidobacterium bifidum probiotic strain on influenza infection in mice. World J. Microbiol. Biotechnol. 2019, 35, 91. [Google Scholar] [CrossRef]
- Song, J.A.; Kim, H.J.; Hong, S.K.; Lee, D.H.; Lee, S.W.; Song, C.S.; Kim, K.T.; Choi, I.S.; Lee, J.B.; Park, S.Y. Oral intake of Lactobacillus rhamnosus M21 enhances the survival rate of mice lethally infected with influenza virus. J. Microbiol. Immunol. Infect. 2016, 49, 16–23. [Google Scholar] [CrossRef]
- Eguchi, K.; Fujitani, N.; Nakagawa, H.; Miyazaki, T. Prevention of respiratory syncytial virus infection with probiotic lactic acid bacterium Lactobacillus gasseri SBT2055. Sci. Rep. 2019, 9, 4812. [Google Scholar] [CrossRef]
- Laursen, R.P.; Hojsak, I. Probiotics for respiratory tract infections in children attending day care centers-a systematic review. Eur. J. Pediatr. 2018, 177, 979–994. [Google Scholar] [CrossRef]
- Wang, Y.; Li, X.; Ge, T.; Xiao, Y.; Liao, Y.; Cui, Y.; Zhang, Y.; Ho, W.; Yu, G.; Zhang, T. Probiotics for prevention and treatment of respiratory tract infections in children: A systematic review and meta-analysis of randomized controlled trials. Medicine 2016, 95, e4509. [Google Scholar] [CrossRef]
- Luoto, R.; Ruuskanen, O.; Waris, M.; Kalliomäki, M.; Salminen, S.; Isolauri, E. Prebiotic and probiotic supplementation prevents rhinovirus infections in preterm infants: A randomized, placebo-controlled trial. J. Allergy Clin. Immunol. 2014, 133, 405–413. [Google Scholar] [CrossRef]
- Waki, N.; Matsumoto, M.; Fukui, Y.; Suganuma, H. Effects of probiotic Lactobacillus brevis KB290 on incidence of influenza infection among schoolchildren: An open-label pilot study. Lett. Appl. Microbiol. 2014, 59, 565–571. [Google Scholar] [CrossRef] [PubMed]
- Kumpu, M.; Lehtoranta, L.; Roivainen, M.; Rönkkö, E.; Ziegler, T.; Söderlund-Venermo, M.; Kautiainen, H.; Järvenpää, S.; Kekkonen, R.; Hatakka, K.; et al. The use of the probiotic Lactobacillus rhamnosus GG and viral findings in the nasopharynx of children attending day care. J. Med. Virol. 2013, 85, 1632–1638. [Google Scholar] [CrossRef] [PubMed]
- Lehtoranta, L.; Söderlund-Venermo, M.; Nokso-Koivisto, J.; Toivola, H.; Blomgren, K.; Hatakka, K.; Poussa, T.; Korpela, R.; Pitkäranta, A. Human bocavirus in the nasopharynx of otitis-prone children. Int. J. Pediatr. Otorhinolaryngol. 2012, 76, 206–211. [Google Scholar] [CrossRef] [PubMed]
- Lehtoranta, L.; Kalima, K.; He, L.; Lappalainen, M.; Roivainen, M.; Närkiö, M.; Mäkelä, M.; Siitonen, S.; Korpela, R.; Pitkäranta, A. Specific probiotics and virological findings in symptomatic conscripts attending military service in Finland. J. Clin. Virol. 2014, 60, 276–281. [Google Scholar] [CrossRef]
- Wang, B.; Hylwka, T.; Smieja, M.; Surrette, M.; Bowdish, D.M.E.; Loeb, M. Probiotics to Prevent Respiratory Infections in Nursing Homes: A Pilot Randomized Controlled Trial. J. Am. Geriatr. Soc. 2018, 66, 1346–1352. [Google Scholar] [CrossRef]
- Kumpu, M.; Kekkonen, R.A.; Korpela, R.; Tynkkynen, S.; Järvenpää, S.; Kautiainen, H.; Allen, E.K.; Hendley, J.O.; Pitkäranta, A.; Winther, B. Effect of live and inactivated Lactobacillus rhamnosus GG on experimentally induced rhinovirus colds: Randomised, double blind, placebo-controlled pilot trial. Benef. Microbes 2015, 6, 631–639. [Google Scholar] [CrossRef]
- Tapiovaara, L.; Kumpu, M.; Mäkivuokko, H.; Waris, M.; Korpela, R.; Pitkäranta, A.; Winther, B. Human rhinovirus in experimental infection after peroral Lactobacillus rhamnosus GG consumption, a pilot study. Int. Forum Allergy Rhinol. 2016, 6, 848–853. [Google Scholar] [CrossRef]
- Turner, R.B.; Woodfolk, J.A.; Borish, L.; Steinke, J.W.; Patrie, J.T.; Muehling, L.M.; Lahtinen, S.; Lehtinen, M.J. Effect of probiotic on innate inflammatory response and viral shedding in experimental rhinovirus infection—A randomized controlled trial. Benef. Microbes 2017, 8, 207–215. [Google Scholar] [CrossRef]
- West, N.P.; Horn, P.L.; Pyne, D.B.; Gebski, V.J.; Lahtinen, S.J.; Fricker, P.A.; Cripps, A.W. Probiotic supplementation for respiratory and gastrointestinal illness symptoms in healthy physically active individuals. Clin. Nutr. 2014, 33, 581–587. [Google Scholar] [CrossRef]
