Circulating miRNAs as Biomarkers of Obesity and Obesity-Associated Comorbidities in Children and Adolescents: A Systematic Review
Abstract
1. Introduction
2. Materials and Methods
2.1. Search Strategy and Eligibility Criteria
2.2. Data Sources and Search Strategies
2.3. Study Selection Process
2.4. Data Collection Process and Data Items
2.5. Study Quality and Risk of Bias Assessment
3. Results
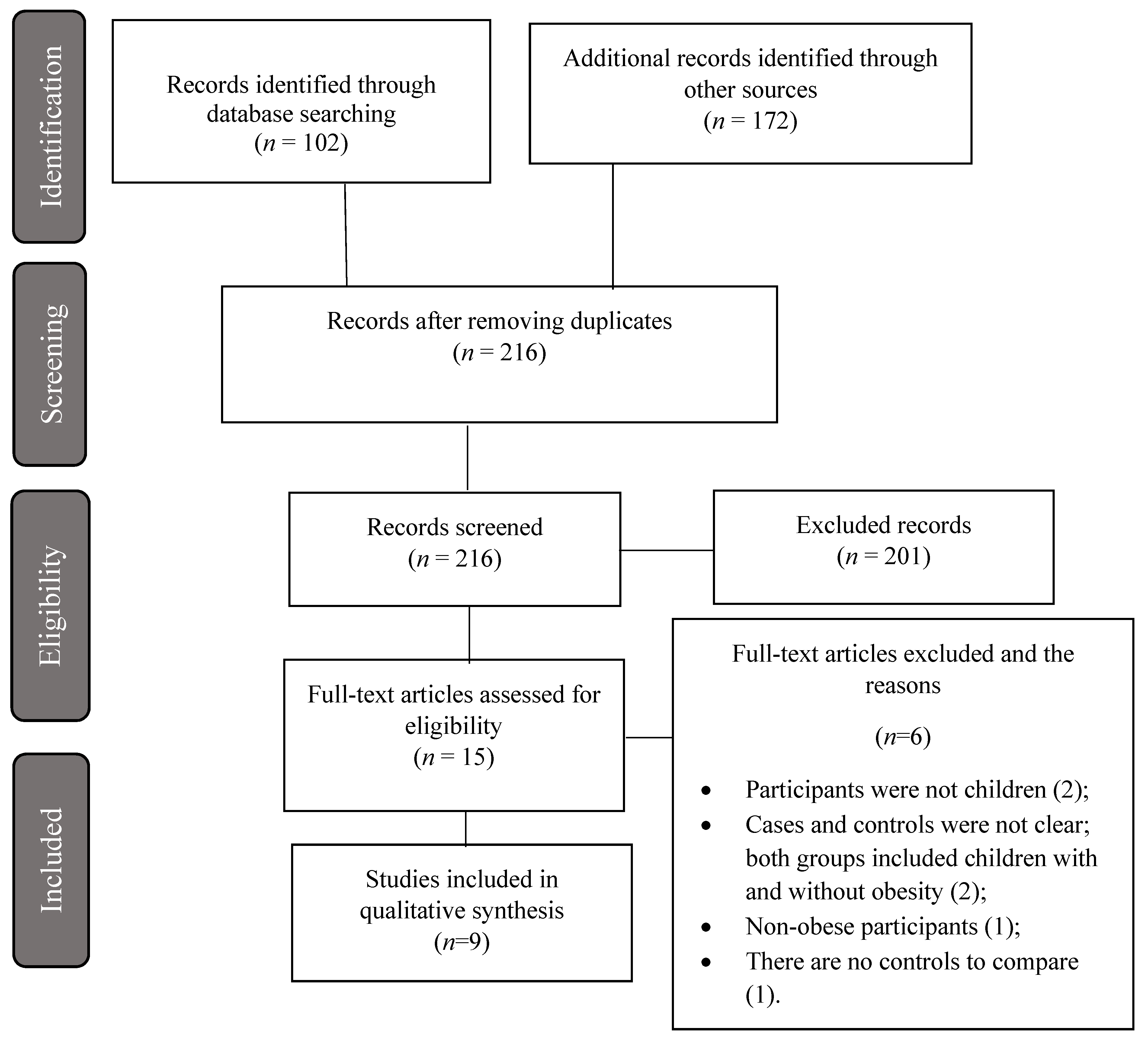
3.1. Study Selection and Characteristics
3.2. Risk of Bias Within Studies
3.3. Results of Individual Studies
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
References
- WHO|Childhood Overweight and Obesity. WHO 2017. Available online: http://www.who.int/dietphysicalactivity/childhood/en/ (accessed on 11 September 2018).