| Study Type and Reference | No. of Included Studies for Meta-Analysis and Analysis Population | Probiotic Effect Compared with Control | |||
|---|---|---|---|---|---|
| RTI Incidence/Risk | RTI Duration | Absence from Daycare/School/Work | Antibiotic Use | ||
| Children | |||||
| Systematic review and meta-analysis | 23 randomized, double-blinded, and placebo-controlled trials | Decreased number of subjects having at least 1 RTI episode (17 RCTs, 4513 children, relative risk 0.89, 95% CI 0.82–0.96, p = 0.004) | No significant difference of illness episode duration between study groups (9 RCTs, 2817 children, (MD 0.60, 95% CI 1.49–0.30, p = 0.19). | Fewer numbers of days absent from daycare/school (8 RCTs, 1499 children, MD 0.94, 95% CI 1.72–0.15, p = 0.02) | NA |
| [80] | 6269 children (0–18 years) | Fewer numbers of days of RTIs per person (6 RCTs, 2067 children, MD 0.16, 95% CI 0.29–0.02, p = 0.03) | |||
| Systematic review and meta-analysis | 15 randomized placebo-controlled trials | Lower number of children with RTI, reduced RTI risk, 5 RCTs, n = 1841, RR 0.78, 95% CI 0.63–0.98, random effect model). | Probiotic consumption had no effect on the duration of RTIs (9 RCTs, n = 3529, MD -0.81 days, 95% CI −1.88–0.25, random effect model). | No effect on the days absent from daycare centers (9 RCTs, n = 3040, MD -0.25 days, 95% CI −0.75–0.24, random effect model). | Reduced the risk of antibiotic use (7 RCTs, n = 2858, RR 0.69, 95% CI 0.49–0.95, random effect model). |
| [79] | 5121 children in daycare settings (3 months to 7 years) | No effect on the risk of at least one URTI (5 RCTs, n = 1711, RR 0.81, 95% CI 0.62–1.05, random effect model) | |||
| All Age Groups | |||||
| Cochrane systematic review and meta-analysis | 10 randomized, placebo-controlled trials | Lower number of participants experiencing episodes of acute URTI by 42–47%: (at least 1 episode: OR 0.58; 95% CI 0.36–0.92, p = 0.022; at least 3 episodes: OR 0.53; 95% CI 0.36–0.80, p = 0.002). | No efficacy when measuring the mean duration of acute URTI episode: MD −0.29; 95% CI −3.71–3.13 (p = 0.87) | NA | Reduced antibiotic prescription rates for acute URTIs: OR 0.67; 95% CI 0.45–0.98, p = 0.04 |
| [6] | 3451 participants (infants to elderly) | Reduction in episode rate ratio of acute URTIs (events per person/year) (rate ratio 0.88; 95% CI 081 to 0.96, p = 0.004) | |||
| Cochrane systematic review and meta-analysis | 12 randomized, placebo-controlled trials | Lower number of participants experiencing episodes of acute URTI by 47% (at least 1 episode: OR 0.53; 95% CI 0.37- 0.76, p value < 0.001; at least 3 episodes: OR 0.53; 95% CI 0.36–0.80, p= 0.002). | Reduced the mean duration of an acute URTI episode by 1.89 days (MD -1.89; 95% CI −2.03–1.75, p < 0.001) | Reduced cold-related school absence (OR 0.10; 95% CI 0.02–0.47 (only one trial) | Reduced antibiotic prescription rates for acute URTIs (OR 0.65; 95% CI 0.45–0.94, p = 0.024) |
| [5] | 3720 participants (children to elderly) | No effect when measuring episode rate ratio (events per person/year) of acute URTI (rate ratio 0.83; 95% CI 0.66–1.05, p = 0.12) | |||
| Systematic review and meta-analysis | 20 randomized controlled trials | Reduced numbers of days of illness per person (standardized MD -0.31 (95% CI −0.41 to −0.22, p < 0.001) | Shortened illness episodes by almost a day (weighted MD -0.77 (95% CI −1.50 to −0.04), p = 0.04) (without an increase in the number of illness episodes) | Reduced numbers of days absent from daycare/school/work (standardized MD -0.17 (95% CI −0.31 to −0.03. p = 0.02), | NA |
| [45] | 3-month-old children to elderly (participant numbers not specified) | ||||
| Study Type and Reference | Randomized Subjects | Probiotic Intervention | Analyzed Viruses | Study Outcomes: Probiotic vs. Placebo |
|---|---|---|---|---|
| Community | ||||
| R DB PC [81] | 94 pre-term infants (2 days–2 months) | L. rhamnosus GG 1 × 109 CFU/day (1–30day) and 2 × 109 CFU/day for 31–60 days or galacto-oligosaccharide or placebo for 60 days | From nasal swab:
| Lower incidence of rhinovirus-induced RTI episodes (p = 0.04). |
| Lower number of rhinovirus findings in acute RTI over 12 months (p = 0.015). | ||||
| No significant difference in the mean duration of symptoms in rhinovirus episodes, severity scores of clinical symptoms in rhinovirus episodes, rhinovirus RNA load during infections, duration of rhinovirus RNA shedding, duration or severity of rhinovirus. | ||||
| R DB PC [84] | 269 otitis-prone children (9 months–5.6 years) | L. rhamnosus GG, L. rhamnosus Lc705, B. breve 99 and Propionibacterium jensenii JS 8–9 × 109 CFU/day of each strain, or placebo in a capsule for 6 months | From nasal swab:
| Lower number of human bocavirus 1 positive sample during the study (6.4% vs. 19.0%, p = 0.039). |
| No effect on rhinovirus/enterovirus occurrence. | ||||
| R DB PC [83] | 97 daycare children (2–6 years) visiting health care practitioner due to RTI | L. rhamnosus GG approximately 108 CFU/day in milk for 28 weeks | From nasal swab:
| Children had less days with respiratory symptoms per month (6.5 vs. 7.2, p < 0.001). |
| No effect on the occurrence of respiratory viruses during the study or respiratory symptoms associated with viral findings. | ||||
| R DB PC [85] | 192 military conscripts (18–30 years) visiting health care practitioner due to RTI | L. rhamnosus GG 5 × 109 CFU/day + B. lactis BB-12 2 × 109 CFU/day in a chewing tablet for either 3 or 6 months | From nasal swab:
| Overall no significant effect on the occurrence of common respiratory viruses. In a subgroup, there was lower occurrence of rhino/enteroviruses after 3 months (5 vs. 15, p < 0.01). |
| Open label, parallel group [82] | 2926 schoolchildren (6–12 years) | L. brevis KB290 in a nutrient drink 6 × 109 CFU/bottle 5 day/week for 8 weeks + no consumption for 8 weeks or vice versa (2 alternate study groups) | Physician diagnosed influenza virus infection | During influenza epidemic, less influenza infections in the group consuming probiotic drink compared with the group not consuming probiotic drink (15.7% vs. 23.9%, p < 0.001) |
| R DB PC [86] | 209 nursing home residents aged ≥65 years | L. rhamnosus GG 2 × 1010 CFU/d in capsule or placebo for 6 months | From nasal swab:
| No statistically significant difference in laboratory confirmed viral respiratory infections. |
| Experimental Virus Challenge | ||||
| R DB PC [87,88] | 59 healthy adults (mean 22–24 years) | L. rhamnosus GG 109 CFU of live or heat-inactivated (by spray-drying) in 100 mL of fruit juice or control juice daily for 6 weeks. | From nasal lavage:
| No significant effect on rhinovirus infection rate. |
| No significant effect on the occurrence and severity of cold symptoms during rhinovirus infection. | ||||
| No significant effect on viral loads. | ||||
| R DB PC [89] | 115 healthy adults with confirmed experimental infection (mean 22–23 years) | B. lactis Bl-04 2 × 109 CFU powder or placebo daily for 32 days | From nasal lavage:
| Reduction in nasal rhinovirus titer and the proportion of subjects shedding virus in nasal secretions (76% vs. 91%, p = 0.04) during the infection. |
| Significantly higher IL-8 levels in nasal lavage prior to infection (90 vs. 58 pg/mL, p = 0.04). Significantly reduced IL-8 response to rhinovirus infection in nasal lavage (p = 0.03). | ||||
| No significant effect on symptom severity/scores or infection rate. | ||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lehtoranta, L.; Latvala, S.; Lehtinen, M.J. Role of Probiotics in Stimulating the Immune System in Viral Respiratory Tract Infections: A Narrative Review. Nutrients 2020, 12, 3163. https://doi.org/10.3390/nu12103163
Lehtoranta L, Latvala S, Lehtinen MJ. Role of Probiotics in Stimulating the Immune System in Viral Respiratory Tract Infections: A Narrative Review. Nutrients. 2020; 12(10):3163. https://doi.org/10.3390/nu12103163
Chicago/Turabian StyleLehtoranta, Liisa, Sinikka Latvala, and Markus J. Lehtinen. 2020. "Role of Probiotics in Stimulating the Immune System in Viral Respiratory Tract Infections: A Narrative Review" Nutrients 12, no. 10: 3163. https://doi.org/10.3390/nu12103163
APA StyleLehtoranta, L., Latvala, S., & Lehtinen, M. J. (2020). Role of Probiotics in Stimulating the Immune System in Viral Respiratory Tract Infections: A Narrative Review. Nutrients, 12(10), 3163. https://doi.org/10.3390/nu12103163