- World Health Organization. Obesity and Overweight. Available online: http://www.who.int/news-room/fact-sheets/detail/obesity-and-overweight (accessed on 22 November 2018).
- Shashaj, B.; Bedogni, G.; Graziani, M.P.; Tozzi, A.E.; DiCorpo, M.L.; Morano, D.; Tacconi, L.; Veronelli, P.; Contoli, B.; Manco, M. Origin of cardiovascular risk in overweight preschool children: A cohort study of cardiometabolic risk factors at the onset of obesity. JAMA Pediatr. 2014, 168, 917–924. [Google Scholar] [CrossRef]
- Meza, M.N.; Carrillo, J.A. Biomarkers, Obesity, and Cardiovascular Diseases. Role Biomark. Med. 2016, 119. [Google Scholar] [CrossRef][Green Version]
- Katsareli, E.A.; Dedoussis, G.V. Biomarkers in the Field of Obesity and Its Related Comorbidities. Expert Opin. Ther. Targets 2014, 18, 385–401. Available online: http://www.tandfonline.com/doi/full/10.1517/14728222.2014.882321 (accessed on 23 November 2018). [CrossRef]
- Bazzini, A.A.; Johnstone, T.G.; Christiano, R.; MacKowiak, S.D.; Obermayer, B.; Fleming, E.S.; E Vejnar, C.; Lee, M.T.; Rajewsky, N.; Walther, T.C.; et al. Identification of small ORFs in vertebrates using ribosome footprinting and evolutionary conservation. EMBO J. 2014, 33, 981–993. [Google Scholar] [CrossRef]
- Ambros, V. The functions of animal microRNAs. Nature 2004, 431, 350–355. [Google Scholar] [CrossRef]
- Wronska, A.; Kurkowska-Jastrzebska, I.; Santulli, G. Application of microRNAs in diagnosis and treatment of cardiovascular disease. Acta Physiol. 2015, 213, 60–83. [Google Scholar] [CrossRef]
- Etheridge, A.; Lee, I.; Hood, L.; Galas, D.; Wang, K. Extracellural microRNA: A new resource of biomarkers. Mutat. Res. 2011, 717, 85–90. Available online: http://www.sciencedirect.com/science/article/pii/S0165614710000921 (accessed on 23 November 2018). [CrossRef]
- Higuchi, C.; Nakatsuka, A.; Eguchi, J.; Teshigawara, S.; Kanzaki, M.; Katayama, A.; Yamaguchi, S.; Takahashi, N.; Murakami, K.; Ogawa, D.; et al. Identification of circulating miR-101, miR-375 and miR-802 as biomarkers for type 2 diabetes. Metabolism 2015, 64, 489–497. [Google Scholar] [CrossRef]
- Ding, Y.; Sun, X.; Shan, P.-F. MicroRNAs and Cardiovascular Disease in Diabetes Mellitus. Biomed. Res. Int. 2017, 2017. Available online: https://www.hindawi.com/journals/bmri/2017/4080364/ (accessed on 23 November 2018). [CrossRef]
- Hilton, C.; Neville, M.J.; Karpe, F. MicroRNAs in adipose tissue: Their role in adipogenesis and obesity. Int. J. Obes. 2013, 37, 325–332. [Google Scholar] [CrossRef]
- Quiat, D.; Olson, E.E.N. MicroRNAs in cardiovascular disease: From pathogenesis to prevention and treatment. J. Clin. Investig. 2013, 123, 11–18. Available online: http://www.jci.org/articles/view/62876?utm_campaign=impact_2013_january&utm_content=short_url&utm_medium=pdf&utm_source=impact (accessed on 25 November 2018). [CrossRef]
- Dehwah, M.A.S.; Xu, A.; Huang, Q. MicroRNAs and type 2 diabetes/obesity. J. Genet. Genom. 2012, 39, 11–18. [Google Scholar] [CrossRef]
- Schardt, C.; Adams, M.B.; Owens, T.; Keitz, S.; Fontelo, P. Utilization of the PICO framework to improve searching PubMed for clinical questions. BMC Med. Inform. Decis. Mak. 2007, 7, 16. [Google Scholar] [CrossRef]
- Downes, M.J.; Brennan, M.L.; Williams, H.C.; Dean, R.S. Development of a critical appraisal tool to assess the quality of cross-sectional studies (AXIS). BMJ Open. 2016, 6, e011458. [Google Scholar] [CrossRef]
- Downes, M.J.; Brennan, M.L.; Williams, H.C.; Dean, R.S. Appraisal tool for Cross-Sectional Studies (AXIS). BMJ Open. 2016. Available online: http://bmjopen.bmj.com/content/bmjopen/6/12/e011458/DC2/embed/inline-supplementary-material-2.pdf?download=true (accessed on 4 December 2018). [CrossRef]
- Effective Public Health Practice Project. Quality Assessment Tool for Quantitative Studies; McMaster University: Toronto, ON, Canada, 2010; pp. 2–5. Available online: http://www.ephpp.ca/tools.html (accessed on 4 December 2018).
- Cui, X.; You, L.; Zhu, L.; Wang, X.; Zhou, Y.; Li, Y.; Wen, J.; Xia, Y.; Wang, X.; Ji, C.; et al. Change in circulating microRNA profile of obese children indicates future risk of adult diabetes. Metabolism 2018, 78, 95–105. [Google Scholar] [CrossRef]
- Ouyang, S.; Tang, R.; Liu, Z.; Ma, F.; Li, Y.; Wu, J. Characterization and predicted role of microRNA expression profiles associated with early childhood obesity. Mol. Med. Rep. 2017, 16, 3799–3806. [Google Scholar] [CrossRef][Green Version]
- Iacomino, G.; Russo, P.; Stillitano, I.; Lauria, F.; Marena, P.; Ahrens, W.; De Luca, P.; Siani, A. Circulating microRNAs are deregulated in overweight/obese children: Preliminary results of the I. Family study. Genes Nutr. 2016, 11, 3–11. [Google Scholar] [CrossRef]
- Can, U.; Buyukinan, M.; Yerlikaya, F.H. The investigation of circulating microRNAs associated with lipid metabolism in childhood obesity. Pediatr. Obes. 2016, 11, 228–234. [Google Scholar] [CrossRef]
- Prats-Puig, A.; Ortega, F.J.; Mercader, J.M.; Moreno-Navarrete, J.M.; Moreno, M.; Bonet, N.; Ricart, W.; Lopez-Bermejo, A.; Fernández-Real, J.M. Changes in Circulating MicroRNAs Are Associated with Childhood Obesity. J. Clin. Endocrinol. Metab. 2013, 98, E1655–E1660. [Google Scholar] [CrossRef]
- Carolan, E.; Hogan, A.E.; Corrigan, M.; Gaotswe, G.; O’Connell, J.; Foley, N.; O’Neill, L.A.; Cody, D.; O’Shea, D. The impact of childhood obesity on inflammation, innate immune cell frequency, and metabolic microRNA expression. J. Clin. Endocrinol. Metab. 2014, 99, 474–478. [Google Scholar] [CrossRef] [PubMed]
- Al-Rawaf, H.A. Circulating microRNAs and adipokines as markers of metabolic syndrome in adolescents with obesity. Clin. Nutr. 2018. Available online: https://www.sciencedirect.com/science/article/pii/S0261561418324622?dgcid=rss_sd_all (accessed on 25 November 2018). [CrossRef] [PubMed]
- Thompson, M.D.; Cismowski, M.J.; Serpico, M.; Pusateri, A.; Brigstock, D.R. Elevation of circulating microRNA levels in obese children compared to healthy controls. Clin. Obes. 2017, 7, 216–221. [Google Scholar] [CrossRef] [PubMed]
- Masotti, A.; Baldassarre, A.; Fabrizi, M.; Olivero, G.; Loreti, M.C.; Giammaria, P.; Graziani, M.P.; Manco, M. Oral glucose tolerance test unravels circulating miRNAs associated with insulin resistance in obese preschoolers. Pediatr. Obes. 2017, 12, 229–238. [Google Scholar] [CrossRef]
- Ortega, F.J.; Mercader, J.M.; Catalán, V.; Moreno-Navarrete, J.M.; Pueyo, N.; Sabater, M.; Gomez-Ambrosi, J.; Anglada, R.; Fernandez-Formoso, J.A.; Ricart, W.; et al. Targeting the circulating microRNA signature of obesity. Clin. Chem. 2013, 59, 781–792. [Google Scholar] [CrossRef]
- Delic, D.; Eisele, C.; Schmid, R.; Luippold, G.; Mayoux, E.; Grempler, R. Characterization of micro-RNA changes during the progression of type 2 diabetes in Zucker diabetic fatty rats. Int. J. Mol. Sci. 2016, 17, 665. [Google Scholar] [CrossRef]
- Esau, C.; Davis, S.; Murray, S.F.; Yu, X.X.; Pandey, S.K.; Pear, M.; Watts, L.; Booten, S.L.; Graham, M.; McKay, R.; et al. miR-122 regulation of lipid metabolism revealed by in vivo antisense targeting. Cell Metab. 2006, 3, 87–98. [Google Scholar] [CrossRef]
- Willeit, P.; Skroblin, P.; Moschen, A.R.; Yin, X.; Kaudewitz, D.; Zampetaki, A.; Barwari, T.; Whitehead, M.; Ramírez, C.M.; Goedeke, L.; et al. Circulating MicroRNA-122 is associated with the risk of new-onset metabolic syndrome and type 2 diabetes. Diabetes 2017, 66, 347–357. [Google Scholar] [CrossRef]
- Miyaaki, H.; Ichikawa, T.; Kamo, Y.; Taura, N.; Honda, T.; Shibata, H.; Milazzo, M.; Fornari, F.; Gramantieri, L.; Bolondi, L.; et al. Significance of serum and hepatic microRNA-122 levels in patients with non-alcoholic fatty liver disease. Liver Int. 2014, 34, e302–e307. [Google Scholar] [CrossRef]
- Yamada, H.; Suzuki, K.; Ichino, N.; Ando, Y.; Sawada, A.; Osakabe, K.; Sugimoto, K.; Ohashi, K.; Teradaira, R.; Inoue, T.; et al. Associations between circulating microRNAs (miR-21, miR-34a, miR-122 and miR-451) and non-alcoholic fatty liver. Clin. Chim. Acta 2013, 424, 99–103. [Google Scholar] [CrossRef]
- Ye, D.; Zhang, T.; Lou, G.; Xu, W.; Dong, F.; Chen, G.; Liu, Y. Plasma miR-17, miR-20a, miR-20b and miR-122 as potential biomarkers for diagnosis of NAFLD in type 2 diabetes mellitus patients. Life Sci. 2018, 208, 201–207. [Google Scholar] [CrossRef]
- Stull, A.J.; Wang, Z.Q.; Zhang, X.H.; Yu, Y.; Johnson, W.D.; Cefalu, W.T. Skeletal muscle protein tyrosine phosphatase 1B regulates insulin sensitivity in African Americans. Diabetes 2012, 61, 1415–1422. [Google Scholar] [CrossRef]
- Wang, R.; Hong, J.; Cao, Y.; Shi, J.; Gu, W.; Ning, G.; Zhang, Y.; Wang, W. Elevated circulating microRNA-122 is associated with obesity and insulin resistance in young adults. Eur. J. Endocrinol. 2015, 172, 291–300. [Google Scholar] [CrossRef]
- Wang, C.C.L.; Goalstone, M.L.; Draznin, B. Molecular Mechanisms of Insulin Resistance That Impact Cardiovascular Biology. Diabetes 2004, 53, 2735–2740. [Google Scholar] [CrossRef]
- Shah, R.; Murthy, V.; Pacold, M.; Danielson, K.; Tanriverdi, K.; Larson, M.G.; Hanspers, K.; Pico, A.; Mick, E.; Reis, J.; et al. Extracellular RNAs are associated with insulin resistance and metabolic phenotypes. Diabetes Care 2017, 40, 546–553. [Google Scholar] [CrossRef]
- Ruderman, N.B.; Carling, D.; Prentki, M.; Cacicedo, J.M. Science in medicine AMPK, insulin resistance, and the metabolic syndrome. J. Clin. Investig. 2013, 123, 2764–2772. [Google Scholar] [CrossRef]
- Salvoza, N.C.; Klinzing, D.C.; Gopez-Cervantes, J.; Baclig, M.O. Association of circulating serum MIR-34a and MIR-122 with dyslipidemia among patients with non-alcoholic fatty liver disease. PLoS ONE 2016, 11, e0153497. [Google Scholar] [CrossRef]
- Liu, C.H.; Ampuero, J.; Gil-Gómez, A.; Montero-Vallejo, R.; Rojas, Á.; Muñoz-Hernández, R.; Gallego-Durán, R.; Romero-Gómez, M. miRNAs in patients with non-alcoholic fatty liver disease: A systematic review and meta-analysis. J. Hepatol. 2018, 69, 1335–1348. [Google Scholar] [CrossRef]
- Trajkovski, M.; Hausser, J.; Soutschek, J.; Bhat, B.; Akin, A.; Zavolan, M.; Heim, M.H.; Stoffel, M. MicroRNAs 103 and 107 regulate insulin sensitivity. Nature 2011, 474, 649–653. [Google Scholar] [CrossRef]
| AUTHOR | Study Aim | Biological Sample | Search for miRNA and Methodology | Participants | ||
|---|---|---|---|---|---|---|
| Type of Search Number of miRNAs Sample Size | Technique/Method | Cases Sample size Sex (girls %) Age BMI | Control Sample size Sex (girls %) Age BMI | |||
| Al-rawaf HA et al. 2018 | To describe the circulating miRNA profile for adolescences and its association with circulating levels of leptin and adiponectin according to specific degree of obesity | Circulating miRNAs (Blood-plasma) | Specific search miRNAs N = 10 N = 150 | qPCR | Obese 100 29 13.87 ± 2.91 years BMI = 26.7 ± 8.2 kg/m2 | Normal weight 50 44 13.8 ± 2.88 years BMI = 17.4 ± 4.3 kg/m2 |
| Cui et al. 2017 | To screen candidate miRNAs as biomarkers for identifying obese children who are at risk of developing diabetes | Circulating miRNAs (Blood-plasma) | Massive - N = 18 Validation miRNAs N = 18 N = 246 | Global miRNAs profiling - Illumina qPCR | Obese N = 100 51.5 61.0 ± 10.4 months BMI = 20.3 ± 2.2 kg/m2 | Normal weight N = 146 49.5 60.4 ± 11.1 months BMI = 15.1 ± 1.06 kg/m2 |
| Ouyang et al. 2017 | To characterize the miRNA profile in PBMC of obese children | Circulating miRNAs (Blood—PBMC) | Massive - N = 12 | Global miRNA Profiling - NanoString nCounter | Obese 50 39.7 ± 2.2 months BMI = 18.5 ± 26kg/m2 | Normal weight 50 39.2 ± 2.3 months BMI = 13.5 ± 15 kg/m2 |
| Thompson et al. 2017 | To Evaluate whether circulating miRNAs that have been associated with NAFLD are altered in children with obesity, compared with healthy controls | Circulating miRNAs (Blood-plasma) | Specific search for miRNAs related to NAFLD miRNAs N = 20 N = 30 | TaqMan RT-qPCR | Obese and NAFLD N = 20 42.8 13.2 ± 3.1 years BMI = 34.7 ± 10.4kg/m2 | Normal weight and non-NAFLD N = 10 60 13.8 ± 2.1 years BMI = 20.1 ± 2.5 kg/m2 |
| Iacomino et al. 2016 | To identify circulating miRNAs potentially associated with early obesity in children | Circulating miRNAs (Blood-plasma) | Specific search miRNAs N = 372 N = 20 | qPCR | Obese and overweight N = 10 40 10.7 ± 1.7 years BMI = 31,7 ± 4.3 kg/m2 | Normal weight N = 10 50 10.5 ± 2.67 years BMI = 16.4 ± 1.7 kg/m2 |
| Masotti et al. 2016 | To investigate the expression profile of circulating miRNA 1) fasting and 2)120min after OGTT in 6 IR obese preschoolers and 6 controls without IR. | Circulating miRNAs (Blood-plasma) | Specific search miRNAs N = 179 N = 12 | qPCR | Obese + IR N = 6 - 4.63 ± 1.82 years BMI = 20.9 ± 2.9 kg/m2 | Obese without IR N = 6 - 4.35 ± 0.85 years BMI = 18.5 ± 1.2 kg/m2 |
| Can et al. 2015 | To examine the relationship between 7 specific miRNAs and lipid metabolism in obese and non-obese children and adolescents. | Circulating miRNAs (Blood-plasma) | Specific search miRNAs N = 7 N = 86 | qPCR | Obese N = 45 57.7 14.71 ± 1.76 years BMI = 41.3 ± 52.9 kg/m2 | Normal weight N = 41 58.5 14.44 ± 1.62 years BMI = 18.9 ± 2.1 kg/m2 |
| Prats-Puig et al 2013. | To examine the dysregulated circulating miRNAs in obese children. | Circulating miRNAs (Blood-plasma) | Massive - N = 10 Validation miRNAs N = 15 N = 125 | Global miRNA profiling – low-density TaqMan arrays (TLDAs) qPCR | Obese N = 5 0 (only boys) 8.8 ± 1.8 years z-BMI = 3.36 ± 0.43 Obese N = 40 55 9.2 ± 1.4 years z BMI = 2.69 ± 0.59 | Normal weight N = 5 0 (only boys) 9.9 ± 1.0 years z-BMI = −0.62 ± 0.3 Lean N = 85 49 9.0 ± 1.6 years z BMI = −0.32 ± 0.71 |
| Carolan et al. 2013 | To investigate sCD163 levels, circulating iNKT frequency, cytokine profile and miR expression in obese and non-obese children. | Circulating miRNAs (Blood-PBMC) | Specific search miRNAs N = 3 N = 49 | qPCR | Obese N = 29 46.4 13.0 ± 3.0 years z-BMI = 3.4 ± 0.5 | Normal weight N = 20 35 12.8 ± 3.2 years z-BMI = 0.2 ± 1.1 |
| miRNAs | Author | Effect Size (Cases vs. Controls) | p | ||
|---|---|---|---|---|---|
| Relative Expression Level | Mean Expression Level | Fold Change | |||
| miR-222 | Al-rawaf HA et al. 2018 | 14.5 vs. 4.5 | - | - | <0.001 |
| Cui et al. 2017 | - | - | >6 | <0.01 | |
| Prats-Puig et al. 2013 | - | 41.08 ± 30.59 vs. 25.43 ± 17.87 | - | 0.001 | |
| miR-142-3p | Al-rawaf HA et al. 2018 | 12 vs. 2.5 | - | - | <0.001 |
| Prats-Puig et al. 2013 | - | 90.31 ± 61.46 vs. 32.30 ± 21.29 | - | <0.0001 | |
| miR-140-5p | Al-rawaf HA et al. 2018 | 13.5 vs. 4 | - | - | <0.001 |
| Prats-Puig et al. 2013 | - | 32.66 ± 18.13 vs. 23.15 ± 17.50 | - | 0.001 | |
| miR-143 | Al-rawaf HA et al. 2018 | 14 vs. 3.5 | - | - | <0.001 |
| Can et al. 2015 | - | 30.5 vs. 115.35 | - | 0.001 | |
| miR-532-5p | Al-rawaf HA et al. 2018 | 8 vs. 17 | - | - | <0.001 |
| Prats-Puig et al. 2013 | - | 10.49 ± 7.75 vs. 5.49 ± 4.28 | - | 0.001 | |
| miR-423-5p | Al-rawaf HA et al. 2018 | 4 vs. 14 | - | - | <0.001 |
| Prats-Puig et al. 2013 | - | 2.16 ± 1.35 vs. 1.13 ± 0.77 | - | <0.0001 | |
| miR-146a | Al-rawaf HA et al. 2018 | 4 vs. 15 | - | - | <0.001 |
| Cui et al. 2017 | - | - | 3.8 | <0.01 | |
| miRNAs | Author | Effect Size: Fold Change (Cases vs Controls) | p |
|---|---|---|---|
| miR-122-5p | Thompson et al. 2017 | 12.48 | <0.0001 |
| miR-34a-5p | 5.09 | <0.0001 | |
| miR-191-5p | 7.21 | <0.0001 | |
| miR-15b-5b | 3.42 | 0.0004 | |
| miR-199a-5p | 17.18 | <0.0001 | |
| miR-222-3p | 2.14 | <0.0001 | |
| miR-223-3p | 6.72 | <0.0001 | |
| miR-181b-5p | 3.29 | 0.0009 | |
| miR-23a-3p | 5.3 | <0.0001 | |
| miR-27b-3p | 6.74 | <0.0001 | |
| miR-21-5p | 4.89 | <0.0001 | |
| miR-451-5p | 1.54 | 0.0404 | |
| miR-192-5p | 3.78 | <0.0001 | |
| miR-16-5p | 1.56 | 0.0064 | |
| miR-29a-3p | 2.81 | <0.0001 | |
| miR-150-5p | 1.79 | 0.0006 | |
| miR-214-5p | 2.73 | 0.0213 | |
| miR-155-5p | 2.63 | 0.0023 | |
| miR-103a-5p | 3.38 | <0.0001 |
| Differences In miRNA Expression (fast). | |||
|---|---|---|---|
| miRNAs | Author | Effect Size: Fold Change (Cases vs. Controls) | p |
| miR-122-5p | Masotti et al. 2016 | 2.82 ± 0.49 | 0.037 |
| miR-34a-5p | 2.41 ± 0.39 | 0.032 | |
| miR-320a | 1.55 ± 0.11 | 0.014 | |
| miR-505-3p | 3.11 ± 0,65 | 0.03 | |
| miR-26b-5b | 1.63 ± 0.17 | 0.02 | |
| miR-146a-5p | 1.48 ± 0.09 | 0.014 | |
| miR-148b-3p | 1.47 ± 0.18 | 0.032 | |
| miR-342-3p | 1.46 ± 0.25 | 0.05 | |
| miR-190a | −3.04 ± 0.39 | 0.032 | |
| miR-200c-3p | −2.78 ± 0.46 | 0.032 | |
| miR-205-5p | −2.60 ± 0.44 | 0.032 | |
| miR-95 | −1.72 ± 0.26 | 0.032 | |
| miR-19a-3p | −1.55 ± 0.21 | 0.032 | |
| miR-660-5p | −1.50 ± 0.19 | 0.032 | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oses, M.; Margareto Sanchez, J.; Portillo, M.P.; Aguilera, C.M.; Labayen, I. Circulating miRNAs as Biomarkers of Obesity and Obesity-Associated Comorbidities in Children and Adolescents: A Systematic Review. Nutrients 2019, 11, 2890. https://doi.org/10.3390/nu11122890
Oses M, Margareto Sanchez J, Portillo MP, Aguilera CM, Labayen I. Circulating miRNAs as Biomarkers of Obesity and Obesity-Associated Comorbidities in Children and Adolescents: A Systematic Review. Nutrients. 2019; 11(12):2890. https://doi.org/10.3390/nu11122890
Chicago/Turabian StyleOses, Maddi, Javier Margareto Sanchez, Maria P. Portillo, Concepción María Aguilera, and Idoia Labayen. 2019. "Circulating miRNAs as Biomarkers of Obesity and Obesity-Associated Comorbidities in Children and Adolescents: A Systematic Review" Nutrients 11, no. 12: 2890. https://doi.org/10.3390/nu11122890
APA StyleOses, M., Margareto Sanchez, J., Portillo, M. P., Aguilera, C. M., & Labayen, I. (2019). Circulating miRNAs as Biomarkers of Obesity and Obesity-Associated Comorbidities in Children and Adolescents: A Systematic Review. Nutrients, 11(12), 2890. https://doi.org/10.3390/nu11122890







